Introduction
Bladder cancer is a common malignancy in urinary system. According to data of Global Cancer Statistics, about 440,000 new cases of bladder cancer are diagnosed while 130,000 patients died from it around the world every year. In China, bladder cancer has the highest incidence in all urinary system tumors and the mortality increased significantly during the past decades [1]. The high recurrence rate of non-invasive bladder cancer [2] and the evident aggressive performance of invasive bladder cancer are the main threaten to patient life. However, the effective therapy for bladder cancer is still deficient because the detailed mechanism underlying bladder cancer is unclear. Therefore, intensive study into the molecular mechanism will promote clinical treatment of bladder cancer.
With the advance of high-throughput transcriptome analyses, most of the human genome has been found to be transcribed into noncoding RNAs. Different from the conventional coding genes, which exert their biological functions by translated into protein molecules, noncoding RNAs play important roles in regulation of various biological process in form of RNA, and do not have the capability to coding proteins [3]. Among the noncoding RNAs, a group of long noncoding RNAs (lncRNAs) with a length of >200 nucleotides have been highlighted in virtue of their large amount and the prominent part they played in intracellular regulation network [4-6]. Recent studies have demonstrated that lncRNAs were closely involved in carcinogenesis and can be used as the potential biomarkers of cancer [7-10]. Besides, several lncRNAs were identified to influence occurrence or progress of bladder cancer, e.g. ncRNA [11], UBC1 [12], GAS5 [13], etc.. However, most of the role of lncRNAs in bladder cancer remains unknown yet.
According to their relative location to adjacent coding genes, lncRNAs can be roughly divided into 5 categories, i.e. antisense lncRNAs, sense lncRNAs, intronic lncRNAs, bidirectional lncRNAs and intergenic lncRNAs [14]. Among them, antisense lncRNAs have been recognized to regulate expression of corresponding coding genes in post-transcriptional level [15] and therefore participate in carcinogenesis by regulation of oncogenes as well as anti-oncogenes. For instance, antisense transcript of coding gene PCNA, PCNA-AS1, was recently shown to increase PCNA mRNA stability and promote hepatocellular tumor growth [16].
Mediator of DNA damage checkpoint protein 1 (MDC1) was an important mediator of the repair of double-strand breaks (DSB). It acts as a strong tumor suppressor through its DNA damage repair function and may be involved in carcinogenesis of bladder cancer. However, antisense transcript of MDC1, named MDC1-AS in the present study, was a novel unknown lncRNA. Studies on the biological function of MDC1-AS and its role in carcinogenesis have not been reported yet.
To investigate the dysregulated lncRNAs in bladder cancer and demonstrate their biological roles, we conducted lncRNA microarray assay using 3 paired bladder cancer tissues. For the identified dysregulated lncRNAs, we performed preliminary screening and selected MDC1-AS as the main subject of the present study. The following molecular assays demonstrated a tumor-suppressor role of this novel antisense lncRNA.
Results
Difference of lncRNA expression profile between bladder cancer tissues and non-cancer tissues
Fold change greater than 2 and P value less than 0.05 between cancer tissues and non-cancer tissues were set as the criteria in filtering differently expressed lncRNAs. In summary, there were 562 up-regulated and 672 down-regulated lncRNAs in bladder cancer tissues compared with adjacent normal tissues, including 152 antisense lncRNAs, 89 sense lncRNAs, 158 intronic lncRNAs, 58 bidirectional lncRNAs and 771 intergenic lncRNAs.
Results of hierarchical clustering analysis on the most significantly dysregulated cancer-related antisense lncRNAs are shown in Figure 1. Through further literature search on the adjacent coding genes, we found that coding gene MDC1 was more deeply involved in carcinogenesis than the others. It was an important participant in DNA damage repair process with established tumor-suppress function. Therefore, in the following study, we focused on antisense lncRNA of MDC1, MDC1-AS (transcript number ENST00000442150), and performed a series of molecular biological assays to deeply investigate role of MDC1-AS in bladder cancer.
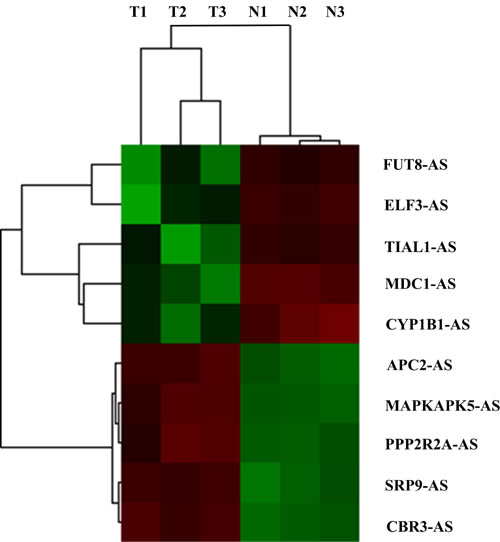
Figure 1: Hierarchical clustering analysis on the most 10 significantly dysregulated cancer-related antisense lncRNAs resulted from microarray assay. All the antisense lncRNAs were named according to their adjacent cancer-related coding genes. T1, T2, T3 and N1, N2, N3 represented for cancerous and paired non-cancerous tissues of samples BC1, BC2 and BC3, respectively.
Dysregulation of MDC1-AS and MDC1 expression level
Thirty-eight pairs of bladder cancer tissues and adjacent noncancerous tissues were used to confirm the aberrant expression level of MDC1-AS obtained from microarray assay by RT-PCR. It should be stated that MDC1-AS RT-PCR data were only available in 32 pairs of tissues because the absolute quantity of MDC1-AS is very few and the transcripts were not stable. Consistently, MDC1-AS levels in cancerous tissues were evidently lower than those in the noncancerous tissues (P = 0.001, as shown in Figure 2A).
In addition to MDC1-AS, levels of coding gene MDC1 were also detected in the 38 pairs of tissues. AS shown in Figure 2B, MDC1 was also down-regulated in bladder cancer tissues (P = 0.025), which was concordant with MDC1-AS.
Spearman’s correlation analysis was performed to determine the correlation between MDC1-AS and MDC1 in statistical level. As a result, expression levels of MDC1-AS and MDC1 in tissues were significantly correlated in a positive direction (P < 0.001, Figure 2C), indicating a potential regulating role of MDC1-AS on the coding gene MDC1.
Furthermore, expression levels of MDC1-AS and MDC1 were also detected in two bladder cancer cell lines T24 and EJ. Consistent with the dysregulated tendency in tissues, both MDC1-AS and MDC1 were decreased in bladder cancer cells, compared with those in normal cell line 293A (Figure 2D).
Down-regulation of coding gene MDC1 was further confirmed in protein level. As a result, bladder cancer tissues showed observably reduced MDC1 protein levels than noncancerous tissues.
In addition, we analyzed the association between MDC1-AS and MDC1 expression levels and clinical factors of bladder cancer patients. However, no significant association was found in this analysis (data not shown).
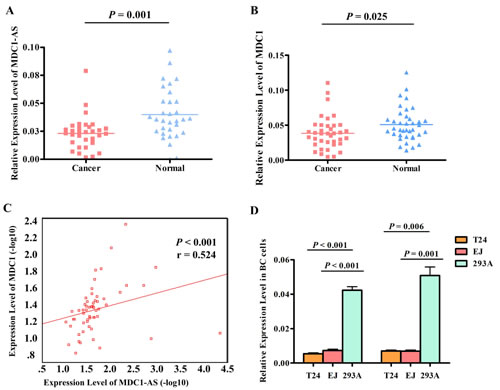
Figure 2: Dysregulation of MDC1-AS and MDC1 expression level. (A) MDC1-AS levels in cancerous tissues were significantly lower than those in the noncancerous tissues. (B) MDC1 was down-regulated in bladder cancerous tissues, compared with that in adjacent normal tissues. (C) Expression levels of MDC1-AS and MDC1 in tissues were significantly correlated in a positive direction. (D) Both MDC1-AS and MDC1 were decreased in bladder cancer cells T24 and EJ, compared with those in normal cell line 293A.
Coding capability prediction and subcellular location of MDC1-AS
Results of coding capability prediction using MDC1-AS sequence suggested that MDC1-AS did not possess the ability to code any protein (Supplementary Figure 1, coding probability of MDC1-AS was 0.138).
EJ cells were separated into nuclear and cytoplasmic fractions to determine the cellular location of both MDC1-AS and MDC1. As presented in Supplementary Figure 2, we found that both MDC1-AS and MDC1 were exist primarily in the nuclear fraction. We attribute this phenomenon to the DNA repair function of MDC1. Besides, the same location of MDC1-AS and MDC1 indicated potential interrelations between them.
Regulating role of MDC1-AS on MDC1 expression level
To investigate whether MDC1-AS regulate expression level of protein coding gene MDC1, we enhanced MDC1-AS expression by transfecting an expression vector containing MDC1-AS into T24 and EJ cells, using pcDNA3.1 empty vector as negative control. Twenty-four hours and 48 h after transfection, levels of MDC1-AS and MDC1 in these two cell lines were detected. Results revealed that MDC1-AS was significantly over-expressed (Figure 3A), indicating a success in our transfection and over-expression of MDC1-AS in bladder cancer cells.
With the over-expression of MDC1-AS, up-regulation of coding gene MDC1 was also observed in both RNA (Figure 3B) and protein levels. Besides, this up-regulation was repeatedly confirmed in two cell lines, suggesting that MDC1-AS may exert a positive effect on expression of MDC1.
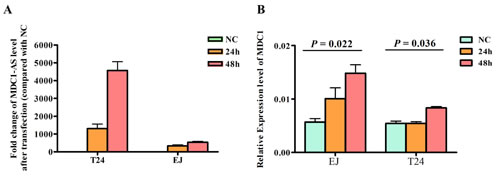
Figure 3: Regulating role of MDC1-AS on MDC1 expression level. (A) Over-expression of MDC1-AS in bladder cancer cells T24 and EJ with transfection of vectors, value of NC group was set as 1 and not presented in this figure. (B) Levels of MDC1 mRNA in cells were significantly increased after transfected with MDC1-AS over-expressed vector for 48h.
Inhibitory role of MDC1-AS on cell proliferation and colony formation
The significant down-regulation of MDC1-AS in bladder cancer tissues and cells prompted us to explore the potential biological functions of MDC1-AS in carcinogenesis. Firstly, CCK-8 assay showed that proliferation of bladder cancer cells EJ and T24 were remarkably inhibited after MDC1-AS expression was enhanced for 24h and 48h, compared with those transfected with NC vectors (Figure 4A and Supplementary Table 2, P < 0.05 in all the experimental groups).
As colony formation was another important indicator for the viability of cells, we further detect effect of MDC1-AS on cells colony formation. In accordance with t he proliferation assay, observations in colony formation assay also indicated that EJ and T24 cells clonogenic survival was evidently suppressed following over-expression of MDC1-AS (Figure 4B and 4C, P = 0.032 for EJ cells and P = 0.007 for T24 cells).
Inhibitory role of MDC1-AS on cell migration and invasion
Following the verification of inhibitory role of MDC1-AS on bladder cancer cells viability, we further investigate whether this lncRNA participant in cells migration and invasion. Compared with EJ and T24 cells transfected with NC vector, cells over-expressing MDC1-AS for 48h showed significantly decreased ability in migration (Figure 4D and 4E, P = 0.006 for EJ cells and P = 0.003 for T24 cells). Besides, the suppression effect on migration was also observed in T24 cells after 24h of transfection (P = 0.018).
Subsequently, in the evaluation of cells invasion, T24 cells exhibited significant impairment of invasion ability after transfected with MDC1-AS for 48h (Figure 4F and 4G, P = 0.007). However, no significant difference was observed in EJ cells.
In addition to the malignant behaviors mentioned above, we also analyzed cell cycle progression in cells treated with MDC1-AS and NC vectors. However, there were no distinct changes in the distribution of cells in each cell cycle phase, indicating that the inhibitory effect of MDC1-AS on cell proliferation was irrelevant to cell cycle arrest (shown in Figure 4H)
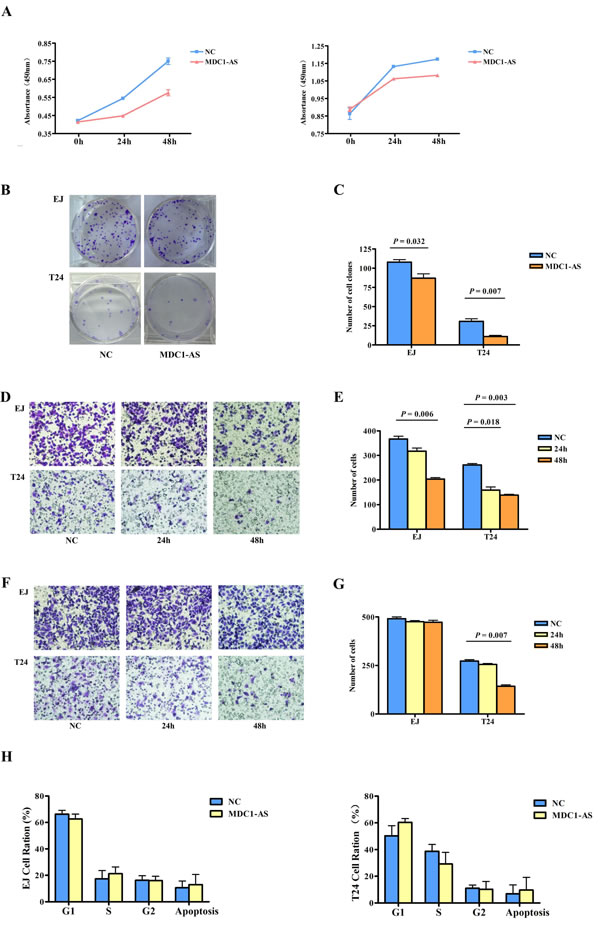
Figure 4: Inhibitory role of MDC1-AS on malignant cell behavior. (A) Proliferation of bladder cancer cells EJ (left) and T24 (right) were significantly decreased after MDC1-AS expression was enhanced for 24h and 48h. (B) Colony formation of EJ and T24 cells was evidently suppressed following MDC1-AS over-expression and (C) Statistical chart of clone number. (D) Migration of EJ and T24 cells were decreased after over-expressing of MDC1-AS and (E) Number of migrated cells. (F) Significant impairment of T24 cell invasion after transfected with MDC1-AS for 48h and (G) Number of invaded cell. (H) Distribution of cells in each cell cycle phase was not changed after MDC1-AS over-expressing.
Knockdown of MDC1 expression by siRNA
In order to determine whether the observed inhibitory effects of MDC1-AS was the consequence of its up-regulation of coding gene MDC1, knockdown of MDC1 expression was achieved by siRNA interference. RT-PCR assay of the interfered cells showed that MDC1 level was markedly decreased, except for si-MDC1-3 (Supplementary Figure 3). In the following study, siMDC1 with the highest inhibition ratio up to 80% (i.e. si-MDC1-1, P = 0.025) was selected. Malignant phenotypes were monitored repeatedly in EJ and T24 cells with both MDC1-AS over-expressing and MDC1 knockdown.
Repeating observation on cell proliferation, colony formation, migration and invasion
In the repeated CCK-8 assay, we found that inhibitory role of MDC1-AS on bladder cancer cells proliferation was evidently weakened with co-transfection of siMDC1. As shown in Figure 5A and Supplementary Table 3, there was no significant change in proliferation ratio of treated EJ and T24 cells, compared with the NC groups.
Similarly, suppression on cell colony formation was also attenuated by siMDC1, that is, MDC1-AS did not have the ability to inhibit colony formation of bladder cancer cells any longer after MDC1 was knockdown (Figure 5B and 5C).
Furthermore, observation on cell migration and invasion were also repeated and time points were selected according to time finding significant differences in the previous study. As a result, no differences were found between co-transfected cells and cells with NC vector (Figure 5D - 5G), indicating that siMDC1 also diminishes suppressing role of MDC1-AS on cells migration and invasion.
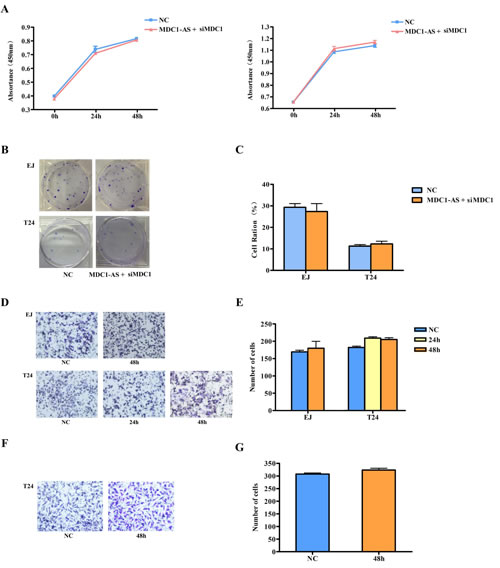
Figure 5: Repeating observation on malignant cell behavior after co-transfected with MDC1-AS and siMDC1. Inhibitory role of MDC1-AS on malignant cell phenotype, i.e. (A) proliferation (the left was EJ cell and the right was T24 cell); (B) and (C) colony formation; (D) and (E) migration; (F) and (G) invasion, was diminished after MDC1 was knockdown.
Discussion
Human Genome Project has revealed that most of the transcriptional output of the whole genome was lncRNAs, which were far more than the coding transcripts and short noncoding RNAs [3]. The huge quantity also indicated fundamental functions of lncRNAs. Studies have demonstrated that an important category of lncRNAs was transcripts from the opposite strand to a protein coding gene, called antisense lncRNAs [15, 17]. Along with the development of relevant researches, antisense lncRNAs were suggested to function as epigenetic regulators of transcription through various mechanisms [17-18]. They have been identified to hybridize with the coding strand or their antisense transcripts, with the consequence of interfering with transcription or stability of mRNA [15]. Several cancer-related genes, such as p15 [19] p53 [20] and Myc [21] have already been demonstrated to be regulated by their antisense lncRNAs. However, determine the detailed mechanism of the large number of antisense lncRNAs with unknown function was still a huge challenge to our understanding of the regulation on human genome.
In the present study, microarray analysis of lncRNAs expression revealed a set of lncRNAs expressing differently between bladder cancer tissues and adjacent normal tissues, indicating the participation of lncRNAs in carcinogenesis of bladder cancer. This result further promoted us to detect deeply into the detailed functions of the abnormally expressed lncRNAs.
MDC1 is an important mediator of the repair of double-strand breaks (DSB) with an amino-terminal forkhead-associated (FHA) domain and a tandem repeat of breast cancer susceptibility gene-1 carboxy terminus (BRCT) domains [22]. In response of DSB damage, MDC1 was activated by ataxia telangiectasia mutated (ATM) protein kinase and recruit DNA damage repair (DDR) factors to DSB sites [23]. It is well known that unrepaired DSB will result in genome instability and promote apoptosis or oncogenesis [24]. Therefore, involvement of DDR factors, including MDC1, in the mechanism of carcinogenesis has aroused great concern. For instance, MDC1-knockout mice presented susceptibility to cancer [25] and lack of MDC1 expression has been observed in various human cancers [26]. In general, MDC1 acts as a strong tumor suppressor through its DDR function.
Given the vital role of antisense lncRNAs on regulating their neighboring coding genes and the definite tumor-suppressing role of MDC1, we focused on MDC1-AS in the following study and hypothesized that this antisense lncRNA may take part in bladder cancer through regulation of MDC1.
Firstly, RT-PCR detection in a larger size of tissues verified the significant down-regulation of lncRNA MDC1-AS and the reduction of MDC1-AS expression was also observed in bladder cancer cells EJ and T24, suggesting that MDC1-AS may act as a tumor suppressor in bladder cancer. Besides, expression of coding gene MDC1 in bladder cancer tissues also showed similar trend with MDC1-AS, compared with those in non-tumor tissues. This observation was consistent with previous studies reporting decreased MDC1 level in cancerous tissues [26-27] and provided further support for tumor suppressing role of MDC1. In Spearman’s correlation analysis, evident correlation was found between expression levels of MDC1-AS and MDC1. Therefore, lncRNA MDC1-AS was likely to exert positive regulatory effect on its neighboring coding gene MDC1. However, we didn’t observe significant associations between MDC1-AS and MDC1 expression level and clinical factors of bladder cancer, we speculated that this may be due to relatively small size of our tissue samples. Further bioinformatic prediction revealed that sequence of MDC1-AS did not code for protein, this result provided further support for the characteristic of MDC1-AS as a long noncoding RNA, in addition to the database information. (http://asia.ensembl.org/Homo_sapiens/Transcript). Subsequently, elevated expression level of MDC1 was observed in MDC1-AS over-expressing cells. These findings suggested that MDC1-AS can up-regulate coding gene MDC1 in both mRNA and protein level, thus verified the assuming role of MDC1-AS mentioned above.
After demonstrating regulatory role of MDC1-AS on tumor suppressing gene MDC1, we detected the influence of it on bladder cancer cells. In a series assays on malignant cell behaviors, cancer cells over-expressing MDC1-AS exhibited significant decreases in proliferation, colony formation, migration and invasion. The observed reduction in malignant cell behavior provided evidence for the tumor suppressing role of MDC1-AS, which were consistent with our conjecture. Therefore, the significant down-regulation of MDC1-AS observed in bladder cancer tissues and cells can be interpreted: cells in cancer tissues showed lower level of MDC1-AS, thus their malignant phenotypes were more evident than those in normal tissues.
At this point, our study has found an up-regulating effect of MDC1-AS on coding gene MDC1, and distinct suppressing roles of this lncRNA on cancer cells. As MDC1 was an identified tumor-suppressing gene, we wondered whether the inhibitory role of MDC1-AS on bladder cancer cells was attribute to up-regulation of MDC1. Therefore, we interfered intracellular expression of MDC1 to investigate on this issue. As a result, cells co-transfected with MDC1-AS and siMDC1 presented similar malignant behavior with NC group, suggesting that inhibitory role of MDC1-AS was remarkably diminished when MDC1 was knockdown. Taken together, it stood to reason that MDC1-AS exhibit an inhibitory role on bladder cancer cells through its up-regulation of a tumor suppressing gene, MDC1.
Recently, more and more studies have investigated into biological effects of genetic variants on lncRNAs. Researchers have found that single nucleotide polymorphisms (SNPs) located in lncRNAs may be associated with cancer risk and modify expression level or biological functions of the lncRNA [28-30]. Therefore, further studies focused on the effects of SNPs in MDC1-AS will be needed to clarify the role of genetic variants on lncRNAs in carcinogenesis of bladder cancer.
There was a limitation of our study. Although we demonstrated the regulation role of MDC1-AS in MDC1 expression, the detailed mechanism underlying this regulation was still not known. According to previous study reported by Faghihi et al., antisense lncRNA may elevate expression level of coding genes through increasing of mRNA stability [31]. Investigation on this issue will be conducted in further studies.
In conclusion, we have identified MDC1-AS as a novel antisense lncRNA through lncRNA microarray approach. This lncRNA was down-regulated in bladder cancer tissues, and exerted a positive effect on expression of coding gene MDC1. In addition, MDC1-AS significantly inhibited malignant phenotype of bladder cancer cells, through up-regulation of MDC1. Further studies should be conducted to investigate the detailed mechanism of our findings.
Material and Methods
Patient specimens and clinical assessments
In general, 41 pairs of bladder cancer tissues and adjacent normal bladder tissues were collected in the present study. All of the samples were obtained from the First Affiliated Hospital of Nanjing Medical University and Jiangsu Province Hospital of TCM in Nanjing, China. All samples were frozen in liquid nitrogen immediately after surgical resections. The hematoxylin and eosin (H&E) stained sections prepared using the cancerous and normal tissues were histologically confirmed by the pathologists. Detailed information of the 3 cases randomly selected to be analyzed in microarray platform is shown in Supplementary Table 1. The informed consent was obtained from all the participants and procedures used in this study were approved by the institutional review boards of Nanjing Medical University.
RNA extraction
Total RNA was extracted from all the tissues samples as well as cancer cells using Trizol reagent (Invitrogen, CA, USA) according to the manufacturer’s instruction. NanoDrop ND-1000 spectrophotometer (OD 260 nm, NanoDrop, Wilmington, DE, USA) was used to assess the quantity of RNA, and standard denaturing agarose gel electrophoresis was used to assess RNA integrity.
LncRNA microarray and data analysis
Genome-wide lncRNA expression assay was performed using Human LncRNA Microarray V2.0, which was manufactured by Arraystar Inc (MD, USA). This microarray covered more than thirty-three thousand lncRNAs identified in human genome. Raw signal intensities of the microarray were normalized by GeneSpring GX v12.0, and low intensity LncRNAs were filtered. Passing Volcano Plot filtering was used to identify differentially expressed LncRNAs with statistical significance between the two groups. The microarray hybridization and collection of expression data were performed by KangChen Bio-tech, Shanghai, China.
Hierarchical clustering analysis
For the important regulating role of antisense lncRNAs on their adjacent coding genes, biological functions and their role in carcinogenesis of the adjacent coding genes were retrieved in literatures and we assumed that those lncRNAs with cancer-related adjacent coding genes were most likely to be involved in the occurrence of malignancies. Unsupervised hierarchical clustering analysis of the cancer-related antisense lncRNAs was described in Supplementary Methods.
Quantitative real-time PCR (qRT-PCR)
cDNA was synthesized from total RNA using M-MLV reverse transcriptase (Invitrogen) according to the manufacturer’s protocol. qRT-PCR with SYBR Green assays (TaKaRa Biotechnology, Dalian, China) was used to determine expression level of selected lncRNA MDC1-AS and its adjacent coding gene MDC1. A detailed description of qRT-PCR was provided in Supplementary Methods.
Transfection of MDC1-AS overexpression vector
The vectors expressing MDC1-AS were prepared by amplifying full length of complementary DNA encoding MDC1-AS and the amplified fragments were then cloned into pcDNA 3.1 vector (Invitrogen, Carlsbad, CA, USA). The amplified fragments were then sequenced to confirm that there were no errors in nucleotides. Bladder cells EJ and T24 were transiently transfected with the MDC1-AS overexpression plasmid using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) transfection reagent according to the protocols. The pcDNA3.1 empty vector was used as negative control (NC).
Observation about malignant behaviors of cancer cells
We conducted a series of assays to determine the effects of lncRNA MDC1-AS on the malignant behaviors of bladder cancer cells, e.g. proliferation, colony formation and migration. Detailed assay conditions were described in Supplementary Methods.
RNA interference
To further detect whether the inhibitory effect of MDC1-AS on cancer cells was attribute to the upregulation of coding gene MDC1, artificial downregulation of MDC1 was achieved by small interfering RNA (siRNA). All the 3 siRNAs were targeting nonoverlapping regions of MDC1 cDNA and therefore only knockdown the targeted MDC1 transcript, not MDC1-AS. After measurement of the interference efficiency, si-MDC1-1 (named siMDC1 in the present study) with the highest efficiency was selected for the following study. Sequence of siMDC1 was provided in Supplementary Methods.
Statistical analysis
Independent t-test was used to compare expression level of lncRNAs obtained by microarray as well as continuous data generated in the following functional studies between two groups. Spearman’s correlation analysis was used to detect the correlationship between level of MDC1-AS and MDC1 in biological tissues. All statistical analyses were performed by SPSS for v.13.0 (SPSS, Chicago, IL). P-values of less than 0.05 were considered significant.
There were some other methods and detailed informations listed in Supplementary Methods.
Funding
This work was partly supported by National Natural Science Foundation of China (81473050, 81230068, 81373091, 81202268 and 81102089), Natural Science Foundation of Jiangsu Province (BK2011773, and BK2011775), the Key Program for Basic Research of Jiangsu Provincial Department of Education (12KJA330002 and 11KJB330002), Jiangsu Provincial Six Talent Peaks Project (2012-SWYY-028, and 2012-WSN-30), The research project of Jiangsu Province of TCM (y13031), Specialized Research Fund for the Doctoral Program of Higher Education (20123234110001), Jiangsu Provincial Science and Technology Innovation Team, and the Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (Public Health and Preventive Medicine).
Acknowledgements
Conception and design: Yao Xue and Meilin Wang; acquisition of data: Gaoxiang Ma and Qiuhan Hua; analysis and interpretation of data: Yao Xue and Gaoxiang Ma; drafting of the manuscript: Yao Xue and Meilin Wang; critical reversion of the manuscript: Zhengdong Zhang and Meilin Wang; statistical analysis: Haiyan Chu and Na Tong; obtaining of funding: Zhengdong Zhang and Meilin Wang; sample support: Lin Yuan, Chao Qin and Changjun Yin; supervision: Zhengdong Zhang.
Conflict of interest
Authors declare no conflicts of interest.
References
1. Parkin DM, Bray F, Ferlay J and Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005; 55(2):74-108.
2. Kim WJ and Bae SC. Molecular biomarkers in urothelial bladder cancer. Cancer Sci. 2008; 99(4):646-652.
3. Mercer TR, Dinger ME and Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009; 10(3):155-159.
4. Moran VA, Perera RJ and Khalil AM. Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res. 2012; 40(14):6391-6400.
5. Tee AE, Ling D, Nelson C, Atmadibrata B, Dinger ME, Xu N, Mizukami T, Liu PY, Liu B, Cheung B, Pasquier E, Haber M, Norris MD, Suzuki T, Marshall GM and Liu T. The histone demethylase JMJD1A induces cell migration and invasion by up-regulating the expression of the long noncoding RNA MALAT1. Oncotarget. 2014; 5(7):1793-1804.
6. Chen B, Yu M, Chang Q, Lu Y, Thakur C, Ma D, Yi Z and Chen F. Mdig de-represses H19 large intergenic non-coding RNA (lincRNA) by down-regulating H3K9me3 and heterochromatin. Oncotarget. 2013; 4(9):1427-1437.
7. Perez DS, Hoage TR, Pritchett JR, Ducharme-Smith AL, Halling ML, Ganapathiraju SC, Streng PS and Smith DI. Long, abundantly expressed non-coding transcripts are altered in cancer. Hum Mol Genet. 2008; 17(5):642-655.
8. Sun M, Jin FY, Xia R, Kong R, Li JH, Xu TP, Liu YW, Zhang EB, Liu XH and De W. Decreased expression of long noncoding RNA GAS5 indicates a poor prognosis and promotes cell proliferation in gastric cancer. BMC Cancer. 2014; 14:319.
9. Takahashi K, Yan IK, Wood J, Haga H and Patel T. Involvement of Extracellular Vesicle long non-coding RNA (linc-VLDLR) in Tumor Cell Responses to Chemotherapy. Mol Cancer Res. 2014.
10. Zhang EB, Kong R, Yin DD, You LH, Sun M, Han L, Xu TP, Xia R, Yang JS, De W and Chen J. Long noncoding RNA ANRIL indicates a poor prognosis of gastric cancer and promotes tumor growth by epigenetically silencing of miR-99a/miR-449a. Oncotarget. 2014; 5(8):2276-2292.
11. Zhu Y, Yu M, Li Z, Kong C, Bi J, Li J and Gao Z. ncRAN, a newly identified long noncoding RNA, enhances human bladder tumor growth, invasion, and survival. Urology. 2011; 77(2):510 e511-515.
12. He W, Cai Q, Sun F, Zhong G, Wang P, Liu H, Luo J, Yu H, Huang J and Lin T. linc-UBC1 physically associates with polycomb repressive complex 2 (PRC2) and acts as a negative prognostic factor for lymph node metastasis and survival in bladder cancer. Biochim Biophys Acta. 2013; 1832(10):1528-1537.
13. Liu Z, Wang W, Jiang J, Bao E, Xu D, Zeng Y, Tao L and Qiu J. Downregulation of GAS5 promotes bladder cancer cell proliferation, partly by regulating CDK6. PLoS One. 2013; 8(9):e73991.
14. Pang KC, Frith MC and Mattick JS. Rapid evolution of noncoding RNAs: lack of conservation does not mean lack of function. Trends Genet. 2006; 22(1):1-5.
15. Katayama S, Tomaru Y, Kasukawa T, Waki K, Nakanishi M, Nakamura M, Nishida H, Yap CC, Suzuki M, Kawai J, Suzuki H, Carninci P, Hayashizaki Y, Wells C, Frith M, Ravasi T, et al. Antisense transcription in the mammalian transcriptome. Science. 2005; 309(5740):1564-1566.
16. Yuan SX, Tao QF, Wang J, Yang F, Liu L, Wang LL, Zhang J, Yang Y, Liu H, Wang F, Sun SH and Zhou WP. Antisense long non-coding RNA PCNA-AS1 promotes tumor growth by regulating proliferating cell nuclear antigen in hepatocellular carcinoma. Cancer Lett. 2014; 349(1):87-94.
17. Morris KV and Vogt PK. Long antisense non-coding RNAs and their role in transcription and oncogenesis. Cell Cycle. 2010; 9(13):2544-2547.
18. Morris KV. Long antisense non-coding RNAs function to direct epigenetic complexes that regulate transcription in human cells. Epigenetics. 2009; 4(5):296-301.
19. Yu W, Gius D, Onyango P, Muldoon-Jacobs K, Karp J, Feinberg AP and Cui H. Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature. 2008; 451(7175):202-206.
20. Mahmoudi S, Henriksson S, Corcoran M, Mendez-Vidal C, Wiman KG and Farnebo M. Wrap53, a natural p53 antisense transcript required for p53 induction upon DNA damage. Mol Cell. 2009; 33(4):462-471.
21. Celano P, Berchtold CM, Kizer DL, Weeraratna A, Nelkin BD, Baylin SB and Casero RA, Jr. Characterization of an endogenous RNA transcript with homology to the antisense strand of the human c-myc gene. J Biol Chem. 1992; 267(21):15092-15096.
22. Xu X and Stern DF. NFBD1/MDC1 regulates ionizing radiation-induced focus formation by DNA checkpoint signaling and repair factors. FASEB J. 2003; 17(13):1842-1848.
23. Kobayashi J, Fujimoto H, Sato J, Hayashi I, Burma S, Matsuura S, Chen DJ and Komatsu K. Nucleolin participates in DNA double-strand break-induced damage response through MDC1-dependent pathway. PLoS One. 2012; 7(11):e49245.
24. Ciccia A and Elledge SJ. The DNA damage response: making it safe to play with knives. Mol Cell. 2010; 40(2):179-204.
25. Minter-Dykhouse K, Ward I, Huen MS, Chen J and Lou Z. Distinct versus overlapping functions of MDC1 and 53BP1 in DNA damage response and tumorigenesis. J Cell Biol. 2008; 181(5):727-735.
26. Bartkova J, Horejsi Z, Sehested M, Nesland JM, Rajpert-De Meyts E, Skakkebaek NE, Stucki M, Jackson S, Lukas J and Bartek J. DNA damage response mediators MDC1 and 53BP1: constitutive activation and aberrant loss in breast and lung cancer, but not in testicular germ cell tumours. Oncogene. 2007; 26(53):7414-7422.
27. Motoyama N and Naka K. DNA damage tumor suppressor genes and genomic instability. Curr Opin Genet Dev. 2004; 14(1):11-16.
28. Wu H, Zheng J, Deng J, Hu M, You Y, Li N, Li W, Lu J and Zhou Y. A genetic polymorphism in lincRNA-uc003opf.1 is associated with susceptibility to esophageal squamous cell carcinoma in Chinese populations. Carcinogenesis. 2013; 34(12):2908-2917.
29. Xue Y, Wang M, Kang M, Wang Q, Wu B, Chu H, Zhong D, Qin C, Yin C, Zhang Z and Wu D. Association between lncrna PCGEM1 polymorphisms and prostate cancer risk. Prostate Cancer Prostatic Dis. 2013; 16(2):139-144, S131.
30. Jin G, Sun J, Isaacs SD, Wiley KE, Kim ST, Chu LW, Zhang Z, Zhao H, Zheng SL, Isaacs WB and Xu J. Human polymorphisms at long non-coding RNAs (lncRNAs) and association with prostate cancer risk. Carcinogenesis. 2011; 32(11):1655-1659.
31. Faghihi MA, Modarresi F, Khalil AM, Wood DE, Sahagan BG, Morgan TE, Finch CE, St Laurent G, 3rd, Kenny PJ and Wahlestedt C. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nat Med. 2008; 14(7):723-730.

