INTRODUCTION
With approximately 1.4 million new cases and 700,000 deaths occurring in 2012, colorectal cancer (CRC) is the third most cancer and the fourth most common cause of death from cancer worldwide [1]. Stage at diagnosis is the most important prognostic factor, with 5-year relative survival ranging from >90% for patients with localized CRC to approximately 10% for patients with distant tumor spread [2, 3]. Randomized trials and observational studies have shown a large potential for reduction of CRC incidence and mortality by endoscopic or stool-based screening tests, such as sigmoidoscopy, colonoscopy, guaiac-based or immunochemical fecal occult blood tests [4–7].
An alternative approach for cancer screening might be blood-based screening tests. Due to their minimally invasive nature and straightforward implementation in routine medical examinations, blood tests might achieve high levels of adherence when applied in population-based screening [8, 9]. For instance, in a study by Adler and colleagues [9], the majority of participants who refused screening colonoscopy preferred a blood-based test (83%) over a stool-based test (15%) when both choices were offered. Autoantibodies against tumor-associated antigens (TAAs) were found to be present in cancer patients’ blood. The mechanism behind the humoral immune responses toward TAAs is complex and not fully understood. Production of autoantibodies could be induced in responses to over-expression, mutations, or abnormal posttranslational modifications of proteins in cancer cells [10].
Autoantibodies as potential biomarkers for early detection of cancer have been intensively studied in previous studies [11-14]. Although the sensitivity of single autoantibodies for cancer detection seems to be low, higher sensitivities might be achieved by joint testing for multiple autoantibodies [11, 14]. Promising candidates were reported in some studies, but most of the findings were based on relatively small sample sizes, and lacked independent validation using prospectively collected samples from screening settings [11, 14]. In regards to CRC, validation of autoantibody markers in large prospective studies was rarely done, with only a few exceptions [15-17]. For instance, Pedersen and colleagues [15] evaluated autoantibodies against MUC1 and MUC4 in 97 prospectively collected CRC samples and matched healthy controls. However, the included CRC patients were recruited from an ovarian cancer screening program in UK rather than a true CRC screening population.
In this study, we evaluated the individual and joint diagnostic performance for CRC and its precursors through antibodies against a panel of 64 predefined autoantigens, and we validated the most promising marker combinations in independent samples of participants recruited in a true CRC screening setting.
RESULTS
Figure 1 provides the flow diagram showing the study population selection for the training set and the validation set. Overall, 380 clinically identified CRC cases were included in our study. After excluding 28 samples with invalid multiplex serology test results, the remaining 352 CRC patients were included as cases in the training set. The independent validation set samples were exclusively sampled from participants enrolled in the BliTz study in 2005-2013. After excluding participants without adequate blood samples, participants who do not represent a true screening setting, and participants with potentially false negative results at screening colonoscopy, 5680 participants were eligible for the sample selection, from whom 417 samples (all CRC case and random sample of advanced adenomas, non-advanced adenomas and controls free of colorectal neoplasms) were selected for our measurement. After further excluding samples with invalid laboratory results, 49 CRC cases, 99 advanced adenomas, 29 non-advanced adenomas and 224 controls free of colorectal neoplasms were included in the analysis. The average time between blood sample withdrawal and screening colonoscopy among these participants was 6.6 days. We randomly selected 124 controls from this screening setting as control group in the training set. The remaining 100 participants were used as controls in the validation set.
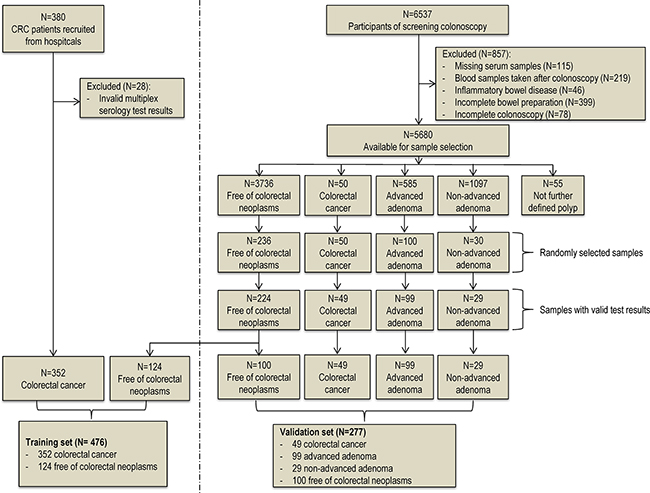
Figure 1: Flow diagram of sample selection procedure of the training set and the validation set.
Table 1 presents the distribution of socio-demographic characteristics of the study population. Compared to controls, more men and older patients were included in the CRC groups of both the training set and the validation set. About half of CRC patients were diagnosed at early stages in both sets. Slightly more than half of the cancers were located in the colon (57.4% in the training set and 53.1% in the validation set). Among advanced adenomas, the proportion of polyps with high grade dysplasia (HGD), polyps with villous architecture without HGD, and large adenomas (≥10mm) with neither HGD nor villous architecture were 12%, 56%, and 32%, respectively.
Table 1: Characteristics of the study population
Training set |
Validation set |
||||||
|---|---|---|---|---|---|---|---|
Group |
CRC (N, %) |
Control (N, %) |
CRC (N, %) |
Advanced adenoma (N, %) |
Non−advanced adenoma (N, %) |
Control (N, %) |
|
Age (years) |
|||||||
<60 |
73(20.8) |
50 (40.3) |
9 (18.4) |
30 (30.3) |
10 (34.5) |
43 (43.0) |
|
60−64 |
49 (14.0) |
30 (24.2) |
14 (28.6) |
27 (27.3) |
7 (24.1) |
22 (22.0) |
|
65−69 |
57 (16.2) |
22 (17.7) |
11(22.4) |
16 (16.2) |
6 (20.7) |
20 (20.0) |
|
≥70 |
172 (49.0) |
22 (17.7) |
15 (30.6) |
26 (26.3) |
6 (20.7) |
15 (15.0) |
|
Mean ± SD |
68.3 ± 11.6 |
62.2 ± 7.4 |
66.3 ± 6.7 |
64.5 ± 7.7 |
63.6 ± 6.5 |
62.0 ± 6.1 |
|
Sex |
|||||||
Male |
202 (57.4) |
56 (45.2) |
34 (69.4) |
50 (50.5) |
17 (58.6) |
45 (45.0) |
|
Female |
150 (42.6) |
68 (54.8) |
15 (30.6) |
49 (49.5) |
12 (41.4) |
55 (55.0) |
|
UICC TNM tumor stage |
|||||||
Tis (0) |
− |
− |
4 (8.2) |
− |
− |
− |
|
I |
96 (27.3) |
− |
18 (36.7) |
− |
− |
− |
|
II |
102 (29.0) |
− |
5 (10.2) |
− |
− |
− |
|
III |
105 (29.8) |
− |
19 (38.8) |
− |
− |
− |
|
IV |
49 (13.9) |
− |
3 (6.1) |
− |
− |
− |
|
CRC location |
|||||||
Colon |
202 (57.4) |
− |
26 (53.1) |
− |
− |
− |
|
Rectum |
150 (42.6) |
− |
22 (44.9) |
− |
− |
− |
|
Unknown |
− |
− |
1 (2.0) |
− |
− |
− |
|
Advanced adenoma subclass |
|||||||
HGD |
− |
− |
− |
12 (12.1) |
− |
− |
|
Villous |
− |
− |
− |
55 (55.6) |
− |
− |
|
Large polyp |
− |
− |
− |
32 (32.3) |
− |
− |
|
Abbreviations: CRC, colorectal cancer; HGD, high grade dysplasia; SD, standard deviation; TNM, tumor-node-metastasis; UICC, Union for International Cancer Control.
A head-to-head comparison of the diagnostic performance of all 64 autoantibody markers for detecting colorectal neoplasms is shown in Supplementary Table 2 and the diagnostic performance of the 21 best performing autoantibodies identified in the training set is presented in Table 2. Using cutoffs yielding 98% specificity in the training set generally resulted in rather low levels of sensitivity in both the training set and the validation set. The majority of markers were found to have lower sensitivities for CRC in the validation set compared to the training set. Overall, there were 21 markers showing sensitivities for early stage CRC ≥4% at 98% specificity in the training set. However 7 of them did not detect any early stage CRC in the validation set (see Table 2). We further examined the joint seropositivity of the top 21 markers among cases of the training set (see Figure 2). There was only very limited co-occurrence beyond chance of the single autoantibody markers. Although the majority of Kappa coefficients were positive (145 out of 210, 69%), 189 out of 210 Kappa coefficients (90%) for pairwise joint occurrence were close to zero with values between −0.10 and +0.20.
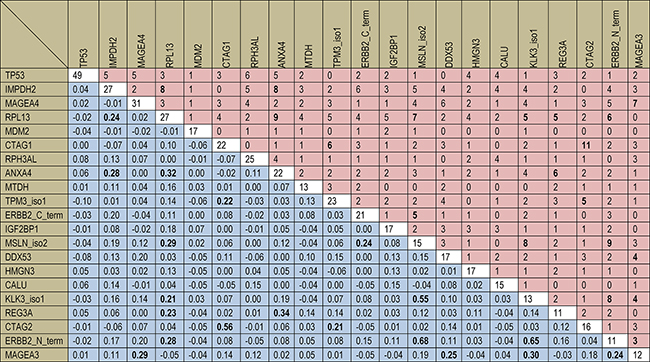
Figure 2: Individual and joint seropositivity of 18 autoantibody markers in the 352 CRC cases of the training set. The diagonal (white cells) and the upper right triangle (red cells) show the absolute numbers of seropositivity for single autoantibody markers and of joint seropositivity for pairwise marker combinations, respectively. The lower left triangle (blue cells) shows Cohen’s Kappa coefficients for pairwise marker combinations. Bold figures indicate Cohens’s Kappa coefficients higher than 0.20 and corresponding absolute numbers of joint seropositivity.
Table 2: Diagnostic performance of the top 21 single autoantibody markers for detecting colorectal neoplasms
Antigen |
cutoff (MFI) |
Training set |
Validation set |
||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
SE for CRC† |
SE for stage I/II CRC† |
SE for stage III/IV CRC† |
SPE† |
SE for CRC† |
SE for stage 0/I/II CRC† |
SE for stage III/IV CRC† |
SE for AA† |
SE for NA† |
SPE† |
||
TP53 |
601 |
14 [11–18] |
13 [9–19] |
15 [10–21] |
98 [93–99] |
8 [3–19] |
7 [2–23] |
9 [3–28] |
1 [0–6] |
0 [0–12] |
100 [96–100] |
IMPDH2 |
47 |
8 [5–11] |
10 [7–15] |
5 [2–9] |
98 [93–99] |
4 [1–14] |
4 [0–18] |
5 [0–22] |
6 [3–13] |
7 [2–22] |
95 [89–98] |
MAGEA4 |
168 |
9 [6–12] |
9 [6–14] |
8 [5–14] |
98 [93–99] |
6 [2–17] |
11 [4–28] |
0 [0–15] |
12 [7–20] |
0 [0–12] |
96 [90–98] |
RPL13 |
170 |
8 [5–11] |
9 [5–13] |
6 [4–12] |
98 [93–99] |
4 [1–14] |
0 [0–12] |
9 [3–28] |
2 [1–7] |
10 [4–26] |
97 [92–99] |
MDM2 |
400 |
5 [3–8] |
8 [5–12] |
1 [0–5] |
98 [93–99] |
2 [0–11] |
4 [0–18] |
0 [0–15] |
1 [0–6] |
7 [2–22] |
98 [93–99] |
CTAG1 |
91 |
6 [4–9] |
7 [4–11] |
6 [3–11] |
98 [93–99] |
2 [0–11] |
0 [0–12] |
5 [0–22] |
6 [3–13] |
3 [0–17] |
95 [89–98] |
RPH3AL |
87 |
7 [5–10] |
7 [4–11] |
8 [5–13] |
98 [93–99] |
0 [0–7] |
0 [0–12] |
0 [0–15] |
6 [3–13] |
3 [0–17] |
91 [84–95] |
ANXA4 |
50 |
6 [4–9] |
6 [4–10] |
6 [4–12] |
98 [93–99] |
4 [1–14] |
4 [0–18] |
5 [0–22] |
2 [1–7] |
7 [2–22] |
99 [95–100] |
MTDH |
440 |
4 [2–6] |
6 [4–10] |
1 [0–4] |
98 [93–99] |
4 [1–14] |
4 [0–18] |
5 [0–22] |
2 [1–7] |
3 [0–17] |
98 [93–99] |
TPM3_iso1 |
424 |
7 [4–10] |
6 [4–10] |
7 [4–12] |
98 [93–99] |
8 [3–19] |
0 [0–12] |
18 [7–39] |
4 [2–10] |
7 [2–22] |
96 [90–98] |
ERBB2_C_term |
90 |
6 [4–9] |
6 [3–10] |
6 [4–12] |
98 [93–99] |
6 [2–17] |
7 [2–23] |
5 [0–22] |
8 [4–15] |
3 [0–17] |
92 [85–96] |
IGF2BP1 |
309 |
5 [3–8] |
6 [3–10] |
4 [2–8] |
98 [93–99] |
2 [0–11] |
4 [0–18] |
0 [0–15] |
2 [1–7] |
3 [0–17] |
99 [95–100] |
MIA |
52 |
4 [3–7] |
5 [3–9] |
3 [1–7] |
98 [94–100] |
2 [0–11] |
4 [0–18] |
0 [0–15] |
1 [0–6] |
3 [0–17] |
98 [93–99] |
DDX53 |
460 |
5 [3–8] |
5 [2–8] |
5 [3–10] |
98 [93–99] |
0 [0–7] |
0 [0–12] |
0 [0–15] |
2 [1–7] |
3 [0–17] |
99 [95–100] |
HMGN3 |
706 |
5 [3–8] |
5 [2–8] |
5 [3–10] |
98 [93–99] |
8 [3–19] |
11 [4–28] |
5 [0–22] |
9 [5–16] |
10 [4–26] |
94 [88–97] |
CALU |
578 |
4 [3–7] |
4 [2–8] |
5 [2–9] |
98 [93–99] |
6 [2–17] |
0 [0–12] |
14 [5–33] |
4 [2–10] |
3 [0–17] |
92 [85–96] |
KLK3_iso1 |
41 |
4 [2–6] |
4 [2–8] |
3 [1–7] |
98 [93–99] |
2 [0–11] |
4 [0–18] |
0 [0–15] |
1 [0–6] |
3 [0–17] |
98 [93–99] |
REG3A |
62 |
3 [2–6] |
4 [2–8] |
2 [1–6] |
98 [93–99] |
0 [0–7] |
0 [0–12] |
0 [0–15] |
1 [0–6] |
3 [0–17] |
99 [95–100] |
CTAG2 |
135 |
5 [3–7] |
4 [2–7] |
6 [3–11] |
98 [93–99] |
4 [1–14] |
7 [2–23] |
0 [0–15] |
3 [1–9] |
0 [0–12] |
93 [86–97] |
ERBB2_N_term |
81 |
3 [2–6] |
4 [2–7] |
3 [1–6] |
98 [93–99] |
4 [1–14] |
4 [0–18] |
5 [0–22] |
3 [1–9] |
3 [0–17] |
97 [92–99] |
MAGEA3 |
220 |
3 [2–6] |
4 [2–7] |
3 [1–7] |
98 [93–99] |
4 [1–14] |
7 [2–23] |
0 [0–15] |
6 [3–13] |
0 [0–12] |
95 [89–98] |
†In % with 95% confidence interval.
Abbreviations: AA, advanced adenoma; NA, non−advanced adenoma, CRC, colorectal cancer; MFI, median fluorescence intensity; SE, sensitivity; SPE, specificity.
Overall, there were only 9 autoantibody markers whose sensitivities for detecting early stage CRC were higher than 5%, ranging from 7% to 11% in the validation set (see Supplementary Table 2). Anti-MAGEA4 was found to have best diagnostic potential for detecting early stage CRC and advanced adenomas, with sensitivities of 11% (95% CI, 4−28%) and 12% (95% CI, 7−20%), respectively, at a specificity of 96%. Anti-TP53 detected 8% of CRC (7% at early stage), but only 1% of advanced adenomas at 100% specificity.
A systematic search for the best n-marker combination was conducted among all the 64 autoantibody markers with the purpose to enhance the diagnostic performance. Results for the best performing 2-, 3-, 4-, 5- and 6-marker combinations are shown in Table 3. Including six or more markers in the combination resulted in specificities much lower than 90% for the best performing marker combinations even in the training set, the analysis was therefore limited to combinations of no more than 6 markers. As expected, sensitivities increased and specificities decreased with increasing numbers of markers included the multi-marker combination. For example, sensitivity for any CRC increased from 22% for the best performing 2-marker panel to 40% for the best performing 6-marker panel in the training set, whereas specificity decreased from 95% to 88%. Interestingly, sensitivity was consistently higher for early stage than for advanced stage CRC. As expected, sensitivities were slightly lower in the validation set, but a sensitivity of 30% (95% CI, 16−48%) for early stage CRC along with a specificity of 85% (95% CI, 77−91%) was still observed for the best performing 6-marker combination. Additionally, it also detected 25% (95% CI, 18−35%) of advanced adenomas, the most important precursors of CRC.
Table 3: Diagnostic performance of best performing two-, three-, four-, five-, and six-marker combinations derived from training set for detecting colorectal neoplasms
Marker combination |
Training set |
Validation set |
||||||||
|---|---|---|---|---|---|---|---|---|---|---|
SE for CRC† |
SE for stage I/II CRC† |
SE for stage III/IV CRC† |
SPE† |
SE for CRC† |
SE for stage 0/I/II CRC† |
SE for stage III/IV CRC† |
SE for AA† |
SE for NA† |
SPE† |
|
Best performing two-marker combination |
||||||||||
TP53+IMPDH2 |
22 (18-26) |
23 (17-29) |
20 (15-28) |
95 (90-98) |
10 (4-22) |
11 (4-28) |
9 (3-28) |
7 (3-14) |
7 (2-22) |
95 (89-98) |
Best performing three-marker combination |
||||||||||
TP53+IMPDH2+MDM2 |
26 (22-31) |
30 (24-37) |
21 (15-29) |
93 (87-96) |
12 (6-32) |
15 (6-32) |
9 (3-28) |
8 (4-15) |
10 (4-26) |
94 (88-97) |
Best performing four-marker combination |
||||||||||
TP53+IMPDH2+MDM2+MAGEA4 |
33 (28-38) |
37 (30-44) |
28 (21-36) |
90 (84-94) |
18 (10-31) |
26 (13-45) |
9 (3-28) |
20 (13-29) |
10 (4-26) |
90 (83-94) |
Best performing five-marker combination |
||||||||||
TP53+IMPDH2+MDM2+MAGEA4+CTAG1 |
38 (33-43) |
43 (36-50) |
32 (25-40) |
89 (82-93) |
20 (11-34) |
26 (13-45) |
14 (5-33) |
24 (17-34) |
14 (5-31) |
87 (79-92) |
Best performing six-marker combination |
||||||||||
TP53+IMPDH2+MDM2+MAGEA4+CTAG1+MTDH |
40 (34-45) |
46 (39-53) |
32 (25-40) |
88 (81-93) |
24 (15-38) |
30 (16-48) |
18 (7-39) |
25 (18-35) |
17 (8-35) |
85 (77-91) |
† In % with 95% confidence interval.
Abbreviations: AA, advanced adenoma; CI, confidence interval; CRC, colorectal cancer; NA, non-advanced adenoma; SE, sensitivity; SPE, specificity.
We further evaluated additional seven 6-marker combinations, whose Youden indices were only slightly lower than the best-performing 6-marker combination in the training set. Details of the diagnostic performance of these combinations are shown in Table 4. Overall, the sensitivities of these 6-marker combinations for detecting CRC ranged from 20% to 29% at specificities ranging from 84% to 86%. The sensitivities for detecting early stage CRC were higher than for detecting advanced stage CRC, and ranged from 26% to 30%. Notably, these panels also exhibited good diagnostic potential for detecting advanced adenomas, with sensitivities ranging from 21% to 27%. Interestingly, four autoantibody markers, anti-TP53, anti-IMPDH2, anti-MDM2 and anti-MAGEA4, were consistently included in all of these seven 6-marker combinations. These are also the makers that were included in the best 4-marker combination which yielded almost the same sensitivity for early stage and advanced adenomas (26% and 20%, respectively), albeit at higher specificity (90%). A graphic presentation of the MFI value distributions of these four markers is shown in Figure 3.
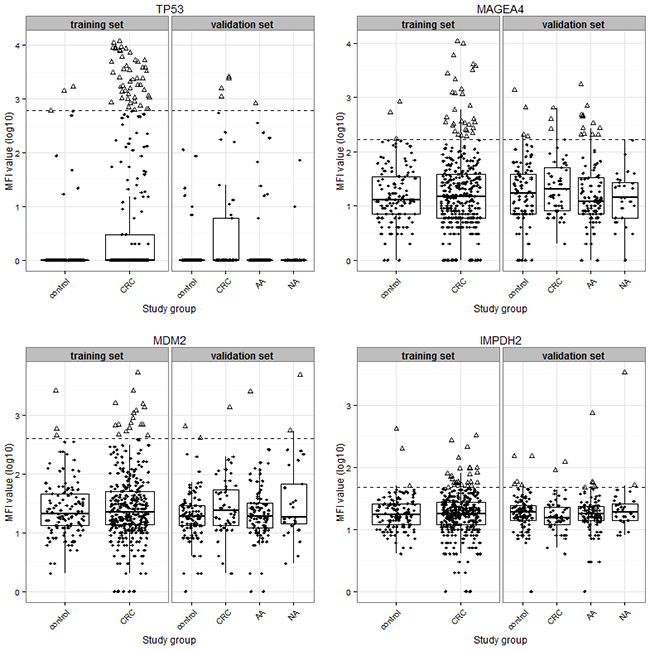
Figure 3: Scatter plot and box plot showing the distribution of median fluorescence intensity (MFI) values (in log10 transformation) of four autoantibody biomarkers (anti-TP53, anti-MAGEA4, anti-MDM2 and anti-IMPDH2) between different study groups. The bottom and top of the box indicate the first (Q1) and third (Q3) quartiles, and the middle line in the box is the median; the upper-limit equals Q3 plus 1.5 times interquartile range (IQR), and the lower-limit equals Q1 minus 1.5 times IQR. The horizontal dashed lines represent the cutoffs of respective biomarkers. Triangles represent samples with MFI values above the respective cutoffs, and solid dots represent samples with MFI values below the respective cutoffs. Abbreviations: AA, advanced adenoma; CRC, colorectal cancer; NA, non-advanced adenoma.
Table 4: Diagnostic performance of the top 8 best 6-marker combinations derived from training set for detecting colorectal neoplasms
Marker combination |
Training set |
Validation set |
||||||||
|---|---|---|---|---|---|---|---|---|---|---|
SE for CRC† |
SE for stage I/II CRC† |
SE for stage III/IV CRC† |
SPE† |
SE for CRC† |
SE for stage 0/I/II CRC† |
SE for stage III/IV CRC† |
SE for AA† |
SE for NA† |
SPE† |
|
Panel 1 |
40 (34-45) |
46 (39-53) |
32 (25-40) |
88 (81-93) |
24 (15-38) |
30 (16-48) |
18 (7-39) |
25 (18-35) |
17 (8-35) |
85 (77-91) |
Panel 2 |
42 (37-48) |
48 (41-55) |
35 (28-43) |
86 (79-91) |
27 (16-40) |
26 (13-45) |
27 (13-48) |
27 (19-37) |
21 (10-38) |
84 (76-90) |
Panel 3 |
39 (34-44) |
44 (37-51) |
32 (25-40) |
89 (82-93) |
24 (15-38) |
30 (16-48) |
18 (7-39) |
21 (14-30) |
21 (10-38) |
86 (78-91) |
Panel 4 |
39 (34-45) |
44 (37-51) |
33 (26-41) |
89 (82-93) |
27 (16-40) |
26 (13-45) |
27 (13-48) |
23 (16-32) |
17 (8-35) |
85 (77-91) |
Panel 5 |
39 (34-45) |
45 (38-52) |
32 (25-40) |
88 (81-93) |
22 (13-36) |
30 (16-48) |
14 (5-33) |
24 (17-34) |
14 (5-31) |
85 (77-91) |
Panel 6 |
41 (36-46) |
46 (39-53) |
34 (27-42) |
87 (80-92) |
20 (11-34) |
26 (13-45) |
14 (5-33) |
26 (19-36) |
14 (5-31) |
86 (78-91) |
Panel 7 |
41 (36-47) |
46 (39-53) |
35 (28-43) |
87 (80-92) |
24 (15-38) |
26 (13-45) |
23 (10-43) |
25 (18-35) |
21 (10-38) |
85 (77-91) |
Panel 8 |
42 (37-47) |
47 (40-54) |
36 (28-44) |
86 (79-91) |
29 (18-42) |
26 (13-45) |
32 (16-53) |
24 (17-34) |
24 (12-42) |
85 (77-91) |
† In % with 95% confidence interval.
Abbreviations: AA, advanced adenoma; CI, confidence interval; CRC, colorectal cancer; NA, non-advanced adenoma; SE, sensitivity; SPE, specificity.
Panel 1: TP53 + IMPDH2 + MDM2 + MAGEA4 + CTAG1+ MTDH
Panel 2: TP53 + IMPDH2 + MDM2 + MAGEA4 + CTAG1 + TPM3_iso1
Panel 3: TP53 + IMPDH2 + MDM2 + MAGEA4 + RPL13 + SAG
Panel 4: TP53 + IMPDH2 + MDM2 + MAGEA4 + TPM3_iso1 + CTAG1
Panel 5: TP53 + IMPDH2 + MDM2 + MAGEA4 + CTAG1 + SAG
Panel 6: TP53 + IMPDH2 + MDM2 + MAGEA4 + CTAG1 + IGF2BP1
Panel 7: TP53 + IMPDH2 + MDM2 + MAGEA4 + CTAG1 + RPL13
Panel 8: TP53 + IMPDH2 + MDM2 + MAGEA4 + TPM3_iso1 + RPL13
DISCUSSION
In this study, we first identified autoantibody markers and their combinations for early detection of CRC and its precursors in a large set of samples, which were then independently validated in prospectively collected samples from a true screening population. Overall, most of the tested autoantibodies showed relatively low sensitivity at cutoff levels yielding high specificity. Among all single markers, anti-MAGEA4 showed the highest sensitivity for detecting early stage CRC (11%, 95% CI, 4−28%) and advanced adenomas (12%, 95% CI, 7−20%) at a specificity of 96% (95% CI, 90−98%). We also explored the potential of multi-marker combinations to enhance diagnostic performance. Sensitivity increased and specificity decreased with increasing numbers of markers included in multi-marker combinations. All marker-combinations had higher sensitivities for early stage than for advanced stage CRC even in the validation set. Four markers, including anti-TP53, anti-MAGEA4, anti-MDM2 and anti-IMPDH2, were consistently included in best performing 4-, 5-, and 6-marker combinations. A combination of these four markers could detect 26% of early stage CRC (95% CI, 13−45%) and 20% of advanced adenomas (95% CI, 13−29%) at a specificity of 90% (95% CI, 83−94%).
Many single autoantibody markers tested in our study were also evaluated in previous studies, mostly using retrospective case-control designs. Apparently substantially better discriminative ability between CRC and healthy controls were reported for some markers. For instance, Kanojia and colleagues [18] measured blood anti-SPAG9 from 54 clinically diagnosed CRC patients and 50 healthy donors using ELISA and Western Blot. They reported a sensitivity of 70% at 100% specificity. However, in our analyses, anti-SPAG9 only detected 2% of the CRC cases at 98% specificity in the training set, and 4% of the CRC cases at 97% specificity in the validation set. Similar apparent discrepancies with previous reports were observed for other markers, such as anti-RPH3AL [19], anti-SEC61B [20], anti-ANXA4 [21].
The apparent discrepancies could be explained by different design issues and different study settings adopted in different studies. Most previous studies recruited participants in clinical settings, i.e., using clinically detected cases and convenience controls or healthy donors who often substantially differed from cases with respect to major characteristics that may be related to the autoantibody levels, such as age and sex [18-21]. Under such circumstances, a number of factors, such as tumor stage [22] and size, or differences in sociodemographic characteristic between cases and controls [23] may affect the validity of study findings. Moreover, most previous studies did not take efforts to adjust for the potential overestimation of diagnostic performance indicators, either through internal validation (bootstrap or cross-validation) or through external validation. In our analyses, a two-step approach was taken including selecting biomarkers in a training set and subsequently validating findings in an independent validation set. Of note, the validation was performed in a true screening setting, i.e., in the ideal target population for CRC screening, with colonoscopic verification of presence or absence of colorectal neoplasms (including adenomas) among all participants. By adopting such a study design and analysis strategy, the pitfalls often encountered in retrospective study designs could be avoided [24, 25]. In addition, we applied multiplex Luminex-based serological assays to measure all 64 autoantibody markers. Previous research results indicated that multiplex serology exhibited increased detection of weak antibody responses compared to ELISA [26], thus possibly providing more accurate results regarding serum autoantibody responses against TAAs.
Definition of cutoff directly affects the sensitivity and specificity of respective markers. In our analysis, the cutoffs of 64 autoantibody markers were set at relatively high levels to ensure very high specificities of each single marker (≥ 98% in the training set). This may explain the somewhat lower sensitivity for the single markers compared to previous reports, but it ensured maintenance of reasonably high specificity of multi-marker combinations. For instance, anti-TP53 is a widely evaluated antibody marker for cancer detection. In the current study, ROC (receiver operating characteristic) analysis showed that the AUC (area under the curve) of anti-TP53 for detecting CRC was 0.60 in the validation set (data not shown in the results). This result was quite similar to results from other studies evaluating anti-TP53 for early detection of CRC in large prospective studies [16, 17].
Among tested single autoantibody markers, anti-MAGEA4 was found to have good diagnostic potential for detecting early stage CRC (sensitivity=11%; 95% CI, 4−28%) and advanced adenomas (sensitivity=12%; 95% CI, 7−20%) in our analysis. MAGEA4 is a member of cancer/testis (CT) antigens, which is expressed in malignant cells and in immune-privileged germ-line cells [27]. It has a putative role of mediating anti-apoptotic functions by interacting with TP53 [28]. Over-expression of MAGEA4 was previously reported in other malignancies, such as lung cancer [29] and ovarian cancer [30]. To our knowledge, our study is the first to indicate that humoral responses to MAGEA4 could be observed in the early stage of CRC and its precursors. Although its diagnostic performance as a single biomarker is quite limited, it seems to have potential to be included in a multi-marker panel in the future.
As genetic or epigenetic alterations leading to aberrantly expression of antigens, such as TP53 [31] and CTAG1 (also known as NY-ESO-1) [32], may occur in various types of cancer, humoral responses to autoantigens evaluated in our study may also be observed in patients with other malignancies rather than in CRC only. To what extent these markers could detect other malignancies should be evaluated in future studies.
The sensitivities for detecting CRC of the included autoantibodies are clearly inferior to detection of CRC by currently recommended tests for early detection of CRC, i.e., faecal immunochemical tests (FITs) for hemoglobin [33, 34], and possibly even inferior to guaiac-based faecal occult blood tests (gFOBTs) [35, 36], probably still the most widely used non-invasive tests globally. Notably, sensitivities for detecting advanced adenomas of our marker combinations (≥20%) were similar to sensitivities reported for FITs [33, 34], and clearly superior to sensitivities reported for gFOBTs35,36 and any of other suggested blood tests that were validated in screening settings. For instance, a recent study by Church and colleagues [37] tested methylated Septin9 in 7941 asymptomatic individuals attending screening colonoscopy, and reported a sensitivity for detecting advanced adenoma of 4.2% (6/130) and 15.7% (24/184) for women and men, respectively, at a specificity of 91.5%.In our study, we deliberately selected antibody markers against TAAs based on their ability to detect early stage CRC rather than any stage CRC. This most likely explains the apparently unusual finding of better diagnostic performance for early stage CRC than for late stage CRC even in the validation set, whereas other biomarker studies typically showed opposite patterns. Another possible explanation might be immune destruction, an emerging hallmark of cancer [38], which occurs in advanced stage of CRC inducing the loss of antibody production. In a screening setting, the vast majority of prevalent preclinical and previously undetected CRC are at an early stage [22, 39]. Early detection and removal of these cancers, along with early detection and removal of the precursors, advanced adenomas, is likely to bear the largest potential for prevention of CRC deaths [5, 40].
It should be stated that the multi-marker panels identified in this study cannot be regarded as a final solution for CRC screening due to their overall limited sensitivity. Nevertheless, given their potential for detecting early stage CRC and especially its most important precursor – advanced adenomas (positive predictive values ranging from 57% to 67%), at high specificities, combining these panels with other promising biomarkers or tests might contribute to the development of clinically useful multi-marker panel in the future. In particular, further research should also explore the possibility of enhanced detection of advanced adenomas by combining FIT with a panel of autoantibody markers. However, additional issues such as compliance to specimen sampling (blood and stool), complexity of conducting multiple tests and cost, need to be taken into consideration for such potential test combinations.
Specific strengths and limitations deserve careful consideration when interpreting our study. Strengths include that we adopted a two-step approach to minimize overestimation of indicators of diagnostic performance. The validation of biomarkers was conducted in a large subset sample of participants of screening colonoscopy, which represent the target population for CRC screening in which the diagnostic performance of biomarkers should be evaluated. Furthermore, we applied the state-of-art technique to measure a large number of autoantibodies simultaneously, enabling a head-to-head comparison of the diagnostic performance of these biomarkers possible. Limitations of our study include the relatively small number of CRC cases in the validation samples, despite the very large screening population recruited, which reflects the very low prevalence of CRC in a true screening population. Moreover, the target antigens evaluated in our measurement were mostly native full-length proteins. Other types of TAAs, such mutated neo-antigens or proteins with abnormal posttranslational modifications were not tested in our study. In addition, the cutoffs of respective autoantibody markers were defined based on MFI values of controls with no significant findings at screening colonoscopy in the training set. Further determination of optimal cutoffs of respective autoantibody markers needs to be validated in larger screening populations.In summary, autoantibodies against TAAs carry potential for early detection of CRC. The multi-marker combinations developed in our study exhibited promising diagnostic performance for detecting a relevant proportion of early stage CRC and its relevant precursors. The combination of the most promising biomarkers identified in our study with additional promising biomarkers or tests might contribute to the development of a useful multi-marker blood-based test for early detection of CRC and its precursors in the future.
MATERIALS AND METHODS
1. Study design and study population
Our analysis is based on the ongoing BliTz study (Begleitende Evaluierung innovativer Testverfahren zur Darmkrebsfrüherkennung), which has been described in detail elsewhere [34, 41]. Briefly, participants of screening colonoscopy are recruited in gastroenterology practices in Southern Germany since November 2005. Patients are recruited at a preparatory visit and invited to donate blood and stool samples for the evaluation of novel CRC screening tests (typically one week) before screening colonoscopy. Colonoscopy records are collected from all participants. In Germany, screening colonoscopies have been offered for CRC screening since October 2002. Only experienced endoscopists are licenced for performing screening colonoscopies. Rigorous standards, including central review of full examination documentation, have been adopted to ensure high quality of screening colonoscopy [39, 42, 43]. The high quality of screening colonoscopy in Germany is reflected in high adenoma detection rates which have steadily increased since the introduction of the screening program [44]. Histopathological examination of polyps is decentrally conducted by certified pathology labs.
For this analysis, the following exclusion criteria were applied to exclude participants without adequate blood samples, participants not representing a true screening setting, and participants with potentially inadequate colonoscopy results: blood samples taken after screening colonoscopy, history of CRC or inflammatory bowel disease, incomplete colonoscopy or insufficient bowel preparation (latter only for controls). From the remaining participants of the BliTz study recruited in 2005-2013 (N=5680), all 50 newly detected CRC cases, as well as random samples of participants with advanced colorectal adenomas (AA, N=100), non-advanced colorectal adenomas (NA, N=30) and with no colorectal neoplasms (controls, N=236) were included in the measurement. Because in a true screening population like ours CRC patients are expected to be on average slightly older and to include a somewhat larger proportion of men, age and sex were not matched between different disease groups to avoid biased estimates of specificity [23].
Given the limited number of CRC cases even in a screening study as large as the BliTz study, we also included 380 CRC cases recruited after diagnosis but before treatment at four hospitals in Southern Germany for the discovery phase of this study. All studies were approved by the ethics committees of the medical faculty of the University of Heidelberg and of the respective state medical boards. Written informed consent was obtained from each participant.
2. Classification of colorectal cancer and advanced adenoma
Based on the colonoscopy reports or hospital records of all participants, invasive CRC was classified into stages Tis (carcinoma in situ) and I–IV according to the UICC (Union for International Cancer Control) TNM (tumor-node-metastasis) stage classification (7th version). In addition, advanced adenomas were subclassified into high grade dysplasia (HGD), villous architecture without HGD, and large adenomas (≥10 mm) with neither HGD nor villous architecture. Relevant information was extracted by two research assistants independently who were blind to the blood test results.
3. Laboratory procedures
3.1 Sample preparation
Blood samples were collected in Sarstedt S-Monovette (Product No. 02.1388) tubes and Vacutainer SST II (Product No. 367953) tubes before bowel preparation for colonoscopy (BliTz study), or prior to large bowel surgery or neoadjuvant therapy (380 CRC cases from the clinical setting). After completion of blood clotting, blood samples were immediately centrifuged at 2000−2500g for 10 minutes and stored at -80°C until analyses. Details on the standard operating procedures have also been described previously [45].
3.2 Multiplex serology
We selected 64 candidate TAAs encoded by 59 genes based on previous autoantibody measurements [46] and two recent systematic reviews [11, 14]. Details on the 64 candidate TAAs are provided in Supplementary Table 1. Autoantibodies to the selected TAAs were measured by multiplex serology, a fluorescent bead-based GST capture immunosorbent assay, as described previously 46, 47]. In short, TAAs were bacterially expressed as GST-X-tag fusion proteins [48], loaded and affinity-purified on glutathione-casein-coupled spectrally distinct fluorescence-labeled polystyrene beads (SeroMap, Luminex Corp., Austin, Tx, USA). A mix of differently loaded bead sets provides an antigen suspension array that is presented to sera. A Luminex analyzer (Luminex Corp., Austin, Tx, USA) distinguishes the bead set by its internal bead-color and quantifies the amount of serum-antibody detected by a secondary goat antihuman IgA, IgM, IgG antibody (Dianova, Hamburg, Germany) and the reporter conjugate streptavidin-R-phycoerythrin. The antibody reactivity is given as median fluorescence intensity (MFI) of at least 100 beads per set measured. Final antigen-specific MFI values were generated by subtraction of GST-tag and individual bead background values.
Sera were sent on dry ice to the Division of Molecular Diagnostics of Oncogenic Infections, DKFZ, Heidelberg and analyzed in a 1:1000 dilution by laboratory staff that was blinded for the case-control status.
Antigen-loading of the beads was controlled via detection of the C-terminal tag. Identity of the antigen loaded on the beads was verified by identifying the encoding plasmids via PCR and sequencing. Serum samples were denominated as invalid due to either insufficient amount of serum or too high GST-background values (>300 MFI). Variation between different assay-plates was controlled by pipetting three control sera on each plate as replicates. For overall 19 replicates, coefficients of variation (CV) were calculated for every antigen with a mean reactivity above 30 MFI. The median (range) CVs for the three samples were 16% (10-24%), 14% (11-21%), and 18% (11-25%), respectively.
4. Statistical analyses
We used a two-step approach with selection of markers and marker combinations in a training set, and validation of the results in an independent validation set. The validation set was defined in such a way that it represents a true screening setting, i.e., only participants from the BliTz study were included. CRC cases recruited in the clinical setting and randomly selected controls from the BliTz study were used for the training set.
Characteristics of the study population were first described and compared between the training set and the validation set. The strategy for selection of autoantibodies and their cutoff levels for predicting presence of colorectal neoplasm were based on the particular character of single autoantibody markers, which typically have rather low sensitivity at very high specificity [11, 14]. Cutoff levels yielding 98% specificity in the training set were determined (cutoffs lower than 30 MFI were replaced by a technical minimum cutoff of 30 MFI to reduce the influence of background noise of the multiplex serology measurement). After applying the cutoffs to respective markers, dichotomous variables indicating the predicted outcome (positive or negative) were generated. Sensitivities of all the 64 autoantibody markers for detecting CRC were calculated. Additionally, multi-marker combinations were also explored to enhance the diagnostic performance. Multi-marker combinations were considered positive, if at least one marker in the combination was positive. A systematic search for the best n-marker combination (starting with n=2) was conducted in the following steps: i) calculate the Youden index, equaling the sum of sensitivity for detecting early stage CRC (UICC TNM stage I/II) and specificity minus 1 for all possible marker combinations; ii) determine the specific n-marker combination which gives the highest index and evaluate its diagnostic performance in the training set; iii) continue the selection process with increasing n, the number of markers included in the combination, until the specificity of the best performing marker combination in the training set falls below 90%. The diagnostic performance of the final marker-combination was then evaluated in the validation set. 95% confidence intervals (95% CIs) of sensitivity and specificity were calculated using the Wilson method [49].
All statistical analyses were performed with the statistical software R version 3.1.1 [50]. All tests were two-sided and p-values of 0.05 or less were considered to be statistically significant.
ACKNOWLEDGMENT
We gratefully acknowledge the excellent cooperation of gastroenterology practices and clinics in patient recruitment and of Labor Limbach in sample collection. We gratefully acknowledge Dr. Katja Butterbach and Ulrike Schlesselmann for their excellent work in laboratory preparation of blood samples and thank Ute Koch and Monika Oppenländer for excellent technical assistance. We also gratefully acknowledge Isabel Lerch, Susanne Köhler, Utz Benscheid, Jason Hochhaus and Maria Kuschel for their contribution in data collection, monitoring and documentation.
CONFLICTS OF INTEREST
The authors disclose no potential conflicts of interest.
GRANT SUPPORT
The BliTz study was partly funded by grants from the German Research Council (DFG, grant No. BR1704/16-1). The autoantibody measurements were supported by iMed - the Helmholtz Initiative on Pesonalized Medicine. The work of Hongda Chen was partly supported by China Scholarship Council (CSC). Inka Zörnig received funding from the Wilhelm Sander Foundation (Grant No. 2008.091.1).
REFERENCES
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J and Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87-108.
2. Siegel R, Desantis C and Jemal A. Colorectal cancer statistics, 2014. CA Cancer J Clin. 2014; 64:104-117.
3. Brenner H, Kloor M and Pox CP. Colorectal cancer. Lancet (London, England). 2014; 383:1490-1502.
4. Hewitson P, Glasziou P, Watson E, Towler B and Irwig L. Cochrane systematic review of colorectal cancer screening using the fecal occult blood test (hemoccult): an update. The American journal of gastroenterology. 2008; 103:1541-1549.
5. Zauber AG, Winawer SJ, O’Brien MJ, Lansdorp-Vogelaar I, van Ballegooijen M, Hankey BF, Shi W, Bond JH, Schapiro M, Panish JF, Stewart ET and Waye JD. Colonoscopic polypectomy and long-term prevention of colorectal-cancer deaths. The New England journal of medicine. 2012; 366:687-696.
6. Brenner H, Stock C and Hoffmeister M. Effect of screening sigmoidoscopy and screening colonoscopy on colorectal cancer incidence and mortality: systematic review and meta-analysis of randomised controlled trials and observational studies. BMJ (Clinical research ed). 2014; 348:g2467.
7. Zorzi M, Fedeli U, Schievano E, Bovo E, Guzzinati S, Baracco S, Fedato C, Saugo M and Dei Tos AP. Impact on colorectal cancer mortality of screening programmes based on the faecal immunochemical test. Gut. 2015; 64:784-790.
8. Symonds EL, Pedersen S, Cole SR, Massolino J, Byrne D, Guy J, Backhouse P, Fraser RJ, LaPointe L and Young GP. Improving Participation in Colorectal Cancer Screening: a Randomised Controlled Trial of Sequential Offers of Faecal then Blood Based Non-Invasive Tests. Asian Pacific journal of cancer prevention. 2015; 16:8455-8460.
9. Adler A, Geiger S, Keil A, Bias H, Schatz P, deVos T, Dhein J, Zimmermann M, Tauber R and Wiedenmann B. Improving compliance to colorectal cancer screening using blood and stool based tests in patients refusing screening colonoscopy in Germany. BMC gastroenterology. 2014; 14:183.
10. Heo CK, Bahk YY and Cho EW. Tumor-associated autoantibodies as diagnostic and prognostic biomarkers. BMB reports. 2012; 45:677-685.
11. Werner S, Chen H, Tao S and Brenner H. Systematic review: Serum autoantibodies in the early detection of gastric cancer. International journal of cancer. 2015; 136:2243-2252.
12. Anderson KS, Cramer DW, Sibani S, Wallstrom G, Wong J, Park J, Qiu J, Vitonis A and LaBaer J. Autoantibody signature for the serologic detection of ovarian cancer. Journal of proteome research. 2015; 14:578-586.
13. Lacombe J, Mange A and Solassol J. Use of autoantibodies to detect the onset of breast cancer. Journal of immunology research. 2014; 2014:574981.
14. Chen H, Werner S, Tao S, Zornig I and Brenner H. Blood autoantibodies against tumor-associated antigens as biomarkers in early detection of colorectal cancer. Cancer letters. 2014; 346:178-187.
15. Pedersen JW, Gentry-Maharaj A, Nostdal A, Fourkala EO, Dawnay A, Burnell M, Zaikin A, Burchell J, Papadimitriou JT, Clausen H, Jacobs I, Menon U and Wandall HH. Cancer-associated autoantibodies to MUC1 and MUC4--a blinded case-control study of colorectal cancer in UK collaborative trial of ovarian cancer screening. International journal of cancer. 2014; 134:2180-2188.
16. Pedersen JW, Gentry-Maharaj A, Fourkala EO, Dawnay A, Burnell M, Zaikin A, Pedersen AE, Jacobs I, Menon U and Wandall HH. Early detection of cancer in the general population: a blinded case-control study of p53 autoantibodies in colorectal cancer. British journal of cancer. 2013; 108:107-114.
17. Werner S, Krause F, Rolny V, Strobl M, Morgenstern D, Datz C, Chen H and Brenner H. Evaluation of a 5-Marker Blood Test for Colorectal Cancer Early Detection in a Colorectal Cancer Screening Setting. Clin Cancer Res. 2016; 22:1725-33
18. Kanojia D, Garg M, Gupta S, Gupta A and Suri A. Sperm-associated antigen 9 is a novel biomarker for colorectal cancer and is involved in tumor growth and tumorigenicity. The American journal of pathology. 2011; 178:1009-1020.
19. Chen JS, Kuo YB, Chou YP, Chan CC, Fan CW, Chen KT, Huang YS and Chan EC. Detection of autoantibodies against Rabphilin-3A-like protein as a potential biomarker in patient’s sera of colorectal cancer. Clinica chimica acta. 2011; 412:1417-1422.
20. Fan CW, Chan CC, Chen KT, Twu J, Huang YS, Han CL, Chen YJ, Yu JS, Chang YS, Kuo YB and Chan EC. Identification of SEC61beta and its autoantibody as biomarkers for colorectal cancer. Clinica chimica acta. 2011; 412:887-893.
21. Chen JS, Chou YP, Chen KT, Hung RP, Yu JS, Chang YS and Chan EC. Detection of annexin A autoantibodies in sera from colorectal cancer patients. Journal of clinical gastroenterology. 2011; 45:125-132.
22. Tao S, Hundt S, Haug U and Brenner H. Sensitivity estimates of blood-based tests for colorectal cancer detection: impact of overrepresentation of advanced stage disease. The American journal of gastroenterology. 2011; 106:242-253.
23. Brenner H, Altenhofen L and Tao S. Matching of controls may lead to biased estimates of specificity in the evaluation of cancer screening tests. Journal of clinical epidemiology. 2013; 66:202-208.
24. Pesch B, Bruning T, Johnen G, Casjens S, Bonberg N, Taeger D, Muller A, Weber DG and Behrens T. Biomarker research with prospective study designs for the early detection of cancer. Biochimica et biophysica acta. 2014; 1844:874-883.
25. Ransohoff DF. How to improve reliability and efficiency of research about molecular markers: roles of phases, guidelines, and study design. Journal of clinical epidemiology. 2007; 60:1205-1219.
26. Waterboer T, Sehr P and Pawlita M. Suppression of non-specific binding in serological Luminex assays. Journal of immunological methods. 2006; 309:200-204.
27. Simpson AJ, Caballero OL, Jungbluth A, Chen YT and Old LJ. Cancer/testis antigens, gametogenesis and cancer. Nature reviews Cancer. 2005; 5:615-625.
28. Marcar L, Maclaine NJ, Hupp TR and Meek DW. Mage-A cancer/testis antigens inhibit p53 function by blocking its interaction with chromatin. Cancer research. 2010; 70:10362-10370.
29. Chapman CJ, Healey GF, Murray A, Boyle P, Robertson C, Peek LJ, Allen J, Thorpe AJ, Hamilton-Fairley G, Parsy-Kowalska CB, MacDonald IK, Jewell W, Maddison P and Robertson JF. EarlyCDT(R)-Lung test: improved clinical utility through additional autoantibody assays. Tumour biology. 2012; 33:1319-1326.
30. Yakirevich E, Sabo E, Lavie O, Mazareb S, Spagnoli GC and Resnick MB. Expression of the MAGE-A4 and NY-ESO-1 cancer-testis antigens in serous ovarian neoplasms. Clinical cancer research. 2003; 9:6453-6460.
31. Vousden KH and Prives C. P53 and prognosis: new insights and further complexity. Cell. 2005; 120:7-10.
32. Gnjatic S, Nishikawa H, Jungbluth AA, Gure AO, Ritter G, Jager E, Knuth A, Chen YT and Old LJ. NY-ESO-1: review of an immunogenic tumor antigen. Advances in cancer research. 2006; 95:1-30.
33. Cubiella J, Castro I, Hernandez V, Gonzalez-Mao C, Rivera C, Iglesias F, Cid L, Soto S, de-Castro L, Vega P, Hermo JA, Macenlle R, Martinez A, Martinez-Ares D, Estevez P, Cid E, et al. Characteristics of adenomas detected by fecal immunochemical test in colorectal cancer screening. Cancer epidemiology, biomarkers & prevention. 2014; 23:1884-1892.
34. Hundt S, Haug U and Brenner H. Comparative evaluation of immunochemical fecal occult blood tests for colorectal adenoma detection. Annals of internal medicine. 2009; 150:162-169.
35. Brenner H and Tao S. Superior diagnostic performance of faecal immunochemical tests for haemoglobin in a head-to-head comparison with guaiac based faecal occult blood test among 2235 participants of screening colonoscopy. European journal of cancer (Oxford, England : 1990). 2013; 49:3049-3054.
36. Parra-Blanco A, Gimeno-Garcia AZ, Quintero E, Nicolas D, Moreno SG, Jimenez A, Hernandez-Guerra M, Carrillo-Palau M, Eishi Y and Lopez-Bastida J. Diagnostic accuracy of immunochemical versus guaiac faecal occult blood tests for colorectal cancer screening. Journal of gastroenterology. 2010; 45:703-712.
37. Church TR, Wandell M, Lofton-Day C, Mongin SJ, Burger M, Payne SR, Castanos-Velez E, Blumenstein BA, Rosch T, Osborn N, Snover D, Day RW, Ransohoff DF, Presept Clinical Study Steering Committee I and Study T. Prospective evaluation of methylated SEPT9 in plasma for detection of asymptomatic colorectal cancer. Gut. 2014; 63:317-325.
38. Hanahan D and Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011; 144:646-674.
39. Pox CP, Altenhofen L, Brenner H, Theilmeier A, Von Stillfried D and Schmiegel W. Efficacy of a nationwide screening colonoscopy program for colorectal cancer. Gastroenterology. 2012; 142:1460-1467.e1462.
40. Brenner H, Chang-Claude J, Rickert A, Seiler CM and Hoffmeister M. Risk of colorectal cancer after detection and removal of adenomas at colonoscopy: population-based case-control study. Journal of clinical oncology. 2012; 30:2969-2976.
41. Brenner H, Tao S and Haug U. Low-dose aspirin use and performance of immunochemical fecal occult blood tests. Jama. 2010; 304:2513-2520.
42. Pox C and Schmiegel W. Colorectal screening in Germany. Zeitschrift fur Gastroenterologie. 2008; 46:S31-32.
43. Pox C, Schmiegel W and Classen M. Current status of screening colonoscopy in Europe and in the United States. Endoscopy. 2007; 39:168-173.
44. Brenner H, Altenhofen L, Kretschmann J, Rosch T, Pox C, Stock C and Hoffmeister M. Trends in Adenoma Detection Rates During the First 10 Years of the German Screening Colonoscopy Program. Gastroenterology. 2015; 149:356-366.e351.
45. Tao S, Haug U, Kuhn K and Brenner H. Comparison and combination of blood-based inflammatory markers with faecal occult blood tests for non-invasive colorectal cancer screening. British journal of cancer. 2012; 106:1424-1430.
46. Zornig I, Halama N, Lorenzo Bermejo J, Ziegelmeier C, Dickes E, Migdoll A, Kaiser I, Waterboer T, Pawlita M, Grabe N, Ugurel S, Schadendorf D, Falk C, Eichmuller SB and Jager D. Prognostic significance of spontaneous antibody responses against tumor-associated antigens in malignant melanoma patients. International journal of cancer. 2015; 136:138-151.
47. Waterboer T, Sehr P, Michael KM, Franceschi S, Nieland JD, Joos TO, Templin MF and Pawlita M. Multiplex human papillomavirus serology based on in situ-purified glutathione s-transferase fusion proteins. Clinical chemistry. 2005; 51:1845-1853.
48. Sehr P, Zumbach K and Pawlita M. A generic capture ELISA for recombinant proteins fused to glutathione S-transferase: validation for HPV serology. Journal of immunological methods. 2001; 253:153-162.
49. Brown LD, Cai TT and DasGupta A. Interval estimation for a binomial proportion (with discussion). Statistical Science. 2001; 16:101-133.
50. R Core Team (2014). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/.

