INTRODUCTION
Multiple myeloma (MM) is a common and life-threatening hematological malignancy and is characterized by uncontrolled growth and accumulation of monoclonal plasma cells. These cells can infiltrate the bone marrow (BM) and typically secrete monoclonal (M) proteins into the peripheral circulation, resulting in related organ and tissue injury (ROTI). The pathogenesis of MM is largely attributed to the interplay between myeloma cells and the in vivo environment, mostly the bone marrow microenvironment (BMME). This interaction occurs throughout the entire disease process, namely, from monoclonal gammopathy of undetermined significance (MGUS), smoldering MM, symptomatic MM, and finally to plasma cell leukemia (PCL) [1]. Conventional therapies such as hematopoietic stem cell transplantation, which mainly focused on myeloma cells, result in a low complete remission (CR) rate and short survival time in MM [2]. The emerging novel therapies, including proteasome inhibitors and immunomodulatory drugs (IMiDs), can influence the BMME of MM and have strikingly improved the survival for MM patients [3–5]. However, resistance to novel drugs tends to be a clinical frustration because nearly all MM patients inevitably relapse or evolve to a refractory stage. In addition to the biobehavioral changes of myeloma cells of resisting drug challenges, BMME such as bone marrow stromal cells (BMSCs), have been found to play a vital role in drug resistance like bortezomib [6]. Thus, more attention needs focused on the crosstalk between myeloma cells and the in vivo environment, which may shed light on understanding drug resistance in MM. Accordingly, it is necessary to find some routine clinical markers that reflect the in vivo environment directly correlated to a high predictive value of DR.
We designed this study using the real world study (RWS) method of using data for decision making that was not collected in randomized clinical trials (RCTs), a concept introduced in 2007 by the International Society for Pharmacoeconomics and Outcomes Research (ISPOR) Task Force [7]. Additionally, due to focal distribution of the disease it is difficult to obtain samples containing myeloma cells and those that are obtained may lack biomarkers that reflect MM [8]. Exosomes, which are released into circulation from all cell types, are lipid bilayer cup-shaped nanovesicles with 30–100 nm in diameter and provide membrane protection for inclusive RNAs and proteins [9]. MicroRNAs (miRNAs), existing naturally as the most biologically stable nucleic acid molecule with only about 19–23 nucleotides, act as fine-tuning regulators of gene expression at post-transcriptional level through a complicated miRNA-mRNA interaction [10]. Until now, emerging studies have suggested that tumor-derived exosomes quantitatively predominate in peripheral blood and exosome-mediated miRNA transduction plays a pivotal role in the dialogue between human tumors and their microenvironment [11]. Thereby, we hypothesized that the profile of exosomal miRNA from peripheral blood, which can be easily available with a minimally invasive procedure, had a predictive value of primary or acquired drug resistance (DR) for MM patients.
RESULTS
Patients’ characteristics
A total of 300 MM patients with 1826 episodes of hospitalizations (with an average of 6.1 per patient per year) were analyzed in our center. The median age of this cohort was 61 years old, while the estimated median OS was 83.9 months. As shown in Table 1, 68.0% (n = 204) of patients with 1682 episodes of hospitalizations received a novel agent-based regimen and were enrolled in the study, among which, 56.4% (n = 115), 32.8% (n = 67) and 10.8% (n = 22) were treated with Bz, thalidomide and lenalidomide, respectively. Meanwhile, 32% (n = 96) of the remaining patients were admitted for clinical trials or supportive treatment. Bz and thalidomide were mainly used as front-line therapy, while lenalidomide as salvage therapy. In total, DR accounted for 53.4% (n = 109) of the entire cohort with the lowest frequency of 36.5% (n = 42) in the Bz group (p = 0.000).
Table 1: Baseline characteristics of the MM patient cohort receiving a novel agent-based regimen
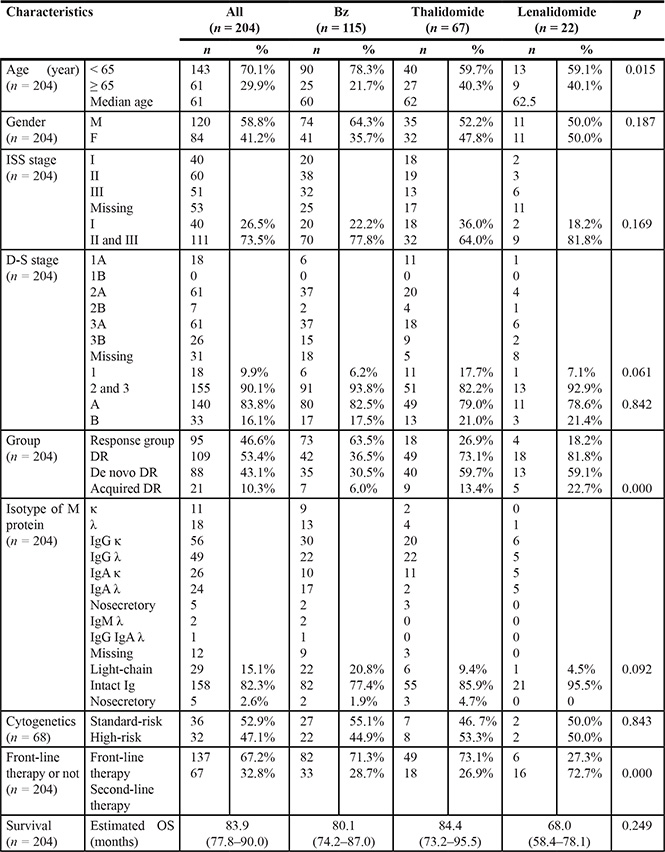
Baseline data predicting DR for MM treated with novel agents
As shown in Tables 2–3, the de novo DR group accounted for 83.38% (n = 35) of the Bz-resistant group, while 73.13% (n = 40) of the thalidomide-resistant group. For the drug resistant cases receiving bortezomib-containing regimens, compared with the response, also called the sensitive group, the serum total light chain ratio was higher than 100 or less than 0.01 RR = 34.286; 95% CI: 3.476–338.2 P = 0.001), CRP was greater than 20 mg/L (RR = 14; 95% CI: 1.23–158.84 P = 0.032), and the second-line usage of bortezomib (RR = 9.667; 95%CI: 1.705–54.87; P = 0.009) was associated with an increased risk for the occurrence of acquired resistance. However, no significant indicators from the routine workup were found between the response and primary resistant groups capable of predicting the efficacy for the usage of either Bz or thalidomide as well as between the response and acquired thalidomide-resistant group.
Table 2: The relationship between the in vivo environment and treatment efficacy of bortezomib for MM
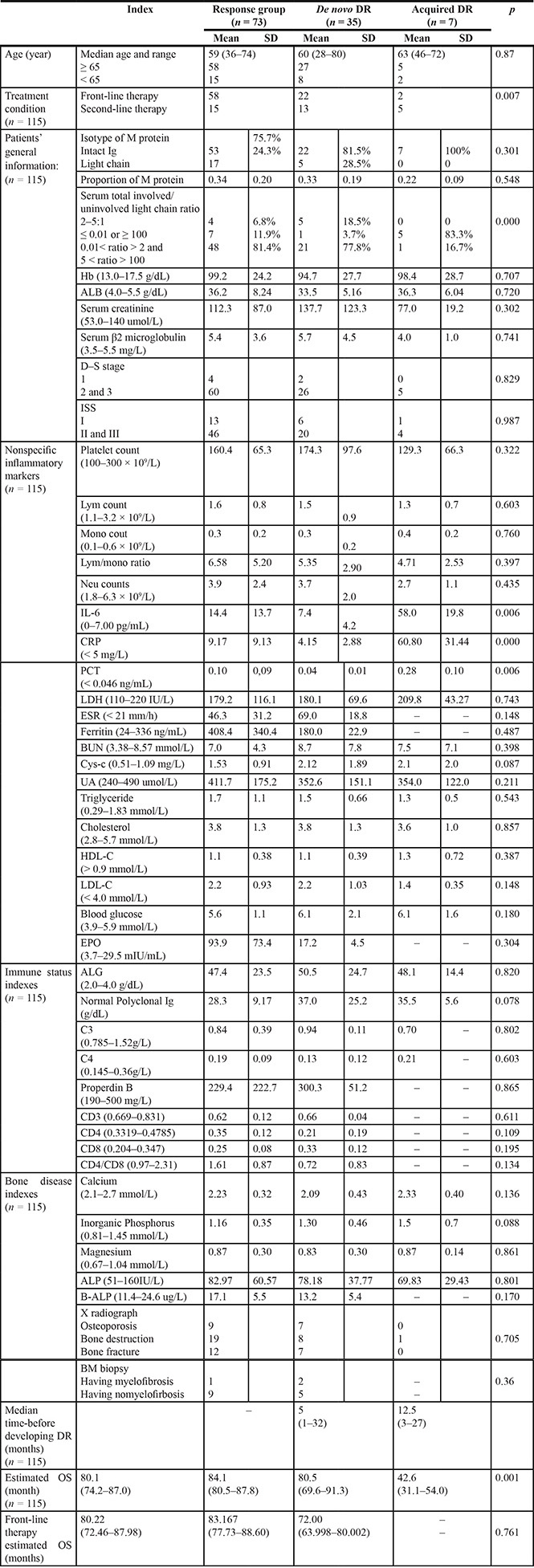
Table 3: The relationship between the internal environment and treatment efficacy of thalidomide for MM
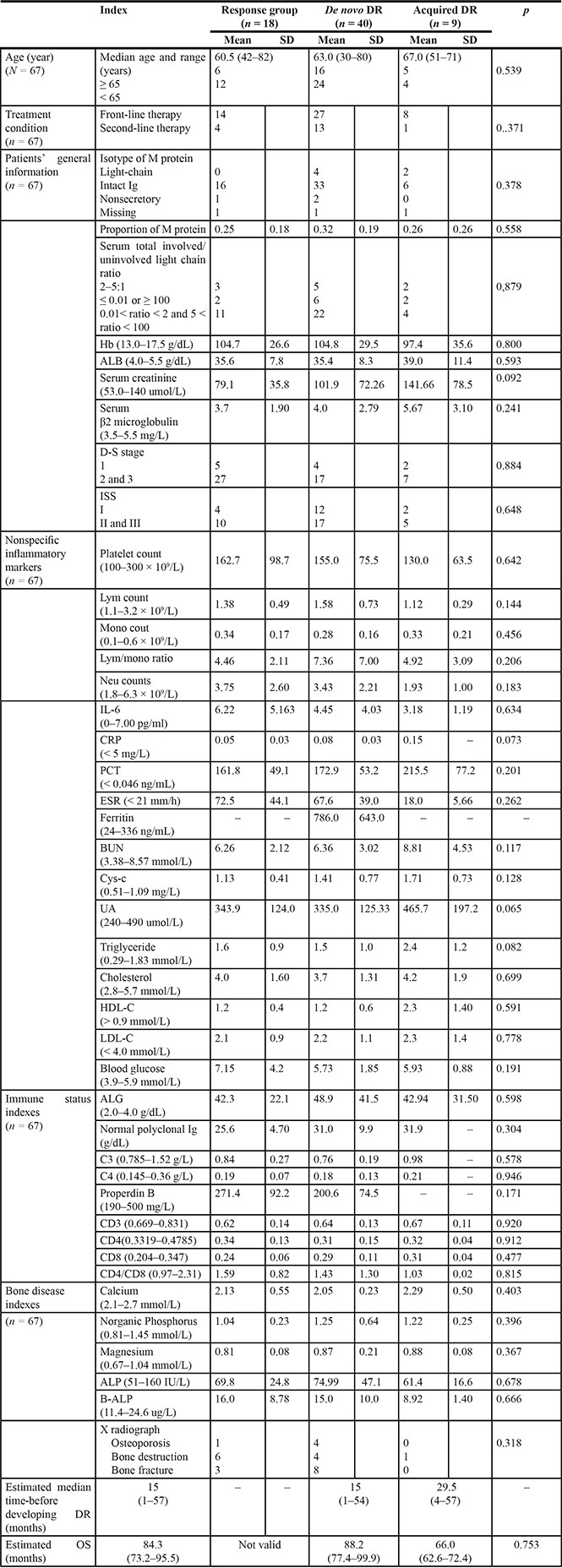
As shown in Table 4, among 68 cases with cytogenetic results in this cohort, a statistically significant difference of a 1q21 gain frequency was found between the response and de novo DR groups (RR:3.472, 95% CI:1.184–10.179, p = 0.034). Moreover, patients with a 1q21 gain in the de novo DR groups were significantly associated with lower levels of serum cholesterol (p = 0.029) and LDL-C (p = 0.024) (shown in Supplementary Table S1).
Table 4: The role of genetic abnormalities in predicting DR for MM in the new agent therapy era
|
|
high-risk cytogenetics |
1q gain |
1p deletion |
13p Deletion |
TP53 deletion |
t (4:14) |
t (11;14) |
t (14;16) |
|---|---|---|---|---|---|---|---|---|---|
All (n = 68) |
Response (n = 37) De novo(n = 24) Acquired (n = 7) p |
13 15 4 0.096 |
12 15 4 0.057 |
0 1 1 0.111 |
12 9 4 0.460 |
1 1 1 0.391 |
2 2 0 0.700 |
0 1 2 0.003 |
0 1 0 0.394 |
Exosomes and exosomal RNAs isolated from culture media and peripheral blood of MM
As shown in Figure 1B, TEM images and NanoSight NS300 revealed the presence of nano-sized, round EMVs, MVs and cup-shaped exosomes isolated from the U-266 cell culture media, serum of MM patients and healthy control. The purified exosomes of less than 100 nm in diameter, which are equipped with exosomal marker proteins CD63 and HSP70 (Figure 1C), were more abundant in the serum of MM patients than in healthy control. As shown in Figure 2A, significant differences of exosomal RNA content were observed between the Bz-resistant group and the Bz-response group (350.17 ± 37.55 ng/μl vs 235.43 ± 3.91 ng/μl, p = 0.033) by Nanodrop analysis and the exosomal RNA concentrations of both groups were obviously higher than 78.21 ± 45.51 ng/μl of the healthy control. Besides, we also isolated circulating RNAs in plasma of patients in the Bz-resistant group and the Bz-response group (Figure 2A–2D and Figure 2A–2F). It showed that the exosomal RNA concentrations of both groups were significantly higher than the circulating RNA concentrations, respectively (data not shown).
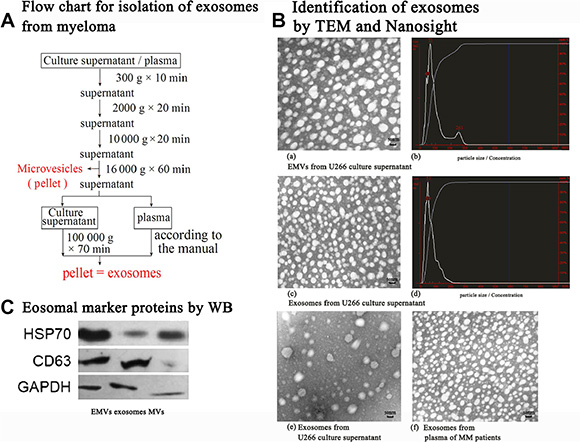
Figure 1: Isolation and validaton of exosomes from both U266 cell line and MM patients. MM-derived exosomes were isolated and purified as depicted in section A. In section B, presence of MVs and exosomes: Both MVs and exosomes were present by TEM (A) and Nanosight (B) following centrifugation of U266 culture supernatant without removal of MVs by 16 000 g × 60 min; Exosomes were present by TEM (C) and Nanosight (D) following centrifugation of U266 culture supernatant with removal of MVs by 16 000 g × 60 min. Processed plasma samples from MM patients based on the protocol in section A, exosomes derived from MM quantitatively predominated (F) than those from the healthy control (E). In section C, EMVs, exosomes and MVs were isolated from the culture supernatant of U266 cells, respectively. Exosomal marker proteins of HSP70 were detected for both exosomes and MVs, while CD63 was mainly observed in exosomes.
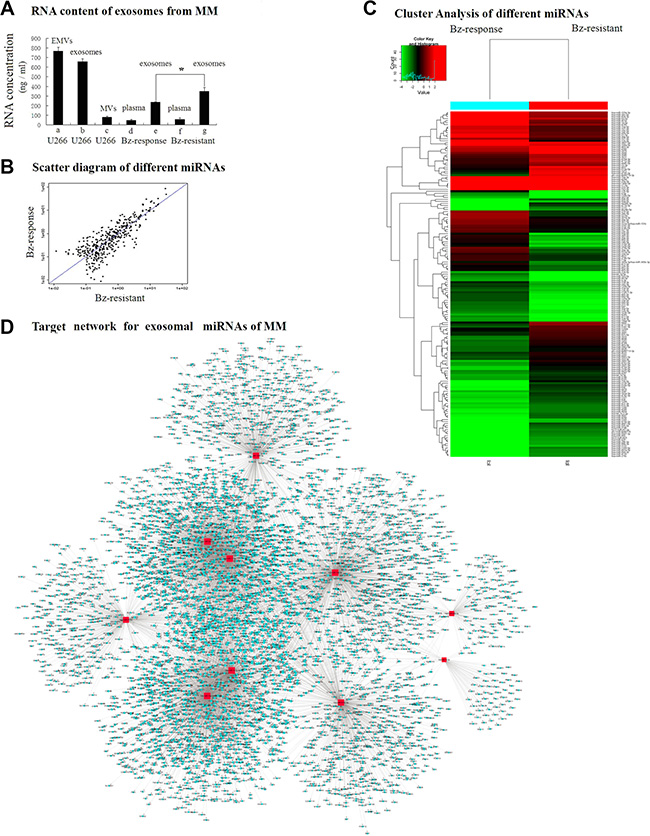
Figure 2: The differential expression of exosome-associated miRNAs in the subgroups of MM. Exosomal RNAs were detected from both the culture supernatant of U266 cells and the plasma samples from MM patients (A). Significant differences (* < 0.05) of exosomal RNA content were observed between the Bz-response group (E) and Bz-resistant group (G). Exosomal RNA content of both groups was much higher than the content of circulating RNAs (D and F) extracted using the miRNeasy Serum/Plasma Kit (Qiagen). In sections (B and C) compared to the Bz-response group, 83 miRNAs were expressed at higher levels and 88 miRNAs were expressed at lower levels in the Bz-resistant group among 3180 miRNAs on the microarray. In section D, a miRNA-RNA synergistic network was constructed with miR-17-5p, miR-20a-5p and miR-15a-5p, miR-16-5p exhibiting more synergism in the Bz-resistant mechanism of MM.
The exosome-associated miRNA expression patterns for predicting Bz-resistance in MM
Among 3180 miRNAs detected on a microarray, profiling data analysis predicted that 83 miRNAs were expressed at higher levels and 88 miRNAs were expressed at lower levels in Bz-resistant group (Figure 2B), and more than 90% of these miRNAs had novel functions. After result confirmation by RT-PCR to remove false positives, we listed the top 10 miRNAs exhibiting the largest changes, overlapped with the top miRNAs based on literature, namely, miR-513a-5p, miR-20b-3p, let-7d-3p were up-regulated and miR-125b-5p, miR-19a-3p, miR-21-5p, miR-20a-5p, miR-17-5p, miR-15a-5p, miR-16-5p were down-regulated. These differentially expressed miRNA families play key roles in post-transcriptional regulation by influencing transcription co-factors, the MAP kinase pathway and ubiquitin conjugating enzyme activity (Figure 2C). Furthermore, a miRNA-RNA synergistic network was constructed to describe the role of multiple exosomal miRNAs in the DR mechanisms of complex post-transcriptional regulations in human MM, which were by several miRNAs rather than a single miRNA. We found that miR-17-5p, miR-20a-5p, miR-15a-5p and miR-16-5p exhibited higher synergistic effects (Figure 2D), indicating a functional complexity where a global central core of the post-transcriptional regulatory network is involved as a Bz-resistant mechanism of MM. Levels of miR-16-5p, miR-15a-5p, miR-20a-5p and miR-17-5p in the Bz-resistant group were 3.91, 1.83 and 2.96, 1.97 folds lower, respectively, than those in the Bz-response group.
DISCUSSION
Early predication of DR has been playing an increasingly important role in holistic treatment for patients with MM, considering that nearly all patients eventually relapse and refractory MM (RRMM) with multidrug resistance is observed even with the use of novel treatment agents. Moreover, almost 20–30% of MM patients have innate DR to Bz [12, 13]. The present study showed that Bz and thalidomide were mainly used as front-line therapies (67.2%, 71.3%, respectively) and lenalidomide as a salvage therapy (72.7%), differing from the management guideline for multiple myeloma (Version4, 2015) published by the National Comprehensive Cancer Network (NCCN). To some extent, this may be associated with an overwhelming price advantage of thalidomide over lenalidomide in the Chinese market and this may generally represent the current MM treatment in China. In this study we explored the potential factors responsible for the occurrence of Bz and thalidomide DR.
Compared with the Bz-response groups, risk predictors for an increased likelihood of acquired resistance were in descending order, namely, abnormal total serum light chain ratio (≤ 0.01 or ≥ 100), higher CRP level (≥ 20 mg/L), and the second-line usage. First, although the free light chain (FLC) ratio has been increasingly used for monitoring MGUS evolving to MM and early evaluation of Bz response, the test of FLC ratio has not been widely available in the majority of Chinese myeloma centers [14, 15]. Under such circumstances, the clinical role of the total light chain ratio in Bz resistance cannot be ignored. The total serum light chain detection in this analysis was consistent with results from a published retrospective study that speculated that reaching a normal total light chain ratio may be sufficient to maintain a stable phase of the disease and confer prolonged event free survival (EFS) [16]. Second, a high level of CRP is known to contribute to a high tumor burden, extensive complications and poor prognosis in MM [17, 18]. In contrast, CRP may protect myeloma cells from dexamethasone and melphalan-induced apoptosis in vitro and in vivo [19]. A shorter time to progression was correlated with an elevated CRP level during the use of Bz-containing therapy, which suggested that a high level of CRP was involved in the rapid progress due to resistance to Bz [20]. Meanwhile, according to the up-to-date data [21, 22], CRP elevation was one of the potential side effects of Bz, especially for male patients older than 60 years old using the drug less than 1 month, but the mechanisms of DR remain largely unknown. Thereby, monitoring CRP changes is essential during Bz usage and lowering CRP to the baseline level may offer a potential way to alleviate the risk of Bz acquired resistance. In reference to the inferiority of the second-line usage of Bz for relapsed disease, the phase III VISTA trial and the phase II RETRIEVE study analyzed the efficacy of Bz retreatment after the initial response to Bz, with a response rate (≥ PR) in the range of 21%–50% [23, 24]. In contrast, Bz was approved by the FDA for treating relapsed MM patients in 2003, this was due, to a large extent, to data from 202 cases for whom previous therapies did not work, including traditional the chemotherapy of thalidomide or hematopoietic stem cell transplantation (HSCT) from 14 centers [25]. The response rate of second-line usage of Bz for RRMM was 59% [26–29]. In this study, the drug resistance rate to second-line usage of Bz was as high as 54.5% (n = 18) for the whole RRMM and 50% for the RRMM previously obtaining the Bz treatment. The clonal evolution theory indicates that there are emerging dominant clones that may now be drug resistance to the same drug [21]. Until now, for RRMM, whether the ineffective agent used previously could be reused for MM has given rise to a heated debate and awaits further investigation.
It has not been possible to assess the impact of the in vivo environment in the subgroup of patients with MM presenting de novo resistance to Bz, as well as both primary and acquired DR to thalidomide. Prospective and larger clinical trials are clearly needed to resolve this issue. The use of precise medicine would be a promising model providing tailored therapies to individual patients especially when the in vivo microenvironment influences the pathological processes via the interaction with MM cancerous components that varies from patient to patient. In addition, no correlation between DR and immune status indicators (bone metabolism and bone disease indicators) has been found in the multivariate analysis, which may be partly explained by the limited number of patients in our cohort and other factors should also be taken into considerations, namely, 1) the sensitivity of the measurements, for example, plain radiographs rather than MRI may show false negative results for myeloma-induced bone lesions, and 2) dynamic parameter tracking for particular individuals.
The number of samples with genetic information has been fairly limited. Among 63 cases with FISH results in the cohort, the gain of 1q21 has been implicated as a risk factor for conferring de novo DR of the new agents. In line with this possibility, increasing evidence has suggested that the 1q21 gain accounted for DR and poor OS [30, 31]. The 1q21 gain is also correlated with lower levels of serum cholesterol and LDL-C. In fact, metabolic pathways in multiple myeloma are essential for the regulation of DR [32], but based on genetic evidence, chromosome 1q21 was associated with type 2 diabetes susceptibility in Hong Kong Chinese [33].
Since there has not been a practical model from the routine workup of MM to effectively predict DR, we aimed to construct a model based on the differential expression of exosomal miRNAs. Given that Bz is the most widely used new drug for MM, exosome-associated miRNA panels, reflecting the crosstalk between MM cells and the in vivo environment, were explored in this study by comparing the Bz-resistance and Bz-response groups. Although in vivo tracking of the exosomes in humans is challenging, a conclusion is that the majority of the exosomes come from myeloma cells because the significant elevation of the total quantification of RNA when comparing both the myeloma subgroups to the healthy control and the resistant group to the response group. The microarray profiling found four exosomal miRNAs (miR-16, miR-15a, miR-20a and miR-17) are at the core of the miRNA-RNA regulatory signaling network predicted by integrating network analysis and Gene Ontology and were confirmed by quantitative real-time PCR. Our previous work and other findings suggest that down-regulation of miR-15a and miR-16 of primary MM cells as well as in MM cell lines could contribute to the DR and progression by the modulation of the bone marrow microenvironment [34, 35]. In addition, miR-20a and miR-17 are also involved in the tumorigenicity of MM [36]. Although miR-15a and miR-16 are located on 13q13.4 and miR-17-92 cluster of chromosome 13q31.3, their expression is independent of the loss of chromosome 13, which is observed in almost half of MM patients and resulted in poor prognosis. Further prospective studies on a larger cohort are required to clarify the potential role of specific exosomal miRNAs, namely, miR-15a and miR-16 with the putative BCL2 target and miR-20a and miR-17 with the putative Myc target.
Our study has the important implication of using serum exosomal miRNAs as drug resistance biomarkers for MM. First, cell-free circulating RNAs have opened a window to assess global alteration of the in vivo disease and an extensive list of circulating RNA-based biomarker candidates have been reported in the recent years [37, 38]. Significantly, in this study from the equivalent blood sample of the same patients, obviously higher concentrations of exosomal RNAs than circulating RNAs were found, suggesting that more effort is necessary to make a comprehensive assessment of the potential roles of exosomal RNAs in human disease. Second, unlike monogenic disorders such as chronic myeloid leukemia with the formation of the BCR-ABL fusion gene [39], MM is a complex polygenic trait shaped by multiple and translocation genetic anomalies, thus leading to a more obvious disease heterogeneity. The advances in systems biology and the development of new molecular tools in the “omics” science, including genomics, Rnomics, miRNomics, proteomics and epigenomics, have made it possible to reveal new findings in relation to DR in MM patients. The expression panel of miRNAs that are highly stable in blood and involved in a complex, multi-faceted network of regulatory interactions suggest that miRNAs may be the preferred option for circulating biomarker for MM in routine clinical practice since mRNAs and proteins are unstable molecules, in constant alteration and dynamic expression of mRNAs and proteins are frequently correlated with the events of each episode taking place within extended periods of time. Although DNAs are biologically stable, their mutations exert a great deal of diversity due to the complexity of assembled genomic clones and biological subtypes among individual MM cases. Our results still warrant further investigations because of the limitation of number of miRNAs microarray in each group and outside of Bz. Moreover, large multi-center studies are needed to explore the more precise exosome-associated microRNA expression models accompanying the development of novel agents.
MATERIALS AND METHODS
Patients
From January 2013 to December 2014, all the hospitalized MM patients in our tertiary hematology center, who received novel agents-based therapies, namely, bortezomib (Bz), thalidomide or lenalidomide, and did not participate in any clinical trials during the corresponding period, were enrolled in this study. This study was performed in accordance with the 1996 Declaration of Helsinki and approved by the ethics committees of West China Hospital, Sichuan University. The diagnosis of MM was established using the International Myeloma Working Group 2003 diagnostic criteria [40]. The cut-off date for follow-up was March 31st, 2015. The median follow-up was 23 months with a median follow-up of 23 months, and 38 patients (19.1%) were lost from follow-up sessions.
Study design and treatment
The overall treatment for patients with MM was divided into three phases: induction, consolidation and maintenance. Thalidomide-base therapy of MPT regimen specifically included melphalan orally 0.25 mg/kg on days 1–4, prednisone orally 1 mg/kg on days 1–4 and thalidomide orally 50–200 mg/day, with over 4–6 weeks as per cycle. Bz-based therapy of BD regimen specifically included Bz intravenously or subcutaneously 1.3 mg/m2 on days 1, 4, 8, 11 and dexamethasone orally or intravenously 20 mg/day on days 1–2, 4–5, 8–9, 11–12 with over 21-days per cycle. Lenalidomide-based therapy of Rd regimen specifically included lenalidomide orally 25 mg/day on days 1–21 and dexamethasone orally 20 mg on days 1, 8, 15 and 22 with over 28-days per cycle. After at least 4 cycles of induction treatment, patients with partial remission or better response underwent consolidation therapy, either autologous stem cell transplant (ASCT) or chemotherapy with the initial regimens, according to their intent and performance status. Subsequently, the patients with good tolerance would be treated with thalidomide (100–150 mg/day) for 1 year for maintenance, if the drug was well tolerated. The role of the in vivo environment was analyzed as contributors to DR, while the data based on the routine workup for MM were collected into 5 aspects as follows: (1) patients’ general information; (2) immune status indicators; (3) bone disease indicators; (4) nonspecific inflammatory markers; (5) genetic profile. More details are found in the Tables.
DR was defined according to the criteria of the 19th annual meeting of European Hematology Association. Briefly, the patients were divided into two groups based on therapeutic outcome: the response group and resistant group. The latter were further divided into the de novo drug resistant group (de novo DR) and the acquired drug resistant group (acquired DR). The de novo DR consisted of the patients who failed to achieve minimal remission (MR) or experienced progressive disease (PD) within 60 days when receiving the novel agent-based therapy for the first time, while the acquired DR consisted of those who developed DR when receiving the novel agent-based therapy and had previously been treated with the drug. The response group represented the patients who acquired partial remission (PR) or complete remission (CR) no matter the first or second line application of the novel agent-based regimen. Duration of response was defined as the time from the date of the first response to the date of PD or death due to PD. Survival time was defined as the time from diagnosis to the last follow-up or death.
Cell lines, cell culture and blood sample collection
The human multiple myeloma U266 cell line was purchased from American Type Culture Collection (ATCC) and cultured in RPMI 1640 containing 10% fetal bovine serum (FBS) at 37°C in 5% CO2. For exosome isolation, the U-266 was prepared with 10% SBI EXO-FBS-50A-1 Exosome-depleted FBS media supplement. Blood samples of the patients who would receive Bz-based regimen were collected into EDTA-coated tubes and in the same day processed under a protocol approved by the Institutional Review Board in Sichuan University, West China Hospital. Blood samples of the patients who preferred to receive the novel drug-based regimen were collected with written informed consent.
Exosome isolation and identification
Extracellular membrane microvesicles (EMVs) are circulating fragments of membrane, including exosomes, which are released from the endosomal compartments with diameters of 30 to 100 nm and microvesicles (MVs), which are shedding from the surface membranes of most cell types with diameters of 50 to 2,000 nm. Until now, even differential ultracentrifugation, the gold standard method for separating and purifying EMVs, is incapable of efficiently distinguishing between exosomes and MVs. Alternatively, based on the principle of aqueous gradient solubility differences between various lipids and nanoparticles, commercial kits are able to capture EMVs and then use either 0.2-micrometer (μm) pore size filters or artificially synthesized molecular sieves to enrich exosomes. In this study, we isolated and purified exosomes from both U266 cell line and MM samples, as depicted in Figure 1A. Then, the size and morphology of EMVs were observed by transmission electron microscope (TEM) (hitachi H-600 Japan) after negatively staining by 2% uranyl acetase. The size distribution of EMVs was traced by NanoSight NS300 (Malvern company, Great Britain) following manufacture protocols. Protein lysates of EMVs were assessed by Coomassie staining and western blotting for identification the expression of exosomal protein marker such as heat shock protein (Hsp70), CD63 and myeloma membrane-associated CD138.
RNA isolation and microRNA array profiling
Serum exosomal samples of 3 subjects were selected and harvested, respectively, from Bz-response and Bz-resistant groups. To extract circulating RNA from plasma samples, the miRNeasy Serum/Plasma Kit (Qiagen, Germany) was used according to the manufacturer’s instructions. Total RNA from cells or exosomes harvested from serum was isolated using TRIzol (Invitrogen, Carlsbad, CA). Quantity and quality of RNA was evaluated by nanodrop spectrophotometer (ND-1000, Nanodrop Technologies). An equal amount of 150 ng RNA from the 3 subjects of the same group was mixed to minimize differences among subjects within a group. Total RNA from both groups was labeled with Hy3™ fluorescent using the miRCURY™ Hy3™/Hy5™ Power labeling kit (Exiqon, Vedbaek, Denmark) following the procedure described by the manufacturer. The Hy3™-labeled samples were mixed and hybridized to the miRCURY™ LNA Array version 7th Generation (Exiqon), covering all human, mouse and rat miRNAs annotated in miRBase 18.0, as well as all viral microRNAs related to these species. In addition, this array contains capture probes for 25 miRPlus™ human miRNAs. The hybridization was performed according to the miRCURY™ LNA array manual. Then, the slides were scanned using the Axon GenePix 4000B microarray scanner (Axon Instruments, Foster City, CA), and scanned images were then imported into GenePix Pro 6.0 software (Axon) for grid alignment and data extraction. Replicated miRNAs were averaged and miRNAs that intensities > = 30 in all samples were chosen for calculating normalization factor. Expressed data were normalized using the Median normalization. After normalization, differentially expressed miRNAs were identified through Fold Change filtering. Hierarchical clustering was performed using MEV software (v4.6, TIGR). MicroRNA array experiments were performed by the Shanghai KangChen Bio-tech Company, Shanghai, China. Real-time PCR was used for confirmation. Differentially expressed miRNAs were identified through volcano plot screening. Cluster analysis was carried out by hierarchical clustering. Finally, a fold change analysis was performed by calculating the ratio between the two groups with a cut-off value of 2-fold changes. The mechanism of miRNA differentially expressed between groups were further explored by Ontology (GO) classification analyses through evaluating the genes affected by the upregulated and downregulated miRNAs, and Kyoto Encyclopedia of Genes and Genomes (KEGG) for pathway analysis were also employed to estimate the functions and pathways of miRNAs target genes.
Statistical analysis
Results were analyzed using SPSS version 13.00 software. For categorical variables, statistical data were described as frequency counts and percentages; for continuous variables, average, medians and ranges were adopted. Comparison of ratio and constituent ratio were using chi-square test. Median and average were computed by using independent samples T-test or one-way ANOVA. Kaplan-Meier method was used to estimate duration of response and OS, and the differences between groups were computed using stratified log-rank tests. All statistical tests were 2-sided, and P < 0.05 was considered statistically significant for all comparisons and confidence intervals refer to 95% boundaries.
ACKNOWLEDGMENTS
Not specified.
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
GRANT SUPPORT
This work was supported by the National Natural Science Foundation of China for Young Scholars grants 81302148 to Dr. Li Zhang.
Authors’ contributions
LZ, LP, TN, and TL initiated the work, designed the studies, and wrote the paper. LZ, BX, HZ, YW, MC, PG, JL, LX, HM, TD and XS performed the experiments and statistical analyses. FW, XZ, BD and CAV provided critical suggestions. The paper was modified by the native English speaker Prof. CAV.
REFERENCES
1. Kyle RA, Durie BG, Rajkumar SV, Landgren O, Blade J, Merlini G, Kröger N, Einsele H, Vesole DH, Dimopoulos M, San Miguel J, Avet-Loiseau H, Hajek R, et al. Monoclonal gammopathy of undetermined significance (MGUS) and smoldering (asymptomatic) multiple myeloma: IMWG consensus perspectives risk factors for progression and guidelines for monitoring and management. Leukemia. 2010; 24:1121–1127.
2. Child JA, Morgan GJ, Davies FE, Owen RG, Bell SE, Hawkins K, Brown J, Drayson MT, Selby PJ. Medical Research Council Adult Leukaemia Working Party. High-dose chemotherapy with hematopoietic stem-cell rescue for multiple myeloma. N Engl J Med. 2003; 348:1875–1883.
3. Kristinsson SY, Anderson WF, Landgren O. Improved long-term survival in multiple myeloma up to the age of 80 years. Leukemia. 2014; 28:1346–1348.
4. Turesson I, Velez R, Kristinsson SY, Landgren O. Patterns of Improved Survival in Patients With Multiple Myeloma in the Twenty-First Century: A Population-Based Study. J Clin Oncol. 2010; 28:830–834.
5. Kumar SK, Dispenzieri A, Lacy MQ, Gertz MA, Buadi FK, Pandey S, Kapoor P, Dingli D, Hayman SR, Leung N, Lust J, McCurdy A, Russell SJ, et al. Continued improvement in survival in multiple myeloma: changes in early mortality and outcomes in older patients. Leukemia. 2014; 28:1122–1128.
6. Hao M, Zhang L, An G, Meng H, Han Y, Xie Z, Xu Y, Li C, Yu Z, Chang H, Qiu L. Bone marrow stromal cells protect myeloma cells from bortezomib induced apoptosis by suppressing microRNA-15a expression. Leuk Lymphoma. 2011; 52:1787–1794.
7. Garrison LP, Neumann PJ, Erickson P, Marshall D, Mullins D. Using real-world data for coverage and payment decisions: The ISPOR real-world data task force report. Value Health. 2007; 10:326–335.
8. Smith AD, Roda D, Yap TA: Strategies for modern biomarker and drug development in oncology. J Hematol Oncol. 2014; 7:70.
9. Xu Zhang, Xiao Yuan, Hui Shi, Lijun Wu, Hui Qian, Wenrong Xu. Exosomes in cancer: small particle, big player. J Hematol Oncol. 2015; 8:83.
10. Srivatsava Naidu, Peter Magee, Michela Garofalo. MiRNA-based therapeutic intervention of cancer. J Hematol Oncol. 2015; 8:68.
11. Challagundla KB, Fanini F, Vannini I, Wise P, Murtadha M, Malinconico L, Cimmino A, Fabbri M. microRNAs in the tumor microenvironment: solving the riddle for a better diagnostics. Expert Rev Mol Diagn. 2014; 14:565–574.
12. Palumbo A, Anderson K. Multiple myeloma. N Engl J Med. 2011; 364:1046–1060.
13. Meads MB, Gatenby RA, Dalton WS. Environment-mediated drug resistance: a major contributor to minimal residual disease. Nat Rev Cancer. 2009; 9:665–674.
14. Graziani MS, Merlini G. Serum free light chain analysis in the diagnosis and management of multiple myeloma and related conditions. Expert Rev Mol Diagn. 2014; 14:55–66.
15. Larsen JT, Kumar SK, Dispenzieri A, Kyle RA, Katzmann JA, Rajkumar SV. Serum free light chain ratio as a biomarker for high-risk smoldering multiple myeloma. Leukemia. 2013; 27:941–946.
16. Giarin MM, Giaccone L, Sorasio R, Sfiligoi C, Amoroso B, Cavallo F, Cipriani A, Palumbo A, Boccadoro M. Serum free light chain ratio, total kappa/lambda ratio, and immunofixation results are not prognostic factors after stem cell transplantation for newly diagnosed multiple myeloma. Clin Chem. 2009; 55:1510–1516.
17. Offidani M, Corvatta L, Polloni C, Piersantelli MN, Galieni P, Visani G, Alesiani F, Catarini M, Brunori M, Burattini M, Centurioni R, Ferranti M, Giuliodori L, et al. Serum C-reactive protein at diagnosis and response to therapy is the most powerful factor predicting outcome of multiple myeloma treated with thalidomide/ anthracycline-based therapy. Clin Lymphoma Myeloma. 2008; 8:294–299.
18. Zahlten-Hinguranage A, Goldschmidt H, Cremer FW, Egerer G, Moehler T, Witte D. Preoperative elevation of serum C—reactive protein is predictive for prognosis in myeloma bone disease after surgery. Br J Cancer. 2006; 95:782–787.
19. Yang J, Wezeman M, Zhang X, Lin P, Wang M, Qian J, Wan B, Kwak LW, Yu L, Yi Q. Human C-reactive protein binds activating Fcgamma receptors and protects myeloma tumor cells from apoptosis. Cancer cell. 2007; 12:252–265.
20. San Miguel J, Bladé J, Boccadoro M, Cavenagh J, Glasmacher A, Jagannath S, Lonial S, Orlowski RZ, Sonneveld P, Ludwig H. A practical update on the use of bortezomib in the management of multiple myeloma. Oncologist. 2006; 11:51–61.
21. http://www.ehealthme.com/ds/velcade/c-reactive+protein+increased.
22. http://factmed.com/report-VELCADE-causing-C-REACTIVE%20PROTEIN%20INCREASED.php.
23. Sood R, Carloss H, Kerr R, Lopez J, Lee M, Druck M, Walters IB, Noga SJ. Retreatment with bortezomib alone or in combination for patients with multiple myeloma following an initial response to bortezomib. Am J Hematol. 2009; 84:657–660.
24. Petrucci MT, Giraldo P, Corradini P, Teixeira A, Dimopoulos MA, Blau IW, Drach J, Angermund R, Allietta N, Broer E, Mitchell V, Bladé J. A prospective, international phase 2 study of bortezomib retreatment in patients with relapsed multiple myeloma. Br J Haematol. 2013; 160:649–659.
25. Keats JJ, Chesi M, Egan JB, Garbitt VM, Palmer SE, Braggio E, Van Wier S, Blackburn PR, Baker AS, Dispenzieri A, Kumar S, Rajkumar SV, Carpten JD, et al. Clonal competition with alternating dominance in multiple myeloma. Blood. 2012; 120:1067–1076
26. Richardson PG, Barlogie B, Berenson J, Singhal S, Jagannath S, Irwin D, Rajkumar SV, Srkalovic G, Alsina M, Alexanian R, Siegel D, Orlowski RZ, Kuter D, et al. A phase 2 study of bortezomib in relapsed, refractory myeloma. N Engl J Med. 2003; 348:2609–2617.
27. Cavo M, Tacchetti P, Patriarca F, Petrucci MT, Pantani L, Galli M, Di Raimondo F, Crippa C, Zamagni E, Palumbo A, Offidani M, Corradini P, Narni F, et al. Bortezomib with thalidomide plus dexamethasone compared with thalidomide plus dexamethasone as induction therapy before, and consolidation therapy after, double autologous stem-cell transplantation in newly diagnosed multiple myeloma: a randomised phase 3 study. Lancet. 2010; 376:2075–2085.
28. Harousseau JL, Attal M, Leleu X, Troncy J, Pegourie B, Stoppa AM, Hulin C, Benboubker L, Fuzibet JG, Renaud M, Moreau P, Avet-Loiseau H. Bortezomib plus dexamethasone as induction treatment prior to autologous stem cell transplantation in patients with newly diagnosed multiple myeloma: results of an IFM phase II study. Haematologica. 2006; 91:1498–1505.
29. Sonneveld P, Schmidt-Wolf IG, van der Holt B, El Jarari L, Bertsch U, Salwender H, Zweegman S, Vellenga E, Broyl A, Blau IW, Weisel KC, Wittebol S, Bos GM, et al. Bortezomib induction and maintenance treatment in patients with newly diagnosed multiple myeloma: results of the randomized phase III HOVON-65/ GMMG-HD4 trial. J Clin Oncol. 2012; 30:2946–2955.
30. An G, Xu Y, Shi L, Shizhen Z, Deng S, Xie Z, Sui W, Zhan F, Qiu L. Chromosome 1q21 gains confer inferior outcomes in multiple myeloma treated with bortezomib but copy number variation and percentage of plasma cells involved have no additional prognostic value. Haematologica. 2014; 99:353–359.
31. Biran N, Malhotra J, Bagiella E, Cho HJ, Jagannath S, Chari A. Patients with newly diagnosed multiple myeloma and chromosome 1 amplification have poor outcomes despite the use of novel triplet regimens. Am J Hematol. 2014; 89:616–620.
32. Maiso P, Huynh D, Moschetta M, Sacco A, Aljawai Y, Mishima Y, Asara JM, Roccaro AM, Kimmelman AC, Ghobrial IM. Metabolic signature identifies novel targets for drug resistance in multiple myeloma. Cancer Res. 2015; 75:2071–2082.
33. Ng MC, So WY, Cox NJ, Lam VK, Cockram CS, Critchley JA, Bell GI, Chan JC. Genome-wide scan for type 2 diabetes loci in Hong Kong Chinese and confirmation of a susceptibility locus on chromosome 1q21-q25. Diabetes. 2004; 53:1609–1613.
34. Hao M, Zhang L, An G, Sui W, Yu Z, Zou D, Xu Y, Chang H, Qiu L. Suppressing miRNA-15a/-16 expression by interleukin-6 enhances drug-resistance in myeloma cells. J Hematol Oncol. 2011; 4:37
35. Sun CY, She XM, Qin Y, Chu ZB, Chen L, Ai LS, Zhang L, Hu Y. miR-15a and miR-16 affect the angiogenesis of multiple myeloma by targeting VEGF. Carcinogenesis. 2013; 34:426–435.
36. Chen L, Li C, Zhang R, Gao X, Qu X, Zhao M, Qiao C. miR-17–92 cluster microRNAs confers tumorigenicity in multiple myeloma. Cancer Lett. 2011; 309:62–70.
37. Rocci A, Hofmeister CC, Geyer S, Stiff A, Gambella M, Cascione L, Guan J, Benson DM, Efebera YA, Talabere T, Dirisala V, Smith EM, Omedè P, et al. Circulating miRNA markers show promise as new prognosticators for multiple myeloma. Leukemia. 2014; 28:1922–1926.
38. Wen-Tao Wang, Yue-Qin Chen. Circulating miRNAs in cancer: from detection to therapy. J Hematol Oncol. 2014; 7:86.
39. Hanfstein B, Shlyakhto V, Lauseker M, Hehlmann R, Saussele S, Dietz C, Erben P, Fabarius A, Proetel U, Schnittger S, Krause SW, Schubert J, Einsele H, et al. Velocity of early BCR-ABL transcript elimination as an optimized predictor of outcome in chronic myeloid leukemia (CML) patients in chronic phase on treatment with imatinib. Leukemia. 2014; 28:1988–1992.
40. International Myeloma Working Group. Criteria for the classification of monoclonal gammopathies, multiple myeloma and related disorders: a report of the International Myeloma Working Group. Br J Haematol. 2003; 121:749–757.

