INTRODUCTION
Nasopharyngeal carcinoma (NPC) is a malignancy derived from nasopharyngeal epithelium and is highly prevalent in Southern China and Southeast Asia. Although radiotherapy can effectively control early stage NPC, the average 5-year survival rate after therapy is less than 30–40% in the patients with advanced NPC [1]. Most seriously, due to no obvious symptoms at early stage, more than 60% of clinical NPC patients are easily overlooked and reach advanced stages [2, 3]. Thus, it is necessary to discover novel therapeutic targets and interventions for effectively treating NPC.
MicroRNAs (miRNAs) are endogenous non-coding RNAs that suppress gene expression post-transcriptionally by targeting the 3’UTRs of specific mRNAs. They have emerged as important regulators of physiology and disease, and presented the important therapeutic potential. Recently, aberrant expression of some miRNAs, such as miR-184 [4], miR-34b [5, 6], miR-92a [7], miR-93 [8], miR-27a [9], miR-101 [10], miR-497 [11], and miR-663 [12], has been linked to various cancers including NPC [13], relating to multiple cellular responses such as cell proliferation, cell-cycle progression and tumorigenesis [14, 15]. Epstein-Barr virus (EBV), as a major etiological factor for NPC, encodes 25 miRNA precursors including 3 BHRF1 pre-miRNAs and 22 BART pre-miRNAs [16, 17]. Investigations gradually disclose that this virus can actively deploy its BART miRNAs to flexibly manipulate various viral and host cellular functions. For example, EBV-miR-BART1 influences multiple metabolism–associated genes in NPC [18]. EBV-miR-BART2 suppresses viral DNA polymerase BALF5 to reduce EBV lytic replication [19]. Both EBV-miR-BART5-5p and miR-BART19-5p inhibit viral oncoprotein LMP1 expression [20], while EBV-miR-BART22 decreases LMP2A [21]. Moreover, EBV-miR-BART15-3p suppresses an apoptosis inhibitor BRUCE to induce apoptosis [22] and miR-BART3* targets a tumor suppressor DICE1 to promote cellular growth and transformation in NPC [23]. These findings open up a promising therapeutic option and encourage more studies to develop novel miRNA-based therapies for NPC.
As a member of BART-miRNA family, EBV-miR-BART7-3p is emerging as one of the most important regulatory factors in NPC. We have recently discovered that highly expressed EBV-miR-BART7-3p was closely associated with NPC metastasis and EMT [24]. Other two investigations reported the high EBV-miR-BART7-3p expression in NPC tissue [25]/plasma samples [26] and its correlation with NPC cell proliferation in vitro [26], suggesting its involvement in NPC tumorigenesis though the underlying molecular mechanism still remains unclear. In the present study, we not only elucidate the effect of this viral miRNA on NPC tumorigenesis in vitro and in vivo and explore the underlying regulatory mechanism, but also further investigate the feasibility of using EBV-miR-BART7-3p as a potential onco-target for therapy against NPC. We successfully construct nano-particles carrying anti-EBV-miR-BART7-3p, and carry out an in vivo tumor therapy. This may help to understand a viral miRNA-dependent NPC tumorigenesis and provide a promising bio-therapeutic target for this disease.
RESULTS
EBV-miR-BART7-3p promoted NPC cell growth and tumorigenesis
To gain an insight into the role of EBV-miR-BART7-3p in the cell growth and tumorigenesis of NPC, we firstly generated two NPC cell lines (CNE1 and 5-8F) stably expressing EBV-miR-BART7-3p (5-8F-BART7-3p and CNE1-BART7-3p) and two corresponding control cells (5-8F-NC and CNE1-NC) (Figure S1A, S1B, see Materials and Methods). EBV-miR-BART7-3p expression levels in these two stable cell lines were in a similar physiological range to NPC tissue samples (Figure S1A, S1B). Subsequently, MTT and colony formation assays showed that this viral miRNA significantly promoted cell growth, proliferation (Figure 1A) and colony formation (Figure 1B). Flow cytometric evaluation displayed that these two stable cell lines had an increased proportion of G2 phase and a decreased proportion of G1 phase relative to the control cell lines (Figure 1C, Figure S2A). Contrarily, after the transfection with EBV-miR-BART7-3p inhibitor (anti-miR), these two stable cell lines presented an obviously reduced cell growth and colony formation (Figure S3A and S3B) as well as an increased G1 phase and fewer cells in G2 phase (Figure S2B and Figure S3C).
Further, we conducted an in vivo tumor formation experiment by subcutaneously injecting 5-8F-BART7-3p or 5-8F-NC cells into nude mice. Notably, in three weeks after implantation, the mice injected with 5-8F-BART7-3p cells appeared to carry larger tumor burdens (Figure 1D, Figure S4) and display relatively higher expression levels of Ki67 and PCNA in tumor tissues relative to the controls (Figure 1E).
We also examined EBV-miR-BART7-3p expression in 40 clinical NPC specimens. EBV-miR-BART7-3p tended to be highly expressed in NPC patients with advanced T classification (T3-T4) (Figure S5A), hinting it may be a late event involving in NPC tumorigenesis.
Collectively, these results indicated that EBV-miR-BART7-3p had the ability to enhance NPC tumorigenesis.
EBV-miR-BART7-3p enhanced cell growth and proliferation via stimulating the PTEN/PI3K/Akt pathway and inducing c-Myc and c-Jun
We previously found that EBV-miR-BART7-3p promoted NPC metastasis and EMT via suppressing Phosphatase and tensin homolog (PTEN) [24]. In this present study, we further discovered a new binding site for this miRNA in the 3’UTR of PTEN (Figure S6A). Luciferase reporter assay showed that EBV-miR-BART7-3p mimic attenuated the fluorescence intensity of reporter vector with wt 3’UTR of PTEN by more than two-fold relative to NC, whereas mut 3’UTR showed no response to mimic. The reduced fluorescence intensity of reporter vector with wt 3’UTR was rescued by anti-miR (Figure S6B). Consistently, upon immunohistochemistry evaluation of PTEN protein expression in tissue samples derived from in vivo tumor formation models, we also found an obvious reduction of PTEN expression in 5-8F-BART7-3p group relative to control (Figure S6C). Theses data provided additional evidence that this miRNA repressed PTEN by directly binding to its 3’UTR (Figure S5B).
PTEN/PI3K/Akt constitutes an important pathway modulating multiple biological processes. To address the mechanism of EBV-miR-BART7-3p-mediated phenotypic changes, we conducted western blotting assay to examine phosphorylated Akt (Ser473), a centrally important effector of this pathway. We observed that EBV-miR-BART7-3p enhanced p-Akt expression via suppressing PTEN expression, whereas anti-miR-BART7-3p rescued its expression (Figure 2A and 2B). More interestingly, the expression levels of c-Myc and c-Jun, two transcriptional factors that favor cell growth and proliferation of cancers [27–29], were also increased accordingly (Figure 2A and 2B), indicating that EBV-miR-BART7-3p stimulated PI3K/Akt/c-myc and c-Jun through suppressing PTEN in NPC cells (Figure 2A and 2B). Following the observation of EBV-miR-BART7-3p-mediated growth promotion, we next evaluated the expression of cell-cycle associated genes in EBV-miR-BART7-3p overexpressing or suppressing cells. Two cell-cycle regulators (CCND1 and CCNE1) were highly expressed in EBV-miR-BART7-3p overexpressing cells and lowly expressed in EBV-miR-BART7-3p suppressing cells, whereas P21CIP1, a cell-cycle inhibitor, was lowly expressed in EBV-miR-BART7-3p overexpressing cells and highly expressed in EBV-miR-BART7-3p suppressing cells (Figure 2A and 2B), suggesting the effects of EBV-miR-BART7-3p on cell cycle process probably through activating Akt/c-myc and c-Jun. Furthermore, we did an expanded qPCR examination of cell-cycle regulators and inhibitors. The levels of cyclins and cyclin-dependent kinases were generally increased more than 2-fold upon EBV-miR-BART7-3p overexpression, whereas the levels of cell cycle inhibitors were decreased at least 2-fold (Figure 2C). The opposite results appeared in 5-8F-BART7-3p and CNE1-BART7-3p cells after treating with anti-BART7-3p (Figure 2C). Therefore, these data indicated that EBV-miR-BART7-3p stimulated the PTEN/PI3K/Akt signaling pathway and induced c-Myc and c-Jun, thereby influencing cell cycle process and eventually promoting cell growth and proliferation of NPC (Figure 2D).
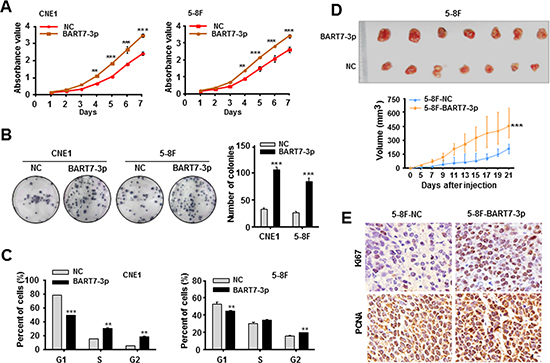
Figure 1: EBV-miR-BART7-3p promotes NPC cell growth and tumorigenesis. (A, B) MTT assay and colony-formation assay were conducted to evaluate cell growth and proliferation in CNE1 and 5-8F cells stably expressing EBV-miR-BART7-3p (BART7-3p for short) and their correspondence NC cells. The data were shown as the mean ± SEM (**P < 0.01, ***P < 0.001). (C) The cell-cycle distribution was detected by the flow cytometry in two stable cell lines as compared with control cells. The data were shown as the mean ± SEM (**P < 0.01, ***P < 0.001). (D) In vivo tumor formation experiment was conducted by the subcutaneous injection of 5-8F-BART7-3p cells and 5-8F-NC cells into seven nude mice respectively. Top, the nude mouse tumors after 21-day inoculation. Bottom, tumor volumes of each group during the tumor growth, plotted as mean values ± SEM (***P < 0.001). (E) The IHC detection of Ki67 and PCNA in the tumor tissues originated from the mouse models.
The siRNA against PTEN (si-PTEN) was also transfected into CNE1 and 5-8F cells. As expected, si-PTEN-transfected CNE1 and 5-8F-cells presented a higher ability to grow and proliferate relative to the control cells (Figure 3A and 3B). Similarly, flow cytometry analysis revealed an increased percentage of G2 phase cells in two NPC cell lines after introducing si-PTEN (Figure 3C). Western blotting assay confirmed a reduced PTEN protein expression in both CNE1 and 5-8F cells after 72 h transfection of si-PTEN (Figure 3D), followed by an obviously induced expression of p-Akt, c-Myc, c-Jun, CCND1 and CCNE1 (Figure 3D). These results supported that knockdown of PTEN mimicked the EBV-miR-BART7-3p-induced tumorigenic phenotype.
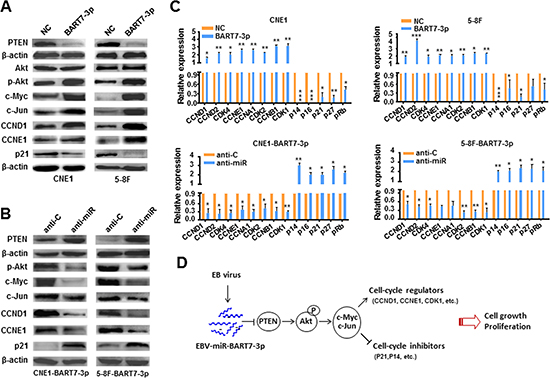
Figure 2: EBV-miR-BART7-3p stimulated PTEN/PI3K/Akt pathway and its downstream signals. (A) The expression levels of Akt, p-Akt, c-Myc, c-Jun, CCND1, CCNE1 and p21 were detected using western blotting in 5-8F-BART7-3p and CNE1-BART7-3p cells or (B) after transfected with anti-miR or anti-C. β-actin served as an internal control. (C) The mRNA expression levels of cell-cycle regulators were analyzed by qPCR in 5-8F-BART7-3p and CNE1-BART7-3p cells before and after transfected with anti-miR. All data were normalized to GAPDH expression and plotted as mean values ± SEM (*p < 0.05, **p < 0.01, ***p < 0.001). (D) The diagram of EBV-miR-BART7-3-mediated signaling pathway for the cell growth and proliferation in NPC.
Silencing of EBV-miR-BART7-3p reduced the in vitro growth of EBV-positive NPC cells
Given that EBV-miR-BART7-3p modulated growth promotion, we further explored whether or not EBV-miR-BART7-3p was a potential therapeutic target for NPC. We firstly analyzed EBV-miR-BART7-3p expression by qPCR in several EBV-positive NPC cell lines. As shown in Figure S7, there was a similar expression level of EBV-miR-BART7-3p in EBV positive NPC cells to the pooled NPC tissues. Of these EBV-positive NPC cell lines, HONE1-EBV cell line was selected as a representative cellular model because it grew better and was suitable for the following in vitro and in vivo experiments [24]. Subsequently, we fabricated a nano-carrier (gold-PEI) using a layer-by-layer method [30] (Figure 4A, see Materials and Methods) to deliver anti-EBV-miR-BART7-3p into HONE1-EBV cells. The size of Gold-PEI nano-carrier was about 20–30 nm as measured by dynamic lighting scatter (DLS) (Figure S8A). The zeta potential of gold-PEI was positive charge (about 20 mV) that facilitated the absorption of anti-miR with negative charge (Figure S8B). The nano-anti-miR displayed an approximately spherical shape with good dispersion and its size was similar to that of gold-PEI observed by TEM (Figure S9). To validate the transfection efficiency at the cellular level, we next investigated the cellular uptake of nano-carrier with FAM-anti-miR using confocal microscopy. There were much more visible green fluorescence particles in HONE-EBV cells transfected with nano-anti-miR (Figure 4B). Furthermore, the inhibition efficiency of nano-anti-miR was evaluated. qPCR revealed that nano-anti-miR effectively inhibited EBV-miR-BART-3p expression level in HONE-EBV cells in the first three days (Figure 4C). After treating with nano-anti-miR, cell growth was accordingly reduced (Figure 4D) and PTEN expression was obviously increased in HONE-EBV while the relevant signals (p-Akt, c-Myc and CCND1) were decreased (Figure 4E).
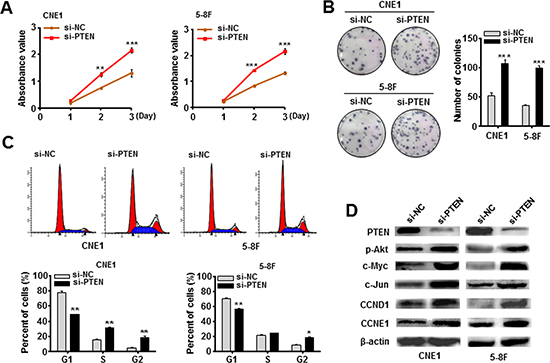
Figure 3: Knockdown of PTEN mimicked the EBV-miR-BART7-3p-induced phenotype in NPC cells. (A, B) Cell growth and proliferation ability was detected by MTT assay and in colony formation assay in CNE1 and 5-8F cells following the treatment of siRNA against PTEN (si-PTEN for short) or si-NC (Control). Data were shown as the mean ± SEM (**P < 0.01, ***P < 0.001). (C) The cell-cycle transition from G1 to S and G2 was evaluated by the flow cytometry after PTEN was silenced in vitro. Data were shown as the mean ± SEM (*P < 0.05, **P < 0.01). (D) The expression levels of PTEN, p-Akt, c-Myc, CCND1 and CCNE1 were analyzed by western blot in indicated cells after treated with si-PTEN. β-actin is internal control.
Silencing of EBV-miR-BART7-3p therapeutically inhibited the tumorigenicity of EBV-positive NPC cells in vivo
We further evaluated the in vivo effectiveness of nano-anti-miR in mice bearing tumors originating from HONE1-EBV cells. The nano-anti-miR was injected into the solid tumor of each mouse per three days according to the results of in vitro experiments. During the 18-day treatment, tumor volume was periodically tested and growth curve was plotted. Consistently, we observed that nano-anti-miR obviously inhibited tumor growth compared with NC control or nano-carrier groups (Figure 5A and 5B). Western blot assay confirmed that PTEN expression was also decreased in tumor tissues, whereas p-Akt, c-Myc and CCND1 were increased after the treatment of nano-anti (Figure 5C).
DISCUSSION
The incidence of viral infection-associated human cancer is rising. Approximately 90% of the people in the world carry latent viruses that can cause various cancers. These viruses majorly include human immunodeficiency virus (HIV), cytomegalovirus (CMV), papilloma virus (HPV), hepatitis B virus (HBV) and EBV [31]. It is estimated that EBV-associated cancers account for about 1.5% of all cancers worldwide, causing about 200,000 cancers worldwide each year, most commonly Burkitt's lymphoma, NPC and about 10% of gastric cancer [32]. Recently, exploring the specific vaccine and therapeutic genes has been becoming a major approach against virus infection [33]. Some tumor-causing viruses such as HPV and HBV have been the subjects of successful vaccination development programs, but there are still no specific treatments or therapeutic genes available for preventing against EBV infection and its associated diseases [31, 34, 35].
EBV is closely associated with NPC. Several EBV-encoded proteins, such as LMP1 [36], LMP2A [37], and BALF3 [38], have been reported to promote the growth and proliferation of NPC cells. However, not all tumor tissues express these viral oncoproteins, suggesting that other viral factors may contribute to NPC aggressiveness. Among the various regulatory mechanisms underlying NPC [39], EBV-encoded miRNAs have been increasingly brought to attention in light of their potential regulatory roles in many aspects [40], including latent virus maintenance [41], escape immune recognition [23], anti-apoptosis [42], and the promotion of invasion and metastasis [43], but there are rare studies on EBV-encoded miRNAs relevant to tumor growth and tumorigenesis. In the present study, the role of EBV-miR-BART7-3p in NPC tumorigenesis was investigated. In vitro and in vivo experiments revealed that this viral miRNA enhanced cell growth, colony formation and cell-cycle progression of NPC cells, provoking further tumorigenesis. Interferential experiments displayed that its functions were reversed upon using specific anti-miR. Clinical sample analysis showed a positive correlation between EBV-miR-BART7-3p expression and NPC T classification in clinical specimens, consistent with our previous observation that this viral miRNA was positively correlated with lymph node metastasis and clinical stage of NPC [24], further highlighting a clinical effect of EBV-miR-BART7-3p on NPC tumorigenesis. These results demonstrate that EBV-miR-BART7-3p may play roles as an oncomir in the development and progression of NPC.
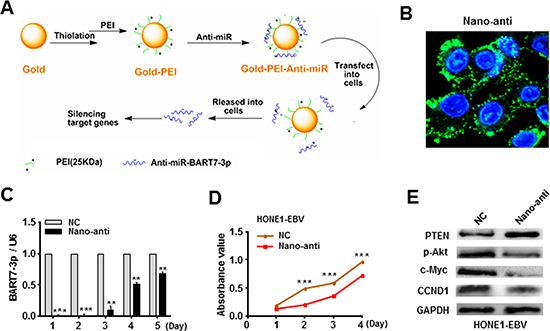
Figure 4: Silencing of EBV-miR-BART7-3p reduced the in vitro growth of EBV-positive NPC cells. (A) A layer-by-layer method was applied to prepare gold-PEI nanoparticles carrying anti-miRNA. (B) The transfection efficiency of nanoparticles carrying anti-miRNA was evaluated by confocal microscopy at cellular level. (C) The inhibition efficiency of nano-anti-miR in HONE1-EBV cells was evaluated by qPCR in different time points. Data were shown as the mean ± SEM (**P < 0.01, ***P < 0.001). (D) Cell growth ability was evaluated by MTT assay in HONE1-EBV cells after transfected with anti-miR or anti-C. Data were shown as the mean ± SEM (***P < 0.001). (E) The expression levels of PTEN, p-Akt, c-Myc and CCND1 were analyzed by western blot in HONE1/EBV cells after transfected with anti-miR or anti-C. β-actin served as an internal control.
To date only a few cellular target genes, such as DICE1 [23], CXCL11 [44], PUMA [42] and E-cadherin [43], have been identified for EBV-encoded miRNAs. As a major tumor suppressor, PTEN has been closely associated with cell proliferation [45], differentiation [46] apoptosis [47], and metastasis [48] in various cancers. We previously demonstrated that PTEN was targeted by EBV-miR-BART7-3p in NPC metastasis and EMT [24]. In the present study, we discovered and experimentally confirmed an additional binding site for this viral miRNA within the 3’UTR of PTEN, and observed that silencing of PTEN mimicked EBV-miR-BART7-3p-induced tumorigenic phenotype, providing additional evidence that EBV-miR-BART7-3p did directly inhibit PTEN in NPC tumorigenesis. It is long believed that PTEN and PI3K/Akt constitute an important pathway modulating multiple biological processes. PTEN is a dual protein/lipid phosphatase which major substrate is PIP3, the product of PI3K. PIP3 can recruit Akt to the membrane where phosphorylation by other kinases (such as PDK1) can activate Akt [49, 50]. Particularly, the activation of PI3K/Akt pathway is always responsible for lowly expressed PTEN in cancers including NPC [51, 52]. Here, our experiments also showed a decrease in PTEN expression and an increase in p-Akt expression in the presence of EBV-miR-BART7-3p, indicating that this pathway was stimulated in EBV-miR-BART7-3p-mediated NPC tumorigensis. Interestingly, it is known that c-Myc and c-Jun transcription factors are critical promoters of cellular proliferation. It is in the active Akt-dependent manner that they can be induced though the regulation is probably indirect [49, 53]. These two transcriptional factors can modulate cell-cycle progression [54, 55], potentiating the transition from the G1 phase into the S phase (G1/S transition) with progression through the cell cycle and further continuing cell proliferation [56]. Of note, in the present study, we observed that c-Myc and c-Jun were induced in the presence of EBV-miR-BART7-3p, and several cell-cycle regulators and inhibitors were influenced accordingly. These data first disclose that a viral miRNA may affect cell-cycle progression and promote cell proliferation through activating PI3K/Akt/ c-myc and c-Jun.
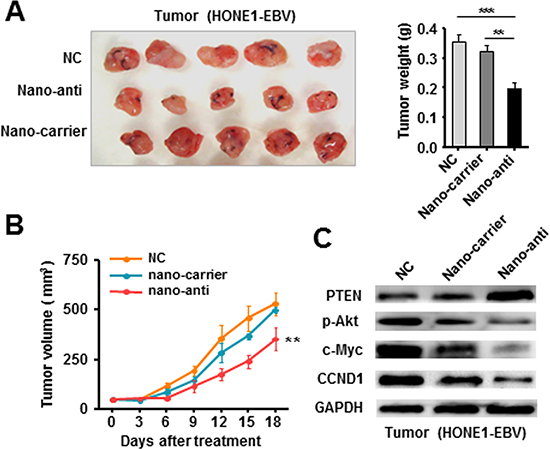
Figure 5: Therapeutically Silencing of EBV-miR-BART7-3p inhibited tumorigenicity of EBV-positive NPC cells in vivo. (A) The in vivo effectiveness of nano-anti-miR was evaluated in xenograft mouse models bearing tumors originating from HONE1-EBV cells. (B) The tumor volume was periodically tested for each mouse and tumor growth curve was plotted. Data were shown as the mean ± SEM (**P < 0.01, ***P < 0.001). (C) The expression of PTEN, p-Akt, c-Myc and CCND1 was analyzed by western blot in the tumor tissues derived from therapeutic and control models respectively. β-actin served as an internal control.
Disease-associated miRNAs have recently become potential targets for therapeutic intervention [57–59]. An inspiring fact is that delivery of a locked nucleic acid (LNA)-modified oligonucliotide complementary to miR-122 has been successfully applied to decrease HCV viremia in primates [60]. This provides evidence of the feasibility of delivering anti-miRNA oligonucleotides for therapy. Nanotechnology has provided a good platform for cancer gene therapy based on nanoparticle unique properties, for example, diverse surface chemistry, appropriate size scale and special pharmacokinetics [61]. The fabrication of nanosized carrier for the delivery of nucleic acid has gained intensive interest because of their potentially clinical application [62]. In the present study, we verified the tumorigenic function of EBV-miR-BART7-3p in NPC, suggesting it was a potential therapeutic target for NPC treatment. Accordingly, we developed a gold-PEI nanocarrier with good transfection efficiency to deliver anti-miR-BART-3p and elicit its potential therapeutic effect. In vitro and In vivo data revealed that cell proliferation and tumor growth was effectively suppressed by anti-EBV-miR-BART7-3p transported by nano-particles and the expression of relevant genes was modulated accordingly, indicating the feasibility of utilizing nano-particles to deliver anti-miR and silence endogenous EBV-miR-BART-3p. More importantly, these results had a clinical implication for cancer treatment involving miRNA intervention by nano-delivery, although there may be several limitations. For example, the intratumoral injection may not fully reflect the curative effect of systemic or intranasal administration in mice. Chemically synthesized nanoparticles may have a certain degree of toxicity and instability that may not be fully evaluated in such a therapeutic experiment done using intratumoral injection. Therefore, our group has been developing a spray that can deliver nano-anti-miR directly into nasal cavity, and is planning to generate the plant-derived nano carrying anti-miR. This may facilitate designing a better in vivo experiment and even clinical application. Moreover, given that multiple EBV BART miRNAs are highly expressed in NPC, a promising nanoparticle-based system carrying multiple antisenses against EBV BART miRNAs is deserved to be established for NPC treatment in the near future.
In summary, we demonstrated that EBV-miR-BART7-3p was a viral oncomir, highly overexpressing in NPC tissues and promoting NPC cell growth, proliferation and tumorigenesis. It stimulated the PTEN/PI3K/Akt pathway and induced c-Myc and c-Jun. We also developed a gold-PEI nanocarrier to deliver anti-miR-BART-3p and elicit its potential therapeutic effect in animal models. Although miRNA-based therapeutics is still in their infancy, our findings are encouraging and EBV BART miRNAs may be potential therapeutic targets for the treatment of patients with NPC.
MATERIALS AND METHODS
Tissue specimens and cell line
A total of 40 NPC specimens (not pretreated with radiotherapy or chemotherapy) with TNM classification and 15 NP specimens were collected and confirmed pathologically in Zhongshan People's Hospital, Guangdong, China for qPCR and clinical data analyses (Table S1 and S2). Staging was performed according to the 1992 Fuzhou NPC staging system of China [63]. T classification (The “T” plus a letter or number 0 to 4) is applied to characterize the size and location of the tumor. Advanced T stage is the late stage (stage T3–4) of NPC relative to the earlier stage (T1 or T2). Clinical tissue studies for research purposes have received patient's informed consents and the approval from the Ethics Committee of Southern Medical University, Guangzhou and Zhongshan People's Hospital, Guangdong, China.
Two EBV-negative NPC cell lines (CNE1, 5-8F) and human 293T cells were obtained from Cancer Research Institute of Southern Medical University, Guangzhou, China. Three EBV-positive NPC cell lines (C666–1-EBV, HONE1-EBV, and HK1-EBV) were kindly offered by Prof. George S. W. Tsao from the University of Hong Kong. All cell lines were cultured in RPMI-1640 (HyClone) with 10% calf serum (Gibco) at 37°C and 5% CO2.
RNA extraction and qPCR
Total RNA was extracted from tissues and cell lines with TRIzol reagent (Invitrogen) according to the user manual. cDNA was synthesized with the PrimeScript RT reagent Kit (TaKaRa). Quantitative PCR analyses were performed with SYBR Premix Ex Taq (TaKaRa) on Mx3005P Stratagene. The primers used are shown in Table S4 and S5. All data were normalized to GAPDH expression and further normalized to negative control unless otherwise indicated. Quantification of EBV-miR-BART7-3p was performed with TaqMan miRNA assays (Applied Biosystems). Mature miRNAs were reverse-transcribed, and real-time PCR was performed using All-in-One™ miRNA qRT-PCR Detection Kit following the manufacturer's protocol (GeneCopoeia™). Data were normalized to small nuclear RNA RNU6B (U6 snRNA) expression. qRT-PCR was conducted for each sample in triplicate. The fold change was calculated through relative quantification (2−ΔΔCt).
Lentivirus production and infection
Lentiviral (GV209, H1-MCS-CMV-EGFP) particles carrying EBV-miR-BART7-3p precursor (BART7-3p for short) and its franking control sequence (NC for short) were constructed by GeneChem, Shanghai, China. The Lentiviral transduction of NPC cells was carried out according to the manufactures’ protocol. CNE1 and 5-8F cells were infected with recombinant Lentiviral transducing units plus 8 mg/ml Polybrene (Sigma-Aldrich) for 2 days, then the EBV-miR-BART7-stably-expressed NPC cells, their corresponding NC cells (GFP+) were sorted with BD FACS AriaTM cell sorter for the following experiments. The expression of EBV-miR-BART7-3p was validated by qPCR (Figure S1A, S1B).
RNA oligoribonucleotides and cell transfection
EBV-miR-BART7-3p mimic and inhibitors (anti-miR) (2′-O-methyl modification) were synthesized by Genepharma (Shanghai, China) (Table S3). PTEN siRNA (h2) (Santa Cruz, sc-44272) and its Control siRNA-A (Santa Cruz, sc-37007) were indicated as si-PTEN and si-NC respectively.
Before transfection, the medium was changed to the RPMI-1640 (HyClone) with 10% fetal bovine serum (Gibco). All cells were maintained in a humidified atmosphere of 95% air and 5% CO2 at 37°C, and seeded on six-well plates (NEST, China) 24 h prior to transfection. Si-PTEN, EBV-miR-BART7-3p mimic or anti-miR was transfected into cells respectively at a final concentration of 50 nmol/L using LipofectamineTM 2000 (Invitrogen, 11668–019) in serum-free conditions. Six hours later, the medium was changed to fresh RPMI-1640 (HyClone) with 10% fetal bovine serum (Gibco).
Cell proliferation analysis
Cell proliferation was analyzed using MTT assay. Briefly, 1 × 103 cells were seeded into a 96-well plate with six replicates for each condition and incubated at 37°C. The cells, stably overexpressing EBV-miR-BART7-3p, were incubated for 1, 2, 3, 4, 5, 6 and 7 days. The cells, infected with anti-miR or si-PTEN, were incubated for 24, 48, and 72 h. 20 ml of MTT (5 mg/ml) (Sigma) was added to each well and incubated for 4 h. At the end of incubation, the media was replaced with 150 ml of dimethyl sulfoxide (DMSO; Sigma). The absorbance value (OD) of each well was measured at 490 nm using a microplate reader. Experiments were performed in triplicate.
Colony formation assay
Cells were plated in six-well culture plates at 2 × 102 cells/well, and the medium was replaced every 3 days. Each cell group had three wells. After incubation for 12 days at 37°C, cells were washed twice with PBS, fixed with 4% paraformaldehyde, and stained with 0.5% crystal violet. The number of colony formation was counted under a microscope.
Cell cycle analysis
For cell cycle analysis, 1 × 105 cells were seeded on 6-well plates in RPMI 1640 containing 10% FBS. After incubation for 48 h, a total of 1 × 106 cells were harvested, rinsed with cold PBS, and fixed with 70% ice-cold ethanol at 4°C overnight. Fixed cells were rinsed with cold PBS followed by incubation with PBS containing 10 mg/ml propidium iodide and 0.5 mg/ml RNase for 30 min at 37°C. The DNA content of labeled cells was acquired using FACS cytometry assay (BD Biosciences). Each experiment was performed in triplicate.
In vivo tumorigenesis in nude mice
Animal experiments were approved by the Ethical Committee of Animal Research at Southern Medical University. The experimental protocol was established according to the associated national guidelines from Ministry of Science and Technology of China.
In vivo tumorigenic ability of EBV-miR-BART7-3p was investigated by tumor xenograft experiment. A total of 1 × 106 5-8F-BART7-3p and 5-8F-NC in 0.2 ml RPMI 1640 medium were subcutaneously injected into the dorsal flanks of 4–6-week-old male BALB/c nu/nu mice. The mice were maintained in a barrier facility on HEPA-filtered racks and fed with an autoclaved laboratory rodent diet. Each experimental group contained seven mice. Tumor size was monitored using a calliper in the process of tumor growth and measured every 3 days. All animal studies were conducted in accordance with the principles and procedures outlined in Southern Medical University Guide for the Care and Use of Animals under assurance number SCXK (Guangdong) 2011-020. After 3 weeks, mice were killed and tumors were dissected. Tumor volumes were calculated as follows: volume = (D × d 2)/2, where D is the longest diameter and d is the shortest diameter.
Dual luciferase assay
293T cells (1×104) were cultured in 24-well plates and co-transfected with 20 nM EBV-miR-BART7-3p mimics or NC, 5 ng of pRL-CMV Renilla luciferase reporter and 30ng of luciferase reporter that contained the wild-type or mutant 3’-UTR of PTEN. For antagonism experiments, cells were also co-transfected with 20 nM anti-miR-BART7-3p or anti-C. Transfections were performed in duplicate and repeated in three independent experiments. 48 h after transfection, the luciferase activities were analyzed with a Dual-Luciferase Reporter Assay System (Promega).
Western blot analysis
Western blotting analyses were performed with standard methods. Briefly, cell pallets were lysed in the radio-immunoprecipitation assay (RIPA) buffer containing protease inhibitors (Sigma-Aldrich) and phosphatase inhibitors (Keygen, China). Proteins were separated by 10% SDS-PAGE gels, and blotted onto PVDF (polyvinylidene difluoride) membrane (Millipore). The membrane was probed with the specific antibodies (Table S6), and then with peroxidase-conjugated secondary antibodies. Beta-actin was used as a protein loading control. The bands were visualized by eECL Western Blot Kit (CWBIO Technology). The images were captured with ChemiDocTM CRS+ Molecular Imager (Bio-Rad).
Immunohistochemical staining
The paraffin sections prepared from in vivo experiments were applied to immunohistochemistry assays for detecting protein expression levels of PTEN, Ki67 and PCNA proteins. The indirect streptavidin-peroxidase method was used as the manufacture's introduction. The immunohistochemically stained tissue sections were reviewed separately by two pathologists. The information of antibodies was shown in table S6.
Preparation of AuNPs-PEI/PEG-antisense (nano-anti)
AuNPs-PEI/PEG-antisense was prepared as previously described method [30]. Briefly, AuNPs with the average diameter of 20 nm were prepared by using the standard reduction of tetrachloroauric (III) acid with sodium citrate. The pH of AuNPs was adjusted to 10, followed by the addition of 11-MUA at a final concentration of 0.1 mg/mL. The stabilized particles were purified twice at 12,000 xg for 25 min and resuspended in 1 mM NaCl. For coating step with PEI/PEG, 1.0 mg/mL PEI/PEG was added to the AuNPs-MUA, and stirred for 30 min. Purification of crude product was performed twice at 12,000 xg for 25 min and the stabilized particles were resuspended in 10 mM NaCl. For coating step with antisense, antisense was added at a final concentration of 2.0 μM to purified PEI/PEG-AuNPs and reacted for 30 min at room temperature.
Treatment experiment on nude mice
In vivo experiment was approved by the Animal Care and Use Committee of Southern Medical University. All mice of 4–5 weeks old and 16–20 g in weight were provided by the Central Animal Facility of Southern Medical University. To establish an NPC mouse model, 6 × 105 HONE1-EBV cells (expressing EBV-miR-BART7-3p) in 0.2 mL buffered saline were subcutaneously injected into the back of nude mice (BALB/C, nu/nu, 4–6 weeks) and tumors were allowed to grow for 10 days. The tumor volume was measured with a caliper every three days and calculated by the formula: Volume = 1/2 × length × width2, where length represented the longest tumor diameter and width represented the shortest tumor diameter. When tumor volume reached 30–100 mm3, the animals were randomized into three groups for therapy testing. Mice were intratumorally injected with saline, nano-carrier (10 uL) and nano-anti (10 μL nano-carrier combined with 8 μL EBV-miR-BART7-3p antisense), respectively. All mice were sacrificed after 18 days and the tumors were carefully dissected and snap frozen for RNA and protein expression analysis.
Statistical analysis
The data are expressed as the mean ± Standard error of the mean (SEM) from at least three independent experiments. Comparisons between two groups were performed using Student's t-test, unless otherwise indicated. Statistical analyses were performed with the SPSS 13.0 statistical software package (SPSS Inc. Chicago, IL, USA). All statistical tests were two-sided, and P < 0.05 was considered to be statistically significant.
ACKNOWLEDGEMENTS
This work was supported by grants from the National Natural Science Foundation of China (No. 81372895, 21204036), Guangzhou Science and Technology research project ( No. S2014J4100149, S2011040003731), and Research Fund for the Doctoral Program of Higher Education of China (No. 20134433110013). No potential conflicts of interest were disclosed.
REFERENCES
1. Sun X, Zeng L, Chen C, Huang Y, Han F, Xiao W, Liu S, Lu T. Comparing treatment outcomes of different chemotherapy sequences during intensity modulated radiotherapy for advanced N-stage nasopharyngeal carcinoma patients. Radiat Oncol. 2013; 8:265.
2. Perri F, Bosso D, Buonerba C, Lorenzo GD, Scarpati GD. Locally advanced nasopharyngeal carcinoma: Current and emerging treatment strategies. World J Clin Oncol. 2011; 2:377–383.
3. Xiao C, Wang L, Jiao Y, Sun K, Qin S, Xu X, Guo J, Zhou J. Long-term results of concurrent chemoradiotherapy for T3/T4 locally advanced nasopharyngeal carcinoma. Mol Clin Oncol. 2013; 1:507–510.
4. Zhen Y, Liu Z, Yang H, Yu X, Wu Q, Hua S, Long X, Jiang Q, Song Y, Cheng C, Wang H, Zhao M, Fu Q, et al. Tumor suppressor PDCD4 modulates miR-184-mediated direct suppression of C-MYC and BCL2 blocking cell growth and survival in nasopharyngeal carcinoma. Cell Death Dis. 2013; 4:e872.
5. Nadal E, Chen G, Gallegos M, Lin L, Ferrer-Torres D, Truini A, Wang Z, Lin J, Reddy RM, Llatjos R, Escobar I, Moya J, Chang AC, et al. Epigenetic inactivation of microRNA-34b/c predicts poor disease-free survival in early-stage lung adenocarcinoma. Clin Cancer Res. 2013; 19:6842–6852.
6. Corney DC, Flesken-Nikitin A, Godwin AK, Wang W, Nikitin AY. MicroRNA-34b and MicroRNA-34c are targets of p53 and cooperate in control of cell proliferation and adhesion-independent growth. Cancer Res. 2007; 67:8433–8438.
7. Zhang G, Zhou H, Xiao H, Liu Z, Tian H, Zhou T. MicroRNA-92a functions as an oncogene in colorectal cancer by targeting PTEN. Dig Dis Sci. 2014; 59:98–107.
8. Lyu X, Fang W, Cai L, Zheng H, Ye Y, Zhang L, Li J, Peng H, Cho WC, Wang E, Marincola FM, Yao K, Cai H, et al. TGFbetaR2 is a major target of miR-93 in nasopharyngeal carcinoma aggressiveness. Mol Cancer. 2014; 13:51.
9. Guo D, Li Q, Lv Q, Wei Q, Cao S, Gu J. MiR-27a Targets sFRP1 in hFOB Cells to Regulate Proliferation, Apoptosis and Differentiation. PLoS One. 2014; 9:e91354.
10. Konno Y, Dong P, Xiong Y, Suzuki F, Lu J, Cai M, Watari H, Mitamura T, Hosaka M, Hanley SJ, Kudo M, Sakuragi N. MicroRNA-101 targets EZH2, MCL-1 and FOS to suppress proliferation, invasion and stem cell-like phenotype of aggressive endometrial cancer cells. Oncotarget. 2014; 5:6049–6062.
11. Xu J, Wang T, Cao Z, Huang H, Li J, Liu W, Liu S, You L, Zhou L, Zhang T, Zhao Y. MiR-497 downregulation contributes to the malignancy of pancreatic cancer and associates with a poor prognosis. Oncotarget. 2014; 5:6983–6993.
12. Yi C, Wang Q, Wang L, Huang Y, Li L, Liu L, Zhou X, Xie G, Kang T, Wang H, Zeng M, Ma J, Zeng Y, et al. MiR-663, a microRNA targeting p21(WAF1/CIP1), promotes the proliferation and tumorigenesis of nasopharyngeal carcinoma. Oncogene. 2012; 31:4421–4433.
13. He ML, Luo MX, Lin MC, Kung HF. MicroRNAs: potential diagnostic markers and therapeutic targets for EBV-associated nasopharyngeal carcinoma. Biochim Biophys Acta. 2012; 1825:1–10.
14. Kovarikova A, Hezova R, Srovnal J, Redova-Lojova M, Slaby O. The Role of MicroRNAs in Molecular Pathology of Esophageal Cancer and Their Potential Usage in Clinical Oncology. Klin Onkol. 2014; 27:87–96.
15. Xu Y, Liu L, Liu J, Zhang Y, Zhu J, Chen J, Liu S, Liu Z, Shi H, Shen H, Hu Z. A potentially functional polymorphism in the promoter region of miR-34b/c is associated with an increased risk for primary hepatocellular carcinoma. Int J Cancer. 2011; 128:412–417.
16. Pfeffer S, Zavolan M, Grasser FA, Chien M, Russo JJ, Ju J, John B, Enright AJ, Marks D, Sander C, Tuschl T. Identification of virus-encoded microRNAs. Science. 2004; 304:734–736.
17. Skalsky RL, Corcoran DL, Gottwein E, Frank CL, Kang D, Hafner M, Nusbaum JD, Feederle R, Delecluse HJ, Luftig MA, Tuschl T, Ohler U, Cullen BR. The viral and cellular microRNA targetome in lymphoblastoid cell lines. PLoS Pathog. 2012; 8:e1002484.
18. Ye Y, Zhou Y, Zhang L, Chen Y, Lyu X, Cai L, Lu Y, Deng Y, Wang J, Yao K, Fang W, Cai H, Li X. EBV-miR-BART1 is involved in regulating metabolism-associated genes in nasopharyngeal carcinoma. Biochem Biophys Res Commun. 2013; 436:19–24.
19. Barth S, Pfuhl T, Mamiani A, Ehses C, Roemer K, Kremmer E, Jaker C, Hock J, Meister G, Grasser FA. Epstein-Barr virus-encoded microRNA miR-BART2 down-regulates the viral DNA polymerase BALF5. Nucleic Acids Res. 2008; 36:666–675.
20. Riley KJ, Rabinowitz GS, Yario TA, Luna JM, Darnell RB, Steitz JA. EBV and human microRNAs co-target oncogenic and apoptotic viral and human genes during latency. Embo J. 2012; 31:2207–2221.
21. Lung RW, Tong JH, Sung YM, Leung PS, Ng DC, Chau SL, Chan AW, Ng EK, Lo KW, To KF. Modulation of LMP2A expression by a newly identified Epstein-Barr virus-encoded microRNA miR-BART22. Neoplasia. 2009; 11:1174–1184.
22. Choi H, Lee H, Kim SR, Gho YS, Lee SK. Epstein-Barr virus-encoded microRNA BART15–3p promotes cell apoptosis partially by targeting BRUCE. J Virol. 2013; 87:8135–8144.
23. Lei T, Yuen KS, Xu R, Tsao SW, Chen H, Li M, Kok KH, Jin DY. Targeting of DICE1 tumor suppressor by Epstein-Barr virus-encoded miR-BART3* microRNA in nasopharyngeal carcinoma. Int J Cancer. 2013; 133:79–87.
24. Cai LM, Lyu XM, Luo WR, Cui XF, Ye YF, Yuan CC, Peng QX, Wu DH, Liu TF, Wang E, Marincola FM, Yao KT, Fang WY, et al. EBV-miR-BART7-3p promotes the EMT and metastasis of nasopharyngeal carcinoma cells by suppressing the tumor suppressor PTEN. Oncogene. 2014; doi: 10.1038/onc.2014.341. [Epub ahead of print].
25. Chang PL, Chang YS, Chen JH, Chen SJ, Chen HC. Analysis of BART7 microRNA from Epstein-Barr virus-infected nasopharyngeal carcinoma cells by capillary electrophoresis. Anal Chem. 2008; 80:8554–8560.
26. Chan JY, Gao W, Ho WK, Wei WI, Wong TS. Overexpression of Epstein-Barr virus-encoded microRNA-BART7 in undifferentiated nasopharyngeal carcinoma. Anticancer Res. 2012; 32:3201–3210.
27. Liang J, Slingerland JM. Multiple roles of the PI3K/PKB (Akt) pathway in cell cycle progression. Cell Cycle. 2003; 2:339–345.
28. Vartanian R, Masri J, Martin J, Cloninger C, Holmes B, Artinian N, Funk A, Ruegg T, Gera J. AP-1 regulates cyclin D1 and c-MYC transcription in an AKT-dependent manner in response to mTOR inhibition: role of AIP4/Itch-mediated JUNB degradation. Mol Cancer Res. 2011; 9:115–130.
29. Scoville DH, Sato T, He XC, Li L. Current view: intestinal stem cells and signaling. Gastroenterology. 2008; 134:849–864.
30. Elbakry A, Zaky A, Liebl R, Rachel R, Goepferich A, Breunig M. Layer-by-layer assembled gold nanoparticles for siRNA delivery. Nano Lett. 2009; 9:2059–2064.
31. Holmes D. The cancer-virus cures. Nat Med. 2014; 20:571–574.
32. Lo AK, Dawson CW, Jin DY, Lo KW. The pathological roles of BART miRNAs in nasopharyngeal carcinoma. J Pathol. 2012; 227:392–403.
33. Lapp S, Pfankuche VM, Baumgartner W, Puff C. Viral oncolysis - can insights from measles be transferred to canine distemper virus?. Viruses. 2014; 6:2340–2375.
34. Fang W, Zhang J, Hong S, Zhan J, Chen N, Qin T, Tang Y, Zhang Y, Kang S, Zhou T, Wu X, Liang W, Hu Z, et al. EBV-driven LMP1 and IFN-gamma up-regulate PD-L1 in nasopharyngeal carcinoma: Implications for oncotargeted therapy. Oncotarget. 2014; 5; 12189–12202.
35. Smith PA, Merritt D, Barr L, Thorley-Lawson DA. An orthotopic model of metastatic nasopharyngeal carcinoma and its application in elucidating a therapeutic target that inhibits metastasis. Genes Cancer. 2011; 2:1023–1033.
36. Xiao L, Hu ZY, Dong X, Tan Z, Li W, Tang M, Chen L, Yang L, Tao Y, Jiang Y, Li J, Yi B, Li B, et al. Targeting Epstein-Barr virus oncoprotein LMP1-mediated glycolysis sensitizes nasopharyngeal carcinoma to radiation therapy. Oncogene. 2014.
37. Kong QL, Hu LJ, Cao JY, Huang YJ, Xu LH, Liang Y, Xiong D, Guan S, Guo BH, Mai HQ, Chen QY, Zhang X, Li MZ, et al. Epstein-Barr virus-encoded LMP2A induces an epithelial-mesenchymal transition and increases the number of side population stem-like cancer cells in nasopharyngeal carcinoma. PLoS Pathog. 2010; 6:e1000940.
38. Chiu SH, Wu CC, Fang CY, Yu SL, Hsu HY, Chow YH, Chen JY. Epstein-Barr virus BALF3 mediates genomic instability and progressive malignancy in nasopharyngeal carcinoma. Oncotarget. 2014; 5:8583–8601.
39. Li HP, Peng CC, Chung IC, Huang MY, Huang ST, Chen CC, Chang KP, Hsu CL, Chang YS. Aberrantly hypermethylated Homeobox A2 derepresses metalloproteinase-9 through TBP and promotes invasion in Nasopharyngeal carcinoma. Oncotarget. 2013; 4:2154–2165.
40. Wang LJ, Chou YF, Chen PR, Su B, Hsu YC, Chang CH, Lee JW. Differential miRNA expression in repeated recurrence of nasopharyngeal carcinoma. Cancer Lett. 2014; 344:188–194.
41. Iizasa H, Wulff BE, Alla NR, Maragkakis M, Megraw M, Hatzigeorgiou A, Iwakiri D, Takada K, Wiedmer A, Showe L, Lieberman P, Nishikura K. Editing of Epstein-Barr virus-encoded BART6 microRNAs controls their dicer targeting and consequently affects viral latency. J Biol Chem. 2010; 285:33358–33370.
42. Choy EY, Siu KL, Kok KH, Lung RW, Tsang CM, To KF, Kwong DL, Tsao SW, Jin DY. An Epstein-Barr virus-encoded microRNA targets PUMA to promote host cell survival. J Exp Med. 2008; 205:2551–2560.
43. Hsu CY, Yi YH, Chang KP, Chang YS, Chen SJ, Chen HC. The Epstein-Barr virus-encoded microRNA MiR-BART9 promotes tumor metastasis by targeting E-cadherin in nasopharyngeal carcinoma. PLoS Pathog. 2014; 10:e1003974.
44. Xia T, O’Hara A, Araujo I, Barreto J, Carvalho E, Sapucaia JB, Ramos JC, Luz E, Pedroso C, Manrique M, Toomey NL, Brites C, Dittmer DP, et al. EBV microRNAs in primary lymphomas and targeting of CXCL-11 by ebv-mir-BHRF1–3. Cancer Res. 2008; 68:1436–1442.
45. Damania P, Sen B, Dar SB, Kumar S, Kumari A, Gupta E, Sarin SK, Venugopal SK. Hepatitis B Virus Induces Cell Proliferation via HBx-Induced microRNA-21 in Hepatocellular Carcinoma by Targeting Programmed Cell Death Protein4 (PDCD4) and Phosphatase and Tensin Homologue (PTEN). PLoS One. 2014; 9:e91745.
46. Gont A, Hanson JE, Lavictoire SJ, Parolin DA, Daneshmand M, Restall IJ, Soucie M, Nicholas G, Woulfe J, Kassam A, Da SV, Lorimer IA. PTEN loss represses glioblastoma tumor initiating cell differentiation via inactivation of Lgl1. Oncotarget. 2013; 4:1266–1279.
47. Liao C, Chen W, Fan X, Jiang X, Qiu L, Chen C, Zhu Y, Wang H. MicroRNA-200c inhibits apoptosis in pituitary adenoma cells by targeting the PTEN/Akt signaling pathway. Oncol Res. 2014; 21:129–136.
48. Wang H, Wu Q, Liu Z, Luo X, Fan Y, Liu Y, Zhang Y, Hua S, Fu Q, Zhao M, Chen Y, Fang W, Lv X. Downregulation of FAP suppresses cell proliferation and metastasis through PTEN/PI3K/AKT and Ras-ERK signaling in oral squamous cell carcinoma. Cell Death Dis. 2014; 5:e1155.
49. Bader AG, Kang S, Zhao L, Vogt PK. Oncogenic PI3K deregulates transcription and translation. Nat Rev Cancer. 2005; 5:921–929.
50. Carnero A, Blanco-Aparicio C, Renner O, Link W, Leal JF. The PTEN/PI3K/AKT signalling pathway in cancer, therapeutic implications. Curr Cancer Drug Targets. 2008; 8:187–198.
51. Trigka EA, Levidou G, Saetta AA, Chatziandreou I, Tomos P, Thalassinos N, Anastasiou N, Spartalis E, Kavantzas N, Patsouris E, Korkolopoulou P. A detailed immunohistochemical analysis of the PI3K/AKT/mTOR pathway in lung cancer: correlation with PIK3CA, AKT1, K-RAS or PTEN mutational status and clinicopathological features. Oncol Rep. 2013; 30:623–636.
52. Foster JS, Fish LM, Phipps JE, Bruker CT, Lewis JM, Bell JL, Solomon A, Kestler DP. Odontogenic ameloblast-associated protein (ODAM) inhibits growth and migration of human melanoma cells and elicits PTEN elevation and inactivation of PI3K/AKT signaling. Bmc Cancer. 2013; 13:227.
53. Xu N, Lao Y, Zhang Y, Gillespie DA. Akt: a double-edged sword in cell proliferation and genome stability. J Oncol. 2012; 2012:15pages.
54. Liang J, Slingerland JM. Multiple roles of the PI3K/PKB (Akt) pathway in cell cycle progression. Cell Cycle. 2003; 2:339–345.
55. Whang YM, Kim YH, Kim JS, Yoo YD. RASSF1A suppresses the c-Jun-NH2-kinase pathway and inhibits cell cycle progression. Cancer Res. 2005; 65:3682–3690.
56. Taira N, Mimoto R, Kurata M, Yamaguchi T, Kitagawa M, Miki Y, Yoshida K. DYRK2 priming phosphorylation of c-Jun and c-Myc modulates cell cycle progression in human cancer cells. J Clin Invest. 2012; 122:859–872.
57. Zheng J, Deng J, Xiao M, Yang L, Zhang L, You Y, Hu M, Li N, Wu H, Li W, Lu J, Zhou Y. A sequence polymorphism in miR-608 predicts recurrence after radiotherapy for nasopharyngeal carcinoma. Cancer Res. 2013; 73:5151–5162.
58. Han K, Xuan P, Ding J, Zhao ZJ, Hui L, Zhong YL. Prediction of disease-related microRNAs by incorporating functional similarity and common association information. Genet Mol Res. 2014; 13:2009–2019.
59. Cai YL, Li J, Lu AY, Zheng YM, Zhong WM, Wang W, Gao JQ, Zeng H, Cheng JR, Tang MZ. Diagnostic Significance of Combined Detection of Epstein-Barr Virus Antibodies, VCA/IgA, EA/IgA, Rta/IgG and EBNA1/IgA for Nasopharyngeal Carcinoma. Asian Pac J Cancer Prev. 2014; 15:2001–2006.
60. Lanford RE, Hildebrandt-Eriksen ES, Petri A, Persson R, Lindow M, Munk ME, Kauppinen S, Orum H. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis C virus infection. Science. 2010; 327:198–201.
61. Chen Y, Zhu X, Zhang X, Liu B, Huang L. Nanoparticles modified with tumor-targeting scFv deliver siRNA and miRNA for cancer therapy. Mol Ther. 2010; 18:1650–1656.
62. Cubillos-Ruiz JR, Sempere LF, Conejo-Garcia JR. Good things come in small packages: Therapeutic anti-tumor immunity induced by microRNA nanoparticles. Oncoimmunology. 2012; 1:968–970.
63. Gao JM, Zeng YX, Cui NJ, Lu TX, Zhao C, Xia YF, Ma J, Xie FY. Staging 915 cases of nasopharyngeal carcinoma after simple radical radiotherapy—checkout of Fuzhou staging system (1992). Ai Zheng. 2005; 24:1165–1172.

