INTRODUCTION
Nasopharyngeal carcinoma (NPC) is a highly invasive and metastatic cancer that is widely prevalent in southern China and Southeast Asia. Although the overall survival rate is approximately 90% in patients with early stage disease, most patients are diagnosed with advanced disease at their first clinic visit, and the survival rate for these patients is less than 50% [1]. Developing a simple and reliable method for diagnosing and monitoring NPC is of great importance for improving patient outcomes [2].
The infection of epithelial cells by Epstein-Barr virus (EBV) is a typical characteristic of NPC cases in high-risk areas [3, 4]. Therefore, substantial effort has been made to develop novel EBV detection approaches utilizing EBV-related markers. Methods for detecting antibody titers against EBV serum antigens, including viral capsid antigen (VCA) and early antigen (EA), and for detecting EBV DNA load in plasma have been developed to assist in NPC diagnosis [5–8]. However, the results of antibody titer tests alone have proven insufficient to accurately diagnose NPC [5]. Additionally, EBV DNA tests appear to be of limited value for diagnosing NPC patients with early stage disease and local recurrence [8].
As an alternative to the more-invasive biopsy method, tumor tissue samples can also be obtained directly from the nasopharynx using a nasopharyngeal (NP) brush or swab. Samples from NP brushes or swabs are considered to directly reflect the local tumor load and disease process. EBV DNA is a popular biomarker [9–16] based on the hypothesis that normal NP tissue is negative for EBV [17]. However, evidence from endemic areas suggests that EBV DNA is also present in samples from normal NP tissue [12, 14, 15]. Our recent study showed that EBV DNA was detectable in 70.6% of NP brush samples from non-NPC controls and in 87.8% of samples from high-risk controls [16]. Another recent study detected significantly elevated EBV DNA loads in NP brush samples from patients with non-NPC head and neck tumors [15].
Previous studies have also demonstrated that a series of EBV-encoded microRNAs (miRNAs) are specifically expressed in NPC tumors, particularly the BamHI A rightward transcript (BART) miRNAs, and may play a role in tumorigenesis [18]. Because EBV gene products should only originate from infected cells, we hypothesized that detecting functional EBV miRNAs could improve the accuracy and efficiency of NPC diagnosis. miRNAs located in the EBV cluster 1 region reportedly exhibit higher expression and lower degrees of variability than miRNAs from the cluster 2 region [18–20]. Five miRNAs, including mir-bart1-5p, mir-bart5, mir-bart6-5p, mir-bart16 and mir-bart17-5p, were selected for testing in our study. However, mir-bart16 was excluded due to its relatively poor performance in our preliminary experiments (data not shown). The aim of this study was to assess the value of detecting EBV miRNAs from NP brush samples for diagnosis and monitoring of NPC.
RESULTS
Characteristics of the study population
Biopsy tissue samples were collected from 122 different patients, including 53 NPC patients, 21 with chronic nasopharyngitis, 24 with lymphoma and 24 with non-NPC head and neck tumors (Figure 1). Samples from patients with lymphoma or non-NPC head and neck tumors were not taken from the nasopharynx region.
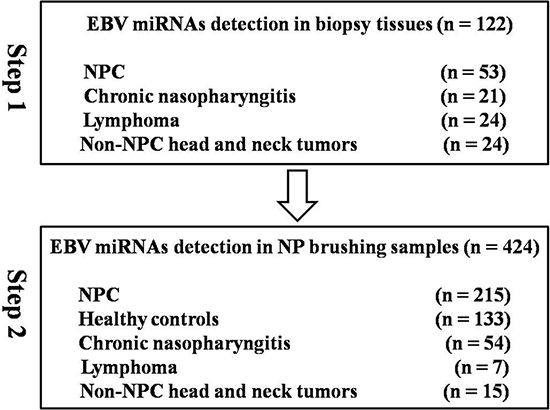
Figure 1: Study workflow.
NP brush samples were collected from 424 different participants. There was no significant difference in age (p = 0.426) or gender (p = 0.095) between case and control groups. In contrast, significant differences in VCA-IgA titer (P < 0.001) and EA-IgA titer (P < 0.001) were observed. TNM staging was performed for the patients using the Chinese 2008 staging system. Additionally, nearly all of the patients were classified as WHO type 3. The detailed characteristics of the study population are presented in Table 1.
Table 1: Characteristics of participants undergoing nasopharyngeal brush sampling
Variable |
NPC |
Controls |
P value |
|---|---|---|---|
Number |
215 |
209 |
|
Age(y) |
0.426 |
||
mean |
47.1 |
47.8 |
|
Gender |
0.095 |
||
male |
157 |
137 |
|
female |
58 |
72 |
|
VCA-IgA titer |
<0.001 |
||
negativity |
20 |
129 |
|
1:10 - 1:40 |
21 |
50 |
|
≥1:80 |
174 |
30 |
|
EA-IgA titer |
<0.001 |
||
negativity |
67 |
195 |
|
1:10 - 1:40 |
134 |
14 |
|
≥1:80 |
14 |
0 |
|
Cancer stage# |
|||
Stage I/II |
15 |
||
Stage III |
86 |
||
Stage IV |
48 |
||
T stage |
|||
T1/T2/T3/T4 |
7/28/81/33 |
||
N stage |
|||
N0/N1/N2/N3 |
6/59/70/14 |
||
M stage |
|||
M0/M1 |
137/12 |
||
Histopathology |
|||
WHO 3 |
2/213 |
#The pathological staging information was evaluated by clinical doctors based on comprehensive results of magnetic resonance (MR), histopathology and clinical symptoms. Some patients were biopsy-diagnosed with NPC in our Cancer Center but subsequently moved to other hospitals for further diagnosis and treatment. For these patients, results such as MR were not collected, while only their histopathology information was obtained. The staging information of 66 patients was lost in this study.
EBV miRNAs were uniquely expressed in biopsy tissues from NPC patients
Four EBV miRNAs (mir-bart1-5p, mir-bart5, mir-bart6-5p and mir-bart17-5p) exhibited high expression in biopsy tissue samples from NPC patients, whereas they were only detected at low levels in non-NPC controls (Figure 2). The median expression values of mir-bart1-5p, mir-bart5, mir-bart6-5p and mir-bart17-5p were 68, 11, 33 and 4.1, while their median expression values in non-NPC controls were 0.002, 0.001, 0.003, and 0.001, respectively. The fold changes of the four miRNAs were 22220, 10255, 7869 and 5416 (all with p < 0.001). The relative expression ranges of the miRNAs in clinical tissue samples are presented in Supplementary Table S1.
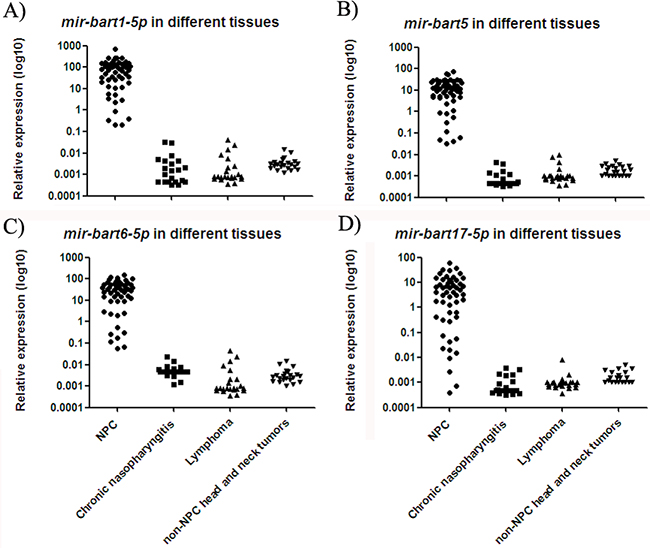
Figure 2: Expressions of four EBV miRNAs in different NP tissue samples. The levels of mir-bart1-5p A. mir-bart5 B. mir-bart6-5p C. and mir-bart17-5p D. were higher in tissue samples from NPC patients as compared to non-NPC controls (p < 0.001).
EBV miRNAs were significantly elevated in nasopharyngeal brush samples from NPC patients
NP brush sampling provided a less invasive pathway for directly collecting samples from the nasopharynx (Supplementary Figure S1). Similar to the results for biopsy tissue samples, relative expression of all four miRNAs was significantly increased in NP brush samples from NPC patients as compared to controls (p < 0.001; Figure 3). The median expression values of mir-bart1-5p, mir-bart5, mir-bart6-5p and mir-bart17-5p were 2.05, 0.84, 1.08 and 0.12, while their median expression values in controls were 0.002, 0.001, 0.002, and 0.001, respectively. The fold changes were 4300, 4840, 4350 and 3700 (all with p < 0.001). The relative expression ranges of the miRNAs in the NP brush samples are presented in Supplementary Table S2.
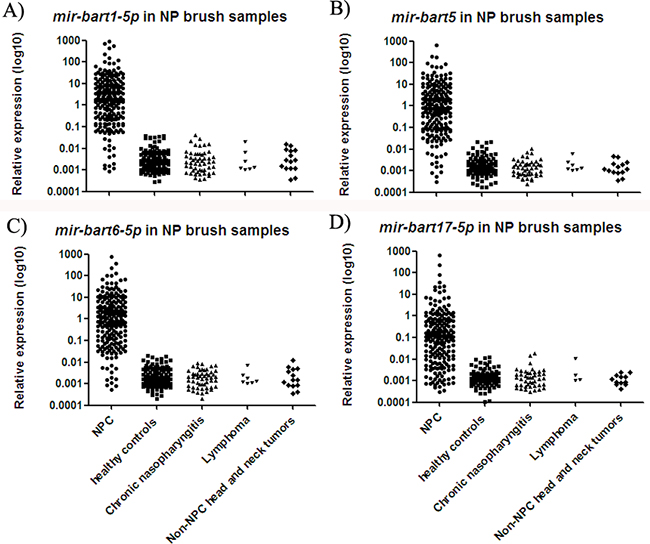
Figure 3: Expressions of four EBV miRNAs in NP brush samples from different participants. The levels of mir-bart1-5p A. mir-bart5 B. mir-bart6-5p C. and mir-bart17-5p D. were significantly increased in the NPC group relative to the controls (p < 0.001).
EBV miRNAs from NP brush samples can be used as biomarkers for diagnosing NPC
Receiver operating characteristic (ROC) analysis revealed that mir-bart1-5p, mir-bart5, mir-bart6-5p and mir-bart17-5p were all valuable biomarkers for differentiating NPC from controls. Area under the curve (AUC) values were 0.98 (95% CI, 0.96 to 0.99), 0.97 (95% CI, 0.95 to 0.99), 0.97 (95% CI, 0.94 to 0.99) and 0.92 (95% CI, 0.89 to 0.95), respectively. At the cut off value (COV) of 0.050 for mir-bart1-5p, the optimal sensitivity and specificity were 93.5% and 100%, respectively. At the COV of 0.023, the optimal sensitivity and specificity for mir-bart5 were 91.2% and 100%; for mir-bart6-5p these values were 91.6% and 100%, respectively. mir-bart17-5p exhibited slightly less accuracy, with 83.7% sensitivity and 92.8% specificity (Table 2). Combining the four miRNAs did not strengthen diagnosis accuracy (data not shown). The correlation coefficients of mir-bart1-5p with mir-bart5, mir-bart6-5p and mir-bart17-5p were 0.93, 0.97 and 0.84, respectively (all with p < 0.001; Supplementary Figure S2).
Table 2: Sensitivity, specificity, PPV and NPV of four EBV miRNAs
miRNAs |
COV |
Case |
Control |
Sensitivity (%) |
Specificity (%) |
PPV (%) |
NPV (%) |
AUC |
||
|---|---|---|---|---|---|---|---|---|---|---|
Median expression |
No. positivity/negativity |
Median expression |
No. positivity/negativity |
|||||||
mir-bart1-5p |
0.050 |
2.05 |
201/14 |
0.002 |
0/209 |
93.5 |
100 |
100 |
93.7 |
0.98 |
mir-bart5 |
0.023 |
0.84 |
196/19 |
0.001 |
0/209 |
91.2 |
100 |
100 |
91.6 |
0.97 |
mir-bart6-5p |
0.023 |
1.08 |
197/18 |
0.002 |
0/209 |
91.6 |
100 |
100 |
92.1 |
0.97 |
mir-bart17-5p |
0.004 |
0.12 |
180/35 |
0.001 |
15/194 |
83.7 |
92.8 |
92.3 |
84.7 |
0.92 |
PPV: positive predictive value
NPV: negative predictive value
mir-bart1-5p levels in NP brush samples were reflective of disease progression
Due to its high correlation with the other three miRNAs, mir-bart1-5p was further analyzed in samples at various stages of disease progression. Levels of mir-bart1-5p in control NP brush samples were significantly lower than in stage I/II NPC samples (p < 0.001; Figure 4A), and samples from the T1 subgroup (p < 0.001; Figure 4B). Stage I/II and T1 subgroup NPC samples also exhibited significantly lower levels of EBV miRNAs when compared with samples from patients with more advanced disease. In contrast, no significant difference was observed in node–positive/negative or metastasis-positive/negative cases (Figure 4C and 4D).
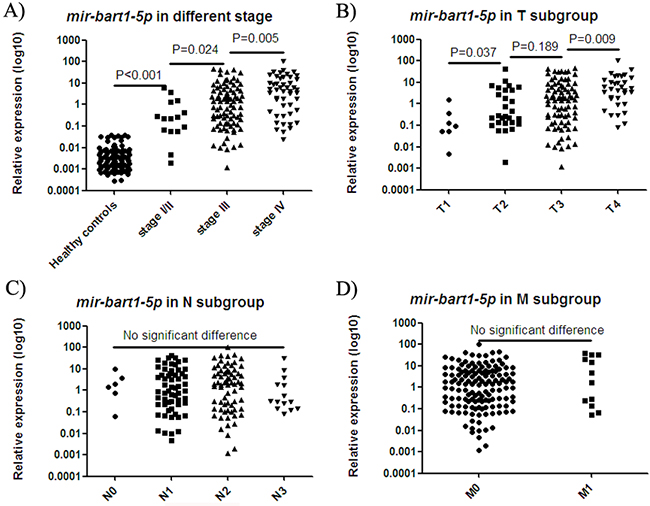
Figure 4: mir-bart1-5p expression in samples at various stages of disease progression. The expression of mir-bart1-5p in the NP brush samples of healthy controls was significantly lower than in stage I/II NPC samples A. and samples from the T1 subgroup B. Stage I/II A. and T1 subgroup B. NPC samples also exhibited significantly lower levels of EBV mir-bart1-5p compared with the levels observed in samples from more advanced disease. No significant difference was observed in node–positive/negative C. or metastasis-positive/negative cases D.
Diagnosis by mir-bart1-5p versus pathological biopsy, VCA-IgA and EA-IgA
We further compared the diagnostic performance of mir-bart1-5p with common diagnostic methods, including histopathological assessment based on initial biopsy sampling, and molecular diagnosis based on VCA-IgA and EA-IgA detection.
In this study, repeated biopsy sampling was conducted in 16 patients. The expression of mir-bart1-5p was above the COV (0.050 copy/RUN6B) in only the NP brush samples. The results of diagnoses by biopsy and mir-bart1-5p in NP brush samples are summarized in Supplementary Table S3. Computed tomography (CT) and magnetic resonance imaging (MRI) results, along with endoscopy images, are shown for a representative patientin Supplementary Figure S3. A slightly higher sensitivity (93.5%) was obtained using mir-bart1-5p detection as compared with initial biopsy (92.6%; Table 3).
Table 3: Diagnostic performance of mir-bart1-5p and other current methods
COV |
Sensitivity (%) |
Specificity (%) |
PPV (%) |
NPV (%) |
|
|---|---|---|---|---|---|
mir-bart1-5p |
0.050 |
93.5 |
100 |
100 |
93.7 |
Initial biopsy |
N/A |
92.6 |
100 |
100 |
92.9 |
VCA-IgA |
≥ 1:80 |
80.9 |
85.6 |
85.3 |
81.3 |
EA-IgA |
≥ 1:10 |
68.8 |
92.8 |
91.4 |
74.4 |
IgA antibody titers for VCA-IgA and EA-IgA are traditional markers used in areas at high-risk for NPC. VCA-IgA exhibited 80.9% sensitivity and 85.6% specificity at COV ≥ 1:80, whereas EA-IgA exhibited 68.8% sensitivity and 92.8% specificity at COV ≥ 1:10 in this study (Table 3).
DISCUSSION
Early and accurate diagnosis of NPC is important for enhancing patient survival. Blood and NP brush tissue-sampling methods, which are less invasive than biopsies, show promise for diagnosing and monitoring NPC through detection of EBV-related biomarkers (such as VCA-IgA titer and EBV DNA load). In this study, detection of EBV mir-bart1-5p in NP brush samples was identified as a novel method for clinically diagnosing and monitoring the progression of NPC. The clinical application of a NP brush sampling/EBV miRNA detection method could help to improve patient outcomes in high-prevalence areas such as southern China.
Utilizing NP brush sampling to detect EBV DNA load has attracted increasing attention in recent years, and various DNA fragments have been detected in multiple studies. For example, a 213-bp region of EBV nuclear antigen-1 (EBNA-1) [14, 15] and a 75-bp fragment of the EBV BamHI-W region [12] are commonly used in detection studies, and have comparable sensitivities and specificities. However, EBV DNA has also been detected in NP brush samples from patients with non-NPC head and neck tumors [15]. EBV-encoded small RNA (EBER) in situ hybridization (ISH) is a useful qualitative technique for identifying EBV infection in cells [21]. EBER ISH results from this study showed that high EBER signaling was only observed in tissue slices from NPC patients, suggesting there was no EBV infection in non-NPC controls (Supplementary Figure S4). Similarly, we observed that the EBV miRNAs tested in this study (mir-bart1-5p, mir-bart5 and mir-bart6-5p) were detected only in NPC patient samples, and not in any of the controls (Figure 3), resulting in a specificity of 100% for NPC diagnosis (Table 2). These results verified our hypothesis that quantitative detection of functional EBV miRNA transcripts could be used to effectively diagnose NPCs. Specific EBV miRNA detection could also potentially be used in combination with EBER ISH to further enhance NPC diagnosis accuracy.
Plasma EBV DNA testing appears to be of limited value when diagnosing patients with early clinical stage NPC [22] or local recurrence [23]. Small tumors may release only low levels of EBV DNA fragments into the blood. By contrast, our findings indicate that EBV miRNAs can be effectively used to diagnose early stage NPC. EBV mir-bart1-5p detected in NP brush samples was used to correctly diagnose 13 NPC cases at stage I/II and 6 cases at T1 in the first sampling. The potential value of EBV mir-bart1-5p for early NPC diagnosis is further illustrated by the following two cases. First, a 49-year-old man presented with minor symptoms and was negative for plasma EBV DNA load and for VCA-IgA and EA-IgA titers. The second patient was a 74-year-old man with a VCA-IgA titer of 320. The histopathological assessments based on initial NP biopsy were NPC-negative for both patients, likely due to no obvious tumor mass under endoscopic examination. The two patients respectively underwent four and two biopsies before a correct pathological diagnosis was achieved. The first patient was diagnosed with T1N2M0, stage III disease, and the second patient was diagnosed with T1N2M1, stage IV disease. In contrast to the other diagnostic techniques used, expression of mir-bart1-5p at initial NP brush sampling was above COV (0.05 copy/RUN6B) in both cases, indicating its unique diagnostic value even for early NPC cases.
We also observed that mir-bart1-5p expression potentially could be used as a biomarker for monitoring NPC progression, in that increasing mir-bart1-5p expression correlated with disease progression (Figure 4A). However, expression was only significantly correlated with the T subgroup of NPC (Figure 4B), not N and M stage (Figure 4C and 4D). NP brush sampling may directly reflect the local tumor load in the nasopharynx.
The nasopharynx is often difficult to access, and pathological diagnosis based on biopsies may lead to false negatives. While initial biopsy sampling resulted in a diagnostic sensitivity of 92.6% among patients in this study (Table 3), 16 out of 215 NPC patients required repeated biopsy sampling in order to obtain a correct pathological confirmation. Notably, the EBV mir-bart1-5p levels in the NP brush samples from these 16 patients were all above the COV, correctly indicating the presence of NPC (Supplementary Table S3). Similarly, in nearly all cases for which VCA-IgA titer produced a false negative diagnosis (38 out of 41), mir-bart1-5p expression was above the COV (data not shown).
Still, there were 14 NPC cases in which NP brush samples tested negative for mir-bart1-5p expression. Most of these cases were diagnosed within the T3 subgroup. mir-bart1-5p detection and EBER ISH were both conducted directly with tissue slices from these patients. The results showed mir-bart1-5p and EBERs were both expressed in nasopharyngeal tumor cells (Supplementary Figure S5). It is possible that NP brush sampling failed to capture tumor cells in these samples, leading to false negative results. Although repeated brush sampling is feasible, further optimization of the sampling method is required. For example, a brush with a shape more suitable to the physiological structure of the nasopharynx should be developed.
In conclusion, quantification of EBV mir-bart1-5p in NP brush samples could be used as an effective supplementary method for NPC diagnosis. Thanks in part to the reduced invasiveness of the described NP brush sampling/EBV miRNA detection NPC diagnostic technique, it has great potential for use in monitoring disease progression in NPC patients and in the screening of high-risk populations.
MATERIALS AND METHODS
Clinical specimens and study design
To identify EBV miRNAs expressed exclusively in NP tissues from NPC patients, 122 clinical tissue samples from patients with chronic nasopharyngitis, lymphoma, non-NPC head and neck tumors, and NPC were obtained from the Tumor Resource Bank of our cancer center. Subsequently, a total of 424 participants undergoing NP brush sampling were recruited from January 2013 to October 2015. Two hundred and ninety one NP brush samples were consecutively collected from participants when they underwent NP biopsy at Sun Yat-Sen University Cancer Center (SYSUCC). Among the participants, 215 exhibited biopsy-positive NPC, 54 were eventually diagnosed with chronic nasopharyngitis, 7 were diagnosed with lymphoma, and 15 had non-NPC tumors. Additionally, 133 NP brush samples from healthy control individuals were collected from Sihui city, a high-risk area of Guangdong province, China. A three ml blood sample was collected from each participant. The Human Ethics Committee of the Sun Yat-Sen University Cancer Center approved this study, and all participants provided informed consent.
Sampling procedures
Experienced specialists and resident trainees conducted NP brush sampling under the guidance of endoscopy. An endoscope was used to evaluate the entire nasopharynx, and images were captured at the site of suspicious tumors. Before biopsy, a NP brush (Copan Diagnostics) was inserted into the nose until the NP cavity was reached. The brush was rotated several times over the NP epithelium at the site of the suspected lesion and then quickly removed. Immediately after sampling, the brush tip (1.5 cm) was cut, placed in one ml of RNAlater (Invitrogen, Carlsbad, CA, USA) and stored at –80°C until use. For healthy participants, NP brush samples were obtained from the lateral pharynx because this was the most common site for disease occurrence.
RNA extraction, reverse transcription (RT) and quantitative polymerase chain reaction (Q-PCR)
Total RNA from biopsy tissue and NP brush samples was extracted using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. The RNA concentration was quantified using a NanoDrop 1000 Spectrophotometer (NanoDrop Technologies, Waltham, MA).
A total of 350 ng RNA (100 ng in tissue samples) was used for reverse transcription with a TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s protocol. Reverse transcription reactions (20 μl volume/sample) were conducted with custom stem-loop primers (Applied Biosystems) specific to the corresponding mature sequence obtained from miRBase (www.miRBase.org). RNU6B was usually used as the endogenous control, and 4 EBV miRNAs were targets. Each reaction contained 0.2 μl dNTPs (100 mM), 1.3 μl MultiScribe Reverse Transcriptase (50 U/μL), 2.0 μl 10X Reverse Transcription Buffer, 0.25 μl RNase Inhibitor (20 U/μL), 8 μl RT primer (1.6 μl/per primer) and 8.25 μl RNA and nuclease-free water (350 ng total RNA). The following parameter values were used to perform the thermal cycles: 16°C for 30 min, then 42°C for another 30 min followed by 85°C for 5 min.
Q-PCR was carried out on the LightCycler®480 instrument (Roche). Each PCR reaction (10 μl volume) included 1 μl of RT product, 5 μl of TaqMan 2X Universal PCR Master Mix, 0.5 μl of 20X TaqMan MicroRNA Assay mix and 3.5 μl nuclease-free water. All reactions were performed in duplicate and the relative expression of each miRNA (copy number/RNU6B) in one sample was determined using the equation 2-ΔCT, where ΔCT = Ct (miRNA) – Ct (RNU6B) (Ct value were the threshold cycle for each miRNA). Thermal cycling was initiated with a denaturation step of 10 min at 95°C, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min.
VCA-IgA and EA-IgA antibody titers in blood specimens
The VCA-IgA and EA-IgA titer of EBV were measured using a commercial kit (Zhongshan Bio-tech Co Ltd., Zhongshan City, China) based on standard techniques described previously [24].
EBERs and mir-bart1-5p in situ hybridization
Paraffin-embedded tissue sections were obtained from the Pathology Department of our cancer center. They included 14 NPC samples with negative mir-bart1-5p expression in NP brushing samples, 7 lymphoma samples, 3 non-NPC other tumor samples and 16 NP tissue samples with chronic nasopharyngitis. The presence of EBV in tumor cells was assessed by in situ hybridization (ISH) using the EBER ISH kit (ZSGB Bio-tech Co Ltd., Beijing, China). A DIG-double labeled locked nucleic acid (LNA)-based probe specific for mir-bart1-5p (Exiqon, Vedbaek, Denmark) was used to detect mir-bart1-5p in tumor cells. ISH was performed according to the manufacturer’s instructions.
Statistical analyses
The Mann-Whitney test was used to compare the nonparametric variables between NPC and control groups. A one-way ANOVA was used to compare the differences in multiple groups. Differences in a demographic variable (e.g., sex) between the cases and all controls were evaluated using a Chi-square test. EBV miRNA correlations were assessed by applying Pearson correlation coefficients and were subjected to two-tailed significance tests; p < 0.05 was considered significant. Receiver operating characteristic (ROC) curves were used to evaluate diagnostic value, and area under the ROC curve (AUC) was used as an accuracy index for evaluating diagnostic performance. We used the curves to select cut-off values (COV) with maximum sensitivity and specificity. The statistical analyses were performed using the SPSS 16 software.
ACKNOWLEDGMENTS
We thank Dr Pei-Yu Huang, Dr Rui Sun, Dr Yi-Jun Hua, Dr Dong-Hua Luo and Dr Qiu-Yan Chen from the Department of Nasopharyngeal Carcinoma, Sun Yat-sen University Cancer Center, for conducting the nasopharyngeal brush sampling.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
GRANT SUPPORT
This work was supported by the National Science & Technology Pillar Program during the Twelfth Five-year Plan Period (NO: 2014BAI09B10), National Science Fund for Distinguished Young Scholars (81325018), the Key Project for International Cooperation and Exchange of the National Natural Science Foundation of China (81220108022), the National Basic Research Program of China (2011CB504303) and the Science and Technology Planning Project of Guangdong Province, China (2011B031800218).
REFERENCES
1. Razak AR, Siu LL, Liu FF, Ito E, O’Sullivan B, Chan K. Nasopharyngeal carcinoma: the next challenges. Eur J Cancer. 2010; 46: 1967–1978.
2. He ML, Luo MX, Lin MC, Kung HF. MicroRNAs: Potential diagnostic markers and therapeutic targets for EBV-associated nasopharyngeal carcinoma. Biochim Biophys Acta. 2012; 1825: 1–10.
3. Niedobitek G, Young LS. Epstein-Barr virus persistence and virus-associated tumours. Lancet. 1994; 343: 333–335.
4. Pathmanathan R, Prasad U, Sadler R, Flynn K, Raab-Traub N. Clonal proliferations of cells infected with Epstein-Barr virus in preinvasive lesions related to nasopharyngeal carcinoma. N Engl J Med. 1995; 333: 693–698.
5. Chan KC, Lo YM. Circulating EBV DNA as a tumor marker for nasopharyngeal carcinoma. Semin Cancer Biol. 2002; 12: 489–496.
6. Lin JC, Wang WY, Chen KY, Wei YH, Liang WM, Jan JS, Jiang RS. Quantification of plasma Epstein-Barr virus DNA in patients with advanced nasopharyngeal carcinoma. N Engl J Med. 2004; 350: 2461–2470.
7. Le QT, Jones CD, Yau TK, Shirazi HA, Wong PH, Thomas EN, Patterson BK, Lee AW, Zehnder JL. A comparison study of different PCR assays in measuring circulating plasma epstein-barr virus DNA levels in patients with nasopharyngeal carcinoma. Clin Cancer Res. 2005; 11: 5700–5707.
8. Yip TT, Ngan RK, Fong AH. Application of circulating plasma/serum EBV DNA in the clinical management of nasopharyngeal carcinoma. Oral Oncol. 2014; 50: 527–538.
9. Sheen TS, Ko JY, Chang YL, Chang YS, Huang YT, Chang Y, Tsai CH, Hsu MM. Nasopharyngeal swab and PCR for the screening of nasopharyngeal carcinoma in the endemic area: a good supplement to the serologic screening. Head Neck. 1998; 20: 732–738.
10. Tune CE, Liavaag PG, Freeman JL, van den Brekel MW, Shpitzer T, Kerrebijn JD, Payne D, Irish JC, Ng R, Cheung RK, Dosch HM. Nasopharyngeal brush biopsies and detection of nasopharyngeal cancer in a high-risk population. J Natl Cancer Inst. 1999; 91: 796–800.
11. Lin SY, Tsang NM, Kao SC, Hsieh YL, Chen YP, Tsai CS, Kuo TT, Hao SP, Chen IH, Hong JH. Presence of Epstein-Barr virus latent membrane protein 1 gene in the nasopharyngeal swabs from patients with nasopharyngeal carcinoma. Head Neck. 2001; 23: 194–200.
12. Tong JH, Tsang RK, Lo KW, Woo JK, Kwong J, Chan MW, Chang AR, van Hasselt CA, Huang DP, To KF. Quantitative Epstein-Barr virus DNA analysis and detection of gene promoter hypermethylation in nasopharyngeal (NP) brushing samples from patients with NP carcinoma. Clin Cancer Res. 2002; 8: 2612–2619.
13. Hao SP, Tsang NM, Chang KP. Screening nasopharyngeal carcinoma by detection of the latent membrane protein 1 (LMP-1) gene with nasopharyngeal swabs. Cancer. 2003; 97: 1909–1913.
14. Stevens SJ, Verkuijlen SA, Hariwiyanto B, Harijadi, Paramita DK, Fachiroh J, Adham M, Tan IB, Haryana SM, Middeldorp JM. Noninvasive diagnosis of nasopharyngeal carcinoma: nasopharyngeal brushings reveal high Epstein-Barr virus DNA load and carcinoma-specific viral BARF1 mRNA. Int J Cancer. 2006; 119: 608–614.
15. Adham M, Greijer AE, Verkuijlen SA, Juwana H, Fleig S, Rachmadi L, Malik O, Kurniawan AN, Roezin A, Gondhowiardjo S, Atmakusumah D, et al. Epstein-Barr virus DNA load in nasopharyngeal brushings and whole blood in nasopharyngeal carcinoma patients before and after treatment. Clin Cancer Res. 2013; 19: 2175–2186.
16. Zheng XH, Lu LX, Li XZ, Jia WH. Quantification of EBV DNA load in nasopharyngeal brushing samples in the diagnosis of nasopharyngeal carcinoma in southern china. Cancer Sci. 2015; 106: 1196–1201
17. Liavaag PG, Cheung RK, Kerrebijn JD, Freeman JL, Irish JC, Dosch HM. The physiologic reservoir of Epstein-Barr virus does not map to upper aerodigestive tissues. Laryngoscope. 1998; 108: 42–46.
18. Lo AK, Dawson CW, Jin DY, Lo KW. The pathological roles of BART miRNAs in nasopharyngeal carcinoma. J Pathol. 2012; 227: 392–403.
19. Cosmopoulos K, Pegtel M, Hawkins J, Moffett H, Novina C, Middeldorp J, Thorley-Lawson DA. Comprehensive profiling of Epstein-Barr virus microRNAs in nasopharyngeal carcinoma. J Virol. 2009; 83: 2357–2367.
20. Wong AM, Kong KL, Tsang JW, Kwong DL, Guan XY. Profiling of Epstein-Barr virus-encoded microRNAs in nasopharyngeal carcinoma reveals potential biomarkers and oncomirs. Cancer. 2012; 118: 698–710.
21. Howe JG, Steitz JA. Localization of Epstein-Barr virus-encoded small RNAs by in situ hybridization. Proc Natl Acad Sci U S A. 1986; 83: 9006–9010.
22. Ji MF, Huang QH, Yu X, Liu Z, Li X, Zhang LF, Wang P, Xie SH, Rao HL, Fang F, Guo X, Liu Q, Hong MH, et al. Evaluation of plasma Epstein-Barr virus DNA load to distinguish nasopharyngeal carcinoma patients from healthy high-risk populations in Southern China. Cancer. 2014; 120: 1353–1360.
23. Leung SF, Lo YM, Chan AT, To KF, To E, Chan LY, Zee B, Huang DP, Johnson PJ. Disparity of sensitivities in detection of radiation-naive and postirradiation recurrent nasopharyngeal carcinoma of the undifferentiated type by quantitative analysis of circulating Epstein-Barr virus DNA1,2. Clin Cancer Res. 2003; 9: 3431–3434.
24. Xu FH, Xiong D, Xu YF, Cao SM, Xue WQ, Qin HD, Liu WS, Cao JY, Zhang Y, Feng QS, Chen LZ, Li MZ, Liu ZW, et al. An epidemiological and molecular study of the relationship between smoking, risk of nasopharyngeal carcinoma, and Epstein-Barr virus activation. J Natl Cancer Inst. 2012; 104: 1396–1410.

