INTRODUCTION
Nasopharyngeal carcinoma (NPC) is a unique head and neck cancer which is highly prevalent in Southern China and throughout Southeast Asia [1], with an incidence of 25–30 cases per 100,000 persons annually [2].NPC pathogenesis is a multistep process driven by an accumulation of genetic alterations, including the loss of tumor suppressor genes and activation of oncogenes [3]. Malignant cell proliferation and apoptosis inhibition are the main factors influencing the development and progression of NPC and lead to the poor overall survival of NPC patients [4, 5]. It is essential to elucidate the molecular mechanisms underlying tumorigenesis and invasiveness in NPC in order to identify novel therapeutic targets and develop new modalities of treatment.
It has been reported that ARHGEF3, a Rho-guanine nucleotide exchange factor (GEF) upregulated in acute myeloid leukemia (AML), modulates AML differentiation through activation of RhoA and pathways directly controlled by small GTPase family proteins [6]. The human gene ARHGEF3 is located at chromosome 3p13-21 and encodes a polypeptide of 526 amino acids with homology to neuroepithelial transforming gene 1 (NET1) [7–9]. ARHGEF3 belongs to the family of Rho-GEFs which specifically activates two members of the Rho-GTPase family, RHOA and RHOB, and accelerates Rho-GTPase activity by conversion of GTP to GDP [10, 11]. Mutations in some members of the GEF family, such as DOCK2, DOCK8 and ARHGEF6, are associated with invasiveness and metastasis of human malignancies [12–15]. However, the potential roles and biological mechanisms of the GEF family gene ARHGEF3 in human cancers have not been studied. To investigate if abnormalities in ARHGEF3 are involved in NPC pathogenesis, we examined ARHGEF3 protein levels in a series of carcinomatous and non-neoplastic human nasopharyngeal cells and tissues, assessed the clinicopathologic/prognostic significance of ARHGEF3 expression in our NPC cohort, and investigated the mechanisms underlying the oncogenic and tumorigenic role of ARHGEF3 in NPC.
We found that high expression of ARHGEF3 in NPCs is important in the acquisition of an aggressive phenotype. Silencing ARHGEF3 in NPC cells was sufficient to inhibit cell growth, migration, and invasion in vitro, while overexpression of ARHGEF3 supported the tumorigenic and metastatic capacities of NPC cells in vivo. Further, we demonstrated that depletion of ARHGEF3 in NPC cells promoted caspase3-induced apoptosis. We also identified the anti-apoptosis factor BIRC8 as a critical downstream target of ARHGEF3. Collectively, our results provide an explanation for the malignant nature of NPC involving ARHGEF3 overexpression and the underlying mechanism that links ARHGEF3 to BIRC8 in NPC cell apoptosis.
RESULTS
Analysis of ARHGEF3 protein levels in NPC cells and nasopharyngeal tissues
We analyzed endogenous ARHGEF3 protein levels in 8 human nasopharyngeal cell lines by Western blotting and found that ARHGEF3 was overexpressed in 5 NPC cell lines (CNE2, SUNE1, 5-8F, 6-10B and C666), while the other 2 NPC lines (CNE1 and HONE1) and the immortalized normal nasopharyngeal cell line NP69 exhibited low ARHGEF3 protein levels (Figure 1A, left). At the same time, we found that ARHGEF3 protein expression was higher in 8 primary NPC tissues, compared with adjacent non-neoplastic nasopharyngeal tissues. But there were no difference between the tumor and adjacent tissues in 1 case. (Figure 1A, right).
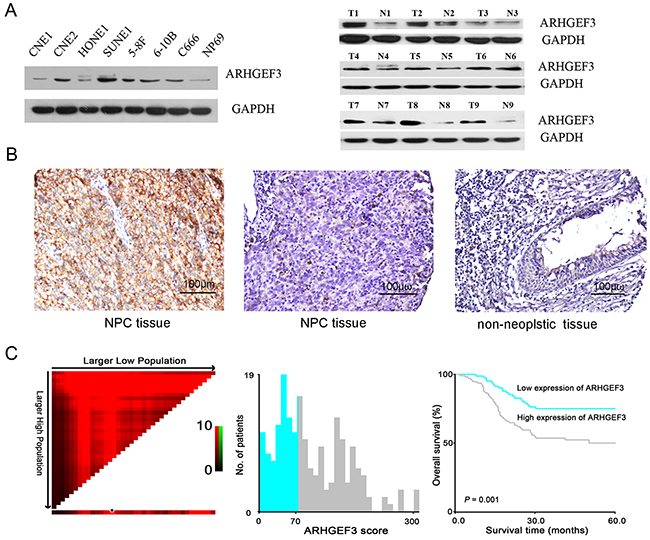
Figure 1: Expression of ARHGEF3 in nasopharyngeal cell lines and tissues and its prognostic significance in nasopharyngeal carcinoma (NPC) patients. A. Western blot showing relative levels of ARHGEF3 protein in 8 nasopharyngeal cell lines (left). ARHGEF3 expression was up-regulated in primary NPC tissues compared with paired non-neoplastic nasopharyngeal mucosa tissues (right). B. Representative immunohistochemistry images showing high expression of ARHGEF3 in one NPC tissue (case 27, left), low expression of ARHGEF3 in another NPC tissue (case 99, middle), and negative expression of ARHGEF3 in a non-neoplastic nasopharyngeal tissue (case 33, right). C. X-tile plots of the prognostic marker ARHGEF3. X-tile analysis was carried out on patient data from the NPC cohort. The plot shows the χ2 log-rank values. Panels depict the cut-off point for high expression (highlighted by the black/white circle; left), a histogram of the entire cohort (middle), and a Kaplan–Meier survival curve (right).
IHC staining of ARHGEF3 expression in NPC tissues and its correlation with NPC patients’ pathological features and survival
Using IHC staining, we observed high expression of ARHGEF3 (Figure 1B, left) in 111 of 192 (57.8%) primary NPC tissues (Table 1). 17 cases of NPC were not informative due to unrepresentative samples or lost samples. We used the whole NPC tissue slides of these cases to improve the limitation of TMA technology in our study. Correlation analysis demonstrated that high expression of ARHGEF3 was positively associated with an increased T status, distant metastasis, and/or a more advanced clinical stage of NPCs (P< 0.05, Table 1). Kaplan-Meier survival curves showed that the mean disease-free survival time in NPC patients with high expression of ARHGEF3 was significantly shorter than in patients with low expression of ARHGEF3 (P=0.001, long-rank test, Figure 1C, Table 2). Multivariate Cox proportional hazards regression analysis demonstrated that high expression of ARHGEF3 was a significant and independent prognostic factor for poor survival of NPC patients (relative risk: 1.709, confidence interval: 1.002-2.913, P=0.049, Table 2).
Table 1: Correlation between the clinicopathological features and expression of ARHGEF3 in NPCs
cases |
ARHGEF3 protein |
|||
|---|---|---|---|---|
Low expression |
High expression |
P value* |
||
Sex |
0.532 |
|||
Female |
57 |
26 (45.6%) |
31 (54.4%) |
|
Male |
135 |
55 (40.7%) |
80 (59.3%) |
|
Age at diagnosis (years) |
0.299 |
|||
≤ 47† |
103 |
47 (45.6%) |
56 (54.4%) |
|
> 47 |
89 |
34 (38.2%) |
55 (61.8%) |
|
Histological classification (WHO) |
0.303 |
|||
Type II |
50 |
18 (36.0%) |
32 (64.0%) |
|
Type III |
142 |
63 (44.4%) |
79 (55.6%) |
|
T classification |
0.044 |
|||
1 |
23 |
10 (43.5%) |
13 (56.5%) |
|
2 |
65 |
31 (47.7%) |
34 (52.3%) |
|
3 |
67 |
32 (47.8%) |
35 (52.2%) |
|
4 |
37 |
8 (21.6%) |
29 (78.4%) |
|
N classification |
0.203 |
|||
0 |
38 |
15 (39.5%) |
23 (60.5%) |
|
1 |
89 |
43 (48.3%) |
46 (51.7%) |
|
2 |
50 |
20 (40.0%) |
30 (60.0%) |
|
3 |
15 |
3 (20.0%) |
12 (80.0%) |
|
Distant metastasis |
0.005 |
|||
0 |
152 |
72 (47.4%) |
80 (52.6%) |
|
1 |
40 |
9 (22.5%) |
31 (77.5%) |
|
Clinical stage |
0.002 |
|||
I |
9 |
3 (33.3%) |
6 (66.7%) |
|
II |
50 |
26 (52.0%) |
24 (48.0%) |
|
III |
83 |
42 (50.6%) |
41 (49.4%) |
|
IV |
50 |
10 (20.0%) |
40 (80.0%) |
|
*Chi-square test; †median age.
Abbreviation: ARHGEF3, Rho guanine nucleotide exchange factor3; NPC, nasopharyngeal carcinoma; T, tumor; N, node.
Table 2: Univariate and multivariate analysis of different prognostic parameters in 192 patients with NPC
Variable |
Univariate analysis |
Multivariate analysis |
|||
|---|---|---|---|---|---|
All cases |
HR (95% CI) |
P value |
HR (95% CI) |
P value |
|
Sex |
0.868 |
||||
Female |
57 |
1.0 |
|||
Male |
135 |
0.959 (0.581-1.581) |
|||
Age at surgery (years) |
0.511 |
||||
≤ 47* |
103 |
1.0 |
|||
> 47 |
89 |
0.856 (0.539-1.360) |
|||
Histological classification (WHO) |
0.126 |
||||
Type II |
50 |
1.0 |
|||
Type III |
142 |
1.577 (0.881-2.824) |
|||
T classification |
<0.0001 |
0.744 |
|||
T1-T2 |
88 |
1.0 |
1.0 |
||
T3-T4 |
104 |
2.465 (1.484-4.094) |
1.106 (0.605-2.020) |
||
N classification |
<0.0001 |
0.050 |
|||
N0-N1 |
127 |
1.0 |
1.0 |
||
N2-N3 |
65 |
2.829 (1.781-4.492) |
1.668 (1.001-2.779) |
||
Distant metastasis |
<0.0001 |
< 0.0001 |
|||
0 |
152 |
1.0 |
1.0 |
||
1 |
40 |
4.462 (2.791-7.135) |
2.602 (1.575-4.298) |
||
Clinical stage |
<0.0001 |
0.034 |
|||
I-II |
59 |
1.0 |
1.0 |
||
III -IV |
133 |
6.291 (2.727-14.515) |
3.103 (1.087-8.859) |
||
ARHGEF3 expression |
0.001 |
0.049 |
|||
Low |
81 |
1.0 |
1.0 |
||
high |
111 |
2.334 (1.395-3.905) |
1.709 (1.002-2.913) |
||
*median age; HR indicates hazards ratio; CI indicates confidence interval.
Abbreviation: NPC, nasopharyngeal carcinoma; ARHGEF3, Rho guanine nucleotide exchange factor3; T, tumor; N, node.
Knockdown of ARHGEF3 suppresses NPC cell growth, migration, and invasion in vitro
The above observations prompted us to explore the biological function of ARHGEF3 in NPC tumorigenesis and progression. The capacity for colony formation was evaluated in two NPC cell lines (CNE2 and SUNE1) that were transfected with siARHGEF3 or control siNC. The efficiency of ARHGEF3 knockdown by siARHGEF3 was examined by Western blotting (Figure 2A). Both ARHGEF3-silenced CNE2 and SUNE1 cells had fewer and smaller colonies than that siNC-transfected cells (Figure 2B), indicating that depletion of ARHGEF3 inhibits growth in NPC cells. Next, the effect of ARHGEF3 levels on NPC cell migration and invasion capacities were characterized by the wound-healing and Matrigel invasion assays, respectively. Knockdown of ARHGEF3 in both CNE2 and SUNE1 cells caused a dramatic suppression of cell migration and invasion abilities as compared to control cells (Figure 2C and 2D).
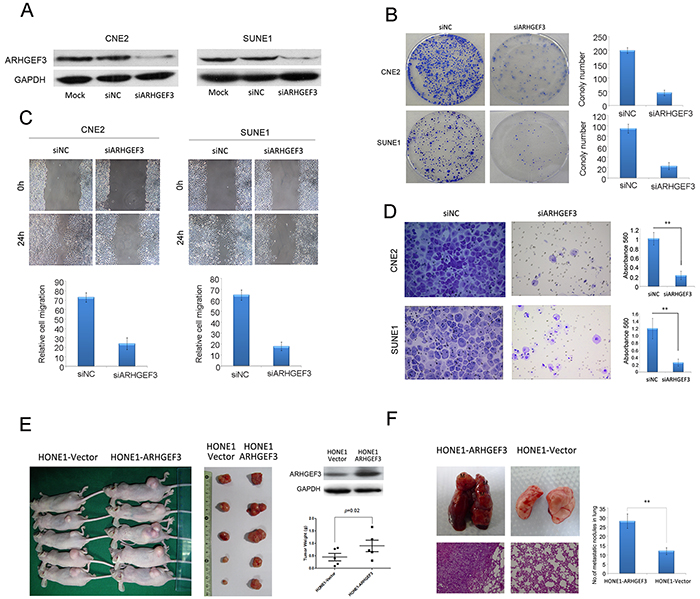
Figure 2: Effect of ARHGEF3 on NPC cells colony formation, cell motility, and invasion in vitro and tumorigenesis and metastasis in vivo. A. Western blot confirming reveals that ARHGEF3 was efficiently knocked down by the treatment of a specific siRNA in CNE2 and SUNE1 NPC cells. B. Representative images of decreased colony formation in monolayer culture induced by knockdown of ARHGEF3 in NPC cells. C. The wound-healing assay shows that knockdown of ARHGEF3 substantially inhibited the migration of CNE2 and SUNE1 cells. D. Transwell invasion assays show that ARHGEF3-silenced CNE2 and SUNE1 cells had lower invasive capacity compared to control cells. Data are the mean ± SD of at least 3 independent experiments. **P< 0.01 by Student’s t test. E. Western blots of ARHGEF3 protein levels in HONE1-ARHGEF3 and HONE1-vector cells (right, upper). Images of the xenograft tumors formed in nude mice injected with HONE1-ARHGEF3 or HONE1-vectorcells (left and right, lower). Weights of xenograft tumors are given as mean ± SD. **, P=0.02 by Student’s t test. F. Representative image of lungs showing metastatic nodules originating from HONE1-ARHGEF3 or control HONE1-vector cells injected into BALB/C-nu athymic nude mice. H&E staining of lung metastatic tumors are shown (left). Quantification of metastatic nodules formed in the lungs ofmice 8 weeks after tail vein injection of HONE1-ARHGEF3 or HONE1-vector cells (n = 5 mice per group; P<0.001, independent Student’s t test, right).
Upregulated expression of ARHGEF3 supports the tumorigenic and metastatic capacities of NPC cells in vivo
To investigate whether levels of ARHGEF3 influence the tumorigenic function of NPC cells in vivo, we first constructed a HONE1-ARHGEF3 cell line which stably overexpressed ARHGEF3 (Figure 2E, right, upper) . Next, HONE1-ARHGEF3 cells were transplanted into the backs of BALB/C-nu athymic nude mice, while HONE1-vector cells were used as a negative control (n = 5 mice per group). Thirty days after cell injection mice were sacrificed and the size and the weight of the subcutaneous tumors were examined. Tumors developed from HONE1-ARHGEF3 cells were significantly larger and heavier (P=0.02) than those arising from control cells (Figure 2E). To investigate if increased expression of ARHGEF3 in NPC cells is causative in an in vivo experimental metastasis model, we injected HONE1-ARHGEF3 or control HONE1-vector cells into the tail vein of BALB/C-nu athymic nude mice (n = 5 mice per group). Eight weeks after injection, mice were killed and metastatic tumor nodules formed in lung and liver were examined. We did not detect tumor nodule formation in the livers of all mice examined, but overexpression of ARHGEF3 significantly increased metastasis in lung (P<0.01, Figure 2F).
Expression level of ARHGEF3 influences the apoptosis of NPC cells in vitro
To further study the effect of ARHGEF3 on NPC cell apoptosis, we transfected CNE2 and SUNE1 cells with either siARHGEF3 or siBIRC8 for 48h and found that knockdown of ARHGEF3 and BIRC8 increased apoptosis in both CNE2 and SUNE1 cells compared with control cells. Then in rescue experiment, three days after the transfection of siARHGEF3, cells were transfected with pcDNA3.1(+)-BIRC8. Finally, the apoptotic assay was performed with flow cytometry. Ectopic expression of BIRC8 in CNE2 and SUNE1 cells with the knockdown of ARHGEF3 reversed the pro-apoptotic function of siARHGEF3 (Figure 3A). We also determined that the levels of active, cleaved caspase-3 were substantially increased in siARHGEF3-CNE2 and siARHGEF3-SUNE1 cells when compared to matched control cells (Figure 3B). These data suggest that the attenuation of ARHGEF3 expression promotes NPC cell apoptosis.
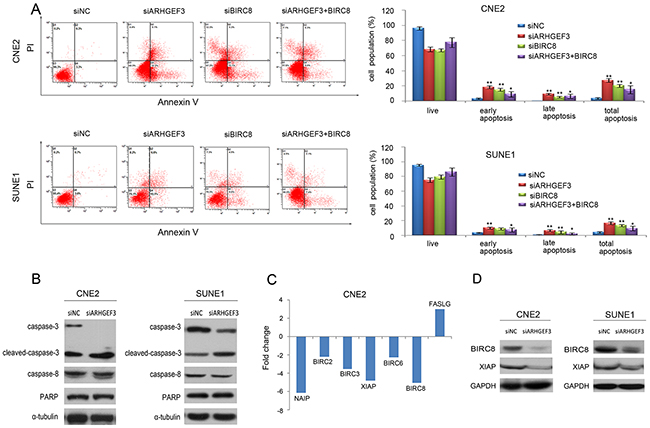
Figure 3: Knockdown of ARHGEF3 promotes cellular apoptosis and regulates apoptosis-associated gene expression in NPC cells. A. A representative image showing that knockdown of ARHGEF3 and BIRC8 in CNE2 and SUNE1 cells significantly increases cell apoptosis compared to control cells (A). At first, CNE2 and SUNE1 cells were transfected with siARHGEF3 or siBIRC8 alone. Then in rescue experiment, three days after the transfection of siARHGEF3, cells were transfected with pcDNA3.1(+)-BIRC8. Finally, the apoptotic assay was performed with flow cytometry. The knockdown of ARHGEF3 and BIRC8 in CNE2 and SUNE1 cells significantly increases cell apoptosis
ARHGEF3 regulates apoptosis-related gene expressions in NPC cells
In an effort to determine the potential downstream targets of ARHGEF3 that are involved in the promotion of NPC cell apoptosis, we compared mRNA expression profiles of siARHGEF3-CNE2 cells with those of control siNC-CNE2 cells using a Human Tumor Apoptosis RT2 Profiler™ PCR Array containing 84 apoptosis-related genes. We identified a total of 6 downregulated genes (NAIP, BIRC2, BIRC3, XIAP, BIRC6, and BIRC8) and 1 upregulated gene (FASLG) in siARHGEF3-transfected CNE2 cells, which showed more than a twofold change in mRNA levels compared to control siNC-CNE2 cells (Figure 3C and Table 3). Downregulation of XIAP and BIRC8 (ILP-2) was further validated by Western blotting assay in CNE2 and SUNE1 cells after ARHGEF3 knockdown (Figure 3D). Further, we found a significant positive correlation between the expression of ARHGEF3 and BIRC8 in our large cohort of NPC tissues (P=0.015, Table 4). There was no significant difference in XIAP expression between the ARHGEF3 high-expressing and low-expressing groups (P = 0.321, Table 4).
Table 3: List of genes differentially expressed in NPC CNE2 cells after ARHGEF3 knockdown using a human tumor apoptosis real-time PCR array
Gene |
Fold Change |
Location |
Function |
|---|---|---|---|
Downregulated genes |
|||
ABL1 |
-1.47 |
9q34.12 |
Induces cell division, adhesion |
AKT1 |
-1.37 |
14q32.33 |
Inhibits cell apoptosis |
BAD |
-1.40 |
11q13.1 |
Induces cell apoptosis |
BAG1 |
-1.68 |
9p13.3 |
Inhibits cell apoptosis |
BAG3 |
-1.83 |
10q26.11 |
Inhibits cell apoptosis |
BAG4 |
-1.45 |
8p11.23 |
Inhibits cell apoptosis |
BAK1 |
-1.73 |
6p21.31 |
Induces cell apoptosis |
BAX |
-1.65 |
19q13.33 |
Induces cell apoptosis |
BCL2A1 |
-1.93 |
15q25.1 |
Inhibits cell apoptosis |
BCL2L1 |
-1.63 |
20q11.21 |
Inhibits cell apoptosis |
BCL2L10 |
-1.03 |
15q21.2 |
Inhibits cell apoptosis |
BCL2L11 |
-1.76 |
2q13 |
Induces cell apoptosis |
BCL2L2 |
-1.91 |
14q11.2 |
Inhibits cell apoptosis |
BFAR |
-1.47 |
16p13.11 |
Inhibits cell apoptosis |
BID |
-1.01 |
22q11.21 |
Induces cell apoptosis |
BIK |
-1.95 |
22q13.2 |
Induces cell apoptosis |
BIRC2 |
-2.22 |
11q22.2 |
Inhibits cell apoptosis |
BIRC3 |
-3.55 |
11q22.2 |
Inhibits cell apoptosis |
BIRC6 |
-2.27 |
2p22.3 |
Inhibits cell apoptosis |
BIRC8 |
-5.04 |
6q21 |
Inhibits cell apoptosis |
BNIP1 |
-1.03 |
5q35.1 |
Inhibits cell apoptosis |
BNIP2 |
-1.69 |
15q22.2 |
Inhibits cell apoptosis |
BNIP3 |
-1.65 |
10q26.3 |
Inhibits cell apoptosis |
BNIP3L |
-1.60 |
8p21.2 |
Inhibits cell apoptosis |
BRAF |
-1.97 |
7q34 |
Inhibits cell apoptosis |
CARD6 |
-1.07 |
5p13.1 |
Induces cell apoptosis |
CARD8 |
-1.67 |
19q13.33 |
Induces cell apoptosis |
CASP3 |
-1.11 |
4q35.1 |
Induces cell apoptosis |
CASP4 |
-1.13 |
11q22.3 |
Induces cell apoptosis |
CASP5 |
-1.58 |
11q22.3 |
Induces cell apoptosis |
CASP6 |
-1.52 |
4q25 |
Induces cell apoptosis |
CASP7 |
-1.56 |
10q25.3 |
Induces cell apoptosis |
CASP8 |
-1.29 |
2q33.1 |
Induces cell apoptosis |
CD40 |
-1.00 |
20q13.12 |
Induces cell apoptosis |
CFLAR |
-1.54 |
2q33.1 |
Inhibits cell apoptosis |
CIDEA |
-1.13 |
18p11.21 |
Induces cell programmed death |
CIDEB |
-1.47 |
14q12 |
Induces cell programmed death |
CRADD |
-1.99 |
12q22 |
Induces cell apoptosis |
DAPK1 |
-1.02 |
9q21.33 |
Induces cell programmed death |
FADD |
-1.66 |
11q13.3 |
Induces cell programmed death |
FAS |
-1.50 |
10q23.31 |
Induces cell programmed death |
GADD45A |
-1.17 |
1p31.3 |
Induces cell apoptosis |
HRK |
-1.01 |
12q24.22 |
Induces cell apoptosis |
IGF1R |
-1.74 |
15q26.3 |
Inhibits cell apoptosis |
LTA |
-1.47 |
6p21.33 |
Induces cell apoptosis |
MCL1 |
-1.86 |
1q21.3 |
Inhibits cell apoptosis |
NAIP |
-6.12 |
5q13.2 |
Inhibits cell apoptosis |
NOL3 |
-1.45 |
16q22.1 |
Inhibits cell apoptosis |
RIPK2 |
-1.58 |
8q21.3 |
Induces cell apoptosis |
CD70 |
-1.88 |
19p13.3 |
T cell activator |
TNFSF8 |
-1.26 |
9q32 |
Induces cell apoptosis |
TP53 |
-1.89 |
17p13.1 |
Induces cell apoptosis |
TP53BP2 |
-1.13 |
1q41 |
Induces cell apoptosis |
TP73 |
-1.27 |
1p36.32 |
Induces cell apoptosis |
TRADD |
-1.38 |
16q22.1 |
Induces cell apoptosis |
TRAF2 |
-1.27 |
9q34 |
Induces cell apoptosis |
TRAF3 |
-1.85 |
14q32.32 |
Induces cell apoptosis |
XIAP |
-4.8 |
Xq25 |
Inhibits cell apoptosis |
B2M |
-1.23 |
15q21.1 |
Immune response |
HPRT1 |
-1.17 |
Xq26.1 |
Nucleotide metabolism |
ACTB |
-1.09 |
7p22.1 |
Cytoskeleton actin |
Upregulated genes |
|||
APAF1 |
1.16 |
12q23.1 |
Induces cell apoptosis |
BCL10 |
1.79 |
1p22.3 |
Induces cell apoptosis |
BCL2 |
1.17 |
18q21.33 |
Induces cell apoptosis |
BCLAF1 |
1.37 |
6q23.3 |
Induces cell apoptosis |
NOD1 |
1.75 |
7p14.3 |
Induces cell apoptosis |
CASP1 |
1.18 |
11q22.3 |
Induces cell apoptosis |
CASP10 |
1.96 |
2q33.1 |
Induces cell apoptosis |
CASP14 |
1.08 |
19p13.12 |
Induces cell apoptosis |
CASP2 |
1.08 |
7q34 |
Induces cell apoptosis |
CASP9 |
1.67 |
1p36.21 |
Induces cell apoptosis |
CD40LG |
1.44 |
Xq26.3 |
Inhibits cell apoptosis |
DFFA |
1.55 |
1p36.22 |
Induces DNA damage |
FASLG |
2.99 |
1q24.3 |
Induces cell apoptosis |
LTBR |
1.77 |
12p13.31 |
Inhibits cell apoptosis |
PYCARD |
1.72 |
16p11.2 |
Induces cell apoptosis |
TNF |
1.74 |
6p21.33 |
Induces cell apoptosis |
TNFRSF10A |
1.11 |
8p21.3 |
Induces cell apoptosis |
TNFRSF10B |
1.08 |
8p21.3 |
Induces cell apoptosis |
TNFRSF11B |
1.66 |
8q24.12 |
Induces cell apoptosis |
TNFRSF1A |
1.06 |
12p13.31 |
Induces cell apoptosis |
TNFRSF21 |
1.40 |
6p21.1 |
Induces cell apoptosis |
TNFRSF25 |
1.09 |
1p36.31 |
Induces cell apoptosis |
CD27 |
1.39 |
12p13.31 |
Induces cell apoptosis |
TNFRSF9 |
1.51 |
1p36.23 |
Induces cell apoptosis |
TNFSF10 |
1.59 |
3q26.31 |
Induces cell apoptosis |
RPL13A |
1.14 |
19q13.3 |
Protein metabolism |
GAPDH |
1.02 |
12p13.31 |
Glycometabolism |
Abbreviation: NPC, nasopharyngeal carcinoma; ARHGEF3, Rho guanine nucleotide exchange factor3.
Table 4: Correlation between expression of ARHGEF3 and that of BIRC8 and XIAP in 192 patients with NPC
All cases |
BIRC8 expression |
XIAP expression |
|||||
|---|---|---|---|---|---|---|---|
Low |
High |
P value* |
Low |
High |
P value* |
||
ARHGEF3 expression |
0.015 |
0.321 |
|||||
Low |
81 |
48 (59.3%) |
33 (40.7%) |
46 (56.8%) |
35 (43.2%) |
||
High |
111 |
46 (41.4%) |
65 (58.6%) |
55 (49.5%) |
56 (50.5%) |
||
*Chi-square test.
Abbreviation: NPC, nasopharyngeal carcinoma; ARHGEF3, Rho guanine nucleotide exchange factor3.
DISCUSSION
ARHGEF3 is a key activator of Rho-GTPases including, in particular RhoA, Rac1, and CDC42, which function as molecular switches in a variety of cellular signaling pathways [16, 17]. Increasing evidence suggests that Rho-GTPases are frequently deregulated during tumor progression, which promotes malignant phenotypes in cancer cells and is correlated with poor patient prognosis [18–20]. However, the molecular status of ARHGEF3 and its potential function in the underlying mechanisms of NPC are unclear.
In the present study, we demonstrated that the majority of human NPC cell lines and tissues expressed high levels of endogenous ARHGEF3 protein compared to control non-neoplastic nasopharyngeal cells and tissues. These findings suggest that upregulated expression of ARHGEF3 provides a selective advantage in NPC pathogenic processes.
Our analyses also found that high ARHGEF3 expression in our NPC cohorts was positively correlated with tumor T status, distant metastasis, and advanced clinical stage, suggesting that high expression of ARHGEF3 facilitates a malignant phenotype in NPC. Further, we observed that expression of ARHGEF3 was a strong and independent prognostic predictor for NPC patients. These findings underscore a potentially important role of ARHGEF3 in the development and progression of NPC, and suggest that examination of ARHGEF3 expression by IHC could be used as an additional tool in identifying those NPC patients at increased risk of tumor growth and/or metastasis.
Our investigation of the mechanisms through which ARHGEF3 regulates NPC cell malignancy demonstrated that knockdown of ARHGEF3 in CNE2 and SUNE1 NPC cells dramatically repressed cell growth, migration, and invasion in vitro. In contrast, ARHGEF3 overexpression in HONE1 NPC cells induced cell tumorigenicity in vivo. Moreover, in a tail vein injection mouse model of cancer metastasis ectopic overexpression of ARHGEF3 in HONE1 cells led to a significant increase in the number of metastatic lung lesions. These data support our emerging view that increased ARHGEF3 expression is a critical molecular event in the process of NPC pathogenesis.
It has been reported that cancer cells exhibit deficiencies in the induction of apoptosis, resulting in accelerated tumor development and reduced responsiveness to anti-cancer therapies [21–23]. Conversely, increased apoptosis in cancer cells may provide therapeutic benefits, especially in apoptosis-defective cancers [24, 25]. In the current study, we found that silencing ARHGEF3 in NPC cells could induce apoptosis, as measured by an increased percentage of annexin V positive cells and increased cleavage of caspases3. Gene expression profiling also revealed that silencing of ARHGEF3 resulted in downregulation of a number of genes, most notably BIRC8. Further, we did observe a significant positive correlation between expression of ARHGEF3 and BIRC8 in our large cohort of NPC clinical tissues.BIRC8 (ILP-2) belongs to the inhibitors of apoptosis protein (IAP) family, which are apoptosis inhibitors that may protect against apoptotic stimuli and suppress apoptosis [26]. It has been reported that BIRC8 exerts its effects by association with an inhibition of specific caspases [27]. In addition, studies have documented that BIRC8 (ILP-2) is a tumor biomarker and promotes cancer progression [28, 29]. These results suggest that ARHGEF3 regulates cell apoptosis via control of BIRC8 expression, which is in turn involved in attenuation of caspases3-induced apoptosis in the pathogenesis of NPC.
Our study demonstrates, for the first time, the protein expression dynamics of ARHGEF3 in a large cohort of clinical NPC tissues. High expression of ARHGEF3 may be important in tumorigenesis and acquisition of a poor prognostic phenotype of human NPC. Importantly, our functional and mechanistic studies suggest an important oncogenic role for ARHGEF3 in the suppression of NPC cell apoptosis by regulating BIRC8 expression and caspases3-induced apoptosis, activities that might be responsible for the development and progression of human NPCs.
MATERIALS AND METHODS
Nasopharyngeal cell lines and tissue specimens
Seven human NPC cell lines (CNE1, CNE2, HONE1, SUNE1, 5-8F, 6-10B and C666) and one immortalized normal nasopharyngeal cell line (NP69) were cultured in RPMI-1640 supplemented with 10% fetal bovine serum. 192 specimens of NPC and 50 specimens of non-neoplastic nasopharyngeal mucosa were collected at Sun Yat-Sen University Cancer Center, Guangzhou, China, between January 1991 and August 2000. A nasopharyngeal tissue microarray (TMA) was then constructed. In addition, 9 pairs of fresh NPC tissues and adjacent non-neoplastic nasopharyngeal mucosa specimens were collected at Guangdong Provincial People’s Hospital (Guangzhou, China) in 2012. None of the NPC patients had received preoperative radiation or chemotherapy before diagnosis. A pathological diagnosis for all specimens was confirmed according to the 2005 WHO histological classification of NPC. Tumor stage was defined according to the criteria of the sixth edition of the TNM classification of the Union for International Cancer Control (UICC, 2002). The study was approved by the Institute Research Medical Ethics Committee of Sun Yan-Sun University Cancer Center (Guangzhou, China).
Western blotting assay
Equal amounts of whole cell and tissue lysates were resolved by SDS-polyacrylamide gel electrophoresis and electrotransferred on a polyvinylidenedifluoride membrane (Pall Corp., Port Washington, NY). The samples were then incubated with primary antibodies against ARHGEF3 (Abgent Laboratories, San Diego, CA), caspase-3, cleaved caspase-3, caspase-8 (BD Biosciences, San Jose, CA), PARP, NAIP(Abcam, Cambridge, UK), BIRC2, BIRC3, XIAP, BIRC6, BIRC8 (Proteintech, Chicago, IL), FASLG (Abnova, Taibei City, Taiwan), α-tubulin (Santa Cruz Biotech, Dallas, TX), or GAPDH (BD Biosciences, San Jose, CA). The immunoreactive signals were detected with an enhanced chemiluminescence reagent kit (Amersham Biosciences, Uppsala, Sweden). The procedures were conducted in accordance with the manufacturer’s instructions.
Immunohistochemistry (IHC) staining
IHC studies were performed using a standard streptavidin biotin-peroxidase complex method. For antigen retrieval, TMA slides were microwave treated in 10 mM citrate buffer (pH 6.0) for 10 min. The TMA slides were incubated with anti-ARHGEF3 (1:100 dilution; Abgent Laboratories, San Diego, CA), in a moist chamber overnight at 4ºC. A negative control was obtained by replacing the primary antibody with a normal rabbit IgG.
A semi-quantitative scoring criterion for IHC of ARHGEF3 was used, in which both staining intensity and positive areas were recorded. A staining index (values 0 to 12) obtained as the intensity of positive staining (negative = 0, weak = 1, moderate = 2, or strong = 3 scores) and the proportion of immunopositive cells of interest (< 25% = 1, 25 to 50% = 2, 51% to 75% = 3, > 75% = 4 scores) were calculated. Because the staining index of expression of ARHGEF3 in all 50 non-neoplastic nasopharyngeal mucosa tissues was no more than 4, we designated staining index scores of 0-4 as “low” expression of ARHGEF3 (Figure 1B right) and staining index scores of 6-12 was as “high” expression of ARHGEF3 (Figure 1B left).The cutoff scores for determining “high” level expression of XIAP and BIRC8 (ILP-2) were determined at staining index scores of >4 and >6, respectively.
Small interfering RNA (siRNA)
CNE2 and SUNE1 cells were cultured in six-well plates. The cells were transfected with anti-ARHGEF3 siRNA or anti-control siRNA (Ambion, Austin, Texas) using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. The gene silencing effect was measured by Western blotting 48 h post-transfection.
Colony formation assay
Five hundred infected cells were placed in a fresh six-well plate and maintained in RPMI1640 containing 10%FBS for 10 days. Colonies were fixed with methanol and stained with 0.1% Giemsa in 20% methanol for 15 minutes.
Wound-healing and matrigel invasion assays
Cell migration was assessed by measuring the movement of cells into a scraped cellular area created by 200μL pipette tip.The spread of wound closure was observed after 48h and photographed under a microscope. We measured the fraction of cell coverage across the line to calculate migration rate.
For invasion assays, 1×104 cells were added to a Matrigel invasion chamber (BD Biosciences, Becton Dickson Labware, Franklin Lakes, NJ) present in the insert of a 24-well culture plate. FBS was added to the lower chamber as a chemoattractant. After 48h, the non-invading cells were gently removed with a moist cotton swab. Invasive cells located on the lower side of the chamber were fixed with paraformaldehyde and then stained with crystal violet, air dried, and photographed. For colorimetric assays, the samples were treated with 150μL 10% acetic acid and absorbance was measured with a spectrophotometer at 560nm.
Plasmid constructs and transfection
ARHGEF3 and BIRC8 complementary DNA (Fulengen, Guangzhou, China) was cloned into a pcDNA3.1 plasmid. HONE1 Cells were transfected with pcDNA-ARHGEF3 or the control plasmid pcDNA3.1(+) using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. For the establishment of the ARHGEF3-HONE1 cell line stably expressing ARHGEF3, the cells were split at a ratio of 1:10 48 h after transfection. Next, cells were maintained in Leibovitz’s L-15 medium containing 200 μg/mL of G418 (Calbiochem, San Diego, CA). After 6 weeks of selection, resistant colonies stably transfected with pcDNA-ARHGEF3 (HONE1 pcDNA-ARHGEF3) or pcDNA3.1(+) [HONE1pcDNA3.1(+)] were pooled.
In vivo tumorigenesis and metastasis assays
For the in vivo assays of subcutaneous tumorigenesis of NPC cells, 1×106 mixed populations of HONE1-ARHGEF3 cells stably overexpressing ARHGEF3 or the control HONE1–vector were injected subcutaneously into the backs of 4-week-old male BALB/C-nu athymic nude mice. At day 30, the mice were sacrificed, the tumors were removed, and tumor weight was calculated.
To evaluate the metastasis potential of NPC cells in vivo, five 4-week-old male BALB/C-nu athymic nude mice in each experimental group were injected with HONE1-ARHGEF3 and HONE1-vector cells, separately. Briefly, each mouse received 2×105 cells via tail vein injection. Eight weeks after cell injection mice were killed and the lungs and the liver were removed from each mouse and fixed with phosphate-buffered neutral formalin. Consecutive tissue sections were made for each block of the tissues, which were then stained with haematoxylin-eosin. Finally, the slides of the lungs and the liver were carefully examined under a microscope. All experimental procedures involving animals were are accordant with the Guidelines for the Care and Use of Laboratory Animals (NIH publications Nos. 80-23, revised 1996).
Apoptosis assay
To assess the rate of apoptosis, transfected cells were harvested and washed twice with cold PBS, and the Annexin V-PI Kit (Nanjing Keygen, Nanjing, China) was used according to the manufacturer’s guidelines. The detection was performed with a FACS Calibur using Cell Quest software (BDIS, San Jose, CA).
Real-time PCR gene array
RNA was extracted from siARHGEF3-CNE2 and siNC-CNE2 cells using Trizol (Invitrogen, Carlsbad, CA) and was cleaned them using the RNeasy MinElute cleanup kit (Qiagen, Valencia, CA). Messenger RNA expression levels were quantified with a Human Tumor Apoptosis RT2 Profiler PCR array (Super Array Bioscience, Frederick, MD). SiARHGEF3-CNE2 cells were compared with control siNC-CNE2 cells using a Human Apoptosis RT2 Profiler PCR array containing 84 key genes involved in programmed cell death.
Statistical analysis
Statistical analysis was performed using the SPSS software package (SPSS Standard version18.0, SPSS Inc). Differences between variables were assessed by the Chi-square test or Fisher’s exact test. For survival analysis, we analyzed all patients with NPC by Kaplan-Meier analysis. A log rank test was used to compare different survival curves. Multivariate survival analysis was performed on all parameters that were found to be significant in univariate analysis using the Cox regression model. Data derived from cell line experiments are presented as mean ±SD and assessed by the two-tailed Student’s t test. P values of < 0.05 were considered statistically significant.
CONFLICTS OF INTEREST
The authors declare that there is no conflict of interest.
GRANT SUPPORT
This work was supported by grants from the Nature Science Foundation of China (No. 81225018 and 81572848), the Science and Technology Planning Project of Guangdong Province, China (No. 2013B021800103)and the Foundation of Guangzhou Science and Technology Innovation Commission, China (No.2014J4100181).
REFERENCES
1. Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005; 55: 74-108.
2. Cao SM, Simons MJ, Qian CN. The prevalence and prevention of nasopharyngeal carcinoma in China. Chin J Cancer. 2011; 30: 114-119.
3. Tong ZT, Cai MY, Wang XG, Kong LL, Mai SJ, Liu YH, Zhang HB, Liao YJ, Zheng F, Zhu W, Liu TH, Bian XW, Guan XY, et al. EZH2 supports nasopharyngeal carcinoma cell aggressiveness by forming a co-repressor complex with HDAC1/HDAC2 and Snail to inhibit E-cadherin. Oncogene. 2012; 31: 583-594.
4. Xiao L, Hu ZY, Dong X, Tan Z, Li W, Tang M, Chen L, Yang L, Tao Y, Jiang Y, Li J, Yi B, Li B, et al. Targeting Epstein-Barr virus oncoprotein LMP1-mediated glycolysis sensitizes nasopharyngeal carcinoma to radiation therapy. Oncogene. 2014; 33: 4568-4578.
5. Razak AR, Siu LL, Liu FF, Ito E, O’Sullivan B, Chan K. Nasopharyngeal carcinoma: the next challenges. Eur J Cancer.2010;46: 1967-1978.
6. D’Amato L, Dell’Aversana C, Conte M, Ciotta A, Scisciola L, Carissimo A, Nebbioso A, Altucci L. ARHGEF3 controls HDACi-induced differentiation via RhoA-dependent pathways in acute myeloid leukemias. Epigenetics.2015;10: 6-18.
7. Thiesen S, Kubart S, Ropers HH, Nothwang HG (2000). Isolation of two novel human RhoGEFs, ARHGEF3 and ARHGEF4, in 3p13-21 and 2q22. Biochem Biophys Res Commun. 273: 364-369.
8. Mullin BH, Prince RL, Dick IM, Hart DJ, Spector TD, Dudbridge F, Wilson SG. Identification of a role for the ARHGEF3 gene in postmenopausal osteoporosis. Am J Hum Genet. 2008; 82: 1262-1269.
9. Hall A. Rho family GTPases. Biochem Soc Trans. 2012; 40: 1378-1382.
10. Arthur WT, Ellerbroek SM, Der CJ, Burridge K, Wennerberg K.XPLN, a guanine nucleotide exchange factor for RhoA and RhoB, but not RhoC. J Biol Chem. 2002; 277: 42964-42972.
11. Mullin BH, Mamotte C, Prince RL, Wilson SG. Influence of ARHGEF3 and RHOA knockdown on ACTA2 and other genes in osteoblasts and osteoclasts. PLoS One. 2014; 9: e98116.
12. Benedetti G, Fokkelman M, Yan K, Fredriksson L, Herpers B, Meerman J, van de Water B, de Graauw M. The nuclear factor kappaB family member RelB facilitates apoptosis of renal epithelial cells caused by cisplatin/tumor necrosis factor alpha synergy by suppressing an epithelial to mesenchymal transition-like phenotypic switch. Mol Pharmacol. 2013; 84: 128-138.
13. Laurin M, Huber J, Pelletier A, Houalla T, Park M, Fukui Y, Haibe-Kains B, Muller WJ, Côté JF. Rac-specific guanine nucleotide exchange factor DOCK1 is a critical regulator of HER2-mediated breast cancer metastasis. Proc Natl Acad Sci USA. 2013; 110: 7434-7439.
14. Matsuoka T, Yashiro M. Rho/ROCK signaling in motility and metastasis of gastric cancer. World J Gastroenterol. 2014; 20: 13756-13766.
15. Zegers MM, Friedl P. Rho GTPases in collective cell migration. Small GTPases. 2014; 5: e28997.
16. Serbanovic-Canic J, Cvejic A, Soranzo N, Stemple DL, Ouwehand WH, Freson K. Silencing of RhoA nucleotide exchange factor, ARHGEF3, reveals its unexpected role in iron uptake. Blood. 2011; 118: 4967-4976.
17. Kristelly R, Gao G, Tesmer JJ. Structural determinants of RhoA binding and nucleotide exchange in leukemia-associated Rho guanine-nucleotide exchange factor. J Biol Chem. 2004; 279: 47352-47362.
18. Cheng KW, Agarwal R, Mills GB. Ras-superfamily GTP-ases in ovarian cancer. Cancer Treat Res. 2009; 149: 229-240.
19. Grise F, Bidaud A, Moreau V. Rho GTPases in hepatocellular carcinoma. Biochim Biophys Acta. 2009; 1795: 137-151.
20. Huh YH, Oh S, Yeo YR, Chae IH, Kim SH, Lee JS, Yun SJ, Choi KY, Ryu JH, Jun CD, Song WK. Swiprosin-1 stimulates cancer invasion and metastasis by increasing the Rho family of GTPase signaling. Oncotarget. 2015; 6: 13060-13071. doi:10.18632/oncotarget.3637.
21. Chow SE, Kao CH, Liu YT, Cheng ML, Yang YW, Huang YK, Hsu CC, Wang JS. Resveratrol induced ER expansion and ER caspase-mediated apoptosis in human nasopharyngeal carcinoma cells. Apoptosis. 2014; 19: 527-541.
22. Lung HL, Man OY, Yeung MC, Ko JM, Cheung AK, Law EW, Yu Z, Shuen WH, Tung E, Chan SH, Bangarusamy DK, Cheng Y, Yang X, et al. SAA1 polymorphisms are associated with variation in antiangiogenic and tumor-suppressive activities in nasopharyngeal carcinoma. Oncogene. 2015; 34: 878-889.
23. Li L, Tao Q, Jin H, van Hasselt A, Poon FF, Wang X, Zeng MS, Jia WH, Zeng YX, Chan AT, Cao Y. The tumor suppressor UCHL1 forms a complex with p53/MDM2/ARF to promote p53 signaling and is frequently silenced in nasopharyngeal carcinoma. Clin Cancer Res. 2010; 16: 2949-2958.
24. Liu Z, Long X, Chao C, Yan C, Wu Q, Hua S, Zhang Y, Wu A, Fang W. Knocking down CDK4 mediates the elevation of let-7c suppressing cell growth in nasopharyngeal carcinoma. BMC Cancer. 2014; 14: 274.
25. Deng Q, Yu X, Xiao L, Hu Z, Luo X, Tao Y, Yang L, Liu X, Chen H, Ding Z, Feng T, Tang Y, Weng X, et al. Neoalbaconol induces energy depletion and multiple cell death in cancer cells by targeting PDK1-PI3-K/Akt signaling pathway. Cell Death Dis. 2013; 4: e804.
26. Shin H, Renatus M, Eckelman BP, Nunes VA, Sampaio CA, Salvesen GS. The BIR domain of IAP-like protein 2 is conformationally unstable: Implications for caspase inhibition. Biochem J. 2005; 385: 1-10.
27. Richter BW, Mir SS, Eiben LJ, Lewis J, Reffey SB, Frattini A, Tian L, Frank S, Youle RJ, Nelson DL, Notarangelo LD, Vezzoni P, Fearnhead HO, Duckett CS. Molecular cloning of ILP-2, a novel member of the inhibitor of apoptosis protein family. Mol Cell Biol. 2001; 21: 4292-4301.
28. Xiang M, Zhou W, Gao D, Fang X, Liu Q. Inhibitor of apoptosis protein-like protein-2 as a novel serological biomarker for breast cancer. Int J MolSci. 2012; 13: 16737-16750.
29. Glodkowska-Mrowka E, Solarska I, Mrowka P, Bajorek K, Niesiobedzka-Krezel J, Seferynska I, Borg K, Stoklosa T. Differential expression of BIRC family genes in chronic myeloid leukaemia--BIRC3 and BIRC8 as potential new candidates to identify disease progression. Br J Haematol. 2014; 164: 740-742.

