Introduction
Ubiquitination is one of the most important post-translation modifications (PTMs) responsible for regulating the stability and activity of modified proteins. For the ubiquitin molecule to be attached to its target protein it initially has to be activated by an E1 enzyme during an ATP-dependent reaction, which is followed by conjugation of ubiquitin by an E2 class enzyme allowing E3 ubiquitin ligase to ubiquitinate target proteins directly or indirectly [1], [2], [3] .
The consequences of ubiquitination depend on the type of chains formed during the process [4]. Proteins can be mono-, multi-mono- or poly-ubiquitinated. Ubiquitin contains seven lysine (K) residues with poly-ubiquitin chains linked through the K6, K11, K27, K29, K33, K48 and K63 residues. Mono-ubiquitination and K63 poly-ubiquitination have been linked to regulating protein activity [5]. K6, K11, K29 and K48 poly-ubiquitin chains control protein stability with K6 and K48 chains targeting proteins for proteosomal degradation, K11 is involved in endoplasmic reticulum mediated degradation pathways and control of cell cycle progression and K29 in regulating lysosomal degradation of proteins [6]. The role of K27 and K33 poly-ubiquitination is less understood however, they have been linked to innate immunity and immune responses.
Deubiquitinating enzymes (DUBs)
Just as every action provokes a reaction all of the major post-translational modifications can be reversed. Enzymes that reverse PTMs are equally important to normal homeostasis as those that initially modify proteins. Protein ubiquitination by the E3 ligases can be reversed by deubiquitinating enzymes [7]. DUBs can be divided into five families; ubiquitin carboxy - terminal hydrolases (UCH), ubiquitin specific proteases (Usp), Otubain/Ovarian tumour - domain containing proteins (OTU), Machado - Joseph Domain (Josephin domain) - containing proteins (MJD) and Jab1/MPN domain associated metalloisopeptidase domain proteins. UCH, Usp, OUT and MJD proteins are all thiol proteases containing an active site cysteine which serves as a nucleophile facilitating the attack on lysine-glycine isopeptide bonds of ubiquitinated proteins. Jab1/MPN domain associated metalloisopeptidase domain proteins differ from the other DUB classes as they utilise a JAMM zinc metalloproteinase domain to break the bond between their target proteins and ubiquitin.
With increased appreciation for the critical importance of ubiquitination in cellular processes, DUBs role in health and disease is becoming a new focus of scientific research. Recent advances in the field uncovered a growing number of DUB substrates. DUBs control the stability and activity of multiple proteins crucial in cellular proliferation and survival including p53 - the guardian of the genome [8], [9], MDM2 [10], androgen receptor (AR) [11], [12], histones [13], [14], PHLPP and PHLPPL Akt phosphatases [15], [16], [17], Notch [18], NF-κB [19], [20], β-catenin [21], [22] and many more. It is now understood that a tight balance between ubiquitination and deubiquitination is required for cellular survival underlying the equal importance of E3s and DUBs.
Epigenetics and DUBs in cancer
All stable, long-term alterations to the transcriptional potential of the cells that are not caused by direct changes to the DNA sequence itself and can be passed to daughter cells are referred to as epigenetics. These changes include DNA methylation, histone modifications and alterations to the activity of repressors and transcription factors. All of these processes result in altered transcriptome and affect cellular growth, survival and homeostasis; as such epigenetics play a crucial role in cancer development and progression [23]. DUBs can regulate all levels of epigenetic changes, by deubiquitinating proteins and changing their activity and / or stability DUBs control the levels of methylases and demethylases, histone proteins and their binding partners, repressors and transcription factors (Figure 1). DUBs can have both activating and repressing effects on gene transcription depending on their target proteins and the type of ubiquitin chain that is being removed. Additionally, DUBs can alter ubiquitin chains rather than remove them completely by for example deubiquitinating K48-linked chains and leaving the target protein in a mono-ubiquitinated state which results in altered protein fate. In this review we will focus on the role of DUBs in epigenetics and resulting cancer development. We will also review the potential therapeutic aspects of targeting DUBs to affect cellular epigenetics.
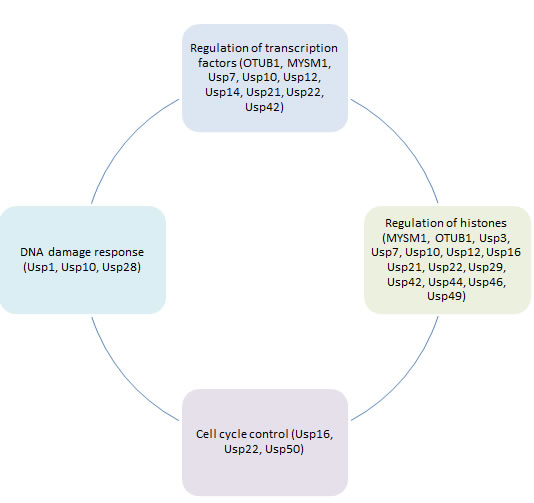
Figure 1: Role of deubiquitinating enzymes in different aspects of cancer epigenetic regulation.
Regulation of epigenetics by deubiquitination
Regulation of transcription factors
Gene expression is controlled by transcription factors that directly bind to DNA. Transcription factors act in complexes with other proteins and DUBs can affect their activity by deubiquitinating transcription factors directly or by targeting their binding partners and altering the stoichiometry of the complex. Usp12 activity towards the androgen receptor (AR) is an example of direct transcription factor deubiquitination [12]. AR is a member of the nuclear receptor superfamily and plays a key role in the transcriptional regulation of numerous genes important in the development of both normal and malignant prostate. AR deregulation is the key feature of prostate cancer (PC) development. Usp12 was reported to deubiquitinate the AR resulting in its increased protein stability and transcriptional activity [12]. As a result Usp12 promotes PC development with protein levels increased in PC patients compared to benign controls. However, AR has multiple ubiquitination sites and it is targeted by a variety of E3s, as a result AR deubiquitination doesn’t always promote transcription. Usp26 can directly bind and deubiquitinate AR, acting as a co-regulator of the AR by reversing AR activation and degradation by MDM2 ubiquitination depending on cellular context highlighting the complexity of post-translational regulation [11]. Similarly, Estrogen Receptor (ER) can also be regulated by deubiquitination, OTUB1 has been identified to target ER-alpha affecting ER target gene transcription and stabilising its protein levels on chromatin [24].
Sometimes it is not clear if DUBs control transcription factors via direct deubiquitination or if their regulation of the transcriptome is more via effects exerted on histones that cause a signalling cascade affecting the transcription factors. This is the case for Usp10, it was initially reported to directly regulate p53 and AR by deubiquitinating them and acting as a co-activator [25]. However, more recent reports indicate that the effects on AR might be indirect via Usp10 activity towards mono-ubiquitinated histone H2A.Z [14]. As a result of Usp10’s role in the regulation of histones and p53 it is frequently overexpressed in breast and brain cancer patients where its levels correlate with survival. In prostate cancer, with increasing grade of disease, the cellular localisation of Usp10 changes with expression becoming predominantly nuclear allowing for higher activity towards H2A.Z. Similarly MYSM-1 has been reported to indirectly regulate the AR. MYSM-1 can activate transcription of AR target genes via its involvement with p300 affecting histone acetylation and deubiquitination and binding of H1 to the nucleosome [26].
P53 is a crucial tumour suppressor acting as a transcription factor responsible for regulating the expression of multiple genes associated with stress responses, cellular survival, growth and homeostasis [27], [28], [29]. P53 can be directly deubiquitinated by Usp42 which reverses its ubiquitination by MDM2 [9]. Usp42 forms a direct complex with p53 and controls its activation in response to cellular stress; as a result it regulates p53-dependent transcription and cell cycle arrest. Usp7 is another DUB important for p53 activity [8]. Usp7 regulates the polycomb complex and factors associated with transcription including MDM2, p53 and FOXO. Usp7 directly deubiquitinates and stabilises p53, it is also necessary for p53 stabilisation by the tumour suppressor ING1 [30]. Additionally, independently of its deubiquitinase activity the DUB Usp7 regulates sequence specific RNA binding of the core domain of p53 thereby stimulating its transcriptional activity and expression of p21 [31]. Consequently, Usp7 inhibition was reported to inhibit cancer cell growth and increase apoptosis.
Some DUBs can control p53 stability independently of their enzymatic activity. One such example is OTUB1 which can inhibit UbcH5. UbcH5 is an MDM2 cognate ubiquitin-conjugating enzyme (E2), as a result of its inhibition p53 ubiquitination is abrogated leading to its increased stability and activity [32], [33]. Through binding to E2 enzyme Ubc13, OTUB1 also regulates the K63 ubiquitination of chromatin induced by DNA damage [34].
GATA3 serves as another example of a transcription factor regulated by deubiquitination. GATA3 is a master regulator of T helper cells (Th2) cell differentiation and function; it controls early T cell development and is one of only 3 genes mutated in over 10% of breast cancers as it plays a key role in mammary gland development [35], [36]. Recent reports demonstrate that GATA3 can be deubiquitinated by Usp21 which rescues it from proteosomal degradation and stabilises GATA3 protein levels [37]. This results in increased transcriptional activity of GATA3 and highlights the role of Usp21 in immune responses with Usp21 protein upregulated in regulatory T cells (Tregs) from asthma patients, however the role of this interaction in carcinogenesis still remains to be established.
Both canonical and non-canonical Wnt signalling pathways are also regulated by deubiquitination. Recent reports identified proteasome associated DUB Usp14 as an oncogene and a positive regulator of Wnt signalling via deubiquitination of Dishevelled (Dvl) [38]. Depletion of Usp14 attenuated downstream Wnt signalling which was further evidenced when correlation between the levels of Usp14 and β-catenin in colon tissues was observed. Usp14 was also up regulated in non-small cell lung carcinoma where high levels correlated with decreased survival and poor prognosis [39]. This was attributed to the oncogenic properties of Usp14 with silencing causing cell cycle arrest as a consequence of β-catenin degradation.
An additional level of control of transcriptional activity mediated by the nuclear receptors is provided by the SAGA (Spt-Ada-Gcn5-acetyl 1 transferase) histone acetyltransferase (HAT) complex. In addition to HAT activity, SAGA has deubiquitinating activity, which is required for the transcriptional activity of nuclear receptors with Usp22 being the active DUB subunit of SAGA that removes ubiquitin from the histones H2A and H2B [40]. As SAGA is a chromatin modifying transcription coactivator complex which regulates the expression of genes related to tumourigenicity and proliferation its levels predict treatment failure and are also used as a marker of recurrence, metastasis and resistance to therapy with levels significantly increased in cancers, including colorectal cancer [41], [42].
Regulation of histones
Mono-ubiquitination of histone H2A is one of the key histone modifications, it is associated with transcriptional repression by the polycomb group proteins and maintenance of the genome integrity. H2A is mono-ubiquitinated at K119 and it is estimated that between five and fifteen percent of the H2A fraction is mono-ubiquitinated at any one time compared to only one percent of H2B. As previously discussed, Usp10 can deubiqutinate the mono-ubiquitinated H2A.Z variant of H2A which results in increased AR activity [14]. However, H2A can also be deubiquitinated by MYSM1 (2A-DUB) [26] and UbpM (Usp16) in a way that is independent of its phosphorylation by CDK1 [43], [44]. UbpM specifically deubiquitinates histone H2A, but not H2B. This deubiquitination is required for dephosphorylation of histone H3 on S10 resulting in chromosome segregation during the mitotic entry. As a result, UbpM silencing decreases cell growth due to defects in mitosis. UbpM similarly controls Hox gene expression via regulation of H2A ubiquitination status. Ubiquitination of H2A is known to contribute to the embryonic stem cell (ESC) pluripotency by repressing lineage-specific gene expression [45]. It has been reported that deubiquitination of H2A by UbpM controls gene expression in ESCs with UbpM binding to the promoter regions of multiple genes in ESCs and regulating the H2A ubiquitination levels. It is now established that UbpM is required for ESCs to differentiate, as in its absence ubiquitinated H2A mediates repression of lineage-specific genes expression abrogating cellular differentiation [45].
Histone ubiquitination plays a vital role in both DNA damage response and repair pathways (Table 1). Increased mono-ubiquitination of H2A, H2B, H3 and H4 has been previously observed upon DNA damage. During the early DNA damage response, DNA-PK, ATM and ATR kinases phosphorylate a fraction of the H2AX variant of H2A, which is commonly referred to as γH2AX [46], [47]. RNF168 is an E3 ligase that ubiquitinates histones H2A and γH2AX during the DNA damage response [48], this ubiquitination can be reversed by Usp3 [49]. Usp3 associates with chromatin and deubiquitinates H2A at K13 and K15 and γH2AX at K118 and K119. As a result Usp3 overexpression has been shown to impair the accumulation of the BRCA1 and 53BP1 repair factors at the DNA damage sites in response to DNA damage and to counteract the activity of RNF168. Consistent with these findings, Usp3 ablation caused accumulation of DNA breaks and activation of DNA damage checkpoint pathways. This pathway can also be reversed by other DUBs, including Usp21 [50], OTUB1, Usp29 and Usp44 [51]. Both Usp44 and Usp29 deubiquitinate H2A with USP44 being recruited to the RNF168-generated ubiquitination products at double stranded break sites [51].
Ubiquitination of both H2A and H2B is equally important for maintaining cellular homeostasis and there are DUBs that specifically target histone H2B, including Usp49 [52] and Usp42 [53]. H2B ubiquitination regulates H3K4 and H3K79 methylation and impacts on the chromatin structure [52]. Usp49 in complex with RVB1 and SUG1 yeast homologues deubiquitinates H2B, this modification is required for efficient co-transcriptional splicing of a large set of exons. Silencing Usp49 induces relatively small changes in gene expression, however alterations in H2B ubiquitination levels caused by Usp49 regulate U1A and U2B association with chromatin and binding to nascent pre-mRNA. Consequently, Usp49 plays a crucial role in co-transcriptional pre-mRNA processing [52]. H2B ubiquitination can additionally be reversed by Usp7 resulting in epigenetic silencing of homeotic genes [54].
Some DUBs are not as specific and have the ability to deubiquitinate both histones H2A and H2B, for example the closely related family of DUBs comprising Usp1, Usp12 and Usp46. All three DUBs require Uaf-1 (WDR48) for their enzymatic activity [55] and additionally Usp12 and Usp46, but not Usp1, activity is further enhanced by binding to WDR20 [56]. Usp12 and Usp46 deubiquitinate both H2A and H2B and Uaf-1 is required for this reaction [13]. Usp22 can also reverse the polycomb complex mediated ubiquitination of H2A and H2B causing multiple changes in gene expression profiles including transcriptional activation of MDM2 and Hox [57], [58]. Recent reports however, indicate that even though Usp22 is active towards both H2A and H2B it preferentially targets H2B for which it is one of the main DUBs.
Cell cycle regulation and DNA damage response
In normal cells most genes have an epigenetically stable transcriptional status. However, some genes are an exception to this rule as their sole purpose is to be responsive to the outside stimuli including growth factors and cellular contact. Those genes are most susceptible to epigenetic changes and their expression is rapidly affected by them. Dysregulation of epigenetics might result in altered expression of these genes leading to cellular transformation and malignancy. As a result, impact of DUBs on cell cycle regulation and DNA damage repair pathways deserves specific attention. The human genome is continuously challenged by both endogenous and exogenous insults potentially damaging the DNA which can result in various types of damage including double and single strand breaks, oxidative lesions and pyrimidine dimers. Cells have developed multiple ways to counteract and repair the DNA damage known as the DNA damage response pathways. These responses can be divided into two main groups; cell-cycle checkpoint activation and DNA repair. Both of these mechanisms are tightly controlled by chromatin remodelling and epigenetics.
As discussed in previous sections multiple DUBs affect cell cycle indirectly by regulating transcription factors that control cell cycle progression, including the AR and p53, and by modulating histones. However, some DUBs regulate cell cycle in a much more direct fashion. DUBs can regulate cell cycle progression by controlling the G2/M checkpoint. Specifically both Usp50 and UbpM contribute towards this process. Usp50, even though it is catalytically inactive, plays a role in cell cycle progression. It associates with Hsp90 and controls Wee1 stability via an Hsp90-dependent mechanism. Usp50 consequently functions as a negative regulator of the G2/M checkpoint [59]. UbpM regulates the same checkpoint but via a different mechanism. Following S552 phosphorylation UbpM translocates to the nucleus and regulates the cell cycle G2/M phase progression and cell proliferation [44]. Additionally, Usp22 directly deubiquitinates TRF1 (TBP(TATA box-binding protein)-related factor 1) to regulate the transcription of cell cycle and apoptosis genes [60] and inhibits the transcriptional activity of p53 by deubiquitinating SIRT1 histone deacetylase [61] and by regulating MDMX stability [62].
Usp1 is a key protein involved in the DNA damage response. Indeed inhibition of the Usp1-Uaf-1 complex sensitises cells to chemotherapy. Usp1 counteracts the mono-ubiquitination of PCNA which prevents recruitment of low fidelity DNA polymerases in the absence of DNA damage [63]. It is also involved in double strand DNA break repair through the homologous recombination pathway. Additionally, Usp1 deubiquitinates and stabilises ID (Inhibitor of DNA binding) proteins 1, 2 and 3, as ID can inhibit differentiation this preserves the undifferentiated state of cells [64]. Usp1 modulates DNA replication, polymerase choice and DNA repair by PCNA and as a result Usp1 knock-out mice are genetically unstable and hypersensitive to DNA damage [65], [66].
Usp10 is involved in DNA damage response control via regulation of the p53 protein. Upon DNA damage, Usp10 is phosphorylated which is essential for Usp10 to relocate to the nucleus, allowing it to stabilise p53 [67]. The DNA damage response is also regulated by the Chk2-p53-PUMA pathway in response to double strand breaks in vivo. This process is tightly controlled by Usp28 which is necessary to stabilise Chk2 and 53BP1 in response to DNA damage and is required for DNA damage induced apoptosis [68]. Usp28 is found to be recruited to double strand breaks and this is dependent on the 53BP1 protein.
Regulation of deubiquitinating enzyme activity
Activity and stability of multiple DUBs is regulated by posttranslational modifications and interactions with other proteins. Commonly enzymes are regulated by phosphorylation, this is also frequent amongst DUBs. To allow its activity towards p53, the Usp10 protein needs to be phosphorylated at T42 and S337 by ATM, this occurs as a result of DNA damage and stabilises Usp10 allowing it to translocate to the nucleus and deubiquitinate p53 [67]. Interestingly in prostate cancer with increased cancer grade and metastasis the expression of Usp10 becomes predominantly nuclear which could possibly be associated with elevated DNA damage. Phosphorylation of Usp10 by ATM is also required for Usp10 antioxidant activity in stress granules [69]. Phosphorylation of UbpM at Ser552 by cyclin-dependent kinase 1 (CDK1) is also needed for its translocation to the nucleus and regulation of the cell cycle G2/M phase progression and cell proliferation but, this phosphorylation is not required for its deubiquitinase activity, substrate specificity and regulation of gene expression [44]. Similarly, Usp7 short isoform (Usp7S) can be phosphorylated by CK2 at S18 leading to its protein stabilisation which subsequently increases the pool of MDM2 and decreases the levels of p53 protein [70]. This is reversed upon irradiation by an ATM dependant phosphatase PPM1G [70]. DUBs can play a role in infection and immunity and their activity can be regulated by bacterial kinases. During Yersinia infection OTUB1 can be phosphorylated by a bacterial kinase YpkA which modulates cellular susceptibility to Yersinia invasion [71].
Many of DUB phosphorylation sites are highly conserved throughout evolution. Yeast homologue of Usp12 (Ubp9) has been shown to be phosphorylated and this phosphorylation was lost upon deletion of both of its co-factors Uaf-1 and WDR20 highlighting its potential role in enzymatic activity [72]. Similarly close family member of Usp12, Usp1 is also phosphorylated at S313 by CDK1 during mitosis, this modification is required for its interaction with Uaf-1 as it lies within the Uaf-1 binding region (amino acids 235-408). Consequently, it is also a pre-requisite for Usp1 DUB activity as complex formation with Uaf-1 is required [73], [74]. Conversely phosphorylation at S42 and S67 have not been attributed any physiological roles to date [73]. S313 of Usp1 lies within both the consensus sequence for CDKs and also Usp1s region 307-330 responsible for APC/C cdh1 mediated Usp1 degradation during G1 phase. Consequently S313 phosphorylation plays a crucial role in maintaining protein stability during mitosis as Usp1 is ubiquitinated during G1 by APC/C cdh1 leading to its proteosomal degradation. Usp1 can be additionally stabilised by CAPNS1 which activates Cdk5 resulting in inhibition of cdh1 subsequently inhibiting Usp1 degradation [75].
Frequently the activity of deubiquitinating enzymes is regulated by interactions with various binding partners. TRAF2 can bind to Usp2a which inhibits its effect on K48 but not K63 linked poly-ubiquitin chains, consequently the ratio between TRAF2 and Usp2a determines cells sensitivity to cell death [76]. Usp10, alongside Usp13, is regulated by binding to Beclin-1 which affects their protein stability, activity and subsequent deubiquitination of target proteins [77]. Usp22 activity is regulated by histone deacetylases, their inhibition abrogates the binding of RNA Polymerase II to Usp22 promoter supressing its transcription [78] .
DUB protein stability has a crucial role in regulating enzymatic activity; some DUBs can auto-regulate themselves others are ubiquitinated/SUMOylated. For example, Usp7 is activated by its own C-terminal domain [79]. Conversely, Usp1 has the ability to auto-cleave itself at a di-glycine motif leading to its degradation [80]. OTUB1 is regulated by mono-ubiquitination at K59 and K109 which is required for it to inhibit the E2 UhcH5 resulting in MDM2 inhibition and p53 activation [33]. Binding of OTUB1 and E2 enzymes, including UbcH5 and Ubc13, also regulates OTUB1 enzymatic activity towards K48 ubiquitin chains [81]. Usp28 on the other hand can be regulated by SUMOylation at the N-terminal domain, this has a negative effect on its deubiquitinase activity [82].
Splicing can also play a role in the regulation of DUBs activity and cellular localisation. Both Usp7 and Usp21 have been reported to undergo alternative splicing. Usp21 short variant lacks the nuclear export sequence (NES) and as a result localises predominantly in the nucleus however, that has no major effect on its enzymatic activity as both full length and variant Usp21 affect H2A ubiquitination to a comparable degree in vitro but variant was observed to be more active in vivo due to its localisation [83].
Table 1: Role of DUBs in histone deubiquitination and the cellular consequences.
DUB |
DUB Family |
Histone substrates |
Cellular processes |
Usp3 |
Usp |
H2A and H2B |
DNA repair and cell cycle progression |
Usp7 |
Usp |
H2A and H2B |
Gene expression, protein stability, cell cycle progression and proliferation |
Usp10 |
Usp |
H2A |
DNA repair and transcription |
Usp12 |
Usp |
H2A and H2B |
Transcription |
UbpM (Usp16) |
Usp |
H2A |
DNA repair, cell cycle progression, differentiation and proliferation |
Usp21 |
Usp |
H2A |
Gene expression and DNA repair |
Usp22 |
Usp |
H2A and H2B |
Gene expression, protein stability, proliferation and cell cycle progression |
Usp29 |
Usp |
H2A |
DNA repair and transcription |
Usp44 |
Usp |
H2A |
DNA repair, cell cycle progression and differentiation |
Usp46 |
Usp |
H2A and H2B |
Transcription |
Usp49 |
Usp |
H2B |
Gene expression and pre-mRNA processing |
MYSM1 |
JAMM |
H2A |
Transcription, haematopoiesis |
OTUB1 |
OTU |
H2A |
DNA repair |
Targeting DUBs
DUBs are key enzymes which regulate cellular growth, survival and homeostasis through multiple pathways including epigenetics, as such aberrations in DUB signalling and activity can play crucial roles in cancer development, progression and metastasis. This is evidenced by multiple carcinogenic agents that exert their effects via the DUB pathway. Carcinogenic properties of nickel compounds have been attributed to the increased ubiquitination of H2A and H2B [84]. However, nickel compounds do not affect histone ubiquitination directly but rather inhibit the DUBs responsible for reversing this process, as such increasing the pool of ubiquitinated histones. As a consequence, targeting DUBs might prove to be a valid strategy for developing novel anti-cancer therapeutics (Table 2).
Table 2: Currently available agents aimed at DUBs discussed in this review.
DUB |
Compound |
Other targeted DUBs |
Reference |
Usp1 |
Pimozide ML323 SJB2-043 and SJB3-019A GW7647 |
Usp2, Usp5, Usp7, Usp8, Usp46 - - - |
[87] [91] [87] |
Usp7 |
HBX 19,818 and HBX 28,258 P5091 and P22077 Pimozide |
Usp47 - Usp1, Usp2, Usp5, Usp8, Usp46 |
[93] [87] |
Usp10 |
Spautin-1 |
Usp13 |
[77] |
Usp14 |
b-AP15 AC17 |
UCHL5 UCHL5 |
[98] [99] |
Usp46 |
Pimozide |
Usp1, Usp2, Usp5, Usp7, Usp8 |
[87] |
Targeting DUBs as an anti-tumourigenic therapeutic strategy has its proof of principle in the use of bortezomib, a broad range inhibitor of the ubiquitin proteasome system, in multiple melanoma treatment [85]. This strategy is however limited by the lack of specificity of bortezomib which results in toxicity. Targeting individual DUBs that play a role in particular cancers is predicted to be a much better strategy. However, targeting single DUBs is a very complex challenge due to the high levels of homology, particularly between the catalytically active domains, and promiscuity. Most DUBs target multiple proteins and additionally many of the DUB targets can be deubiquitinated by more than one DUB. The same DUB can target proteins from the same pathway that exert opposing effects, for example Usp7 can deubiquitinate both p53 and its E3 ligase MDM2 depending on the circumstances. In this context broad spectrum inhibitors targeting closely related families of DUBs that deubiquitinate the same substrates might in fact be a valuable strategy.
As mentioned previously multiple DUBs rely on co-factors, such as WD40 proteins, for their activity (Table 3), [86], [7]. This poses a very exciting drug design opportunity and offers a chance for developing much more specific agents when the binding of two proteins rather than a conserved active domain of one of them is being targeted. As previously discussed, Usp1 is responsible for DNA damage response pathway regulation and its activity is dependent on binding to its cofactor, Uaf-1. This allowed for the identification of more specific agents targeting the Usp1 protein’s interaction with Uaf-1 rather than Usp1 directly, thus circumventing the potential issue of sequence homology and conserved domains within similar DUBs. Three Usp1 inhibitors explored this strategy, Pimozide [87], ML323 [88], [89], [90] and GW7647 [87] each targeting the Usp1-Uaf-1 complex in a non-competitive manner. Usp1 known targets include the Fanconi anaemia complex proteins, FANC1 and FANCD2, PCNA and the inhibitor of DNA binding (ID) transcription factors [87]. As Usp1-null mice have been shown to be hypersensitive to DNA damage it is likely that targeting Usp1 could increase the sensitivity of cancer cells to DNA damaging agents [66]. Multiple Usp1 inhibitors have been identified via both compound library screening and drug development programs. Therapeutic inhibition of Usp1 was previously reported to re-sensitise cisplatin-resistant non-small-cell lung cancer cells to the drug [87]. This result was confirmed by an observed increase in the mono-ubiquitination of PCNA and FANCD2 upon Pimozide treatment which indicated successful targeting of Usp1. Later reports supported this observation with the ML323 compound decreasing the cells ability to repair the DNA damage and potentiating the effects of cisplatin [88]. The authors proposed synthetic lethality as an explanation for this observation; because the DNA damage repair pathway is inhibited upon Usp1 targeting then damage induced by cisplatin is not being repaired by the cells, consequently resulting in cell death. More recently developed compounds SJB2-043 and SJB3-019A inhibited the salvaging of ID1, ID2, ID3, FANC1 and FANCD2 proteins by Usp1 from proteosomal degradation and as a result caused an increase in cell death and sensitisation to DNA damaging agents [91].
Usp7 is one of the most extensively researched DUBs due to its role in regulating multiple key proteins including p53, MDM2, PTEN, FOXO, the polycomb complex and histones [92]. During cancer development, Usp7 plays an oncogenic role as it promotes cellular survival. Most recent attempts at targeting Usp7 include development of P5091, HBX 19,818 and HBX 28,258 inhibitors [93]. All of these compounds were shown to abrogate the effects of Usp7 on p53, they also inhibit the enhanced tumourigenicity of claspin caused by Usp7 activity and increase apoptosis in cancer cells [94]. Promising data arose from the application of P5091 and its second-generation derivative P22077 in an investigation where both compounds were capable of increasing survival when used as single agents in a xenograft study. Additionally, when combined with standard treatments synergistic effects and re-sensitisation were observed for both compounds in multiple melanoma [95] and neuroblastoma [96], respectively. These results are very promising as resistance is one of the major challenges of present cancer management. As such, developing compounds focussed upstream of the present therapeutic targets could help to both avoid developing future resistance and also allow to re-sensitise patients resistant to the available DNA damaging agents by exploring synthetic lethality and combining them with inhibitors of DUBs responsible for DNA damage repair.
Table 3: List of DUBs discussed in this review that rely on binding to WD40 proteins.
DUB |
WD40 |
Cellular processes |
Usp1 |
WDR48 (Uaf-1) |
DNA damage response, Fanconi anaemia pathway, homologous recombination, cellular differentiation and Akt signalling |
Usp3 |
WDTC1 |
DNA repair and cell cycle progression |
Usp7 |
BUB3, WDR21A, RAE1 |
Gene expression, protein stability, cell cycle progression and proliferation |
Usp12 |
WDR20, WDR26, WDR48, WDR77, DMWD |
Transcription, Notch signalling and Akt signalling |
Usp22 |
TAF5L |
Gene expression, protein stability, proliferation and cell cycle progression |
Usp42 |
WDR18 |
Transcription, stress response and cell-cycle progression. When fused to RUNX1 it is involved in pathogenesis of acute leukaemia |
Usp44 |
TBL2 |
DNA repair, cell cycle progression and differentiation |
Usp46 |
WDR20, WDR26, WDR48, WDR77, DMWD |
Transcription, nervous system development and Akt signalling |
Usp49 |
COPA |
Gene expression and pre-mRNA processing |
Usp50 |
PRPF4 |
Cell cycle progression |
Conclusions and future prospects
Discoveries over recent years clearly indicate that ubiquitination and deubiquitination regulate cellular homeostasis and as a result deregulation of these processes can promote cancer development and progression. DUBs can affect carcinogenesis through multiple cellular pathways with epigenetics being the main example. Deubiquitination of histones, transcription factors and their co-factors plays a major role in regulating epigenetics by DUBs. As a result DUBs could be valuable therapeutic targets in oncology. Although this field is still relatively novel, recent advances indicate that targeting DUBs could be an efficient strategy. Novel inhibitors aimed at both Usp1 and Usp7 have been shown to re-sensitise cells to known therapeutic agents and to have a therapeutic effect when used independently in both the cellular setting and in some xenograft models. DUBs might present a challenge for compound design as their active domains are often conserved and many of them are Cys-dependent making targeting more complex. Recently, Usp1 inhibitors have been discovered which inhibit Usp1 by targeting its interaction with Uaf-1, a protein binding partner required for Usp1’s enzymatic activity, rather than by targeting the active domain directly. This is a very exciting direction which could offer a solution to DUB inhibitors design providing both specificity and decreased toxicity. Multiple DUBs rely on binding to other proteins for their activity and stability and many of those interacting partners are WD40 proteins just like Uaf-1 (Table 3), it does however remain to be established if this strategy will prove therapeutically advantageous.
ACKNOWLEDGMENTS
The authors would like to thank Dr Olivier Binda for his help. This work was supported by Prostate Cancer UK, Medical Research Council, Joint Research Executive Scientific Committee and Cancer Research UK.
Grant support
Prostate Cancer UK #PG09-23 (U.L.M and C.N.R), Joint Research Executive Scientific Committee #JG/ML/0414 (U.L.M) and Cancer Research UK #C27826/A15994 (C.N.R).
conflicts of interest
Authors have no conflict of interest.
References
1. Villamil MA, Liang Q and Zhuang Z. The WD40-repeat protein-containing deubiquitinase complex: catalysis, regulation, and potential for therapeutic intervention. Cell biochemistry and biophysics. 2013; 67:111-126.
2. Blumenfeld N, Gonen H, Mayer A, Smith CE, Siegel NR, Schwartz AL and Ciechanover A. Purification and characterization of a novel species of ubiquitin-carrier protein, E2, that is involved in degradation of non-”N-end rule” protein substrates. The Journal of biological chemistry. 1994; 269:9574-9581.
3. Ciechanover A, Heller H, Elias S, Haas AL and Hershko A. ATP-dependent conjugation of reticulocyte proteins with the polypeptide required for protein degradation. Proceedings of the National Academy of Sciences of the United States of America. 1980; 77:1365-1368.
4. Komander D. The emerging complexity of protein ubiquitination. Biochemical Society transactions. 2009; 37:937-953.
5. Martinez-Forero I, Rouzaut A, Palazon A, Dubrot J and Melero I. Lysine 63 polyubiquitination in immunotherapy and in cancer-promoting inflammation. Clinical cancer research : an official journal of the American Association for Cancer Research. 2009; 15:6751-6757.
6. Kulathu Y and Komander D. Atypical ubiquitylation - the unexplored world of polyubiquitin beyond Lys48 and Lys63 linkages. Nature reviews Molecular cell biology. 2012; 13:508-523.
7. Sowa ME, Bennett EJ, Gygi SP and Harper JW. Defining the Human Deubiquitinating Enzyme Interaction Landscape. Cell. 2009; 138:389-403.
8. Li M, Chen D, Shiloh A, Luo J, Nikolaev AY, Qin J and Gu W. Deubiquitination of p53 by HAUSP is an important pathway for p53 stabilization. Nature. 2002; 416:648-653.
9. Hock AK, Vigneron Am Fau - Carter S, Carter S Fau - Ludwig RL, Ludwig Rl Fau - Vousden KH and Vousden KH. Regulation of p53 stability and function by the deubiquitinating enzyme USP42. EMBO J. 2011; 30:4921-4930.
10. Li M, Brooks CL, Kon N and Gu W. A dynamic role of HAUSP in the p53-Mdm2 pathway. Molecular cell. 2004; 13:879-886.
11. Dirac AM and Bernards R. The deubiquitinating enzyme USP26 is a regulator of androgen receptor signaling. Molecular cancer research : MCR. 2010; 8:844-854.
12. Burska UL, Harle VJ, Coffey K, Darby S, Ramsey H, O’Neill D, Logan IR, Gaughan L and Robson CN. Deubiquitinating enzyme Usp12 is a novel co-activator of the Androgen Receptor. The Journal of biological chemistry. 2013; 288:32641-32650.
13. Joo HY, Jones A, Yang C, Zhai L, Smith ADt, Zhang Z, Chandrasekharan MB, Sun ZW, Renfrow MB, Wang Y, Chang C and Wang H. Regulation of histone H2A and H2B deubiquitination and Xenopus development by USP12 and USP46. The Journal of biological chemistry. 2011; 286:7190-7201.
14. Draker R, Sarcinella E and Cheung P. USP10 deubiquitylates the histone variant H2A.Z and both are required for androgen receptor-mediated gene activation. Nucleic acids research. 2011; 39:3529-3542.
15. Li X, Stevens PD, Yang H, Gulhati P, Wang W, Evers BM and Gao T. The deubiquitination enzyme USP46 functions as a tumor suppressor by controlling PHLPP-dependent attenuation of Akt signaling in colon cancer. Oncogene. 2013; 32:471-478.
16. McClurg UL, Summerscales EE, Harle VJ, Gaughan L and Robson CN. Deubiquitinating enzyme Usp12 regulates the interaction between the androgen receptor and the Akt pathway. Oncotarget. 2014; 5:7081-7092.
17. Zhiqiang Z, Qinghui Y, Yongqiang Z, Jian Z, Xin Z, Haiying M and Yuepeng G. USP1 regulates AKT phosphorylation by modulating the stability of PHLPP1 in lung cancer cells. Journal of cancer research and clinical oncology. 2012; 138:1231-1238.
18. Moretti J, Chastagner P, Liang CC, Cohn MA, Israel A and Brou C. The ubiquitin-specific protease 12 (USP12) is a negative regulator of notch signaling acting on notch receptor trafficking toward degradation. The Journal of biological chemistry. 2012; 287:29429-29441.
19. Kovalenko A, Chable-Bessia C, Cantarella G, Israel A, Wallach D and Courtois G. The tumour suppressor CYLD negatively regulates NF-kappaB signalling by deubiquitination. Nature. 2003; 424:801-805.
20. Trompouki E, Hatzivassiliou E, Tsichritzis T, Farmer H, Ashworth A and Mosialos G. CYLD is a deubiquitinating enzyme that negatively regulates NF-kappaB activation by TNFR family members. Nature. 2003; 424:793-796.
21. Taya S, Yamamoto T, Kanai-Azuma M, Wood SA and Kaibuchi K. The deubiquitinating enzyme Fam interacts with and stabilizes beta-catenin. Genes to cells : devoted to molecular & cellular mechanisms. 1999; 4:757-767.
22. Zhao B, Schlesiger C, Masucci MG and Lindsten K. The ubiquitin specific protease 4 (USP4) is a new player in the Wnt signalling pathway. Journal of cellular and molecular medicine. 2009; 13:1886-1895.
23. Gronbaek K, Hother C and Jones PA. Epigenetic changes in cancer. APMIS. 2007; 115:1039-1059.
24. Stanisic V, Malovannaya A, Qin J, Lonard DM and O’Malley BW. OTU Domain-containing ubiquitin aldehyde-binding protein 1 (OTUB1) deubiquitinates estrogen receptor (ER) alpha and affects ERalpha transcriptional activity. The Journal of biological chemistry. 2009; 284:16135-16145.
25. Faus H, Meyer HA, Huber M, Bahr I and Haendler B. The ubiquitin-specific protease USP10 modulates androgen receptor function. Molecular and cellular endocrinology. 2005; 245:138-146.
26. Zhu P, Zhou W, Wang J, Puc J, Ohgi KA, Erdjument-Bromage H, Tempst P, Glass CK and Rosenfeld MG. A histone H2A deubiquitinase complex coordinating histone acetylation and H1 dissociation in transcriptional regulation. Molecular cell. 2007; 27:609-621.
27. Hasty P and Christy BA. p53 as an intervention target for cancer and aging. Pathobiology of aging & age related diseases. 2013; 3.
28. Guseva NV, Rokhlin OW, Bair TB, Glover RB and Cohen MB. Inhibition of p53 expression modifies the specificity of chromatin binding by the androgen receptor. Oncotarget. 2012; 3:183-194.
29. Marouco D, Garabadgiu AV, Melino G and Barlev NA. Lysine-specific modifications of p53: a matter of life and death? Oncotarget. 2013; 4:1556-1571.
30. Thalappilly S, Feng X, Pastyryeva S, Suzuki K, Muruve D, Larocque D, Richard S, Truss M, von Deimling A, Riabowol K and Tallen G. The p53 tumor suppressor is stabilized by inhibitor of growth 1 (ING1) by blocking polyubiquitination. PloS one. 2011; 6:e21065.
31. Sarkari F, Sheng Y and Frappier L. USP7/HAUSP promotes the sequence-specific DNA binding activity of p53. PloS one. 2010; 5:e13040.
32. Sun XX, Challagundla KB and Dai MS. Positive regulation of p53 stability and activity by the deubiquitinating enzyme Otubain 1. EMBO J. 2012; 31:576-592.
33. Li Y, Sun XX, Elferich J, Shinde U, David LL and Dai MS. Monoubiquitination is critical for ovarian tumor domain-containing ubiquitin aldehyde binding protein 1 (Otub1) to suppress UbcH5 enzyme and stabilize p53 protein. The Journal of biological chemistry. 2014; 289:5097-5108.
34. Sato Y, Yamagata A, Goto-Ito S, Kubota K, Miyamoto R, Nakada S and Fukai S. Molecular basis of Lys-63-linked polyubiquitination inhibition by the interaction between human deubiquitinating enzyme OTUB1 and ubiquitin-conjugating enzyme UBC13. The Journal of biological chemistry. 2012; 287:25860-25868.
35. Yagi R, Zhu J and Paul WE. An updated view on transcription factor GATA3-mediated regulation of Th1 and Th2 cell differentiation. International immunology. 2011; 23:415-420.
36. Chou J, Provot S and Werb Z. GATA3 in development and cancer differentiation: cells GATA have it! Journal of cellular physiology. 2010; 222:42-49.
37. Zhang J, Chen C, Hou X, Gao Y, Lin F, Yang J, Gao Z, Pan L, Tao L, Wen C, Yao Z, Tsun A, Shi G and Li B. Identification of the E3 deubiquitinase ubiquitin-specific peptidase 21 (USP21) as a positive regulator of the transcription factor GATA3. The Journal of biological chemistry. 2013; 288:9373-9382.
38. Jung H, Kim BG, Han WH, Lee JH, Cho JY, Park WS, Maurice MM, Han JK, Lee MJ, Finley D and Jho EH. Deubiquitination of Dishevelled by Usp14 is required for Wnt signaling. Oncogenesis. 2013; 2:e64.
39. Wu N, Liu C, Bai C, Han YP, Cho WC and Li Q. Over-Expression of Deubiquitinating Enzyme USP14 in Lung Adenocarcinoma Promotes Proliferation through the Accumulation of beta-Catenin. International journal of molecular sciences. 2013; 14:10749-10760.
40. Zhang XY, Varthi M, Sykes SM, Phillips C, Warzecha C, Zhu W, Wyce A, Thorne AW, Berger SL and McMahon SB. The putative cancer stem cell marker USP22 is a subunit of the human SAGA complex required for activated transcription and cell-cycle progression. Molecular cell. 2008; 29:102-111.
41. Glinsky GV. Genomic models of metastatic cancer: functional analysis of death-from-cancer signature genes reveals aneuploid, anoikis-resistant, metastasis-enabling phenotype with altered cell cycle control and activated Polycomb Group (PcG) protein chromatin silencing pathway. Cell cycle (Georgetown, Tex). 2006; 5:1208-1216.
42. Liu YL, Yang YM, Xu H and Dong XS. Aberrant expression of USP22 is associated with liver metastasis and poor prognosis of colorectal cancer. Journal of surgical oncology. 2011; 103:283-289.
43. Joo HY, Zhai L, Yang C, Nie S, Erdjument-Bromage H, Tempst P, Chang C and Wang H. Regulation of cell cycle progression and gene expression by H2A deubiquitination. Nature. 2007; 449:1068-1072.
44. Xu Y, Yang H, Joo HY, Yu JH, Smith ADt, Schneider D, Chow LT, Renfrow M and Wang H. Ubp-M serine 552 phosphorylation by cyclin-dependent kinase 1 regulates cell cycle progression. Cell cycle (Georgetown, Tex). 2013; 12:3219-3227.
45. Yang W, Lee YH, Jones AE, Woolnough JL, Zhou D, Dai Q, Wu Q, Giles KE, Townes TM and Wang H. The histone H2A deubiquitinase Usp16 regulates embryonic stem cell gene expression and lineage commitment. Nature communications. 2014; 5:3818.
46. Burma S, Chen BP, Murphy M, Kurimasa A and Chen DJ. ATM phosphorylates histone H2AX in response to DNA double-strand breaks. The Journal of biological chemistry. 2001; 276:42462-42467.
47. Stiff T, O’Driscoll M, Rief N, Iwabuchi K, Lobrich M and Jeggo PA. ATM and DNA-PK function redundantly to phosphorylate H2AX after exposure to ionizing radiation. Cancer research. 2004; 64:2390-2396.
48. Li T, Guan J, Huang Z, Hu X and Zheng X. RNF168-mediated H2A neddylation antagonizes ubiquitylation of H2A and regulates DNA damage repair. Journal of cell science. 2014; 127:2238-2248.
49. Sharma N, Zhu Q, Wani G, He J, Wang QE and Wani AA. USP3 counteracts RNF168 via deubiquitinating H2A and gammaH2AX at lysine 13 and 15. Cell cycle (Georgetown, Tex). 2014; 13:106-114.
50. Nakagawa T, Kajitani T, Togo S, Masuko N, Ohdan H, Hishikawa Y, Koji T, Matsuyama T, Ikura T, Muramatsu M and Ito T. Deubiquitylation of histone H2A activates transcriptional initiation via trans-histone cross-talk with H3K4 di- and trimethylation. Genes & development. 2008; 22:37-49.
51. Mosbech A, Lukas C, Bekker-Jensen S and Mailand N. The deubiquitylating enzyme USP44 counteracts the DNA double-strand break response mediated by the RNF8 and RNF168 ubiquitin ligases. The Journal of biological chemistry. 2013; 288:16579-16587.
52. Zhang Z, Jones A, Joo HY, Zhou D, Cao Y, Chen S, Erdjument-Bromage H, Renfrow M, He H, Tempst P, Townes TM, Giles KE, Ma L and Wang H. USP49 deubiquitinates histone H2B and regulates cotranscriptional pre-mRNA splicing. Genes & development. 2013; 27:1581-1595.
53. Hock AK, Vigneron AM and Vousden KH. Ubiquitin-specific peptidase 42 (USP42) functions to deubiquitylate histones and regulate transcriptional activity. The Journal of biological chemistry. 2014; 289:34862-34870.
54. van der Knaap JA, Kumar BR, Moshkin YM, Langenberg K, Krijgsveld J, Heck AJ, Karch F and Verrijzer CP. GMP synthetase stimulates histone H2B deubiquitylation by the epigenetic silencer USP7. Molecular cell. 2005; 17:695-707.
55. Cohn MA, Kee Y, Haas W, Gygi SP and D’Andrea AD. UAF1 is a subunit of multiple deubiquitinating enzyme complexes. The Journal of biological chemistry. 2009; 284:5343-5351.
56. Kee Y, Yang K, Cohn MA, Haas W, Gygi SP and D’Andrea AD. WDR20 regulates activity of the USP12 x UAF1 deubiquitinating enzyme complex. The Journal of biological chemistry. 2010; 285:11252-11257.
57. Zhang XY, Pfeiffer HK, Thorne AW and McMahon SB. USP22, an hSAGA subunit and potential cancer stem cell marker, reverses the polycomb-catalyzed ubiquitylation of histone H2A. Cell cycle (Georgetown, Tex). 2008; 7:1522-1524.
58. Cole AJ, Clifton-Bligh RJ and Marsh DJ. Ubiquitination and cancer: Histone H2B monoubiquitination - roles to play in human malignancy. Endocrine-related cancer. 2014; 22:19-33.
59. Aressy B, Jullien D, Cazales M, Marcellin M, Bugler B, Burlet-Schiltz O and Ducommun B. A screen for deubiquitinating enzymes involved in the G(2)/M checkpoint identifies USP50 as a regulator of HSP90-dependent Wee1 stability. Cell cycle (Georgetown, Tex). 2010; 9:3815-3822.
60. Atanassov BS, Evrard YA, Multani AS, Zhang Z, Tora L, Devys D, Chang S and Dent SY. Gcn5 and SAGA regulate shelterin protein turnover and telomere maintenance. Molecular cell. 2009; 35:352-364.
61. Lin Z, Yang H, Kong Q, Li J, Lee SM, Gao B, Dong H, Wei J, Song J, Zhang DD and Fang D. USP22 antagonizes p53 transcriptional activation by deubiquitinating Sirt1 to suppress cell apoptosis and is required for mouse embryonic development. Molecular cell. 2012; 46:484-494.
62. Ding F, Bao C, Tian Y, Xiao H, Wang M, Xie X, Hu F and Mei J. USP22 Promotes NSCLC Tumorigenesis via MDMX Up-Regulation and Subsequent p53 Inhibition. International journal of molecular sciences. 2014; 16:307-320.
63. Huang TT, Nijman SM, Mirchandani KD, Galardy PJ, Cohn MA, Haas W, Gygi SP, Ploegh HL, Bernards R and D’Andrea AD. Regulation of monoubiquitinated PCNA by DUB autocleavage. Nature cell biology. 2006; 8:339-347.
64. Williams SA, Maecker HL, French DM, Liu J, Gregg A, Silverstein LB, Cao TC, Carano RA and Dixit VM. USP1 deubiquitinates ID proteins to preserve a mesenchymal stem cell program in osteosarcoma. Cell. 2011; 146:918-930.
65. Kim JM, Parmar K, Huang M, Weinstock DM, Ruit CA, Kutok JL and D’Andrea AD. Inactivation of murine Usp1 results in genomic instability and a Fanconi anemia phenotype. Developmental cell. 2009; 16:314-320.
66. Park E, Kim JM, Primack B, Weinstock DM, Moreau LA, Parmar K and D’Andrea AD. Inactivation of Uaf1 causes defective homologous recombination and early embryonic lethality in mice. Molecular and cellular biology. 2013; 33:4360-4370.
67. Yuan J, Luo K, Zhang L, Cheville JC and Lou Z. USP10 regulates p53 localization and stability by deubiquitinating p53. Cell. 2010; 140:384-396.
68. Zhang D, Zaugg K, Mak TW and Elledge SJ. A role for the deubiquitinating enzyme USP28 in control of the DNA-damage response. Cell. 2006; 126:529-542.
69. Takahashi M, Higuchi M, Matsuki H, Yoshita M, Ohsawa T, Oie M and Fujii M. Stress granules inhibit apoptosis by reducing reactive oxygen species production. Molecular and cellular biology. 2013; 33:815-829.
70. Khoronenkova SV, Dianova, II, Ternette N, Kessler BM, Parsons JL and Dianov GL. ATM-dependent downregulation of USP7/HAUSP by PPM1G activates p53 response to DNA damage. Molecular cell. 2012; 45:801-813.
71. Edelmann MJ, Kramer HB, Altun M and Kessler BM. Post-translational modification of the deubiquitinating enzyme otubain 1 modulates active RhoA levels and susceptibility to Yersinia invasion. The FEBS journal. 2010; 277:2515-2530.
72. Kouranti I, McLean JR, Feoktistova A, Liang P, Johnson AE, Roberts-Galbraith RH and Gould KL. A global census of fission yeast deubiquitinating enzyme localization and interaction networks reveals distinct compartmentalization profiles and overlapping functions in endocytosis and polarity. PLoS Biol. 2010; 8:e1000471.
73. Villamil MA, Liang Q, Chen J, Choi YS, Hou S, Lee KH and Zhuang Z. Serine phosphorylation is critical for the activation of ubiquitin-specific protease 1 and its interaction with WD40-repeat protein UAF1. Biochemistry. 2012; 51:9112-9123.
74. Cotto-Rios XM, Jones MJ and Huang TT. Insights into phosphorylation-dependent mechanisms regulating USP1 protein stability during the cell cycle. Cell cycle (Georgetown, Tex). 2011; 10:4009-4016.
75. Cataldo F, Peche LY, Klaric E, Brancolini C, Myers MP, Demarchi F and Schneider C. CAPNS1 regulates USP1 stability and maintenance of genome integrity. Molecular and cellular biology. 2013; 33:2485-2496.
76. Mahul-Mellier AL, Datler C, Pazarentzos E, Lin B, Chaisaklert W, Abuali G and Grimm S. De-ubiquitinating proteases USP2a and USP2c cause apoptosis by stabilising RIP1. Biochimica et biophysica acta. 2012; 1823:1353-1365.
77. Liu J, Xia H, Kim M, Xu L, Li Y, Zhang L, Cai Y, Norberg HV, Zhang T, Furuya T, Jin M, Zhu Z, Wang H, Yu J, Li Y, Hao Y, et al. Beclin1 controls the levels of p53 by regulating the deubiquitination activity of USP10 and USP13. Cell. 2011; 147:223-234.
78. Xiong J, Xu X, Zhou X, Liu J, Gong Z, Wu P and Li W. USP22 transcriptional activity is negatively regulated by the histone deacetylase inhibitor trichostatin A. Molecular medicine reports. 2014; 10:3343-3347.
79. Faesen AC, Dirac AM, Shanmugham A, Ovaa H, Perrakis A and Sixma TK. Mechanism of USP7/HAUSP activation by its C-terminal ubiquitin-like domain and allosteric regulation by GMP-synthetase. Molecular cell. 2011; 44:147-159.
80. Piatkov KI, Colnaghi L, Bekes M, Varshavsky A and Huang TT. The auto-generated fragment of the Usp1 deubiquitylase is a physiological substrate of the N-end rule pathway. Molecular cell. 2012; 48:926-933.
81. Wiener R, DiBello AT, Lombardi PM, Guzzo CM, Zhang X, Matunis MJ and Wolberger C. E2 ubiquitin-conjugating enzymes regulate the deubiquitinating activity of OTUB1. Nature structural & molecular biology. 2013; 20:1033-1039.
82. Zhen Y, Knobel PA, Stracker TH and Reverter D. Regulation of USP28 deubiquitinating activity by SUMO conjugation. The Journal of biological chemistry. 2014; 289:34838-34850.
83. Okuda H, Ohdan H, Nakayama M, Koseki H, Nakagawa T and Ito T. The USP21 short variant (USP21SV) lacking NES, located mostly in the nucleus in vivo, activates transcription by deubiquitylating ubH2A in vitro. PloS one. 2013; 8:e79813.
84. Ke Q, Ellen TP and Costa M. Nickel compounds induce histone ubiquitination by inhibiting histone deubiquitinating enzyme activity. Toxicology and applied pharmacology. 2008; 228:190-199.
85. D’Arcy P and Linder S. Molecular pathways: translational potential of deubiquitinases as drug targets. Clinical cancer research : an official journal of the American Association for Cancer Research. 2014; 20:3908-3914.
86. Faesen Alex C, Luna-Vargas Mark PA, Geurink Paul P, Clerici M, Merkx R, van Dijk Willem J, Hameed Dharjath S, El Oualid F, Ovaa H and Sixma Titia K. The differential modulation of USP activity by internal regulatory domains, interactors and eight ubiquitin chain types. Chemistry & biology. 2011; 18:1550-1561.
87. Chen J, Dexheimer TS, Ai Y, Liang Q, Villamil MA, Inglese J, Maloney DJ, Jadhav A, Simeonov A and Zhuang Z. Selective and cell-active inhibitors of the USP1/ UAF1 deubiquitinase complex reverse cisplatin resistance in non-small cell lung cancer cells. Chemistry & biology. 2011; 18:1390-1400.
88. Liang Q, Dexheimer TS, Zhang P, Rosenthal AS, Villamil MA, You C, Zhang Q, Chen J, Ott CA, Sun H, Luci DK, Yuan B, Simeonov A, Jadhav A, Xiao H, Wang Y, et al. A selective USP1-UAF1 inhibitor links deubiquitination to DNA damage responses. Nature chemical biology. 2014; 10:298-304.
89. Dexheimer TS, Rosenthal AS, Luci DK, Liang Q, Villamil MA, Chen J, Sun H, Kerns EH, Simeonov A, Jadhav A, Zhuang Z and Maloney DJ. Synthesis and structure-activity relationship studies of N-benzyl-2-phenylpyrimidin-4-amine derivatives as potent USP1/UAF1 deubiquitinase inhibitors with anticancer activity against nonsmall cell lung cancer. Journal of medicinal chemistry. 2014; 57:8099-8110.
90. Dexheimer TS, Rosenthal AS, Liang Q, Chen J, Villamil MA, Kerns EH, Simeonov A, Jadhav A, Zhuang Z and Maloney DJ. (2010). Discovery of ML323 as a Novel Inhibitor of the USP1/UAF1 Deubiquitinase Complex. Probe Reports from the NIH Molecular Libraries Program. (Bethesda MD: NIH Molecular Libraries Programme).
91. Mistry H, Hsieh G, Buhrlage SJ, Huang M, Park E, Cuny GD, Galinsky I, Stone RM, Gray NS, D’Andrea AD and Parmar K. Small-molecule inhibitors of USP1 target ID1 degradation in leukemic cells. Molecular cancer therapeutics. 2013; 12:2651-2662.
92. Nicholson B and Suresh Kumar KG. The multifaceted roles of USP7: new therapeutic opportunities. Cell biochemistry and biophysics. 2011; 60:61-68.
93. Reverdy C, Conrath S, Lopez R, Planquette C, Atmanene C, Collura V, Harpon J, Battaglia V, Vivat V, Sippl W and Colland F. Discovery of specific inhibitors of human USP7/HAUSP deubiquitinating enzyme. Chemistry & biology. 2012; 19:467-477.
94. Lill JR and Wertz IE. Toward understanding ubiquitin-modifying enzymes: from pharmacological targeting to proteomics. Trends in pharmacological sciences. 2014; 35:187-207.
95. Chauhan D, Tian Z, Nicholson B, Kumar KG, Zhou B, Carrasco R, McDermott JL, Leach CA, Fulcinniti M, Kodrasov MP, Weinstock J, Kingsbury WD, Hideshima T, Shah PK, Minvielle S, Altun M, et al. A small molecule inhibitor of ubiquitin-specific protease-7 induces apoptosis in multiple myeloma cells and overcomes bortezomib resistance. Cancer cell. 2012; 22:345-358.
96. Fan YH, Cheng J, Vasudevan SA, Dou J, Zhang H, Patel RH, Ma IT, Rojas Y, Zhao Y, Yu Y, Zhang H, Shohet JM, Nuchtern JG, Kim ES and Yang J. USP7 inhibitor P22077 inhibits neuroblastoma growth via inducing p53-mediated apoptosis. Cell death & disease. 2013; 4:e867.
97. Altun M, Kramer HB, Willems LI, McDermott JL, Leach CA, Goldenberg SJ, Kumar KG, Konietzny R, Fischer R, Kogan E, Mackeen MM, McGouran J, Khoronenkova SV, Parsons JL, Dianov GL, Nicholson B, et al. Activity-based chemical proteomics accelerates inhibitor development for deubiquitylating enzymes. Chemistry & biology. 2011; 18:1401-1412.
98. D’Arcy P, Brnjic S, Olofsson MH, Fryknas M, Lindsten K, De Cesare M, Perego P, Sadeghi B, Hassan M, Larsson R and Linder S. Inhibition of proteasome deubiquitinating activity as a new cancer therapy. Nat Med. 2011; 17:1636-1640.
99. Zhou B, Zuo Y, Li B, Wang H, Liu H, Wang X, Qiu X, Hu Y, Wen S, Du J and Bu X. Deubiquitinase inhibition of 19S regulatory particles by 4-arylidene curcumin analog AC17 causes NF-kappaB inhibition and p53 reactivation in human lung cancer cells. Molecular cancer therapeutics. 2013; 12:1381-1392.

