INTRODUCTION
Melanoma is a highly curable malignancy in early stages but unfortunately holds devastating consequences when metastases develop, with a decline in 5-year survival rate from 98% for localized to only 16% for metastatic melanoma [1]. The site of distant metastasis is an important independent predictor of survival with patients harboring visceral melanoma metastases showing the worst prognostic behavior [1, 2]. Lung is the second most common site for metastatic spread and the annual probability of developing lung metastatic melanoma (LMMs) progressively increases from 10% at 5 years to 17% at 15 years after the resection of the primary tumor [3]. Until recently, systemic chemotherapies (dacarbazine), hydroxyurea, or immunotherapy with high-dose interleukin-2 (IL-2) were the only treatment options approved by the U.S. Food and Drug Administration (FDA) for patients with advanced melanoma [4–6]. These systemic treatments provide little, if any, attested survival benefit for patients harboring LMM, with surgery remaining, unfortunately only for few eligible patients, the best treatment to improve overall survival (OS) [7, 8]. Recent progresses in the understanding of melanoma pathogenesis have allowed the identification of both immunotargeting agents, as ipilimumab, and actionable driver mutations in several genes potentially exploitable for therapeutic purposes [9–11]. The BRAF gene, encoding for a protein member of the mitogen-activated-protein-kinase family (MAPK) [12], is mutated in approximately half of all melanomas, and its mutations, commonly occurring at codon 600 (BRAFV600) [9, 13], have been exploited to develop drugs (vemurafenib, dabrafenib and trametinib) effective against BRAF-driven metastatic melanoma [13–17]. In addition to BRAF, NRAS mutations, mostly affecting codon 61, are found in 10–15% of melanomas and are involved in mutagenic activation of the MAPK pathway [18–20]. They can be targeted by MEK162, a strong MEK1/2-inhibitor [21]. Melanomas have also been reported to over express CKIT whose mutations are found in 1.7% of cutaneous melanomas, 23% of acral melanomas, and 15.6% of mucosal melanomas [22, 23]. Emerging evidence suggests that melanomas with CKIT activation may respond to specific targeted agents [24–26]. Finally, epidermal-growth-factor receptor (EGFR) copy number alterations have been found in primary cutaneous malignant melanomas and their presence has been associated with poor prognosis [27, 28].
Despite of the observation that the lungs are a frequent site of visceral metastases from melanoma, very limited and scattered data are available on the molecular status of pulmonary lesions. The molecular characterization of the above-mentioned genes in a lung metastatic setting could help in the decision to select the best possible therapeutic option among novel tailored biologic therapies or surgical treatment. Based on the above observations, the aim of the present study was to determine the frequency of alterations of the aforementioned genes and possible clinical and pathological correlations in a well characterized cohort of patients affected by lung metastases from advanced melanoma.
RESULTS
Patients and samples characteristics
A total of 25 lung metastasis specimens collected after surgery from 25 patients with LMM were analyzed. The cohort included 10 females (40%) and 15 males (60%). Median age at first diagnosis of primary melanoma was 53 years (range 18–76). Females were significantly younger than males at diagnosis of both primary melanoma and lung metastases (mean age at primary 44 ± 14 yrs vs 58 ± 14, p = 0.02; mean age at LM 50 ± 15 vs 63 ± 14, p = 0.04) (Table 1). On CT scan, a single pulmonary nodule was observed in 18 cases (72%) while multiple nodules (n = 2–4) were observed in the remaining 7 cases (28%). All patients were surgically treated with curative intent and all of them were free of extra-pulmonary disease and regional lymph node involvement. Lung wedge resection was the commonest surgical procedure (14 cases) whereas lobectomy was performed in 11 patients. Median Breslow thickness (available for 21 patients) was 2.3 mm (range 0.75–5 mm) and the trunk was the most frequent primary location (trunk 72%, arm/leg 20%, other sites 8%) (p = 0.01). The median metastasis size was 1.6 cm (range 0.5–7 cm) (Table 1).
Table 1: Patient demographics and clinical characteristics of primary melanoma and of MLMs
Parameter |
All |
BRAF |
NRAS |
WT/WT |
2-Group |
3-Group |
Gender |
||||||
Female |
10 (40%) |
5 (42%) |
2 (43%) |
3 (36%) |
||
Male |
15 (60%) |
8 (58%) |
3 (57%) |
4 (64%) |
||
Age at primary tumor (yrs) |
||||||
Mean ± ds |
52 ± 16 |
50 ± 16 |
55 ± 22 |
55 ± 12 |
||
Median |
53 |
47 |
62 |
58 |
||
Range |
18–76 |
25–71 |
18–76 |
35–70 |
||
≥60 yrs |
9 |
4 |
3 |
2 |
||
<60 yrs |
16 |
9 |
2 |
5 |
||
Female (mean ± ds) |
44 ± 14 |
41 ± 9 |
40 ± 31 |
51 ± 14 |
0.02i |
|
Male (mean ± ds) |
58 ± 14 |
56 ± 17 |
65 ± 12 |
58 ± 12 |
||
Age at MLM (yrs) |
||||||
Mean ± ds |
58 ± 16 |
56 ± 16 |
59 ± 20 |
59 ± 11 |
||
median |
61 |
54 |
67 |
61 |
||
range |
23–80 |
32–76 |
23–80 |
42–73 |
||
≥60 yrs |
13 |
6 |
3 |
4 |
||
<60 yrs |
12 |
7 |
2 |
3 |
||
Female (mean ± ds) |
50 ± 15 |
48 ± 11 |
45 ± 31 |
57 ± 11 |
0.04i |
|
Male (mean ± ds) |
63 ± 14 |
62 ± 17 |
68 ± 13 |
61 ± 12 |
||
Number of LMM |
||||||
Single lesions |
18 (72%) |
9 (69%) |
5 (100%) |
4 (57%) |
||
Multiple lesions (n = 2–4) |
7 (28%) |
4 (31%) |
- |
3 (43%) |
||
Surgical procedure |
||||||
Lung wedge resection |
14 (56%) |
7 (54%) |
3 (60%) |
4 (57%) |
||
Lobectomy |
11 (44%) |
6 (46%) |
2 (40%) |
3 (43%) |
||
Primary site |
||||||
Trunk |
18 (72%) |
8 (62%) |
4 (80%) |
6(86%) |
0.01ii |
|
Arm/Leg |
5 (20%) |
3 (23%) |
1 (20%) |
- |
||
Other |
2 (8%) |
2 (15%) |
- |
1(14%) |
||
Breslow Thickness (mm) |
||||||
Assessed |
21 |
11 |
5 |
5 |
||
Not Assessed |
4 |
2 |
- |
2 |
||
Mean ± ds |
2.3 ± 0.9 |
2.3 ± 0.1 |
2.1 ± 0.5 |
2.7 ± 0.6 |
0.1iii |
0.5 |
Median |
2.3 |
2.3 |
2.1 |
2.4 |
||
Range |
0.75–5 |
0.75–5 |
1.5–2.8 |
2.3–3.2 |
||
Size of MLM (cm) |
||||||
Mean ±ds |
2.3 ± 1.8 |
1.9 ± 1.1 |
3.3 ± 2.2 |
2.2 ± 2.2 |
0.2 |
|
Median |
1.6 |
2 |
4.2 |
1.2 |
||
Range |
0.5–7 |
0.5–4.5 |
0.8–5.5 |
0.8–7 |
||
<1.6 cm (%) |
13 (52%) |
6(46%) |
2 (40%) |
5 (71%) |
||
mean |
1.0 ± 03 |
0.9 ± 0.3 |
0.9 ± 0.1 |
1.0 ± 0.2 |
||
1.6cm (%) |
12 (48%) |
7(54%) |
3(60%) |
2 (29%) |
||
mean |
3.6 ± 1.8 |
2.7 ± 0.9 |
4.9 ± 0.7 |
3.9 ± 2.8 |
0.006iv |
0.03i |
iFemale vs Male (All)
iiTrunk vs Arm/Leg vs Other (All);
iiiNRAS vs WT
ivBRAF vs NRAS
vBRAF vs NRAS vs WT
BRAF/NRAS/CKIT mutation frequencies
BRAF was altered in 13 lung metastases (52%) with all exhibiting a BRAFV600E mutation type (Table 2). NRAS mutations were detected in 5 specimens (20%), 4 NRASQ61R and 1 NRASQ61K, and were mutually exclusive with BRAF aberrations. A total of 7 lesions (28%) were wild type for both oncogenes (WT/WT). C-KIT mutations were not found (Table 2).
Table 2: BRAF/NRAS/CKIT mutation frequency
Molecular pattern |
N° cases |
(%) |
BRAF mutation |
13 |
52 |
V600E |
13 |
100 |
NRAS mutation |
5 |
20 |
Q61R |
4 |
80 |
Q61K |
1 |
20 |
WT/WT |
7 |
28 |
CKIT mutation |
0 |
0 |
EGFR gene and chromosome 7 abnormalities
FISH analysis was carried out for 17 of 25 specimens. Tumors were classified into 3 groups as described in the methods section. A normal signal was observed in 9 lesions (53%) while aberrant ones were detected in 8 cases (47%) (Table 3, Figure 1). In particular, 3 of 17 cases showed chromosome 7 copy-number-gain (CNG) (17.5%) with a normal EGFR/cep7 ratio, while EGFR specific copy number alterations were recorded in 5 of 17 cases (29, 5%), 3 with EGFR amplification (17.5%) and 2 exhibiting a deletion of the EGFR signal (12%). The three cases with specific EGFR amplification were all BRAF mutated, therefore, concurrent BRAF and EGFR abnormalities were present in 3 of 13 BRAF positive cases (23%). No correlation was observed between NRAS mutation and EGFR copy number alteration (Table 3).
Table 3: FISH analysis
FISH signal |
n (%) |
Normal EGFR/Chr 7 ratio |
Aberrant EGFR/Chr 7 ratio |
||
Disomy n (%) |
CNG/Polysomy n (%) |
Amplification n (%) |
Deletion n (%) |
||
Normal |
9 (53) |
9 (100) |
- |
- |
- |
BRAF |
4 (44) |
4 (100) |
- |
- |
- |
NRAS |
1 (11) |
1(100) |
- |
- |
- |
WT/WT |
4 (44) |
4(100) |
- |
- |
- |
Aberrant |
8 (47) |
- |
3 (37.5) |
3 (37.5) |
2 (25) |
BRAF |
5 (60) |
- |
1(20) |
3 (44.5) |
1 (11) |
NRAS |
1(12.5) |
- |
1(100) |
- |
- |
WT/WT |
2 (25) |
- |
1 (25) |
- |
1 (25) |
tot |
17 |
9 (53) |
3(17.5) |
3(17.5) |
2 (12) |
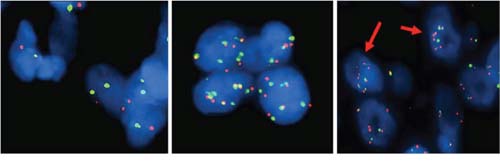
Figure 1: FISH analysis of Chromosome 7 and EGFR copy number alterations. EGFR gene-specific probe was labeled with Spectrum Orange (appearing as red signals) and chromosome 7 centromeric probe (Cep7) was labeled with Spectrum Green (appearing as green signals), cell nuclei were stained with blue fluorescent DAPI. Cells with chromosome 7 disomy (left), chromosome 7 polysomy (middle) and EGFR copy number amplification (right, red arrows). Original magnification X100.
Patient demographics and pathological characteristics based on BRAF, NRAS and EGFR gene alterations
The cohort was divided into three distinct subgroups based on BRAF and NRAS mutation status: BRAF mutated (from now on referred to as BRAF), NRAS mutated (from now on NRAS) and double wild-type (from now on WT/WT). BRAF patients were slightly younger than NRAS and WT/WT; mean age at primary melanoma and at LM was respectively 50 ± 16 and 56 ± 16 years for BRAF, 55 ± 22 and 59 ± 20 years for NRAS and 55 ± 12 and 59 ± 11 years in WT/WT patients (Table 1). NRAS lesions tended to be larger (mean diameter 3.3 ± 2.2 cm) compared with BRAF (mean diameter 1.9 ± 1.1 cm, p = 0.2) but not with respect to WT patients (2.2 ± 2.2 cm; p = 0.4). At a diameter cut-off of 1.6 cm, corresponding to the median size of the lesions, the difference among NRAS a BRAF became statistically significant (p = 0.006) (Table 1). Gender distribution, age, site and Breslow thickness of primary melanoma was not statistically associated with BRAF or NRAS mutational status. EGFR copy number alterations were not associated with any patient demographics or pathological characteristics.
Survival analysis
Tumor mutation status was correlated with time to lung metastasis (TLM), overall survival (OS) from primary melanoma as well as relapse-free survival (RFS) and OS after metastasis resection. The median TLM for the 25 pts cohort was 48 months (range 24–192 months) (Table 4). After metastasectomy, 13 of 25 patients (52%) showed recurrence within 60 months from surgery and the median relapse free-survival was 40 months (range 6–156). Median OS after metastasectomy was 52 months (range 12–144) with a 96%, 68% and 28% respective 1, 3 and 5 years survival rate. Median OS from primary melanoma was 108 months (range 60–240) (Table 4).
Table 4: Survival analysis
Parameter |
All |
BRAF |
NRAS |
WT/WT |
NRAS-WT |
2-Group |
3-Group |
TLM (mo) |
|||||||
Mean |
64 |
76 |
48 |
53 |
68 |
0.4 |
|
Median |
48 |
48 |
48 |
48 |
48 |
||
Range |
24–192 |
24–192 |
24–60 |
24–108 |
24–192 |
||
Rate LMFI ≤ 5yrs |
72% |
61% |
100% |
71% |
65% |
||
Rate LMFI > 5yrs |
40% |
29% |
0% |
19% |
35% |
||
RFS (mo) |
|||||||
Mean |
49 |
48 |
30 |
64 |
53 |
i |
0.3 |
Median |
40 |
36 |
24 |
48 |
42.5 |
||
Range |
6–156 |
6–156 |
10–52 |
30–132 |
6–156 |
||
OS from LM (mo) |
|||||||
Mean |
62 |
63 |
37 |
77 |
68 |
ii |
0.1 |
Median |
52 |
48 |
36 |
70 |
60 |
||
Range |
12–168 |
12–156 |
24–52 |
24–144 |
12–168 |
||
OS from LM Rate |
|||||||
1 yrs |
96% |
92% |
100% |
100% |
95% |
||
3 yrs |
68% |
69% |
40% |
86% |
75% |
||
5 yrs |
28% |
31% |
0% |
29% |
30% |
||
OS from primary (mo) |
|||||||
Mean |
125 |
138 |
84 |
130 |
136 |
iii |
0.2 |
Median |
108 |
120 |
84 |
96 |
120 |
||
Range |
60–240 |
96–240 |
60–96 |
72–240 |
72–240 |
||
TLM: time to lung metastasis; RFS: relapse free-survival; OS: overall survival; NRAS-WT: BRAF plus WT/WT; mo=month
*(i) NRAS vs WT/WT p = 0.1; NRAS vs NRAS-WT p = 0.08
(ii) BRAF vs NRAS p = 0.09; NRAS vs WT/WT p = 0.057; NRAS vs NRAS-WT p = 0.027
(iii) BRAF vs NRAS p = 0.0018; NRAS vs NRAS-WT p = 0.0008
**BRAF vs NRAS vs WT/WT
No significant differences in terms of relapse-free-survival after primary melanoma (TLM) or after metastasectomy (RFS) were recorded among the NRAS, BRAF and WT/WT subgroups respect to each other (Table 4, Figure 2A and 2B). Then, we evaluated NRAS-mutated versus NRAS-WT (BRAF-mutated and WT/WT combined) and found a borderline significance in mean RFS after metastasectomy (NRAS-mutated versus NRAS-WT; 30 vs 53 months, p = 0.08) (Table 4). Same trend was evident in terms of a shorter OS from metastasectomy in NRAS patients with respect to NRAS-WT (BRAF plus WT/WT combined) (p = 0.09) that became significant when OS from the primary tumor was considered (p = 0.017) (Table 4; Figure 3A and 3B). Moreover, the multivariate Cox regression analysis, revealed that NRAS mutation was the only predictive factor of adverse survival (p = 0.039, OR = 5.5 (1.1–27.6). No difference in survival was observed based on specific EGFR copy number changes.
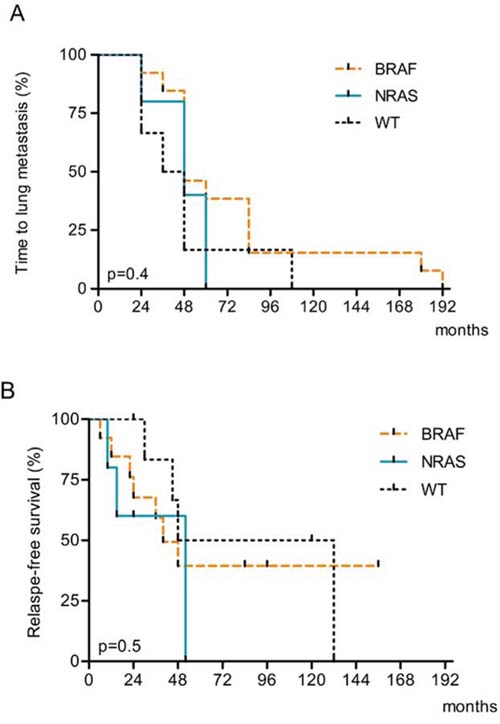
Figure 2: Kaplan-Meier estimates of A. Time to lung metastasis (TLM) and B. Relapse-free survival (RFS) after lung metastasis resection according to BRAF and NRAS mutation status.
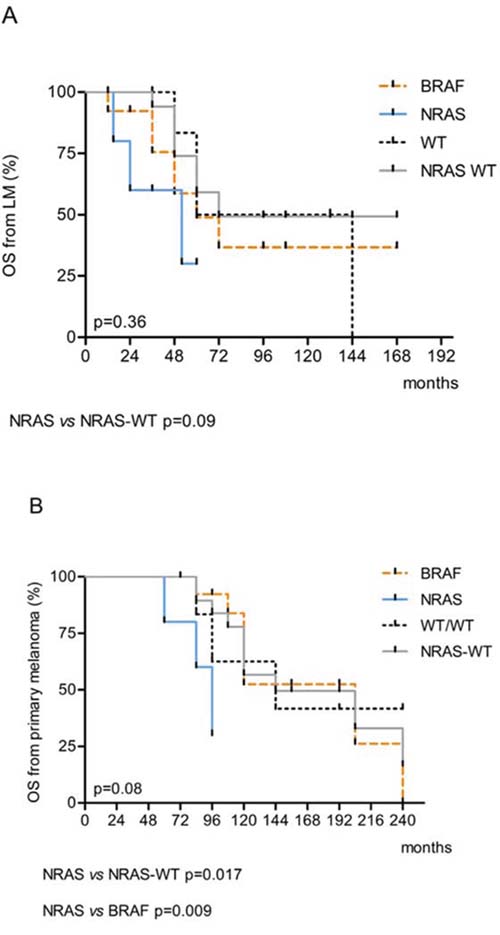
Figure 3: Kaplan-Meier estimates of A. Overall Survival (OS) from lung metastasis and B. OS from primary tumor according to BRAF and NRAS mutations.
DISCUSSION
In this study we performed the molecular appraisal of BRAF, NRAS, CKIT and EGFR genes altogether in a highly selected cohort of melanoma patients with lung metastases, that had been surgically treated with curative intent. No systemic or radiation therapies had been administered to the entire cohort before or after surgery. The frequency of BRAF (52%) and NRAS (20%) mutations was in line with the rates reported for other visceral melanoma metastatic sites [29–34] while no CKIT gene alterations were found. This was in accord with the main derivation of our cohort from primary melanoma of the trunk where only low frequency (2%) of CKIT mutations have been previously documented [22–23]. Also in agreement with previous studies, with respect to age at onset of both, primary and metastasis, females were significantly younger than males regardless of BRAF and NRAS mutation status [30–34]. Breslow thickness of primary melanoma was not significantly associated with BRAF or NRAS gene alterations.
Chromosome 7/EGFR copy number alterations have been studied in primary and metastatic melanoma and speculated to play a role in disease progression [35]. In line with these data our FISH analysis revealed a 47% overall rate of EGFR/chromosome 7 aberrations. Intriguingly, 3 of the 13 BRAF positive cases (23%) displayed a concomitant EGFR copy number amplification. This observation can be relevant in the light of the following data that link EGFR with biologic-therapy resistance: (i) ectopic expression of EGFR in melanoma cells is sufficient to cause resistance to PLX4032 (vemurafenib), a specific small-molecule BRAF inhibitor [36]; (ii) a rapid feedback activation of EGFR can support continued proliferation in the presence of BRAF (V600E) inhibition [36]; (iii) EGFR inhibition blocked proliferation and invasion of BRAFV600E resistant melanoma [37]; (iv) demethylation of EGFR regulatory DNA elements has been observed in cutaneous melanomas resistant to BRAF inhibition [38] and finally (v) a melanoma subtype with intrinsic resistance to BRAF inhibition has been identified that was associated with differential EGFR and ERBB3 expression [39]. With respect to the above observations one could therefore speculate that EGFR amplification might explain, at least in part, the lack of response observed in approximately 20% of patients with melanoma harboring BRAFV600E mutations.
Patients with pulmonary metastases show a better survival rate than individuals affected by metastases to other visceral sites [40] and a growing number of studies have shown that pulmonary metastectomy dramatically improves survival [8, 41–44]. Accordingly, in our study, the survival rate registered for the all cohort of LMM patients was excellent with a median relapse-free and overall survival after surgery of respectively 40 and 52 months that was slightly higher than the ones reported before [8, 45]. Our results probably reflects the accurate selection of patients eligible for surgical excision and/or an intrinsic more favorable outcome linked to a less aggressive disease biology. In this context, our survival analyses intriguingly indicated a trend to a worse prognosis for NRAS mutated patients. In particular, compared with BRAF and WT individuals, all NRAS patients developed lung metastases within 5 years from the primary, tended to carry pulmonary lesions of larger size and showed shorter survival time after lung metastasis resection (Table 4). In addition, the mean OS evaluation from primary melanoma to death or last follow-up, revealed significant differences among NRAS and BRAF patients, as well as among NRAS and their wild-type counterparts and, finally, the multivariate Cox regression analysis showed that NRAS mutation was the only predictive factor of shorter survival from primary melanoma. An adverse prognostic role of NRAS mutations has been already speculated in previous studies. Patients with NRAS mutant melanomas had thicker tumors at presentation and these tumors had greater rates of mitosis than BRAF mutant and WT melanoma [30, 46, 47]. Shorter survival from initial melanoma diagnosis for NRAS patients compared to WT has been reported [30, 46, 47], and NRAS mutation status has been identified as an independent predictor of shorter survival after a diagnosis of stage IV melanoma [46]. From a therapeutic standpoint novel exciting options exist for improving the overall survival of NRAS-driven melanomas. In particular, MEK162, a MEK1/2-inhibitor, was recently proposed as the first type of target therapy to show activity in patients with NRAS-dependent melanoma and, in addition, individuals with similar molecular characteristics were found to be uniquely sensitive to CMET inhibition, thus providing a new rationale for therapeutic targeting of CMET in this patient cohort [48, 49]. Moreover, it was recently reported that a local administration of low-dose IL-2 through inhalation (lh-IL-2) might offer an effective and safe treatment option for lung metastases in melanoma patients and lh-IL-2 may have a prophylactic potential to prevent recurrence to the lung after pulmonary melanoma metastasectomy [50]. Intriguingly, in an independent study, NRAS status was suggested as a new candidate biomarker for selecting melanoma patients for high-dose interleukin-2 treatment (HD IL-2) [51].
This work has one major limitation consisting in the small number of patients enrolled during a 12 years period. This small accrual was mainly due to the extreme rarity of application of the intervention of surgery for the treatment of melanoma lung metastases, and the strict parameters utilized to select the population under molecular study including the lung metastasis as the first manifestation of metastatic disease and the absence of systemic and radiation treatment. Notwithstanding this weakness, we believe that the present investigation, based on a small but carefully selected cohort of pulmonary metastatic melanoma, represents a useful reference and provides several clues concerning the lung as a specific site of melanoma disease that may be highly relevant to identify melanoma patients with diverse prognostic features and therapeutic options.
MATERIALS AND METHODS
Tumor samples and patients
The study cohort consisted of 25 pulmonary metastasis specimens obtained by lobectomy (11) or wedge resection (14) from 25 patients affected by melanoma who underwent metastasectomy with curative intent between years 2000 and 2012 at the San Camillo-Forlanini Hospitals. All patients were free of extra-pulmonary disease and lymph node involvement and, in all cases, pulmonary metastasis was the first sign of metastatic disease. Informed consent was acquired from all patients and the study was approved by the Local Institutional Ethics Committee. For all individuals information concerning the site of primary tumor and date of first ever diagnosis was acquired from medical records. Breslow thickness of the primary tumor was available for 21 cases. No patient underwent systemic or radiation therapy before or after surgery.
DNA extraction and mutation analysis
4 μm thick sections were lightly stained with haematoxylin and manually microdissected by an expert pathologist (LM). Genomic DNA was extracted using the QIAamp DNA FFPE Tissue kit (Qiagen, Germany) following the manufacturer's protocol and subjected to PCR amplification of BRAF (exon 15), NRAS (exon 3) and CKIT (exon 9, 11, 13 and 17). Mutations were tested by Sanger sequencing using a BigDye Terminator Cycle Sequencing Kit (Applied Biosystems, Austin TX, USA) and an ABI PRISM 310 Genetic Analyzer (Applied Biosystems). Sequencing reactions were performed in forward and reverse orientation.
Fluorescence in situ hybridization analysis
Fluorescence in situ hybridization was executed by dual colour FISH assay using 7 centromere-(cep7, SpectrumGreen) and locus specific 7p12 (EGFR locus, SpectrumOrange) probes from Vysis (Vysis Inc. IL, USA). FISH was carried out according to the manufacturer's protocol. An average of 50–100 nuclei were screened per each sample. Categories of FISH abnormalities were defined as follows: (i) EGFR gene deletion: EGFR copy number was less than chromosome 7 centromere (cep7) in more than 15% of nuclei; (ii) Chromosome 7 copy number gain (CNG): EGFR/cep7 ratio =1, but the cep7 signals were > 2 per nucleus in more than 15% of nuclei; (iii) EGFR amplification: EGFR/cep7 ratio > 2.2 in more than 15% of screened nuclei.
Statistical analysis
Statistical analyses were performed using STATISTICA 7 software (Stat Soft, Inc., Tulsa, Okla., USA). Continuous variables were reported as mean, median, and standard deviation while categorical variables as number (n) and percentage (%). For all patients, clinical and pathological features were tested for association with BRAF and NRAS mutation status using t-test and non parametric Mann Whitney or Kruskal Wallis tests for continuous variable and Fisher’s exact test for categorical variables. Time to lung metastasis (TLM) was defined as the time from primary melanoma to diagnosis of lung metastasis, while relapse-free survival (RFS) was estimated from metastasectomy to first evidence of relapse. Analysis was also made comparing overall survival (OS) from the date of diagnosis of primary melanoma or lung metastasis to death/last follow-up. Survival analyses were performed with the Kaplan-Meier survival curve and compared using the Log-Rank test (Graphpad 5 software). The Cox proportional hazards model was used to determine the significance of variables in predicting adverse factor for survival introducing in the model patient’s age, gender, BRAF and NRAS mutations. All statistical tests were performed 2-sided and a p-value of < 0.05 was considered statistically significant.
ACKNOWLEDGMENTS
The cost of publication of this work was covered by the Ca-The-Dra (Cancer Therapy Drawing) project based at Villa Margherita Hospital, Rome, Italy
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.
REFERENCES
1. American Cancer Society. Cancer facts & figures 2014. Available at: http://www.cancer.org/acs/groups/content/@research/documents/webcontent/acspc-042151.pdf. Accessed March 7, 2014.
2. Tsao H, Atkins MB, Sober AJ. Management of cutaneous melanoma. N Engl J Med. 2004; 351:998–1012.
3. Filosso PL, Sandri A, Ruffini E, Lausi PO, Bruna MC, Oliaro A. Surgical Treatment of Pulmonary Metastases from Melanoma: Emerging Options, Melanoma in the Clinic. Diagnosis, Management and Complications of Malignancy, Prof. Mandi Murph(Ed.), 2011, ISBN 978-953-307-571-6.
4. Garbe C, Eigentler TK, Keilholz U, Hauschild A, Kirkwood JM. Systematic review of medical treatment in melanoma: current status and future prospects. Oncologist. 2011; 16:5–24.
5. Mansfield AS, Markovic SN. Novel therapeutics for the treatment of metastatic melanoma. Future Oncol. 2009; 5:543–57.
6. Olszanski AJ. Current and future roles of targeted therapy and immunotherapy in advanced melanoma. J Manag Care Spec Pharm. 2014; 20:346–56.
7. Oliaro A, Filosso PL, Bruna MC, Mossetti C, Ruffini E. Pulmonary metastasectomy for melanoma. J Thorac Oncol. 2010; 5:S187–91.
8. Neumann HB, Patel A, Hanlon C, Wolchok JD, Houghton AN, Coit DG: Stage IV melanoma and pulmonary metastases: factors predictive of survival. Ann. Surg. Oncol. 2007; 14:2847–2853.
9. Gray-Schopfer V, Wellbrock C, Marais R. Melanoma biology and new targeted therapy. Nature. 2007; 445:851–857.
10. Hodi FS, O’Day SJ, McDermott DF, et al. Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med. 2010; 363:711–23.
11. Cohen C, Zavala-Pompa A, Sequeira JH, Shoji M, Sexton DG, Cotsonis G, Cerimele F, Govindarajan B, Macaron N, Arbiser JL. Mitogen-actived protein kinase activation is an early event in melanoma progression. Clin Cancer Res. 2002; 8:3728–33.
12. Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, et al. Mutations of the BRAF gene in human cancer. Nature 2002; 417:949–54.
13. Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, et al. BRIM-3 Study Group. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med. 2011; 364:2507–16.
14. Hauschild A, Grob JJ, Demidov LV, Jouary T, Gutzmer R, Millward M, Rutkowski P, Blank CU, Miller WH Jr, Kaempgen E, Martín-Algarra S, Karaszewska B, Mauch C, et al. Dabrafenib in BRAF-mutated metastatic melanoma: a multicentre, open-label, phase 3 randomised controlled trial. Lancet. 2012; 380:358–65.
15. Falchook GS, Lewis KD, Infante JR, Gordon MS, Vogelzang NJ, DeMarini DJ, Sun P, Moy C, Szabo SA, Roadcap LT, Peddareddigari VG, Lebowitz PF, Le NT, et al. Activity of the oral MEK inhibitor trametinib in patients with advanced melanoma: a phase 1 dose-escalation trial. Lancet Oncol. 2012; 13:782–9.
16. Menzies AM, Long GV. Dabrafenib and Trametinib, Alone and in Combination for BRAF Mutant Metastatic Melanoma. Clin Cancer Res. 2014; 20:2035–43.
17. Robert C, Thomas L, Bondarenko I, O'Day S, Weber J, Garbe C, Lebbe C, Baurain JF, Testori A, Grob JJ, Davidson N, Richards J, Maio M, et al. Ipilimumab plus dacarbazine for previously untreated metastatic melanoma. N Engl J Med. 2011; 364:2517–26.
18. Curtin JA, Fridlyand J, Kageshita T, Patel HN, Busam KJ, Kutzner H, Cho KH, Aiba S, Bröcker EB, LeBoit PE, Pinkel D, Bastian BC. Distinct sets of genetic alterations in melanoma. N Engl J Med. 2005; 353:2135–47.
19. Palmieri G, Capone M, Ascierto ML, Gentilcore G, Stroncek DF, Casula M, Sini MC, Palla M, Mozzillo N, Ascierto PA. Main roads to melanoma. J Transl Med. 2009; 7:86.
20. Colombino M, Capone M, Lissia A, Cossu A, Rubino C, De Giorgi V, Massi D, Fonsatti E, Staibano S, Nappi O, Pagani E, Casula M, Manca A, et al. BRAF/NRAS mutation frequencies among primary tumors and metastases in patients with melanoma. J Clin Oncol. 2012; 30:2522–9.
21. Ascierto PA, Schadendorf D, Berking C, Agarwala SS, van Herpen CM, Queirolo P, Blank CU, Hauschild A, Beck JT, St-Pierre A, Niazi F, Wandel S, Peters M, et al. MEK162 for patients with advanced melanoma harbouring NRAS or Val600 BRAF mutations: a non-randomised, open-label phase 2 study. Lancet Oncol. 2013; 14:249–56.
22. Smalley KSM, Vernon K, Weber S, Weber J. Review: c-Kit signaling as the driving oncogenic event in sub-groups of melanomas. histology and histopathology cellular and molecular biology. J Pathol. 2009; 29:643–5.
23. Beadling C, Jacobson-Dunlop E, Hodi FS, Le C, Warrick A, Patterson J, Town A, Harlow A, Cruz F 3rd, Azar S, Rubin BP, Muller S, West R, et al. KIT gene mutations and copy number in melanoma subtypes. Clin Cancer Res. 2008; 14:6821–8.
24. Kluger HM, Dudek AZ, McCann C, Ritacco J, Southard N, Jilaveanu LB, Molinaro A, Sznol M. A Phase 2 Trial of Dasatinib in Advanced Melanoma. Cancer. 2011; 117:2202–8.
25. Hodi FS, Friedlander P, Corless CL, Heinrich MC, Mac Rae S, Kruse A, Jagannathan J, Van den Abbeele AD, Velazquez EF, Demetri GD, Fisher DE. Major response to imatinib mesylate in KIT-mutated melanoma. J Clin Oncol. 2008; 26:2046–51.
26. Lyle M, Long GV. Diagnosis and Treatment of KIT-Mutant Metastatic Melanoma. J Clin Oncol. 2013; 31:3176–81.
27. Boone B, Jacobs K, Ferdinande L, Taildeman J, Lambert J, Peeters M, Bracke M, Pauwels P, Brochez L. EGFR in melanoma: clinical significance and potential therapeutic target. J Cutan Pathol. 2011; 38:492–502.
28. Rákosy Z, Vízkeleti L, Ecsedi S, Vokó Z, Bégány A, Barok M, Krekk Z, Gallai M, Szentirmay Z, Adány R, Balázs M. EGFR gene copy number alterations in primary cutaneous malignant melanomas are associated with poor prognosis. Int J Cancer. 2007; 121:1729–37.
29. Greaves WO, Verma S, Patel KP, Davies MA, Barkoh BA, Galbincea JM, Yao H, Lazar AJ, Aldape KD, Medeiros LJ, Luthra R. Frequency and Spectrum of BRAF Mutations in a Retrospective, Single-Institution Study of 1112 Cases of Melanoma. J Mol Diagn. 2013; 15:220–6.
30. Devitt B, Liu W, Salemi R, Wolfe R, Kelly J, Tzen CY, Dobrovic A, McArthur G. Clinical outcome and pathological features associated with NRAS mutation in cutaneous melanoma. Pigment Cell Melanoma Res. 2011; 24:666–672.
31. Carlino MS, Haydu LE, Kakavand H, Menzies AM, Hamilton AL, Yu B, Ng CC, Cooper WA, Thompson JF, Kefford RF, O’Toole SA, Scolyer RA, Long GV. Correlation of BRAF and NRAS mutation status with outcome, site of distant metastasis and response to chemotherapy in metastatic melanoma. Br J Cancer. 2014; 111:292–299.
32. Edlundh-Rose E, Egyházi S, Omholt K, Månsson-Brahme E, Platz A, Hansson J, Lundeberg J. NRAS and BRAF mutations in melanoma tumours in relation to clinical characteristics: a study based on mutation screening by pyrosequencing. Melanoma Res. 2006; 16:471–8.
33. Ugurel S, Thirumaran RK, Bloethner S, Gast A, Sucker A, Mueller-Berghaus J, Rittgen W, Hemminki K, Becker JC, Kumar R, Schadendorf D. B-RAF and N-RAS mutations are preserved during short time in vitro propagation and differentially impact prognosis. PLoS One. 2007; 2:e236.
34. El-Osta H, Falchook G, Tsimberidou A, Hong D, Naing A, Kim K, Wen S, Janku F, Kurzrock R. BRAF Mutations in Advanced Cancers: Clinical Characteristics and Outcomes. PLoS One. 2011; 6:e25806.
35. Udart M, Utikal J, Krahn GM, Peter RU. Chromosome 7 aneusomy. A marker for metastatic melanoma? Expression of the epidermal growth factor receptor gene and chromosome 7 aneusomy in nevi, primary malignant melanomas and metastases. Neoplasia. 2001; 3:245–54.
36. Prahallad A, Sun C, Huang S, Di Nicolantonio F, Salazar R, Zecchin D, Beijersbergen RL, Bardelli A, Bernards R. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature. 2012; 483:100–103.
37. Girotti MR, Pedersen M, Sanchez-Laorden B, Viros A, Turajlic S, Niculescu-Duvaz D, Zambon A, Sinclair J, Hayes A, Gore M, Lorigan P, Springer C, Larkin J, et al. Inhibiting EGF receptor or SRC family kinase signaling overcomes BRAF inhibitor resistance in melanoma. Cancer Discov. 2013; 3:158–67.
38. Wang J, Huang SK, Marzese DM, Hsu SC, Kawas NP, Chong KK, Long GV, Menzies AM, Scolyer RA, Izraely S, Sagi-Assif O, Witz IP, Hoon DS. Epigenetic changes of EGFR play an important role in BRAF inhibitor resistant cutaneous melanomas. J Invest Dermatol. 2015; 135:532–41.
39. Dugo M, Nicolini G, Tragni G, Bersani I, Tomassetti A, Colonna V, Del Vecchio M, De Braud F, Canevari S, Anichini A, Sensi M. A melanoma subtype with intrinsic resistance to BRAF inhibition identified by receptor tyrosone kinases gene-driven classification. Oncotarget. 2015; 6:5118–5133.
40. Balch CM, Gershenwald JE, Soong SJ, Thompson JF, Atkins MB, Byrd DR, Buzaid AC, Cochran AJ, Coit DG, Ding S, Eggermont AM, Flaherty KT, Gimotty PA, et al. Final version of 2009 AJCC melanoma staging and classification. J Clin Oncol. 2009; 27:6199–6206.
41. Petersen RP, Hanish SI, Haney JC, Miller CC 3rd, Burfeind WR Jr, Tyler DS, Seigler HF, Wolfe W, D’Amico TA, Harpole DH Jr. Improved survival with pulmonary metastasectomy: an analysis of 1720 patients with pulmonary metastatic melanoma. J Thorac Cardiovasc Surg. 2007; 133:104–110.
42. Schuhan C, Muley T, Dienemann H, Pfannschmidt J. Survival after pulmonary metastasectomy in patients with malignant melanoma. Thorac Cardiovasc Surg. 2011; 59:158–162.
43. Younes R, Abrao FC, Gross J. Pulmonary metastasectomy for malignant melanoma: prognostic factors for long-term survival. Melanoma Res. 2013; 23:307–311.
44. Chua TC, Scolyer RA, Kennedy CW, Yan TD, McCaughan BC, Thompson JF. Surgical management of melanoma lung metastasis: an analysis of survival outcomes in 292 consecutive patients. Ann Surg Oncol. 2012; 19:1774–1781.
45. Rolland Gyulai, Zsolt Kádár, Zsuzsanna Lengyel. Surgery and Staging of Melanoma, Melanoma - Current Clinical Management and Future Therapeutics, Prof Mandi Murph (Ed.) 2015, ISBN: 978-953-51-2036-0.
46. Jakob JA, Bassett RL Jr, Ng CS, Curry JL, Joseph RW, Alvarado GC, Rohlfs ML, Richard J, Gershenwald JE, Kim KB, Lazar AJ, Hwu P, Davies MA. NRAS mutation status is an independent prognostic factor in metastatic melanoma. Cancer. 2012; 118:4014–23.
47. Ellerhorst JA, Greene VR, Ekmekcioglu S, Warneke CL, Johnson MM, Cooke CP, Wang LE, Prieto VG, Gershenwald JE, Wei Q, Grimm EA. Clinical Correlates of NRAS and BRAF Mutations in Primary Human Melanoma. Clin Cancer Res. 2011; 17:229–35.
48. Chattopadhyay C, Ellerhorst JA, Ekmekcioglu S, Greene VR, Davies MA, Grimm EA. Association of activated c-Met with NRAS-mutated human melanomas. Int. J. Cancer. 2012; 131:E56–E65.
49. Thumar J, Shahbazian D, Aziz SA, Jilaveanu LB, Kluger HM. MEK targeting in N-RAS mutated metastatic melanoma. Mol Cancer. 2014; 4:13–45.
50. Posch C, Weihsengruber F, Bartsch K, Feichtenschlager V, Sanlorenzo M, Vujic I, Monshi B, Ortiz-Urda S, Rappersberger K. Low-dose inhalation of interleukin-2 bio-chemotherapy for the treatment of pulmonary metastases in melanoma patients. Br J Cancer. 2014; 110:1427–32.
51. Joseph RW, Sullivan RJ, Harrell R, Stemke-Hale K, Panka D, Manoukian G, Percy A, Bassett RL, Ng CS, Radvanyi L, Hwu P, Atkins MB, Davies MA. Correlation of NRAS mutations with clinical response to high-dose IL-2 in patients with advanced melanoma. J Immunother. 2012; 35:66–72.

