INTRODUCTION
The term “pseudogene” comes from the Greek word “pseudo” meaning false. Pseudogenes are also identified as “genomic fossils”[1]. They are outdated DNA sequences that lack protein coding ability because of the presence of frame shift mutations and early or delayed stop codons, even though they look like functional genes [2]. They are considered nonfunctional relatives of ancestral functional genes that might have lost their function during evolution [3]. Pseudogenes have been found in plants [4], bacteria [5], yeast [6], insects [7], nematodes [8] and mammals [9]. Based on their origins, pseudogenes have been classified into: (i) Processed pseudogenes - produced by mRNA retrotransposition [10]; (ii) Duplicated pseudogenes (called unprocessed pseudogenes) - originated from the duplication of functional genes that than become nonfunctional because of acquired mutations [10]; (iii) Unitary or Disabled pseudogenes - originated by mutations in the functional protein coding sequences [2].
Nowadays, pseudogene origin, evolution and function are only partially understood. The first paper about the biological role of a pseudogene was published about 16 years ago [11]. In fact, Korneev et al. reported that the neuronal nitric oxide synthase pseudogene worked as natural antisense in regulating neuronal nitric oxide synthase gene expression. However, recent studies have shown more functional roles for pseudogenes, associating them to long non-coding RNAs (lncRNAs) family [10, 12-14]. In fact, Poliseno et al. reported that PTEN pseudogene 1 (PTENpg1) is transcribed in human tissues and cancers and harbor microRNA (miRNA) response elements (MREs) for some of the same miRNAs that target its corresponding protein-coding gene, PTEN [12]. By sequestering miRNAs that would modulate PTEN, the corresponding pseudogene derepresses the protein-coding genes from miRNA regulation [12]. In this way transcripts could talk each other by competing for the same miRNAs, acting as competitive endogenous RNAs (ceRNAs) [15, 16]. In addition, Johnsson et al. characterized two PTENpg1 antisense RNA isoforms, alpha and beta. The alpha isoform acts on the PTEN promoter inhibiting PTEN transcription by epigenetic mechanisms. On the contrary, the beta isoform directly interacts with PTENpg1 RNA, which influences PTEN protein by changing PTENpg1 stability and miRNA decoy activity [17]. Therefore, the overexpression of PTENpg1 sustains PTEN expression acquiring oncosuppressive functions [12].
Moreover, the human BRAF pseudogene (BRAFP1) has been recently found overexpressed in various tumor types, suggesting that it may contribute to cancer development. Karret et al. demonstrated the ceRNA role of both mouse Braf-rs1 pseudogene (Braf-rs1)and its human ortholog, BRAFP1, eliciting the expression of BRAF and the activation of MAPK cascade both in vitro and in vivo. Indeed, miRNA bioinformatic analysis showed that murine Braf-rs1 and B-Raf share 53 miRNAs, equally human BRAFP1 and BRAF share 40 miRNAs. Thus, the BRAF pseudogene, via sequestration of common miRNAs, may work as a decoy for BRAF in mice and humans then upregulating BRAF and promoting MAPK signaling and tumorigenesis. Furthermore, mice overexpressing Braf-rs1 develop an aggressive tumor similar to the human diffuse large B cell lymphoma. In addition, several transcriptional or genomic aberrations of BRAFP1 were frequently found in multiple human cancers, including B cell lymphomas [18]. Taken together, pseudogenes are deeply involved in ceRNA hypothesis and give rise to large-scale controlling system across the transcriptome, critically increasing the functional data of human genome and acquiring main roles in physiological and pathological conditions [19].
Besides pseudogene-derived small RNAs have been demonstrated to have a role in chromatin repression [20]. Latest evidences show contribution of pseudogenes in regulating development and disease by encoding peptides or proteins [21-23]. Interestingly, Kandouz et al. detected the expression of Cx43 pseudogene (psiCx43) in several cancer cell lines demonstrating its translationability in a protein of 43 kDa. Moreover, the psiCx43 protein overexpression was able to induce translational inhibition of Cx43 acting as a posttranscriptional regulator of Cx43, whose expression in cancer slows growth and renders the cells more sensitive to cytotoxic chemotherapeutics [23]. Finally, it has been reported that pseudogenes produce small interfering RNAs (siRNAs) in African Trypanosoma brucei and suppress several kinds of functional protein-coding genes through RNA interference pathway [24].
The family of High-Mobility Group A (HMGA) is composed of four proteins, HMGA1a, HMGA1b, HMGA1c, encoded by HMGA1 gene at the end of alternative splicing, and HMGA2, encoded by the HMGA2 gene [25]. HMGA1 is located on chromosome 6p21 in humans and in the t-complex locus on mouse chromosome 17, whereas HMGA2 is found on chromosome 12q13-15 in humans and at the pigmy locus on chromosome 10 in mice [26]. HMGA1 and HMGA2 genes are well conserved through the species, in fact, only few differences have been found between the human and the murine HMGA sequence [26]. They are non-histone chromosomal proteins, also identified as “architectural transcriptional factors” since they do not show a direct transcriptional activity, but modify the chromatin structure thanks to their DNA-binding domains, called “AT hooks”, by which they bind the DNA minor groove at AT-rich nucleotide sequences modulating the gene transcription [27-29]. HMGA proteins are expressed at low levels in adult tissues, but their expression is copious during embryogenesis [26], suggesting their important role in development. Indeed, the phenotypic study of Hmga1 knock out mice showed that this protein has a critical role in different aspects of development [26, 30]. In particular, cardiac hypertrophy and type 2 diabetes were reported in Hmga1-null and heterozygous mice meaning that a correct quantity of HMGA1 protein is necessary for cardiomyocytic cell growth and regulation of the insulin pathway [26, 31-33]. In fact, the downregulation of HMGA1 protein leads to a reduced insulin receptor (INSR) expression in patients with insulin resistance and type 2 diabetes. The recovery of HMGA1 levels improved INSR gene transcription, restoring both expression of insulin receptor protein on cell surface and insulin-binding capacity [31].
It is worth noting that HMGA1 has been found abundantly expressed in all human neoplastic tissues analyzed, including, prostate [34-36], colon [37-39], breast [40-42], gastric [43-45], lung [46-48], testis [49-51], pancreas [52-54], ovary [55-57], thyroid carcinomas [58-60] and also in some forms of leukemia [61-63]. Importantly, HMGA1 expression level has been correlated with an advanced stage, occurrence of distant metastases and reduced survival in colorectal carcinomas [64-67]. To further sustain the HMGA1 function in cancer development, its expression levels have been associated with histologic grade of breast and ovarian carcinomas, where HMGA1 expression gradually enhances from no expression in normal breast tissue, to modest expression in hyperplastic lesions to overexpression in ductal carcinomas [67, 68], and augments from faintly expressed in ovarian carcinomas with low invasive potential to extremely expressed in invasive carcinomas [55, 69]. Importantly, HMGA overexpression plays a causal role in cell transformation. Indeed, their upregulation is able to transform rat fibroblasts [70] and human epithelial breast cells [71] and the block of HMGA1 protein expression prevented thyroid cell transformation induced by Kirsten Murine Sarcoma Virus, and it induced cell death into human thyroid anaplastic cell lines [72, 73]. Furthermore, the silencing of HMGA1 expression in colon cancer stem cells restores normal stem cell characteristics, reducing sphere-forming efficiency and recovering the asymmetric division pattern [39]. Finally, HMGA1 transgenic mice develop several benign or malignant neoplasias, such as GH/PRL-secreting pituitary adenomas, T-cell acute lymphoblastic leukemia and T/NK lymphomas [26, 37, 74].
The molecular processes involved in cell transformation induced by the HMGA genes are based on their capacity to positively or negatively control the expression of genes and miRNAs, small noncoding RNAs engaged in gene regulation [75, 76] and implicated in the regulation of cellular proliferation, invasion and apoptosis [77-79].
The upregulation of the HMGA genes in cancer may occur through oncofetal transcriptional mechanisms, which have not been elucidated yet. It is known that the high expression of HMGA1 in cancer cells needs a close cooperation between SP1 family elements and AP1 proteins, stimulated by the activation of Ras GTPase cascade [80]. Furthermore, recent studies have demonstrated the miRNAs HMGA proteins regulation by binding its 3’ untranslated region (UTR), provoking mRNA degradation or inhibition of its translation [81, 82]. In particular, several studies reported a strong HMGA regulation by miRNAs in pituitary adenomas (mir-15, mir-16, miR-34b, mir-214, miR-326, miR-432, miR-548c-3p, miR-570, miR-603 and mir-761) [83-85], in thyroid carcinomas (let-7) [86], in breast cancer (mir-26a, miR-33b) [87, 88]. Moreover, the loss of HMGA2 3’UTR, commonly found in benign tumors of mesenchymal origin, abolishes the inhibition of HMGA2 expression by several miRNAs [89, 90], leading to HMGA2 protein overexpression that accounts for neoplastic transformation.
HMGA1 pseudogenes
The analysis of the human genome by bioinformatic database revealed the presence of eight processed HMGA1 pseudogenes (HMGA1Ps): HMGA1-p, HMGA1P1, HMGA1P2, HMGA1P3, HMGA1P4, HMGA1P5, HMGA1P6, and HMGA1P7 (Table 1).
Table 1: HMGA1 Pseudogenes family
Gene |
Location |
Function |
Main mutations |
Reference |
HMGA1P1 |
Xp21.3 |
Competitor protein for HMGA1 with different post-translational modifications.* |
arg25, thr53, arg57, ser64 |
91-95,99 |
HMGA1P2 |
4q13.3 |
Competitor protein for HMGA1 with different post-translational modifications.* |
arg57, arg59 |
94,99 |
HMGA1P3 |
12q24.11 |
Truncated form of HMGA1 with all molecular activities mentioned above.* |
arg59, c-terminal tail deletion |
94,96,97,99 |
HMGA1P4 |
9q34.11 |
- |
- |
- |
HMGA1P5 |
10q22.2 |
HMGA1 non-homologous peptide* |
- |
- |
HMGA1P6 |
13q12.12 |
Sustains the overexpression of several cancer-related genes by ceRNA mechanism |
Stop codon |
100-102 |
HMGA1P7 |
6q23.2 |
Sustains the overexpression of several cancer-related genes by ceRNA mechanism. |
Start codon |
100-102 |
HMGA1-p |
2p13.2 |
Competes with HMGA1 3’ UTR for the binding to αCP1 RNA stability factor. |
Few mutations |
106 |
*The function of these pseudogenes still needs to be validated.
HMGA1P1 and HMGA1P2
HMGA1P1 and HMGA1P2 pseudogenes, classified as processed pseudogenes, are located on Xp21.3 and 4q13.3 chromosome, respectively. They are not conserved during the evolution, but are only found in human genome. There are few mutations that distinguish HMGA1P1 and HMGA1P2 from HMGA1a. These changes in DNA cause few errors in protein sequence that importantly do not affect their translationability. Indeed, our preliminary data show that expressing vectors for HMGA1P1 and HMGA1P2 are able to code for proteins detectable by western blotting analysis. In fact, lysates from HMGA1-null cells transfected with the both vectors were positive to HMGA1 antibodies, which recognize the N-terminal aminoacids shared by HMGA1P1, HMGA1P2 and HMGA1. As shown in Figure 1, some HMGA1P1 and HMGA1P2 mutations hit aminoacidic residues that are frequently modified at post-translational level along the HMGA1 protein. Indeed, HMGA1P1 is mutated at position 25 where a tryptophan residue substitutes an arginine, within the first AT-hook of HMGA1, which has been shown to be a major site of modification in tumor cells [91]. In fact, Sgarra et al. demonstrated that the arginine residue 25 is strictly related to the execution of programmed cell death in tumor cell lines [92]. HMGA1P1 is also mutated at threonine residue 53 that is substituted with a lysine residue. Interestingly, threonine 53 was previously known as the main site of phosphorylation by cdc2 kinase during the cell cycle [93]. Compared with unphosphorylated protein, stoichiometric phosphorylation of recombinant human HMGA1 by cdc2 kinase strongly decreases the binding to DNA. Moreover, the HMGA1 protein arginine residue at position 57, along the second AT-hook, is replaced by a glutamine in HMGA1P1. It has been reported that PRMT6 methylates HMGA1 at the level of arginine 57, which is involved in the affinity for DNA binding and also in protein-protein interaction, thus implying an important role for arginine methylation in modulating HMGA functions [94]. Finally, HMGA1P1 brings a mutation at serine 64 where it shows an arginine residue. This is a Protein Kinase C (PKC) phosphorylation site [95]. Phosphorylation of HMGA1 by PKC resulted in a reduction of DNA-binding affinity as compared with that caused by the phosphorylation with cdc2 kinase, which phosphorylates threonine 53 [91, 95]. Therefore, HMGA1 could be additively phosphorylated by cdc2 kinase and PKC, and the resulting doubly phosphorylated protein exhibits a strong reduction in binding affinity [91, 95].
HMGA1P2 is mutated at arginine residues 57 and 59 where it shows a tryptophan and a glutamine residue, respectively (Figure 1). As well as arginine 57 methylation, it has been reported that HMGA1 arginine 59 is methylated by PRMT6 modulating its ability to bind to DNA and also the protein-protein affinity [94]. Therefore, if HMGA1P1 and HMGA1P2 pseudogenes coded for proteins, they could represent a sort of competitor proteins for HMGA1 wild-type with different post-translational modifications, altering HMGA1 properties in chromatin remodeling and protein-protein interactions.
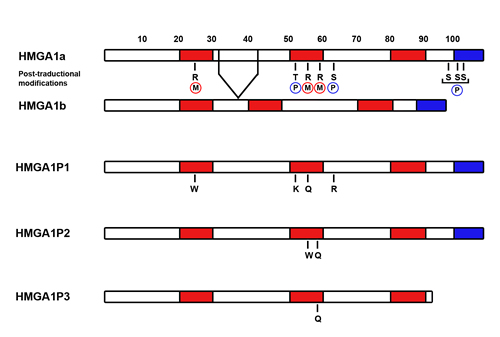
Figure 1: Structure of HMGA1Ps and their main mutations with respect to HMGA1 proteins. Diagrams illustrating the domain structures of HMGA1a, HMGA1b, HMGA1P1, HMGA1P2 and HMGA1P3. Known post-translational modifications of human HMGA1a and HMGA1b proteins impaired in HMGA1Ps are highlighted (Phosphorylation in blue, methylation in red). The three AT-hooks are in red and the acidic tail in blue.
HMGA1P3
HMGA1P3 pseudogene, only found in human genome, is classified as processed pseudogene and it is located on chromosome 12q24.11. Even though classified as non-coding RNA, it has only four aminoacidic mutations along the protein sequence compared to HMGA1 sequence, without affecting its translationability (Figure 1). Interestingly, arginine 59 is replaced with a glutamine residue as mentioned above for HMGA1P2 [95]. Moreover, HMGA1P3 pseudogene lacks the C-terminal acidic tail that is a feature of HMGA proteins [26]. In fact, it has been revealed that HMGA1a and HMGA1b are phosphorylated by Casein kinase II (CK2) on three serines situated in the C-terminal tail (S98, S101 and S102) [96, 97]. Moreover, it is believed that the HMGA C-terminal tail may be important in modulating protein-protein interactions [97] and could be involved in enhancing transcription factor activity, but the role of these phosphorylations has been not completely uncovered yet [97]. Finally, expression of a truncated Hmga1b gene,
without both the acidic tail and the 3’ UTR, significantly enhances growth rate and impairs adipocytic differentiation, also suggesting that the Hmga1/T mutant works in a contrasting manner [98]. Indeed, transgenic mice overexpressing the HMGA1 wild-type protein showed a reduction of the fat tissue in contrast with the obese phenotype of the Hmga1/T, mice even though there are no data that could explain why the wild-type and the truncated form of Hmga1 operate in opposite ways [98].
Given this scenario, if HMGA1P3 pseudogene coded for protein, it could represent a truncated form of HMGA1 wild-type with all molecular activities mentioned above.
Interestingly, previous studies showed that HMGA1P1, HMGA1P2 and HMGA1P3 can be affected by chromosomal rearrangements in benign human tumors [99]. In particular, significantly higher frequency of chromosomal breaks within the chromosomal bands containing these pseudogenes were observed in uterine leiomyomas, lipomas, pleomorphic adenomas, and pulmonary chondroid hamartomas [99]. This study unveils the existence of an interesting pseudogene activation mechanism in tumor, since they could translocate, after chromosomal rearrangements, under a promoter region or within a functional gene, then coding for new fusion proteins.
However, no studies have evaluated HMGA1P1, HMGA1P2 and HMGA1P3 expression in human normal and malignant tissues where their possible deregulated expression might have consequences on the function of the wild type HMGA1 protein and then influence cancer progression.
HMGA1P4 and HMGA1P5
The non-coding RNA HMGA1P4 pseudogene is classified as processed pseudogene and is located on the human chromosome 9q34.11. Differently from the above mentioned pseudogenes, HMGA1P4 genomic sequence shows low homology with HMGA1. Moreover, further bioinformatics analysis confirms its untranslationability. Therefore, it could not be classified either as ceRNA or as peptide related to HMGA1.
Another processed pseudogene related to HMGA1 is HMGA1P5. It is present only in humans and located on the chromosome 10q22.2. As HMGA1P4, HMGA1P5 has low homology along HMGA1 sequence and it may code for a peptide not related to HMGA1 protein. At the moment there are no published studies about these pseudogenes.
HMGA1P6 and HMGA1P7
The processed pseudogenes HMGA1P6 and HMGA1P7 are sited on 13q12.12 and 6q23.2 chromosome, respectively. They are not conserved through the evolution, but are present only in human genome [100-102]. These pseudogenes have high sequence homology with HMGA1 both in the 5’ and 3’ UTRs and in the coding region (Figure 2). A missense mutation of the start methionine codon avoids HMGA1P7 mRNA translation whereas HMGA1P6 bears a mutation in the stop codon, which is postponed several aminoacidic residues downstream, producing a non-translatable mRNA [100-102]. In the homology sequences, among HMGA1, HMGA1P6 and HMGA1P7, we retrieved conserved seed matches for miRNAs that have been predicted (miR-103, miR-142-3p, miR-370, and miR-432) or previously validated (miR-15 [83], miR-16 [83], miR-26a [103], miR-214 [104], miR-548c-3p [84] and miR-761 [104]) able to target the HMGA1 gene (Figure 2).
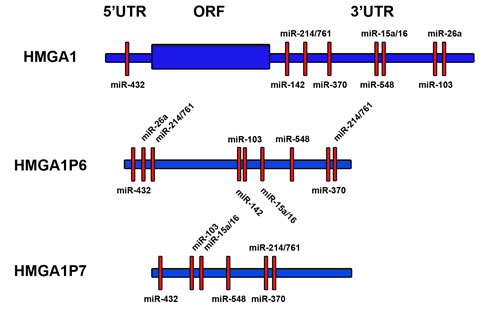
Figure 2: HMGA1P6 and HMGA1P7 mRNA sequence shares HMGA1-targeting miRNAs. HMGA1 (top), HMGA1P6 (middle) and HMGA1P7 (bottom) mRNA sequences are shown in blue. HMGA1-targeting miRNA seed matches (red boxes) within the high homology regions are shared among HMGA1, HMGA1P6 and HMGA1P7.
It has been reported that both HMGA1P6 and HMGA1P7 act as decoys for HMGA1-targeting miRNAs. In fact, their overexpression enhances HMGA1 protein levels whereas their knocking down results in the reduction of HMGA1 mRNA and protein amounts (Figure 3) [100-102].
Consistently, these HMGA1Ps have also oncogenic action by preventing apoptosis and enhancing cell proliferation and migration [100, 101]. Indeed, overexpression of HMGA1P6 or HMGA1P7 increases the growth rate and migration of different cell lines, contributing to tumor development [100-102]. Moreover, the generation of HMGA1P6 or HMGA1P7 transgenic mice confirms their oncogenic activity. In fact, mouse embryonic fibroblasts (MEFs) obtained from HMGA1P6 or HMGA1P7 [100] transgenic mice grow more rapidly and senesce later than their wild-type counterparts. Remarkably, in HMGA1P6 and HMGA1P7 overexpressing cells and MEFs we detected the upregulation of several cancer-related genes such as High Mobility Group A2 (HMGA2), Enhancer of Zeste Homolog 2 (EZH2), Vascular Endothelial Growth Factor (VEGF), and Ephrin Type-A Receptor 3 (Epha3), with respect to the control cells [100]. This happens because of shared miRNAs targeting HMGA1P6, HMGA1P7, HMGA1 and other cancer related genes. Therefore, high HMGA1 gene or its pseudogene expression allows to increase other oncogene protein levels then contributing to cancer progression. Finally, a direct correlation among HMGA1, HMGA1P6 and HMGA1P7 expression in a group of human thyroid and ovary tumors has been shown [97-99]. Indeed, papillary thyroid carcinomas (PTC), which are fine differentiated and weakly aggressive, express low levels of HMGA1P6, HMGA1P7 and HMGA1. On the contrary, anaplastic thyroid carcinomas (ATC), which are one of the most malignant human cancers, express very high HMGA1Ps levels that, moreover, correlated with HMGA1 protein levels [100]. Similar results were obtained in human ovarian carcinomas [100] and in endometrial carcinomas, where the HMGA1P6 and HMGA1P7 expression correlates with the malignancy rate. Interestingly, HMGA1P6 and HMGA1P7 were also overexpressed in human pituitary adenomas where the HMGA proteins play a critical role in their development [105]. In particular, HMGA1P6 and HMGA1P7 expression significantly correlates with HMGA1 mRNA in somatotropic and nonfunctioning pituitary adenomas. Moreover, functional studies show that the enforced expression HMGA1P6 and HMGA1P7 enhances the proliferation of a pituitary adenoma cell line. Therefore, HMGA1P6 and HMGA1P7 overexpression contributes to keep high HMGA1 protein levels enhancing, then, its oncogenic ability.
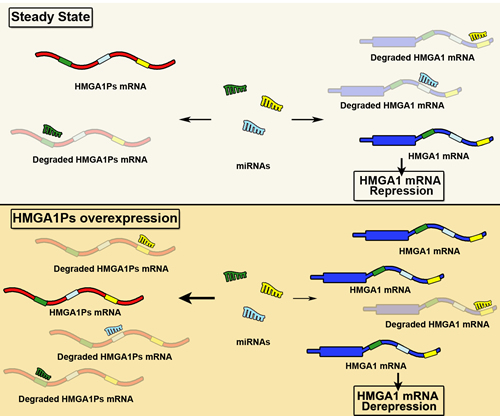
Figure 3: miRNA decoy function of HMGA1Ps. In the steady state, equilibrium exists between the miRNAs and their targets HMGA1 and HMGA1Ps. By contrast, the overexpression of HMGA1P6 and HMGA1P7 results in fewer miRNAs free to bind to HMGA1, and thus HMGA1 levels increase.
HMGA1-p
HMGA1-p is located on chromosome 2p13.2. Its expression is able to induce destabilization of HMGA1 mRNA [83]. Indeed, it has been demonstrated that the HMGA1-p RNA competes with HMGA1 3’ UTR for a critical RNA stability factor, the alpha C-binding protein (αCP1) [106]. The HMGA1-p was found overexpressed in diabetic patients then causing a significant destabilization of HMGA1 mRNA with consequent loss of INSR expression, which is regulated by HMGA1, then generating the insulin resistance phenotype (Figure 4). Moreover, targeted knockdown of HMGA1-p mRNA results in an increase of HMGA1 mRNA stability and expression levels, with a parallel correction in cell-surface INSR expression and insulin binding capacity [106]. Therefore, this study established a novel mechanistic linkage between HMGA1-p pseudogene expression and type 2 diabetes mellitus.
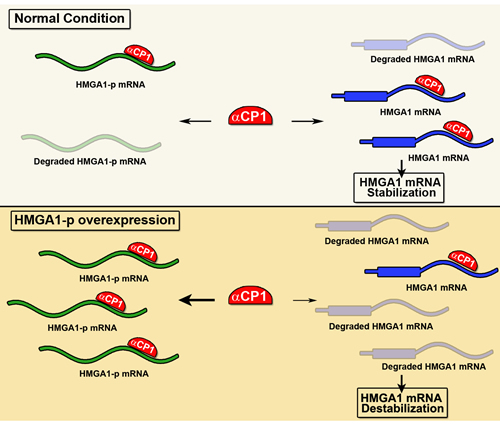
Figure 4: HMGA1-p function model. In normal condition, the RNA-binding protein αCP1 stabilizes HMGA1 mRNA by binding to its 3’ UTR. In diabetes, the HMGA1-p overexpressed transcript competes with HMGA1 mRNA for the binding to αCP1, increasing the degradation of HMGA1 mRNA.
Conclusions and Perspectives
The mammalian genome contains an high number of pseudogenes (about 20,000 in humans) [107, 108] more than those present in other organisms. The biological meaning of pseudogenes was completely obscure until few years ago, whereas recent studies have shown their critical role in regulating gene transcription mainly functioning as decoy for miRNAs, and also evidenced a role of pseudogenes in carcinogenesis [109-116]. Interestingly, we have identified, by bioinformatic search, eight pseudogenes for the HMGA1 gene whose expression is a feature of human malignancies with a key function in promoting cancer progression. From the analysis of the HMGA1P sequences it comes up that they could be able to regulate HMGA1 expression and function. Indeed, HMGA1P6 and HMGA1P7 act on the stability of HMGA1 mRNA or by protecting them from miRNAs able to target this gene, whereas HMGA1-p competes with HMGA1 3’ UTR for a critical RNA stability factor, the αCP1. Conversely, HMGA1P1, HMGA1P2 and HMGA1P3 could represent a sort of competitor proteins for HMGA1 wild-type with different post-translational modifications, altering HMGA1 properties in chromatin remodeling and protein-protein interactions. So far, the role of HMGA1-p in type 2 diabetes and HMGA1P6 and HMGA1P7 in the progression of some human neoplasias appears well documented, but further analysis of their expression in embryonic and adult tissues, and in human carcinomas is required to be deeper investigated. Recently, it has been observed that transgenic mice overexpressing either HMGA1P6 and HMGA1P7, develop lymphomas, infiltrating different organs likely working as ceRNAs for their oncogenic related genes.
Therefore, HMGA1Ps represent an epigenetic event, as well as miRNAs, able to regulate HMGA1 activity, and then play a critical role in all the processes such as cancer progression, development, metabolism and many other function in which HMGA1 is involved. The involvement of HMGA1 in all these important cellular processes likely accounts for the need of its fine regulation by using different molecular approaches. Interestingly, recent studies unveil a correlation between HMGA1P6 and HMGA1P7 and some clinico-pathological features, opening the perspective of using the evaluation of HMGA1Ps expression as diagnostic and prognostic marker, and maybe also in tumor classification. Therefore, the studies summarized here rehabilitate the HMGA1Ps from “junk” to a multifunctional pseudogene family that needs to be extensively studied.
Acknowledgments
This work was supported by grants from Associazione Italiana per la Ricerca sul Cancro-AIRC (IG 11477), “Progetto di Interesse strategico Invecchiamento (PNR-CNR Aging Program) PNR-CNR 2012-2014” and CNR Epigenomics Flagship Project “EPIGEN”.
Conflicts of Interest
There is no conflict of interest.
REFERENCES
1. Lafontaine I, Dujon B. Origin and fate of pseudogenes in Hemiascomycetes: a comparative analysis. BMC Genomics. 2010;11:260.
2. Mighell AJ, Smith NR, Robinson PA, Markham AF. Vertebrate pseudogenes. FEBS Letters. 2000; 468:109-14.
3. Balakirev ES, Ayala FJ. Pseudogenes: are they “junk” or functional DNA? Annu Rev Genet. 2003;37:123-51.
4. Loguercio LL, Wilkins TA. Structural analysis of a hmg-coA-reductase pseudogene: Insights into evolutionary processes affecting the hmgr gene family in allotetraploid cotton (Gossypium hirsutum L.). Curr Genet. 1998;34:241-9.
5. Ochman H, Davalos L. The nature and dynamics of bacterial genomes. Science. 2006;311:1730-3.
6. Harrison P, Kumar A, Lan N, Echols N, Snyder M, Gerstein M. A small reservoir of disabled ORFs in the yeast genome and its implications for the dynamics of proteome evolution. J Mol Biol. 2002;316:409-19.
7. Ramos-Onsins S, Aguadé M. Molecular evolution of the Cecropin multigene family in Drosophila: Functional genes vs. pseudogenes. Genetics. 1998;150:157-71.
8. Harrison PM, Kumar A, Lang N, Snyder M, Gerstein M. A question of size: the eukaryotic proteome and the problems in defining it. Nucleic Acids Res. 2002;30:1083-90.
9. Zhang Z, Gerstein M. Large-scale analysis of pseudogenes in the human genome. Current Opinion in Genetics and Development. 2004;14: 328-35.
10. Li W, Yang W, Wang XJ. Pseudogenes: Pseudo or Real Functional Elements? J Genet Genomics. 2013;40:171-7.
11. Korneev SA, Park JH, O’Shea M. Neuronal expression of neural nitric oxide synthase (nNOS) protein is suppressed by an antisense RNA transcribed from an NOS pseudogene. J Neurosci. 1999;19:7711-20.
12. Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465:1033-8.
13. Pink RC, Wicks K, Caley DP, Punch EK, Jacobs L, Carter DRF. Pseudogenes: pseudo-functional or key regulators in health and disease? RNA. 2011;17:792-8.
14. Peng H, Ishida M, Li L, Saito A, Kamiya A, Iacob R, Dima S, Alexandrescu ST, Grigorie R. Pseudogene INTS6P1 regulates its cognate gene INTS6 through competitive binding of miR-17-5p in hepatocellular carcinoma. Oncotarget. 2015;6:56666-77. doi 10.18632/oncotarget.3290.
15. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146:353-8.
16. Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505:344-52.
17. Johnsson P, Ackley A, Vidarsdottir L, Lui W-O, Corcoran M, Grandér D, Morris K V. A pseudogene long-noncoding-RNA network regulates PTEN transcription and translation in human cells. Nat Struct Mol Biol. 2013;20:440-6.
18. Karreth FA, Reschke M, Ruocco A, Ng C, Chapuy B, Léopold V, Sjoberg M, Keane TM, Verma A, Ala U, Tay Y, Wu D, Seitzer N, et al. The BRAF Pseudogene Functions as a Competitive Endogenous RNA and Induces Lymphoma In Vivo. Cell. 2015;161:319-32.
19. Karreth FA, Ala U, Provero P, Pandolfi PP. Pseudogenes as competitive endogenous RNAs: target prediction and validation. Methods Mol Biol. 2014;1167:199-212.
20. Guo X, Lin M, Rockowitz S, Lachman HM, Zheng D. Characterization of human pseudogene-derived non-coding RNAs for functional potential. PLoS One. 2014;9:e93972.
21. Marques AC, Tan J, Lee S, Kong L, Heger A, Ponting CP. Evidence for conserved post-transcriptional roles of unitary pseudogenes and for frequent bifunctionality of mRNAs. Genome Biol. 2012;13:R102.
22. Bertrand N, Castro DS, Guillemot F. Proneural genes and the specification of neural cell types. Nat Rev Neurosci. 2002;3:517-30.
23. Kandouz M, Bier A, Carystinos GD, Alaoui-Jamali MA, Batist G. Connexin43 pseudogene is expressed in tumor cells and inhibits growth. Oncogene. 2004;23:4763-70.
24. Wen Y-Z, Zheng L-L, Liao J-Y, Wang M-H, Wei Y, Guo X-M, Qu L-H, Ayala FJ, Lun Z-R. Pseudogene-derived small interference RNAs regulate gene expression in African Trypanosoma brucei. Proc Natl Acad Sci U S A. 2011;108:8345-50.
25. Johnson KR, Lehn DA, Reeves R. Alternative processing of mRNAs encoding mammalian chromosomal high-mobility-group proteins HMG-I and HMG-Y. Mol Cell Biol. 1989;9:2114-23.
26. Fusco A, Fedele M. Roles of HMGA proteins in cancer. Nat Rev Cancer. 2007;7:899-910.
27. Grosschedl R, Giese K, Pagel J. HMG domain proteins: Architectural elements in the assembly of nucleoprotein structures. Trends in Genetics. 1994;10:94-100.
28. Thanos D, Maniatis T. The high mobility group protein HMG I(Y) is required for NF-kappa B-dependent virus induction of the human IFN-beta gene. Cell. 1992;71:777-89.
29. Sepe R, Formisano U, Federico A, Forzati F, Bastos AU, Angelo DD, Cacciola NA, Fusco A, Pallante P. CBX7 and HMGA1b proteins act in opposite way on the regulation of the SPP1 gene expression. Oncotarget. 2015;6:2680-92. doi: 10.18632/oncotarget.2777.
30. Federico A, Forzati F, Esposito F, Arra C, Palma G, Barbieri A, Palmieri D, Fedele M, Pierantoni GM, De Martino I, Fusco A. Hmga1/Hmga2 double knock-out mice display a “superpygmy” phenotype. Biol Open. 2014;3:372-8.
31. Foti D, Chiefari E, Fedele M, Iuliano R, Brunetti L, Paonessa F, Manfioletti G, Barbetti F, Brunetti A, Croce CM, Fusco A, Brunetti A. Lack of the architectural factor HMGA1 causes insulin resistance and diabetes in humans and mice. Nat Med. 2005;11:765-73.
32. Chiefari E, Tanyolaç S, Paonessa F, Pullinger CR, Capula C, Iiritano S, Mazza T, Forlin M, Fusco A, Durlach V, Durlach A, Malloy MJ, Kane JP, et al. Functional variants of the HMGA1 gene and type 2 diabetes mellitus. JAMA. 2011;305:903-12.
33. Brunetti A, Chiefari E, Foti D. Recent advances in the molecular genetics of type 2 diabetes mellitus. World J Diabetes. 2014;5:128-40.
34. Tamimi Y, Van Der Poel HG, Denyn MM, Umbas R, Karthaus HF, Debruyne FM, Schalken JA. Increased expression of high mobility group protein I(Y) in high grade prostatic cancer determined by in situ hybridization. Cancer Res. 1993;53:5512-6
35. Tamimi Y, Van Der Poel HG, Karthaus HF, Debruyne FM, Schalken JA. A retrospective study of high mobility group protein I(Y) as progression marker for prostate cancer determined by in situ hybridization. Br J Cancer. 1996;74: 573-8.
36. Wei JJ, Wu X, Peng Y, Shi G, Olca B, Yang X, Daniels G, Osman I, Ouyang J, Hernando E, Pellicer A, Rhim JS, Melamed J, et al. Regulation of HMGA1 expression by MicroRNA-296 affects prostate cancer growth and invasion. Clin Cancer Res. 2011;17:1297-305.
37. Belton A, Gabrovsky A, Bae YK, Reeves R, Iacobuzio-Donahue C, Huso DL, Resar LM. HMGA1 induces intestinal polyposis in transgenic mice and drives tumor progression and stem cell properties in colon cancer cells. PLoS One. 2012;7:e30034.
38. Bush BM, Brock AT, Deng JA, Nelson RA, Sumter TF. The Wnt/β-catenin/T-cell factor 4 pathway up-regulates high-mobility group A1 expression in colon cancer. Cell Biochem Funct. 2013;31:228-36.
39. Puca F, Colamaio M, Federico A, Gemei M, Bastos AU, Vecchio L Del, Pece S, Fusco A. HMGA1 silencing restores normal stem cell characteristics in colon cancer stem cells by increasing p53 levels Oncotarget. 2014;5:3234-45. doi: 10.18632/oncotarget.1914.
40. Pegoraro S, Ros G, Piazza S, Sommaggio R, Ciani Y, Rosato A, Sgarra R, Del Sal G, Manfioletti G. HMGA1 promotes metastatic processes in basal-like breast cancer regulating EMT and stemness. Oncotarget. 2013;4:1293-308. doi: 10.18632/oncotarget.1136.
41. Sepe R, Piscuoglio S, Quintavalle C, Perrina V, Quagliata L, Formisano U, Terracciano LM, Fusco A, Pallante P. HMGA1 overexpression is associated with a particular subset of human breast carcinomas. J Clin Pathol. 2015. pii: jclinpath-2015-202907.
42. Maurizio E, Wiśniewski JR, Ciani Y, Amato A, Arnoldo L, Penzo C, Pegoraro S, Giancotti V, Zambelli A, Piazza S, Manfioletti G, Sgarra R. Translating Proteomic Into Functional Data: An High Mobility Group A1 (HMGA1) Proteomic Signature Has Prognostic Value in Breast Cancer. Mol Cell Proteomics. 2016 ;15:109-23
43. Nam ES, Kim DH, Cho SJ, Chae SW, Kim HY, Kim SM, Han JJ, Shin HS, Park YE. Expression of HMGI(Y) associated with malignant phenotype of human gastric tissue. Histopathology. 2003 May;42:466-71.
44. Akaboshi S, Watanabe S, Hino Y, Sekita Y, Xi Y, Araki K, Yamamura K, Oshima M, Ito T, Baba H, Nakao M. HMGA1 is induced by Wnt/beta-catenin pathway and maintains cell proliferation in gastric cancer. Am J Pathol. 2009;175:1675-85.
45. Jun KH, Jung JH, Choi HJ, Shin EY, Chin HM. HMGA1/HMGA2 protein expression and prognostic implications in gastric cancer. Int J Surg. 2015;24:39-44.
46. Hillion J, Wood LJ, Mukherjee M, Bhattacharya R, Di Cello F, Kowalski J, Elbahloul O, Segal J, Poirier J, Rudin CM, Dhara S, Belton A, Joseph B, Zucker S, Resar LMS. Upregulation of MMP-2 by HMGA1 promotes transformation in undifferentiated, large-cell lung cancer. Mol Cancer Res. 2009;7:1803-12.
47. Zhang Z, Wang Q, Chen F, Liu J. Elevated expression of HMGA1 correlates with the malignant status and prognosis of non-small cell lung cancer. Tumour Biol. 2015;36:1213-9.
48. Pallante P, Sepe R, Puca F, Fusco A. High mobility group a proteins as tumor markers. Front Med (Lausanne). 2015;2:15.
49. Franco R, Esposito F, Fedele M, Liguori G, Pierantoni GM, Botti G, Tramontano D, Fusco A, Chieffi P. Detection of high-mobility group proteins A1 and A2 represents a valid diagnostic marker in post-pubertal testicular germ cell tumours. J Pathol. 2008;214:58-64.
50. Esposito F, Boscia F, Gigantino V, Tornincasa M, Fusco A, Franco R, Chieffi P. The high-mobility group A1-estrogen receptor β nuclear interaction is impaired in human testicular seminomas. J Cell Physiol. 2012;227:3749-55.
51. Chieffi P. Recent advances in molecular and cell biology of testicular germ-cell tumors. Int Rev Cell Mol Biol. 2014;312:79-100.
52. Trapasso F, Sarti M, Cesari R, Yendamuri S, Dumon KR, Aqeilan RI, Pentimalli F, Infante L, Alder H, Abe N, Watanabe T, Viglietto G, Croce CM, Fusco A. Therapy of human pancreatic carcinoma based on suppression of HMGA1 protein synthesis in preclinical models. Cancer Gene Ther. 2004;11:633-41.
53. Piscuoglio S, Zlobec I, Pallante P, Sepe R, Esposito F, Zimmermann A, Diamantis I, Terracciano L, Fusco A, Karamitopoulou E. HMGA1 and HMGA2 protein expression correlates with advanced tumour grade and lymph node metastasis in pancreatic adenocarcinoma. Histopathology. 2012;60:397-404.
54. Hillion J, Smail SS, Di Cello F, Belton A, Shah SN, Huso T, Schuldenfrei A, Nelson DM, Cope L, Campbell N, Karikari C, Aderinto A, Maitra A, Huso DL, Resar LM. The HMGA1-COX-2 axis: a key molecular pathway and potential target in pancreatic adenocarcinoma. Pancreatology. 2012;12:372-9.
55. Masciullo V, Baldassarre G, Pentimalli F, Berlingieri MT, Boccia A, Chiappetta G, Palazzo J, Manfioletti G, Giancotti V, Viglietto G, Scambia G, Fusco A. HMGA1 protein over-expression is a frequent feature of epithelial ovarian carcinomas. Carcinogenesis. 2003;24:1191-8.
56. Treff NR, Dement GA, Adair JE, Britt RL, Nie R, Shima JE, Taylor WE, Reeves R. Human KIT ligand promoter is positively regulated by HMGA1 in breast and ovarian cancer cells. Oncogene. 2004;23:8557-62.
57. Liu Y, Wang Y, Zhang Y, Fu J, Zhang G. Knockdown of HMGA1 expression by short/small hairpin RNA inhibits growth of ovarian carcinoma cells. Biotechnol Appl Biochem. 2012;59:1-5.
58. Czyz W, Balcerczak E, Jakubiak M, Pasieka Z, Kuzdak K, Mirowski M. HMGI(Y) gene expression as a potential marker of thyroid follicular carcinoma. Langenbecks Arch Surg. 2004;389:193-7.
59. Frasca F, Rustighi A, Malaguarnera R, Altamura S, Vigneri P, Del Sal G, Giancotti V, Pezzino V, Vigneri R, Manfioletti G. HMGA1 inhibits the function of p53 family members in thyroid cancer cells. Cancer Res. 2006;66:2980-9.
60. Martinez Hoyos J, Ferraro A, Sacchetti S, Keller S, De Martino I, Borbone E, Pallante P, Fedele M, Montanaro D, Esposito F, Cserjesi P, Chiariotti L, Troncone G, Fusco A. HAND1 gene expression is negatively regulated by the High Mobility Group A1 proteins and is drastically reduced in human thyroid carcinomas. Oncogene. 2009;28:876-85.
61. Pierantoni GM, Agosti V, Fedele M, Bond H, Caliendo I, Chiappetta G, Lo Coco F, Pane F, Turco MC, Morrone G, Venuta S, Fusco A. High-mobility group A1 proteins are overexpressed in human leukaemias. Biochem J. 2003;372:145-50.
62. Roy S, Di Cello F, Kowalski J, Hristov AC, Tsai HL, Bhojwani D, Meyer JA, Carroll WL, Belton A, Resar LM. HMGA1 overexpression correlates with relapse in childhood B-lineage acute lymphoblastic leukemia. Leuk Lymphoma. 2013;54:2565-7.
63. Mahajan L, Pandit H, Madan T, Gautam P, Yadav AK, Warke H, Sundaram CS, Sirdeshmukh R, Sarma PU, Kishore U, Surolia A. Human surfactant protein D alters oxidative stress and HMGA1 expression to induce p53 apoptotic pathway in eosinophil leukemic cell line. PLoS One. 2013;8:e85046.
64. Abe N, Watanabe T, Sugiyama M, Uchimura H, Chiappetta G, Fusco A, Atomi Y. Determination of high mobility group I(Y) expression level in colorectal neoplasias: a potential diagnostic marker. Cancer Res. 1999 59:1169-74.
65. Chiappetta G, Manfioletti G, Pentimalli F, Abe N, Di Bonito M, Vento MT, Giuliano A, Fedele M, Viglietto G, Santoro M, Watanabe T, Giancotti V, Fusco A. High mobility group HMGI(Y) protein expression in human colorectal hyperplastic and neoplastic diseases. Int J Cancer. 2001;91:147-51.
66. Huang ML, Chen CC, Chang LC. Gene expressions of HMGI-C and HMGI(Y) are associated with stage and metastasis in colorectal cancer. Int J Colorectal Dis. 2009;24(11):1281-6.
67. Rogalla P, Drechsler K, Kazmierczak B, Rippe V, Bonk U, Bullerdiek J. Expression of HMGI-C, a member of the high mobility group protein family, in a subset of breast cancers: relationship to histologic grade. Mol Carcinog. 1997;19:153-6.
68. Chiappetta G, Botti G, Monaco M, Pasquinelli R, Pentimalli F, Di Bonito M, D’Aiuto G, Fedele M, Iuliano R, Palmieri EA, Pierantoni GM, Giancotti V, Fusco A. HMGA1 protein overexpression in human breast carcinomas: correlation with ErbB2 expression. Clin Cancer Res. 2004;10:7637-44.
69. Peters DG, Kudla DM, Deloia JA, Chu TJ, Fairfull L, Edwards RP, Ferrell RE. Comparative gene expression analysis of ovarian carcinoma and normal ovarian epithelium by serial analysis of gene expression. Cancer Epidemiol Biomarkers Prev. 2005;14:1717-23.
70. Wood LJ, Maher JF, Bunton TE, Resar LMS. The oncogenic properties of the HMG-I gene family. Cancer Res. 2000;60:4256-61.
71. Reeves R, Edberg DD, Li Y. Architectural transcription factor HMGI(Y) promotes tumor progression and mesenchymal transition of human epithelial cells. Mol Cell Biol. 2001;21:575-94.
72. Scala S, Portella G, Fedele M, Chiappetta G, Fusco a. Adenovirus-mediated suppression of HMGI(Y) protein synthesis as potential therapy of human malignant neoplasias. Proc Natl Acad Sci U S A. 2000;97:4256-61.
73. Berlingieri MT, Pierantoni GM, Giancotti V, Santoro M, Fusco A. Thyroid cell transformation requires the expression of the HMGA1 proteins. Oncogene. 2002;21:2971-80.
74. Xu Y, Sumter TF, Bhattacharya R, Tesfaye A, Fuchs EJ, Wood LJ, Huso DL, Resar LMS. The HMG-I oncogene causes highly penetrant, aggressive lymphoid malignancy in transgenic mice and is overexpressed in human leukemia. Cancer Res. 2004;64:3371-5.
75. Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T. Identification of tissue-specific MicroRNAs from mouse. Curr Biol. 2002;12:735-9.
76. Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015;15:321-33.
77. Carleton M, Cleary MA, Linsley PS. MicroRNAs and cell cycle regulation. Cell Cycle. 2007;6:2127-32.
78. Ma L, Weinberg R a. MicroRNAs in malignant progression. Cell Cycle. 2008;7:570-2.
79. Su Z, Yang Z, Xu Y, Chen Y, Yu Q. MicroRNAs in apoptosis, autophagy and necroptosis. Oncotarget. 2015;6: 8474-90. doi: 10.18632/oncotarget.3523.
80. Cleynen I, Huysmans C, Sasazuki T, Shirasawa S, Van De Ven W, Peeters K. Transcriptional control of the human high mobility group A1 gene: Basal and oncogenic Ras-regulated expression. Cancer Res. 2007;67:4620-9.
81. Bartel DP. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell. 2004; 116:281-97.
82. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857-66.
83. Palmieri D, D’Angelo D, Valentino T, De Martino I, Ferraro A, Wierinckx A, Fedele M, Trouillas J, Fusco A. Downregulation of HMGA-targeting microRNAs has a critical role in human pituitary tumorigenesis. Oncogene. 2012; 31:3857-65.
84. D’Angelo D, Palmieri D, Mussnich P, Roche M, Wierinckx A, Raverot G, Fedele M, Croce CM, Trouillas J, Fusco A. Altered microRNA expression profile in human pituitary GH adenomas: Down-regulation of miRNA targeting HMGA1, HMGA2, and E2F1. J Clin Endocrinol Metab. 2012;97:E1128-38.
85. D’Angelo D, Esposito F, Fusco A. Epigenetic Mechanisms Leading to Overexpression of HMGA Proteins in Human Pituitary Adenomas. Front Med. 2015;2:39.
86. Pallante P, Battista S, Pierantoni GM, Fusco A. Deregulation of microRNA expression in thyroid neoplasias. Nat Rev Endocrinol. 2014;10:88-101.
87. Zhao X, Yuan Q, Mu D, Sun D, Bo Q, Pan G, Li G, Cui T. MicroRNA-26a inhibits proliferation by targeting high mobility group AT-hook 1 in breast cancer. 2015;8:368-73.
88. Lin Y, Liu AY, Fan C, Zheng H, Li Y, Zhang C, Wu S, Yu D, Huang Z, Liu F, Luo Q, Yang CJ, Ouyang G. MicroRNA-33b Inhibits Breast Cancer Metastasis by Targeting HMGA2, SALL4 and Twist1. Sci Rep. 2015;5:9995.
89. Hebert C, Norris K, Scheper MA, Nikitakis N, Sauk JJ. High mobility group A2 is a target for miRNA-98 in head and neck squamous cell carcinoma. Mol Cancer. 2007;6:5.
90. Yong SL, Dutta A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 2007;21:1025-30.
91. Zhang Q, Wang Y. High mobility group proteins and their post-translational modifications. Biochim Biophys Acta. 2008;1784:1159-66.
92. Sgarra R, Diana F, Bellarosa C, Dekleva V, Rustighi A, Toller M, Manfioletti G, Giancotti V. During apoptosis of tumor cells HMGA1a protein undergoes methylation: Identification of the modification site by mass spectrometry. Biochemistry. 2003;42:3575-85.
93. Nissen MS, Langan TA, Reevest R. Phosphorylation by cdc2 kinase modulates DNA binding activity of high mobility group I nonhistone chromatin protein. J Biol Chem. 1991;266:19945-52.
94. Sgarra R, Lee J, Tessari MA, Altamura S, Spolaore B, Giancotti V, Bedford MT, Manfioletti G. The AT-hook of the chromatin architectural transcription factor high mobility group A1a is arginine-methylated by protein arginine methyltransferase 6. J Biol Chem. 2006;281:3764-72.
95. Reeves R. Molecular biology of HMGA proteins: Hubs of nuclear function. Gene. 2001;277:63-81.
96. Currie RA. Functional interaction between the DNA binding subunit trimerization domain of NF-Y and the high mobility group protein HMG-I(Y). J Biol Chem. 1997;272:30880-8.
97. Sgarra R, Maurizio E, Zammitti S, Lo Sardo A, Giancotti V, Manfioletti G. Macroscopic differences in HMGA oncoproteins post-translational modifications: C-terminal phosphorylation of HMGA2 affects its DNA binding properties. J Proteome Res. 2009;8:2978-89.
98. Pierantoni GM, Battista S, Pentimalli F, Fedele M, Visone R, Federico A, Santoro M, Viglietto G, Fusco A. A truncated HMGA1 gene induces proliferation of the 3T3-L1 preadipocytic cells: a model of human lipomas. Carcinogenesis. 2003;24:1861-9.
99. Rogalla P, Blank C, Helbig R, Wosniok W, Bullerdiek J. Significant correlation between the breakpoints of rare clonal aberrations in benign solid tumors and the assignment of HMGIY retropseudogenes. Cancer Genet Cytogenet. 2001;130:51-6.
100. Esposito F, Martino M De, Petti MG, Forzati F, Tornincasa M, Federico A, Arra C, Pierantoni GM, Fusco A. HMGA1 pseudogenes as candidate proto-oncogenic competitive endogenous RNAs. Oncotarget. 2014;5:8341-54. doi: 10.18632/oncotarget.2202.
101. Esposito F, De Martino M, Forzati F, Fusco A. HMGA1-pseudogene overexpression contributes to cancer progression. Cell Cycle. 2014;13:3636-9.
102. Esposito F, De Martino M, D’Angelo D, Mussnich P, Raverot G, Jaffrain-Rea M-L, Fraggetta F, Trouillas J, Fusco A. HMGA1 -pseudogene expression is induced in human pituitary tumors. Cell Cycle. 2015;14:1471-5.
103. Li W, Yuan Y, Huang L, Qiao M, Zhang Y. Metformin alters the expression profiles of microRNAs in human pancreatic cancer cells. Diabetes Res Clin Pract. 2012;96:187-95.
104. Sehic A, Risnes S, Khuu C, Khan Q-E-S, Osmundsen H. Effects of in vivo transfection with anti-miR-214 on gene expression in murine molar tooth germ. Physiol Genomics. 2011;43:488-98.
105. Fedele M, Pierantoni GM, Visone R, Fusco A. Critical role of the HMGA2 gene in pituitary adenomas. Cell Cycle. 2006;5:2045-8.
106. Chiefari E, Iiritano S, Paonessa F, Le Pera I, Arcidiacono B, Filocamo M, Foti D, Liebhaber S a, Brunetti A. Pseudogene-mediated posttranscriptional silencing of HMGA1 can result in insulin resistance and type 2 diabetes. Nat Commun. 2010;1:40.
107. Hirotsune S, Yoshida N, Chen A, Garrett L, Sugiyama F, Takahashi S, Yagami K, Wynshaw-Boris A, Yoshiki A. An expressed pseudogene regulates the messenger-RNA stability of its homologous coding gene. Nature. 2003;423:91-6.
108. Kalyana-Sundaram S, Kumar-Sinha C, Shankar S, Robinson DR, Wu YM, Cao X, Asangani I a., Kothari V, Prensner JR, Lonigro RJ, Iyer MK, Barrette T, Shanmugam A, Dhanasekaran SM, Palanisamy N, Chinnaiyan AM. Expressed pseudogenes in the transcriptional landscape of human cancers. Cell. 2012;149:1622-34.
109. Sorge J, Gross E, West C, Beutler E. High level transcription of the glucocerebrosidase pseudogene in normal subjects and patients with Gaucher disease. J Clin Invest. 1990;86:1137-41.
110. Renaudie F, Yachou AK, Grandchamp B, Jones R, Beaumont C. A second ferritin L subunit is encoded by an intronless gene in the mouse. Mamm Genome. 1992;2:143-9.
111. Chakrabarti R, McCracken JB, Chakrabarti D, Souba WW. Detection of a functional promoter/enhancer in an intron-less human gene encoding a glutamine synthetase-like enzyme. Gene. 1995;153:163-9.
112. Thiele H, Berger M, Skalweit A, Thiele BJ. Expression of the gene and processed pseudogenes encoding the human and rabbit translationally controlled turnout protein (TCTP). Eur J Biochem. 2000;267:5473-81.
113. Suo G, Han J, Wang X, Zhang J, Zhao Y, Zhao Y, Dai J. Oct4 pseudogenes are transcribed in cancers. Biochem Biophys Res Commun. 2005;337:1047-51.
114. Zhang J, Wang X, Li M, Han J, Chen B, Wang B, Dai J. NANOGP8 is a retrogene expressed in cancers. FEBS J. 2006;273:1723-30.
115. Poliseno L. Pseudogenes: Newly Discovered Players in Human Cancer. Sci Signal. 2012;5:re5-re5.
116. Poliseno L, Marranci A, Pandolfi PP. Pseudogenes in Human Cancer. Front Med. 2015;2:68.

