INTRODUCTION
Taxanes and anthracyclines are widely used for the treatment of breast cancer yet highly toxic to patients [1–4]. At present there is no biological marker or assay available to identify which subset of patients will benefit from chemotherapy although numerous molecules have been investigated with limited success [4–12]. Research from our group [8] linked the predictive effect of CEP17 in vivo to chromosome instability (CIN), which itself is predictive of anthracycline benefit in the BR9601 trial [8]. We recently derived a four gene signature that was predictive of anthracycline benefit and demonstrated that patients with low tumour CIN4 scores benefited from anthracycline treatment significantly more than those with high CIN4 scores (HR 0.37, 95% CI 0.20-0.56, p=0.001) [13]. Given that most patients now receive taxane based chemotherapy we sought to extend the validation of this marker into trials including taxanes.
The NCIC Clinical Trials Group (CTG) MA.21 trial compared dose-intense cyclophosphamide, epirubicin and fluorouracil (CEF) to dose-dense, dose-intense epirubicin, cyclophosphamide with added paclitaxel (EC/T) and to a standard at the time of doxorubicin, cyclophosphamide and paclitaxel (AC/T). Analyses comparing the two epirubicin arms of the trial (CEF vs EC/T) did not demonstrate benefit from the addition of taxane (EC/T vs CEF) in terms of relapse free survival (RFS) (HR 0.85, 95% CI 0.64-1.22, p=0.46) [14].
Our present study, we evaluated the predictive effect of the CIN signatures in patients in the NCIC CTG MA.21 trial and hypothesized that CIN signatures are associated with clinical outcome and resistance to taxane therapy.
RESULTS
Characteristics
Table 1 shows the baseline characteristics of the MA.21 CIN study population, whereas Figure 1 illustrates the total number of cases available for analysis; 342 RNA samples from CEF arm and 336 RNA samples from the ECT arm were successfully analysed on the NanoString platform. High CIN4, CIN25 and CIN70 scores were defined as above the median and were previously described [15–13].
Table 1: Baseline patient and tumour characteristics from the MA.21 study
Baseline characteristics |
CEF 342 (100%) |
ECT 336 (100%) |
|---|---|---|
Age |
||
≤39 |
56 (16.4%) |
59 (17.6%) |
40-49 |
159 (46.5%) |
146 (43.4%) |
50-59 |
126 (36.8%) |
128 (38.1%) |
60-69 |
1 (0.3%) |
3 (0.9%) |
Median |
||
Menopausal status |
||
Post-menopausal |
102 (29.8%) |
101 (30.1%) |
Pre-menopausal |
240 (70.2%) |
235 (69.9%) |
#of positive axillary nodes |
||
0 |
109 (31.9%) |
102 (30.4%) |
1-3 |
136 (39.8%) |
146 (43.5%) |
4-10 |
79 (23.0%) |
72 (21.4%) |
>10 |
18 (5.3%) |
16 (4.7%) |
Tumour stage |
||
T1 |
108 (31.6%) |
114 (34.2%) |
T2 |
200 (58.5%) |
177 (53.2%) |
T3 |
30 (8.8%) |
37 (11.1%) |
T4 |
4 (1.1%) |
5 (1.5%) |
Missing |
3 |
|
ER Status |
||
Positive |
195 (57.0%) |
196 (58.3%) |
Negative |
147 (43.0%) |
140 (41.7%) |
CIN4 |
||
High |
161 (47.1%) |
164 (48.9%) |
Low |
181 (52.9%) |
173 (51.1%) |
CIN25 |
||
High |
183 (53.5%) |
186 (55.2%) |
Low |
159 (46.5%) |
151 (44.8%) |
CIN70 |
||
High |
181 (52.9%) |
186 (55.2%) |
Low |
161 (47.1%) |
151 (44.8%) |
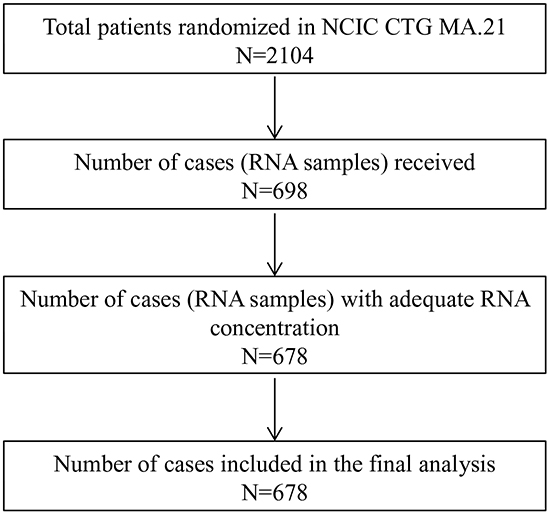
Figure 1: REMARK diagram for the evaluation of chromosome instability (CIN) gene expression signatures in the NCIC CTG MA.21 trial.
Correlation of CIN signatures and clinicopathological parameters with clinical outcomes
In univariate analysis using continuous clinico-pathological biomarkers, high CIN4 was associated with ER positivity (p<0.0001) and grade (p<0.001), whereas CIN25 and CIN70 were associated with younger age (p=0.01 and p=0.001, respectively), increase in the number of positive axillary nodes (p<0.0001) and ER negativity (p<0.0001) (Table 2).
Table 2: Correlations between CIN scores and conventional pathological markers
CIN4 |
CIN25 |
CIN70 |
|
|---|---|---|---|
Age |
NS |
-0.128 |
-0.131 |
Positive axillary node |
NS |
-0.246 |
-0.226 |
Tumour size |
NS |
NS |
NS |
Grade |
-0.195 |
0.528 |
0.500 |
ER |
0.092 |
-0.430 |
-0.413 |
HER2 |
NS |
NS |
NS |
CIN signatures as a prognostic marker for RFS
The prognostic impact of the CIN signatures were tested on the entire cohort, irrespective of allocated adjuvant chemotherapy. No statistically significant association was observed between CIN4 or CIN70 scores and RFS (HR: 1.11, 95% CI 0.79-1.55, p=0.548 and HR: 0.73, 95% CI 0.52-1.02, p=0.07, (Figure 2A) respectively). In contrast, tumours with high CIN25 scores were associated with increased RFS (HR: 1.52, 95%CI 2.13-10.64, p=0.02) (Figure 2B). After multivariate analysis and adjustment for nodal status, grade, size, age, HER2 and ER status, both high CIN25 and CIN70 scores were significantly associated with an increased RFS (HR 1.76, 95%CI 1.07-2.86, p=0.0018 and HR 1.59, 95%CI 1.12-2.25, p=0.0096 respectively).
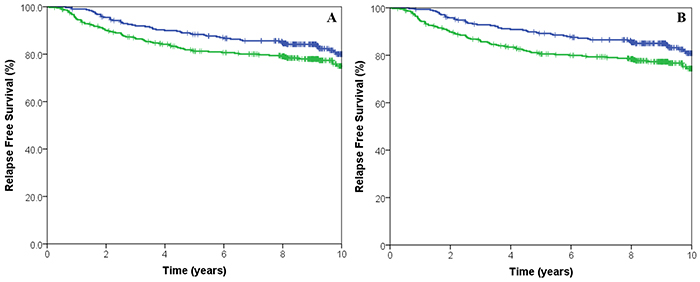
Figure 2: Kaplan-Meier survival curves for low CIN70 score (green line) and high CIN70 score (blue line) for relapse free survival. A. Kaplan-Meier survival curves for low CIN25 score (green line) and high CIN25 score (blue line) for relapse free survival B.
CIN signatures as predictive markers of taxane benefit
Subsequent analysis focused on the differential impact of multigene signatures on RFS between patients receiving anthracycline plus taxane (EC/T) therapy and those given anthracycline treatment (CEF) alone. Univariate analysis demonstrated no significant difference in benefit from ECT versus CEF treatment between patients whose tumours had high (HR 0.79, 95%CI 0.51-1.23, p=0.30) or low CIN70 expression (HR 1.30, 95% CI 0.76-2.21, p=0.34) (Table 3). Similarly there was no differential benefit from ECT treatment between patients whose tumours had high (HR 0.78, 95%CI 0.51-1.20, p=0.26) or low CIN25 expression (HR 1.35, 95% CI 0.78-2.34, p=0.28).
Table 3: Hazard ratios for relapse free survival comparing cyclophosphamide, epirubicin and fluorouracil (CEF) with cyclophosphamide, epirubicin and paclitaxel (EC/T) by biomarker status
Relapse Free Survival |
||||||
|---|---|---|---|---|---|---|
Low Biomarker |
High Biomarker |
Treatment*Marker |
||||
HR |
95% CI |
HR |
95% CI |
HR |
Test for Interaction P |
|
CIN70 |
1.30 |
0.76-2.21 |
0.79 |
0.51-1.23 |
1.43 |
0.355 |
CIN25 |
1.35 |
0.78-2.34 |
0.78 |
0.51-1.20 |
1.57 |
0.263 |
CIN4 |
0.64 |
0.39-1.03 |
1.53 |
0.91-2.57 |
0.49 |
0.066 |
The hazard ratio for treatment by marker interaction of CIN25 and CIN70 before correction for clinical variables was 0.58 (95% CI 0.29-1.17, p=0.128) and 0.61 (95% CI 0.31-1.122, p=0.166), respectively. After correction for size, nodal status, ER status, HER2, grade, CIN4 and treatment, the hazard ratio was 0.64 (95% CI 0.29-1.40, p=0.263) and 0.70 (95% CI 0.33-1.50, p=0.355) for CIN25 and CIN70, respectively. However, a trend was noted in patients whose tumours had low CIN4 gene expression scores with an increase in RFS (HR: 0.64, 95% CI 0.39-1.03, p=0.06) when treated with EC/T compared to patients treated with CEF (Figure 3). There was no statistical significant difference in survival in patients that has high CIN4 gene expression scores (HR: 1.54, 95% CI 0.1-2.57, p=0.107). The hazard ratio for treatment by marker interaction of CIN4 before correction for clinical variables was 0.49 (95% CI 0.25-0.97, p=0.04) for RFS. After correction for size, nodal status, ER status, HER2, grade, CIN4 and treatment, the hazard ratio was 0.49 (95% CI 0.23-1.05, p=0.066).
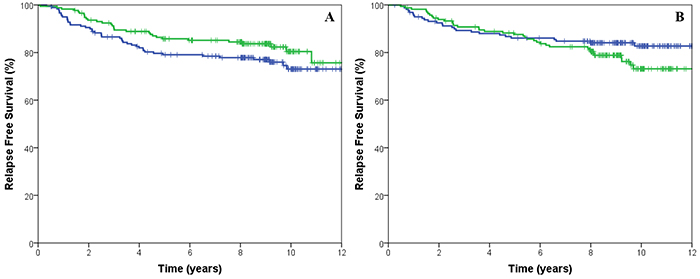
Figure 3: Kaplan-Meier survival curves for cyclophosphamide, epirubicin and fluorouracil (CEF, blue) treated with cyclophosphamide, epirubicin and paclitaxel (EC/T, green line) for relapse free survival stratified by low CIN4 expression. A. and high CIN4 expression B.
DISCUSSION
CIN is associated with poor prognosis in patients with solid tumours [16–15]. We have previously shown that CIN is associated with sensitivity to anthracycline treatments [8–13] while others have demonstrated that CIN is associated with taxane resistance [17]. Since modern chemotherapy treatment of breast cancer includes the addition of a taxane, we examined the role of three CIN gene signatures in the NCIC CTG MA.21 breast cancer trial that compares taxane versus no-taxane containing arms. Our study has demonstrated that patients with low expression of the CIN4 gene signature exhibited greater benefit from the EC/T treatment regimen versus the CEF regimen. This is in contradiction to previous work in ovarian cancer suggesting a CIN70 gene expression signature was predictive of resistance to taxanes and sensitivity to carboplatin treatment [17]. However, it is unclear if our current result reflects the addition of a taxane to the treatment regimen or simply extending the duration of the chemotherapy regimen. Both arms included a cumulative dose of 720mg/m2 of epirubicin however; the EC/T arm had an additional four cycles of paclitaxel [14]. Therefore, our result could reflect either the addition of paclitaxel or the need for extended chemotherapy. However, there is accumulating evidence to support the former. For example, the UK TACT study investigated whether sequential docetaxel after anthracycline chemotherapy would improve patient outcome compared to standard chemotherapy of a similar duration. This study demonstrated no difference in disease free survival between the FEC-D group and the control group (HR 0.95, 95% CI 0.85-1.08, p=0.44) [18]. Meta-analyses performed by the EBCTCG group demonstrated the incorporation of taxanes into an anthracycline regimen resulted in the reduction of the risk of recurrence and survival at 8 years [19]. This analysis recognised there was variation in trial designs when comparing the effects of additional cycles of just a taxane to a constant background chemotherapy regimen. However, in the taxane trials with different number of cycles the 8-years breast cancer mortality was 21.1% for the taxane groups versus 23.9% for the control groups. In contrast in trials examining the effects of a taxane regimen with the same number of cycles non-taxane chemotherapy demonstrated the 5-year a small but significant reduction in breast cancer mortality [19]. These results would indicate that the addition of a taxane to the chemotherapy regimen improves survival and the CIN4 is predictive of taxane benefit.
In agreement with other studies, we have demonstrated that the CIN25 and CIN70 signature is an independent prognostic biomarker in breast cancer [15–20]. We performed our analysis on the NanoString platform, which has been shown to be a robust and reproducible platform that can easily be incorporated into the pathology laboratory setting [21]. The identification of methods that predict breast cancer clinical outcome is being studied in numerous labs around the world. There are a number of prognostic expression arrays available such as Oncotype DX [22], Mammaprint [23] and Prosigna [21] that predict patient outcome in estrogen receptor (ER) positive patients; however, none of them are tailored towards the ER negative population. The CIN25 and CIN70 gene expression signatures were derived irrespective of ER status and contain genes, such as TPX2, TOP2A, MAD2L1, CCNB1, CDC20 and PTTG1 that are involved in spindle assembly checkpoint (SAC) as well as DNA damage checkpoint genes [15]. Therefore, the CIN25 and CIN70 signatures are credible candidates for use in the prognostic setting.
Ultimately, although there are a number of prognostic markers available, but there are no predictive signatures associated with specific chemotherapies to level I evidence. We had previously demonstrated the CIN4 signature to be predictive of anthracycline benefit [13], in this study we demonstrate that CIN4 has potential to be predictive for benefit from the addition of a taxane to chemotherapy treatment.
MATERIALS AND METHODS
Patients
The MA.21 (ClinicalTrials.gov (NCT00014222)) trial recruited 2104 pre- and post-menopausal women with histologically confirmed node positive or high risk node negative adenocarcinoma of the breast treated with either total or partial mastectomy. Patients were randomised to one of three regimens to receive: 1) 6 cycles of CEF (epirubicin (60mg/m2) and fluorouracil (500mg/m2) given intravenously on days 1 and 8 and oral cyclophosphamide (75mg/m2) on days 1-14) every 28 days, 2) 6 cycles of EC/T (epirubicin (120mg/m2) and cyclophosphamide (830mg/m2) on day 1 and filgrastim (G-CSF) subcutaneously (SC) on days 2-13 followed by 4 cycles of paclitaxel (175mg/m2)) every 21 days, 3) 4 cycles of AC/T (doxorubicin (60mg/m2) and cyclophosphamide (600mg/m2) on day 1 every 21 days followed by 4 cycles of paclitaxel (175mg/m2)) every 21 days. Treatment in all arms continued in the absence of disease progression or unacceptable toxicity; patient characteristics are shown in Table 1. The primary end point for the MA.21 trial was relapse-free survival (RFS) defined as the time from randomization to the time of recurrence of the primary disease. The protocol was approved by central and local ethics committees, and each patient provided written informed consent prior to randomization. For the current analysis, tissue blocks were retrieved for RNA extraction.
RNA extraction
Total RNA from FFPE tissue samples (2 x 10µM sections) was extracted using the RecoverAll Total Nucleic Acid Isolation kit (Life Technologies) according to the manufacturer’s protocol and concentrations were determined using the NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies).
Gene expression analysis
RNA (200ng) was used with the nCounter system, according to the manufacturer’s protocol (NanoString ® Technologies, Seattle, WA, USA). In brief, 5µl of RNA was hybridized at 96°C overnight with the NanoString Codeset. Probes for the analysis were synthesized by NanoString technologies and included probes for the 93 genes of interest and 6 normalising genes (Supplementary Table S1); all 99 genes were assayed simultaneously in multiplexed reactions. After probe hybridizations and NanoString nCounter digital reading, counts for each RNA species analyzed. The nCounter CodeSet contains two types of built-in controls: positive controls (spiked RNA at various concentrations to assess the overall assay performance) and negative controls (probes for background calculation). Raw mRNA abundance count data were pre-processed using the NanoStringNorm R package (v1.1.19) using normalization factors derived from the geometric mean of housekeeping genes, mean of negative controls and geometric mean of positive controls [24]. The CIN4/25/70 scores were developed and analysed as described in [13].
Statistics
The SAS (SAS Institute Inc., Cary, NC, USA, version 9.2) statistical package was used for statistical analysis. Kaplan-Meier curves of survival were used for estimation of relapse free survival (RFS), whereas log-rank test was used to compare RFS among different groups of patients. The Cox proportional hazards model was used to estimate hazard ratios for relapse. When comparing outcomes between the treatment arms within the groups of patients identified by biomarker expression, p-values were not calculated for sub-groups to avoid multiple testing and bias where one group was much smaller than the other. The Cox model was instead used to identify statistically significant interactions (p<0.05) between biomarkers and outcome on the different treatments (treatment by marker interaction), in models that also included biomarker status (marker effect) and treatment, as covariates. In exploratory analyses a value of p<0.05 was considered statistically significant as the Bonferroni correction was not applied.
ACKNOWLEDGMENTS
This work was supported by the funding from OICR. We thank the government of Ontario for funding, which is provided through the Ontario Ministry of Research and Innovation.
CONFLICTS OF INTEREST
All authors state that they have no financial conflicts of interest.
REFERENCES
1. Swain SM, Whaley FS, and Ewer MS. Congestive heart failure in patients treated with doxorubicin: a retrospective analysis of three trials. Cancer. 2003; 97: 2869-2879.
2. Mercuro G, Cadeddu C, Piras A, Dessi M, Madeddu C, Deidda M, Serpe R, Massa E, and Mantovani G. Early epirubicin-induced myocardial dysfunction revealed by serial tissue Doppler echocardiography: correlation with inflammatory and oxidative stress markers. Oncologist. 2007; 12: 1124-1133.
3. Jones LW, Haykowsky MJ, Swartz JJ, Douglas PS, and Mackey JR. Early breast cancer therapy and cardiovascular injury. J.Am.Coll.Cardiol. 2007; 50: 1435-1441.
4. Bartlett JM, A'Hern R, Piper T, Ellis IO, Dowsett M, Mallon EA, Cameron DA, Johnston S, Bliss JM, Ellis P, and Barrett-Lee PJ. Phosphorylation of AKT pathway proteins is not predictive of benefit of taxane therapy in early breast cancer. Breast Cancer Res Treat. 2013; 138: 773-781.
5. Martin M, Romero A, Cheang MC, Lopez Garcia-Asenjo JA, Garcia-Saenz JA, Oliva B, Roman JM, He X, Casado A, de la Torre J, Furio V, Puente J, Caldes T et al . Genomic predictors of response to doxorubicin versus docetaxel in primary breast cancer. Breast Cancer Res.Treat. 2011; 128: 127-136.
6. Pritchard KI, Messersmith H, Elavathil L, Trudeau M, O'Malley F, and Dhesy-Thind B. HER-2 and topoisomerase II as predictors of response to chemotherapy. J Clin Oncol. 2008; 26: 736-744.
7. Pritchard KI, Shepherd LE, O'Malley FP, Andrulis IL, Tu D, Bramwell VH, and Levine MN. HER2 and responsiveness of breast cancer to adjuvant chemotherapy. 2006; 354: 2103-2111.
8. Munro AF, Twelves C, Thomas JS, Cameron DA, and Bartlett JM. Chromosome instability and benefit from adjuvant anthracyclines in breast cancer. Br J Cancer. 2012; 107: 71-74.
9. Bartlett JM, Munro AF, Dunn JA, McConkey C, Jordan S, Twelves CJ, Cameron DA, Thomas J, Campbell FM, Rea DW, Provenzano E, Caldas C, Pharoah P et al . Predictive markers of anthracycline benefit: a prospectively planned analysis of the UK National Epirubicin Adjuvant Trial (NEAT/BR9601). Lancet Oncol. 2010; 11: 266-274.
10. Torikoshi Y, Gohda K, Davis ML, Symmans WF, Pusztai L, Kazansky A, Nakayama S, Yoshida T, Matsushima T, Hortobagyi GN, Ishihara H, Kim SJ, Noguchi S et al . Novel functional assay for spindle-assembly checkpoint by cyclin-dependent kinase activity to predict taxane chemosensitivity in breast tumor patient. J.Cancer. 2013; 4: 697-702.
11. Noguchi S. Predictive factors for response to docetaxel in human breast cancers. Cancer Sci. 2006; 97: 813-820.
12. Bartlett JM, Nielsen TO, Gao D, Gelmon KA, Quintayo MA, Starczynski J, Han L, Burnell MJ, Levine MN, Chen BE, Shepherd LE, and Chapman JW. TLE3 is not a predictive biomarker for taxane sensitivity in the NCIC CTG MA.21 clinical trial. Br.J.Cancer. 2015; 113: 722-728.
13. Spears M, Yousif F, Lyttle N, Boutros PC, Munro AF, Twelves C, Pritchard KI, Levine MN, Shepherd L, and Bartlett JM. A four gene signature predicts benefit from anthracyclines: evidence from the BR9601 and MA.5 clinical trials. Oncotarget. 2015; 6:31693-701. doi: 10.18632/oncotarget.5562.
14. Burnell M, Levine MN, Chapman JA, Bramwell V, Gelmon K, Walley B, Vandenberg T, Chalchal H, Albain KS, Perez EA, Rugo H, Pritchard K, O'Brien P et al . Cyclophosphamide, epirubicin, and Fluorouracil versus dose-dense epirubicin and cyclophosphamide followed by Paclitaxel versus Doxorubicin and cyclophosphamide followed by Paclitaxel in node-positive or high-risk node-negative breast cancer. J.Clin.Oncol. 2010; 28: 77-82.
15. Carter SL, Eklund AC, Kohane IS, Harris LN, and Szallasi Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat Genet. 2006; 38: 1043-1048.
16. Habermann JK, Doering J, Hautaniemi S, Roblick UJ, Bundgen NK, Nicorici D, Kronenwett U, Rathnagiriswaran S, Mettu RK, Ma Y, Kruger S, Bruch HP, Auer G et al . The gene expression signature of genomic instability in breast cancer is an independent predictor of clinical outcome. Int J Cancer. 2009; 124: 1552-1564.
17. Swanton C, Nicke B, Schuett M, Eklund AC, Ng C, Li Q, Hardcastle T, Lee A, Roy R, East P, Kschischo M, Endesfelder D, Wylie P et al . Chromosomal instability determines taxane response. Proc Natl Acad Sci U S A. 2009; 106: 8671-8676.
18. Ellis P, Barrett-Lee P, Johnson L, Cameron D, Wardley A, O'Reilly S, Verrill M, Smith I, Yarnold J, Coleman R, Earl H, Canney P, Twelves C et al . Sequential docetaxel as adjuvant chemotherapy for early breast cancer (TACT): an open-label, phase III, randomised controlled trial. Lancet. 2009; 373: 1681-1692.
19. Peto R, Davies C, Godwin J, Gray R, Pan HC, Clarke M, Cutter D, Darby S, McGale P, Taylor C, Wang YC, Bergh J, Di LA et al . Comparisons between different polychemotherapy regimens for early breast cancer: meta-analyses of long-term outcome among 100,000 women in 123 randomised trials. Lancet. 2012; 379: 432-444.
20. Birkbak NJ, Eklund AC, Li Q, McClelland SE, Endesfelder D, Tan P, Tan IB, Richardson AL, Szallasi Z, and Swanton C. Paradoxical relationship between chromosomal instability and survival outcome in cancer. Cancer Res. 2011; 71: 3447-3452.
21. Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J et al . Supervised risk predictor of breast cancer based on intrinsic subtypes. J.Clin.Oncol. 2009; 27: 1160-1167.
22. Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL et al . A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N.Engl.J.Med. 2004; 351: 2817-2826.
23. van 't Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C et al . Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002; 415: 530-536.
24. Waggott D, Chu K, Yin S, Wouters BG, Liu FF, and Boutros PC. NanoStringNorm: an extensible R package for the pre-processing of NanoString mRNA and miRNA data. Bioinformatics. 2012; 28: 1546-1548.

