Introduction
Squamous cell carcinoma of the lung (LSCC) is the second most common histological subtype of non-small cell lung cancer (NSCLC) having smoking habit as the most recognized risk factor [1, 2].
LSCC currently represents 30% of lung cancers in men and 20% in women [3]. Its incidence has been decreasing worldwide since 1979 even though an apparent increase in LSCC incidence has been recently registered, probably as a consequence of a widespread use of immunohistochemistry for classifying lung cancer histotypes and the subsequent decrease in not otherwise specified NSCLC diagnosis [4]. Squamous cell carcinoma (SCC) is the most frequent histotype among the head and neck tumors. In the latest ten years a substantial global variation in epidemiology of head and neck squamous cell carcinoma (HNSCC) has been observed: laryngeal and oral SCC incidences are strongly associated with changes in smoking habit, while oropharyngeal cancers are increasing especially in young people and in some European and Asian nations mainly due to human papilloma virus (HPV) infection [5, 6]. Many studies have also detected HPV in NSCLC, ranging widely from 0 to 78%, thus suggesting a potential carcinogenic role of HPV infection in at least a subset of lung cancer cases. However, recent comprehensive analyses reported that the variability in HPV detection rates in lung cancer (especially adenocarcinoma and LSCC) could be a function of the geographical origin of the studies. HPV detection was more prevalent in Asia and South America compared to Europe, Australia, and North America [7, 8]. A recent study by Chang et al. supports this finding showing no HPV detection in a North American population of NSCLC [9].
LSCC and HNSCC may present as synchronous or metachronous tumors, also in relationship with common pathogenesis [10]. They also share morphological microscopic features, making differential diagnosis on metastatic sites highly challenging. Currently, the differentiation between distant metastasis and second primary tumors is made on the basis of clinical criteria. Molecular characterization could be useful in this setting, as shown in a small study in which loss of heterozygosity and TP53 mutation analysis have been successfully used for differential diagnosis [11].
Molecular characterization is key in order to develop targeted therapies for solid tumors. Recently, wide genotyping of SCC of the lung and the head and neck has been performed, highlighting specific molecular traits. As far as head and neck is concerned, a different genetic profile according to HPV infection has been depicted. The molecular characterization of these diseases could potentially open new perspectives of tailored treatments for specific subsets of patients.
Common molecular alterations in smoke-related squamous cell carcinomas
Genetic alterations and common pathogenesis
LSCC and HNSCC are clinically and genetically heterogeneous diseases, but they share a number of molecular characteristics suggesting similar biology and pathogenesis.
The two diseases recognize smoke as a major risk factor, thus implying common elements both in pathogenesis and in the pattern of genetic alterations. Tobacco-related carcinogenesis is a typical model of complex multi-step carcinogenesis characterized by larger numbers and complexity of DNA alterations in these two tumors compared to other solid malignancies. Tobacco smoke is implicated in LSCC and HNSCC pathogenesis in different ways. In addition to the well-known mechanism of DNA adducts induction, several other mechanisms have been described, with a relevant role also for epigenetic alterations and micro-RNA deregulation (Figure 1). More than sixty known carcinogens have been detected in cigarette smoke; among these, tobacco-specific N-nitrosamines, polycyclic aromatic hydrocarbons, and aromatic amines are currently recognized as the strongest tumorigenic substances [12]. The majority of the smoke-related carcinogens require metabolic activation to react with DNA and cause DNA adducts formation (Figure 1). Transitions and transversions at CpG sites are typically found in genes commonly altered in smoking patients, such as KRAS, TP53 and RB [13-15]. Interestingly, the pattern of mutations in HNSCC is different according to HPV positivity. HPV-negative tumors, where carcinogenesis is mainly smoke-related, show transversions at CpG sites more frequently than HPV-positive tumors, known to be led by virus-mediated carcinogenesis [12]. On the contrary, mutations at Tp*Cp sites are more common in the latter group. More recent studies demonstrated that nicotine and its oncogenic derivatives are unable to initiate tumorigenesis, but they promote the subsequent steps of carcinogenesis: tumor growth, cell proliferation, migration, invasion, evasion of apoptosis, epithelial-to-mesenchymal transition, tumor angiogenesis and immune-response down-regulation (Figure 1). These effects are mediated especially by the binding to the nicotinic acetylcholine receptors (nAChR) and subsequent activation of multiple signaling cascades such as JAK/STAT, Ras/Raf/MAPK, and PI3K/AKT. In addition, nicotine and nitrosamines have been shown to induce cell-cycle progression, by the overexpression of cyclins and decreased levels of cyclin-dependent kinase (Cdk) inhibitors. These mechanisms mediated by tobacco components on already initiated tumors could also explain the increased probability of resistance to chemotherapy and radiotherapy reported in patients still smoking during cancer treatment [16].
The major genomic determinants related to smoke exposure concern mainly tumor suppressor genes, including TP53, CDKN2A and RB1 [13]. In particular, about 85% of smoke-related LSCC and HNSCC present loss of function in the tumor suppressor gene TP53 while loss of function of CDKN2A is found in about 20% of cases [12, 17]. A large amount of alterations affects genes directly involved in squamous cell differentiation implicating their dysregulation as major driver of SCC carcinogenesis. Typical molecular features of squamous cell differentiation are loss of chromosome 3p and gain of chromosome 3q [18-20]. Also inactivating mutations of the NOTCH family members genes have been associated with squamous differentiation [21]. In addition, LSCC and HNSCC are characterized by frequent alterations of oncogenic pathways typically involved in many solid tumors such as the PI3K/AKT, the NFKB1 and the Hippo pathway [12, 22-24]. Multiplatform genomic and proteomic analysis confirmed that LSCC and HNSCC share common molecular features and in particular they were classified as a unique subtype together with a subset of bladder cancer, another solid tumor typically related to smoke exposure [25]. A more recent work identified a set of 8 genes altered with significantly different frequencies in SCC originating from different organs compared to non-SCC, suggesting the existence of a “squamousness” signature [26]. The Cancer Genome Atlas (TCGA) published the results of the comprehensive multi-platform genomic characterization of a large cohort of LSCC and HNSCC in 2012 and 2015, respectively. These data unveiled main molecular alterations in the two diseases and provided further compelling evidence of the similarity of the two neoplasms in terms of genetic and epigenetic landscape [12, 17].
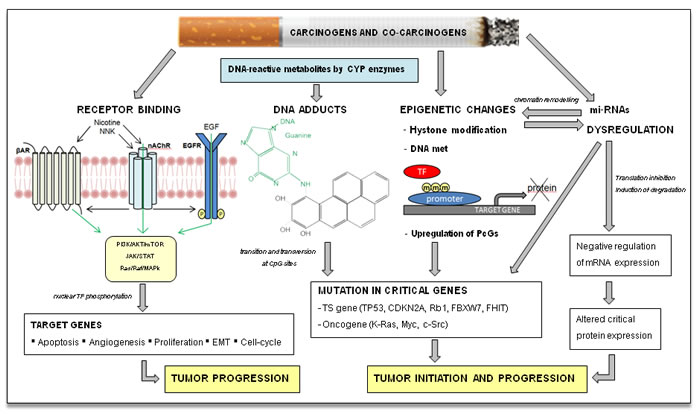
Figure 1: Tobacco smoke carcinogenesis. The main recognized mechanism of tobacco initiation of carcinogenesis is the formation of DNA adducts by polycyclic aromatic hydrocarbons and aromatic amines, with consequent deregulation of critical oncogenes and/or tumor suppressor (TS) genes. Tobacco smoke can influence protein expression of squamous cell cancer patients through induction of reversible epigenetic changes and miRNAs dysregulation; these pathways are also inter-related. Nicotine and its oncogenic derivatives are unable to initiate carcinogenesis, but can promote tumor growth through activation of multiple kinase cascade, acting directly on nicotinic acetylcholine receptor (nAChR) or indirectly on parallel cell surface receptors. βAR, beta adrenergic receptor kinase; CDKN2A, cyclin-dependent kinase inhibitor 2A; CpG, C-phosphate-G; c-Src, cellular Src kinase; CYP, cytochrome p450; FBXW7, F-box and WD repeat domain containing 7; FHIT, fragile histidine triad; m, methyl-CpG; Myc, myelocytomatosis oncogene; nAchR, Nicotinic acetylcholine receptor; NNK, nicotine-derived nitrosamine ketone; PcGs, polycomb-group protein; Rb1, retinoblastoma1; TF, transcription factor.
Molecular subtypes classification
Based on genomic characterization, different molecular subtypes of LSCC and HNSCC have been defined and validated by the evaluation of the dominant gene expression pattern (Figure 2). The milestone of this kind of study is that expression subtypes should be reproducible from a statistical and biological point of view, associated with different clinical pattern and corresponding to different cellular type with different biological behavior, thus paving the way to further studies on different pathogenesis and therapeutic approach. The four mRNA expression subtypes of LSCC are named classical, basal, secretory, and primitive [27], while four different classifications have been proposed for HNSCC [28-31] (Figure 2).
This review focuses on the molecular subtypes associated with smoke exposure and, in the paragraph below, on those related to HPV infection.
In both malignancies and according to all classification of HNSCC, the classical subtype presents the highest percentage of current smokers and it does not show appreciable similarity to any specific normal cell type. This molecular subgroup of SCC is characterized by the highest overall methylation and chromosomal instability, as well as overexpression of genes involved in xenobiotic metabolism, indicating a sustained response to cigarette smoke. Among HNSCC, the classical subtype occurs especially in laryngeal tumors, consistently with the correlation with tobacco exposure [12].
Moreover, broad amplification of chromosome 3q, widely recognized as associated with SCC, particularly defines the classical subtype and results in the overexpression of three oncogenes: SOX2, TP63 and PIK3CA [32]. SOX2 gene is a key transcription factor that promotes normal squamous differentiation and it is overexpressed in about 20% of LSCC. In SCC, it works as a lineage-survival oncogene, probably even activating additional pathways controlled in early pluripotent cells [33]. The transcription factor TP63 gene is expressed as multiple isoforms with different functions, including the truncated amino-deleted isoform and the full length one. The short deltaN isoform (also called p40) is known to be the most commonly expressed isoform in squamous tumors, and it functions as an oncogene, promoting growth and survival by competing for p53 binding sites [20]. As a matter of fact, p63 overexpression, evaluated by immunohistochemistry, is one of the main markers used by the pathologists to define squamous cell differentiation in addition to specific morphology [34]. The PIK3CA gene is a key regulator of cell growth, metabolism, and survival and it is responsible for PI3K activity and phosphorylated Akt expression. PIK3CA activating mutations and copy number gains occur independently of each other, and either molecular event may be sufficient to drive the cell towards tumorigenesis by promoting cell growth and invasion [35, 36]. In SCC, alterations of the NFR2-KEAP1/CUL3 pathway are also frequently reported. In different studies about LSCC and HNSCC, mutations and copy number alterations of NRF2 and KEAP1 and/or deletion or mutation of CUL3 have been detected, especially in the classical subtype [12, 17, 37]. NRF2 and KEAP1/CUL3 encode for proteins that bind to each other and have been shown to promote the cell response to oxidative damage. Indeed, the transcription factor Nrf2 mediates gene induction of numerous cytoprotective enzymes in response to environmental and endogenously derived oxidative/electrophilic agents. On the other hand, under physiological conditions, low cellular concentrations of Nrf2 are maintained by proteasomal degradation through a Cullin3-Keap1 system. Hence, mutations in KEAP1 and CUL3 are mutually exclusive with mutations in NRF2, and each of these leads to a constitutive activation of Nrf2, promoting cancer cell survival [38-40].
Overall, the classical subtype is enriched in genes involved in regulation of oxidant stress and glutathione metabolism, xenobiotic metabolism, and secretion, such as GPX2, ALDH3A1, AKR1C1/3, TXNRD1, and GSTM3 [27, 29, 30].
Even though the classical subtype can be clearly associated with smoke exposure and genetic alterations associated with tobacco-related damage, the presence of different patterns of genetic alterations does not exclude smoke exposure. On the contrary, also in the other molecular subtypes most of the patients were former smokers, although exhibiting molecular alterations in different pathways [28-31]. Figure 2 shows the characteristics of molecular subtypes in HNSCC according to different classifications, highlighting the subtypes corresponding to the LSCC classification.
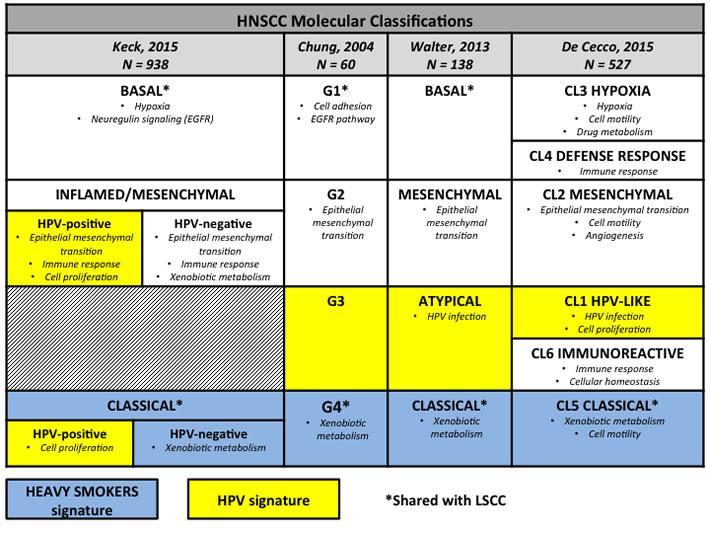
Figure 2: Molecular subtypes in head and neck squamous cell carcinomas. The figure summarizes the molecular subtypes of head and neck squamous cell carcinomas according to the four different molecular classifications available in the literature with the main dysregulated functional pathways for each subgroup. We highlighted molecular subtypes associated with human papillomavirus infection by yellow boxes, those associated with smoking habits by blue boxes, and we marked by * the subgroups corresponding to lung squamous cell carcinoma expression subtypes. HNSCC, head and neck squamous cell carcinoma; HPV human papillomavirus; LSCC, lung squamous cell carcinoma.
Squamous head and neck cancers in HPV-positive patients: a new molecular entity
Although HNSCC is widely viewed as comprised of two distinct clinical and biological entities, the genetic landscape of HPV-positive tumors remains unclear since data from large HPV-positive cohorts are still missing. The first study on molecular alterations in HNSCC including a large proportion of HPV-positive tumors (N = 51 samples, 42.5% of cases) has been published in 2015 [37]. This comprehensive genomic analysis confirmed the hypothesis of distinct mutational and copy-number profiles between HPV-positive and HPV-negative tumors, as suggested by previous studies on small HPV-positive series [12, 41, 42]. Some critical issues should be considered in discussing the results. First, a considerable proportion of HPV-positive patients reported significant smoking history. In addition, HPV-positive status confers a favorable prognosis, but also smoke exposure is independently associated with survival and a significantly increased risk of death is associated with each additional pack-year of tobacco smoke. The effect of tobacco on clinical outcome is similar among patients with HPV-positive tumor (hazard ratio, 1.01; 95% CI, 1.00 to 1.02) and HPV-negative cases (hazard ratio, 1.01; 95% CI: 1.0-1.03) [43]. This finding is likely to be related to the four-to-six-fold higher mutational burden detected in such HPV-positive patients compared to HPV-positive tumors without smoking history. These additional genetic alterations are linked to tobacco-related carcinogens, as confirmed by the presence of KRAS and TP53 mutations. Thus, HPV-positive tumors in smoking patients and HPV-negative tumors share molecular alterations such as 3q amplification, PI3K signaling [44], NOTCH aberrations [45] and SMAD signaling. However, HPV-positive tumors are more enriched in mutations/copy number variations in oncogenes compared to HPV-negative patients [46].
HPV infection is implicated in HNSCC tumorigenesis through different mechanisms. The best known element is the overexpression of viral E6-E7 proteins that act as oncoproteins promoting p53 or Rb degradation, activation of hTERT, neutralization of the inhibitory effect by CDK inhibitors on cell cycle; thus, HPV infection activates the process of carcinogenesis by deregulating fundamental cellular pathways [47]. Moreover, beyond the impairment of immune surveillance in order to promote viral persistence in HPV-infected cells, different immune evasion strategies are involved in HPV-driven tumors including the recruitment of immune cells with immunosuppressive properties [48].
HPV DNA is present in tumor cells both as integrated into the human genome and as episome, nonintegrated into the genome. A recent study evaluated the effect on carcinogenesis of the integration of the viral genome in 25 cases of HPV-positive HNSCC. The data suggest that, beyond the expression of viral oncoproteins E6 and E7, HPV promotes oncogenesis by the alteration of host genome at the sites of integration. The effects of viral DNA integration into human genome include disruption of tumor suppressor genes, oncogene amplifications, interchromosomal rearrangements and altered methylation. Examples of genes altered by viral DNA integration are RAD51B, leading to loss of DNA-repair function, and NR4A2, a nuclear transcription factor which acts as an oncogene, especially by inhibiting apoptosis [49].
Recently Gaykalova et al. confirmed a different underlying biology of HPV-positive and HPV-negative tumors by evaluating the activity of key transcription factors; their data suggested that HPV-positive and HPV-negative tumors show specifically different patterns of alterations in transcription factor pathways, including AP1, STATs, NF-κB and p53 [50].
The most commonly altered pathway in HPV-positive tumors is PI3K/AKT that is involved in regulating the signaling of different processes such as apoptosis, metabolism, cell proliferation, and cell growth. Mutations and/or copy number variations of the catalytic domain (PIK3CA) of PI3K are present in about 20-30% of cases, while, more rarely, mutations occur in other genes of the same pathway: PTEN, AKT1, PIK3R1, TSC1, and TSC2, thus making the pathway a potential therapeutic target. Other potentially druggable alterations of particular interest are FGFR2/3 mutations occurring in about 25% of HPV-positive cases, while in HPV-negative patients FGFR1 amplifications are more common [37].
Interestingly HPV-positive tumors, especially in never/light-smoker patients, are enriched also in alteration to DNA damage repair pathway including the genes BRCA1, ATM, BRCA1, BRCA2, FANCG, FANCA, and FANCD2 [37]. We can speculate that inefficient DNA repair capacity in these tumors may in part explain the increased sensitivity to chemotherapy and platinum-based treatment, specifically. Specific data on this issue are missing, anyway.
Moreover, wide genome analyses highlighted the higher presence of genetic alterations in immune-related genes (HLA-A and HLA-B) and this point may be directly related to viral tumorigenesis [12, 37].
Finally, two genes are exclusively altered in HPV-positive HNSCC: DDX3X, involved in RNA processing, and CYLD, a multifunctional deubiquitinase involved in negative regulation of NF-κB signaling [37]. In particular, CYLD correlates also with HPV-related cervical cancer, thus suggesting the idea of a specific role in virus-related carcinogenesis. In preclinical models, under hypoxic conditions, the HPV-encoded E6 protein promotes inactivation and degradation of the CYLD tumor suppressor, resulting in prolonged activation of NF-κB pathway [51].
In previous molecular classification HPV-related tumors have been mainly included in the atypical subgroup (Figure 2); however the overall small number of HPV-positive patients included in the analysis affected the final classification, probably impairing the identification of specific features in the HPV-related subgroup. The more recent molecular nomenclature of HNSCC by Keck et al. included also a large cohort of HPV-positive patients in order to provide a comprehensive overview of HPV-negative as well as HPV-positive HNSCC. According to this classification, two distinct subgroups of HPV-positive tumors fall in the classical and in the mesenchymal subtype, respectively (Figure 2). HPV-positive patients without smoking history share main molecular alterations with the HPV-negative counterpart with the exception of smoking-associated pathways. Indeed, in both subtypes around 80% of HPV-negative patients were heavy smokers versus 40% of HPV-positive patients. The classical subtype is characterized by the overexpression of cell-cycle genes and cell division cycle protein kinase (“proliferative signature”), while the main theme of the inflamed/mesenchymal subtype is the expression of immune response genes related to the enrichment of cytotoxic T-cell infiltration. This subgroup also shows the expression of mesenchymal genes and downregulation of epithelial differentiation markers while tumors are histologically poorly differentiated. These two HPV-related subtypes exhibit significant differences in terms of morphology and molecular pattern and, importantly, the mesenchymal subtype is characterized by a better survival compared to classical HPV-positive tumors. This finding may explain the clinical heterogeneity observed between HPV-positive patients with different response to treatment and highlights potential clinical application of molecular characterization and subclassification among HPV-positive HNSCC [30].
In the latest classification based on a meta-analysis approach combining multiple datasets, even if the HPV status was not available, one molecular cluster has been defined as HPV-like. This subgroup shows the up-regulation of genes related to HPV infection and cell proliferation with a significant enrichment in oropharyngeal cases and it is characterized by the best outcome in terms of prognosis compared to other clusters [31].
Squamous cell carcinoma of the lung in never-smoker patients: to be explored
Currently, few data are available concerning molecular profiling in LSCC of never smokers. LSCC in non-smokers is an exceptional finding and it is often supposed to be related to professional exposure and related carcinogenesis. Nevertheless, in high-volume thoracic surgery centers, LSCC cases are observed even in non-smokers without known professional exposure.
In the TCGA project, as well as in another cohort of East Asians patients [52], most patients (96%) reported a history of tobacco use and no conclusions may be drawn for non-smokers [17]. One of the few studies evaluating genomic alterations in different histological subtypes and according to smoking status has been conducted in Chinese lung cancer patients. The authors investigated a spectrum of driver genes, including EGFR, KRAS, c-Met, PIK3CA, BRAF, STK11, PTEN, EML4-ALK fusion gene, DDR2, and FGFR2. They found only EGFR (8.0%), c-Met (2.8%), and PIK3CA (2.6%) alterations in the non-smoker LSCC subgroup, without alterations of other genes analyzed. This finding suggests that this subgroup recognizes pathogenic mechanisms that differ from known driver gene alterations of lung adenocarcinoma and SCC of the smokers [53]. Consistently, another analysis on 185 Chinese patients confirmed the detection of EGFR mutations among LSCC tumors ranging from 6% to 17%, with high correlation with the absence of smoking exposure [54].
In another small series of Chinese LSCC patients, the authors evaluated targetable alterations according to gender. In this population, the percentage of smoking patients is significantly lower among females compared to males (5.3% versus 90%). However no significant difference in the mutational frequencies of EGFR, KRAS, PTEN, ALK, or FGFR1 have been observed between males and females. Only PIK3CA mutations were significantly less common in females, and thus in never smokers, than in males, but the small sample of this study (38 females and 40 males) limits the interpretation of the results [55].
In conclusion, mutational landscape and consequently underlying biology in never-smoker LSCC tumors are still largely unexplored. Probably, wide genome characterization of LSCC samples from non-smoker patients could provide interesting data and potentially open new therapeutic perspectives.
Molecular characterization and therapeutic perspectives
Targeted therapy
In the latest years several efforts have been made to introduce genomic tests and targeted therapies in cancer treatment [56-59]. However, SCC is a deadly disease for which platinum-based chemotherapy is still the mainstay of treatment. Currently, cetuximab, an anti-EGFR monoclonal antibody, is the only approved targeted compound for the management of HNSCC [60, 61]. Regarding LSCC, erlotinib is the only targeted treatment used in clinical practice and has been studied in patient population without specific molecular characterization; two different EGFR antibodies and another EGFR-HER2 tyrosine kinase inhibitor also have been investigated in phase III trials, here again in unselected patients [62-64].
The two main goals of comprehensive genomic characterization are to improve the knowledge on pathogenesis and to identify driver somatic alterations for which targeted therapies already exist or are under evaluation [65, 66]. As reported in Table 1, various targeted agents are in different stages of development on the basis of current genome-wide sequencing and copy number data.
Currently, the PI3K pathway is one of the most promising therapeutic targets for both malignancies, on the basis of the high rate of PIK3CA mutations and of the presence of alterations in other genes involved in this pathway. Active clinical trials evaluating PI3K/AKT/mTOR targeted agents, including PIK3CA and mTOR inhibitors, are ongoing in both diseases; however, no significant clinical benefit has been demonstrated to date, even though strong preclinical rationale is widely recognized [67]. Other frequently altered targets in SCC are members of the FGFR family, characterized by the amplification of FGFR1 or mutations of FGFR2/3, which might be targeted by specific agents in development through early phase clinical trials [68-70]. Furthermore, mutations in the DDR2 kinase gene in about 4% of LSCC have been documented; such mutations lead to cellular transformation making this tyrosine kinase gene a promising target [71]. Although DDR2 has not been a major focus of drug development efforts, a recent analysis showed that ABL kinase inhibitors such as imatinib, nilotinib, and dasatinib have activity against DDR2 [72] and potential benefit of these agents in this setting are under investigation.
On the basis of available preclinical data, the design of biomarker-driven, multi-arm, randomized trials is likely to be a promising approach to promote the efficient development of targeted therapies, although a recent study has highlighted that the rarity of driver mutations may limit the feasibility of this approach [73]. According to this strategy, the National Cancer Institute has designed a series of clinical studies on various tumors by selection of patients through the identification of actionable molecular alterations with next-generation sequencing (NGS) technologies; statistical analysis has been planned for deeming the experimental agent an improvement over standard therapy. Regarding LSCC, patients with advanced disease progressing after first-line therapy receive their treatment according to molecular profile resulted from a NGS panel of 250 selected genes. Standard second-line chemotherapy is compared to targeted agent in the presence of potentially druggable alterations or to immunotherapy in the absence of driver mutations [74].
In the present review, rather then listing all the predictive biomarkers suitable for the evaluation of new targeted compounds in the management of HNSCC and SCC, we would like to highlight some key issues. First, as reported above, tobacco-related SCC, especially LSCC and HPV-negative HNSCC, are genetically complex malignancies and it is unlikely that most cases may benefit from targeted monotherapy. The rationale for the development of targeted therapy lay on the evidence of tumor cells addicted to one or few genes for maintenance of the malignant phenotype; on the contrary, malignancies characterized by many genetic and epigenetic alterations may potentially benefit from multiple biological agents targeting different drivers. A new potential model for the development of new targeted treatment specifically concerns tumors predominantly enriched in tumor suppressor genes, in which a “synthetic lethality” approach could be promising [75]. According to this strategy, a synthetic lethal relationship is seen when a cancer specific gene mutation combined with the therapeutic targeting of another pathway results in the selective killing of mutated cancer cells without affecting the normal ones. The most exciting results of this approach are exemplified by the treatment with PARP inhibitors of BRCA1-2 mutated ovarian cancers. Tumor cells carrying the mutations of the tumor suppressor genes BRCA1 or BRCA2 are defective in homologous recombination, and thus are sensitive to the inhibition of PARP activity, another key pathway involved in DNA repair [76, 77]. Combination of chemotherapy and targeted therapy or immunotherapy represents an alternative promising strategy in this setting.
Another important issue is the availability of tumor tissue for pathological and molecular studies during the course of the disease. Locally-advanced HNSCC could receive concurrent chemo-radiation therapy or induction therapy and patients without clinical complete response could undergo salvage surgery; residual neck disease may be present in as many as 30-60% of patients after the completion of chemo-radiation treatment [78-81]. As the neoadjuvant setting in breast cancer [82], this condition represents a suitable setting for evaluating new compounds because of the possibility to analyze tumor tissue before and after treatment as well as to obtain and compare anatomical and functional imaging.
In HNSCC treatment, another key point to consider is HPV status; currently, the HPV-status is associated with an improved outcome but it has no current relevance in terms of clinical management [83]. The most recent molecular platforms showed no significant difference in terms of frequency of mutations between HPV-negative and HPV-positive tumors, but a different molecular profile [37, 46]. These emerging data also showed a higher frequency of oncogene alterations and a lack of common tumor suppressor gene mutations in HPV-positive patients, making them more suitable for targeted therapy experimental approach. More broadly, molecular subtypes identified in both HNSCC and LSCC according to gene-expression data could be useful to design clinical studies with new compounds. Recently, in order to investigate individualized treatment for patients carrying different molecular subtypes of LSCC, Wu D. and colleagues evaluated the response to 24 drugs at eight dosages among 17 cell lines including all four LSCC subtypes. In this drug-response experiment, the authors demonstrated that the majority of cell lines responded to five of the 24 drugs (Panobinostat, 17-AAG, Irinotecan, Topotecan, and Paclitaxel) and the secretive subtype was significantly less sensitive to the drugs tested probably because of lower proliferation score than other subtypes [84]. Also in the latest molecular classification of HNSCC, the authors tested the hypothesis that each molecular subtype might have specific sensitivity to different drugs and they demonstrated a statistically significant difference for patients belonging to different subgroups [31].
Immunotherapy
The chance to enhance immunological response of the host against tumors is at the basis of important advances in the treatment of solid tumors, from the successful story of immunotherapy in melanoma to the most recent success in NSCLC [85-87]. PD-1 is an immune checkpoint receptor expressed on activated T cells, dampening the immune response; PD-1 pathway blockade with monoclonal antibodies restores tumor-specific T cell-mediated immunity [88].
Recently the anti-PD-1 antibody nivolumab has been approved by Food and Drug Administration (FDA) and European Medicines Agency (EMA) for the treatment of previously treated metastatic LSCC according to the results of the phase III trial showing a significantly improvement in median overall survival: 9.2 months with nivolumab compared to 6.0 months with docetaxel (hazard ratio: 0.59; 95% CI: 0.44 to 0.79; P < 0.001), with prolonged persistence of clinical benefit in responders [89].
Molecular predictive markers of sensitivity to immunotherapy are still under investigation; in this trial nivolumab improved survival regardless of PD-L1 expression, while in non-squamous setting the expression of PDL-1, whatever cut-off used, demonstrated to be predictive of benefit from immunotherapy with respect to chemotherapy [90]. On the other hand, Herbst and colleagues observed that the responses to the anti-PD-L1 MPDL3280A were associated with high expression of PD-L1 especially by tumor-infiltrating immune cells [91]. Interestingly, most trials evaluating anti-PD-1 or anti-PD-L1 antibodies suggested that former and current smokers might preferentially benefit from immunotherapy [92]. This finding could be related to the higher overall mutational burden in these patients resulting in more tumor neoantigens and increased immunogenicity. Recently, whole-exome sequencing of NSCLC cases treated with the anti-PD1 pembrolizumab, revealed that higher non-synonymous mutation burden, higher neoantigen burden, and molecular smoking signature, characterized by high nucleotide transversion rate, molecular smoking signature (high nucleotide transversion rate) were associated with improved objective response, durable clinical benefit and progression-free survival [93]. Similarly, another study showed that tumors with genetic mismatch repair defects, which are associated with a high degree of mutational burden, respond better to PD-1 inhibitor compared with tumors proficient in mismatch repair. In this trial, clinical benefit was observed across tumors with mismatch repair deficiency, including cancer of the colon, stomach, uterus, duodenum, prostate, and bile ducts, providing compelling evidence that neoantigens in tumor with high mutational burden may represent a potential immune-target [94].
Similarly, several trials with immunotherapeutic agents are ongoing in HNSCC, as well as the analysis of predictive biomarkers, including HPV status [95]. To date, the most advanced results are from the KEYNOTE-012 expansion cohort study evaluating pembrolizumab in recurrent or metastatic HNSCC. The drug demonstrated a clinically meaningful overall response rate of 18.2% in this setting, in both HPV-positive and HPV-negative patients [96].
Molecular characterization according to the above discussed subtypes may also have a role in predicting the response to immunotherapy [27-31]; indeed, the molecular subtypes characterized by alterations in pathways involved in the immune response against cancer (Figure 2) could be suitable for new immunotherapeutic approaches. Supporting this idea, the inflamed phenotype signature, according to the classification by Keck et al., has shown to be a strong predictor of clinical benefit from anti-PD-1 treatment for HNSCC patients enrolled in KEYNOTE-012 study [96]. Moreover, therapeutic vaccines against HPV are under evaluation (NCT01493154) also in association with immunomodulatory therapies to increase sensitivity in poorly immunogenic tumors by restoring cytotoxic T cell activity [97].
Table 1: Targeted agents in clinical development (active phase II and III trials) in patients with lung squamous cell carcinoma and head and neck squamous cell carcinoma
Target |
SCC |
Frequency of genetic alteration |
Targeted drug |
Phase of development |
PIK3 |
LSCC |
16% |
LY3023414 GDC-0032 |
II II/III |
HNSCC |
16-56% |
BKM120 |
II |
|
mTOR |
LSCC |
Sirolimus LY3023414 AZD2014 (TORC 1/2) MLN0128 (TORC 1/2) |
I/II II II II |
|
CDK4/6 |
LSCC |
Abemaciclib |
II |
|
LSCC and HNSCC |
Palbociclib isethionate |
II/III II |
||
FGFR (Mut/Ampl) |
LSCC |
2-7% |
AZD4547 Dovitinib |
II/III II |
FGFR (Mut/Ampl) -VEGFR |
LSCC |
2-7% |
Nintedanib Lucitanib |
II II |
HNSCC |
1-24% |
Pazopanib |
II |
|
HGF/c-MET |
LSCC |
Rilotumumab Capmatinib |
II/III II |
|
ALK |
LSCC |
AP26113 |
II |
|
PARP |
LSCC |
Veliparib |
III |
|
EGFR (Mut/Ampl) |
LSCC |
9% |
Icotinib Nimotuzumab Necitumumab |
II II I/II |
HNSCC |
6-15% |
Afatinib Cetuximab plus dasatinib Erlotinib Panitumumab |
III II II II |
|
HER3 |
LSCC |
2% |
MM-121 |
II |
pan-HER |
HNSCC |
HM781-36B |
II |
|
XPO1 |
LSCC HNSCC |
Selinexor |
II |
|
Hsp27 |
LSCC |
Apatorsen |
II |
LSCC, lung squamous cell carcinoma; HNSCC, head and neck squamous cell carcinoma; Mut, mutation; Ampl, amplification
Conclusions
Molecular characterization of LSCC and HNSCC is one of the emerging burning issues in cancer translational research. On the basis of recent wide genetic characterization, the complexity and heterogeneity of genetic alterations in the two diseases represent a new fascinating biological model. The potential therapeutic implications stem from data of targeted therapy in other solid tumors, in particular lung adenocarcinomas, and from recent successful results of immunotherapy in LSCC. New molecularly-driven clinical trials of targeted therapy in selected subsets of patients, in parallel with the investigation about the association of conventional and targeted therapy or immunotherapy are warranted in order to test potential synergistic effects aiming to long-lasting disease control in advanced stage.
Conflicts of interest
None.
References
1. Maier H, Dietz A, Gewelke U, Heller WD and Weidauer H. Tobacco and alcohol and the risk of head and neck cancer. The Clinical investigator. 1992; 70:320-327.
2. Kenfield SA, Wei EK, Stampfer MJ, Rosner BA and Colditz GA. Comparison of aspects of smoking among the four histological types of lung cancer. Tobacco control. 2008; 17:198-204.
3. Lortet-Tieulent J, Soerjomataram I, Ferlay J, Rutherford M, Weiderpass E and Bray F. International trends in lung cancer incidence by histological subtype: adenocarcinoma stabilizing in men but still increasing in women. Lung cancer. 2014; 84:13-22.
4. Meza R, Meernik C, Jeon J and Cote ML. Lung cancer incidence trends by gender, race and histology in the United States, 1973-2010. PloS one. 2015; 10:e0121323.
5. Chaturvedi AK, Engels EA, Anderson WF and Gillison ML. Incidence trends for human papillomavirus-related and -unrelated oral squamous cell carcinomas in the United States. Journal of clinical oncology. 2008; 26:612-619.
6. Zaravinos A. An updated overview of HPV-associated head and neck carcinomas. Oncotarget. 2014; 5:3956-3969. doi: 10.18632/oncotarget.1934.
7. Syrjanen K. Detection of human papillomavirus in lung cancer: systematic review and meta-analysis. Anticancer research. 2012; 32:3235-3250.
8. Ragin C, Obikoya-Malomo M, Kim S, Chen Z, Flores-Obando R, Gibbs D, Koriyama C, Aguayo F, Koshiol J, Caporaso NE, Carpagnano GE, Ciotti M, Dosaka-Akita H, Fukayama M, Goto A, Spandidos DA, et al. HPV-associated lung cancers: an international pooled analysis. Carcinogenesis. 2014; 35:1267-1275.
9. Chang SY, Keeney M, Law M, Donovan J, Aubry MC and Garcia J. Detection of human papillomavirus in non-small cell carcinoma of the lung. Human pathology. 2015; 46:1592-1597.
10. Griffioen GH, Louie AV, de Bree R, Smit EF, Paul MA, Slotman BJ, Leemans CR and Senan S. Second primary lung cancers following a diagnosis of primary head and neck cancer. Lung cancer. 2015; 88:94-99.
11. Geurts TW, van Velthuysen ML, Broekman F, van Huysduynen TH, van den Brekel MW, van Zandwijk N, van Tinteren H, Nederlof P, Balm AJ and Brakenhoff RH. Differential diagnosis of pulmonary carcinoma following head and neck cancer by genetic analysis. Clinical cancer research. 2009; 15:980-985.
12. Cancer Genome Atlas N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015; 517:576-582.
13. Hecht SS. Tobacco carcinogens, their biomarkers and tobacco-induced cancer. Nature reviews Cancer. 2003; 3:733-744.
14. Pfeifer GP, Denissenko MF, Olivier M, Tretyakova N, Hecht SS and Hainaut P. Tobacco smoke carcinogens, DNA damage and p53 mutations in smoking-associated cancers. Oncogene. 2002; 21:7435-7451.
15. Riely GJ, Marks J and Pao W. KRAS mutations in non-small cell lung cancer. Proceedings of the American Thoracic Society. 2009; 6:201-205.
16. Schaal C and Chellappan SP. Nicotine-mediated cell proliferation and tumor progression in smoking-related cancers. Molecular cancer research : MCR. 2014; 12:14-23.
17. Cancer Genome Atlas Research N. Comprehensive genomic characterization of squamous cell lung cancers. Nature. 2012; 489:519-525.
18. Patmore HS, Cawkwell L, Stafford ND and Greenman J. Unraveling the chromosomal aberrations of head and neck squamous cell carcinoma: a review. Annals of surgical oncology. 2005; 12:831-842.
19. Singh B, Gogineni SK, Sacks PG, Shaha AR, Shah JP, Stoffel A and Rao PH. Molecular cytogenetic characterization of head and neck squamous cell carcinoma and refinement of 3q amplification. Cancer research. 2001; 61:4506-4513.
20. Massion PP, Taflan PM, Jamshedur Rahman SM, Yildiz P, Shyr Y, Edgerton ME, Westfall MD, Roberts JR, Pietenpol JA, Carbone DP and Gonzalez AL. Significance of p63 amplification and overexpression in lung cancer development and prognosis. Cancer research. 2003; 63:7113-7121.
21. Wang NJ, Sanborn Z, Arnett KL, Bayston LJ, Liao W, Proby CM, Leigh IM, Collisson EA, Gordon PB, Jakkula L, Pennypacker S, Zou Y, Sharma M, North JP, Vemula SS, Mauro TM, et al. Loss-of-function mutations in Notch receptors in cutaneous and lung squamous cell carcinoma. Proceedings of the National Academy of Sciences of the United States of America. 2011; 108:17761-17766.
22. Harvey KF, Zhang X and Thomas DM. The Hippo pathway and human cancer. Nature reviews Cancer. 2013; 13:246-257.
23. Fresno Vara JA, Casado E, de Castro J, Cejas P, Belda-Iniesta C and Gonzalez-Baron M. PI3K/Akt signalling pathway and cancer. Cancer treatment reviews. 2004; 30:193-204.
24. Van Waes C. Nuclear factor-kappaB in development, prevention, and therapy of cancer. Clinical cancer research. 2007; 13:1076-1082.
25. Hoadley KA, Yau C, Wolf DM, Cherniack AD, Tamborero D, Ng S, Leiserson MD, Niu B, McLellan MD, Uzunangelov V, Zhang J, Kandoth C, Akbani R, Shen H, Omberg L, Chu A, et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell. 2014; 158:929-944.
26. Schwaederle M, Elkin SK, Tomson BN, Carter JL and Kurzrock R. Squamousness: Next-generation sequencing reveals shared molecular features across squamous tumor types. Cell cycle. 2015; 14:2355-2361.
27. Wilkerson MD, Yin X, Hoadley KA, Liu Y, Hayward MC, Cabanski CR, Muldrew K, Miller CR, Randell SH, Socinski MA, Parsons AM, Funkhouser WK, Lee CB, Roberts PJ, Thorne L, Bernard PS, et al. Lung squamous cell carcinoma mRNA expression subtypes are reproducible, clinically important, and correspond to normal cell types. Clinical cancer research. 2010; 16:4864-4875.
28. Chung CH, Parker JS, Karaca G, Wu J, Funkhouser WK, Moore D, Butterfoss D, Xiang D, Zanation A, Yin X, Shockley WW, Weissler MC, Dressler LG, Shores CG, Yarbrough WG and Perou CM. Molecular classification of head and neck squamous cell carcinomas using patterns of gene expression. Cancer cell. 2004; 5:489-500.
29. Walter V, Yin X, Wilkerson MD, Cabanski CR, Zhao N, Du Y, Ang MK, Hayward MC, Salazar AH, Hoadley KA, Fritchie K, Sailey CJ, Weissler MC, Shockley WW, Zanation AM, Hackman T, et al. Molecular subtypes in head and neck cancer exhibit distinct patterns of chromosomal gain and loss of canonical cancer genes. PloS one. 2013; 8:e56823.
30. Keck MK, Zuo Z, Khattri A, Stricker TP, Brown CD, Imanguli M, Rieke D, Endhardt K, Fang P, Bragelmann J, DeBoer R, El-Dinali M, Aktolga S, Lei Z, Tan P, Rozen SG, et al. Integrative analysis of head and neck cancer identifies two biologically distinct HPV and three non-HPV subtypes. Clinical cancer research. 2015; 21:870-881.
31. De Cecco L, Nicolau M, Giannoccaro M, Daidone MG, Bossi P, Locati L, Licitra L and Canevari S. Head and neck cancer subtypes with biological and clinical relevance: Meta-analysis of gene-expression data. Oncotarget. 2015; 6:9627-9642. doi: 10.18632/oncotarget.3301.
32. Qian J and Massion PP. Role of chromosome 3q amplification in lung cancer. Journal of thoracic oncology. 2008; 3:212-215.
33. Bass AJ, Watanabe H, Mermel CH, Yu S, Perner S, Verhaak RG, Kim SY, Wardwell L, Tamayo P, Gat-Viks I, Ramos AH, Woo MS, Weir BA, Getz G, Beroukhim R, O’Kelly M, et al. SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nature genetics. 2009; 41:1238-1242.
34. Mukhopadhyay S and Katzenstein AL. Subclassification of non-small cell lung carcinomas lacking morphologic differentiation on biopsy specimens: Utility of an immunohistochemical panel containing TTF-1, napsin A, p63, and CK5/6. The American journal of surgical pathology. 2011; 35:15-25.
35. Yamamoto H, Shigematsu H, Nomura M, Lockwood WW, Sato M, Okumura N, Soh J, Suzuki M, Wistuba, II, Fong KM, Lee H, Toyooka S, Date H, Lam WL, Minna JD and Gazdar AF. PIK3CA mutations and copy number gains in human lung cancers. Cancer research. 2008; 68:6913-6921.
36. Scheffler M, Bos M, Gardizi M, Konig K, Michels S, Fassunke J, Heydt C, Kunstlinger H, Ihle M, Ueckeroth F, Albus K, Serke M, Gerigk U, Schulte W, Topelt K, Nogova L, et al. PIK3CA mutations in non-small cell lung cancer (NSCLC): genetic heterogeneity, prognostic impact and incidence of prior malignancies. Oncotarget. 2015; 6:1315-1326. doi: 10.18632/oncotarget.2834.
37. Seiwert TY, Zuo Z, Keck MK, Khattri A, Pedamallu CS, Stricker T, Brown C, Pugh TJ, Stojanov P, Cho J, Lawrence MS, Getz G, Bragelmann J, DeBoer R, Weichselbaum RR, Langerman A, et al. Integrative and comparative genomic analysis of HPV-positive and HPV-negative head and neck squamous cell carcinomas. Clinical cancer research. 2015; 21:632-641.
38. Padmanabhan B, Tong KI, Ohta T, Nakamura Y, Scharlock M, Ohtsuji M, Kang MI, Kobayashi A, Yokoyama S and Yamamoto M. Structural basis for defects of Keap1 activity provoked by its point mutations in lung cancer. Molecular cell. 2006; 21:689-700.
39. Ohta T, Iijima K, Miyamoto M, Nakahara I, Tanaka H, Ohtsuji M, Suzuki T, Kobayashi A, Yokota J, Sakiyama T, Shibata T, Yamamoto M and Hirohashi S. Loss of Keap1 function activates Nrf2 and provides advantages for lung cancer cell growth. Cancer research. 2008; 68:1303-1309.
40. Shibata T, Ohta T, Tong KI, Kokubu A, Odogawa R, Tsuta K, Asamura H, Yamamoto M and Hirohashi S. Cancer related mutations in NRF2 impair its recognition by Keap1-Cul3 E3 ligase and promote malignancy. Proceedings of the National Academy of Sciences of the United States of America. 2008; 105:13568-13573.
41. Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, Kryukov GV, Lawrence MS, Sougnez C, McKenna A, Shefler E, Ramos AH, Stojanov P, Carter SL, Voet D, Cortes ML, et al. The mutational landscape of head and neck squamous cell carcinoma. Science. 2011; 333:1157-1160.
42. Lechner M, Frampton GM, Fenton T, Feber A, Palmer G, Jay A, Pillay N, Forster M, Cronin MT, Lipson D, Miller VA, Brennan TA, Henderson S, Vaz F, O’Flynn P, Kalavrezos N, et al. Targeted next-generation sequencing of head and neck squamous cell carcinoma identifies novel genetic alterations in HPV+ and HPV- tumors. Genome medicine. 2013; 5:49.
43. Ang KK, Harris J, Wheeler R, Weber R, Rosenthal DI, Nguyen-Tan PF, Westra WH, Chung CH, Jordan RC, Lu C, Kim H, Axelrod R, Silverman CC, Redmond KP and Gillison ML. Human papillomavirus and survival of patients with oropharyngeal cancer. The New England journal of medicine. 2010; 363:24-35.
44. Lui VW, Hedberg ML, Li H, Vangara BS, Pendleton K, Zeng Y, Lu Y, Zhang Q, Du Y, Gilbert BR, Freilino M, Sauerwein S, Peyser ND, Xiao D, Diergaarde B, Wang L, et al. Frequent mutation of the PI3K pathway in head and neck cancer defines predictive biomarkers. Cancer discovery. 2013; 3:761-769.
45. Agrawal N, Frederick MJ, Pickering CR, Bettegowda C, Chang K, Li RJ, Fakhry C, Xie TX, Zhang J, Wang J, Zhang N, El-Naggar AK, Jasser SA, Weinstein JN, Trevino L, Drummond JA, et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science. 2011; 333:1154-1157.
46. Chung CH, Guthrie VB, Masica DL, Tokheim C, Kang H, Richmon J, Agrawal N, Fakhry C, Quon H, Subramaniam RM, Zuo Z, Seiwert T, Chalmers ZR, Frampton GM, Ali SM, Yelensky R, et al. Genomic alterations in head and neck squamous cell carcinoma determined by cancer gene-targeted sequencing. Annals of oncology. 2015; 26:1216-1223.
47. Tommasino M. The human papillomavirus family and its role in carcinogenesis. Seminars in cancer biology. 2014; 26:13-21.
48. Grabowska AK and Riemer AB. The invisible enemy - how human papillomaviruses avoid recognition and clearance by the host immune system. The open virology journal. 2012; 6:249-256.
49. Parfenov M, Pedamallu CS, Gehlenborg N, Freeman SS, Danilova L, Bristow CA, Lee S, Hadjipanayis AG, Ivanova EV, Wilkerson MD, Protopopov A, Yang L, Seth S, Song X, Tang J, Ren X, et al. Characterization of HPV and host genome interactions in primary head and neck cancers. Proceedings of the National Academy of Sciences of the United States of America. 2014; 111:15544-15549.
50. Gaykalova DA, Manola JB, Ozawa H, Zizkova V, Morton K, Bishop JA, Sharma R, Zhang C, Michailidi C, Considine M, Tan M, Fertig EJ, Hennessey PT, Ahn J, Koch WM, Westra WH, et al. NF-kappaB and stat3 transcription factor signatures differentiate HPV-positive and HPV-negative head and neck squamous cell carcinoma. International journal of cancer. 2015; 137:1879-89.
51. An J, Mo D, Liu H, Veena MS, Srivatsan ES, Massoumi R and Rettig MB. Inactivation of the CYLD deubiquitinase by HPV E6 mediates hypoxia-induced NF-kappaB activation. Cancer cell. 2008; 14:394-407.
52. Kim Y, Hammerman PS, Kim J, Yoon JA, Lee Y, Sun JM, Wilkerson MD, Pedamallu CS, Cibulskis K, Yoo YK, Lawrence MS, Stojanov P, Carter SL, McKenna A, Stewart C, Sivachenko AY, et al. Integrative and comparative genomic analysis of lung squamous cell carcinomas in East Asian patients. Journal of clinical oncology. 2014; 32:121-128.
53. An SJ, Chen ZH, Su J, Zhang XC, Zhong WZ, Yang JJ, Zhou Q, Yang XN, Huang L, Guan JL, Nie Q, Yan HH, Mok TS and Wu YL. Identification of enriched driver gene alterations in subgroups of non-small cell lung cancer patients based on histology and smoking status. PloS one. 2012; 7:e40109.
54. Zhang Q, Zhu L and Zhang J. Epidermal growth factor receptor gene mutation status in pure squamous-cell lung cancer in Chinese patients. BMC cancer. 2015; 15:88.
55. Qiong Z, Na WY, Bo W, Zhu Z, Ling P, Bo MH, Min TY, Lei Z, Na HD, Bo Z, Fang LJ and Seng ZS. Alterations of a spectrum of driver genes in female Chinese patients with advanced or metastatic squamous cell carcinoma of the lung. Lung cancer. 2015; 87:117-121.
56. Bonanno L, Favaretto A, Rugge M, Taron M and Rosell R. Role of genotyping in non-small cell lung cancer treatment: current status. Drugs. 2011; 71:2231-2246.
57. Shtivelman E, Hensing T, Simon GR, Dennis PA, Otterson GA, Bueno R and Salgia R. Molecular pathways and therapeutic targets in lung cancer. Oncotarget. 2014; 5:1392-1433. doi: 10.18632/oncotarget.1891.
58. Schmitz S, Ang KK, Vermorken J, Haddad R, Suarez C, Wolf GT, Hamoir M and Machiels JP. Targeted therapies for squamous cell carcinoma of the head and neck: current knowledge and future directions. Cancer treatment reviews. 2014; 40:390-404.
59. Reck M, Heigener DF, Mok T, Soria JC and Rabe KF. Management of non-small-cell lung cancer: recent developments. Lancet. 2013; 382:709-719.
60. Vermorken JB, Mesia R, Rivera F, Remenar E, Kawecki A, Rottey S, Erfan J, Zabolotnyy D, Kienzer HR, Cupissol D, Peyrade F, Benasso M, Vynnychenko I, De Raucourt D, Bokemeyer C, Schueler A, et al. Platinum-based chemotherapy plus cetuximab in head and neck cancer. The New England journal of medicine. 2008; 359:1116-1127.
61. Bonner JA, Harari PM, Giralt J, Azarnia N, Shin DM, Cohen RB, Jones CU, Sur R, Raben D, Jassem J, Ove R, Kies MS, Baselga J, Youssoufian H, Amellal N, Rowinsky EK, et al. Radiotherapy plus cetuximab for squamous-cell carcinoma of the head and neck. The New England journal of medicine. 2006; 354:567-578.
62. Pirker R, Pereira JR, Szczesna A, von Pawel J, Krzakowski M, Ramlau R, Vynnychenko I, Park K, Yu CT, Ganul V, Roh JK, Bajetta E, O’Byrne K, de Marinis F, Eberhardt W, Goddemeier T, et al. Cetuximab plus chemotherapy in patients with advanced non-small-cell lung cancer (FLEX): an open-label randomised phase III trial. Lancet. 2009; 373:1525-1531.
63. Thatcher N, Hirsch FR, Luft AV, Szczesna A, Ciuleanu TE, Dediu M, Ramlau R, Galiulin RK, Balint B, Losonczy G, Kazarnowicz A, Park K, Schumann C, Reck M, Depenbrock H, Nanda S, et al. Necitumumab plus gemcitabine and cisplatin versus gemcitabine and cisplatin alone as first-line therapy in patients with stage IV squamous non-small-cell lung cancer (SQUIRE): an open-label, randomised, controlled phase 3 trial. The lancet oncology. 2015; 16:763-774.
64. Soria JC, Felip E, Cobo M, Lu S, Syrigos K, Lee KH, Goker E, Georgoulias V, Li W, Isla D, Guclu SZ, Morabito A, Min YJ, Ardizzoni A, Gadgeel SM, Wang B, et al. Afatinib versus erlotinib as second-line treatment of patients with advanced squamous cell carcinoma of the lung (LUX-Lung 8): an open-label randomised controlled phase 3 trial. The lancet oncology. 2015; 16:897-907.
65. Martin D, Abba MC, Molinolo AA, Vitale-Cross L, Wang Z, Zaida M, Delic NC, Samuels Y, Lyons JG and Gutkind JS. The head and neck cancer cell oncogenome: a platform for the development of precision molecular therapies. Oncotarget. 2014; 5:8906-8923. doi 10.18632/oncotarget.2417.
66. Perez-Moreno P, Brambilla E, Thomas R and Soria JC. Squamous cell carcinoma of the lung: molecular subtypes and therapeutic opportunities. Clinical cancer research. 2012; 18:2443-2451.
67. Isaacsson Velho PH, Castro G, Jr. and Chung CH. Targeting the PI3K Pathway in Head and Neck Squamous Cell Carcinoma. American Society of Clinical Oncology educational book / ASCO American Society of Clinical Oncology Meeting. 2015:123-128.
68. Liao RG, Jung J, Tchaicha J, Wilkerson MD, Sivachenko A, Beauchamp EM, Liu Q, Pugh TJ, Pedamallu CS, Hayes DN, Gray NS, Getz G, Wong KK, Haddad RI, Meyerson M and Hammerman PS. Inhibitor-sensitive FGFR2 and FGFR3 mutations in lung squamous cell carcinoma. Cancer research. 2013; 73:5195-5205.
69. Jiang T, Gao G, Fan G, Li M and Zhou C. FGFR1 amplification in lung squamous cell carcinoma: a systematic review with meta-analysis. Lung cancer. 2015; 87:1-7.
70. Weiss J, Sos ML, Seidel D, Peifer M, Zander T, Heuckmann JM, Ullrich RT, Menon R, Maier S, Soltermann A, Moch H, Wagener P, Fischer F, Heynck S, Koker M, Schottle J, et al. Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Science translational medicine. 2010; 2:62ra93.
71. Hammerman PS, Sos ML, Ramos AH, Xu C, Dutt A, Zhou W, Brace LE, Woods BA, Lin W, Zhang J, Deng X, Lim SM, Heynck S, Peifer M, Simard JR, Lawrence MS, et al. Mutations in the DDR2 kinase gene identify a novel therapeutic target in squamous cell lung cancer. Cancer discovery. 2011; 1:78-89.
72. Day E, Waters B, Spiegel K, Alnadaf T, Manley PW, Buchdunger E, Walker C and Jarai G. Inhibition of collagen-induced discoidin domain receptor 1 and 2 activation by imatinib, nilotinib and dasatinib. European journal of pharmacology. 2008; 599:44-53.
73. Lopez-Chavez A, Thomas A, Rajan A, Raffeld M, Morrow B, Kelly R, Carter CA, Guha U, Killian K, Lau CC, Abdullaev Z, Xi L, Pack S, Meltzer PS, Corless CL, Sandler A, et al. Molecular profiling and targeted therapy for advanced thoracic malignancies: a biomarker-derived, multiarm, multihistology phase II basket trial. Journal of clinical oncology. 2015; 33:1000-1007.
74. Herbst RS, Gandara DR, Hirsch FR, Redman MW, LeBlanc M, Mack PC, Schwartz LH, Vokes E, Ramalingam SS, Bradley JD, Sparks D, Zhou Y, Miwa C, Miller VA, Yelensky R, Li Y, et al. Lung Master Protocol (Lung-MAP)-A Biomarker-Driven Protocol for Accelerating Development of Therapies for Squamous Cell Lung Cancer: SWOG S1400. Clinical cancer research. 2015; 21:1514-1524.
75. Polyak K and Garber J. Targeting the missing links for cancer therapy. Nature medicine. 2011; 17:283-284.
76. Wiggans AJ, Cass GK, Bryant A, Lawrie TA and Morrison J. Poly(ADP-ribose) polymerase (PARP) inhibitors for the treatment of ovarian cancer. The Cochrane database of systematic reviews. 2015; 5:CD007929.
77. Deeks ED. Olaparib: first global approval. Drugs. 2015; 75:231-240.
78. Hamoir M, Ferlito A, Schmitz S, Hanin FX, Thariat J, Weynand B, Machiels JP, Gregoire V, Robbins KT, Silver CE, Strojan P, Rinaldo A, Corry J and Takes RP. The role of neck dissection in the setting of chemoradiation therapy for head and neck squamous cell carcinoma with advanced neck disease. Oral oncology. 2012; 48:203-210.
79. Adelstein DJ, Li Y, Adams GL, Wagner H, Jr., Kish JA, Ensley JF, Schuller DE and Forastiere AA. An intergroup phase III comparison of standard radiation therapy and two schedules of concurrent chemoradiotherapy in patients with unresectable squamous cell head and neck cancer. Journal of clinical oncology. 2003; 21:92-98.
80. Forastiere AA, Goepfert H, Maor M, Pajak TF, Weber R, Morrison W, Glisson B, Trotti A, Ridge JA, Chao C, Peters G, Lee DJ, Leaf A, Ensley J and Cooper J. Concurrent chemotherapy and radiotherapy for organ preservation in advanced laryngeal cancer. The New England journal of medicine. 2003; 349:2091-2098.
81. Pointreau Y, Garaud P, Chapet S, Sire C, Tuchais C, Tortochaux J, Faivre S, Guerrif S, Alfonsi M and Calais G. Randomized trial of induction chemotherapy with cisplatin and 5-fluorouracil with or without docetaxel for larynx preservation. Journal of the National Cancer Institute. 2009; 101:498-506.
82. DeMichele A, Yee D, Berry DA, Albain KS, Benz CC, Boughey J, Buxton M, Chia SK, Chien AJ, Chui SY, Clark A, Edmiston K, Elias AD, Forero-Torres A, Haddad TC, Haley B, et al. The Neoadjuvant Model Is Still the Future for Drug Development in Breast Cancer. Clinical cancer research. 2015; 21:2911-2915.
83. Chung CH and Schwartz DL. Impact of HPV-related head and neck cancer in clinical trials: opportunity to translate scientific insight into personalized care. Otolaryngologic clinics of North America. 2012; 45:795-806.
84. Wu D, Pang Y, Wilkerson MD, Wang D, Hammerman PS and Liu JS. Gene-expression data integration to squamous cell lung cancer subtypes reveals drug sensitivity. British journal of cancer. 2013; 109:1599-1608.
85. Luke JJ and Ott PA. PD-1 pathway inhibitors: the next generation of immunotherapy for advanced melanoma. Oncotarget. 2015; 6:3479-3492. doi: 10.18632/oncotarget.2980.
86. Champiat S, Ileana E, Giaccone G, Besse B, Mountzios G, Eggermont A and Soria JC. Incorporating immune-checkpoint inhibitors into systemic therapy of NSCLC. Journal of thoracic oncology. 2014; 9:144-153.
87. Martin SD, Coukos G, Holt RA and Nelson BH. Targeting the undruggable: Immunotherapy meets personalized oncology in the genomic era. Annals of oncology. 2015; 26:2367-74.
88. Pico de Coana Y, Choudhury A and Kiessling R. Checkpoint blockade for cancer therapy: revitalizing a suppressed immune system. Trends in molecular medicine. 2015; 21:482-491.
89. Brahmer J, Reckamp KL, Baas P, Crino L, Eberhardt WE, Poddubskaya E, Antonia S, Pluzanski A, Vokes EE, Holgado E, Waterhouse D, Ready N, Gainor J, Aren Frontera O, Havel L, Steins M, et al. Nivolumab versus Docetaxel in Advanced Squamous-Cell Non-Small-Cell Lung Cancer. The New England journal of medicine. 2015; 373:123-135.
90. Paz-Ares LH, L.; Borghaei, H.; Spigel, DR.; Steins, M.; Ready, N.; Chow, LQM.; Vokes, EE.; Felip, E.; Holgado, E.; Barlesi, F.; Kohlhaeufl, M.; Rodriguez, O.; Burgio, MA.; Fayette, J.; Gettinger, SN.; Harbison, C.; Dorange, C.; Finckenstein, FG.; Brahmer, JR. Phase III, randomized trial (CheckMate 057) of nivolumab (NIVO) versus docetaxel (DOC) in advanced non-squamous cell (non-SQ) non-small cell lung cancer (NSCLC). Journal of clinical oncology. 2015; 33(N° 15_suppl).
91. Herbst RS, Soria JC, Kowanetz M, Fine GD, Hamid O, Gordon MS, Sosman JA, McDermott DF, Powderly JD, Gettinger SN, Kohrt HE, Horn L, Lawrence DP, Rost S, Leabman M, Xiao Y, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014; 515:563-567.
92. Gettinger SN, Horn L, Gandhi L, Spigel DR, Antonia SJ, Rizvi NA, Powderly JD, Heist RS, Carvajal RD, Jackman DM, Sequist LV, Smith DC, Leming P, Carbone DP, Pinder-Schenck MC, Topalian SL, et al. Overall Survival and Long-Term Safety of Nivolumab (Anti-Programmed Death 1 Antibody, BMS-936558, ONO-4538) in Patients With Previously Treated Advanced Non-Small-Cell Lung Cancer. Journal of clinical oncology. 2015; 33:2004-2012.
93. Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, Ibrahim F, Bruggeman C, Gasmi B, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015; 348:124-128.
94. Le DT, Uram JN, Wang H, Bartlett BR, Kemberling H, Eyring AD, Skora AD, Luber BS, Azad NS, Laheru D, Biedrzycki B, Donehower RC, Zaheer A, Fisher GA, Crocenzi TS, Lee JJ, et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. The New England journal of medicine. 2015; 372:2509-2520.
95. Sacco AG and Cohen EE. Current Treatment Options for Recurrent or Metastatic Head and Neck Squamous Cell Carcinoma. Journal of clinical oncology. 2015; 33:3305-13.
96. Seiwert TH, RI.; Gupta, S.; Mehra, R.; Tahara, M.; Berger, R.; Lee, SH, Burtness, B.; Le, DT.; Heath, K.; Blum, A.; Dolled-Filhart, M.; Emancipator, K.; Pathiraja, K.; Cheng, JD.; Chow, LQ. Antitumor activity and safety of pembrolizumab in patients (pts) with advanced squamous cell carcinoma of the head and neck (SCCHN): preliminary results from KEYNOTE-012 expansion cohort. Journal of clinical oncology. 2015; 33(N° 15_suppl).
97. Pai S, Smith D, Peng S, Ishida E, Akpeng B, Hung CF and Wu TC. (2015). Therapeutic HPV vaccine increases sensitivity of poorly immunogenic tumor to anti-PD-1 monotherapy. 5th International Conference on Innovative Approaches in Head and Neck Oncology 2015. (Nice, France).

