INTRODUCTION
MicroRNAs (miRNAs) are endogenous small (~22 nucleotides) non-coding RNAs that repress target gene expression by complementary binding to the 3’-untranslated region (3’-UTR) of messenger RNAs (mRNAs), leading to translational repression or mRNA cleavage [1, 2]. The miRNAs play important roles in various biological functions such as cell proliferation and survival, DNA repair, and immune response [3–5]. Evidence indicates that miRNAs are critically involved in the development and progression of diverse human cancers as tumor suppressors and/or oncogenes depending on the tissue type and the presence of specific targets [5–8].
Many studies have suggested that single nucleotide polymorphisms (SNPs) in miRNA target sites are associated with the risk and the prognosis of diverse types of cancer, including lung cancer [9–12]. Most of the studies investigated miRNA binding sites identified by computational prediction methods such as miRanda and TargetScan [13–15]. However, those were developed to predict miRNA-mRNA binding based primarily on the complementarity to seed sequence, being biased towards canonical seed pairings at 3’ UTR, and may have frequent false prediction [13–15]. Recently, crosslinking, ligation, and sequencing of hybrids (CLASH) provided direct experimental observation of transcriptome-wide miRNA-target pairs, revealing that the interactions occurred more frequently in coding sequence than 3’ UTR and the majority of miRNA-target bindings were noncanonical [13].
Based on the important roles of miRNA network in carcinogenesis, we hypothesized that polymorphisms in miRNA target sites may influence miRNA-mRNA binding and consequently the expression of target genes, thereby influencing the response and survival after chemotherapy in lung cancer. To test this hypothesis, we selected SNPs in miRNA binding sites using CLASH data and evaluated their association with the clinical outcome of first line paclitaxel-cisplatin chemotherapy in non-small cell lung cancer (NSCLC).
RESULTS
Patient characteristics and clinical predictors
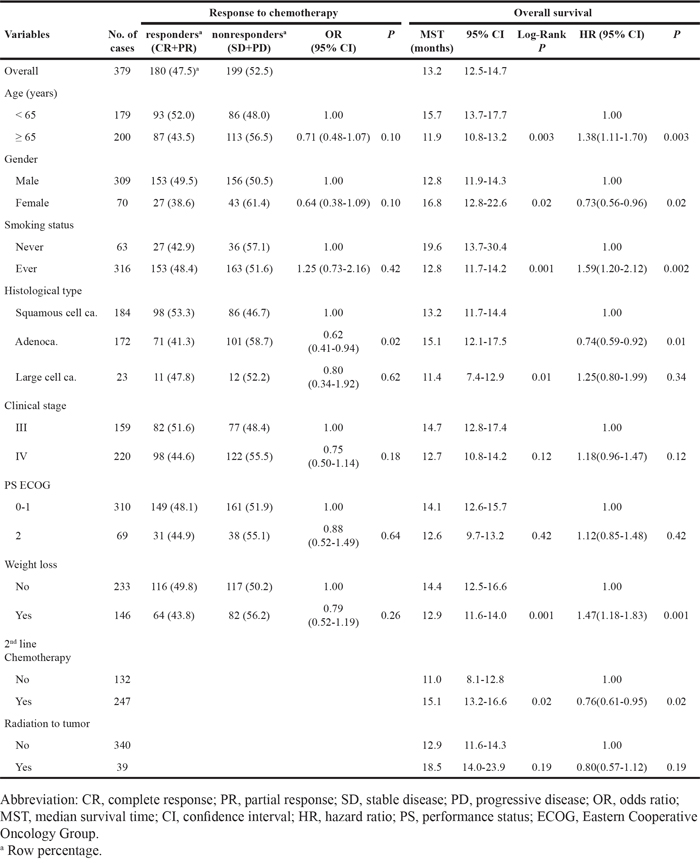
The clinico-pathological characteristics and the association with chemotherapy response and OS are shown in Table 1. The overall response rate was 47.5%. We observed events (deaths) in 347 of the 379 patients (91.6%) and median survival time (MST) was 13.2 months (95% CI = 12.5–14.7 months). The response to chemotherapy was not associated with any of the clinical characteristics. Age, gender, smoking status, tumor histology, weight loss, and second line chemotherapy were significantly associated with the OS (Table 1).
Table 1: Univariate analysis for response to chemotherapy and overall survival by clinical variables
Associations between SNPs and clinical outcomes
Among the 100 SNPs genotyped, 80 were analyzed in this study after excluding 5 SNPs with genotype failure and 15 showing deviation from Hardy-Weinberg equilibrium (P < 0.05). The SNP ID, gene information, miRNA, and minor allele frequencies are shown in Supplementary Table 1. Of the 80 SNPs analyzed, 16 SNPs listed in Table 2 were significantly associated with chemotherapy response and/or survival. Among these, ANAPC1 rs3814026C>T, ETS2 rs461155A>G, SORBS1 rs7081076C>A and POLR2A rs2071504C>T were found to be associated with both chemotherapy response and survival (adjusted OR [aOR] = 0.61, 95% CI = 0.37–1.00, P = 0.05; adjusted HR [aHR] = 1.41, 95% CI = 1.10–1.82, P = 0.008, under recessive model, respectively; aOR = 1.34, 95% CI = 1.00–1.81, P = 0.05; aHR = 0.80, 95% CI = 0.69–0.94, P = 0.006, under additive model, respectively; aOR = 2.32, 95% CI = 1.28–4.24, P = 0.006; aHR = 0.67, 95% CI = 0.49–0.91, P = 0.009, under dominant model; aOR = 0.17, 95% CI = 0.04–0.78, P = 0.02; aHR = 1.83, 95% CI = 1.03–3.26, P = 0.04, under recessive model, respectively; Table 3 and Figure 1).
Table 2: Summary of sixteen SNPs and response to chemotherapy and overall survival
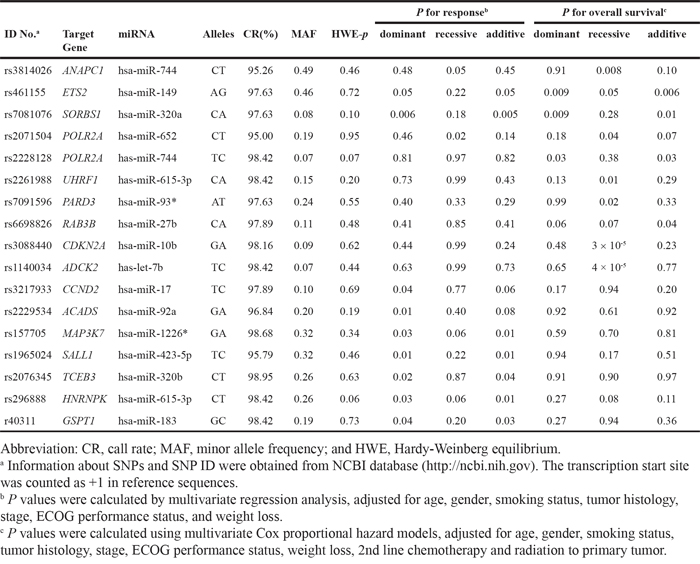
Table 3: Genotypes of ANAPC1, ETS2, SORBS1, and POLR2A polymorphisms and their associations with the response to chemotherapy and overall survival
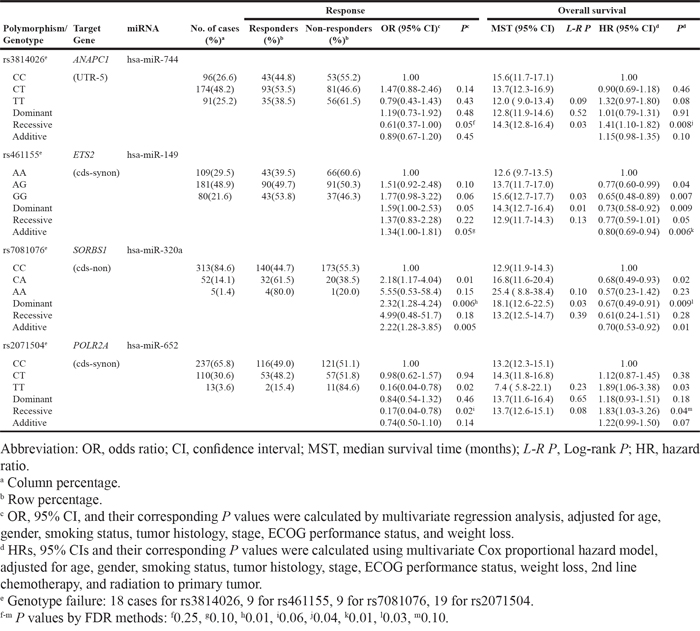
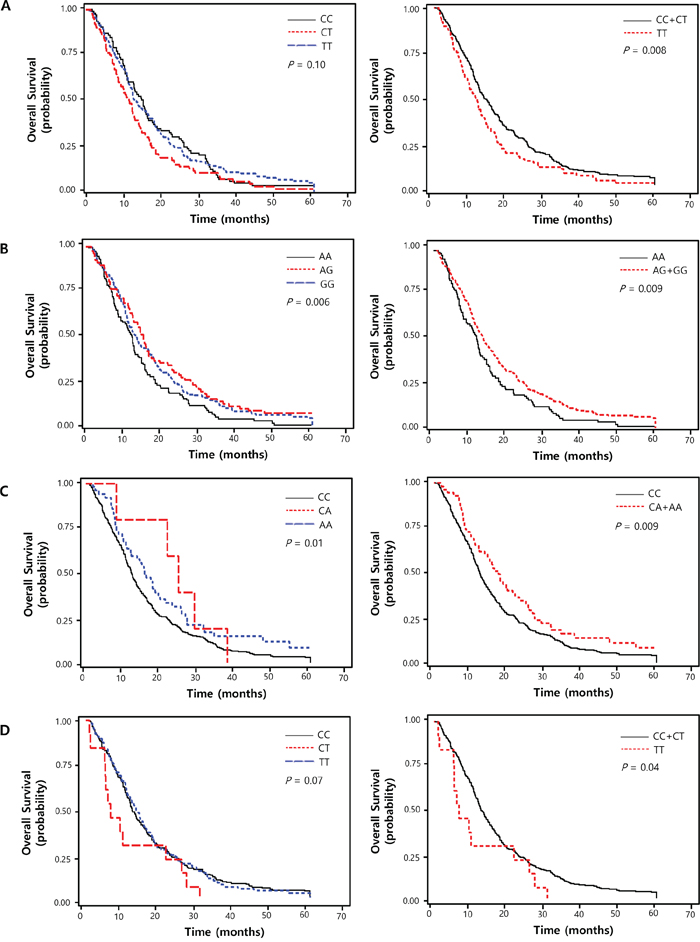
Figure 1: Kaplan-Meier plot of overall survival curves according to (A) ANAPC1 rs3814026C>T, (B) ETS2 rs461155A>G, (C) SORBS1 rs7081076C>A and (D) POLR2A rs2071504C>T genotypes. P values in multivariate Cox proportional hazard model.
Effect of SNPs in microRNA target sites on mRNA expression
To identify the functional effect of ANAPC1 rs3814026C>T, ETS2 rs461155A>G, SORBS1 rs7081076C>A and POLR2A rs2071504C>T, we evaluated the relationship between the genotypes of those SNPs and mRNA expression of each gene in tumor and paired non-malignant lung tissues. As shown in Figure 2A, ANAPC1 expression level was significantly higher (P = 2 × 10−11), and ETS2 and SORBS1 expression level was significantly lower in tumor tissues than in non-malignant tissues (P = 5 × 10−4 and 1 × 10−8), respectively. However, POLR2A expression level was not different between tumor and normal tissues. Notably, ETS2 rs461155A>G was significantly associated with decreased ETS2 mRNA expression in both tumor and paired normal tissues (Ptrend = 4 × 10−7, and 3 × 10−4, respectively, Figure 2B). The difference among genotypes was not observed in ANAPC1, SORBS1 and POLR2A expression (data not shown).
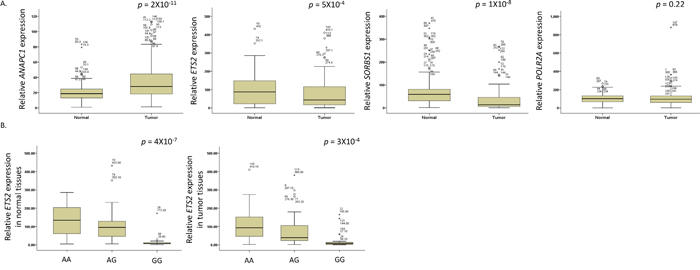
Figure 2: (A) The mRNA expression levels of ANAPC1, ETS2, SORBS1, and POLR2A genes and (B) ETS2 mRNA expression by the rs461155A>G genotypes, in tumor and non-malignant lung tissues. The ETS2 mRNA expression levels and the association with the rs461155A>G genotypes were examined in tumor (30 AA, 62 AG, and 23 GG) and paired non-malignant lung (37 AA, 67 AG, and 21 GG) tissues. The mRNA expression levels of each gene were normalized with that of β-actin gene. The horizontal lines within the boxes represent the median values; the upper and lower boundaries of the boxes represent 75th and 25th percentiles, respectively; the upper and lower bars indicate the largest and smallest observed values, respectively, except outliers. P values, Student’s t-test.
Effect of SNPs in miRNA target sites on miRNA binding
To investigate whether ETS2 rs461155A>G, a synonymous SNP at coding sequence, modulates the binding of miR-149, and thereby alter the expression of ETS2 gene, we generated psiCHECK™-2-ETS2 constructs containing rs461155A>G and co-transfected the constructs into A549 and H1299 cells with miR-149. CLASH data showed that the binding between ETS2 and miR-149 was noncanonical. The rs461155 caused an A-to-G change at the position which pairs with nucleotide 11 of miR-149, outside the seed region (Figure 3A). As shown in Figure 3B and C, the Renilla luciferase activity was significantly decreased in ETS2 rs461155G compared with ETS2 rs461155A in both A549 and H1299 cells (P = 0.02 and 0.05, respectively), which was consistent with the mRNA expression. These results suggest that rs461155A>G may lead to decreased ETS2 expression by altering the binding of miR-149 to ETS2 mRNA.
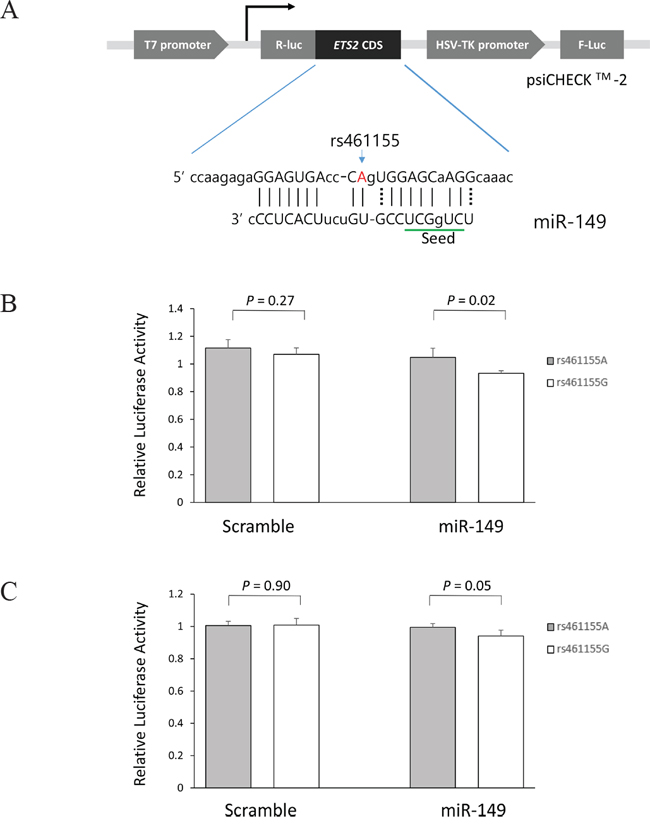
Figure 3: Functional analysis of the ETS2 rs461155A>G by dual luciferase reporter assay. (A) Schematic representation of reporter plasmids containing ETS2 rs461155A>G, and the complementarity between miR-149 and ETS2 coding sequence targeted. Renilla luciferase assay for miRNA effect on rs461155A>G polymorphism using (B) and H1299 cells (C) Cells were co-transfected with hsa-miR-149-5p, and psiCHECK2 plasmid containing ETS2 296 bp coding sequence fragments with A or G allele. Renilla luciferase activity was normalized to firefly luciferase activity and data are presented relative to the Mock control. Each bar represents mean ± SE from two independent experiments carried out in quadruplicate. P-value, Student’s t-test. R-luc, Renilla luciferase; F-luc, firefly luciferase; HSV-TK, Herpes simplex virus thymidine kinase.
DISCUSSION
We investigated the associations between SNPs in miRNA target sites and the treatment outcomes of 1st line paclitaxel-cisplatin chemotherapy, to identify genetic variations that affect the clinical outcomes in NSCLC. Using the CLASH data that provides experimentally proved transcriptome-wide miRNA-target pairs, 80 potentially functional SNPs in miRNA binding sites of cancer-related genes were tested for this study. Four SNPs, ANAPC1 rs3814026C>T, ETS2 rs461155A>G, SORBS1 rs7081076C>A, and POLR2A rs2071504C>T, could predict both chemotherapy response and survival. Notably, functional evidence for ETS2 rs461155A>G was provided by ETS2 mRNA expression in clinical samples, which was further supported by in vitro assays using lung cancer cell lines. These findings suggest that the SNPs, especially ETS2 rs461155A>G, could be used as predictive/prognostic markers for advanced stage NSCLC receiving paclitaxel-cisplatin chemotherapy. This is the first study investigating the effect of miRNA binding site polymorphisms on chemotherapy outcome in lung cancer.
In the present study, four SNPs in miRNA binding sites were associated with chemotherapy response and survival in patients with advanced stage NSCLC. Among those, the ETS2 rs461155A>G was strongly correlated with decreased ETS2 mRNA expression level in both tumor and normal lung tissues. In addition, in vitro luciferase assay suggested that the ETS2 rs461155 A-to-G change could alter binding efficiency of miR-149 to ETS2 mRNA, leading to decreased ETS2 mRNA expression level and better clinical outcomes of lung cancer. However, the biologic mechanism of the observed association between the SNP and the clinical outcomes remains unclear because the role of ETS2 in lung cancer has not been fully elucidated. ETS2 is a transcription factor belonging to the ETS family proteins which control gene expression by binding to over 200 target genes with GGAA/T ETS binding site. ETS2 controls the expression of genes that are critical for a wide range of biological processes including cellular proliferation, differentiation, transformation, angiogenesis, apoptosis, and Ras signal transduction [16–18], and is also required for telomerase activity [19]. It has been reported that ETS2 functions as both oncogene [18, 20–23] and tumor-suppressor gene [24–27] in different types of malignancies. Accumulating evidence suggests that the same ETS factor may function either as a positive or as a negative regulator of target gene transcription [16, 28]. Based on the target DNA sequence, binding of an ETS protein near other transcription factors or additional tissue-specific factors and the consequent interaction between them may result in either synergistic activation or repression of specific target genes, suggesting promoter and cell context specificity [29]. Recently, Kabbout et al. reported that ETS2 exerts a tumor suppressor function in NSCLC pathogenesis through inhibition of the MET proto-oncogene, and that low ETS2 expression by immunohistochemistry was a significant predictor of shorter time to recurrence after curative resection in early stage NSCLC [27]. In the present study, however, ETS2 rs461155 variant G allele, which correlated to decreased expression level of ETS2 mRNA compared with A allele, was associated with better chemotherapy response and survival of the patients with advanced NSCLC. Given the different settings of the two studies, one possible explanation is that the discordant observation could be attributable to the effect of ETS2 on its target genes involved in DNA repair or cell cycle regulation, because alteration of those genes may lead to discordant outcomes in terms of cancer progression and chemotherapy response [30–33]. For example, DNA repair genes have been linked to protection against development and progression of cancer, thereby better prognosis, whereas they have also been associated with resistance to platinum-based anticancer drugs and poor prognosis [30–31]. Further investigation is needed to understand the role of ETS2 in lung cancer and the mechanism of its dissimilar effects on the prognosis of patients in different stages undergoing different anticancer therapy.
Using CLASH data for the association study, this study provides a unique opportunity to evaluate SNPs in miRNA targets outside 3’ UTR and in noncanonical miRNA-mRNA binding in terms of functional and clinical consequences in NSCLC. Although the four SNPs were associated with chemotherapy response and survival, considering the borderline CIs of some associations and multiple comparisons issue with a modest sample size, the impact of the SNPs on chemotherapy outcomes would be marginal. However, our study provides functional evidence for the association of ETS2 rs461155A>G with clinical outcomes by showing its strong correlation with ETS2 mRNA expression in clinical samples as well as in vitro assays using lung cancer cell lines, which supports the credibility of the association. Further studies are warranted to validate our results in a larger population with diverse ethnicity.
In conclusion, this study suggests that the four SNPs, especially ETS2 rs461155A>G, could be used as biomarkers predicting the clinical outcomes of NSCLC patients treated with first-line paclitaxel-cisplatin chemotherapy. Future studies are needed to confirm our findings and to understand the biologic function of ETS2 in the development and progression of lung cancer.
MATERIALS AND METHODS
Study population and chemotherapy
The study population has been described in our previous study [34]. In brief, 379 patients with stage III or IV NSCLC treated with at least two cycles of paclitaxel-cisplatin chemotherapy as a first-line treatment at Kyungpook National University Hospital (KNUH) in Daegu, Korea between August 2005 and December 2008. Genomic DNA samples from the patients were provided by the National Biobank of Korea, KNUH, which is supported by the Ministry of Health, Welfare and Family Affairs. Patients who underwent concurrent chemoradiotherapy were excluded to avoid the confounding effect of radiation on chemotherapy response. The chemotherapy regimen consisted of paclitaxel 175 mg/m2 administered i.v. over three hours, and cisplatin 60 mg/m2 infused over one hours on day 1, every three weeks. Treatment was discontinued upon disease progression, major toxicities, or according to patient decision or physician discretion. Assessment of tumor response was performed by computed tomography scan every two cycles. Responses were assessed using Response Evaluation Criteria in Solid Tumors [35]. The best overall response for each patient was reported and all responses were reviewed by an independent radiologist. Patients with a complete response (CR) or a partial response (PR) were defined as responders, and patients having stable disease (SD) or progressive disease (PD) were defined as nonresponders. For the assessment of survival outcomes, overall survival (OS), defined as the time between the date of chemotherapy start and the date of death or last follow-up, were recorded. There were 184 squamous cell carcinomas (SCC), 172 adenocarcinomas (AC), 3 large cell carcinomas, and 20 cases classified as unspecified NSCLC. Fifty nine patients had stage IIIA disease, 100 stage IIIB, and 220 stage IV. Written informed consent was obtained from all patients and this study was approved by the Institutional Review Board of the KNUH.
SNP selection and genotyping
We searched for all the potentially functional polymorphisms in miRNA target sites using PolymiRTS database 3.0 (http://compbio.uthsc.edu/miRSNP; ref. 36), and selected 24,027 SNPs in experimentally validated miRNA target sites by downloading data from CLASH experiment which has been integrated in PolymiRTS database 3.0 (last modified in July 2013). Among those, 1,574 SNPs in cancer-related genes were selected using a downloaded list of cancer genes from the CancerGenes database (http://cbio.mskcc.org/cancergenes; ref. 37). Finally 100 SNPs with the minor allele frequency (MAF) ≥ 0.05 in the HapMap JPT data were collected after excluding those in linkage disequilibrium (LD, r2 ≥ 0.8). Genotyping was performed using Sequenom MassARRAY® iPLEX assay (Sequenom Inc., San Diego, CA). For validation of genotyping, approximately 5% of samples were randomly selected and genotyped again with a restriction fragment length polymorphism assay by a different investigator and the results were 100% concordant.
RNA preparation and quantitative reverse transcription-PCR
Anaphase promoting complex subunit 1 (ANAPC1), V-ets avian erythroblastosis virus E26 oncogene homolog 2 (ETS2), Sorbin and SH3 domain containing 1 (SORBS1), and Polymerase (RNA) II (DNA directed) polypeptide A, 220kDa (POLR2A) mRNA expression was measured by quantitative reverse transcription-PCR. Total RNA was isolated from paired tumor and nonmalignant lung tissues of 154 NSCLC patients who underwent surgery in Kyungpook National University Medical Center between September 2011 and August 2014 using TRIzol (Invitrogen, Carlsbad, CA) and reverse transcribed using the QuantiTect reverse transcription kit (QIAGEN, Hilden, Germany). Real time-PCR was performed for each gene and beta-actin with QuantiFast SYBR Green PCR Master Mix (QIAGEN) in a LightCycler 480 (Roche Applied Science, Mannheim, Germany) using the following primers: ANAPC1 forward, 5′-CGCGTCCCGAGTTATACAG-3′; ANAPC1 reverse, 5′-TCTCGACCAAAAGGAACAAATTC-3′; ETS2 forward, 5′-CAGATGTTCCCCAAGTCTCG-3′; ETS2 reverse, 5′-GTCGTGGTCTTTGGGAGTC-3′; SORBS1 forward, 5′-CAGTTCAGAGCCCCAGTTG-3′; SORBS1 reverse, 5′-CTTTGTCTTGCCCATTGCTG-3′; POLR2A forward, 5′-CAAGTATGGCATGGAGATCCC-3′; POLR2A reverse, 5′-TGGTTCCAAGGTGTCATGG-3′; beta-actin forward, 5′-TTGTTACAGGAAGTCCCT TGCC-3′; beta-actin reverse, 5′-ATGCTATCACC TCCCCTGTGT-3′. Each test was carried out according to standard protocol. Relative each gene expression was calculated following normalization with human beta actin. The relative mRNA expression were normalized with β-actin expression and then calculated by the 2−ΔΔCT method [38].
Cloning of the luciferase reports gene and dual luciferase assay
We investigated whether miR-149, which was reportedly bind to ETS2 mRNA in CLASH experiment [13] and known to be expressed in lung cancer [39, 40], changes the ETS2 expression level on its binding sites including rs461155A>G by luciferase report assay. The psiCHECK™-2 vector (Promega, Madison, WI, USA) was used to construct luciferase reporter plasmids. Briefly, the putative target site ETS2 coding sequence containing rs461155A or rs461155G was synthesized by PCR from human genomic DNA and cloned into the psiCHECK™-2 vector. The construct was confirmed by DNA sequencing. The NSCLC cell lines, H1299 and A549, were purchased from Korean Cell Line Bank (KCLB), Seoul, Korea, and authenticated by KCLB using short tandem repeat DNA fingerprinting. The cells were plated in 24-well plates at a density of 5x104 cells/well. The following day, cells were transfected with construct vector and 30 nM miScript miRNA Mimic (Syn-hsa-miR-149-5p; QIAGEN, Hilden, Germany) or AllStars negative control siRNA (QIAGEN, Hilden, Germany) based on the manufacturer’s instructions. After 24 h of incubation, Firefly and Renilla luciferase activities were measured using the Dual-Luciferase® Reporter Assay System (Promega, Madison, WI, USA). The Renilla luciferase values were normalized using firefly luciferase activity and data are presented relative to the Mock control. All experiments were performed twice in quadruplicate.
Statistical analysis
Hardy-Weinberg equilibrium was tested using a goodness-of-fit χ2 test with 1 degree of freedom. The genotypes for each SNP were analyzed as a three-group categorical variable, and those were also grouped according to the dominant and recessive model. The association between clinical variables or genotypes and chemotherapy response was tested by odds ratio (OR) and 95% confidence intervals (CIs) using unconditional logistic regression analysis. The survival estimates were calculated using the Kaplan-Meier method. The difference in OS according to different clinical variables or genotypes was compared using log-rank tests. Cox’s proportional hazard regression model was used for the multivariate survival analyses. The hazard ratio (HR) and 95% confidence interval (CI) were also estimated. A cut-off p-value of 0.05 was adopted for all the statistical analyses. The statistical data were obtained using Statistical Analysis System for Windows, version 9.2 (SAS Institute, Cary, NC, USA).
ACKNOWLEDGMENTS
This work was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (NRF-2013R1A1A1009443), in part by the National Research Foundation of Korea (NRF) grant funded by the Korea government (2014R1A5A2009242).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R. Control of translation and mRNA degradation by miRNAs and siRNAs. Genes & development. 2006; 20:515–524.
2. Brodersen P, Voinnet O. Revisiting the principles of microRNA target recognition and mode of action. Nat Rev Mol Cell Biol. 2009; 10:141–148.
3. Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic acids research. 2006; 34:D140–144.
4. Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005; 120:15–20.
5. Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer. 2006; 6:259–269.
6. Fabbri M, Ivan M, Cimmino A, Negrini M, Calin GA. Regulatory mechanisms of microRNAs involvement in cancer. Expert Opin Biol Ther. 2007; 7:1009–1019.
7. Guan P, Yin Z, Li X, Wu W, Zhou B. Meta-analysis of human lung cancer microRNA expression profiling studies comparing cancer tissues with normal tissues. J Exp Clin Cancer Res. 2012; 31:54.
8. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006; 6:857–866.
9. Landi D, Gemignani F, Naccarati A, Pardini B, Vodicka P, Vodickova L, Novotny J, Försti A, Hemminki K, Canzian F, Landi S. Polymorphisms within micro-RNA-binding sites and risk of sporadic colorectal cancer. Carcinogenesis. 2008; 29:579–584.
10. Chin LJ, Ratner E, Leng S, Zhai R, Nallur S, Babar I, Muller RU, Straka E, Su L, Burki EA, Crowell RE, Patel R, Kulkarni T, et al. A SNP in a let-7 microRNA complementary site in the KRAS 3’ untranslated region increases non-small cell lung cancer risk. Cancer research. 2008; 68:8535–8540.
11. Xu J, Yin Z, Gao W, Liu L, Yin Y, Liu P, Shu Y. Genetic Variation in a MicroRNA-502 Minding Site in SET8 Gene Confers Clinical Outcome of Non-Small Cell Lung Cancer in a Chinese Population. PloS one. 2013; 8:e77024.
12. Teo MT, Landi D, Taylor CF, Elliott F, Vaslin L, Cox DG, Hall J, Landi S, Bishop DT, Kiltie AE. The role of microRNA-binding site polymorphisms in DNA repair genes as risk factors for bladder cancer and breast cancer and their impact on radiotherapy outcomes. Carcinogenesis. 2012; 33:581–586.
13. Helwak A, Kudla G, Dudnakova T, Tollervey D. Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell. 2013; 153:654–665.
14. Liu C, Mallick B, Long D, Rennie WA, Wolenc A, Carmack CS, Ding Y. CLIP-based prediction of mammalian microRNA binding sites. Nucleic acids research. 2013; 41:e138.
15. Ørom UA, Lund AH. Experimental identification of microRNA targets. Gene. 2010; 451:1–5.
16. Hsu T, Trojanowska M, Watson DK. Ets proteins in biological control and cancer. Journal of cellular biochemistry. 2004; 91:896–903.
17. Seth A, Watson DK. ETS transcription factors and their emerging roles in human cancer. European journal of cancer. 2005; 41:2462–2478.
18. Yang BS, Hauser CA, Henkel G, Colman MS, Van Beveren C, Stacey KJ, Hume DA, Maki RA, Ostrowski MC. Ras-mediated phosphorylation of a conserved threonine residue enhances the transactivation activities of c-Ets1 and c-Ets2. Molecular and cellular biology. 1996; 16:538–547.
19. Xu D, Dwyer J, Li H, Duan W, Liu JP. Ets2 maintains hTERT gene expression and breast cancer cell proliferation by interacting with c-Myc. The Journal of biological chemistry. 2008; 283:23567–23580.
20. Sementchenko VI, Schweinfest CW, Papas TS, Watson DK. ETS2 function is required to maintain the transformed state of human prostate cancer cells. Oncogene. 1998; 17:2883–2888.
21. Neznanov N, Man AK, Yamamoto H, Hauser CA, Cardiff RD, Oshima RG. A single targeted Ets2 allele restricts development of mammary tumors in transgenic mice. Cancer research. 1999; 59:4242–4246.
22. Ge Y, LaFiura KM, Dombkowski AA, Chen Q, Payton SG, Buck SA, Salagrama S, Diakiw AE, Matherly LH, Taub JW. The role of the proto-oncogene ETS2 in acute megakaryocytic leukemia biology and therapy. Leukemia. 2008; 22:521–529.
23. Ito Y, Miyoshi E, Takeda T, Sakon M, Ihara S, Tsujimoto M, Matsuura N. Ets-2 overexpression contributes to progression of pancreatic adenocarcinoma. Oncology reports. 2002; 9:853–857.
24. Sussan TE, Yang A, Li F, Ostrowski MC, Reeves RH. Trisomy represses Apc(Min)-mediated tumours in mouse models of Down’s syndrome. Nature. 2008; 451:73–75.
25. Foos G, Garcia-Ramirez JJ, Galang CK, Hauser CA. Elevated expression of Ets2 or distinct portions of Ets2 can reverse Ras-mediated cellular transformation. The Journal of biological chemistry. 1998; 273:18871–18880.
26. Munera J, Cecena G, Jedlicka P, Wankell M, Oshima RG. Ets2 regulates colonic stem cells and sensitivity to tumorigenesis. Stem cells. 2011; 29:430–439.
27. Kabbout M, Garcia MM, Fujimoto J, Liu DD, Woods D, Chow CW, Mendoza G, Momin AA, James BP, Solis L, Behrens C, Lee JJ, Wistuba II, et al. ETS2 mediated tumor suppressive function and MET oncogene inhibition in human non-small cell lung cancer. Clinical cancer research. 2013; 19:3383–3395.
28. Kar A, Gutierrez-Hartmann A. Molecular mechanisms of ETS transcription factor-mediated tumorigenesis. Critical reviews in biochemistry and molecular biology. 2013; 48:522–543.
29. Li R, Pei H, Watson DK. Regulation of Ets function by protein - protein interactions. Oncogene. 2000; 19:6514–6523.
30. Wei Q, Frazier ML, Levin B. DNA repair: a double-edged sword. Journal of the National Cancer Institute. 2000; 92:440–441.
31. Yu D, Zhang X, Liu J, Yuan P, Tan W, Guo Y, Sun T, Zhao D, Yang M, Liu J, Xu B, Lin D. Characterization of functional excision repair cross-complementation group 1 variants and their association with lung cancer risk and prognosis. Clinical cancer research. 2008; 14:2878–2886.
32. Roberts PJ, Bisi JE, Strum JC, Combest AJ, Darr DB, Usary JE, amboni WC, Wong KK, Perou CM, Sharpless NE. Multiple roles of cyclin-dependent kinase 4/6 inhibitors in cancer therapy. Journal of the National Cancer Institute. 2012; 104:476–487.
33. McClendon AK, Dean JL, Rivadeneira DB, Yu JE, Reed CA, Gao E, Farber JL, Force T, Koch WJ, Knudsen ES. CDK4/6 inhibition antagonizes the cytotoxic response to anthracycline therapy. Cell cycle. 2012; 11:2747–2755.
34. Lee SY, Kang HG, Yoo SS, Kang YR, Choi YY, Lee WK, Choi JE, Jeon HS, Shin KM, Oh IJ, Kim KS, Lee J, Cha SI, et al. Polymorphisms in DNA repair and apoptosis-related genes and clinical outcomes of patients with non-small cell lung cancer treated with first-line paclitaxel-cisplatin chemotherapy. Lung cancer. 2013; 82:330–339.
35. Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, Dancey J, Arbuck S, Gwyther S, Mooney M, Rubinstein L, Shankar L, Dodd L, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). European journal of cancer. 2009; 45:228–247.
36. Bhattacharya A, Ziebarth JD, Cui Y. PolymiRTS Database 3.0: linking polymorphisms in microRNAs and their target sites with human diseases and biological pathways. Nucleic acids research. 2014; 42:D86–91.
37. Higgins ME, Claremont M, Major JE, Sander C, Lash AE. CancerGenes: a gene selection resource for cancer genome projects. Nucleic acids research. 2007; 35:D721–726.
38. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT Method. Methods. 2001; 25:402–408.
39. Tay Y, Zhang J, Thomson AM, Lim B, Rigoutsos I. MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature. 2008; 455:1124–1128.
40. Molina-Pinelo S, Gutierrez G, Pastor MD, Hergueta M, Moreno-Bueno G, Garcia-Carbonero R, Nogal A, Suárez R, Salinas A, Pozo-Rodríguez F, Lopez-Rios F, Agulló-Ortuño MT, Ferrer I, et al. MicroRNA-dependent regulation of transcription in non-small cell lung cancer. PloS one. 2014; 9:e90524.

