INTRODUCTION
Epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors (TKIs) became the first-line treatment for patients with non-small cell lung cancer (NSCLC) harboring mutant EGFR gene [1]. Most of these patients are highly responsive to treatment with EGFR-TKIs. However, about 20–30% patients with EGFR activating mutations show primary resistance to EGFR-TKIs [2]. Therefore, identifying the prognostic biomarkers of primary resistance to EGFR-TKIs in these patients is urgently needed.
Bcl-2-like protein 11 (BIM) is a BH3-only proapoptotic member of the Bcl-2 protein family [3]. Up-regulation of BIM correlated with gefitinib-induced apoptosis in gefitinib-sensitive EGFR-mutant lung cancer cells [4]. In addition, knockdown of BIM expression by RNA interference eliminated erlotinib-induced cell killing in vitro [5]. Addition of a BH3 mimetic significantly enhanced killing of NSCLC cells by gefitinib [6]. Furthermore, high level of BIM expression was a marker of longer progression-free survival (PFS) in EGFR-mutant NSCLC treated with erlotinib [7]. Therefore, BIM might be a biomarker of survival in EGFR-mutant NSCLC.
Recently, King Pan Ng and colleagues identified a common intronic deletion polymorphism in BIM [8]. This BIM deletion polymorphism was absent in individuals from African and European populations, but was found in about 12% of Asian population [8]. They demonstrated that patients with EGFR-mutant NSCLC harboring BIM deletion polymorphism showed significant inferior responses to TKIs than patients without this polymorphism [9]. This finding was confirmed by several studies [9–11]. However, Lee et al. suggested that BIM deletion polymorphism was not predictive of PFS for EGFR-TKIs [12]. The aim of this meta-analysis was to summarize all the available evidence and determine the predictive role of BIM deletion polymorphism for EGFR-TKIs in NSCLC.
RESULTS
Literature search
The process of identifying studies is shown in Figure 1. A total of 95 publications were identified in the initial search, and 1 publication was identified from other source. Based on screening of titles or abstracts, 78 records were excluded. Full text articles were retrieved only for 18 publications and assessed for eligibility. Of these 18 publications, 13 publications were excluded. Finally, 5 studies were included in this meta-analysis.
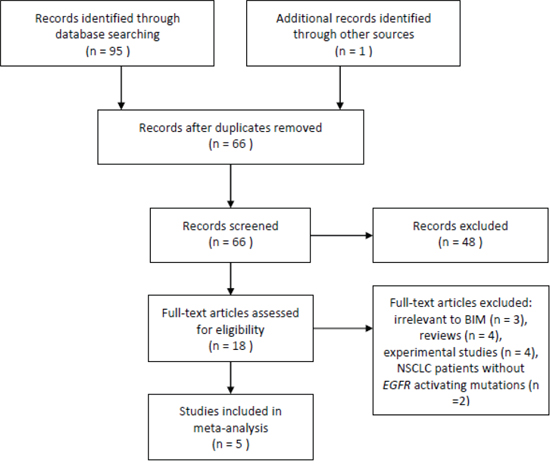
Figure 1: Flow of study identification, inclusion, and exclusion.
Study characteristics
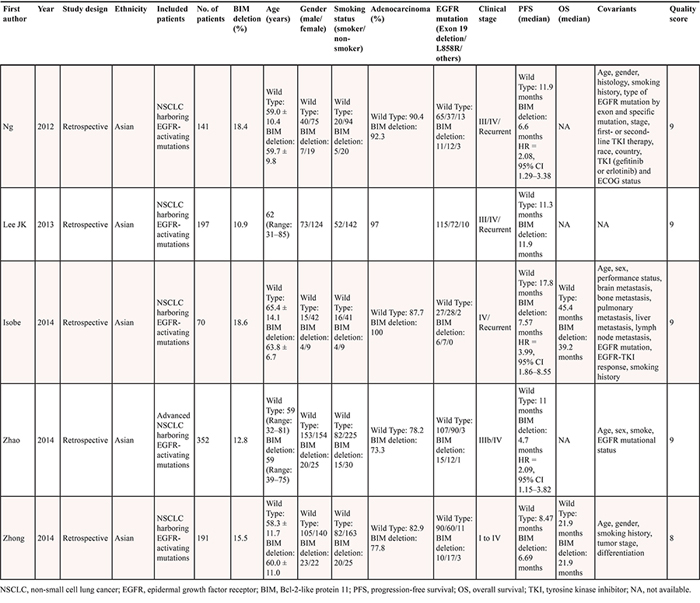
Five retrospective cohort studies were included. All of the studies were conducted in Asian population (n = 951). Four studies included advanced NSCLC patients, and one study included NSCLC from early stage to advanced stage. Five studies reported the status of EGFR mutations, while two studies did not provide this information. All studies were assessed by Newcastle–Ottawa Scale (NOS). The quality scores ranged from 8 to 9, suggesting that the methodological quality was high. The characteristics of each study are presented in Table 1.
Table 1: Characteristics of the included studies
Quantitative data synthesis
All the studies reported the data of PFS. Except the study by Lee JK, all the studies suggested that BIM deletion polymorphism was associated with reduced PFS. Three studies reported adjusted hazard ratios (HRs) and 95% confidence intervals (Cis). Compared with BIM wild type, BIM deletion polymorphism was predictive of shorter PFS in NSCLC patients who were treated with EGFR-TKIs (adjusted HR = 2.38, 95% CI 1.66–3.41, P < 0.001; Figure 2). No significant heterogeneity was observed (I2 = 11%).
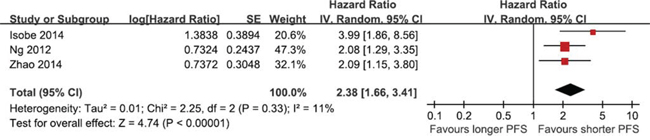
Figure 2: The prognostic role of BIM deletion polymorphism on PFS.
DISCUSSION
This systematic review and meta-analysis found that BIM deletion polymorphism was associated with a poor response to EGFR-TKIs in NSCLC patients. NSCLC patients with BIM deletion polymorphism exhibited shorter PFS when they received EGFR-TKIs.
As for overall survival (OS), two studies provided the median OS in BIM deletion polymorphism group and BIM wild type group. One studies indicated that patients with BIM deletion polymorphism had shorter OS than did those without BIM deletion polymorphism, while another study did not confirm this result. All the two studies did not provided statistical results, thus we did not do meta-analysis of OS.
The BIM deletion polymorphism contained a deletion of a 2903 bp fragment in intron 2. This deletion resulted in the preferential splicing of exon 3 over exon 4, which generated a BIM isoform that lacked the BH3 [8], thereby causing EGFR-TKIs resistant. They also reported that the addition of BH3-mimetic drugs could restore TKIs sensitivity [8]. Recently, Nakagawa et al. suggested the histone deacetylase (HDAC) inhibitor vorinostat could circumvent EGFR-TKI resistance in EGFR-mutant NSCLC cell lines and in xenograft models [13]. The combination of EGFR-TKIs plus a BH3-mimetic drug or vorinostat should be considered for the NSCLC patients with BIM deletion polymorphism in the future.
This meta-analysis had some advantages. First, this was the first meta-analysis which assessed the association between BIM deletion polymorphism and efficacy of EGFR-TKIs. Second, no significant heterogeneity was found in this meta-analysis. Third, the quality of the included studies was high. However, the limitations should also be acknowledged. First, there were only five studies included in this meta-analysis. Although all the studies reported PFS, only two studies provided OS. Thus, it was still unclear whether BIM deletion polymorphism was a prognostic marker of OS. Second, all of the studies were retrospective design, which were prone to bias (e.g., recall and selection bias). Therefore, prospective studies should be designed to validate the results of this meta-analysis. Third, we could not perform subgroup analyses by age, gender, smoking status, and EGFR mutations due to the insufficient data.
In conclusion, the BIM deletion polymorphism was significantly associated with poor response in NSCLC patients who received EGFR-TKIs treatment. The BIM deletion polymorphism could be a prognostic biomarker of EGFR-TKIs resistance in NSCLC.
MATERIALS AND METHODS
Literature search
PubMed, Embase, and the Cochrane Register of Controlled Trials were searched for relevant studies published up to 20 Apr. 2015. The following terms were used: (“NSCLC” or “lung cancer” or “non-small cell lung cancer”) and (“Bcl-2-like protein 11” or “Bcl-2-like 11” or BIM or BCL2L11”. No language restrictions were imposed. References from relevant articles, including review papers, were also reviewed.
Study selection
Studies were included in the meta-analysis if they fulfilled the following inclusion criteria: 1) study design: cohort studies; 2) population: NSCLC patients with EGFR activating mutations; 3) intervention: EGFR-TKIs; 4) primary outcome: the effect of BIM deletion polymorphism on PFS. Abstract, case reports, review articles, experimental studies and commentary articles were excluded.
Data collection and methodological quality assessment
The following data were extracted from each study: the first author, publication year, study design, ethnicity, included patients, number of patients, the percent of BIM deletion, sex, age, smoking status, the percent of adenocarcinoma, EGFR mutations, clinical stage of patients, reported PFS, and covariates controlled for multivariable analysis. Two reviewers independently extracted the relevant data. Any disagreement was resolved by consensus in meetings with all investigators.
Two reviewers completed the quality assessment independently. The NOS was used to evaluate the methodological quality, which scored studies by the selection of the study groups, the comparability of the groups, and the ascertainment of the outcome of interest [14]. We considered a study awarded 0–3, 4–6, or 7–9 as a low-, moderate-, or high-quality study, respectively. Discrepancies were resolved by consensus and discussion.
Statistical analysis
We estimated the HR with 95% CI for primary outcome. The multivariable-adjusted HRs with 95% CIs were pooled in our analysis. We used a random effects model. Statistical heterogeneity among studies was evaluated using the Q and I2 statistics. Publication bias was investigated by funnel plot if more than 10 studies were included. All statistical analyses were performed with Revman 5.1 software (Nordic Cochrane Center, Copenhagen, Denmark).
GRANT SUPPORT
This study was supported by grant No. 13DZ1930601 from Industry-Academia-Research projection of Shanghai and International scientific and technological cooperation projection (No. 2015DFA31810).
CONFLICTS OF INTEREST
The authors have no conflicts of interest to report.
REFERENCES
1. Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H, Gemma A, Harada M, Yoshizawa H, Kinoshita I, Fujita Y, Okinaga S, Hirano H, et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med. 2010; 362:2380–8.
2. Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT, Saijo N, Sunpaweravong P, Han B, Margono B, Ichinose Y, Nishiwaki Y, Ohe Y, Yang JJ, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med. 2009; 361:947–57.
3. Youle RJ, Strasser A. The BCL-2 protein family: opposing activities that mediate cell death. Nat Rev Mol Cell Biol. 2008; 9:47–59.
4. Costa DB, Halmos B, Kumar A, Schumer ST, Huberman MS, Boggon TJ, Tenen DG, Kobayashi S. BIM mediates EGFR tyrosine kinase inhibitor-induced apoptosis in lung cancers with oncogenic EGFR mutations. PLoS Med. 2007; 4:1669–79; discussion 1680.
5. Gong Y, Somwar R, Politi K, Balak M, Chmielecki J, Jiang X, Pao W. Induction of BIM is essential for apoptosis triggered by EGFR kinase inhibitors in mutant EGFR-dependent lung adenocarcinomas. PLoS Med. 2007; 4:e294.
6. Cragg MS, Kuroda J, Puthalakath H, Huang DC, Strasser A. Gefitinib-induced killing of NSCLC cell lines expressing mutant EGFR requires BIM and can be enhanced by BH3 mimetics. PLoS Med. 2007; 4:1681–89; discussion 1690.
7. Costa C, Molina MA, Drozdowskyj A, Giménez-Capitán A, Bertran-Alamillo J, Karachaliou N, Gervais R, Massuti B, Wei J, Moran T, Majem M, Felip E, Carcereny E, et al. The impact of EGFR T790M mutations and BIM mRNA expression on outcome in patients with EGFR-mutant NSCLC treated with erlotinib or chemotherapy in the randomized phase III EURTAC trial. Clin Cancer Res. 2014; 20:2001–10.
8. Ng KP, Hillmer AM, Chuah CT, Juan WC, Ko TK, Teo AS, Ariyaratne PN, Takahashi N, Sawada K, Fei Y, Soh S, Lee WH, Huang JW, et al. A common BIM deletion polymorphism mediates intrinsic resistance and inferior responses to tyrosine kinase inhibitors in cancer. Nat Med. 2012; 18:521–8.
9. Isobe K, Hata Y, Tochigi N, Kaburaki K, Kobayashi H, Makino T, Otsuka H, Sato F, Ishida F, Kikuchi N, Hirota N, Sato K, Sano G, et al. Clinical significance of BIM deletion polymorphism in non-small-cell lung cancer with epidermal growth factor receptor mutation. J Thorac Oncol. 2014; 9:483–7.
10. Zhao M, Zhang Y, Cai W, Li J, Zhou F, Cheng N, Ren R, Zhao C, Li X, Ren S, Zhou C, Hirsch FR. The Bim deletion polymorphism clinical profile and its relation with tyrosine kinase inhibitor resistance in Chinese patients with non-small cell lung cancer. Cancer. 2014; 120:2299–307.
11. Zhong J, Li ZX, Zhao J, Duan JC, Bai H, An TT, Yang XD, Wang J. Analysis of BIM 11 (BCL-2 like 11 gene) deletion polymorphismin Chinese non-small cell lung cancer patients. Thoracic Cancer. 2014; 5:509–516.
12. Lee JK, Shin JY, Kim S, Lee S, Park C, Kim JY, Koh Y, Keam B, Min HS, Kim TM, Jeon YK, Kim DW, Chung DH, et al. Primary resistance to epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors (TKIs) in patients with non-small-cell lung cancer harboring TKI-sensitive EGFR mutations: an exploratory study. Ann Oncol. 2013; 24:2080–7.
13. Nakagawa T, Takeuchi S, Yamada T, Ebi H, Sano T, Nanjo S, Ishikawa D, Sato M, Hasegawa Y, Sekido Y, Yano S. EGFR-TKI resistance due to BIM polymorphism can be circumvented in combination with HDAC inhibition. Cancer Res. 2013; 73:2428–34.
14. Wells GA, Shea B, O’Connell D, Peterson J, Welch V, Losos M, Tugwell P. The Newcastle-Ottawa Scale (NOS) for assessing the quality of nonrandomised studies in meta-analyses. URL: http://www.ohri.ca/programs/clinical_epidemiology/oxford.asp.

