INTRODUCTION
The RUNX proteins belong to a family of transcription factors (RUNX1, 2 and 3) known to play crucial roles in hematopoiesis, osteogenesis and neurogenesis [1]. RUNX1 activity has been comprehensively study in physiological and tumor contexts [2, 3]. It has a runt domain and is able to bind a common TG (T/C)GGT consensus binding site, inducing proliferation in a context-dependent manner [4–6]. Moreover, RUNX1 is considered as a multifaceted protein that associates with diverse partners to direct different biological outcomes [2]. In particular Runx1 has both transcriptional activation and inhibition domains that allow it to bind a plethora of co-factors, such as the tumor suppressor gene Foxp3, which in turn modulates Runx1’s regulatory effect [7, 8]. Sakaguchi and colleagues showed that Foxp3 transcription factor inhibits Runx1 transcriptional activity by protein-protein interaction promoting the suppressive function of regulatory T cells [8]. In human breast cancer, Runx1 activity is still matter of debate and little is known about its direct role in breast cancer progression [9–12]. Interestingly, Ferrari and colleagues have shown using multivariate analysis that high expression of RUNX1 correlates with poor prognosis in triple negative human breast cancer and strongly suggest that Runx1 could be used as an independent prognostic marker in this subgroup of human breast cancer [13].
Foxp3 is an X-linked tumor suppressor gene expressed in the normal mammary gland, but downregulated, mutated or cytoplasmically localized in mammary tumor cells [14–18]. It has been shown that in normal cells, wild-type FOXP3 is bound to the promoter and transcriptionally repress human epidermal growth factor receptor Her-2/Neu and miR-146a promoters, transcriptionally repressing expression of these mediators that are involved in mammary tumorigenesis [15, 19]. Moreover, FOXP3 overexpression in human cancer cell lines was shown to repress tumor growth and metastasis [15, 19, 20].
This context encouraged us to investigate if Runx1 is able to modulate gene expression in mammary tumor cells, and whether this activity could be modulated by Foxp3 in normal epithelial cells. To achieve this goal, we investigated Runx1 and Foxp3 participation in the regulation of expression of two tumor associated genes, Rspo3 [21, 22] and GJA1 [23, 24], which are known modulators of breast tumor cell growth (positively and negatively, respectively). Both promoter regions possess Runx1 binding sites, but no Foxp3-binding regions were detected in their proximity. Runx1 is able to promote RSPO3 gene expression and inhibit GJA1 gene expression on tumor epithelial cells, depending on Foxp3 availability. Our results show, for the first time, that Foxp3 thwarts Runx1 activity through physical interaction in mammary epithelial cells. Furthermore, these data suggest that Runx1 might modulate mammary gland tumorigenesis depending on Foxp3 expression levels unraveling a new mechanism of gene expression regulation on mammary epithelial cells.
RESULTS
Runx1 activates RSPO3 oncogene expression in tumor cells
R-spondin protein 3 (RSPO3) belongs to a family of secreted proteins that strongly potentiates Wnt/βcatenin signaling [25, 26] and regulates tissue patterning and differentiation [27, 28]. In particular, RSPO3 has been described as a potent oncogene due to its ability to transform and generate mammary tumors in vivo after inoculation of RSPO3-transduced epithelial mammary cells [22]. Furthermore, we and other laboratories, described that MMTV-induced mammary gland tumors express high levels of RSPO3 compared with virgin normal mammary gland [21, 22]. To address the question of how this oncogene expression is differentially regulated in normal and tumor mammary epithelial cells, we analyze the promoter region of RSPO3. In silico analysis of Rspo3 promoter region (1500 bp upstream from +1 transcription start site) revealed three putative binding sites for the transcription factor Runx1: two of high affinity (TG (T/C) GGT) and one of low affinity (AGTGGT) (Supplementary Table 1). While, no Foxp3 binding sites (A/GTAAACAA) were found.
We then investigated the potential role of Runx1 in the regulation of Rspo3 gene expression, in the LM3 cell line, which was derived from a spontaneous BALB/c mouse mammary tumor [29]. LM3 cells can generate metastatic tumors when inoculated into syngeneic mice [30]. The LM3 cell line expresses detectable levels of Rspo3 mRNA (Supplementary Figure 1) and a transcriptionally active form of Runx1, which binds to the consensus sequence found in the Rspo3 promoter region (Figure 1A–1B and Figure 2B). In the gel shift assay the signal intensity decreases when cold oligonucleotide is included in the reaction (Figure 1B, 1C lane versus 32P lane) showing the specificity of the DNA-protein binding. Furthermore, when nuclear extracts were co-incubated with the labelled probe and an anti-Runx1 antibody, the intensity of the band decreased (Figure 1B, 1AB lane versus 32P lane), probably because the antibody interferes with Runx1 DNA binding domain. These results suggest that endogenous Runx1 is able to bind its putative binding site in the rspo3 promoter.
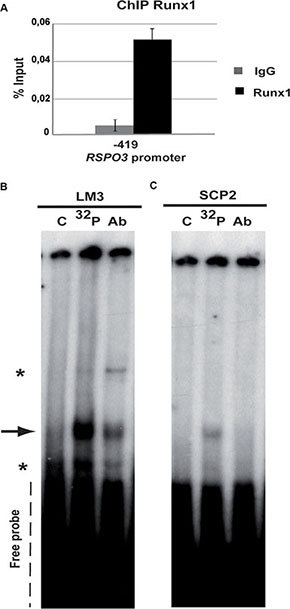
Figure 1: Runx1 binds to Rspo3 promoter. (A) ChIP assays were performed on LM3 cells using specific ChIP-grade Runx1 antibody or control IgG antibody. Specific primers were designed for targeting Runx1 high affinity binding site in the Rspo3’s promoter region. Bar graph shows mean and standard deviation (SD) of three independent experiments each of them performed by triplicate. Primers for Gapdh promoter region were used as negative control with no amplification product. (B–C) Gel shift assays were performed on LM3 (B) or SCp2 (C) nuclear extracts using an oligoprobe containing Runx1 consensus sequence included in the Rspo3 promoter region (–490bp) (lane 32P and lane Ab). This band showed lower intensity when cold oligonucleotides were included in the reaction (lane C). Asterisks in the Figure show unspecific binding. 32P: phospho-labeled oligoprobe, Ab: anti-Runx1 antibody and C unlabeled oligoprobe.
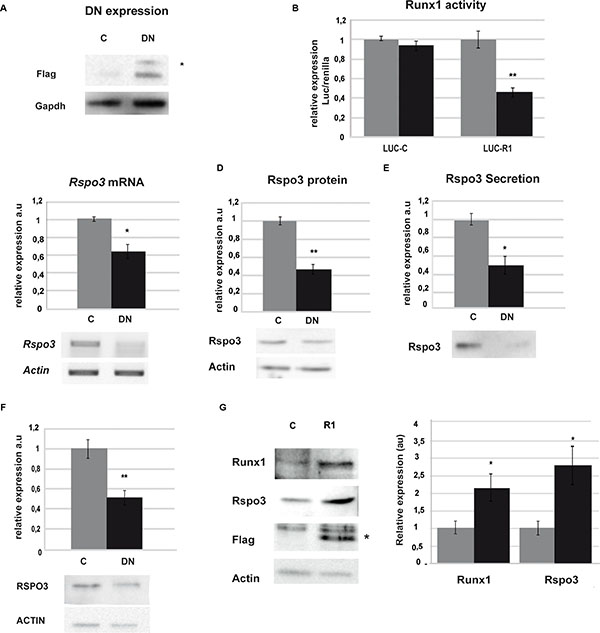
Figure 2: Runx1 regulates Rspo3 expression. (A–F) LM3 and MDA-MB-231 cells were transfected with DN or C. (A) Western blot (WB) showing tag flag expression in LM3 cells 48 h after transfection. Asterisk indicates specific band. (B) Luciferase assays on LM3 cells 48 h after co-transfection with DN or C and a reporter vector of Runx1 activity (Luc-R1) or empty vector, as control (Luc-C). Bar graph showing mean and SD of three independent experiments (p = 0,0052) (C–D) Rspo3 mRNA levels by semi-quantitative RT-PCR (C) and protein by WB (D) on LM3 cells 48 h after transfection. Bar graph showing mean and SD of three independent experiments (p = 0.014 and (p = 0,007). (E) Rspo3 protein secretion levels by WB of conditioned media generated by LM3 cells 48 h after transfection. Values were normalized to the number of cells after transfection. Bar graph showing mean and SD of three experiments (p = 0.021). (F) RSPO3 protein levels by WB analysis on MDA-MB-231 48 h after transfection. Bar graph showing mean and SD of three independent experiments (p = 0.0047). (G) SCP2 cells were transfected with a vector containing full Runx1 cDNA (R1) or empty vector as control (C) and WB analysis of Runx1 (first line), Rspo3 (second line), Flag tag (third line) and Actb (fourth line, used as loading control) were performed. Asterisk indicates specific band. Bar graph shows WB quantification, C in grey columns and R1 in black columns. Bar graph showing mean and SD of three independent experiments (p = 0,042).
To evaluate if the observed DNA/Runx1 interaction is biologically relevant for Rspo3 expression, we altered Runx1 expression levels in tumor and normal cell lines and evaluated Rspo3 expression and cell behaviour changes. Runx1 transcriptional activity was reduced by expression of the dominant-negative (DN) form of Runx1 in LM3 and MDA-DB-231 tumor cells [31]. We observed a significant reduction of Runx1 transcriptional activity in DN/Runx1 transfected tumor cells (Figure 2A and 2B), which resulted in a significant downregulation of Rspo3 expression and secretion (Figure 2C–2E: LM3 cell line and 2F: MDA-MB-231 cell line). On the other hand, we transfected SCp2 non-tumor epithelial mammary cells with an expression vector containing the full length cDNA sequence of Runx1 down-stream of a CMV-promoter [32]. Figure 2G shows that overexpression of Runx1 in these cells induced significant upregulation of Rspo3 expression. These experiments demonstrate that Runx1 is able to bind to Rspo3 promoter and triggers the expression of this oncogene in mammary epithelial cells.
Runx1 and Foxp3 physically interact in normal mammary epithelial cells
It has been previously shown that Foxp3 can interact with Runx1 and block its transcriptional activity in regulatory T cells [8]. Normal mammary epithelial cells express higher functional levels of Foxp3 compared to tumor cells [15] and comparable levels of endogenous Runx1 mRNA (Supplementary Figure 1A). To investigate if Foxp3 is able to modulate Runx1 transcriptional activity in normal mammary cells, we first explored whether endogenous Runx1 and Foxp3 proteins physically interact in these cells. To address this question we performed co-immunoprecipitation assays in which total protein extracts from SCp2 cells were incubated with anti-Foxp3 antibody and the precipitated proteins were immunoblotted with an anti-Runx1 antibody. Figure 3A shows a band of 50KDa corresponding to Runx1 molecular weight in the IP lane (co-immunoprecipitated proteins). Furthermore, Figure 3B shows that endogenous Runx1 and Foxp3 are concentrated in the nuclear compartments. In addition, partial co-localization of these two proteins in SCp2 cell nuclei is observed (Figure 3B). Therefore, these studies indicate that Runx1 and Foxp3 proteins physically interact in this mammary epithelial cell line.
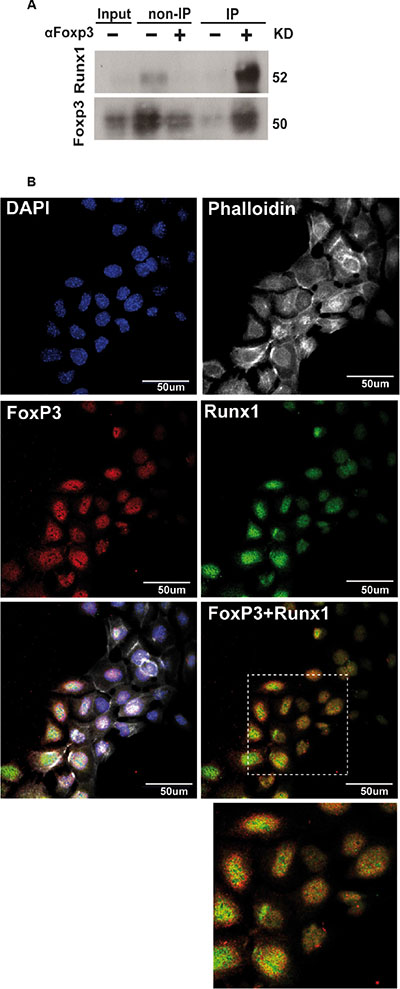
Figure 3: Runx1 physically interacts with Foxp3 in normal mammary epithelial cells. (A) Total protein extracts from SCp2 cells were incubated with anti-Foxp3 antibody and subsequent precipitation products were analyzed with anti-Runx1 antibody by WB analysis. IP (+) lane shows Runx1 co-immunoprecipitated with Foxp3. (B) Immunofluorescence of SCp2 cells shows subcellular localization of Runx1 (green) and Foxp3 (red) by confocal microscopy. Merge and inset figures show yellow dots representing co-localization of Runx1 and Foxp3 proteins. Magnification bar: 50 μm.
Foxp3 blocks Runx1 transcriptional activity in mammary epithelial cells
The SCp2 cell line showed visible mRNA levels of Foxp3, while Rspo3 endogenous expression was undetectable (Supplementary Figure 1A). Figure 1C also shows that in these cells, differently to what we have observed in LM3 cells, a lesser Runx1 binding to its consensus sequences. To study whether endogenous Foxp3 might be responsible for inhibition of Runx1 binding to the DNA, we evaluated Runx1 transcriptional activity after transfecting SCp2 cells with either Foxp3 (siF) or control (siC) siRNAs. Figure 4A and 4B show that Foxp3 is downregulated in SCp2 siF-transfected cells, while Runx1 expression levels are not modified (Figure 4A). Knockdown of Foxp3 in SCp2 cells led to a decrease in p21 expression (Figure 4A), a known Foxp3 target gene [33] and significantly prompted SCp2 proliferation, previously described in vivo in [33] (Figure 4D). Importantly, our results show that Foxp3 downregulation resulted in a significantly enhanced Runx1 transcriptional activity (Figure 4C) and Rspo3 expression induction (Figure 4E and 4F).
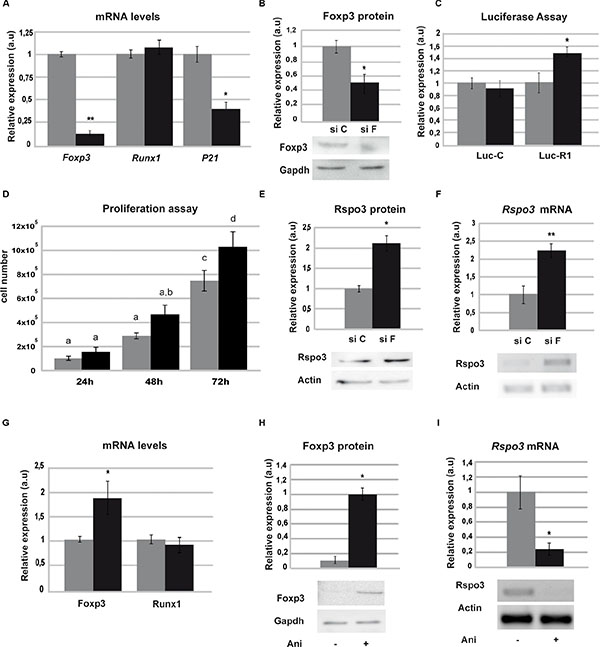
Figure 4: Foxp3 blocks Runx1 transcriptional activity. SCp2 cells were transfected with specific Foxp3 siRNA (siF) or scrambled control (siC) and mRNA or total protein extracts were analyzed. (A) Downregulation of Foxp3 levels (p = 0.004) (left), Runx1 (middle) and p21 (right) (p = 0.042) expression by RT-qPCR. (B) Downregulation of Foxp3 levels by WB (p = 0.013), Gapdh expression was used as loading control. (C) SCp2 cells were co-transfected with Luc-R1 or Luc-C and luciferase assay was performed (p = 0.028). (D) Proliferation assay was performed by counting cell number after siF and siC treatment (ANOVA p < 0.0001). Different letters means significant differences p < 0.05 by Bonferroni contrast. (E–F) Rspo3 expression by WB (p = 0.016) (E) and by PCR (p = 0.0059) (F) upon siF or siC treatment. Actd expression was used as loading control. Images were analyzed with image J software. In each case histogram shows mean and SD of three independent experiments, siC in grey bars and siF in black bars. (G–I) LM3 cells were incubated with Anisomycin and total protein or RNA extracts were analyzed 2 h after treatment. (G) RT-qPCR of Foxp3 (p = 0.04) (left) and Runx1 (right) were normalized to actin levels in Anisomycin treated (black bars) and control (gray bars) cells. (H) WB of Foxp3 with (black bars) or without (gray bars) Anisomycin and Gapdh was used as loading control (p = 0.022). (I) Rspo3 expression by RT-PCR. Histogram shows semi-quantification of the intensity of the band from three independent assays performed with vehicle (gray bars) or Anisomycin (black bars) (p = 0.012).
In addition, to determine whether induction of Foxp3 expression may result in inhibition of Runx1 target genes expression in mammary tumor cells (LM3 cells), which show low levels of Foxp3 (Supplementary Figure 1), were treated with low doses of anisomycin which is able to upregulate Foxp3 expression in mammary epithelial cell lines (Figure 4G) by c-Jun and ATF-2 activation [34]. Figure 4G–4I) shows that anisomycin treatment of LM3 cells caused a decrease in Rspo3 expression (4H–4I) without affecting the levels of Runx1 mRNA (4G).
Taken together, these experiments indicate that Foxp3 might be able to impair Runx1 transcriptional activity in normal mammary epithelial cells. In tumor cells Foxp3 low expression would explain high transcriptional activity of Runx1 on Rspo3 promoter. Furthermore, these data strongly suggest that Foxp3 and Runx1 protein interaction could determine Runx1 tumor promotion transcriptional activity in mammary epithelial cells. Moreover, our data propose a new tumor suppressor role for Foxp3 in mammary epithelial cells.
Impaired Runx1 activity reduces migration of breast tumor cells and upregulates GJA1 gene expression
As tumor metastasis is the main cause of morbidity and mortality, we explored the ability of Runx1 to promote tumor aggressiveness. To address this question we studied migration capacity of DN/Runx1 (DN) transfected tumor cells (LM3 and MDA-MB-231) by in vitro scrape-wound closure assays. Our results show that Runx1 inhibition resulted in a lower ability to fill the area by cell migration (Figure 5A and 5B). In addition, reduction in Runx1 transcriptional activity was also accompanied by a reduced number of cells (Figure 5C) but no changes in apoptosis levels (Figure 5D), indicating that Runx1 might be involved in cell cycle regulation without inducing cell death.
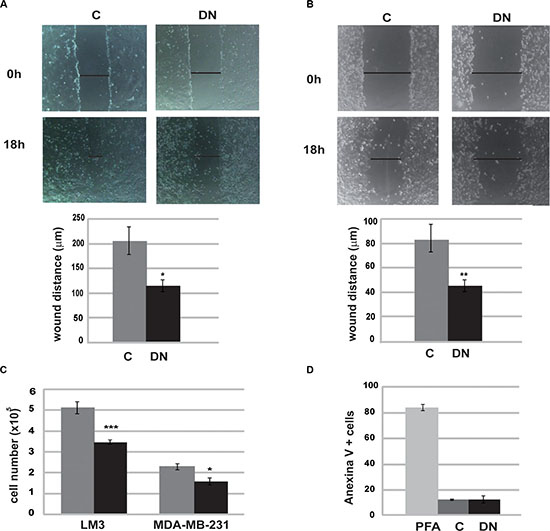
Figure 5: Runx1 is required for tumor cell migration. LM3 cells (A) and MDA-MB-231 cells (B) were subjected to wound healing assays for 18 h after transfection with DN or C expression vectors. Time points: 0 h and 18 h. A representative assay (up) and histograms (down) show mean and SD of three independent experiments (LM3 p = 0.02; MDA p = 0,0035). (C) LM3 and MDA-MB-231 cell number were assessed before and after transfection with DN or C expression vectors. Histogram shows mean and SD of three independent experiments (LM3 p = 0.0007; MDA p = 0,03) Axis X is 105 cells. (D) After transfection with DN or C expression vectors, LM3 cells were incubated with Annexin V-FITC and PI. Apoptosis was measure by flow citometry. PFA 4% was used as positive control.
Since Runx1 is a potent regulator of gene transcription in different cell types [1, 4, 5, 7, 35], we hypothesized that the effect observed when tumor cells expressed the DN/Runx1 might have a wider effect on genome expression of breast cancer cells. Therefore, we performed an in silico analysis that provided us a list of Runx1 probable target genes that were classified by their biological functions using Gene Ontology database. Only those previously described in breast cancer were selected (MS Recouvreux, unpublished data). This approach revealed that Runx1 is potentially able to regulate Gja1 (Supplementary Table 1) among other genes (MS Recouvreux et al. unpublished data). Interestingly, GJA1 has been described as a tumor-suppressor gene, which is able to reduce migration and proliferation of MCF7 and MDA-MB-231 breast cancer cells [24, 36]. Moreover, differential expression of GJA1 has been recently described as a potential positive prognostic marker for a clinically relevant stratification of breast cancer [23]. LM3 cells show lower levels of Gja1 mRNA compared with SCp2 cell line (Supplementary Figure 1B). By ChIP assays we demonstrate that Runx1 is able to bind GJA1 promoter region in MDA-MB-231 (Figure 6A). Moreover, DN/Runx1-transfected tumor cells showed a significant upregulation of GJA1 in this cell line demonstrating an inhibitory role of Runx1 on GJA1 promoter (Figure 6B). These results strongly suggest that Runx1 is able to inhibit GJA1 gene expression in human breast tumor cells. To show that FOXP3 is interfering with RUNX1 transcriptional inhibitory activity on GJA1 gene, we proceed to knockdown Foxp3 expression in SCp2 cells. To evaluate if Foxp3 is also able to interfere with Runx1 transcriptional inhibitory activity, we evaluated GJA1 expression in Foxp3 knockdown cells. Interestingly, reduced Foxp3 gene expression in SCp2 cell line showed a significant reduction of Gja1 gene expression compared with transfected control cells (Figure 6C). Therefore, we propose that GJA1 is a target of Runx1 transcriptional inhibitory activity, which is repressed by Foxp3 in mammary epithelial cells.
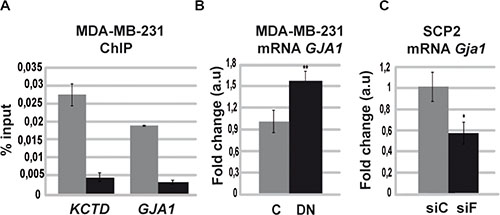
Figure 6: Runx1 downregulates GJA1 gene expression in the absence of Foxp3. (A) ChIP assays were performed on MDA-MB-231 cells using specific ChIP-grade RUNX1 antibody or control IgG. Primers targeting RUNX1 high affinity binding site included in the GJA1’s promoter region were designed. Histogram shows mean and SD of three independent experiments performed by triplicate. Primers for GAPDH promoter region were used as negative control with no amplification product. Primers for KCTD promoter region were used as positive control previously described in [58] (KCTD: Potassium Channel Tetramerization Domain). Anti-Runx1 antibody is in grey bars and Anti-IgG antibody is in black bars. (B) GJA1 mRNA levels by qPCR of MDA-MB-231 transfected cells with DN or C expression vector. Histograms represent mean and SD of three independent experiments (p = 0.004). (C) SCp2 cells were transfected with specific Foxp3 siRNA (siF) or scrambled control (siC) and GJA1 gene expression was measured by RT-qPCR. Histograms represent mean and SD of three independent experiments (p = 0.03).
Collectively, these observations indicate that a decreased in the availability of Foxp3 could cause a decisive increase in Runx1 transcriptional activity, which may alter migration abilities of tumor cells by binding and regulating it’s downstream target genes such as Rspo3 and Gja1. These data strongly suggest that Runx1/Foxp3 interaction could be a new mechanism of gene expression regulation during breast cancer development.
DISCUSSION
In this study we show for the first time that endogenous Runx1 and Foxp3 physically interact in mammary epithelial cells leading to the inhibition of Runx1 transcriptional activity. Furthermore, we demonstrate that Runx1 is able to actively promote oncogene Rspo3 and prevent GJA1 gene expression in mammary tumor cell lines.
Although Runx1 participation has been widely described in the hematopoietic system and leukemia, during the last years different studies have indicated that this transcription factor could be also involved in other cancer types [11]. Particularly, Runx1 has been identified as a significant controller of tumorigenesis in various epithelial cancers [4, 5, 37]; however, its contribution in breast cancer development is still under debate [9]. Experimentally, Ras-mediated transformation of mammary epithelial cells (MCF10A) has been shown to be associated with loss of Runx1 expression [10]. Other report by Wang and colleagues indicated that RUNX1 knockdown causes hyperproliferation and abnormal morphogenesis in MCF10A-5E cell clone [12]. These authors also show that RUNX1 downregulation is associated with compensatory upregulation of FOXO1, since RUNX1-depleted cells require normal FOXO function for proliferation [12]. Our work shows by lost-of-function assays that Runx1 is able to modulate mammary gene expression towards a cancer behavior fate. Our data indicates that reduced Runx1 transcriptional activity downregulates Rspo3 oncogene expression, upregulates GJA1 tumor suppressor gene expression, significantly delays tumor cell migration and wanes cell number, in a susceptible manner to Foxp3’s stalling activity. These results suggest that Runx1 could be acting as a tumor enhancer depending on Foxp3 availability. The difference observed between our study and the ones described above could be explained by different genetic background and/or because of the strategy used. While Wang and colleagues abolished RUNX1 expression in an immortalized cell line (MCF10-A-5E) [12], we reduced RUNX1 transcriptional activity in tumor cell lines (LM3 and MDA-DB-231) and upregulated its endogenous functional availability in normal epithelial cell line (SCp2) by RUNX1 overexpression or Foxp3 knockdown. It would be interesting to unravel in which steps of tumor induction and/or promotion is Runx1 and Foxp3 interplay defining cell fate.
Other authors also described this controversy analyzing RUNX1 expression by different techniques in human breast tumor samples [13, 38, 39]. Ferrari and colleagues performed the first comprehensive characterization of RUNX1 protein expression in breast cancer by immunohistochemistry using a large cohort of human samples [13]. They showed that high Runx1 protein expression correlated with poor prognosis in triple negative breast cancer patients [13]. On the other hand, large-scale sequencing studies in human breast cancer revealed frequent RUNX1 mutations and deletions [38, 39]. This observation could be explained by the consideration that if Runx1 is used as a prognostic marker should be assessed not only by gene or protein expression levels but also by its functional activity. In addition to these considerations that might modulate Runx1 activity in the nucleus here we show new evidence that RUNX1 might be sequestered by Foxp3 blocking its activity as a transcriptional regulator. Several studies on leukemia have already shown that cell context and cellular dosage are relevant for RUNX1 activity [6, 40, 41]. In line with this and due to the multiple roles that RUNX1 displays, inducing (as for Rspo3) or repressing (as for GJA1) gene expression, probably depending on the interaction with other regulatory proteins Foxp3 expression/localization could be additionally used as a collaborative marker for prognosis [6]. The hypothesis that RUNX1 could affect different steps in tumor promotion has also been described in skin cancer where it was found that RUNX1 is important for both, tumor initiation and progression, while it had been previously shown that RUNX1 was not essential for adult tissue maintenance [42].
Here we provide evidence that Runx1 could be relevant for breast cancer cell survival and migration, supporting the idea that Runx1 could act as a tumor enhancer. This phenomenon has previously been described in ovarian [4] and skin cancer [5], among others epithelial neoplasias [3, 35]. In addition a relevant role of Runx1 in physiological hematopoietic cell migration and adhesion has been reported [43]. Interestingly, it has been shown that Runx1 depletion reduced prostate stem cell number without significant differences in apoptotic and necrotic cell death [44]. During the revision process of this manuscript Browne et al. has published experimental data that support part of our results [45].
We also demonstrate that GJA1 mRNA levels are significantly upregulated when Runx1 transcriptional activity is reduced. The participation of GJAs in cancer has been studied in vivo and in vitro and results strongly suggest that these proteins are less expressed during tumor promotion enhancing migration and proliferation [46–49]. Moreover, Teleki and colleagues suggest that GJA1 and GJA6 could be used as potential positive and negative prognostic markers, respectively, for a clinically relevant stratification of breast cancers [23]. Furthermore, during the writing of this paper Giuliano and colleagues showed, using circulating and disseminated tumor cells from breast cancer patient-derived xenograft-bearing mice, that high GJA1 expression could be used to predict distant metastasis-free survival [50].
The results showed herein indicate that endogenous Foxp3 is able to physically interact with endogenous Runx1 in mammary epithelial cells, modulating its transcriptional activity. Ono and colleagues describe for the first time a physical interaction of Runx1/Foxp3 in normal regulatory T cells [8]. Our data show that downregulation of Foxp3 expression in mammary epithelial cells induces Runx1 transcriptional activity, which then upregulates RSPO3 and reduces GJA1 gene expression without increasing RUNX1 expression. Likewise, we demonstrate that upregulation of Foxp3 expression in mammary tumor cells reduces Rspo3 gene expression. It has been proposed that FOXP3 participates in breast cancer development as a tumor suppressor factor directly bound to its oncogene and miR-146a promoter targets [15, 19]. Our in silico analyses revealed an absence of Foxp3 binding site in RSPO3 and GJA1 promoter regions. It has been shown that prognostic value of FOXP3 in breast cancer depends on localization of this protein as well as HER2 and ER expression [17]. Zuo and colleagues showed a high proportion of somatic mutations and deletions in FOXP3 gene in human breast cancer cells, which may include nuclear localization signals [15]. In agreement with our experimental data, accumulating evidence point out that FOXP3 activity is coordinated with multiple transcriptional regulators and Foxp3 localization may also depend on its molecular partners [51, 52]. Taken together these evidences suggest that Foxp3 localization could also define Runx1 activity in mammary epithelial cells [18]. Furthermore, our data strongly suggests this interaction as a new tumor suppressor mechanism used by Foxp3 to reduce breast tumor promotion.
These results strongly suggest that Runx1/Foxp3 physical interaction in mammary epithelial cells is a relevant mechanism to regulate gene expression that controls cell fate. In vivo experiments will allow us to test this hypothesis evaluating tumor initiation, stemness and metastatic models considering tumor microenvironment. Finally, facilitate Runx1/Foxp3 interaction could be use to improve anti-tumor strategies in breast cancer therapy.
MATERIALS AND METHODS
Cell lines
LM3 is a mouse mammary cancer cell line derived from a spontaneous malign adenocarcinoma from a BALB/c strain in the animal care area of Oncology Institute Angel H. Roffo (Argentina). This cell line was kindly given by Elisa Bal De Kier, Oncology Institute Angel H. Roffo. [29, 30, 53, 54]. Cells were grown in MEM (Gibco). SCp2 is a mouse mammary immortalized cell line kindly given by Dr. Mina Bissell, Lawrence Berkeley National Laboratory, Berkeley, USA [32]. Cells were routinely grown in DMEM/F12 (Gibco) supplemented with 2% FBS, 5 μg/ml insulin. MDA-MB-231 [55] (ATCC® HTB- 26™) is a human breast cancer cell line routinely grown in RPMI (Gibco). All cell line were grown with 10% FBS (Internegocios SA), unless specified, 100 U/ml of penicillin and 100 μg/ml of streptomycin. Cells were kept at low passages and store at liquid nitrogen.
In silico promoter analysis
We used database sites available on line (http://www.genomatix.de/ and http://www.cbrc.jp/research/db/TFSEARCH.html) to determine the presence of putative binding sites for the transcription factor Runx1 (TGTGGT of high affinity) at the promoter region of Rspo3 and Gja1 promoter region (–5000 bp up stream from +1 transcription site, Supplementary Table 1). No Foxp3 binding sites (consensus sequence: A/GTAAACAA) were found.
Transient transfection assays and treatments
For transient transfections, 3 × 105 cells were plated in medium without antibiotics and 24 h later were transfected with different vectors or siRNA as indicated: pCDNA3-DN: vector containing a dominant negative version of Runx1 (DN) subcloned from pMSCV2.2 using Bgl II and EcoR I enzymes; pCDNA3: control empty vector (C); pGL3-RORyT: a reporter plasmid containing the promoter of RORyT with putative sites for Runx1 upstream luciferase gene (Luc-R1) kindly given by V Lazarevic [56] (NIH, USA); pGL3: control vector without Runx1 binding sites (Luc-C); siRNA and Stealth RNAi™ siRNA Duplex oligoribonucleotide (Invitrogen) Foxp3: oligonucleotide containing a specific sequence to block Foxp3 transcription (siF); siRNA Control: oligonucleotide with scrambled sequence (siC) (sequences upon request). We used β-Gal vector as transfection control and TK-renilla vector for luciferase assays. Transfection assays were performed using Lipofectamine 2000 (Invitrogen) and OptiMEM (Life Technologies). After 48 h cells were harvested in lysis buffer. Luciferase levels were normalized to renilla expression measured with Promega luciferase/renilla kit according to the manufacturer protocol. β-Gal activity was measured by absorbance at 515 nm in spectrophotometer (BIORAD). For proliferation assay cells were harvested at 24, 48 or 72 hours after siRNA transfection. Cells were stained with bromophenol blue for viability and counted in Neubauer chamber.
Dominant negative version of Runx1 (DN) was generated introducing a stop codon at 826 cDNA sequence position included in the transactivation domain. This results in early translation termination and a short version of Runx1 behaving as a dominant negative over Runx1 during homodimerization.
For Foxp3 overexpression, LM3 cells were incubated with 0.05 μg/ul of Anisomycin (SIGMA) or vehicle (ethanol) as previously described [34].
Immunofluorescence and confocal microscopy
SCp2 cells were fixed with 4% paraformaldehyde for 15 minutes at room temperature (RT), washed in PBS-tween 0.05% (PBS-T). Cells were permeabilized with 0.1% of Triton X-100 for 10 minutes at RT, and blocked with 2% BSA in PBS-T (blocking solution) for 1 h. After blocking, cells were incubated for 1 h at RT with anti-FOXP3 (SC-28705) or anti-Runx1 antibodies (SC-8564) in blocking solution. Cells were washed in PBS and incubated for 1 h at RT with anti goat-Cy5 (Santa Cruz) and anti rabbit-Cy3 (Santa Cruz), Dapi and phalloidin (Life Technologies) in blocking solution. After washes in PBS, coverslips were mounted with gelvatol. Images were taken at LSM510 Meta confocal microscope (Carl Zeiss).
Co-immunoprecipitation assay
For immunoprecipitation SCp2 were washed, lysed, and pre-cleared with 20 μl A/G plus Agarose (Agarose, Santa Cruz) at 4ºC for 30 minutes, and centrifuged at 1000 g for 5 minutes. Supernatant was incubated with or without 1 μl of Foxp3 primary antibody (SC-28705) and 20 μl of Agarose at 4ºC. Immunoprecipitates were pelleted by centrifugation (1000 g, 5 minutes, 4ºC), washed with PBS, and prepared for western blot (WB).
Chromatin immunoprecipitation assays
LM3 and MDA-MD-231 cell lines were used for chromatin immunoprecipitation (ChIP) assays. ChIP were performed as previously described [57] using anti-RUNX1 (Abcam 23980) and anti-IgG (Abcam 46540, negative control). Quantification of ChIP was performed by real-time polymerase chain reaction (qPCR) using Stratagene Mx3000P™ instrument. The fold enrichment of target sequence in the immunoprecipitated compared with input fractions was calculated using the comparative Ct (the number of cycles required to reach a threshold concentration) method with the equation 2(DCt (ref)-DCt (t)). Each of these values were corrected by mouse GAPDH gene and referred as relative abundance over time zero. Primers sequences are available on request.
Protein analysis
Total proteins were extracted from mammary tissue in lysis buffer supplemented with protease cocktail set I (Calbiochem). Proteins were resolved in 10 or 12% polyacrylamide gel, and transferred to a poly-vinylidine fluoride membrane. Primary antibodies: β-actin (SC-1616-R), Foxp3 (Abcam 14551), Runx1 (Abcam 61753), RSPO3 (R & D MAB41201), GAPDH (Abcam 9483), FLAG (Sigma F3165). Secondary antibodies: rabbit IgG (SC-2004), mouse IgG (SC-2005), rat IgG (SC-2006). The immune reactive protein bands were detected using chemiluminescence system (ECL_Plus system; GE) and the Fuji Film Image Reader (LAS-1000 Fuji Film). Densitometric analysis of protein levels was performed with ImageJ 1.34s software (Wayne Rasband, NIH). In each case obtained values were normalized as indicated in each experiment.
RNA analysis
RNA was extracted from cellular lysates with Trizol reagent (Invitrogen) according to the manufacturer’s instructions. For reverse transcription, 1 μg of total RNA was used. Retrotranscription was performed at 37ºC for 60 minutes followed by 15 minutes at 72ºC. For the quantitative real-time RT-PCR (qRT-PCR) 1 μl cDNA was used. All reactions were placed at Stratagene Mx3000P™ Instrument. Primers are available upon request. Reactions were run for 35 cycles under the following conditions: 15 seconds at 94ºC, 20 seconds at 58ºC–65ºC (according to the primers), and 20 seconds at 72ºC. The amplification of unique products in each reaction was verified by melting curve. Expression level of each gene was normalized to actin expression level using delta Ct method and specific primers. Means and standard deviation (SD) from at least three experiments were calculated and shown as fold changes with respect to the control. RSPO3 gene expression was analyzed by RT-PCR using MJ Research MiniCycler™ or Lambeth Multigene getting the same results on both machines. PCR products were visualized on 2% gel with ethidium bromide. Rspo3 expected band is 400 bp and 200 bp for Actin.
In vitro wound-healing repair assays
Cells were grown in normal growth media to monolayer confluence. Scratches were performed with a tip, and cell debris was removed by washing with PBS. Width of each wounded area was measured using grids at three marked positions photographed (40x magnification) at the indicated times. Cell migration was quantified as the distance covered by cells in wound-healed surface from the marked positions. Results are expressed as mean ± SD of three independent experiments.
Apoptosis assays
After 48 h of Lipofectamine transfection with Runx1 dominant negative (DN), control (C) expression vectors used or after 24 h incubation with 4% PFA, cells were incubated with Annexin V-FITC and Propidium Iodide (BD) for 10 minutes on ice. Apoptosis was analyzed by flow cytometer (BD FACS-Aria II). Data were processed using FlowJo software (Tree Star). Percentage of early plus late apoptotic cells was calculated as the ratio between Annexin-V FITC/PI- stained cells and total number of cells.
Statistical analysis
Results were expressed as mean with SD as indicated in Figure legends. Differences were regarded as significant at p < 0.05. Statistical analyses were performed by Infostat software (Di Rienzo J.A., et al. InfoStat versión 2012. Grupo InfoStat, FCA, Universidad Nacional de Córdoba, Argentina).
ACKNOWLEDGMENTS
The authors would like to thanks Elisa Bal De Kier (Oncology Institute Angel H. Roffo) for providing LM3 cell line Mina Bissell (Lawrence Berkeley National Laboratory, Berkeley, USA) for providing SCp2 cell line, Vanja Lazarevic for providing pGL3-RORyT construction (Bethesda MD, NIH, USA); Omar A Coso (University of Buenos Aires) for providing some of the antibodies used for confocal microscopy, Adali Pecci for anisomycin supply, Victoria Slomiansky for confocal microscopy assistance, Paula Barrionuevo for FACS assistance, and Meera Saxena (Unibasel, Basel, Swiss) for English manuscript corrections.
Author contributions
MSR, ENG, ECK and NR contributed to experimental design. MSR (Protein analisys, RNA analisys, proliferation assays, apoptosis assays and statistical analisys), EG (Co-IP, transfection experiments, in silico analysis, EMSA), LRV (ChIP), PNE (in silico analysis), CSL (migration assays) were responsible for performing experiments; NR, SR, EG, LC, ECK, JMT and PNE contributed to data analysis. LC contributed with DN/Runx1 and overexpression Runx1 vectors and result discussion. SR, ECK and NR contributed to manuscript preparation. SR and NR designed the figures. NR had the original hypothesis, wrote the manuscript, participated in study design and coordination. All authors have read and approved the final manuscript.
Abbreviations
ChIP, chromatin immunoprecipitation; DN, dominant-negative form of Runx1; EMSA, Electromobility shift assay; GJA1, Gap Junction protein Alpha 1; IP, immunoprecipitated; RSPO3, R spondin 3; siRNA, small interfering RNA; siF, siRNA for Foxp3; siC, siRNA control; SKP2, S-phase kinase-associated protein-2.
FUNDINGS
This work was supported by research grants awarded to NR from: Susan Komen For the Cure (PDF0707437), from Agencia Nacional de Promociones Científicas y Técnicas (BID-PICT2007–00740), from Roemmers Foundation for young Researchers (2010–2011), from University of Buenos Aires (UBACYT-20020090300023) and from CONICET (PIP 2012–2014).
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
REFERENCES
1. Chuang LS, Ito K, Ito Y. RUNX family: Regulation and diversification of roles through interacting proteins. International journal of cancer. 2013; 132:1260–1271.
2. Ito Y, Bae SC, Chuang LS. The RUNX family: developmental regulators in cancer. Nature reviews Cancer. 2015; 15:81–95.
3. Scheitz CJ, Lee TS, McDermitt DJ, Tumbar T. Defining a tissue stem cell-driven Runx1/Stat3 signalling axis in epithelial cancer. The EMBO journal. 2012; 31:4124–4139.
4. Keita M, Bachvarova M, Morin C, Plante M, Gregoire J, Renaud MC, Sebastianelli A, Trinh XB, Bachvarov D. The RUNX1 transcription factor is expressed in serous epithelial ovarian carcinoma and contributes to cell proliferation, migration and invasion. Cell cycle. 2013; 12:972–986.
5. Hoi CS, Lee SE, Lu SY, McDermitt DJ, Osorio KM, Piskun CM, Peters RM, Paus R, Tumbar T. Runx1 directly promotes proliferation of hair follicle stem cells and epithelial tumor formation in mouse skin. Molecular and cellular biology. 2010; 30:2518–2536.
6. Blyth K, Cameron ER, Neil JC. The RUNX genes: gain or loss of function in cancer. Nature reviews Cancer. 2005; 5:376–387.
7. Friedman AD. Cell cycle and developmental control of hematopoiesis by Runx1. Journal of cellular physiology. 2009; 219:520–524.
8. Ono M, Yaguchi H, Ohkura N, Kitabayashi I, Nagamura Y, Nomura T, Miyachi Y, Tsukada T, Sakaguchi S. Foxp3 controls regulatory T-cell function by interacting with AML1/Runx1. Nature. 2007; 446:685–689.
9. Janes KA. RUNX1 and its understudied role in breast cancer. Cell cycle. 2011; 10:3461–3465.
10. Kadota M, Yang HH, Gomez B, Sato M, Clifford RJ, Meerzaman D, Dunn BK, Wakefield LM, Lee MP. Delineating genetic alterations for tumor progression in the MCF10A series of breast cancer cell lines. PloS one. 2010; 5:e9201.
11. Taniuchi I, Osato M, Ito Y. Runx1: no longer just for leukemia. The EMBO journal. 2012; 31:4098–4099.
12. Wang L, Brugge JS, Janes KA. Intersection of FOXO- and RUNX1-mediated gene expression programs in single breast epithelial cells during morphogenesis and tumor progression. Proceedings of the National Academy of Sciences of the United States of America. 2011; 108:E803–812.
13. Ferrari N, Mohammed ZM, Nixon C, Mason SM, Mallon E, McMillan DC, Morris JS, Cameron ER, Edwards J, Blyth K. Expression of RUNX1 Correlates with Poor Patient Prognosis in Triple Negative Breast Cancer. PloS one. 2014; 9:e100759.
14. Douglass S, Ali S, Meeson AP, Browell D, Kirby JA. The role of FOXP3 in the development and metastatic spread of breast cancer. Cancer metastasis reviews. 2012; 31:843–854.
15. Zuo T, Wang L, Morrison C, Chang X, Zhang H, Li W, Liu Y, Wang Y, Liu X, Chan MW, Liu JQ, Love R, Liu CG, et al. FOXP3 is an X-linked breast cancer suppressor gene and an important repressor of the HER-2/ErbB2 oncogene. Cell. 2007; 129:1275–1286.
16. Droeser RA, Obermann EC, Wolf AM, Wallner S, Wolf D, Tzankov A. Negligible nuclear FOXP3 expression in breast cancer epithelial cells compared with FOXP3-positive T cells. Clinical breast cancer. 2013; 13:264–270.
17. Takenaka M, Seki N, Toh U, Hattori S, Kawahara A, Yamaguchi T, Koura K, Takahashi R, Otsuka H, Takahashi H, Iwakuma N, Nakagawa S, Fujii T, et al. FOXP3 expression in tumor cells and tumor-infiltrating lymphocytes is associated with breast cancer prognosis. Molecular and clinical oncology. 2013; 1:625–632.
18. Merlo A, Casalini P, Carcangiu ML, Malventano C, Triulzi T, Menard S, Tagliabue E, Balsari A. FOXP3 expression and overall survival in breast cancer. Journal of clinical oncology. 2009; 27:1746–1752.
19. Liu R, Liu C, Chen D, Yang WH, Liu X, Liu CG, Dugas CM, Tang F, Zheng P, Liu Y, Wang L. FOXP3 controls an miR-146/NFkappaB negative feedback loop that inhibits apoptosis in breast cancer cells. Cancer research. 2015.
20. Zhang C, Xu Y, Hao Q, Wang S, Li H, Li J, Gao Y, Li M, Li W, Xue X, Wu S, Zhang Y, Zhang W. FOXP3 suppresses breast cancer metastasis through down-regulation of CD44. International journal of cancer. 2015; 137:1279–1290. doi: 10.1002/ijc.29482.
21. Gattelli A, Zimberlin MN, Meiss RP, Castilla LH, Kordon EC. Selection of early-occurring mutations dictates hormone-independent progression in mouse mammary tumor lines. Journal of virology. 2006; 80:11409–11415.
22. Theodorou V, Kimm MA, Boer M, Wessels L, Theelen W, Jonkers J, Hilkens J. MMTV insertional mutagenesis identifies genes, gene families and pathways involved in mammary cancer. Nature genetics. 2007; 39:759–769.
23. Teleki I, Szasz AM, Maros ME, Gyorffy B, Kulka J, Meggyeshazi N, Kiszner G, Balla P, Samu A, Krenacs T. Correlations of differentially expressed gap junction connexins Cx26, Cx30, Cx32, Cx43 and Cx46 with breast cancer progression and prognosis. PloS one. 2014; 9:e112541.
24. Talhouk RS, Fares MB, Rahme GJ, Hariri HH, Rayess T, Dbouk HA, Bazzoun D, Al-Labban D, El-Sabban ME. Context dependent reversion of tumor phenotype by connexin-43 expression in MDA-MB231 cells and MCF-7 cells: role of beta-catenin/connexin43 association. Experimental cell research. 2013; 319:3065–3080.
25. Ohkawara B, Glinka A, Niehrs C. Rspo3 binds syndecan 4 and induces Wnt/PCP signaling via clathrin-mediated endocytosis to promote morphogenesis. Developmental cell. 2011; 20:303–314.
26. Hao HX, Xie Y, Zhang Y, Charlat O, Oster E, Avello M, Lei H, Mickanin C, Liu D, Ruffner H, Mao X, Ma Q, Zamponi R, et al. ZNRF3 promotes Wnt receptor turnover in an R-spondin-sensitive manner. Nature. 2012; 485:195–200.
27. Aoki M, Mieda M, Ikeda T, Hamada Y, Nakamura H, Okamoto H. R-spondin3 is required for mouse placental development. Developmental biology. 2007; 301:218–226.
28. Kazanskaya O, Ohkawara B, Heroult M, Wu W, Maltry N, Augustin HG, Niehrs C. The Wnt signaling regulator R-spondin 3 promotes angioblast and vascular development. Development. 2008; 135:3655–3664.
29. Urtreger A, Ladeda V, Puricelli L, Rivelli A, Vidal M, Delustig E, Joffe E. Modulation of fibronectin expression and proteolytic activity associated with the invasive and metastatic phenotype in two new murine mammary tumor cell lines. International journal of oncology. 1997; 11:489–496.
30. Quaglino A, Schere-Levy C, Romorini L, Meiss RP, Kordon EC. Mouse mammary tumors display Stat3 activation dependent on leukemia inhibitory factor signaling. Breast cancer research. 2007; 9:R69.
31. Li D, Sinha KK, Hay MA, Rinaldi CR, Saunthararajah Y, Nucifora G. RUNX1-RUNX1 homodimerization modulates RUNX1 activity and function. The Journal of biological chemistry. 2007; 282:13542–13551.
32. Desprez PY, Hara E, Bissell MJ, Campisi J. Suppression of mammary epithelial cell differentiation by the helix-loop-helix protein Id-1. Molecular and cellular biology. 1995; 15:3398–3404.
33. Liu R, Wang L, Chen G, Katoh H, Chen C, Liu Y, Zheng P. FOXP3 up-regulates p21 expression by site-specific inhibition of histone deacetylase 2/histone deacetylase 4 association to the locus. Cancer research. 2009; 69:2252–2259.
34. Liu Y, Wang Y, Li W, Zheng P, Liu Y. Activating transcription factor 2 and c-Jun-mediated induction of FoxP3 for experimental therapy of mammary tumor in the mouse. Cancer research. 2009; 69:5954–5960.
35. Planaguma J, Diaz-Fuertes M, Gil-Moreno A, Abal M, Monge M, Garcia A, Baro T, Thomson TM, Xercavins J, Alameda F, Reventos J. A differential gene expression profile reveals overexpression of RUNX1/AML1 in invasive endometrioid carcinoma. Cancer research. 2004; 64:8846–8853.
36. McLachlan E, Shao Q, Wang HL, Langlois S, Laird DW. Connexins act as tumor suppressors in three-dimensional mammary cell organoids by regulating differentiation and angiogenesis. Cancer research. 2006; 66:9886–9894.
37. Inoue K, Ito Y. Neuroblastoma cell proliferation is sensitive to changes in levels of RUNX1 and RUNX3 protein. Gene. 2011; 487:151–155.
38. Banerji S, Cibulskis K, Rangel-Escareno C, Brown KK, Carter SL, Frederick AM, Lawrence MS, Sivachenko AY, Sougnez C, Zou L, Cortes ML, Fernandez-Lopez JC, Peng S, et al. Sequence analysis of mutations and translocations across breast cancer subtypes. Nature. 2012; 486:405–409.
39. Ellis MJ, Ding L, Shen D, Luo J, Suman VJ, Wallis JW, Van Tine BA, Hoog J, Goiffon RJ, Goldstein TC, Ng S, Lin L, Crowder R, et al. Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature. 2012; 486:353–360.
40. Goyama S, Huang G, Kurokawa M, Mulloy JC. Posttranslational modifications of RUNX1 as potential anticancer targets. Oncogene. 2014; 0.
41. Goyama S, Schibler J, Cunningham L, Zhang Y, Rao Y, Nishimoto N, Nakagawa M, Olsson A, Wunderlich M, Link KA, Mizukawa B, Grimes HL, Kurokawa M, et al. Transcription factor RUNX1 promotes survival of acute myeloid leukemia cells. The Journal of clinical investigation. 2013; 123:3876–3888.
42. Osorio KM, Lee SE, McDermitt DJ, Waghmare SK, Zhang YV, Woo HN, Tumbar T. Runx1 modulates developmental, but not injury-driven, hair follicle stem cell activation. Development. 2008; 135:1059–1068.
43. Lie ALM, Marinopoulou E, Li Y, Patel R, Stefanska M, Bonifer C, Miller C, Kouskoff V, Lacaud G. RUNX1 positively regulates a cell adhesion and migration program in murine hemogenic endothelium prior to blood emergence. Blood. 2014; 124:e11–20.
44. Kim W, Barron DA, San Martin R, Chan KS, Tran LL, Yang F, Ressler SJ, Rowley DR. RUNX1 is essential for mesenchymal stem cell proliferation and myofibroblast differentiation. Proceedings of the National Academy of Sciences of the United States of America. 2014; 111:16389–16394.
45. Browne G, Taipaleenmaki H, Bishop NM, Madasu SC, Shaw LM, van Wijnen AJ, Stein JL, Stein GS, Lian JB. Runx1 is associated with breast cancer progression in MMTV-PyMT transgenic mice and its depletion in vitro inhibits migration and invasion. Journal of cellular physiology. 2015; 230:2522–2532.
46. Shao Q, Wang H, McLachlan E, Veitch GI, Laird DW. Down-regulation of Cx43 by retroviral delivery of small interfering RNA promotes an aggressive breast cancer cell phenotype. Cancer research. 2005; 65:2705–2711.
47. Simpson KJ, Selfors LM, Bui J, Reynolds A, Leake D, Khvorova A, Brugge JS. Identification of genes that regulate epithelial cell migration using an siRNA screening approach. Nature cell biology. 2008; 10:1027–1038.
48. Naus CC, Laird DW. Implications and challenges of connexin connections to cancer. Nature reviews Cancer. 2010; 10:435–441.
49. Plante I, Stewart MK, Barr K, Allan AL, Laird DW. Cx43 suppresses mammary tumor metastasis to the lung in a Cx43 mutant mouse model of human disease. Oncogene. 2011; 30:1681–1692.
50. Giuliano M, Herrera S, Christiny P, Shaw C, Creighton CJ, Mitchell T, Bhat R, Zhang X, Mao S, Dobrolecki LE, Al-Rawi A, Chen F, Veneziani BM, et al. Circulating and disseminated tumor cells from breast cancer patient-derived xenograft-bearing mice as a novel model to study metastasis. Breast cancer research. 2015; 17:3.
51. Lam EW, Brosens JJ, Gomes AR, Koo CY. Forkhead box proteins: tuning forks for transcriptional harmony. Nature reviews Cancer. 2013; 13:482–495.
52. Zhou Z, Song X, Li B, Greene MI. FOXP3 and its partners: structural and biochemical insights into the regulation of FOXP3 activity. Immunologic research. 2008; 42:19–28.
53. Puricelli L, Colombo LL, Bal de Kier Joffe E, de Lustig ES. Invasiveness in vitro of two mammary adenocarcinoma tumors with different metastasizing ability. Invasion & metastasis. 1984; 4:238–246.
54. Hoijman E, Rocha-Viegas L, Kalko SG, Rubinstein N, Morales-Ruiz M, Joffe EB, Kordon EC, Pecci A. Glucocorticoid alternative effects on proliferating and differentiated mammary epithelium are associated to opposite regulation of cell-cycle inhibitor expression. Journal of cellular physiology. 2012; 227:1721–1730.
55. Cruciger QV, Pathak S, Cailleau R. Human breast carcinomas: marker chromosomes involving 1q in seven cases. Cytogenetics and cell genetics. 1976; 17:231–235.
56. Lazarevic V, Chen X, Shim JH, Hwang ES, Jang E, Bolm AN, Oukka M, Kuchroo VK, Glimcher LH. T-bet represses T (H)17 differentiation by preventing Runx1-mediated activation of the gene encoding RORgammat. Nature immunology. 2011; 12:96–104.
57. Rocha-Viegas L, Villa R, Gutierrez A, Iriondo O, Shiekhattar R, Di Croce L. Role of UTX in RAR-mediated Gene Regulation in Leukemia. Molecular and cellular biology. 2014.
58. Stender JD, Kim K, Charn TH, Komm B, Chang KC, Kraus WL, Benner C, Glass CK, Katzenellenbogen BS. Genome-wide analysis of estrogen receptor alpha DNA binding and tethering mechanisms identifies Runx1 as a novel tethering factor in receptor-mediated transcriptional activation. Molecular and cellular biology. 2010; 30:3943–3955.

