INTRODUCTION
Gastric cancer (GC), one of the serious threats to human health contributing to a malignant tumor in the digestive system, is currently the third most common cause of deaths associated with cancer worldwide, accounting for approximately 9% of cancer-related deaths [1]. Although survival advantages can be achieved by surgical resection, combined with adjuvant/post-operative chemotherapy and/or radiotherapy in patients with early stage GC, almost two-thirds GC patients suffer from locally advanced or metastatic diseases, with poor survival outcomes of a median of 10 months period [2]. However, only a handful of new chemotherapeutic agents or molecular-targeted therapies have been developed. Therefore, further investigations in the underlining mechanisms of incidence and progression, as well as determining the development of prognosis and treatment biomarkers, will provide a comprehensive picture of their impacts on survival and invaluable reference allowing new agents/therapies.
Recently, Brungs D et al. systematically reviewed the current evidence for cancer stem cell (CSC) in GC [3], which the evidence includes superior tumor initiation, growth, and metastatic potential to that of other tumor cells [4]. It is reported that CSC is also responsible for the renewal of tumor mass after systemic treatment and development of sub-clones with more resistance to chemotherapeutics [5]. So far, it has been a challenge to identify a CSC population within a tumor. The methods for CSC identification and isolation mainly depend on side population (SP) assay and combinations of cell surface markers expression [3]. Recently, many protein are reported to be the putative gastric cancer CSC markers, such as CD26 [6], CD44 [7], ALDH1 [8], and CD133 [9], and Lgr5, which was reported to be co-localized with other CSC markers and may be functionally-associated [10].
Aldehyde dehydrogenase (ALDH) is a family of intracellular enzymes that are involved in cellular detoxification, differentiation, and drug resistance by oxidation of cellular aldehydes [11]. ALDH1, a marker of CSC in a variety of cancers, plays as a modulator for cell proliferation and stem cell differentiation, as well as resistance to chemotherapeutic agents [12]. ALDH1 positivity in diffuse-type lymph node metastasis was significantly higher than that in primary, predicting that CSC markers are important in tumor invasion and metastasis and may be prognostic markers in GC patients [13]. However, it is found that the expression of ALDH1 and REG4 can be reduced by TGF-β signaling pathway, resulting in decreasing ALDH1+ cell population size and tumorigenic capacity of diffuse-type gastric carcinoma-initiating cells [14]. CSC markers, such as ALDH1, LGR5 and CD166 were expressed in very low levels in normal human gastric mucosa or young rat gastric mucosa, compared with in H. pylori gastritis and gastric adenocarcinoma as well as in normal gastric mucosa in aged rats [15].
Although ALDH1 has been identified as a reliable marker of GC and other solid tumors, the functions of different ALDH1 isoforms in contribution to ALDH1 activities and prognostic value are still unclear. Recently, Wu et al. and You et al. reported the distinct prognostic values of ALDH1 isoenzymes in breast cancer and non-small-cell lung cancer (NSCLC), respectively [16, 17], providing a comprehensive illustration of the prognostic value of individual ALDH1 isoenzymes. In this study, we extended the research field to gastric cancers, to determine the expression pattern of ALDH1 isoenzymes in gastric cancer versus. normal tissues and distinct prognostic value of them.
RESULTS
Different transcription levels of ALDH1 isoforms in gastric cancers
Up to now, 6 sub-members were reported of the ALDH1 family in Figure 1 [16]. ONCOMINE analysis of 6 ALDH1 sub-members in cancer vursus. normal samples showed that ALDH1A1 was significantly downregulated in different types of GC in different datasets. In Cho’s dataset, the transcription levels of ALDH1A1 in both gastric intestinal type adenocarcinoma and diffuse gastric adenocarcinoma were lower than that in gastric tissues (Fold changes were −5.046 and −3.278, respectively) (Figure 2A and 2B) [18]. The ALDH1A1 mRNA level was also decreased in diffuse gastric adenocarcinoma (Figure 2C) and gastric intestinal type adenocarcinoma (Figure 2D), compared with gastric mucosa [19, 20]. In another dataset with only 27 samples, the ALDH1A1 mRNA level in GC was downregulated, compared with both gastric mucosa and gastric tissue (Figure 2E) [21].
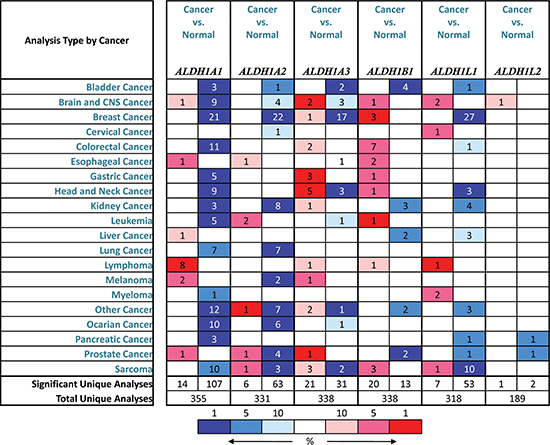
Figure 1: Different ALDH1 isoform mRNA expression in different tumor types. This graphic showed the numbers of datasets with statistically significant mRNA overexpression (red) or downexpression (blue) of the target gene (cancer vs. normal tissue). The p value threshold is 0.01. The number in each cell represents the number of analyses that meet the threshold within those analysis and cancer types. The gene rank was analyzed by percentile of target gene in the top of all genes measured in each research. Cell color is determined by the best gene rank percentile for the analyses within the cell.
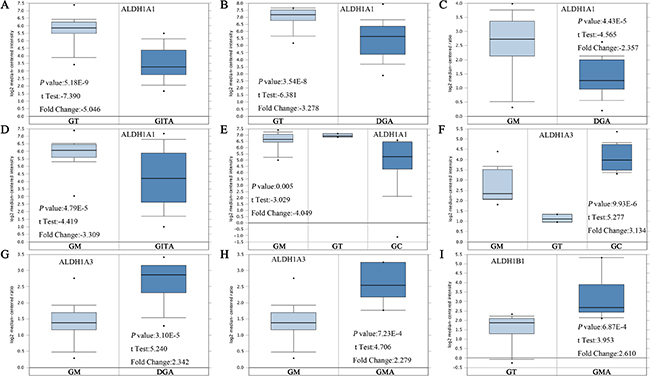
Figure 2: ALDH1 isoforms analysis in gastric cancer (ONCOMINE database). Box plots derived from gene expression data in ONCOMINE comparing expression of a specific ALDH1 isoform in normal and GC tissue. The p value was set up at 0.01 and fold change was defined as 2. (A–E) Comparison of ALDH1A1 mRNA expression. (F–H) Comparison of ALDH1A3 mRNA expression. (I) Comparison of ALDH1B1 mRNA expression [18–21]. Abbreviation: gastric tissue (GT), gastric intestinal type adenocarcinoma (GITA), diffuse gastric adenocarcinoma (DFA), gastric mucosa (GM), gastric cancer (GC), gastric mixed adenocarcinoma (GMA).
In contrast, overexpressions of ALDH1A3 and ALDH1B1 were found in GC compared with normal tissues (Figure 1). In Wang’s dataset, the ALDH1A3 mRNA level was significantly upregulated in GC, with 3.134 fold change (p = 9.93E-6) (Figure 2F) [21]. Consistently, in another dataset with 132 samples from patients with different types of gastric adenocarcinoma, the mRNA level of ALDH1A3 was over 2 up-fold change than in gastric mucosa (Figure 2G and 2H) [18]. However, only one dataset showed the significant fold change (2.610, p = 6.87E-4) of ALDH1B1 mRNA levels between cancer and normal tissue (Figure 2I).
The prognostic value of different ALDH1 isoenzymes in gastric cancer
As previously indicated [16, 17], among all the six ALDH1 isoenzymes, the recently reported ALDH1L2 isoform by Krupenko NI et al., was not found in www.kmplot.com, was not recruited in the screening pool [22].
Survival curves were plotted in www.kmplot.com for all GC patients (n = 876), gastric intestinal type adenocarcinoma (n = 320), diffuse gastric adenocarcinoma (n = 241), HER2-negative GC (n = 532), and HER2-positive GC (n = 344). As the sample size was too small in patients with gastric mixed adenocarcinoma (n = 32), we did not analyze the survival curves in this group. We assessed the prognostic effect of the mRNA expression of ALDH1A1 (the Affymetrix IDs is 212224_at), ALDH1A2 (the Affymetrix IDs is 207016_s_at), ALDH1A3 (the Affymetrix IDs is 203180_at), ALDH1B1 (the Affymetrix IDs is 209646_×_at) and ALDH1L1 (the Affymetrix IDs is 205208_at) in www.kmplot.com.
In Figure 3, increased mRNA levels of ALDH1A1 and ALDH1B1 expressions were associated with longer OS, i.e. better prognosis of GC patients, with HR = 0.77 (0.65–0.91), p = 0.0021 (Figure 3A) and HR = 0.58 (0.48–0.7), p = 0.0000 (Figure 3D), respectively. In contrast to other isoenzymes, high mRNA levels was found to be correlated to worsen OS in GC patients for ALDH1A2 (HR = 1.5 [1.22–1.85], p = 0.0001) (Figure 3B). Interestingly, high mRNA level of ALDH1A3 and ALDH1L1 were also significantly associated with worsen OS in all GC patients, with HR = 1.5 (1.26–1.79), p = 0.0000 (Figure 3C) and HR = 1.89 (1.59–2.32), p = 0.0000 (Figure 3E).
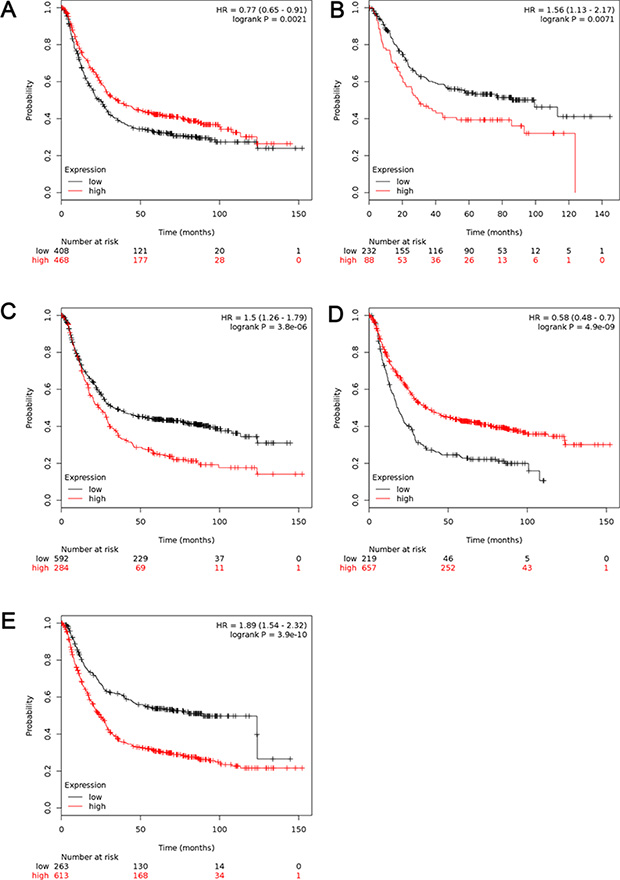
Figure 3: The prognostic value of mRNA level of ALDH1 isoenzymes in all GC patients (n = 876). (A) ALDH1A1 (212224_st) (B) ALDH1A2 (207016_s_at). (C) ALDH1A3 (203180_at). (D) ALDH1B1 (209646_×_at). (E) ALDH1L1 (205208_at). Data was analyzed using Kaplan-Meier plotter.
For further analysis, the patients have been sub-grouped by their types of cancer. High transcriptional expression of ALDH1A1 was associated with longer OS in gastric intestinal type adenocarcinoma, with HR = 0.6 (0.44–0.83), p = 0.0018 (Figure 4A), but not in diffuse gastric adenocarcinoma (Figure 4B). As another good predictor, high level of ALDH1B1 predicted better OS in gastric intestinal type adenocarcinoma, with HR = 0.57 (0.41–0.78), p = 0.0000 (Figure 4G) and worse OS in diffuse gastric adenocarcinoma, with HR = 1.92 (1.46–2.51), p = 0.0000 (Figure 4H). Interestingly, for ALDH1A2 (Figure 4C and 4D), ALDH1A3 (Figure 4E and 4F) and ALDH1L1 (Figure 4I and 4J), high mRNA level was found to be significantly correlated to worsen OS in both gastric intestinal type adenocarcinoma and diffuse gastric adenocarcinoma, with the same trend as in all GC patients.
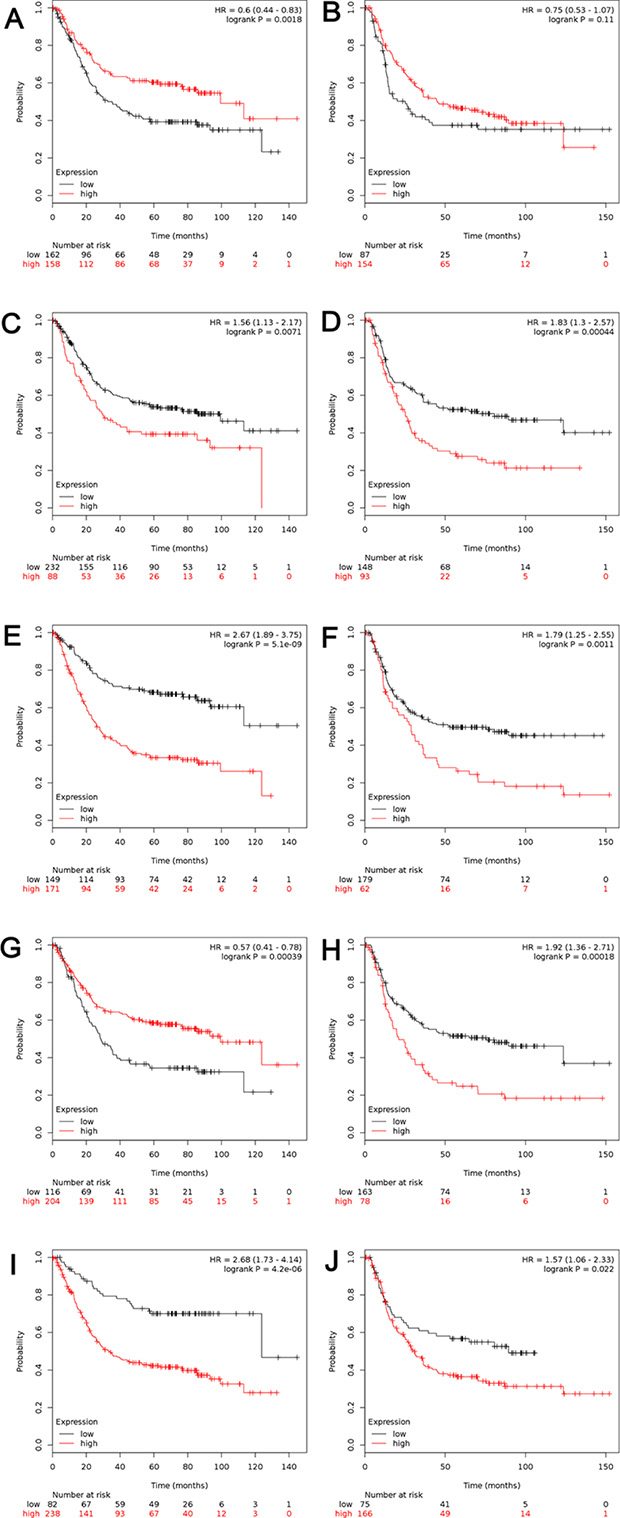
Figure 4: The prognostic value of mRNA level of ALDH1 isoenzymes in patients with gastric intestinal adenocarcinoma (n = 320) and diffuse gastric adenocarcinoma (n = 241). (A, B) ALDH1A1 (212224_st) (C, D) ALDH1A2 (207016_s_at). (E, F) ALDH1A3 (203180_at). (G, H) ALDH1B1 (209646_×_at). (I, J) ALDH1L1 (205208_at). (A, C, E, G, I) Survival curves in patients with gastric intestinal adenocarcinoma (n = 320). (B, D, F, H, J) Survival curves in patients with diffuse gastric adenocarcinoma (n = 241). Data was analyzed using Kaplan-Meier plotter.
HER2 status in GC was reported as a potential target for individual therapy [23]. So, the survival curves have been stratified by HER2 status to determine who might benefit from targeted therapy (Table 1). Although exhibiting the similar expression trend, upregulated ALDH1A1 was significantly associated with better OS in HER2-negative GC, with HR = 0.7 (0.56–0.88), p = 0.0018, not in HER2-positive GC (Supplementary Figure S1A and S1B). Increased ALDH1B1 was only significantly correlated with better OS in HER2-positive GC, with HR = 0.55 (0.44–0.69), p = 0.0000, but not in HER2-negative GC (Supplementary Figure S1G and S1H). Interestingly, except ALDH1A1 and ALDH1B1, other three isoenzymes all predicted worse OS with high expression in both HER2-negative and HER2-positive GC (Table 1).
Table 1: The Hazard ratio (HR) of ALDH1 isoenzymes in HER2-negative and HER2-positive GC patients
|
HER2-negative |
HER2-positive |
||||
|---|---|---|---|---|---|---|
|
HR |
95% CI |
P value |
HR |
95% CI |
P value |
ALDH1A1 |
0.7 |
0.56–0.88 |
0.0018 |
0.89 |
0.69–1.16 |
0.4 |
ALDH1A2 |
1.47 |
1.18–1.84 |
0.0007 |
1.5 |
1.09–2.07 |
0.013 |
ALDH1A3 |
1.68 |
1.34–2.11 |
0.0000 |
1.6 |
1.23–2.07 |
0.0004 |
ALDH1B1 |
0.8 |
0.64–1.01 |
0.064 |
0.58 |
0.44–0.76 |
0.0000 |
ALDH1L1 |
1.92 |
1.46–2.51 |
0.0000 |
1.42 |
1.1–1.85 |
0.0072 |
HR: Hazard ratio.
CI: Confidence Interval.
DISCUSSION
Gastric cancer (GC), mainly developing from the innermost lining of the stomach, presents the highest mortality rate among all digestive tract malignancies, mainly because of chemoradiotherapy resistance and distant metastasis [1]. It is crucial to illustrate the pathogenesis of GC, as well as to find novel prognostic strategies, early diagnostic tools and effective therapeutic approaches. On the other hand, strong evidences supported that CSC possess unique properties, such as uncontrolled growth and high tumorigenic and migration potential. CSCs are responsible for tumor initiation, progression, metastasis and chemoradiotherapy resistance. Among the identified CSC markers, ALDH1 is one found in a broad cancer spectrum. However, there are few reports available to analyze the expression of ALDH1 in GC, not to mention the different ALDH1 isoforms. In our study, we analyzed and discussed distinct roles of ALDH1 isoenzymes in prognostic value of GC.
ALDH1 mainly functions as a retinoic acid enzyme, catalyzing and conversing retinol to retinoic acid. As ALDH1A1 can inactivate integral chemotherapy agents, particularly cyclophosphamide, it is indicated that high ALDH1A1 expression in tumors may contribute to chemotherapy resistance of tumors [24]. Condello et al. recently reported that an ALDH1A1-specific inhibitor was used to block ovarian cancer cell proliferation and survival [25]. Recently, Li et al. reported that the overexpression of ALDH1A1 in protein level was correlated with poor OS and recurrence-free survival (RFS) in GC [26]. However, we found that the mRNA level of ALDH1A1 showed opposite prognostic value, viz. transcripts of ALDH1A1 were down-regulated in all types of GC and high mRNA expression of ALDH1A1 predicted better prognosis. Not surprisingly, Liu et al. concluded that high expression of ALDH1A1 transcripts may be an independent predictor of a favorable outcome in patients with triple-negative breast cancer, possibly due to the process of sample collection as suggested [27]. However, in our study, the datasets from different laboratories showed consistent results, indicating that the mRNA level of ALDH1A1 may be a good prognostic marker in GC, which could be opposite to their protein levels.
ALDH1B1, a mitochondrial ALDH, is another potential good prognostic marker in GC, sharing 65% homology in peptide sequence with cytosolic ALDH1A1 in human. ALH1B1 catalytically metabolizes a wide range of aldehyde substrates, including acetaldehyde and products of lipid peroxidation (LPO), and is activated in ethanol metabolism [28]. In colon cancer, ALDH1B1 is dramatically upregulated [29], and by up-regulating Wnt/β-Catenin, Notch and PI3K/Akt signaling pathways, ALDH1B1 can promote colon cancer tumorigenesis, providing a novel target to prevent or treat colon cancer [30]. The mRNA level of ALDH1B1 was higher in gastric mixed adenocarcinoma than in gastric tissues in Cho’s research dataset [18]. However, in Kaplan-Meier plotter analysis, the ALDH1B1 in this study was found to be a good prognostic marker for OS in patients with all types of GC and intestinal type gastric adenocarcinoma, a poor prognostic marker for OS in patients with diffuse gastric adenocarcinoma. With the preliminary results, more research would be helpful to explore the mechanism of ALDH1B1 in GC.
ALDH1A2 was reported as a candidate tumor suppressor in prostate cancer, downregulated on early stage of human prostate cancer [31, 32]. Controversially, an in vitro study showed that K562 leukemia and H1299 lung cancer cell with ALDH1A2 overexpression exhibited higher cell proliferation rates, higher clonal efficiency, and increased drug resistance to 4-hydroperoxycyclophosphamide and doxorubicin [33]. In this study, we first reported that high ALDH1A2 transcription might predict poorer survival in GC patients, in which may involve the synthesis of retinoic acid pathway.
ALDH1A3 may promote progress and metastasis of various cancers, such as prostate cancer [34], gallbladder cancer [35], breast cancer [36], ovarian cancer [37] and NSCLC [38], as well as gastric cancer in our study, and be identified as the biomarkers for poor prognosis. ALDH1A3 responded to androgen dihydrotestosterone treatment and increased the oxidation of retinal to RA in human prostate cancer cells [34], and was supposed to be the predominant isoenzyme responsible for ALDH activity and tumorigenicity in most NSCLC [38]. As low expression of ALDH1A3 may be due to dysregulation of hypermethylation status of ALDH1A3 promoter [39] or suppression of miR-125a/b [40], the mechanism of ALDH1A3 in GC may involve differential retinoic acid signaling and hypermethylation of ALDH1A3 promoter, providing new therapy approaches.
ALDH1L1 (FDH, a folate metabolic enzyme with tumor suppressor-like properties) is cell-specifically expressed and involved in central nervous system development and reduced proliferation [41]. Interestingly, ALDH1L1 was reduced in clinically aggressive compared with Pilocytic [42]. ALDH1L1 could inhibit cancer cell motility via dephosphorylation of cofilin by PP1 and PP2A in folate-specific manner [43]. In hepatocellular carcinoma, decreased ALDH1L1 is associated with a poor prognosis [44]. Not surprisingly, in our results, the ALDH1L1 was indicated as a potential poor prognostic biomarker for GC, and that the promoter methylation may be a major mechanism controlling ALDH1L1 levels in human cancers [45].
In ONCOMINE datasets, ALDH1A3 expressed lower in GC than in normal tissues. Although no researches illustrated significant correlations of ALDH1A2 or ALDH1L1 with GC in ONCOMINE datasets, high transcriptions of ALDH1A2, ALDH1A3 and ALDH1L1 isoenzymes were found to be correlated with shorter OS in all GC, no matter the type of GC or HER2 status in the patients. GC affect more male than female, so we analyzed the survival in different gender (Supplementary Figure S2) and found that these results were independent of gender, supporting that ALDH1A2, ALDH1A3 and ALDH1L1 were the main contributors for ALDH1 activities in GC and potential prognostic markers and treated targets for GC. The results provide better understanding of the heterogeneity and complexity in the molecular biology of GC, which is the basement of more accurate prognosis and development of new treatment targets.
MATERIALS AND METHODS
ONCOMINE analysis
The mRNA levels of different ALDH1 isoforms in different cancers were analyzed by ONCOMINE gene expression array datasets (www.oncomine.org), which is an online cancer microarray database to facilitate discovery from genome-wide expression analyses [46]. In this study, we compared the clinical specimens of cancer vs. normal control datasets, using a Students’ t-test to generate a p-value. The p value was set up at 0.01 and fold change was defined as 2, whereas the data type was restricted to mRNA. Significant correlations can be found in different researches of GC, showed in typical figures.
The kaplan-meier plotter
The prognostic value of the mRNA expression of ALDH1 isoforms in GC was assessed using the Kaplan-Meier plotter (www.kmplot.com), which is also an online database, including gene expression data and clinical data [47]. Up to now, Kaplan-Meier plotter is capable to assess the effect of 54, 675/22, 277 genes on survival of 10, 188 clinical cancer samples, including 4, 142 breast, 1, 648 ovarian, 2, 437 lung and 1, 065 gastric cancer patients. The patient samples were divided into two groups, according to the expression of ALDH1 isoforms (high vs. low expression). We analyzed the overall survival of the patients with GC using a Kaplan-Meier survival plot. The JetSet best probe set of 5 ALDH1 sub-members (ALDH1A1, ALDH1A2, ALDH1A3, ALDH1B1, and ALDH1L1) were entered into the database (http://kmplot.com/analysis/index.php? p = service and cancer = gastric) obtain Kaplan-Meier plots in which the number-at-risk is indicated below the main plot. The hazard ratio (HR) with 95% confidence intervals and log rank p value was calculated and displayed on the webpage. HER2 status was determined using the gene chip probe set 216836_s_at as described before [48].
ACKNOWLEDGMENTS AND FUNDING
This work was supported by National Natural Science Foundation of China (No. 81501539), Natural Science Foundation of Guangdong Province, China (No. 2015A030310211), and Youth Innovative Talent Project of Universities in Guangdong (No. 2014KQNCX078) and Personnel Training Project from the Education Department of Guangdong Province (Jing Liu), and Technology Planning Project of Shantou, China ([2014] No. 62 of Shantou Technology). Special appreciation would be given to Mr. Kwan Chun Kit for manuscript editing.
CONFLICTS OF INTEREST
Disclose any potential conflicts of interest.
REFERENCES
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108.
2. Park JY, von Karsa L, Herrero R. Prevention strategies for gastric cancer: a global perspective. Clinical endoscopy. 2014; 47:478–489.
3. Brungs D, Aghmesheh M, Vine KL, Becker TM, Carolan MG, Ranson M. Gastric cancer stem cells: evidence, potential markers, and clinical implications. J Gastroenterol. 2015. [Epub ahead of print].
4. Vermeulen L, de Sousa e Melo F, Richel DJ, Medema JP. The developing cancer stem-cell model: clinical challenges and opportunities. Lancet Oncol. 2012; 13:e83–89.
5. Reya T, Morrison SJ, Clarke MF, Weissman IL. Stem cells, cancer, and cancer stem cells. Nature. 2001; 414:105–111.
6. Pang R, Law WL, Chu AC, Poon JT, Lam CS, Chow AK, Ng L, Cheung LW, Lan XR, Lan HY, Tan VP, Yau TC, Poon RT, et al. A subpopulation of CD26+ cancer stem cells with metastatic capacity in human colorectal cancer. Cell Stem Cell. 2010; 6:603–615.
7. Takaishi S, Okumura T, Tu S, Wang SS, Shibata W, Vigneshwaran R, Gordon SA, Shimada Y, Wang TC. Identification of gastric cancer stem cells using the cell surface marker CD44. Stem Cells. 2009; 27:1006–1020.
8. Jiang F, Qiu Q, Khanna A, Todd NW, Deepak J, Xing L, Wang H, Liu Z, Su Y, Stass SA, Katz RL. Aldehyde dehydrogenase 1 is a tumor stem cell-associated marker in lung cancer. Mol Cancer Res. 2009; 7:330–338.
9. Ricci-Vitiani L, Lombardi DG, Pilozzi E, Biffoni M, Todaro M, Peschle C, De Maria R. Identification and expansion of human colon-cancer-initiating cells. Nature. 2007; 445:111–115.
10. Wu C, Xie Y, Gao F, Wang Y, Guo Y, Tian H, Li Y, Fan W. Lgr5 expression as stem cell marker in human gastric gland and its relatedness with other putative cancer stem cell markers. Gene. 2013; 525:18–25.
11. Moreb J, Schweder M, Suresh A, Zucali JR. Overexpression of the human aldehyde dehydrogenase class I results in increased resistance to 4-hydroperoxycyclophosphamide. Cancer Gene Ther. 1996; 3:24–30.
12. Nishikawa S, Konno M, Hamabe A, Hasegawa S, Kano Y, Ohta K, Fukusumi T, Sakai D, Kudo T, Haraguchi N, Satoh T, Takiguchi S, Mori M, et al. Aldehyde dehydrogenase high gastric cancer stem cells are resistant to chemotherapy. Int J Oncol. 2013; 42:1437–1442.
13. Wakamatsu Y, Sakamoto N, Oo HZ, Naito Y, Uraoka N, Anami K, Sentani K, Oue N, Yasui W. Expression of cancer stem cell markers ALDH1, CD44 and CD133 in primary tumor and lymph node metastasis of gastric cancer. Pathol Int. 2012; 62:112–119.
14. Katsuno Y, Ehata S, Yashiro M, Yanagihara K, Hirakawa K, Miyazono K. Coordinated expression of REG4 and aldehyde dehydrogenase 1 regulating tumourigenic capacity of diffuse-type gastric carcinoma-initiating cells is inhibited by TGF-beta. J Pathol. 2012; 228:391–404.
15. Levi E, Sochacki P, Khoury N, Patel BB, Majumdar AP. Cancer stem cells in Helicobacter pylori infection and aging: Implications for gastric carcinogenesis. World journal of gastrointestinal pathophysiology. 2014; 5:366–372.
16. Wu S, Xue W, Huang X, Yu X, Luo M, Huang Y, Liu Y, Bi Z, Qiu X, Bai S. Distinct prognostic values of ALDH1 isoenzymes in breast cancer. Tumour Biol. 2015; 36:2421–2426.
17. You Q, Guo H, Xu D. Distinct prognostic values and potential drug targets of ALDH1 isoenzymes in non-small-cell lung cancer. Drug design, development and therapy. 2015; 9:5087–5097.
18. Cho JY, Lim JY, Cheong JH, Park YY, Yoon SL, Kim SM, Kim SB, Kim H, Hong SW, Park YN, Noh SH, Park ES, Chu IS, et al. Gene expression signature-based prognostic risk score in gastric cancer. Clin Cancer Res. 2011; 17:1850–1857.
19. Chen X, Leung SY, Yuen ST, Chu KM, Ji J, Li R, Chan AS, Law S, Troyanskaya OG, Wong J, So S, Botstein D, Brown PO. Variation in gene expression patterns in human gastric cancers. Mol Biol Cell. 2003; 14:3208–3215.
20. D’Errico M, de Rinaldis E, Blasi MF, Viti V, Falchetti M, Calcagnile A, Sera F, Saieva C, Ottini L, Palli D, Palombo F, Giuliani A, Dogliotti E. Genome-wide expression profile of sporadic gastric cancers with microsatellite instability. Eur J Cancer. 2009; 45:461–469.
21. Wang Q, Wen YG, Li DP, Xia J, Zhou CZ, Yan DW, Tang HM, Peng ZH. Upregulated INHBA expression is associated with poor survival in gastric cancer. Med Oncol. 2012; 29:77–83.
22. Krupenko NI, Dubard ME, Strickland KC, Moxley KM, Oleinik NV, Krupenko SA. ALDH1L2 is the mitochondrial homolog of 10-formyltetrahydrofolate dehydrogenase. J Biol Chem. 2010; 285:23056–23063.
23. Yi JH, Kang JH, Hwang IG, Ahn HK, Baek HJ, Lee SI, Lim DH, Won YW, Ji JH, Kim HS, Rha SY, Oh SY, Lee KE, et al. A Retrospective Analysis for Patients with HER2 Positive Gastric Cancer who were Treated with Trastuzumab-based Chemotherapy: In the Perspectives of Ethnicity and Histology. Cancer Res Treat. 2015. [Epub ahead of print].
24. Moreb JS, Schweder M, Gray B, Zucali J, Zori R. In vitro selection for K562 cells with higher retrovirally mediated copy number of aldehyde dehydrogenase class-1 and higher resistance to 4-hydroperoxycyclophosphamide. Hum Gene Ther. 1998; 9:611–619.
25. Condello S, Morgan CA, Nagdas S, Cao L, Turek J, Hurley TD, Matei D. beta-Catenin-regulated ALDH1A1 is a target in ovarian cancer spheroids. Oncogene. 2015; 34:2297–2308.
26. Li XS, Xu Q, Fu XY, Luo WS. ALDH1A1 overexpression is associated with the progression and prognosis in gastric cancer. BMC Cancer. 2014; 14:705.
27. Liu Y, Baglia M, Zheng Y, Blot W, Bao PP, Cai H, Nechuta S, Zheng W, Cai Q, Shu XO. ALDH1A1 mRNA expression in association with prognosis of triple-negative breast cancer. Oncotarget. 2015; 6:41360–41369. doi: 10.18632/oncotarget.6023.
28. Stagos D, Chen Y, Brocker C, Donald E, Jackson BC, Orlicky DJ, Thompson DC, Vasiliou V. Aldehyde dehydrogenase 1B1: molecular cloning and characterization of a novel mitochondrial acetaldehyde-metabolizing enzyme. Drug Metab Dispos. 2010; 38:1679–1687.
29. Chen Y, Orlicky DJ, Matsumoto A, Singh S, Thompson DC, Vasiliou V. Aldehyde dehydrogenase 1B1 (ALDH1B1) is a potential biomarker for human colon cancer. Biochem Biophys Res Commun. 2011; 405:173–179.
30. Singh S, Arcaroli J, Chen Y, Thompson DC, Messersmith W, Jimeno A, Vasiliou V. ALDH1B1 Is Crucial for Colon Tumorigenesis by Modulating Wnt/beta-Catenin, Notch and PI3K/Akt Signaling Pathways. PLoS One. 2015; 10:e0121648.
31. Kim H, Lapointe J, Kaygusuz G, Ong DE, Li C, van de Rijn M, Brooks JD, Pollack JR. The retinoic acid synthesis gene ALDH1a2 is a candidate tumor suppressor in prostate cancer. Cancer Res. 2005; 65:8118–8124.
32. Touma SE, Perner S, Rubin MA, Nanus DM, Gudas LJ. Retinoid metabolism and ALDH1A2 (RALDH2) expression are altered in the transgenic adenocarcinoma mouse prostate model. Biochem Pharmacol. 2009; 78:1127–1138.
33. Moreb JS, Ucar D, Han S, Amory JK, Goldstein AS, Ostmark B, Chang LJ. The enzymatic activity of human aldehyde dehydrogenases 1A2 and 2 (ALDH1A2 and ALDH2) is detected by Aldefluor, inhibited by diethylaminobenzaldehyde and has significant effects on cell proliferation and drug resistance. Chem Biol Interact. 2012; 195:52–60.
34. Trasino SE, Harrison EH, Wang TT. Androgen regulation of aldehyde dehydrogenase 1A3 (ALDH1A3) in the androgen-responsive human prostate cancer cell line LNCaP. Exp Biol Med (Maywood). 2007; 232:762–771.
35. Yang ZL, Yang L, Zou Q, Yuan Y, Li J, Liang L, Zeng G, Chen S. Positive ALDH1A3 and negative GPX3 expressions are biomarkers for poor prognosis of gallbladder cancer. Dis Markers. 2013; 35:163–172.
36. Marcato P, Dean CA, Pan D, Araslanova R, Gillis M, Joshi M, Helyer L, Pan L, Leidal A, Gujar S, Giacomantonio CA, Lee PW. Aldehyde dehydrogenase activity of breast cancer stem cells is primarily due to isoform ALDH1A3 and its expression is predictive of metastasis. Stem Cells. 2011; 29:32–45.
37. Saw YT, Yang J, Ng SK, Liu S, Singh S, Singh M, Welch WR, Tsuda H, Fong WP, Thompson D, Vasiliou V, Berkowitz RS, Ng SW. Characterization of aldehyde dehydrogenase isozymes in ovarian cancer tissues and sphere cultures. BMC Cancer. 2012; 12:329.
38. Shao C, Sullivan JP, Girard L, Augustyn A, Yenerall P, Rodriguez-Canales J, Liu H, Behrens C, Shay JW, Wistuba, II, Minna JD. Essential role of aldehyde dehydrogenase 1A3 for the maintenance of non-small cell lung cancer stem cells is associated with the STAT3 pathway. Clin Cancer Res. 2014; 20:4154–4166.
39. Zhang W, Yan W, You G, Bao Z, Wang Y, Liu Y, You Y, Jiang T. Genome-wide DNA methylation profiling identifies ALDH1A3 promoter methylation as a prognostic predictor in G-CIMP- primary glioblastoma. Cancer Lett. 2013; 328:120–125.
40. Chen J, Chen Y, Chen Z. MiR-125a/b regulates the activation of cancer stem cells in paclitaxel-resistant colon cancer. Cancer Invest. 2013; 31:17–23.
41. Anthony TE, Heintz N. The folate metabolic enzyme ALDH1L1 is restricted to the midline of the early CNS, suggesting a role in human neural tube defects. The Journal of comparative neurology. 2007; 500:368–383.
42. Rodriguez FJ, Giannini C, Asmann YW, Sharma MK, Perry A, Tibbetts KM, Jenkins RB, Scheithauer BW, Anant S, Jenkins S, Eberhart CG, Sarkaria JN, Gutmann DH. Gene expression profiling of NF-1-associated and sporadic pilocytic astrocytoma identifies aldehyde dehydrogenase 1 family member L1 (ALDH1L1) as an underexpressed candidate biomarker in aggressive subtypes. Journal of neuropathology and experimental neurology. 2008; 67:1194–1204.
43. Oleinik NV, Krupenko NI, Krupenko SA. ALDH1L1 inhibits cell motility via dephosphorylation of cofilin by PP1 and PP2A. Oncogene. 2010; 29:6233–6244.
44. Chen XQ, He JR, Wang HY. Decreased expression of ALDH1L1 is associated with a poor prognosis in hepatocellular carcinoma. Med Oncol. 2012; 29:1843–1849.
45. Oleinik NV, Krupenko NI, Krupenko SA. Epigenetic Silencing of ALDH1L1, a Metabolic Regulator of Cellular Proliferation, in Cancers. Genes Cancer. 2011; 2:130–139.
46. Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, Barrette T, Pandey A, Chinnaiyan AM. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004; 6:1–6.
47. Gyorffy B, Surowiak P, Budczies J, Lanczky A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS One. 2013; 8:e82241.
48. Gyorffy B, Benke Z, Lanczky A, Balazs B, Szallasi Z, Timar J, Schafer R. RecurrenceOnline: an online analysis tool to determine breast cancer recurrence and hormone receptor status using microarray data. Breast Cancer Res Treat. 2012; 132:1025–1034.

