INTRODUCTION
Prostate cancer is the most common non-skin cancer among US men with 220,800 new cases estimated in 2015 and more than 27,500 projected deaths [1]. Recently it has been shown that addition of docetaxel chemotherapy prolongs survival in this stage [2]. Despite this advance, emergence of castration-resistant prostate cancer (CRPC) is inevitable and is treated with systemic chemotherapy with docetaxel [3-7] and several novel systemic anti-cancer therapies [8-17]. Unfortunately, predictive biomarkers for response, efficacy or toxicity to traditional or novel treatments in advanced prostate cancer therapeutics are lacking and the practice continues to be based on best clinical estimates. Molecular classifiers of disease outcome or therapeutic benefit and toxicity are needed for individualizing therapeutic choices.
Body fluid-based biomarkers are appealing in advanced prostate cancer because they are less invasive and easily accessible. Cell free DNA (cfDNA)-based somatic aberrations in plasma of cancer patients have been extensively reported [15, 18, 19]. In advanced prostate cancer patients tumor-derived plasma cfDNA is detected in hormone sensitive and castrate resistant stages [20]. It remains unclear if genomic profiling to detect tumor cfDNA aberrations in urine is feasible and clinically relevant for developing as predictive biomarkers. To determine somatic genomic changes we performed whole-genome sequencing and analyzed copy number variations in matched urine specimens of advanced prostate cancer patients previously sequenced for plasma cfDNA [20]. We first evaluated the urine genome abnormality (UGA) algorithm based on genome-wide copy number variation (CNVs) to determine association with treatment response and clinical outcomes in patients receiving standard advanced prostate cancer treatments. We then compared urine cfDNA-based CNVs with previously reported plasma cfDNA CNVs [20]. Our data show that urine cfDNAs may generate comparable results to plasma cfDNA in CNVs analysis and may have clinical application in predicting treatment response and clinical outcomes.
RESULTS
Patients’ clinical characteristics
Matched urine specimens for patients with previous cfDNA sequencing of plasma specimens were available for 9 of 10 hormone sensitive prostate cancer (HSPC) patients and all ten patients with castrate resistant prostate cancer (CRPC) disease. For this study we selected these samples for urine cfDNA purification. Patient characteristics for these two advanced cancer cohorts are presented in Table 1. Each subject had two serial urine specimens collected before and after initiating stage-specific treatments. All patients in the HSPC sub-cohort underwent continuous ADT and patients with CRPC received docetaxel chemotherapy in addition to ADT as standard stage-specific treatments. The mean time between two sample collections in the HSPC was 128 days; the mean time between two sample collections in the CRPC was 112.4 days. The median follow-up was 64.00 months (40.93-69.13months) and 20.97 months (range 6.77-72.83 months) for HSPC and CRPC cohorts, respectively.
Table 1: Clinical characteristics of 19 advanced prostate cancer patients
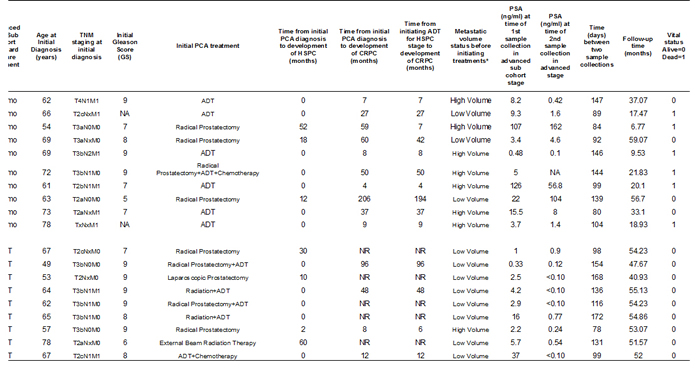
*High volume metastatic disease is defined as the presence of non-nodal visceral metastasis or 4 or more skeletal lesions with at least one outside the axial skeleton
Abbreviations: NR: Not Reached
ADT: Androgen Deprivation Therapy
PCA: Prostate Cancer
NA: Not available
Urine cfDNA yield and quality
To assess cfDNA yield, we tested three different kits using one single urine sample. We found that average cfDNA yields from 15 ml urine were 5.63ng, 6.46ng and 13.27ng for Zymo, Norgen and Analytik, respectively (Supplementary Figure 1). The Analytik kit generated approximately two fold more cfDNA than the two other kits. Due to relatively high yield, 2ng cfDNAs extracted using the Analytik kit was directly used for sequencing library construction. However, qualities of the sequencing libraries made from Analytik-derived cfDNA were extremely poor in three separate evaluation tests as determined by lack of featured library fragment band at ~300-310bp. Meanwhile, cfDNAs derived from Zymo kit generated consistent high quality sequencing library in three separate evaluation tests (Supplementary Figure 2).
Urine cfDNA and sequencing library quality
The final cfDNA yield from 15ml urine samples ranged from undetectable ( < 0.02ng/ul) to 1.6 ng/ul in 10ul elution buffer. Among 19 patients with both pre and post-treatment urine specimen, cfDNAs were detectable in 33 of the 38 samples. cfDNA yields from the remaining five samples were too low for measurement. For the 33 samples with total cfDNA > 0.25ng, sequencing libraries were prepared with final concentration of library DNAs between 0.878 and 3.490ng/ul. High sensitivity DNA chip showed multiple library fragments with peak size at ~300bp (Supplementary Figure 2). Whole genome sequencing generated approximately 7.6 million raw reads (ranging from 4.3 to 15.2) and 6.9 million mappable reads (ranging from 3.7 to 14.0). Corresponding mappable reads ranged from 77 to 93 percent of raw reads. The mean read count was ~134 per 60kb genomic window (Table 2).
Table 2: Statistics of whole genome sequencing
Sample ID |
Raw Reads |
Mappable Reads |
Percent Mapped |
Reads/60kb |
1001U2 |
8,559,863 |
7,780,937 |
90.90 |
151 |
1002U1 |
5,834,360 |
5,359,747 |
91.87 |
104 |
1002U2 |
6,905,307 |
6,218,374 |
90.05 |
120 |
1003U1 |
8,466,006 |
6,717,216 |
79.34 |
130 |
1003U2 |
8,249,703 |
6,668,418 |
80.83 |
129 |
1004U1 |
5,695,192 |
5,239,912 |
92.01 |
101 |
1004U2 |
6,523,691 |
6,056,921 |
92.85 |
117 |
1005U1 |
8,028,779 |
7,146,997 |
89.02 |
138 |
1010U1 |
6,941,555 |
6,376,244 |
91.86 |
123 |
1010U2 |
8,066,311 |
7,392,163 |
91.64 |
143 |
1014U1 |
5,925,269 |
5,434,275 |
91.71 |
105 |
1014U2 |
5,709,509 |
5,264,081 |
92.20 |
102 |
1015U1 |
6,757,096 |
6,173,896 |
91.37 |
119 |
1015U2 |
5,082,468 |
4,680,443 |
92.09 |
91 |
1017U1 |
7,268,083 |
6,383,512 |
87.83 |
124 |
1017U2 |
8,751,507 |
7,962,944 |
90.99 |
154 |
1028U1 |
4,366,638 |
3,758,949 |
86.08 |
73 |
1028U2 |
6,505,105 |
5,927,164 |
91.12 |
115 |
1040U1 |
7,715,099 |
6,887,111 |
89.27 |
133 |
1040U2 |
8,481,528 |
7,888,688 |
93.01 |
153 |
1050U1 |
6,635,918 |
6,006,158 |
90.51 |
116 |
1043U1 |
7,398,308 |
6,791,284 |
91.80 |
131 |
1050U2 |
10,320,989 |
9,167,473 |
88.82 |
177 |
1059U1 |
15,250,498 |
14,061,397 |
92.20 |
272 |
1059U2 |
6,303,913 |
5,865,752 |
93.05 |
114 |
1060U1 |
10,515,071 |
9,691,459 |
92.17 |
188 |
1060U2 |
7,294,862 |
6,713,772 |
92.03 |
130 |
1080U1 |
9,525,919 |
8,793,277 |
92.31 |
170 |
1080U2 |
8,771,271 |
7,816,538 |
89.12 |
151 |
1084U1 |
10,930,339 |
10,145,924 |
92.82 |
196 |
1098U2 |
7,507,997 |
5,835,747 |
77.73 |
113 |
1104U1 |
5,670,174 |
5,231,642 |
92.27 |
101 |
1104U2 |
7,542,611 |
6,868,682 |
91.07 |
133 |
Mean |
7,681,847 |
6,918,397 |
90.06 |
134 |
Overall urine cfDNA genomic abnormalities
To evaluate genomic abnormality, for each genomic bin, log2 ratio between read counts from urine cfDNA and lymphocyte-derived genomic DNA (gDNA) in the same patient was calculated. Fragmentation-based CNV analysis showed that cfDNA genomic abnormalities were detectable in all 19 patients tested. A greater number of genomic abnormalities were observed in the CRPC sub-cohort undergoing chemotherapy than in the HSPC cohort receiving ADT alone. Four of 10 CRPC patients (1003, 1004, 1014, 1017) and 2 of 9 HSPC patients (1050 and 1059) were observed to have specific genomic abnormalities. Of the 33 sequenced cfDNA specimens 14 patients had paired pre- and post-treatment cfDNA detectable abnormalities while five patients had either a pre- or post-treatment analysis. Among the 14 patients with paired samples, seven belonged to the HSPC sub-cohort and other seven to the CRPC sub-cohort. For these 14 patients, we performed unsupervised clustering analysis using log2 ratios in each genomic window and found that 11 pairs were clustered together (Supplementary Figure 3). Among these, some samples (such as patients 1050 and 1104) demonstrated significant CNV intensity differences between pre- and post-treatments. By comparing cfDNA-based CNVs from urine and plasma in matched patient samples, we observed consistent tumor-associated CNVs, although some differences of the log2 ratios in the two specimen types was observed (Figure 1).
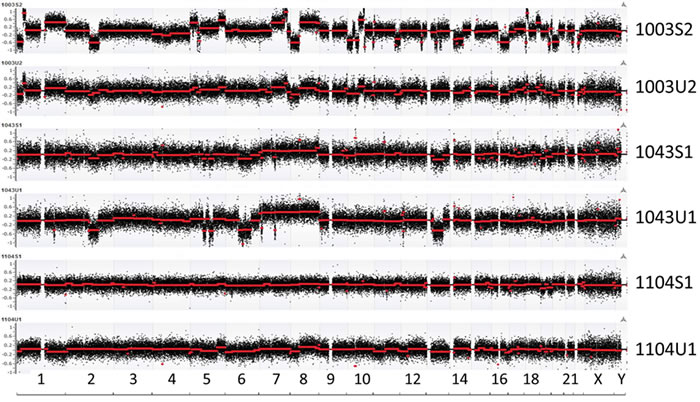
Figure 1: Overall view of genomic abnormalities in plasma (S) and urine (U). Log2 ratio-based segmentations across the human genome are shown with red line indicating averaged segments. Similar patterns of genomic abnormalities are seen in matched plasma and urine samples. “U1” represents the first urine sample findings for a particular patient identifier which is denoted by a four digit number. Similarly, “U2” represents the second serial urine specimen for the same patient; “S1” represents the first plasma specimen, while “S2” represents the second serial plasma sample.
Genomic abnormalities at specific loci
To further define genomic abnormalities in urine, we performed detailed analysis at chromosomal regions with putative and frequent aberrations in prostate cancer. Among these, the androgen receptor (AR) genomic region is most frequently amplified in CRPC patients. To examine the amplification status, we zoomed to the genomic region containing the AR and observed AR locus amplification in five of ten CRPC cases (#1003, #1005, #1010, #1017, and #1043) but none in nine HSPC cases. Although the amplicon boundaries varied they all contained whole AR gene. Another common genomic aberration in prostate cancer is at TMPRSS2 locus where frequent rearrangements create various fusion genes. We observed urine TMPRSS2 genomic variations in four cases with CRPC (#1003, #1005, #1014 and #1017) and two cases with HSPC (#1040, and #1098). The breakpoints for two genomic losses occurred at the two gene (ERG and TMPRSS2) regions forming the TMPRSS2-ERG fusion gene. The third most common genomic abnormality observed in prostate cancer is PTEN deletion. We found the PTEN loss in four cases of our CRPC sub-cohort (#1002, #1005, #1043 and #1060) and one case of HSPC (#1080) in the urine cfDNAs. Additionally, we found NOTCH1 locus amplification in one CRPC patients (#1014) and four HSPC patients (#1050, #1059, #1084 and #1098). Most of these abnormalities in urine cfDNAs were also observed and previously reported in the matched plasma cfDNAs [20] (Figure 2 and Table 3).
Table 3: Loss or gain of common prostate cancer-related genes
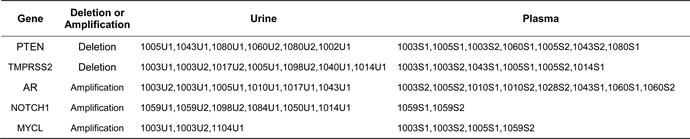
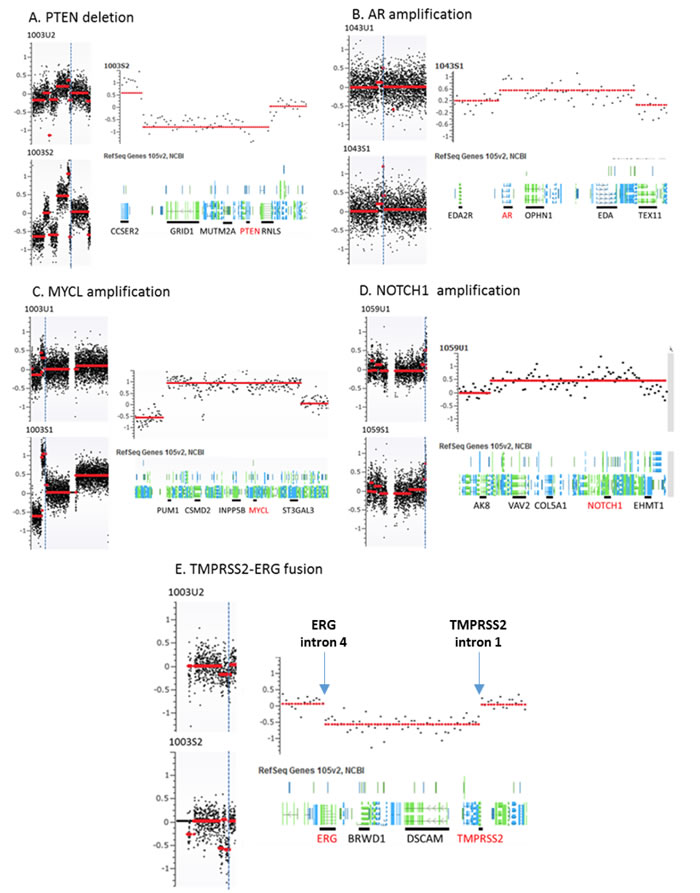
Figure 2: Representative genomic abnormalities detected at specific chromosomal loci in urine and matched plasma samples. Panel A.-E. represent PTEN deletion at chr 10, AR amplification at chr X, MYCL amplification at chr 1 NOTCH1 amplification at chr 9, and TMPRSS2-ERG fusion at chr 21. For each panel of A-E, left side shows chromosome level genomic changes and right side shows gene level genomic change at selected locus. 1003U2 and 1003S3 are urine and plasma cfDNAs from patient 1003, respectively. Vertical dot blue lines indicate the locations of these chromosomal aberrations.
Other chromosomal regions were also frequently altered in the tested samples with most having at least one common deletion or amplification per chromosome. From the common regions, we further defined the minimally overlapped regions which were involved in amplifications at 8q24.3, 9q34.3, 11p15.5 and 14q11.2 and deletions at 4q35.2, 5q31.3, 7q36.3, 12q24.33, and 16p11.2 (Table 4). Among those, seven regions including 5q31.3, 7q36.3, 8q24.3, 9q34.3, 11p15.54, 14q11.2 and 16p11.2 have been reported to be associated with prostate cancer [21-26]. Meanwhile, gene mutations at these loci have also been reported in prostate cancer tissues [27-29]. In addition, frequent “amplification” at TCRA locus was observed in most urine samples. Because of extensive rearrangements (deletions) at TCRA locus during T cell development, lymphocyte-derived gDNA may harbor partial deletions at this locus. Consequently, using such gDNAs as controls to normalize cfDNA may generate false positive amplification at this locus (Figure 3).
Table 4: Co-deletion or co-amplification segment of minimal overlap region
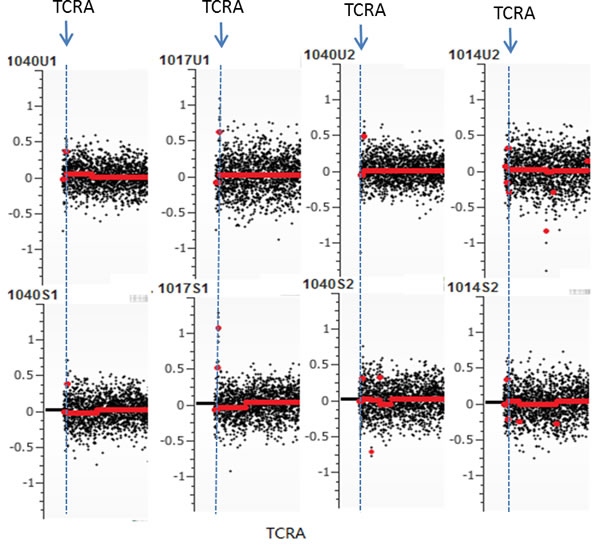
Figure 3: False positive (pseudo) amplification at TCRA locus. Due to frequent rearrangements (deletions) of T cell-derived DNA at TCRA locus, use of lymphocyte DNA as normalization control may create pseudo copy number gain.
Urine genomic abnormality (UGA) score and its clinical utility
Previously, we calculated a plasma genomic abnormality (PGA) score based on multiple genomic abnormalities in plasma as a potential biomarker for association with treatment response and survival [20]. A similar UGA-based classifier was developed for which we modified the calculation of the previously reported PGA score algorithm. UGA score based inter and intra patient variations (for the 14 paired specimens) were observed (Figure 4) and UGA scores in the pre-treatment group were higher in patients with high volume disease than low volume disease although it did not reach statistical significance (p = 0.16) (Figure 5). To evaluate if a genomic abnormality change occurred after initiating treatments and was associated with clinical outcomes, the previously reported TEff (treatment effect) index an algorithmic score which compares the percent differences between pre and post-treatment genome abnormality scores was used [20]. Kaplan-Meier survival analysis showed that a higher TEff index was significantly associated with poor survival (p < 0.04) in CRPC cohort (Figure 6A and 6B). In the HSPC cohort, the UGA and/or the PGA based TEff index did not show an association with change in TEff index and progression to castrate resistance, although a higher TEff index was detected to have a statistically non-significant trend with longer progression time to castration resistance (Figure 6C).
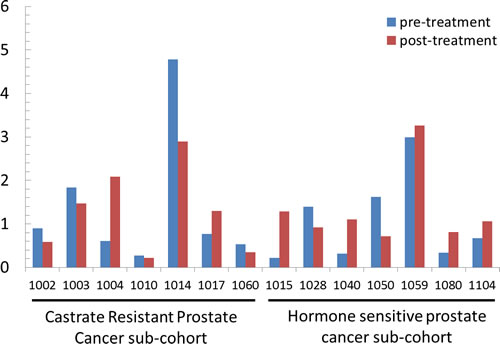
Figure 4: UGA score of 14 paired samples with pre- and post-stage specific therapies. The UGA scores demonstrate inter- and intra-patient variations. Y-axis: Urine Genome Abnormality score.
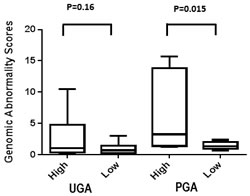
Figure 5: UGA and PGA score differences between high and low volume prostate cancer patients. Average UGA score before treatment is lower in low volume patients than in high volume patients. Average PGA score before treatment is significantly lower in low volume patients than in high volume patients.
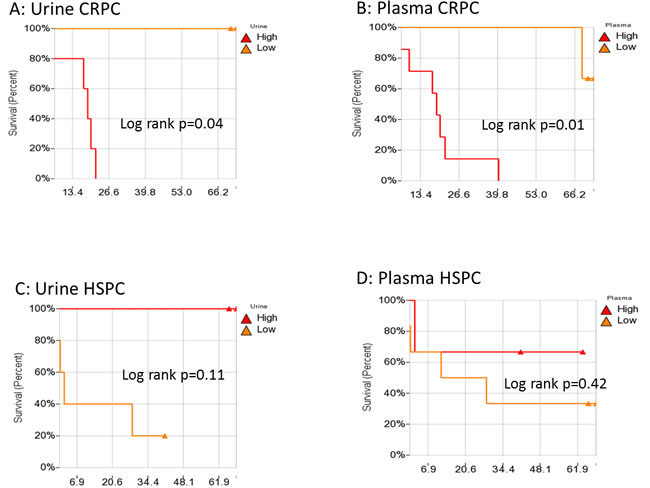
Figure 6: Kaplan-Meier analysis for the association of urine Teff A. and plasma Teff B. with overall survival in CRPC, and the association of urine Teff C. and plasma Teff D. with disease progression to CRPC. TEff: Treatment Efficacy index .
Treatment-associated genomic abnormalities
To examine treatment-associated genomic alterations, we generated log2 ratios between pre- and post-treatment specimens directly from scaled read counts at each genomic window and performed segmentation analysis for treatment-related genomic gain or loss. A total of 34 genomic loci with copy number changes were observed in the post treatment specimens. We then defined minimal overlap regions at each locus and identified commonly shared regions that covered nine genes (ZNRF3, RNF43, LGR4, NCOR1, ZBTB16, MYC, FGFR1, KRAS and STK11) (Supplementary Figure 4 and Table 5). For example, after treatment, the genomic region covering LGR4 was amplified in two cases of HSPC (#1080 and #1104), and the genomic region covering ZBTB16 was deleted in two cases of CRPC (#1014 and #1060). Copy number changes in the remaining seven gene regions were found in both advanced HSPC and CRPC urine specimens.
Table 5: Treatment-related genomic regions and genes
Chromosome |
Location |
Gene |
Sample ID |
Chr22 |
29,427,573- 29,453,476 |
ZNRF3 |
1003S2/S1(C*), 1050U2/U1(H*) |
Chr17 |
56,431,037- 56,494,931 |
RNF43 |
1050U2/U1(H), 1014U2/U1(C) |
Chr11 |
27,387,508- 27,494,338 |
LGR4 |
1104U2/U1(H), 1080U2/U1(H) |
Chr17 |
15,933,864- 16,101,195 |
NCOR1 |
1060S2/S1(C), 1050U2/U1(H) |
Chr11 |
113,933,133-114,126,702 |
ZBTB16 |
1060S2/S1(C), 1014U2/U1(C) |
Chr8 |
128,748,449-128,753,674 |
MYC |
1104U2/U1(H), 1060S2/S1(C) |
Chr7 |
55,177,416- 55,279,262 |
FGFR1 |
1104U2/U1(H), 1060S2/S1(C), 1080S2/S1(H) |
Chr12 |
25,358,180- 25,403,854 |
KRAS |
1104U2/U1(H), 1060S2/S1(C), 1050U2/U1(H) |
Chr19 |
1,205,798- 1,228,434 |
STK11 |
1104S2/S1(H), 1004U2/U1(C),1015U2/U1(H) |
*C represents CRPC patients and H represents HSPC patients.
DISCUSSION
cfDNA in blood has been extensively reported and proposed as biomarkers for cancer diagnosis, prognosis and treatment efficacy estimation. It is known that a small amount of cfDNA in blood passes into urine after renal filtration and tumor specific sequences are detectable in cfDNA isolated from urine [30, 31]. However a systematic determination of somatic genomic abnormalities in urine cfDNAs evaluated by high throughput sequencing technology in prostate cancer has not been performed [15, 19, 32]. Several challenges have limited this determination including a lack of precise knowledge on factors that may impact levels of urine cfDNA as cfDNA in urine is not as stable as in blood. In previous reports urine cfDNA profiling using PCR-based detection of candidate tumor-associated genes indicates that, an optimized and uniform method for cfDNA detection in urine that prevents degradation during extraction and storage should also include adequate volumes of specimens [33, 34]. Factors that may influence urine cfDNA detection include processing time of the urine samples after patient donation, the use of preservatives while processing, and the time of the urine samples in room temperature before storage in -80oC, and urine volumes.
We used a rigorous and uniform sample processing protocol for collecting 15ml of urine specimens and were able to detect cfDNA by Qubit instrument in most samples. Since the type of kit used for cfDNA extraction may impact yield and quality in this study we evaluated three commercial kits to identify association of extraction kit with cfDNA quality and yield. Although cfDNA yield using the Analytik kit was the highest, we were not able to apply the resultant cfDNA to generate high quality sequencing libraries. The Zymo kit generated relatively low yield but high quality sequencing libraries were consistently observed even at extremely low input of 0.25ng suggesting that selection of cfDNA extraction kit and thorough examination of cfDNA quality are important variables to consider for ensuring the subsequent success of sequencing library preparation and subsequent data analysis.
We were able to detect urine CNVs in all samples with adequate cfDNA quality and quantity, although the extent of detectable CNV per sample was stage-dependent with higher CNVs observed for CRPC patients than in HSPC stage. This finding is not surprising considering that mutations increase with progression in advanced prostate stages. CNV levels mirrored disease volume regardless of stage. We also observed lower level of detectable CNVs in urine specimens than in plasma (Figure 5), indicating a possible effect of renal filtration on urine cfDNA contents in concordance with previous reports in other tumor types [30, 31]. However, it does not appear to decrease the ability to detect specific somatic genomic changes . For example, shared specific genomic aberrations were observed in both plasma and urine cfDNAs at loci of PTEN, TMPRSS2 and AR (Figure 2 and Supplementary Figure 5). These results suggest that both urine and plasma fractions can be used for developing liquid biopsy based biomarkers in advanced prostate cancer.
In this limited data set, urine cfDNA changes were also explored as predictive biomarkers by examining CNV changes after initiating treatment and were able to identify treatment-associated CNV changes at nine gene loci. Of interest, majority of gene loci identified have been reported to be aberrant in prostate cancer biology, such as copy number changes after treatment in RNF43 and ZNRF3 loci. These two closely related single membrane spanning molecules have revealed receptor-like functionalities of a ligand-binding ectodomain. Combined with the intracellular architecture and activity of an E3 ligase, the two genes may be implicated in the modulation of Wnt signalling [35]. Post treatment copy number changes were also detectable in LGR4 and MYC proto oncogene loci. LGR4 has been reported to function in mammary gland development and mammary stem cells by activating Sox2 via the Wnt/β-catenin/Lef1 signaling pathway [36] and MYC proto-oncogene is frequently deregulated in prostate cancers, activating genetic programs that orchestrate biological processes to promote growth and proliferation [37].
Detection of cfDNA and the tumor-specific genomic aberrations in urine appears feasible and enhances the choices for developing liquid biopsy programs in advanced stage prostate cancer as predictive and prognostic classifiers. The approach adopted in our study performed with a limited number of samples for developing such classifiers is agnostic of specific gene/region changes and uses an algorithmic summation of the most common genetic abnormalities in urine. Since the mutational landscape of advanced prostate cancer is heterogeneous [38] this approach is likely to account for multiple genomic changes in tumor biology as a result of treatment effect. Due to small sample size, however, our findings are preliminary and need to be confirmed in larger cohorts of clinically annotated specimens. In addition, current whole genome-based sequencing technology is not sensitive to detect low level genomic abnormality. Its low sequencing depth makes detection of genomic rearrangement difficult. Regardless, with rapid advances in high throughput sequencing technology, sensitive detection of low level tumor-associated cfDNAs in body fluids will become feasible [39]. Urine cfDNA-based genomic abnormality tests may have the potential to provide a measurable classifier that can be used to assess treatment response and clinical outcomes in advanced prostate cancer patients.
MATERIALS AND METHODS
Patient methods
Urine specimens were obtained from advanced prostate cancer patients in metastatic hormone sensitive and metastatic castrate resistant stages. Patients were enrolled in a prospectively collected, institutional review board (IRB) approved study at a tertiary hospital while undergoing stage-specific standard of care treatments. Informed consent was obtained from all patients enrolled in the registry. The primary purpose of the registry is for developing blood and urine-based classifiers of disease and treatment outcomes in this patient population while patients receive standard of care treatments. Twenty cases (ten hormone sensitive and ten castrate resistant stage patients) were selected for this study with each patient having two serial urine samples. Each patient provided the first of the two urine specimens before initiating stage specific treatment and a second specimen after starting treatments. All cases selected for this study had matched plasma cfDNA sequencing performed previously [20]. All urine specimens were collected at the same time as the plasma collections. Initial processing of all urine specimens was performed uniformly within 45 minutes of receiving the sample from the patient. An initial centrifugation at 600g for 10 minutes was followed by storage of the urine and pellet in -80oC. No urine specimen underwent any freeze-thaw cycles other than at the time of extraction of cfDNA. Peripheral blood mononuclear cell-derived germline DNA (gDNA) was collected at the same time as the plasma and urine specimens. Clinical outcomes of patients undergoing this prospective specimen banking was performed retrospectively as previously described [20].
Isolation of cell free DNA (cfDNA)
To determine the best urine cfDNA extraction kit, we tested three different commercial products using a single urine sample. The kits included Extract-all Urine DNA kit (Zymo research corp., CA, USA), Urine DNA isolation kit (Norgen Biotek Corp., Ontario, Canada), and PME free-circulating DNA Extraction kit (Analytik Jena Innuscreen GmbH, Berlin, Germany). After thawing the urine sample, it was placed on ice immediately and then centrifuging of 15 ml urine was performed at 3000rpm for 15 minutes. The supernatant was used for DNA extraction according to each manufacturers’ protocol. cfDNA was eluted in 30ul elution buffer and concentration was measured using Qubit Fluorometer (Life Technology, Carlsbad, CA).
DNA extraction and sequencing library preparation
After an initial evaluation of the yield and quality of cfDNA from the three commercial kits, the Zymo research urine DNA Kit (Zymo Research, Irvine, CA) was selected to extract cfDNAs from 15 ml according to the manufacturer’s instructions. The extracted DNA was eluted in 10ul water. 1ul DNA eluent was quantified using Qubit. The remaining was stored at -80°C until preparation of sequencing libraries. For each patient germline DNA (gDNA) was also extracted and quantified. Sequencing DNA libraries were prepared for the urine cfDNA using a ThruPLEX DNA-Seq Kit (Rubicon Genomics, Inc. Ann Arbor, MI). 24 indexed libraries were pooled for single-read sequencing on a HiSeq2000 Sequencing System (Illumina, San Diego, CA).
Copy number variation (CNV) calculation
Raw sequencing data (fastq files) were first mapped to the human genome (hg19) (DNASTAR, Madison, WI). Read counts from the mapped sequence files were then binned into 60kb windows (total 51672 genomic bins) and adjusted to the global mean count for each sample. The read count ratio in each genomic bin was calculated by dividing cfDNA with peripheral blood mononuclear cell germline DNA (gDNA) in the same patient [20]. The resulting ratios were further transformed with log2 and corrected for GC content [40]. The fully normalized log2 ratios in genomic bins were subjected to segmentation using the copy number analysis method (CNAM) algorithm (Golden Helix, Bozeman, MT).
Urine genome abnormality (UGA) score algorithm calculation and comparison with plasma genome abnormality score (PGA)
To quantify genomic abnormality we modified the previously reported methodology for calculating global genomic abnormalities in plasma (plasma genome abnormality, PGA) score [20]. This was performed by summing the most significant log2 ratios in top 95-99% genomic bins. For the current study, we modified the genome abnormality calculation by summing log2 ratios of ten most significant genomic segments. To generate the UGA score, we first mapped raw sequencing data (fastq files) to the human genome (hg19) (DNASTAR, Madison, WI). We then binned read counts from the mapped sequence files into 60kb windows (total 51607 genomic bins) and rescaled to the global mean count for each sample. To account for constitutional CNVs, we divided read count in each bin from urine cfDNA by the one in the same bin from gDNA to generate a normalized read count ratio. The resulting ratio was transformed by log2 transformation and GC content correction [40]. Finally, we performed segmentation analysis using the copy number analysis method (CNAM) algorithm (Golden Helix, Bozeman, MT). Due to highly repeated sequences in centromeres, we removed genomic regions containing centromeres and their surrounding +/-1Mb regions. We also excluded small genomic segments containing ≤ 4 bin windows (4x60kb). From the remaining segments, we summed the top ten most significant segment values (using absolute numbers) and defined the summarized number as Urine Genomic abnormality (UGA) score. For consistency and comparability the previously reported PGA score [20] was also re-analyzed in the same manner as the UGA. A higher score being indicative of a greater tumor DNA fraction in the cfDNA. To quantify a treatment response index in each patient, we defined the TEff (Treatment Efficacy) index as the log2 ratio of UGA (or PGA) scores between the pre- and post-treatments: TEff index = log2 (prePGA/postPGA) x100.
Statistical analysis
For defining hormone sensitive and castrate resistant stage in this hospital based registry a uniform definition was used as reported previously [20, 41]. Briefly, for the CRPC cohort, overall survival was recorded from the date of first plasma collection after ADT failure to death or last follow-up. For the HSPC cohort, disease progression was recorded from the date of first plasma collection at initiation of ADT to disease progression or last follow-up. To evaluate association of the UGA score with overall survival in the CRPC sub-cohort, time from developing castrate resistance to death was considered and Kaplan-Meier analysis was performed for the UGA score and TEff index associations with overall survival (prognostic classifier). For the HSPC cohort time from initiating androgen deprivation therapy (ADT) for hormone sensitive stage to development of castrate resistance was obtained (predictive classifier). We dichotomized each sub-cohort into two risk groups using median UGA score or TEff index as a cut-off. A P-value of ≤ 0.05 was considered statistically significant for all statistical analyses.
Acknowledgments and Funding
This study was supported by Joseph and Gail Gassner Development funds for prostate cancer research to MK, and Advancing a Healthier Wisconsin Fund (#5520227) and National Institute of Health (R01CA157881) to LW.
Conflicts of Interest
The authors declare that there is no conflict of interests.
References
1. Siegel RL, Miller KD and Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015; 65:5-29.
2. Sweeney CJ, Chen Y-H, Carducci M, Liu G, Jarrard DF, Eisenberger M, Wong Y-N, Hahn N, Kohli M and Cooney MM. Chemohormonal therapy in metastatic hormone-sensitive prostate cancer. New England Journal of Medicine. 2015; 373:737-746.
3. Petrylak DP, Tangen CM, Hussain MHA, Lara PN, . J., Jones JA, Taplin ME, Burch PA, Berry D, Moinpour C, Kohli M, Benson MC, Small EJ, Raghavan D and Crawford ED. Docetaxel and estramustine compared with mitoxantrone and prednisone for advanced refractory prostate cancer. New England Journal of Medicine. 2004; 351:1513-1520.
4. Tannock IF, de Wit R, Berry WR, Horti J, Pluzanska A, Chi KN, Oudard S, Theodore C, James ND, Turesson I, Rosenthal MA, Eisenberger MA and Investigators T. Docetaxel plus prednisone or mitoxantrone plus prednisone for advanced prostate cancer. New England Journal of Medicine. 2004; 351:1502-1512.
5. Denis LJ, Keuppens F, Smith PH, Whelan P, de Moura JL, Newling D, Bono A and Sylvester R. Maximal androgen blockade: final analysis of EORTC phase III trial 30853. EORTC Genito-Urinary Tract Cancer Cooperative Group and the EORTC Data Center. Eur Urol. 1998; 33:144-151.
6. Eisenberger MA, Blumenstein BA, Crawford ED, Miller G, McLeod DG, Loehrer PJ, Wilding G, Sears K, Culkin DJ and Thompson Jr IM. Bilateral orchiectomy with or without flutamide for metastatic prostate cancer. New England Journal of Medicine. 1998; 339:1036-1042.
7. Maximum androgen blockade in advanced prostate cancer: an overview of the randomised trials. Prostate Cancer Trialists’ Collaborative Group. Lancet. 2000; 355:1491-1498.
8. De Bono JS, Logothetis CJ, Molina A, Fizazi K, North S, Chu L, Chi KN, Jones RJ, Goodman OB, Saad F, Staffurth JN, Mainwaring P, Harland S, Flaig TW, Hutson TE, Cheng T, et al. Abiraterone and Increased Survival in Metastatic Prostate Cancer. New England Journal of Medicine. 2011; 364:1995-2005.
9. Kantoff PW, Higano CS, Shore ND, Berger ER, Small EJ, Penson DF, Redfern CH, Ferrari AC, Dreicer R and Sims RB. Sipuleucel-T immunotherapy for castration-resistant prostate cancer. New England Journal of Medicine. 2010; 363:411-422.
10. de Bono JS, Oudard S, Ozguroglu M, Hansen S, Machiels J-P, Kocak I, Gravis G, Bodrogi I, Mackenzie MJ and Shen L. Prednisone plus cabazitaxel or mitoxantrone for metastatic castration-resistant prostate cancer progressing after docetaxel treatment: a randomised open-label trial. The Lancet. 2010; 376:1147-1154.
11. Ryan CJ, Smith MR, de Bono JS, Molina A, Logothetis CJ, de Souza P, Fizazi K, Mainwaring P, Piulats JM and Ng S. Abiraterone in metastatic prostate cancer without previous chemotherapy. New England Journal of Medicine. 2013; 368:138-148.
12. Cabot RC, Harris NL, Rosenberg ES, Shepard J-AO, Cort AM, Ebeling SH, McDonald EK, Scher HI, Fizazi K and Saad F. Increased survival with enzalutamide in prostate cancer after chemotherapy. New England Journal of Medicine. 2012; 367:1187-1197.
13. Snedecor SJ, Carter JA, Kaura S and Botteman MF. Denosumab versus zoledronic acid for treatment of bone metastases in men with castration-resistant prostate cancer: a cost-effectiveness analysis. Journal of medical economics. 2012; 16:19-29.
14. Fizazi K, Carducci M, Smith M, Damiao R, Brown J, Karsh L, Milecki P, Shore N, Rader M, Wang H, Jiang Q, Tadros S, Dansey R and Goessl C. Denosumab versus zoledronic acid for treatment of bone metastases in men with castration-resistant prostate cancer: a randomised, double-blind study. Lancet. 2011; 377:813-822.
15. Crowley E, Di Nicolantonio F, Loupakis F and Bardelli A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol. 2013; 10:472-484.
16. Kohli M and Tindall DJ. New Developments in the Medical Management of Prostate Cancer. Mayo Clin Proc. 2010; 85:77-86.
17. Attard G, Reid AH, A’Hern R, Parker C, Oommen NB, Folkerd E, Messiou C, Molife LR, Maier G, Thompson E, Olmos D, Sinha R, Lee G, Dowsett M, Kaye SB, Dearnaley D, et al. Selective inhibition of CYP17 with abiraterone acetate is highly active in the treatment of castration-resistant prostate cancer. Journal of clinical oncology. 2009; 27:3742-3748.
18. Diaz LA, Jr. and Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014; 32:579-586.
19. Heitzer E, Ulz P and Geigl JB. Circulating tumor DNA as a liquid biopsy for cancer. Clin Chem. 2015; 61:112-123.
20. Xia S, Kohli M, Du M, Dittmar RL, Lee A, Nandy D, Yuan T, Guo Y, Wang Y, Tschannen MR, Worthey E, Jacob H, See W, Kilari D, Wang X, Hovey RL, et al. Plasma genetic and genomic abnormalities predict treatment response and clinical outcome in advanced prostate cancer. Oncotarget. 2015; 6:16411-21. doi: 10.18632/oncotarget.3845.
21. Witte JS, Goddard KA, Conti DV, Elston RC, Lin J, Suarez BK, Broman KW, Burmester JK, Weber JL and Catalona WJ. Genomewide scan for prostate cancer-aggressiveness loci. American journal of human genetics. 2000; 67:92-99.
22. Kim JH, Dhanasekaran SM, Mehra R, Tomlins SA, Gu W, Yu J, Kumar-Sinha C, Cao X, Dash A, Wang L, Ghosh D, Shedden K, Montie JE, Rubin MA, Pienta KJ, Shah RB, et al. Integrative analysis of genomic aberrations associated with prostate cancer progression. Cancer research. 2007; 67:8229-8239.
23. Tindall EA, Severi G, Hoang HN, Ma CS, Fernandez P, Southey MC, English DR, Hopper JL, Heyns CF, Tangye SG, Giles GG, Hayes VM and Australian Prostate Cancer B. Comprehensive analysis of the cytokine-rich chromosome 5q31.1 region suggests a role for IL-4 gene variants in prostate cancer risk. Carcinogenesis. 2010; 31:1748-1754.
24. Porkka KP, Tammela TLJ, Vessella RL and Visakorpi T. RAD21 and KIAA0196 at 8q24 are amplified and overexpressed in prostate cancer. Genes, Chromosomes and Cancer. 2004; 39:1-10.
25. Tuupanen S, Turunen M, Lehtonen R, Hallikas O, Vanharanta S, Kivioja T, Björklund M, Wei G, Yan J and Niittymäki I. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nature genetics. 2009; 41:885-890.
26. Saramäki OR, Porkka KP, Vessella RL and Visakorpi T. Genetic aberrations in prostate cancer by microarray analysis. International journal of cancer. 2006; 119:1322-1329.
27. Ellen TP, Ke Q, Zhang P and Costa M. NDRG1, a growth and cancer related gene: regulation of gene expression and function in normal and disease states. Carcinogenesis. 2008; 29:2-8.
28. Wang Z, Li Y, Banerjee S, Kong D, Ahmad A, Nogueira V, Hay N and Sarkar FH. Down-regulation of Notch -1 and Jagged-1 inhibits prostate cancer cell growth, migration and invasion, and induces apoptosis via inactivation of Akt, mTOR, and NF-B signaling pathways. J Cell Biochem. 2010; 109:726-736.
29. Bin Hafeez B, Adhami VM, Asim M, Siddiqui IA, Bhat KM, Zhong W, Saleem M, Din M, Setaluri V and Mukhtar H. Targeted knockdown of Notch1 inhibits invasion of human prostate cancer cells concomitant with inhibition of matrix metalloproteinase-9 and urokinase plasminogen activator. Clinical cancer research. 2009; 15:452-459.
30. Botezatu I, Serdyuk O, Potapova G, Shelepov V, Alechina R, Molyaka Y, Ananev V, Bazin I, Garin A, Narimanov M, Knysh V, Melkonyan H, Umansky S and Lichtenstein A. Genetic analysis of DNA excreted in urine: a new approach for detecting specific genomic DNA sequences from cells dying in an organism. Clin Chem. 2000; 46:1078-1084.
31. Lichtenstein AV, Melkonyan HS, Tomei LD and Umansky SR. Circulating nucleic acids and apoptosis. Annals of the New York Academy of Sciences. 2001; 945:239-249.
32. De Mattos-Arruda L, Cortes J, Santarpia L, Vivancos A, Tabernero J, Reis-Filho JS and Seoane J. Circulating tumour cells and cell-free DNA as tools for managing breast cancer. Nature reviews Clinical oncology. 2013; 10:377-389.
33. Emile JF, Diamond EL, Helias-Rodzewicz Z, Cohen-Aubart F, Charlotte F, Hyman DM, Kim E, Rampal R, Patel M, Ganzel C, Aumann S, Faucher G, Le Gall C, Leroy K, Colombat M, Kahn JE, et al. Recurrent RAS and PIK3CA mutations in Erdheim-Chester disease. Blood. 2014; 124:3016-3019.
34. Diamond EL, Abdel-Wahab O, Pentsova E, Borsu L, Chiu A, Teruya-Feldstein J, Hyman DM and Rosenblum M. Detection of an NRAS mutation in Erdheim-Chester disease. Blood. 2013; 122:1089-1091.
35. Zebisch M and Jones EY. ZNRF3/RNF43-A direct linkage of extracellular recognition and E3 ligase activity to modulate cell surface signalling. Progress in biophysics and molecular biology. 2015; 118:112-8.
36. Nakata S, Phillips E and Goidts V. Emerging role for leucine-rich repeat-containing G-protein-coupled receptors LGR5 and LGR4 in cancer stem cells. Cancer management and research. 2014; 6:171.
37. Hsieh AL, Walton ZE, Altman BJ, Stine ZE and Dang CV. MYC and metabolism on the path to cancer. Semin Cell Dev Biol. 2015.
38. Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM, Montgomery B, Taplin ME, Pritchard CC, Attard G, Beltran H, Abida W, Bradley RK, Vinson J, Cao X, Vats P, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015; 161:1215-1228.
39. Kato S and Janku F. Cell-free DNA as a novel marker in cancer therapy. Biomarkers in medicine. 2015; 9:703-712.
40. Diskin SJ, Li M, Hou C, Yang S, Glessner J, Hakonarson H, Bucan M, Maris JM and Wang K. Adjustment of genomic waves in signal intensities from whole-genome SNP genotyping platforms. Nucleic Acids Res. 2008; 36:e126.
41. Huang X, Yuan T, Liang M, Du M, Xia S, Dittmar R, Wang D, See W, Costello BA, Quevedo F, Tan W, Nandy D, Bevan GH, Longenbach S, Sun Z, Lu Y, et al. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant prostate cancer. Eur Urol. 2015; 67:33-41.

