INTRODUCTION
Long non-coding RNAs (lncRNAs) are now recognized as a major component of the human transcriptome, but the vast majority of these molecules remain to be functionally annotated [1, 2]. Nonetheless, key regulatory roles in fundamental cellular processes are already evident for several well characterized lncRNAs [3 – 5], and dysregulated expression of these may contribute to the pathogenesis of major diseases, notably cancer [3, 6, 7]. Consequently, the lncRNAs may offer new targets for the development of novel diagnostics and therapeutics.
The growth arrest-specific 5 (GAS5) gene is of particular relevance in this regard. GAS5 encodes lncRNA in addition to a diverse range of other non-coding RNAs, including small nucleolar RNAs, PIWI-interacting RNAs and possibly micro RNAs (miRNAs) [8–10]. GAS5 lncRNA is down-regulated in multiple cancers [11], including breast cancer [12]. In such cancers, clinico-pathological characteristics show inverse correlations with GAS5 lncRNA levels, and low GAS5 lncRNA levels are often predictive of poor prognosis [11]. A tumour suppressor role for GAS5 lncRNA is further indicated by its inhibition of tumour growth in xenograft models of breast and other cancers [11, 13].
At the cellular level, GAS5 lncRNA promotes growth arrest and/or apoptosis of multiple cell types [11], including hormone-sensitive and –insensitive breast cancer cells [12–14], which is likely to account for its tumour suppressor function. Since the action of radiation therapy and many chemotherapeutics depends on the engagement of the apoptotic machinery [15, 16], this is likely to be significant from a therapeutic perspective. Indeed, down-regulation of GAS5 lncRNA levels attenuates apoptosis induction by a broad range of treatments; for most, cell death shows a direct quantitative relationship with GAS5 lncRNA levels in breast and other cancer cells [14, 17, 18]. Consequently, enhancing cellular GAS5 lncRNA levels in tumour tissue may not only suppress the growth of such tumours but also enhance tumour cell killing by therapeutic agents, thereby improving patient outcomes.
One way to achieve this therapeutic goal may be to target the physiological mechanism that mediates the accumulation of GAS5 lncRNA levels in growth-arrested cells. GAS5 possesses a 5′-terminal oligopyrimidine (5′-TOP) sequence, therefore its translation is promoted by mTOR, which has high activity in actively growing cells [8, 19]. Because the GAS5 open reading frame is short and does not encode a functional protein, this in turn targets transcripts for degradation by nonsense-mediated decay (NMD), resulting in low cellular levels of GAS5 lncRNA [8, 19]. Conversely, inhibition of cell growth and mTOR activity prevents the active translation of GAS5 transcripts. Since degradation through NMD is dependent on active translation of the RNA concerned, GAS5 lncRNA accumulates upon growth arrest [8, 19]. While mTOR inhibitors increase GAS5 lncRNA levels in hormone-sensitive breast cancer cells, they are ineffective in triple-negative breast cancer (TNBC) cells and other hormone-independent cancer cells [14, 20], so that alternative, mTOR inhibitor-independent, approaches are required to activate this key pathway across a broad range of cancer subtypes.
Several molecular mechanisms of action have been proposed for GAS5 lncRNA, which offer possibilities in this respect. Firstly, it interacts with and riborepresses certain members of the steroid nuclear receptor superfamily, and thereby modulates the transcription of genes regulating apoptosis and the cell cycle. A stem-loop structure within GAS5 lncRNA 3′-terminal sequence (exon 12 –encoded), which serves as a hormone response element mimic (HREM), is required for this interaction [21]. Secondly, it can act as a miRNA sponge, since it binds to and modulates the levels of onco-miR21; a distinct (exon 4-derived) GAS5 sequence is required for this activity [13]. Functional analysis of mutated GAS5 lncRNA sequence has revealed that while apoptosis induction in lymphoid cells is only partially dependent on the GAS5 HREM sequence, more complete dependence is observed in breast and prostate cancer cells [22], implicating riborepression as the major mechanism by which GAS5 lncRNA induces the death of these cells. Novel oligonucleotide therapy, based on the GAS5 lncRNA HREM sequence, may therefore be feasible, but it is not yet clear if this sequence alone is sufficient to mediate apoptosis induction in epithelial cells, or if additional GAS5 lncRNA sequence is required. In order to address these questions, we have examined if the GAS5 HREM sequence alone is sufficient to promote the apoptosis of breast cancer cells, both basally and upon the application of an apoptotic stimulus, and how the resulting cellular responses compare with those of mature GAS5 lncRNA.
RESULTS
DNA oligonucleotides corresponding to the GAS5 HREM induce apoptosis in breast cancer cell lines
Nucleofection of a plasmid encoding mature GAS5 lncRNA induces apoptosis in hormone-sensitive and -insensitive breast cancer cell lines [12 – 14], and wild-type HREM sequence is required for this activity [22]. In initial experiments, we therefore examined if DNA oligonucleotides based on the wild-type GAS5 HREM sequence alone also induced apoptosis in a range of breast cancer cell lines; negative controls were nucleofected with DNA oligonucleotides either with stem complementarity but lacking the GAS5 HRE consensus (stem loop or SL control) or with scrambled GAS5 sequence (Scram control). Parallel nucleofection of a plasmid encoding GAS5 lncRNA (or empty plasmid vector as the respective control) was performed in some experiments, as a positive control.
In hormone-sensitive MCF7 cells, the apoptotic rate and culture viability were similar for cells nucleofected with either the Scram or SL DNA oligonucleotides (Figure 1A & 1B); levels were indistinguishable from those of mock-transfected cells (data not shown). Nucleofection of the HREM DNA oligonucleotide approximately doubled the apoptotic rate (Figure 1A) and produced a corresponding decrease in culture viability (Figure 1B) relative to the control DNA oligonucleotides. The extent of apoptosis induction by the HREM oligonucleotide (Figure 1A) was similar to that for MCF7 cells transfected with a plasmid encoding GAS5 lncRNA (Figure 1C), whereas the latter construct produced a more sustained loss of short-term culture viability (Figure 1D) than the HREM oligonucleotides (Figure 1B). Similar results were obtained for the other hormone-sensitive cell line, T-47D, studied here. Thus, in this cell line, the HREM oligonucleotide produced an approximate doubling of the basal apoptotic rate (Figure 1E), similar to the GAS5 lncRNA plasmid construct (Figure 1G), and a corresponding reduction in culture viability (Figure 1F) which was less sustained than for the GAS5 lncRNA plasmid construct (Figure 1H).
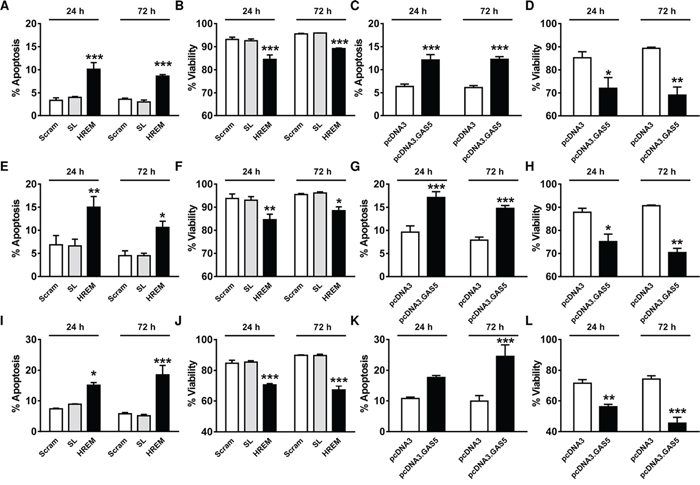
Figure 1: Effect of the GAS5 HREM DNA oligonucleotide on the basal survival of breast cancer cells. Cells (n = 4 cultures) were transfected with the GAS5 HREM DNA oligonucleotide (HREM) or scrambled (scram) or stem loop (SL) control DNA oligonucleotides; transfections were also conducted with the plasmids pcDNA3.GAS5 (encodes mature GAS5 lncRNA) and pcDNA3 (empty control vector) as controls. Cells were harvested at 24 h post-transfection for assessment of cell survival, and re-plated for re-assessment after a further 48 h. The GAS5 HREM oligonucleotide increased apoptosis and decreased cell viability in MCF7 (panels A & B, respectively), T-47D (panels E & F, respectively) and MDA-MB-231 (panels I & J, respectively) cells, similar to the action of GAS5 lncRNA (panels C & D, respectively for MCF7 cells; G & H, respectively for T-47D cells; and K & L, respectively, for MDA-MB-231 cells). *P < 0.05, **P < 0.01 and ***P < 0.001 versus scram and SL for HREM or versus pcDNA3 for pcDNA3.GAS5 (two-way ANOVA and Bonferroni’s MCT).
GAS5 lncRNA has previously been shown to induce apoptosis in hormone-insensitive breast cancer cells in addition to hormone-sensitive cell lines [17, 18]. Accordingly, in the TNBC line, MDA-MB-231, the GAS5 HREM DNA oligonucleotide also increased apoptosis (Figure 1I) and reduced culture viability (Figure 1J), similar to the effects of GAS5 lncRNA (Figure 1K & 1L).
The GAS5 HREM DNA oligonucleotide reduces the clonogenic activity of breast cancer cell lines
To determine the long-term consequences of treatment with the HREM oligonucleotide for cell survival, clonogenic growth assays were performed. Transfection of the HREM DNA oligonucleotide reduced the colony forming ability of MCF7, T-47D and MDA-MB-231 cells by 20 – 30% (Figure 2A, 2C & 2E, respectively) i.e., slightly less effectively than GAS5 lncRNA (25 – 40% inhibition, Figure 2B, 2D & 2F, respectively).
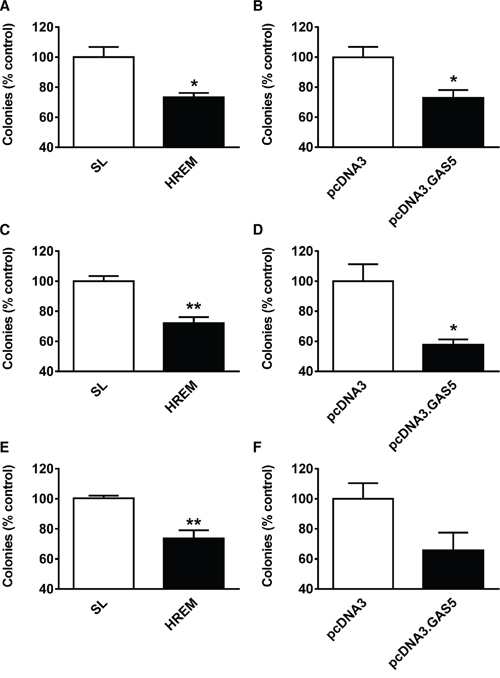
Figure 2: Effect of the GAS5 HREM DNA oligonucleotide on the clonogenic activity of breast cancer cells. Cells (n = 4 cultures) were transfected with the GAS5 HREM DNA oligonucleotide (HREM) or scrambled (scram) or stem loop (SL) control DNA oligonucleotides; transfections were also conducted with the plasmids pcDNA3.GAS5 (encodes mature GAS5 lncRNA) and pcDNA3 (empty control vector) as positive controls. Cells were harvested at 24 h post-transfection and an equal portion of each re-plated at low cell density for assessment of colony forming ability; data are expressed relative to the respective control. The GAS5 HREM oligonucleotide reduced the number of colonies formed by MCF7 (panel A), T-47D (panel C) and MDA-MB-231 (panel E) cells, similar to the action of GAS5 lncRNA (panels B, D & F, respectively). *P < 0.05 and **P < 0.01 versus SL for HREM or versus pcDNA3 for pcDNA3.GAS5 (Student’s t-test).
The GAS5 HREM DNA oligonucleotide promotes apoptosis induction upon DNA damage in breast cancer cell lines
In addition to modulating the basal apoptotic rate, GAS5 lncRNA promotes apoptosis induction in breast cells by a range of treatments, including chemotherapeutic agents and physical stimuli, such as irradiation with UV-C light, which activates the DNA damage response [12 – 14]. To test whether the HREM oligonucleotide exerts similar activity to GAS5 lncRNA in this respect, breast cancer cells were nucleofected with either the HREM or control (Scram and SL) oligonucleotides and then, after 24 h, cells were irradiated with UV-C light at dosages to induce apoptosis in ca. 50% cells. Parallel transfections were conducted with plasmid constructs ± GAS5 lncRNA, as a positive control. Prior transfection with the HREM oligonucleotide enhanced UV-C-induced apoptosis and correspondingly decreased culture viability in MCF7 (Figure 3A & 3B, respectively), T-47D (Figures 3E & 3F, respectively) and MDA-MB-231 (Figures 3I & 3J, respectively) cells. In general, the magnitude of these effects were similar to those of GAS5 lncRNA (Figure 3C & 3D for MCF7, Figure 3G & 3H for T-47D, and Figure 3K & 3L for MDA-MB-231), further highlighting the similar activities of the GAS5 HREM and GAS5 lncRNA sequences.
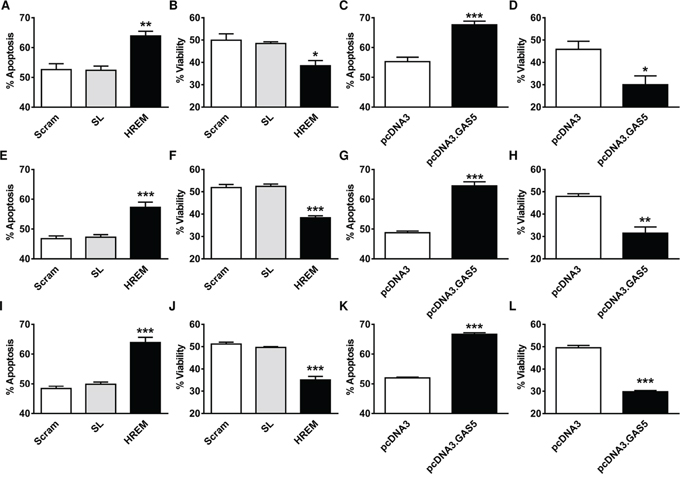
Figure 3: Effect of the GAS5 HREM DNA oligonucleotide on UV-C-induced cell death of breast cancer cells. Cells (n = 4 cultures) were transfected with the GAS5 HREM DNA oligonucleotide (HREM) or scrambled (scram) or stem loop (SL) control DNA oligonucleotides; transfections were also conducted with the plasmids pcDNA3.GAS5 (encodes mature GAS5 lncRNA) and pcDNA3 (empty control vector) as positive controls. Cells were harvested at 24 h post-transfection, irradiated with UV-C light and re-plated for assessment of cell survival after a further 48 h. The GAS5 HREM oligonucleotide increased UV-C induced apoptosis and the associated loss of cell viability in MCF7 (panels A & B, respectively), T-47D (panels E & F, respectively) and MDA-MB-231 (panels I & J, respectively) cells, similar to the action of GAS5 lncRNA (panels C & D, respectively for MCF7 cells; G & H, respectively for T-47D cells; and K & L, respectively, for MDA-MB-231 cells). *P < 0.05, **P < 0.01 and ***P < 0.001 versus scram and SL for HREM or versus pcDNA3 for pcDNA3.GAS5 (one-way ANOVA and Bonferroni’s MCT).
Sequence specificity of GAS5 HREM action
Two guanine-containing pairs (G549 and G559) within the HREM of GAS5 lncRNA are essential for its interaction with the DNA binding domain of the GR [22]. Mutation of one of these (G549A) is sufficient to inhibit the GR/GAS5 lncRNA interaction and to prevent the induction of apoptosis by GAS5 lncRNA in breast and prostate cell lines [22]. We therefore examined if induction of apoptosis by the GAS5 HREM oligonucleotide exhibited similar sequence specificity by employing a modified DNA oligonucleotide carrying the equivalent mutation. In MCF7 cells, the oligonucleotide corresponding to the wild-type GAS5 HREM sequence induced apoptosis (Figure 4A) and reduced short-term culture viability (Figure 4B) compared with the SL control oligonucleotide, whereas the effects of the oligonucleotide with mutated HREM sequence were indistinguishable from those of the SL control (Figure 4A & 4B).
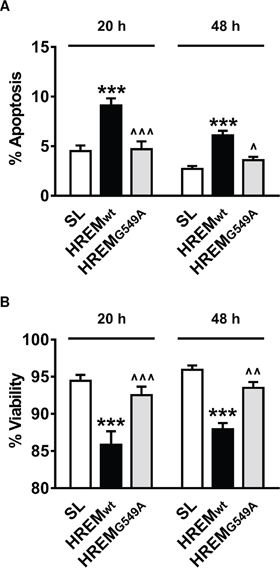
Figure 4: Sequence specificity of the GAS5 HREM DNA oligonucleotide. MCF7 cells (n = 5 cultures) were transfected with either the wild-type GAS5 HREM DNA oligonucleotide (HREMwt), a mutated GAS5 HREM DNA oligonucleotide (HREMG549A) or the standard stem loop (SL) control DNA oligonucleotides. Cells were harvested at 20 and 48 h post-transfection, for assessment of cell survival. Only the HREMwt oligonucleotide increased apoptosis (panel A) and decreased cell viability (panel B) relative to the SL control oligonucleotide. ***P < 0.001 versus SL, and ^P < 0.05, ^^P < 0.01 and ^^^P < 0.001 for HREMG549A versus HREMwt (two-way ANOVA and Bonferroni’s MCT).
The GAS5 HREM oligonucleotide restores cell death inhibited by low endogenous GAS5 lncRNA expression
Endogenous GAS5 lncRNA levels are reduced in breast tumour versus adjacent normal tissue and in hormone-insensitive versus hormone-sensitive breast cancer cells [12 – 14]. In hormone-sensitive breast cancer cells, siRNA-mediated knockdown of GAS5 lncRNA attenuates responses to a range of pro-apoptotic treatments, including UV-C irradiation [13, 14]. To investigate if the HREM DNA oligonucleotide can overcome such resistance consequent upon low endogenous GAS5 lncRNA levels, we used siRNA to silence GAS5 expression in MCF7 cells prior to transfection of the HREM (and control) oligonucleotide, then examined cellular responses ± irradiation with UV-C light.
The GAS5 siRNA markedly reduced GAS5 lncRNA levels immediately prior to oligonucleotide nucleofection (Figure 5A) but this had no effect on basal apoptosis (Figure 5B) or cell survival (Figure 5C) at 20 h post-transfection of either the SL or HREM oligonucleotide; the HREM oligonucleotide produced the expected changes of similar magnitude in these parameters, irrespective of the endogenous GAS5 lncRNA levels. Cells were then treated ± UV-C light then studied after a further 48 h culture; at this time point, the mock-irradiated controls demonstrated a similar pattern of change in apoptosis (Figure 5D) and cell viability (Figure 5E) as at 20 h post-transfection (in comparison to Figure 5B & 5C, respectively), notwithstanding the expected decline in the proportion of cells undergoing active apoptosis. As regards UV-C irradiation, prior knockdown of GAS5 lncRNA clearly attenuated apoptosis induction (Figure 5F) and loss of culture viability (Figure 5G), as judged from the responses of cells which had received the SL control oligonucleotide. Crucially however, prior transfection with the HREM oligonucleotide enhanced UV-C-induced apoptosis (Figure 5F) and the associated loss of culture viability (Figure 5G), irrespective of the level of endogenous GAS5 lncRNA. Notably, the proportions of apoptotic (and viable) cells were similar for the NC-HREM and #4-HREM treatment groups, demonstrating that the HREM oligonucleotide can prevent the attenuation of cell death caused by reduced GAS5 lncRNA levels.
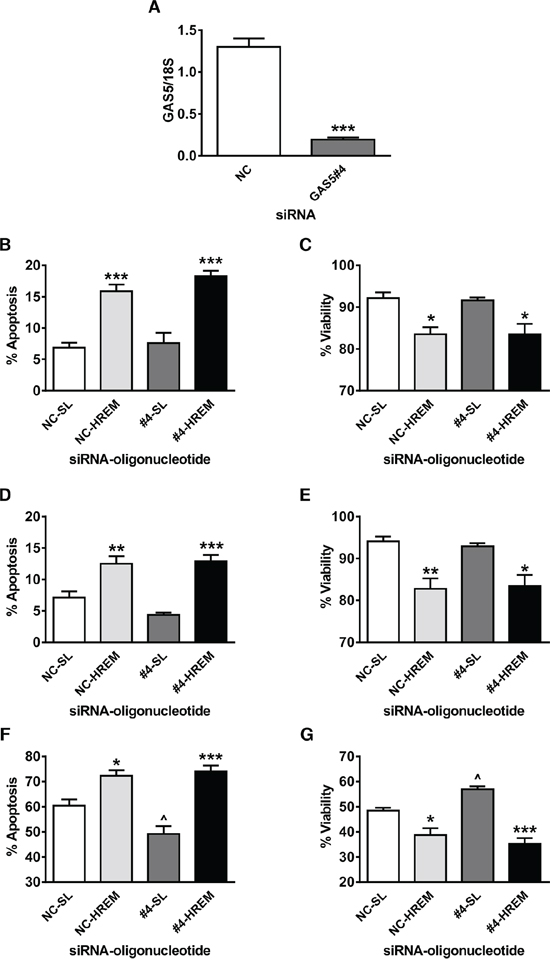
Figure 5: Effect of the GAS5 HREM DNA oligonucleotide on basal and UV-C-induced cell death after silencing of endogenous GAS5 expression in MCF7 cells. Cells (n = 4 cultures) were transfected with either GAS5 siRNA (GAS5#4; targets exon 12 sequence) or negative control (NC) siRNA and, after 24 h, transfected with either the GAS5 HREM DNA oligonucleotide (HREM) or stem loop (SL) control DNA oligonucleotides. After 20 h, cells were irradiated with UV-C light, then plated for assessment of cell survival after a further 48 h. The GAS5#4 siRNA markedly reduced GAS5 lncRNA levels at 24 h post-transfection (panel A) i.e., immediately prior to nucleofection of DNA oligonucleotides. The HREM oligonucleotide induced apoptosis (panel B) and decreased culture viability (panel C) at 20 h post-transfection, irrespective of endogenous GAS5 lncRNA levels; a similar pattern was found for apoptosis (panel D) and culture viability (panel E) in mock-irradiated controls (i.e. at 68 h post-oligonucleotide transfection). Silencing of GAS5 attenuated UV-C induced apoptosis (panel F) and the associated loss of culture viability (panel G) at 48 h post-irradiation, but had no effect cell death induction by the HREM oligonucleotide. Panel A: ***P < 0.001 versus cells transfected with NC siRNA (Student’s t-test). Panels B – H: *P < 0.05, **P < 0.01 & ***P < 0.001 versus cells transfected with SL oligonucleotide; and ^P < 0.05 versus cells transfected with NC siRNA (one-way ANOVA and Bonferroni’s MCT).
The GAS5 HREM DNA oligonucleotide has no effect on endogenous GAS5 lncRNA levels
GAS5 lncRNA exerts pro-apoptotic effects in breast cancer cell lines as does the GAS5 HREM DNA oligonucleotide (Figure 1A, 1E & 1I). To address the postulate that the GAS5 HREM oligonucleotide may exert its pro-apoptotic activity via changes in endogenous levels of GAS5, RT-qPCR analysis was performed on MCF7, T-47D and MDA-MB-231 cell lines at 20 h post-transfection of control and GAS5 HREM DNA oligonucleotides. In all cases, GAS5 lncRNA levels were similar between cells transfected with the GAS5 HREM and control oligonucleotides (Figure 6), excluding an indirect role for GAS5 lncRNA per se in GAS5 HREM oligonucleotide action.
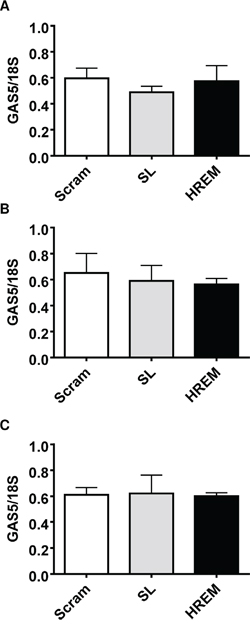
Figure 6: Effect of the GAS5 HREM DNA oligonucleotide on endogenous GAS5 lncRNA levels in breast cancer cells. Cells (n = 4 cultures) were transfected with either the GAS5 HREM DNA oligonucleotide (HREM) or scrambled (scram) or stem loop (SL) control DNA oligonucleotides and cells were harvested at 24 h post-transfection for determination of GAS5 lncRNA levels by RT-qPCR. The GAS5 HREM oligonucleotide had no effect on GAS5 lncRNA levels in MCF7 (panel A), T-47D (panel B) and MDA-MB-231 (panel C).
Chemically modified GAS5 HREM oligonucleotides also induce apoptosis
Since experiments so far had employed a HREM DNA oligonucleotide, although the naturally occurring HREM is present within lncRNA, we wished to examine the efficacy of a HREM RNA oligonucleotide. Since RNA is prone to degradation, especially within exonuclease-rich cellular environments, we additionally examined several oligonucleotides with chemical modifications to the backbone that improve stability in vivo [23], i.e., phosphorothioate RNA [RNA(PT)] and locked nucleic acid (LNA) oligonucleotides. In these screening studies, nucleofection of both unmodified RNA and RNA(PT) HREM oligonucleotides stimulated the basal apoptosis of MCF7 cells, whereas the LNA oligonucleotide was less effective and did not reach statistical significance (Figure 7A). Short-term culture viability was also reduced by the GAS5 HREM RNA and RNA(PT) oligonucleotides (Figure 7B). As regards long-term survival however, only the GAS5 HREM RNA(PT) oligonucleotide was able to reduce the colony forming ability of MCF7 cells to a degree that was statistically significant (Figure 7C).
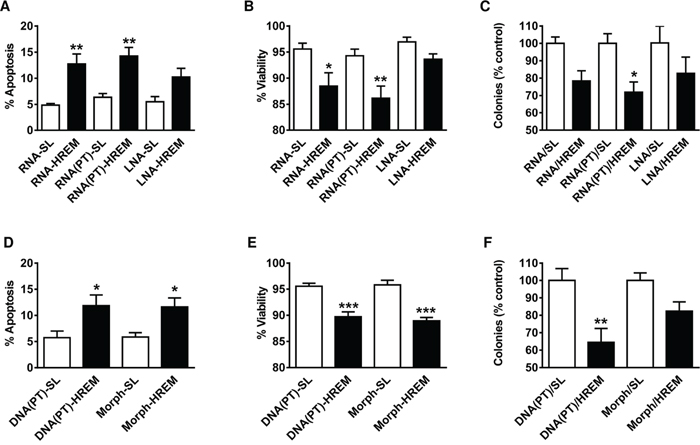
Figure 7: Effect of chemically modified GAS5 HREM RNA and DNA oligonucleotides on the basal survival of MCF7 cells. Cells (n = 4 cultures) were transfected with a range of chemically modified GAS5 HREM and corresponding stem loop (SL) control oligonucleotides comprising: unmodified RNA (RNA-HREM & RNA-SL); phosphorothioate RNA (RNA(PT)-HREM & RNA(PT)-SL); locked nucleic acid (LNA-HREM & LNA-SL); phosphorothioate DNA (DNA(PT)-HREM & DNA(PT)-SL); and morpholinos (morph-HREM & morph-SL). Cells were harvested at 20 h post-transfection for assessment of cell survival, and an equal portion of each re-plated at low cell density for assessment of colony forming ability; the latter data are expressed relative to the respective control. Of the RNA-based oligonucleotides, the LNA-HREM effect did not reach statistical significance but both the RNA-HREM and the RNA(PT)-HREM oligonucleotides increased apoptosis (panel A) and decreased cell viability (panel B); the inhibition of clonogenic growth reached statistical significance for only the RNA(PT)-HREM oligonucleotide (panel C). Both the DNA(PT)-HREM and morph-HREM oligonucleotides also increased apoptosis (panel D) and decreased cell viability (panel E), but the inhibition of clonogenic growth reached statistical significance for only the DNA(PT)-HREM (panel F). *P < 0.05, **P < 0.01 and ***P < 0.001 versus the corresponding SL control (one-way ANOVA and Bonferroni’s MCT).
Given these findings, it was of interest to determine if a phosphorothioate DNA [DNA(PT)] form of the GAS5 HREM also induces apoptosis in MCF7 cells. In addition, a morpholino HREM oligonucleotide was included in this screen, as the utility of this chemical modification has already been demonstrated for other types of therapeutic oligonucleotide (notably antisense oligonucleotides) [23]. Both chemically modified forms of the HREM DNA oligonucleotide induced apoptosis (Figure 7D) and reduced short-term viability (Figure 7E) in MCF7 cells. However, only the DNA(PT) oligonucleotide reduced the long-term survival of MCF7 cells to a degree that was statistically significant (Figure 7F). Notably, the extent of this reduction was similar to that of the RNA(PT) form of the GAS5 HREM oligonucleotide (compare Figure 7C & 7F).
DISCUSSION
Currently, there is a major need to develop new targeted therapies for breast and other cancers. In the case of breast cancer, this is especially true for hormone-insensitive forms of disease, such as TNBC, which is often characterised by highly malignant behaviour and resistance to conventional chemotherapy [24]. Much effort is being focussed on the development of drugs which directly target oncoproteins [24, 25], while increasing knowledge of the importance of lncRNAs in health and disease may offer opportunities for the design of entirely novel oligonucleotide-based oncotherapies. Towards this goal, we have exploited recent knowledge regarding the structure/function relationship of the relatively well characterized GAS5 lncRNA [22], an apoptosis-promoting molecule and putative tumour suppressor [11], and demonstrate here for the first time that oligonucleotides based on the HREM sequence alone are sufficient to induce the apoptosis of breast cells. This work is of more general significance, since it provides the first proof-of-principle of using partial sequences to mimic the action of tumour suppressive lncRNA molecules and, as such, points to an innovative approach for oligonucleotide-based oncotherapies of potential clinical importance.
GAS5 lncRNA levels are down-regulated in a wide range of tumours, including breast cancer; in many cancers low levels of expression are prognostic of poor patient survival [11]. The basis for this tumour suppressor function has been ascribed to the apoptosis-promoting activity of GAS5 lncRNA, which has itself been shown to reside within the HREM portion of the GAS5 lncRNA molecule in breast and other cells [21, 22]. The present study, which employs independent assays of apoptotic morphology, short-term culture viability and long-term clonogenic survival, clearly demonstrates that the HREM portion alone of the GAS5 lncRNA molecule induces cell death in three different breast cancer cell lines, including both hormone-sensitive and –insensitive breast cancer cells. Notably, for all cells, the extent of apoptosis induction and inhibition of growth were similar for both the GAS5 HREM oligonucleotide and GAS5 lncRNA.
The GAS5 HREM oligonucleotide also enhanced the action of an external apoptotic stimulus, UV-C irradiation. For both the partial and full length GAS5 lncRNA sequences, this effect was additive rather than synergistic, as previously noted for the latter sequence [12, 14]. Of direct clinical relevance, reduced endogenous GAS5 lncRNA expression, such as occurs in breast cancer and many other tumours [11 – 14], attenuates cellular responses to apoptotic inducers, including UV-C and various chemotherapeutic agents [13, 14]; the GAS5 HREM was able to restore breast cancer cell sensitivity, as demonstrated here for UV-C irradiation. Thus in cancers with deficient GAS5 expression, the GAS5 HREM oligonucleotide may have therapeutic potential, not only by acting as an apoptotic inducer per se, but also by restoring the effectiveness of more conventional oncotherapies on therapy-resistant cancer cells.
An important question relates to the mechanism of action of the GAS5 HREM oligonucleotide in promoting the apoptosis of breast cancer cells. Our current working hypothesis is that the partial HREM sequence acts as a decoy for members of the steroid hormone nuclear receptor superfamily, as has been demonstrated for GAS5 lncRNA [21, 22]. In order to test this hypothesis we examined the effect of a potentially inactivating mutation in the HREM on its cytotoxic effects. These experiments demonstrate that the HREM oligonucleotide has identical sequence specificity to GAS5 lncRNA. Thus, mutation of a single base (G549A) of the HREM of GAS5 lncRNA diminishes steroid receptor binding and ablates apoptosis induction in breast cancer cells [22], and an identical mutation of the GAS5 HREM oligonucleotide also prevents its induction of apoptosis in MCF7 cells (Figure 4), indicating that the mechanism of induction of apoptosis in breast cancer cells by the HREM is the same as that of GAS5 lncRNA. It is as yet unknown which particular member(s) of the steroid hormone nuclear receptor superfamily interact with GAS5 lncRNA in breast cells, particularly in relation to apoptosis induction; consequently, we have been unable to address this issue for the GAS5 HREM oligonucleotide. In this regard, the GAS5 HREM does not interact with estrogen receptors [21, 22], which are crucial for mediating pro-survival signalling by estrogen in breast cells [26]. While the GAS5 HREM interacts with progesterone receptor [21, 22], this interaction is unlikely to play a major role in breast cancer cell survival, since both GAS5 lncRNA [14] and the GAS5 HREM sequence induce apoptosis in TNBC cells which are devoid of progesterone receptors. On the other hand, a more indirect mechanism of action for the GAS5 HREM oligonucleotide, centring on endogenous GAS5 lncRNA per se, can be excluded, since: i. cellular levels of the latter were unperturbed following transfection of the GAS5 HREM oligonucleotide; and ii. siRNA-mediated silencing of GAS5 lncRNA had no effect on the apoptosis-inducing activity of the GAS5 HREM oligonucleotide in MCF7 cells.
These findings prove that for GAS5 lncRNA, it is feasible to employ a partial sequence to mimic a desirable function of the parental lncRNA and thereby point to the possibility of novel oligonucleotide mimic-based oncotherapeutic approaches. With this in mind, a range of commonly used chemical modifications which are known to improve oligonucleotide stability in vivo [23], were tested for the HREM oligonucleotide in functional assays. Following nucleofection, most modified HREM oligonucleotides were able to induce apoptosis and decrease short-term culture viability, although this effect was not statistically significant for the locked nucleic acid form. In terms of inhibition of long-term cell survival, phosphorothioate forms of RNA and DNA oligonucleotides appeared optimal. These findings indicate that certain, but not all, modifications to the oligonucleotide backbone are well tolerated in functional terms by the GAS5 HREM oligonucleotide.
Mimic oligonucleotides may have widespread applicability as our understanding of the structure/function relationships of lncRNA molecules increases. A particular advantage of such therapies is the ability to mimic the activity of tumour suppressor genes, which are often down-regulated in tumours, whereas current protein-targeted, drug development approaches are biased towards the inhibition of oncogenic proteins, which are typically overexpressed in tumours. Furthermore, the effectiveness of partial sequences, as seen above, makes oligonucleotide-based approaches more amenable, since these are inherently less complex and therefore more practical clinically than gene therapy approaches. Examples of other therapeutic approaches that exploit emerging knowledge of lncRNA biology and are currently under development include siRNA, other RNAi and antisense oligonucleotide approaches for targeting oncogenic lncRNAs, antagoNAT antisense oligonucleotides for targeting antisense lncRNAs to up-regulate specific mRNAs/proteins, and small molecule inhibitor (including oligonucleotides) approaches [23, 27–29]; the oligonucleotide mimic approach proposed here is likely to be compatible with and complementary to these. Indeed, the challenges for clinical translation, in terms of in vivo stability, tumour delivery and cellular uptake, for example, are likely to be similar for all these technologies and are currently the focus of active investigation.
MATERIALS AND METHODS
Cell lines and culture conditions
This study employed hormone-sensitive (MCF7 and T-47D) and –insensitive (MDA-MB-231; TNBC cells) breast cancer cell lines. All cultures were generated from secondary stocks of cells which had been frozen down within two weeks receipt from ATCC-LGC Promochem (Teddington, Middlesex, UK), and cultures were passaged for a maximum period of eight weeks before replacement with fresh cell stocks. Culture medium was R-10 medium, comprising RPMI-1640 supplemented with 2 mM L-glutamine, 1 mM sodium pyruvate, 10 mM HEPES, 10% fetal bovine serum and 50 μg/ml gentamicin, and cells were cultured at 37 °C in a humidified incubator with 5% CO2. Serum was from Labtech International Ltd (Uckfield, UK) and the remaining cell culture reagents were from Sigma-Aldrich Company Ltd (Gillingham, UK).
Nucleofection of oligonucleotides and plasmid DNA
DNA oligonucleotides (Eurofins Genomics, Ebersberg, Germany) were studied in the majority of experiments and comprised the wild-type GAS5 HREM sequence (5′- CAGTGGTCTTTGTAGACTGCCTG-3′), a mutated GAS5 HREM (5′- CAGTAGTCTTTGTAGACTGCCTG -3′), a control sequence which retains stem complementarity but lacks the GAS5 HRE consensus (stem loop control; 5′- CTGATGGTCTTTGTAGACCATCA-3′) and scrambled sequence (5′- TGTTGGCTTGTCACGCATGCGTCT-3′). Equivalent stem loop control and wild-type GAS5 HREM sequences were also employed in experiments with RNA oligonucleotides (Eurofins Genomics, Ebersberg, Germany), DNA and RNA phosporothioate-modified oligonucleotides (GE Healthcare Dharmacon Inc, Little Chalfont, UK) and morpholino oligonucleotides (Gene Tools LLC, Philomath, USA). Sequences of locked nucleic acid oligonucleotides (Exiqon, Vedbaek, Denmark) containing selective phosphorothioate backbone modifications (as indicated by *) were: 5′-C+T+G+A+T+G*G*T*C*T*T*T*G*T+A+G+A+C+C*A*T*C*A-3′ for the stem loop control; and C+A+G+T+G+GT*C*T*T*T*G*T*A*G+A+C+T+G+C*C*T*G-3′ for the GAS5 HREM. Cells (2 × 106 in 0.1 ml Ingenio electroporation solution [Mirus Bio LLC, Madison, WI, USA]) were nucleofected with 5 pmol oligonucleotide or 2 μg plasmid (either pcDNA3/GAS5—encodes mature GAS5 lncRNA—or empty pcDNA3 vector as control; positive control experiments) using programmes E-014, X-005 and X-013 for MCF7, T-47D and MDA-MB-231 cells, respectively, and cells were plated in 3 ml R-10 medium in 6-well plates.
RNA interference by siRNA
MCF7 cells were transfected with Ambion Silencer Select GAS5 siRNA (product code n272331; targets exon 12; Life Technologies, Paisley, UK) or negative control (NC) siRNA (product code AM4611) using HiPerFect reagent (Qiagen, Crawley, UK) and a fast forward protocol. This particular GAS5 siRNA (termed GAS5 siRNA #4) has previously been shown to silence GAS5 expression in MCF7 cells using an RNAiFect protocol [14]. Briefly, trypsinized cells (2 × 105) were seeded in 2.3 ml R-10 medium per well of a 6-well plate. Complexes were prepared by mixing 12 pmol siRNA in 0.1 ml Opti-MEM-I medium with 12 μl HiPerFect reagent and, after 15 min, these were added to cells. Cells were then cultured for 20 – 24 h prior to nucleofection with the indicated oligonucleotides.
Induction of cell death and cell survival assays
At 20 h post-nucleofection with oligonucleotides, cells were trypsinized and resuspended at 2 × 105 cells/ml in R-10 medium. A sample was taken for RNA isolation, and cells were routinely seeded at 1.6 × 105 cells/well of a 12-well plate. In experiments employing ultraviolet-C (UV-C) irradiation for apoptosis induction, this was performed immediately prior to plating, as described elsewhere [17]; the dose used was 40 J/m2 for all cell lines; controls were mock-irradiated. Cells were cultured for a further 48 h, then adherent cells were trypsinized and combined with non-adherent cells for analysis of parameters of cell survival.
Apoptosis was routinely determined by assessment of nuclear morphology of cells by fluorescence microscopy, after staining with acridine orange (25 μg/ml); cells containing condensed or fragmented chromatin were scored as apoptotic. Cell viability was determined by dye-exclusion assay by counting of nigrosin blue (0.1% (w/v)) stained samples using a haemocytometer and light microscopy. To determine long-term survival of cells, a clonogenic assay was performed: cells were replated in R-10 medium supplemented with 10 % (v/v) cell-conditioned medium in 6-well plates, cultured for 3 weeks and the number of colonies formed was determined post-staining with crystal violet.
Real time RT-PCR (RT-qPCR)
Total RNA was isolated with TRIzol, DNase-treated, then reverse transcribed by random hexamer priming using SuperScript II Reverse Transcriptase [14]. Real-time PCR was conducted on the resulting cDNA using a SensiFast Probe Hi-ROX kit (Bioline, London, IK) and TaqMan Gene Expression Assays (assay codes Hs99999901_m1 for 18S and Hs03464472_m1 for GAS5; Life Technologies, Paisley, UK); assays usually contained 10 ng sample cDNA in a final volume of 25 μl and were run on an ABI Prism Sequence Detection System model 7000. Standards (0.2 – 60 ng cDNA from untransfected MCF7 cells) were included with each assay; the resulting standard curves of threshold cycle (CT) value versus log input standard cDNA were used to calculate input amounts of samples from their respective CT values, and data were expressed relative to 18S rRNA.
Statistical analyses
Data are presented as the mean ± SEM; the number of observations (n) refers to different transfected samples, each from a separate culture of cells. Data analysis was either by an unpaired Student’s t-test (when comparing two groups only), by one-way analysis of variance, with Bonferroni’s multiple comparison test (MCT), or by two-way analysis of variance, with Bonferroni’s post tests, as indicated in the text. Homogeneity of variance was checked by Bartlett’s test and, where necessary, data were transformed (log or square root) prior to analysis. Statistical analyses were performed using GraphPad Prism v4.03.
ACKNOWLEDGMENTS AND FUNDING
This work was supported by grants from Breast Cancer Now.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Hangauer MJ, Vaughn IW, McManus MT. Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS genetics. 2013; 9:e1003569
2. Mattick JS, Rinn JL. Discovery and annotation of long noncoding RNAs. Nature structural & molecular biology. 2015; 22: 5-7.
3. Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Molecular cancer. 2011; 10: 38.
4. Kung JT, Colognori D, Lee JT. Long noncoding RNAs: past, present, and future. Genetics. 2013; 193: 651-669.
5. Yang L, Froberg JE, Lee JT. Long noncoding RNAs: fresh perspectives into the RNA world. Trends in biochemical sciences. 2014; 39: 35-43.
6. Gutschner T, Diederichs S. The hallmarks of cancer: a long non-coding RNA point of view. RNA biology. 2012; 9: 703-719.
7. Cheetham SW, Gruhl F, Mattick JS, Dinger ME. Long noncoding RNAs and the genetics of cancer. British journal of cancer. 2013; 108: 2419-2425.
8. Smith CM, Steitz JA. Classification of gas5 as a multi-small-nucleolar-RNA (snoRNA) host gene and a member of the 5’-terminal oligopyrimidine gene family reveals common features of snoRNA host genes. Molecular and cellular biology. 1998; 18: 6897–6909.
9. Brameier M, Herwig A, Reinhardt R, Walter L, Gruber J. Human box C/D snoRNAs with miRNA like functions: Expanding the range of regulatory RNAs. Nucleic acids research. 2011; 39: 675–686.
10. He X, Chen X, Zhang X, Duan X, Pan T, Hu Q, Zhang Y, Zhong F, Liu J, Zhang H, Luo J, Wu K, Peng G, Luo H, Zhang L et al. An LncRNA (GAS5)/SnoRNA-derived piRNA induces activation of TRAIL gene by site-specifically recruiting MLL/COMPASS-like complexes. Nucleic acids research. 2015; 43: 3712–3725.
11. Pickard MR, Williams GT. Molecular and cellular mechanisms of action of tumour suppressor GAS5 lncRNA. Genes (Basel). 2015; 6: 484-499.
12. Mourtada-Maarabouni M, Pickard MR, Hedge VL, Farzaneh F, Williams GT. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene. 2009; 28: 195-208.
13. Zhang Z, Zhu Z, Watabe K, Zhang X, Bai C, Xu M, Wu F, Mo YY. Negative regulation of lncRNA GAS5 by miR-21. Cell death and differentiation. 2013; 20: 1558-1568
14. Pickard MR, Williams GT. Regulation of apoptosis by long non-coding RNA GAS5 in breast cancer cells: Implications for chemotherapy. Breast cancer research and treatment. 2014; 145: 359–370.
15. McKenzie S, Kyprianou N. Apoptosis evasion: the role of survival pathways in prostate cancer progression and therapeutic resistance. Journal of cellular biochemistry. 2006; 97: 18-32.
16. Indran IR, Tufo G, Pervaiz S, Brenner C. Recent advances in apoptosis, mitochondria and drug resistance in cancer cells. Biochimica et biophysica acta. 2011; 1807: 735-745.
17. Pickard MR, Mourtada-Maarabouni M, Williams GT. Long non-coding RNA GAS5 regulates apoptosis in prostate cancer cell lines. Biochimica et biophysica acta. 2013; 1832: 1613-1623.
18. Dong S, Qu X, Li W, Zhong X, Li P, Yang S, Chen X, Shao M, Zhang L. The long non-coding RNA, GAS5, enhances gefitinib-induced cell death in innate EGFR tyrosine kinase inhibitor-resistant lung adenocarcinoma cells with wide-type EGFR via downregulation of the IGF-1R expression. Journal of hematology & oncology. 2015; 8: 43.
19. Williams GT, Farzaneh F. Are snoRNAs and snoRNA host genes new players in cancer? Nature reviews cancer. 2012; 12: 84-88.
20. Yacqub-Usman K, Pickard MR, Williams GT. Reciprocal regulation of GAS5 lncRNA levels and mTOR inhibitor action in prostate cancer cells. The prostate. 2015; 75: 693-705.
21. Kino T, Hurt DE, Ichijo T, Nader N, Chrousos GP. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Science signaling. 2010; 3: ra8.
22. Hudson WH, Pickard MR, de Vera IM, Kuiper EG, Mourtada-Maarabouni M, Conn GL, Kojetin DJ, Williams GT, Ortlund EA. Conserved sequence-specific lincRNA-steroid receptor interactions drive transcriptional repression and direct cell fate. Nature Communications. 2014; 5: 5395.
23. Chan JH, Lim S, Wong WS. Antisense oligonucleotides: from design to therapeutic application. Clinical and experimental pharmacology and physiology. 2006; 33: 533-540.
24. Engebraaten O, Vollan HK, Børresen-Dale AL. Triple-negative breast cancer and the need for new therapeutic targets. The American journal of pathology. 2013; 183: 1064-1074.
25. Mohamed A, Krajewski K, Cakar B, Ma CX. Targeted therapy for breast cancer. The American journal of pathology. 2013; 183: 1096-1112.
26. Lewis-Wambi JS, Jordan VC. Estrogen regulation of apoptosis: how can one hormone stimulate and inhibit? Breast cancer research. 2009; 11: 206.
27. Roberts TC, Wood MJ. Therapeutic targeting of non-coding RNAs. Essays in biochemistry. 2013; 54: 127-145.
28. Wahlestedt C. Targeting long non-coding RNA to therapeutically upregulate gene expression. Nature reviews drug discovery. 2013; 12: 433-446.
29. Fatima R, Akhade VS, Pal D, Rao SM. Long noncoding RNAs in development and cancer: potential biomarkers and therapeutic targets. Molecular and cellular therapies. 2015; 3: 5.

