Introduction
Lung cancer accounts for about 13% of all cancer diagnoses and remains the leading cause of death by cancer in the world [1], with almost 70% of patients diagnosed with locally advanced or metastatic disease at presentation [1, 2]. Non-small cell lung cancer (NSCLC) accounts for approximately 85% of all cases of lung cancer and is associated with poor prognosis [2]. The 5-year overall survival rate for NSCLC across all stages is only 21% and is even lower (~5%) for stages IIIB and IV [1, 3].
Oncogenic ‘driver mutations’ have now been identified in various subsets of NSCLC [4-6]. Of these drivers, somatic mutations in epidermal growth factor receptor (EGFR) and anaplastic lymphoma kinase (ALK) [5-7] are the most frequently described. In 2007, researchers identified the presence of a chimeric ALK protein with fibroblast-transforming properties that was formed following fusion of the echinoderm microtubule-associated protein-like 4 (EML4) and ALK genes [6]. EML4-ALK subverts intracellular signaling pathways to promote tumor cell survival and growth [8]. The overall incidence of ALK gene rearrangement in NSCLC ranges between 0.4% and 13.4%, and is similar in both Asian and Western populations [9]. This discovery resulted in the accelerated development and approval by the U.S. Food and Drug Administration (FDA) of the ALK-targeting tyrosine kinase inhibitors (TKIs) crizotinib (Xalkori®, Pfizer, New York, USA) in 2011, and ceritinib (Zykadia™, Novartis, Basel, Switzerland) in 2014 to treat patients with metastatic NSCLC who express the abnormal ALK gene [10, 11].
The true therapeutic benefit of novel molecules targeting the mutant ALK fusion protein in NSCLC relies on identifying the right patient population for treatment, and on detecting the emergence of tumor resistance. The American Society of Clinical Oncology (ASCO) endorsed the joint College of American Pathologists (CAP)/International Association for the Study of Lung Cancer (IASLC)/Association for Molecular Pathology (AMP) clinical practice guideline on EGFR and ALK molecular testing for patients with lung cancer, which holds that an ALK fluorescent in-situ hybridization (FISH) assay using dual-labeled break-apart probes is the preferred testing methodology to detect ALK gene rearrangement [12].
The accuracy of testing nonetheless depends on the quality of tumor biopsies. Approximately 50% of NSCLC patients who undergo re-biopsy for determination of resistance after first-line chemotherapy have insufficient/non-diagnostic biopsy specimens or cytology samples available for molecular testing [13]. Biopsy is invasive and repeating the procedure is not always feasible due to safety concerns and general unwillingness of patients, among other reasons [14]. Furthermore, lung adenocarcinomas are heterogeneous with a diverse and ever-evolving genetic and epigenetic makeup that contributes towards treatment resistance [15]. These barriers to biopsy collectively pose a challenge to track oncogene activity in real-time over the course of treatment. There is a need for a minimally invasive assay for tumor molecular profiling and continuous treatment monitoring in order to provide timely and tailored cancer treatment.
Circulating tumor cells (CTCs) released from the primary tumor site into the circulation represent a potential means of non-invasively isolating tumor cells for ALK FISH testing and other molecular characterizations. Recent data supports the role of these renegade cells as seeds of cancer metastases [16, 17]. They may recapitulate the phenotypic heterogeneity and molecular signatures of the primary tumor, as well as that of metastatic lesions [18-20]. While their presence and prevalence in blood are often associated with poor prognosis [21], CTCs may hold further relevance as an alternative tumor source, which can complement existing tissue-based diagnostic tests, especially when biopsy material is absent or inadequate. In patients with lung adenocarcinoma, hypothesis-generating studies have strongly suggested that ALK status could be determined based on testing of CTCs, with comparable results as testing of tumor tissues [22, 23].
The key challenge of CTC-based testing is the enrichment and isolation of these cells within an acceptable timeframe. Various technical approaches have been used to isolate these CTCs [19, 24, 25]. They can be broadly categorized into antibody affinity-based, imaging-based and size-based techniques [26]. The only current US FDA-approved CTC capturing technology utilizes EpCAM immunomagnetic means to isolate EpCAM-positive CTCs for prognostic purposes and would inadvertently miss out on EpCAM-negative CTCs [27, 28]. As a predictive biomarker for treatment monitoring and molecular analysis, it is pertinent to ensure reliable and reproducible isolation of CTCs of different phenotypic and molecular subtypes for various pre-and post-treated patient cohorts [25]. An increasing body of evidence suggests that non-immunomagnetic-based CTC technologies can reliably retrieve a comprehensive population of CTCs for molecular subtyping [19, 28].
Here, we evaluated the feasibility of an antibody-independent CTC isolation system using lung adenocarcinomas that have been tested ALK positive as a model to examine the concordance patterns between CTCs and tumor tissue, and to determine whether CTCs were reproducibly detectable in circulation. We further explored the potential use of CTCs in lung cancer, as a surrogate for molecular testing of the primary tumor for ALK gene rearrangement.
Results
Study group
We prospectively recruited 27, mostly late-stage NSCLC patients, 14 of whom had ALK-rearranged and 12 had wild-type ALK, determined from the initial biopsy diagnoses. One patient in the cohort, who was not from Singapore, had an unknown ALK status due to incomplete referral records. Sixty percent of patients were males. All ALK-positive patients were non-smokers. Five healthy donors (three males and two females), aged between 18-55 years old with no history of cancer, were also recruited into the control cohort. As ALK translocation in NSCLC patients is strongly correlated with a non-smoker or light smoker status [29], these healthy donors were non-smokers. The clinicopathologic features of the study group are summarized in Table 1.
Table 1: Clinicopathological characteristics of patients enrolled in this study
Patient characteristics |
Cases (%) N= 27 |
Age, years |
32–76 |
Sex |
|
Male |
16 (59.3%) |
Female |
11 (40.7%) |
Smoking history |
|
Non-smoker |
16 (59.3%) |
Smoker |
5 (18.5%) |
Ex-smoker |
5 (18.5%) |
No info |
1 (3.7%) |
Clinical staging |
|
IB |
1 (3.7%) |
IIIA |
1 (3.7%) |
IIIB |
2 (7.4%) |
IV |
23 (85.2%) |
Histological subtype |
|
ALK-positive |
14 (51.9%) |
Adenocarcinoma (NSCLC) |
11 (40.7%) |
Unknown subtype (NSCLC) |
3 (11.1%) |
ALK-negative |
12 (44.4%) |
Adenocarcinoma (NSCLC) |
5 (18.5%) |
Unknown subtype (NSCLC) |
7 (25.9%) |
ALK status unknown |
1 (3.7%) |
Adenocarcinoma (NSCLC) |
1 (3.7%) |
Abbreviations: ALK, anaplastic lymphoma kinase; NSCLC, non-small cell lung cancer
Histopathological analysis of tumor tissues
In the ALK-positive group, three out of 14 tumor tissues exhibited morphology that is associated with ALK rearrangements [30]. Histological preparations showed solid adenocarcinoma with signet ring cells (Figure 1A and 1B), and cribrifom adenocarcinoma with focal squamoid cells (not shown). Immunohistochemical (IHC) studies showed strong and diffuse nuclear reaction for Thyroid Transcription Factor-1 (TTF-1). This finding confirmed the diagnosis of adenocarcinoma of lung origin in this particular setting (Figure 1C). Some tumors also showed focal reaction to periodic acid-Schiff with diastase (PAS-D) within mucin vacuoles, which is a general feature of adenocarcinomas, as opposed to squamous cell carcinomas (Figure 1D) [30].
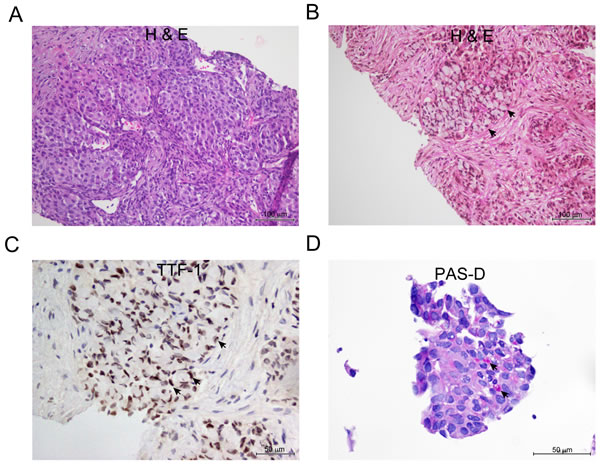
Figure 1: Representative appearance of NSCLC adenocarcinoma with signet ring cells features. A. H&E stain showing solid nests of tumor cells B. Solid with signet ring cells (arrow ) C. Thyroid transcription factor-1 (TTF-1) IHC stain showing strong nuclear reaction in the signet ring cells. D. Solid tumor showing focal positive reaction for (PAS-D) within mucin vacuoles (arrow).
Concordance in ALK rearrangement pattern between CTCs and tumor
Following FISH testing on all tumor samples in the cohort, it was found that ALK-positive tumors harbored ALK rearrangements with various patterns of abnormality (Table 2). The majority of the tumor samples harbored the one fusion (F) and one split orange (R) and green (G) signal (Figure 2A). The tumor from Patient P5 presented various ALK rearrangement patterns such as 1F1R1G, 2F1R, 2F2R, 1F1R and 1F2R.
FISH testing was subsequently performed on CTCs that were enriched and isolated from the matched blood samples. Data showed that ALK rearrangement patterns (majority 1F1R1G) observed in primary tumor tissues were recapitulated on most of the ALK-positive CTCs, giving an overall concordance rate of over 90% based on the 1F1R1G fusion pattern (Table 2). In Patient P5 (Table 2), the CTCs were able to recapitulate three out of five ALK rearrangement patterns observed in the tumor tissue.
Table 2: Concordance of ALK rearrangement patterns between CTC and tumor in patients with ALK-positive NSCLC
Case number of patients with ALK-positive NSCLC |
ALK rearranged/ total cells scored (Tumor) |
ALK rearrangement patterns (% of tumor cells observed with respective patterns) |
|
Tumor |
CTC |
||
P1 |
61/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P2 |
45/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P3 |
79/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P4 |
30/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P5 |
61/100 |
1F1R1G (4.9%) 2F1R (34.4%) 2F2R (3.3%) 1F1R (49.2%) 1F2R (8.2%) |
1F1R1G (50%) 2F1R (37.5%) 2F2R (12.5%) |
P6 |
72/100 |
1F1R1G (29.2%) 1F1R (70.8%) |
1F1R1G (100%) |
P7 |
81/100 |
1F1R (100%) |
1F1R (75%) 1F1R1G (25%) |
P8 |
55/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P9 |
77/100 |
1F1R (31.2%) 1F1R1G (68.8%) |
1F1R (100%) |
P10 |
100/100 |
1F1R (100%) |
1F1R (50%) 1F1R1G (50%) |
P11 |
62/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P12 |
77/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P13 |
45/100 |
1F1R1G (100%) |
1F1R1G (100%) |
P14 |
Not available |
Not available |
1F1R1G (100%) |
Abbreviations: ALK, anaplastic lymphoma kinase; CTC, circulating tumor cells; NSCLC, non-small cell lung cancer.
We further observed an overexpression of vimentin in the tumor samples, along with the control bronchiolar epithelium (Figure 2B). However, loss of E-cadherin was not obvious in these samples.
The number of ALK-positive rearranged CTCs retrieved from ALK-positive patients was significantly enriched compared with ALK-negative patients (p < 0.0001) and healthy donors (p = 0.0003) (Figure 3).
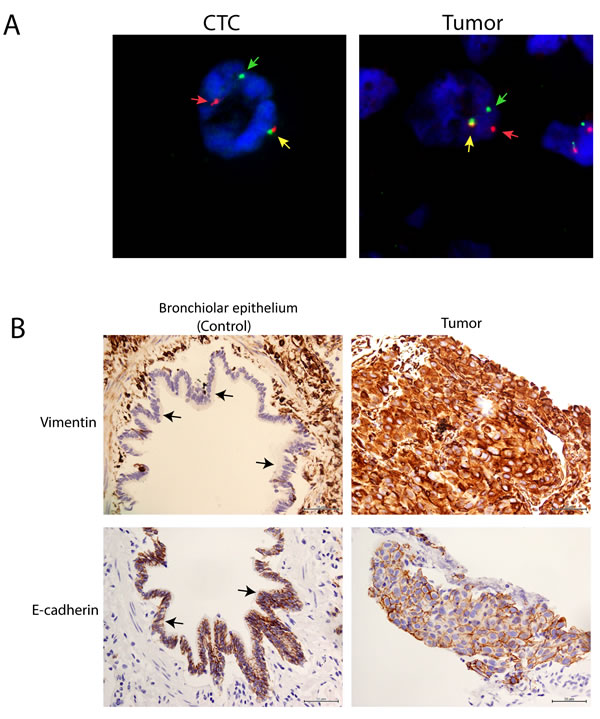
Figure 2: High concordance of ALK FISH rearrangements patterns between CTCs and tumors in NSCLC adenocarcinoma patients. A. Representative ALK FISH rearrangement patterns in CTCs and tumors showing 1F1R1G rearrangement patterns. Yellow, red and green arrows represent fusion (F), orange (R) and green (G) fluorescent signals. B. Representative vimentin (upper panel) and E-cadherin (lower panel) IHC in tumor and control bronchiolar epithelium (black arrow).
Establishment and validation of ALK break-apart probes cutoff in ALK-negative samples
ALK testing by FISH in NSCLC tumor tissues without ALK rearrangement may detect rearrangement-positive patterns (i.e. split patterns or isolated 3’ patterns) in a fraction of cells [31-33], likely because of truncation artefact caused by tissue sectioning, or perhaps a stochastic genomic alteration that does not indicate a specific gene fusion. ALK FISH testing in formalin-fixed, paraffin-embedded (FFPE) NSCLC tumor tissues has a ‘false positive’ cutoff value of 15% to allow for the best separation between ALK-rearranged and ALK wild-type cells [31, 33]. However, it is not possible to apply this guideline in the ALK FISH testing on CTCs because the number of CTCs in any given blood sample would be too low.
In our study, we established and validated the ‘false positive’ cutoff for ALK FISH in CTCs using 12 blood samples from NSCLC ALK-negative patients and five blood samples from healthy donors (Supplementary Data). Results from the ALK-negative NSCLC cohort scored a median of two or less positive cells (range 0-2 cells/1.88 mL blood). The result concurred with the numbers observed for healthy blood samples. In fact, no statistical difference in ALK-positive cell counts was observed between ALK-negative NSCLC cohort and healthy donors (p = 0.0973) (Figure 3). This data established the ‘false-positive’ cutoff for ALK break-apart probes in CTCs at ≤ two cells per 1.88 mL blood.
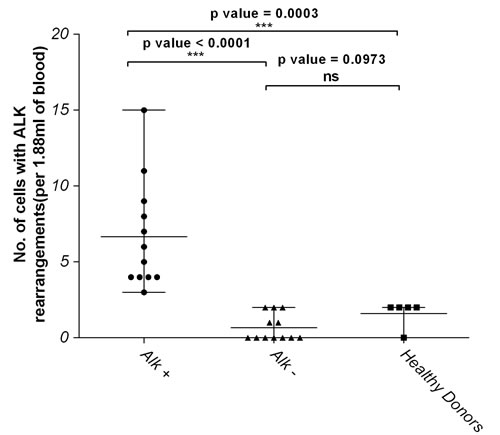
Figure 3: Number of cells with ALK rearrangements in ALK-positive NSCLC patients is significantly higher compared to ALK-negative and healthy donors. Graph represents statistical analyses of the data on Table S1 using the non-parametric two-tailed t-test. NS represents not significant while p value <0.05 were considered significant.
Potential clinical applications
Sequential CTC enumeration and FISH was performed on blood samples from a patient with no accessible tissue for ALK FISH testing. The index case was a never smoker male diagnosed with NSCLC. A transthoracic needle aspiration biopsy was performed on the right hilar mass to obtain a specimen for histological analysis. The hematoxylin and eosin (H&E) stain showed one small cluster of NSCLC cells with strong nuclear reaction for TTF-1 favoring adenocarcinoma. Unfortunately, his diagnostic tissue was exhausted and no further molecular profiling could be performed. He did not respond to EGFR tyrosine kinase inhibitor (TKI) therapy and had a short duration of response to pemetrexed and cisplatin. Re-biopsy of the lung and liver tumors was considered but the patient declined due to the risk of bleeding. He consented to blood sampling instead; the sample was subsequently processed as described in the Methods section.
At baseline, six CTCs displaying a 1F1R1G pattern were isolated and met the necessary cutoffs for ALK-positivity (Figure 4A). A trial of crizotinib was commenced. Confirmatory scans done 3 months after completion of treatment demonstrated good partial response in the liver and minor response in the primary lung tumor, based on RECIST criteria (Figure 4A). Sequential CTC counts dropped to three cells displaying the similar ALK rearranged pattern as the baseline. He continued on crizotinib but unfortunately, his disease progressed in the liver and the brain 5 months after treatment initiation (Figure 4A). A post-progression blood sample showed additional ALK rearrangement patterns present in his CTCs, which differed from the baseline patterns. New ALK rearrangement patterns such as 2R2G and 1F1R appeared, in addition to 1F1R1G, which was previously present (Figure 4B). The number of ALK-positive CTCs also increased from three to eleven CTCs per 1.88 mL of blood post-progression.
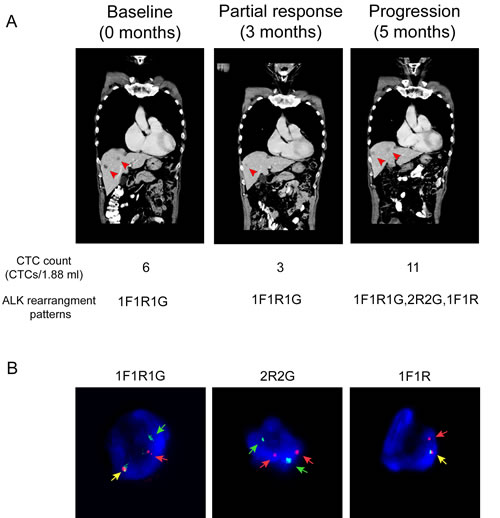
Figure 4: An index case suggests that ALK-rearranged CTCs could have clinical application as a diagnostic biomarker to monitor crizotinib treatment and response. A. CT scan taken at baseline, partial response and progression time points showing presence of metastatic tumor in liver (arrow). CTC counts and ALK rearrangement patterns for each time point is indicated in the lower panel. B. Representative images showing 1F1R1G, 2R2G and 1F1R ALK rearrangement patterns following progression on crizotinib treatment. Yellow, red and green arrows represent fusion (F), orange (R) and green (G) fluorescent signals.
Discussion
We successfully captured CTCs using an antibody-independent CTC isolation system. CTCs were enriched from the blood samples collected from 27 NSCLC patients, 14 of whom were ALK-positive. Three of the cases exhibited solid with signet ring cells pattern associated with ALK positivity [34-37]. Overall, CTCs isolated from the ALK-positive patient cohort were above detectable levels, even among previously treated patients.
The presence of ALK rearrangement in CTCs was previously analyzed and reported by French groups using the Isolation by Size of Epithelial Tumor (ISET) system, which is also an antibody-free system (Rarecells, Paris, France) [22, 23]. They characterized 18 ALK-positive lung tumor and CTC samples showing high concordance in ALK rearrangement among European patient cohort. In agreement with our data reported in this study, ‘false-positive’ signals were similarly observed on CTCs from ALK-negative samples and a 4 cell/mL cutoff was eventually established [22, 38].
In a similarly designed study, Pailler et al. [22] described high concordance in ALK rearrangement patterns between the CTC and tumor biopsies in 18 ALK-positive and 14 ALK-negative patients with metastatic NSCLC. This percentage of concordance is in agreement with our own results. They reported that all ALK-positive NSCLC patients in their cohort had 4 or more ALK-rearranged CTCs per mL of blood. The study did not include healthy donors to establish a ‘false positive’ cutoff for ALK FISH testing of CTCs. They further reported that CTCs harboring the 1F1R1G ALK rearrangement patterns is associated with epithelial-mesenchymal transition (EMT) phenotype [22].
Another study had reported that the EMT phenotype (represented by loss of E-cadherin and expression of vimentin) was more common in ALK-rearranged tumors than other genotypes (38.9%, 19.1%, 26.9% and 14.6% of ALK-rearranged, EGFR-mutated, K-ras mutated and triple negative tumors, respectively; p = 0.015) [39]. Separately, expression of vimentin alone was detected in 49.30% of ALK-rearranged tumors while loss of E-cadherin was detected in 71.30% [39]. In our study, we also observed an overexpression of vimentin in the tumor samples in comparison with the control bronchiolar epithelial tissue. However, the loss of E-cadherin was not obvious in our tumor samples, which suggested that some of the tumor cells retained their epithelial characteristics within a heterogeneous population of cells. The predominance of this particular ALK rearrangement pattern in our patient’s CTCs is therefore consistent with the observation above suggesting that these tumors and their CTCs may be favoring the EMT pathway.
This study further highlights the utility of antibody-independent microfluidic isolation systems for the isolation and downstream characterization of CTCs compared with immunomagnetic antibody-dependent systems. While the numbers of CTCs isolated here are small and may present substrate limitations to downstream characterization of CTC, it should be noted that the current numbers were derived from <2 mL of blood, as opposed to existing systems which use up to 7.5 mL of blood or more. In addition, we have previously demonstrated that there is an association between CTC number and the volume of blood processed [19]. This suggests a limitation that can be easily overcome.
The index case presented here raises the possibility that CTC enumeration based on ALK FISH may be associated with treatment response with crizotinib by imaging. The appearance of additional ALK rearrangement patterns following progression with crizotinib treatment exhibited a double split in both ALK alleles giving rise to the 2R2G ALK rearrangement pattern. The additional copy of the oncogenic ALK may have contributed to disease progression despite treatment with an ALK inhibitor. This observation is worthy of further inquiry, because while the presence of ALK copy number gain is correlated with crizotinib resistance, as previously reported by Doebele et al. [40], and in vitro studies have identified potential resistance mutations in the ALK gene, for example L1196M, G1269A, S1206Y and G1202R [40, 41], limited analysis of post-progression biopsies of tumors from a phase 1 study of LDK378 suggested that these secondary resistance mutations or gene amplification do not account for a majority of resistance cases [11]. Hence, further work with paired re-biopsies and sequential CTC collection may assist understanding of resistance mechanisms in ALK-driven tumors.
Conclusions drawn from our study are limited by its relative small patient population, as was Pallier’s study [22]. Nonetheless, the converging trend of both studies’ findings is indicative of the utility and potential of CTCs as an alternate target of ALK testing in lung cancer and informs the development of CTC-based technology. More importantly, these studies provide the basis for subsequent, large-scale validation studies.
In summary, high concordance of ALK rearrangement patterns in CTCs and tumors as assessed by ALK FISH testing indicates that CTCs may have utility as a non-invasive surrogate diagnostic tool and may be useful in the longitudinal follow-up for resistance profiling. The availability of a non-invasive tool would improve efforts to guide ALK-rearranged targeted treatment in NSCLC, especially in cases without tissue availability. Further efforts at downstream CTC characterization and culture following enrichment are ongoing.
Materials and Methods
Patient recruitment and blood samples
Patients with confirmed NSCLC were recruited into this trial. They were naïve for ALK-targeted TKI treatment, but may have received other forms of chemotherapy. Once informed consent was secured from these patients, their blood samples were processed for CTC analysis. The clinical sample collection protocols were reviewed and approved by SingHealth Centralised Institutional Review Board. Clinicopathological information was also recorded for these patients. Blood samples from healthy donors were used as controls in this study.
Histopathology
H&E was done using the Leica ST5010 XL automated stainer (Leica Biosystems, Wetzlar, Germany) while periodic PAS-D staining was done using the Ventana BenchMark Special Stains automated stainer (Ventana Medical Systems Inc, Tucson, Arizona, USA), following the respective standard protocols. Histological diagnoses were made based on the World Health Organization (WHO) classification [42]. IHC labeling was performed on the Ventana BenchMark Ultra autostainer (Ventana Medical Systems Inc, Tucson, Arizona, USA) using the UltraView detection kit and proprietary Standard CC1 (SC1) pre-treatment sets. The antibodies used with their dilution and pre-treatments were as follows: TTF-1 (Novacastra NCL-TTF-1, clone SPT24, SC1, dilution 1:30), vimentin (DAKO M0725, clone V9, SC1, 1:100) and E-cadherin (Dako M3612, clone MCH-38, SC1, 1:30) antibodies. Histopathology data was reviewed by pathologists who had been accredited by the College of American Pathologists (CAP).
CTC enrichment
Peripheral blood was collected using K2 EDTA vacutainer® blood collection tube (BD, Singapore) and processed within 24 hours. Subsequently, 7.5 mL of whole blood was incubated with red blood cell (RBC) lysis buffer (G-Biosciences, USA) according to manufacturer’s recommendations. Lysed RBCs in the supernatant were discarded after centrifugation. Remaining cell pellet containing CTCs was resuspended in ClearCell® resuspension buffer prior to CTC enrichment using the ClearCell® FX system (Clearbridge BioMedics, Singapore), according to manufacturer’s instruction.
The ClearCell® FX system is an automated CTC enrichment system driven by the CTChip® FR1, a microfluidic biochip to isolate CTCs based on size, deformability and inertia. The isolation principle takes advantage of the inherent Dean vortex flows present in curvilinear channels for CTC enrichment, termed Dean Flow Fractionation (DFF) [43]. The enriched CTC sample output was equally divided into four portions.
Fluorescent in-situ hybridization
Four µm thick FFPE tumor tissue sections were mounted on positively charged slides and deparaffinized.
FISH was subsequently performed using the US FDA-approved Vysis ALK Break Apart FISH Probe Kit (Abbott Molecular, Abbott Park, Des Plaines, IL, USA). The 5’ ALK probe was labeled with SpectrumGreen™ (G) and the 3’ ALK probe with SpectrumOrange™ (R). ALK FISH for FFPE tissues were considered positive if at least 15 % of the tumor cells showed abnormal break apart signals as detailed in the IVD Vysis ALK Break Apart FISH Probe Kit and by Camidge et al. [33]. A cell is interpreted as having a split pattern (ALK-positive) when the 5’ (G) and 3’ (R) signals are separated by two or more signal diameters. Cells lacking both fluorescent signals were not evaluated.
One portion (one-quarter) of the enriched CTCs was fixed in Shandon CytoSpinTM Collection Fluid (ThermoFisher Scientific, USA) overnight at 4°C. The sample was deposited onto positively charged glass slides by cytospin (800 rpm, 5 mins). All the cells on the slides were analyzed for the ALK break-apart signal at 1000X magnification. The scorers analyzing the ALK break-apart signal on CTCs were blinded to the ALK rearrangement patterns on the tumor samples, as well as whether the cell isolated was from patients or healthy controls.
The remaining three portions of the enriched CTCs were stored at 4°C under validated conditions for future molecular testing.
Data analysis
Statistical analyses of the data were performed in GraphPad Prism version 5.00 (GraphPad Software, San Diego, CA, USA). A non-parametric two-tailed, t-test (Mann-Whitney) was used for computing statistical significances. p value of less than 0.05 were considered significant.
Acknowledgments
The authors wish to acknowledge Clearbridge BioMedics, Singapore for providing the CTC capturing technology and technical support. The authors would also like to express sincere gratitude to patients and healthy donors for participating in this study.
FUNDING
This study was supported by National Medical Research Council grants (CSA040/2012 and CS-IRG1225/2009).
CONFLICTS OF INTEREST
Chwee Teck Lim is an advisor with Clearbridge Biomedics, Singapore. Man Chun Leong is an employee of Clearbridge Biomedics, Singapore. All other authors declare that they do not have any conflicts.
Editorial note
This paper has been accepted based in part on peer-review conducted by another journal and the authors’ response and revisions as well as expedited peer-review in Oncotarget.
References
1. American Cancer Society. Cancer Facts & Figures 2015. Atlanta: American Cancer Society; 2015. Accessed on 12 October 2015. Available from: http://www.cancer.org/research/cancerfactsstatistics/cancerfactsfigures2015/index
2. Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA. Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc. 2008; 83: 584-594.
3. Non-small cell lung cancer survival rate by stage. Am. Cancer Soc. Atlanta; 2015. Accessed on 14 March 2015. Available from: http://www.cancer.org/cancer/lungcancer-non-smallcell/detailedguide/non-small-cell-lung-cancer-survival-rates
4. Lira ME, Kim TM, Huang D, Deng S, Koh Y, Jang B, Go H, Lee SH, Chung DH, Kim WH, Schoenmakers EFPM, Choi Y La, Park K, et al. Multiplexed gene expression and fusion transcript analysis to detect ALK fusions in lung cancer. J Mol Diagn. 2013; 15: 51-61.
5. Paez JG, Jänne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004; 304: 1497-500.
6. Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S, Fujiwara S, Watanabe H, Kurashina K, Hatanaka H, Bando M, Ohno S, Ishikawa Y, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature. 2007; 448: 561-566.
7. Sequist L V, Lynch TJ. EGFR tyrosine kinase inhibitors in lung cancer: an evolving story. Annu Rev Med. 2008; 59: 429-442.
8. Roskoski R Jr. Anaplastic lymphoma kinase (ALK): structure, oncogenic activation, and pharmacological inhibition. Pharmacol Res. 2013; 68: 68-94.
9. Solomon B, Varella-Garcia M, Camidge DR. ALK gene rearrangements: a new therapeutic target in a molecularly defined subset of non-small cell lung cancer. J Thorac Oncol. 2009; 4: 1450-1454.
10. Forde PM, Rudin CM. Crizotinib in the treatment of non-small-cell lung cancer. Expert Opin Pharmacother. 2012; 13: 1195-1201.
11. Shaw AT, Kim D-W, Mehra R, Tan DSW, Felip E, Chow LQM, Camidge DR, Vansteenkiste J, Sharma S, De Pas T, Riely GJ, Solomon BJ, Wolf J, et al. Ceritinib in ALK-rearranged non-small-cell lung cancer. N Engl J Med. 2014; 370: 1189-1197.
12. Lindeman NI, Cagle PT, Beasley MB, Chitale DA, Dacic S, Giaccone G, Jenkins RB, Kwiatkowski DJ, Saldivar JS, Squire J, Thunnissen E, Ladanyi M. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J Thorac Oncol. 2013; 8: 823-859.
13. Reck M, Hermes A, Tan E-H, Felip E, Klughammer B, Baselga J. Tissue sampling in lung cancer: a review in light of the MERIT experience. Lung Cancer. 2011; 74: 1-6.
14. Chouaid C, Dujon C, Do P, Monnet I, Madroszyk A, Le Caer H, Auliac JB, Berard H, Thomas P, Lena H, Robinet G, Baize N, Bizieux-Thaminy A, et al. Feasibility and clinical impact of re-biopsy in advanced non small-cell lung cancer: a prospective multicenter study in a real-world setting (GFPC study 12-01). Lung Cancer. 2014; 86: 170-173.
15. De Bruin E, McGranahan N, Yates L, Jamal-Hanjani M, Salm M, Mitter R, Shafi S, Murugaesu N, Rowan A, Gerlinger M, Wedge D, Horswell S, Varela I, et al. Intratumor heterogeneity in non-small cell lung cancer inferred by multi-region exome sequencing. [abstract]. In: Proceedings of the 105th Annual Meeting of the American Association for Cancer Research; 2014 Apr 5-9; San Diego, CA. Philadelphia (PA): AACR; Cancer Res 2014; 74(19 Suppl): Abstract nr 983.
16. Fidler IJ. The pathogenesis of cancer metastasis: the “seed and soil” hypothesis revisited. Nat Rev Cancer. 2003; 3: 453-458.
17. Mendoza M, Khanna C. Revisiting the seed and soil in cancer metastasis. Int J Biochem Cell Biol. 2009; 41: 1452-1462.
18. Alix-Panabier̀es C, Pantel K. Circulating tumor cells: liquid biopsy of cancer. Clin Chem. 2013; 59: 110-118.
19. Khoo BL, Warkiani ME, Tan DSW, Bhagat AAS, Irwin D, Lau DP, Lim AST, Lim KH, Krisna SS, Lim WT, Yap YS, Lee SC, Soo RA, et al. Clinical validation of an ultra high-throughput spiral microfluidics for the detection and enrichment of viable circulating tumor cells. PLoS One. 2014; 9: e99409.
20. Bednarz-Knoll N, Alix-Panabières C, Pantel K. Clinical relevance and biology of circulating tumor cells. Breast Cancer Res. 2011; 13: 228.
21. Müller V, Riethdorf S, Rack B, Janni W, Fasching PA, Solomayer E, Aktas B, Kasimir-Bauer S, Pantel K, Fehm T. Prognostic impact of circulating tumor cells assessed with the CellSearch SystemTM and AdnaTest BreastTM in metastatic breast cancer patients: the DETECT study. Breast Cancer Res. 2012; 14: R118.
22. Pailler E, Adam J, Barthélémy A, Oulhen M, Auger N, Valent A, Borget I, Planchard D, Taylor M, André F, Soria JC, Vielh P, Besse B, et al. Detection of circulating tumor cells harboring a unique ALK rearrangement in ALK-positive non-small-cell lung cancer. J Clin Oncol. 2013; 31: 2273-2281.
23. Ilie M, Long E, Butori C, Hofman V, Coelle C, Mauro V, Zahaf K, Marquette CH, Mouroux J, Paterlini-Bréchot P, Hofman P. ALK-gene rearrangement: a comparative analysis on circulating tumour cells and tumour tissue from patients with lung adenocarcinoma. Ann Oncol. 2012; 23: 2907-2913.
24. Naoe M, Ogawa Y, Morita J, Omori K, Takeshita K, Shichijyo T, Okumura T, Igarashi A, Yanaihara A, Iwamoto S, Fukagai T, Miyazaki A, Yoshida H. Detection of circulating urothelial cancer cells in the blood using the CellSearch system. Cancer. 2007; 109: 1439-1445.
25. Vona G, Sabile A, Louha M, Sitruk V, Romana S, Schütze K, Capron F, Franco D, Pazzagli M, Vekemans M, Lacour B, Bréchot C, Paterlini-Bréchot P. Isolation by size of epithelial tumor cells. Am J Pathol. 2000; 156: 57-63.
26. Yu M, Stott S, Toner M, Maheswaran S, Haber DA. Circulating tumor cells: approaches to isolation and characterization. J Cell Biol. 2011; 192: 373-382.
27. Chinen LTD, de Carvalho FM, Rocha BMM, Aguiar CM, Abdallah EA, Campanha D, Mingues NB, de Oliveira TB, Maciel MS, Cervantes GM, Dettino ALA, Soares FA, Paterlini-Bréchot P, et al. Cytokeratin-based CTC counting unrelated to clinical follow up. J Thorac Dis. 2013; 5: 593-599.
28. Hofman V, Ilie MI, Long E, Selva E, Bonnetaud C, Molina T, Vénissac N, Mouroux J, Vielh P, Hofman P. Detection of circulating tumor cells as a prognostic factor in patients undergoing radical surgery for non-small-cell lung carcinoma: comparison of the efficacy of the CellSearch AssayTM and the isolation by size of epithelial tumor cell method. Int J Cancer. 2011; 129: 1651-1660.
29. Yang P, Kulig K, Boland JM, Erickson-Johnson MR, Oliveira AM, Wampfler J, Jatoi A, Deschamps C, Marks R, Fortner C, Stoddard S, Nichols F, Molina J, et al. Worse disease-free survival in never-smokers with ALK+ lung adenocarcinoma. J Thorac Oncol. 2012; 7: 90-97.
30. Srodon M, Westra WH. Immunohistochemical staining for thyroid transcription factor-1: a helpful aid in discerning primary site of tumor origin in patients with brain metastases. Hum Pathol. 2002; 33: 642-645.
31. Yoshida A, Tsuta K, Nitta H, Hatanaka Y, Asamura H, Sekine I, Grogan TM, Fukayama M, Shibata T, Furuta K, Kohno T, Tsuda H. Bright-field dual-color chromogenic in situ hybridization for diagnosing echinoderm microtubule-associated protein-like 4-anaplastic lymphoma kinase-positive lung adenocarcinomas. J Thorac Oncol. 2011; 6: 1677-1686.
32. Yoshida, A, Varella-Garcia M. Fluorescence in situ Hybridization (FISH). In: Tsao, Ming Sound, Hirsch, Fred R YY, editor. IASLC ATLAS ALK Test Lung Cancer. 10th ed. Internation Association for the study of Lung Cancer, Aurora,Co; 2013. page 23-4. Accessed on 12 October 2015. Available from: https://www.iaslc.org/publications/iaslc-atlas-alk-testing-lung-cancer
33. Camidge DR, Kono SA, Flacco A, Tan AC, Doebele RC, Zhou Q, Crino L, Franklin WA, Varella-Garcia M. Optimizing the detection of lung cancer patients harboring anaplastic lymphoma kinase (ALK) gene rearrangements potentially suitable for ALK inhibitor treatment. Clin Cancer Res. 2010; 16: 5581-5590.
34. Klempner SJ, Cohen DW, Costa DB. ALK translocation in non-small cell lung cancer with adenocarcinoma and squamous cell carcinoma markers. J Thorac Oncol. 2011; 6: 1439-1440.
35. Qu Y, Che N, Zhao D, Zhang C, Su D, Zhou L, Zhang L, Wang C, Zhang H, Wei L. The clinicopathological significance of ALK rearrangements and KRAS and EGFR mutations in primary pulmonary mucinous adenocarcinoma. Tumour Biol. 2015; 36: 6417-6424.
36. Nishino M, Klepeis VE, Yeap BY, Bergethon K, Morales-Oyarvide V, Dias-Santagata D, Yagi Y, Mark EJ, Iafrate AJ, Mino-Kenudson M. Histologic and cytomorphologic features of ALK-rearranged lung adenocarcinomas. Mod Pathol. 2012; 25: 1462-1472.
37. Shaw AT, Yeap BY, Mino-Kenudson M, Digumarthy SR, Costa DB, Heist RS, Solomon B, Stubbs H, Admane S, McDermott U, Settleman J, Kobayashi S, Mark EJ, et al. Clinical features and outcome of patients with non-small-cell lung cancer who harbor EML4-ALK. J Clin Oncol. 2009; 27: 4247-4253.
38. Faugeroux V, Pailler E, Auger N, Taylor M, Farace F. Clinical utility of circulating tumor cells in ALK-positive non-small-cell lung cancer. Front Oncol. 2014; 4: 281.
39. Kim H, Jang SJ, Chung DH, Yoo SB, Sun P, Jin Y, Nam KH, Paik JH, Chung JH. A comprehensive comparative analysis of the histomorphological features of ALK-rearranged lung adenocarcinoma based on driver oncogene mutations: frequent expression of epithelial-mesenchymal transition markers than other genotype. PLoS One. 2013; 8: e76999.
40. Doebele RC, Pilling AB, Aisner DL, Kutateladze TG, Le AT, Weickhardt AJ, Kondo KL, Linderman DJ, Heasley LE, Franklin WA, Varella-Garcia M, Camidge DR. Mechanisms of resistance to crizotinib in patients with ALK gene rearranged non-small cell lung cancer. Clin Cancer Res. 2012; 18: 1472-1482.
41. Katayama R, Shaw AT, Khan TM, Mino-Kenudson M, Solomon BJ, Halmos B, Jessop NA, Wain JC, Yeo AT, Benes C, Drew L, Saeh JC, Crosby K, et al. Mechanisms of acquired crizotinib resistance in ALK-rearranged lung cancers. Sci Transl Med. 2012; 4: 120ra17.
42. Travis WD, Brambilla E, Burke, AP, Marx A, Nicholson A. Lyon: IARC Press; 2015 World Health Organization Classification of Tumours of the Lung, Pleura, Thymus and Heart.
43. Hou HW, Warkiani ME, Khoo BL, Li ZR, Soo R a, Tan DS-W, Lim W-T, Han J, Bhagat AAS, Lim CT. Isolation and retrieval of circulating tumor cells using centrifugal forces. Sci Rep. 2013; 3: 1259.

