Introduction
Gene fusions involving members of the neurotrophic-tyrosine receptor kinase (NTRK) family are established oncogenic drivers across various adult and paediatric cancers [1]. Large real-world studies have reported an NTRK gene fusion prevalence of 0.3% among patients with solid tumors, although this varies by cancer type [2–5]. These fusions are commonly detected in rare cancers such as infantile (congenital) fibrosarcoma and secretory breast carcinoma, with prevalences over 90% [6, 7]. Conversely, they are rare in more common solid tumors such as lung adenocarcinoma (0.10–0.23%) [3, 4, 8], breast cancer (0.13–0.39%) [2, 3, 8], colorectal cancer (0.20–0.31%) [2, 4, 8], bladder cancer (0.34%) [2], and thyroid cancer (1.6–6.0%) [2–4, 8, 9].
The NTRK family includes three genes – NTRK1, NTRK2 and NTRK3 – encoding the transmembrane tropomyosin receptor kinase (TRK) A, B, and C proteins, respectively [10]. Fusion involving the 3’ kinase region of the NTRK gene with the 5’ region of its fusion partner gene (arising from intra-chromosomal or inter-chromosomal rearrangement) results in the formation of a chimeric protein with a kinase domain that is constitutively activated or overexpressed [11, 12]. This, in turn, causes downstream stimulation of cellular proliferation via the RAS/RAF/MAPK pathway [10].
Recent years have seen the landmark, tumor-agnostic regulatory approval of TRK inhibitors (larotrectinib and entrectinib) as treatment for patients with solid tumors harboring an NTRK gene fusion [1], with larotrectinib currently approved across 48 countries. These targeted therapies bind to the kinase domain inhibiting the rearranged chimeric oncoproteins, and have been associated with objective response rates of 71% (larotrectinib) [13] and 54% (entrectinib) [14] in clinical trials of patients with thyroid cancer harboring NTRK gene fusions. Genomic profiling of tumors to identify patients eligible for these drugs is not, however, routine in clinical practice. Estimates of the frequency of NTRK gene fusion in large cohorts of patients with solid tumors come from a limited number of studies. Furthermore, the challenge of recruiting affected patients for follow-up, when these fusions are largely seen in rare cancers, has led to a knowledge gap regarding prognosis, survival, and therapeutic outcome [10] with most current knowledge based on selected patients in clinical trials. We aimed to address this gap by evaluating patients with papillary thyroid cancer (PTC) from Finland using biobank stored samples as a source of genomic data linked to longitudinal hospital electronic health records (EHRs). To our knowledge, this has not been undertaken previously in the context of NTRK gene fusions. We selected PTC for this present study due to its relatively high prevalence of NTRK gene fusions compared with other cancer entities [2, 8], with a reported prevalence between 1.2% and 6.7% [10, 15–18], higher in paediatrics (19.2%–26.0%) [18, 19]. The study objectives were to determine the frequency of NTRK gene fusion in patients with PTC in Finland, and to describe co-occurring genetic alternations, clinical characteristics, disease progression, and treatment pathways in NTRK gene fusion positive patients.
Results
Patients with NTRK gene fusion
A total of 6/389 (1.5%) of the PTC cohort tested positive for NTRK gene fusion. Genomic and other characteristics of these six patients are shown in Table 1; the vital status for all was confirmed by data from Statistics Finland. Three patients (numbers 1, 2 and 6) were diagnosed with PTC outside of the period 2005–2017, and therefore did not have complete EHR data and were not included in the subcohort. Consequently, the subcohort contained 3/234 patients positive for NTRK gene fusion (patients 3, 4, and 5 in Table 1). Four samples had tested positive upon immunohistochemistry (IHC) testing, with three confirmed as positive following NGS; no IHC negative samples were subsequently confirmed as positive (Table 2).
Table 1: Demographic and clinical characteristics of patients with PTC diagnosed between 1993 and 2018 and positive for NTRK gene fusion (n = 6)
| Patient 1* | Patient 2* | Patient 3 | Patient 4 | Patient 5 | Patient 6* | |
|---|---|---|---|---|---|---|
| Demographics | ||||||
| Year of thyroid cancer diagnosis | 1993 | 1993 | 2009 | 2009 | 2011 | 2018 |
| Age group at cancer diagnosis (years)† | 50–54 | 55–59 | 40–44 | 30–34 | 25–29 | 55–59 |
| Lifestyle factors/comorbidities | ||||||
| BMI (kg/m2) | N/A | N/A | 22.4 | 25.8 | 22.6 | 22.98 |
| Smoking status | N/A | N/A | Non-smoker | Current smoker | Non-smoker | Non-smoker |
| Comorbidities | N/A | N/A | None | None | None | None |
| CCI at thyroid cancer diagnosis | N/A | N/A | 1 | 0 | 0 | 0 |
| Tumor characteristics | ||||||
| Stage at diagnosis (AJCC) | NA | NA | Unknown | Unknown | I | Unknown |
| Stage at diagnosis (TNM) | NA | NA | T1N0Mx | T1N1Mx | T1N1aM0 | T1bNxMx |
| Genetic characteristics | ||||||
| NTRK gene | NTRK3 | NTRK3 | NTRK3 | NTRK3 | NTRK3 | NTRK3 |
| NTRK gene fusion partner | EML4 | EML4 | EML4 | RBPMS | ETV6 | ETV6 |
| Genomic co-alteration | None | None | None | RB1 deletion | NSG2::KIF5B | BRAF-V600E detected in the other lobe |
| Cancer treatments | ||||||
| Radiotherapy | N/A | N/A | No | No | No | No |
| Radioactive iodine ablation (RAI-131) | N/A | N/A | Yes | Yes | Yes | No |
| Time to first treatment (months) | N/A | N/A | 13.5 | 4.0 | 2.6 | N/A |
| Number of treatments | N/A | N/A | 1 | 2 | 2 | N/A |
| Chemotherapy | N/A | N/A | No | No | No | No |
| Observation length (years) | 25.8 | 25.2 | 9.8 | 9.1 | 7.0 | 0.8 |
| Vital status at end of follow-up‡ | Alive | Alive | Alive | Alive | Alive | Alive |
Table 2: NTRK gene fusion results from IHC and NGS testing of samples from the subcohort (N = 234)
| NGS positive, n | NGS negative, n | |
|---|---|---|
| IHC positive | 3 | 1 |
| IHC negative | 0 | 228 |
Of the six patients positive for NTRK gene fusion in the study cohort, all tested positive for NTRK3. The STAR-Fusion analyses confirmed the presence of in-frame junction reads in the correct orientation in all cases. Also, in all cases, the NTRK kinase domain was preserved and the ‘left’ and ‘right’ breakpoints of the two genes were identified. Finally, in two of these six cases, reads corresponding to the reciprocal translocation were additionally identified. The gene fusion partner was EML4 in three patients, ETV6 in two patients and RBPMS in one patient. Of the two patients with a ETV6 gene fusion partner, one contained an established PTC oncogenic BRAF-V600E mutation (note, for this patient, ETV6::NTRK3 fusion was determined from a sample from one lobe of the thyroid gland, while the BRAF mutation was determined from a separate sample obtained from the other lobe of the thyroid gland a month later). The other patent with an ETV6 gene fusion partner also exhibited a NSG2::KIF5B fusion of unclear clinical relevance. The patient with the RBPMS gene fusion partner also harbored a RB1 deletion as a genomic co-alteration. Among the six patients positive for NTRK gene fusion, 50% were female, two were in the 55–59 age group at cancer diagnosis, and one each were in the following in age groups: 50–54 years, 40–44 years, 30–34 years, and 25–29 years. Cancer stage was known for one patient who had early-stage disease. Comorbidities at PTC diagnosis were not documented for any patient.
Performance of the IHC assay
Because a large number of subcohort member samples were assessed by both NGS and IHC (Table 2), we were able to evaluate the analytical performance of the latter, taking the NGS findings as the gold standard [20]. Using the best performance criterium “tumor cytoplasm score >1” as the threshold gave a sensitivity of 100.00%, a specificity of 99.57%, a positive predictive value of 75%, a negative predictive value of 100%, and an accuracy of 99.57%. The cytoplasm was the subcellular tumor compartment predominantly stained (mostly weakly). No sample was scored as category ‘3’, and no sample showed perinuclear tumor cell pan-TRK staining. Two samples showed nuclear staining and both samples were NGS positive. However, while the overall accuracy of such a threshold definition would be the same (data not shown), its sensitivity would be lower because one of three NGS positive cases would not be detected. Thus, the threshold defined above was preferred from a clinical perspective. NGS testing also revealed that the two cases with nuclear staining were those where NTRK3 was fused to the transcription factor ETV6 (Patient 5) and to RBPMS (Patient 4) involved in maturation of mRNA, respectively. The case without nuclear staining was the patient where NTRK3 was fused to EML4 (Patient 3), which is located in the nucleus and encodes a protein associated with the mitotic spindle. Thus, as also noted by others [8], the biological role of the fusion partner can influence the subcellular localization of the NTRK fusion chimeric protein.
Characteristics of the subcohort
Of the 234 patients in the subcohort, just over a quarter were male (26%), mean age at thyroid cancer diagnosis was 53 years, and median Charlson Comorbidity Index (CCI) score at diagnosis was 2.0. Characteristics stratified by NTRK gene fusion status (positive/wild type) are shown in Table 3; however, comparisons between the two groups were limited based on there being only three patients in the NTRK fusion positive group. Among patients with NTRK wildtype, 69% were treated with radioactive iodine ablation, 16% with radiotherapy and 12% with chemotherapy, and 86% were still alive at the end of follow-up (median 7.2 years’ observation). Among the three patients in the NTRK fusion positive group, all three (100%) received radioactive iodine ablation (none received chemotherapy, radiotherapy or a TRK inhibitor), and all were alive at the end of follow-up (median 9.12 years’ observation, range 8–10 years).
Table 3: Characteristics of the 234 patient subcohort diagnosed from 2005–2017 with PTC and with NTRK gene fusion data and linked EHR data
| Characteristic | NTRK gene fusion N = 3 n (%) | NTRK wild type N = 231 n (%) |
|---|---|---|
| Age at thyroid cancer diagnosis | ||
| Median (IQR) | 31.0 (15.0) | 56.0 (23.0) |
| 18–39 | 2 (66.7) | 51 (22.1) |
| 50–54 | 1 (33.3) | 60 (26.0) |
| ≥55 | 0 (0) | 120 (52.0) |
| Sex | ||
| Female | 1 (33.3) | 173 (74.9) |
| Male | 2 (66.7) | 58 (25.1) |
| BMI, kg/m2 | ||
| <30 (non-obese) | 3 (100.0) | 117 (50.7) |
| ≥30 (obese) | 0 (0.0) | 42 (18.2) |
| Missing | 0 (0.0) | 72 (31.2) |
| Smoking status | ||
| Current | 1 (33.3) | 45 (19.5) |
| Former | 0 (0.00) | 35 (15.15) |
| Never | 2 (66.7) | 117 (50.7) |
| Missing | 0 (0.0) | 34 (14.7) |
| Charlson Comorbidity Index at diagnosis | ||
| Median (IQR) | 0.0 (1.0) | 2.0 (3.0) |
| 0–2 | 3 (100) | 116 (50.2) |
| 3–4 | 0 (0.0) | 81 (35.1) |
| ≥5 | 0 (0.0) | 34 (14.7) |
| Cancer stage at diagnosis, AJCC* | ||
| I | 1 (33.3) | 69 (29.9) |
| II | 0 (0.0) | 9 (3.9) |
| III | 0 (0.0) | 2 (0.9) |
| IV | 0 (0.0) | 2 (0.9) |
| Unknown | 2 (66.7) | 149 (64.5) |
| Cancer treatments | ||
| Radiotherapy | 0 (0.0) | 36 (15.6) |
| Radioactive iodine ablation | 3 (100.0) | 159 (68.8) |
| Chemotherapy | 0 (0.0) | 28 (12.1) |
| Procedures*, median (IQR) | ||
| Total | 46 (29.0) | 33 (39.0) |
| Before PTC diagnosis | 6 (3.0) | 6 (16.0) |
| After PTC diagnosis | 40 (32.0) | 25 (26.0) |
| Hospitalizations (mean number per patient) | ||
| Visit | 215 (71.7) | 16,669 (72.2) |
| Ward | 14 (4.7) | 1447 (6.3) |
| Deaths/survival | ||
| Length of observation while alive (median, IQR) | 9.12 (2.8) | 7.23 (5.3) |
| Total deaths | 0 (0.0) | 32 (13.9) |
| Deaths within 5 years | 0 (0.0) | 16 (6.9) |
| Death within 10 years | 0 (0.0) | 30 (13.0) |
DISCUSSION
In this population-based study among patients diagnosed with PTC and with an archived PTC tissue sample in the Auria Biobank, the frequency of NTRK gene fusion was 1.5% (6/389). All six patients with an NTRK gene fusion tested positive for NTRK3, and the most common gene fusion partner was EML4 (three patients), followed by ETV6 (two patients) and RBPMS (one patient). We have also demonstrated the ability of biobank–EHR linkage for longitudinal follow-up of these patients enabling their characterization, and evaluation of treatment patterns and survival.
Our study comprising 389 patients and 456 tissue samples provided substantial coverage of patients with thyroid cancers in the Hospital District of Southwest Finland 1993–2018 because there were 722 new thyroid cancer diagnoses in this region during this period [21]. The observed 1.5% frequency of NTRK gene fusion is similar to the prevalence seen in the US Cancer Genome Atlas project [15], where 1.2% (6/484) of patients with PTC (excluding clinically aggressive tumors) harbored an NTRK gene fusion. Both of these estimates are slightly lower than those reported in other large cohorts of patients with PTC from the Czech Republic (6.7%, 57/846) [10], Taiwan (23.%, 12/525) [17], China (3.3%, 12/355) [16], and the Middle East (6.0%, 9/315) [18]. Frequencies of NTRK gene fusions from smaller cohorts have similarly varied by geographical region with estimates of 12.1% (4/33) in Poland [22], 11.8% (9/76) in Italy [23], 5.3% (1/19) in France [24], and 0% (0/14) in China [25]. All six patients positive for NTRK gene fusion in our study harbored rearrangement of the NTRK3 gene, which has been found to be the most frequent NTRK gene fusion in other PTC cohorts [9, 10, 15, 17, 18, 26]. NTRK1 gene fusions are also established in thyroid cancer, albeit less commonly than NTRK3 gene fusions [3, 10]; for example, Pekova et al. [10, 16] found that NTRK3 gene fusions were five times more common than NTRK1 gene fusions in their cohort of 846 patients with PTC. No patient in our study cohort had a NTRK2 gene fusion, consistent with the lack of reported cases in thyroid cancer in the literature. The NTRK3 fusion partners found in our study – EML4, ETV6, and RBPMS – have all been previously documented in the literature [10, 15–17, 19, 26, 27], especially ETV6::NTRK3 gene fusions, seemingly the most common detected NTRK gene fusion detected in other PTC cohorts [10, 15, 26]. Other NTRK gene fusions reported in other studies include SQSTM1::NTRK3 [10, 16, 26], ERC1::NTRK3 [17], IRF2BP2::NTRK1 [10, 16], TPR::NTRK1 [10, 19, 26], TPM3::NTRK1 [10], and SQSTM1::NTRK1 [10, 26]. Interestingly, one of the two patients in our cohort with a ETV6::NTRK3 gene fusion had a BRAF-V600E genomic alteration detected from a tissue sample originating from the other lobe of the thyroid gland than the sample from which the ETV6::NTRK3 fusions was determined. Pekova et al. [10] similarly describes a case where a patient with multifocal PTC harboring an ETV6::NTRK3 gene fusion in a nodule in the right thyroid lobe had a BRAF-V600E mutation detected in a nodule in the left thyroid lobe. While intratumor heterogeneity is a well-known phenomenon [28, 29], it is important to note there was no co-occurrence of the two oncogenes NTRK gene fusion and BRAF mutation in the same lobe of the thyroid gland.
Following the introduction of TRK inhibitors for the treatment of patients with solid tumors harboring an NTRK gene fusion, observational cohorts of patients receiving these drugs are required to evaluate treatment outcomes in real-world settings. None of the six patients with PTC positive for NTRK gene fusion in the cohort received TRK inhibitor therapy because they did not have metastatic disease; additionally, these drugs were not approved in Finland at that time. Outside of clinical trials, the effectiveness of larotrectinib in patients with PTC harboring a NTRK gene fusion has thus far been demonstrated in case studies [30–32]. Our present study demonstrates the ability to create a clinicogenomic dataset through harnessing biobank sample derived genomic data, linked EHRs and vital status records, and to describe and follow-up such patients to evaluate natural history, real-world treatment outcomes, and prognosis. It shows the potential to conduct larger studies of this kind in future involving the evaluation of real-world TRK treatment outcomes. These could be among larger cohorts of patients with PTC (through ongoing identification of more NTRK fusion patients) as well as potentially among patients with other cancer types. The median follow-up of the three patients with an NTRK gene fusion in the subcohort at the time of the analysis was over 9 years, which indicates a good length of longitudinal data to observe long-term outcomes, treatment patterns and prognosis. However, as the availability of EHRs (rather than paper-based from records) were largely available from 2005 onwards, this led to a smaller subcohort being described.
Strengths of this study are the population-based sample representative of the region of Finland from which it was drawn meaning our findings have good generalizability, and the long follow-up. The information included in the data sources enabled the description of a wide range of patient characteristics including tumor features and treatments received. Turku University Hospital covers cancer care in the region, and the hospital EHR database includes both inpatient and outpatient data, thus the level of data completeness on disease stage and cancer treatments was high in the subcohort. Another strength is the high sensitivity (100%) and specificity (99.57%) of the pan-TRK IHC for detecting NTRK gene fusions, showing that the IHC was able to detect the same number as fusions as NGS, and indicating that NTRK gene fusion misclassification was likely to be minimal. In a previous study that included 571 patients with thyroid cancer, pan-TRK IHC demonstrated 81.8% sensitivity and 100% specificity for the 38 cases where IHC was undertaken [8]. Additionally, sensitivities of 75–95% and specificities of 82–100% for pan-TRK IHC across various tumor types have been reported in the literature [4, 8, 33, 34]. Further, to the best of our knowledge, our subcohort is the largest thyroid cancer cohort yet described where all FFPE samples underwent pan-TRK IHC and NGS confirmation. However, clinical adoption of IHC requires setup of the assay and availability of an experienced pathologist. The main weakness of the study was that the small number of patients testing positive for NTRK gene fusion prevented the identification of other potential gene fusion partners and limited meaningful comparisons with wild-type NTRK patients.
In conclusion, this study showed the frequency of NTRK gene fusions in patients with PTC to be 1.5%. Furthermore, the creation of a clinicogenomic dataset based on genomic data from the Auria biobank linked to longitudinal EHRs and vital statistics is a feasible method of conducting a real-world observational study of patients with PTC, and potentially those with other cancer types, who harbor an NTRK gene fusion.
Materials and Methods
Study design and data source
This was a retrospective cohort study from Finland using tumor genomic data from patients with PTC held in the Auria Biobank linked to hospital EHR and vital status records. These data sources have previously been successful in determining biomarker expression among patients with malignant mesothelioma, and head and neck squamous cell carcinoma, as well as in evaluating their characteristics, prognosis and survival [35, 36]. The Auria Biobank is situated in the Turku region of Finland and was founded as a joint institution between the University of Turku and the hospital districts of Southwest Finland, Satakunta and Vaasa [37]. At the end of 2018, the population in the area of Hospital District of Southwest Finland was 481,478, with an average age of 42.2 years for males and 44.9 years for females, and with the majority being of Finnish descent [38]. Patients with cancer in this population are primarily treated at Turku University Hospital.
Launched in 2014, the Auria Biobank stores human biological samples collected for future health/medical research with the donor’s consent. In addition, the biobank stores diagnostic tissue samples that were collected before 1 September 2013 and legally transferred from hospital pathology collections (based on a public announcement procedure with a statement that the samples and information may be used for biobank research, according to Finnish law). These samples are linkable at the individual patient level to hospital EHRs in Turku University Hospital, which contain information on a wide range of patient and tumor characteristics, as described in Supplementary Table 1. Further, the hospital vital status records have high validity and inform the national Statistics Finland vital statistics database. Minimal migration that occurs out of the Turku region, combined with further linkage to the Statistics Finland national vital database, provides the potential for longitudinal observation of patients with minimal loss to follow-up. For this present study, use of the thyroid cancer tumor samples, and other patient data, were approved by Auria Biobank’s Scientific Steering Committee under Decisions AB18-6900 and AB18-9957, Hospital District of Southwest Finland (research permission T278/2018) and by Statistics Finland (research permission TK-53-448-20).
Study cohort and subcohort
Identification of the PTC study cohort and subcohort is depicted in Figure 1. We included all individuals aged ≥18 years with a histologically confirmed diagnosis of PTC in the database of the Department of Pathology, Turku University Hospital (between 1993 and mid-2018) who also had at least one formalin-fixed paraffin-embedded (FFPE) tumor sample available and sufficient for research in the Auria Biobank (a total of 456 FFPE tumor samples from 389 patients). Subsequently, we identified a subcohort of 234 patients whose PTC was diagnosed between 2005 (when the hospital EHR data became most complete) and 2017. Of the 155/389 patients not included in the subcohort, the majority (144 patients) were diagnosed between 1993 and 2004 and so had an incomplete EHR, and 11 patients had incomplete follow-up due to being diagnosed during 2018.
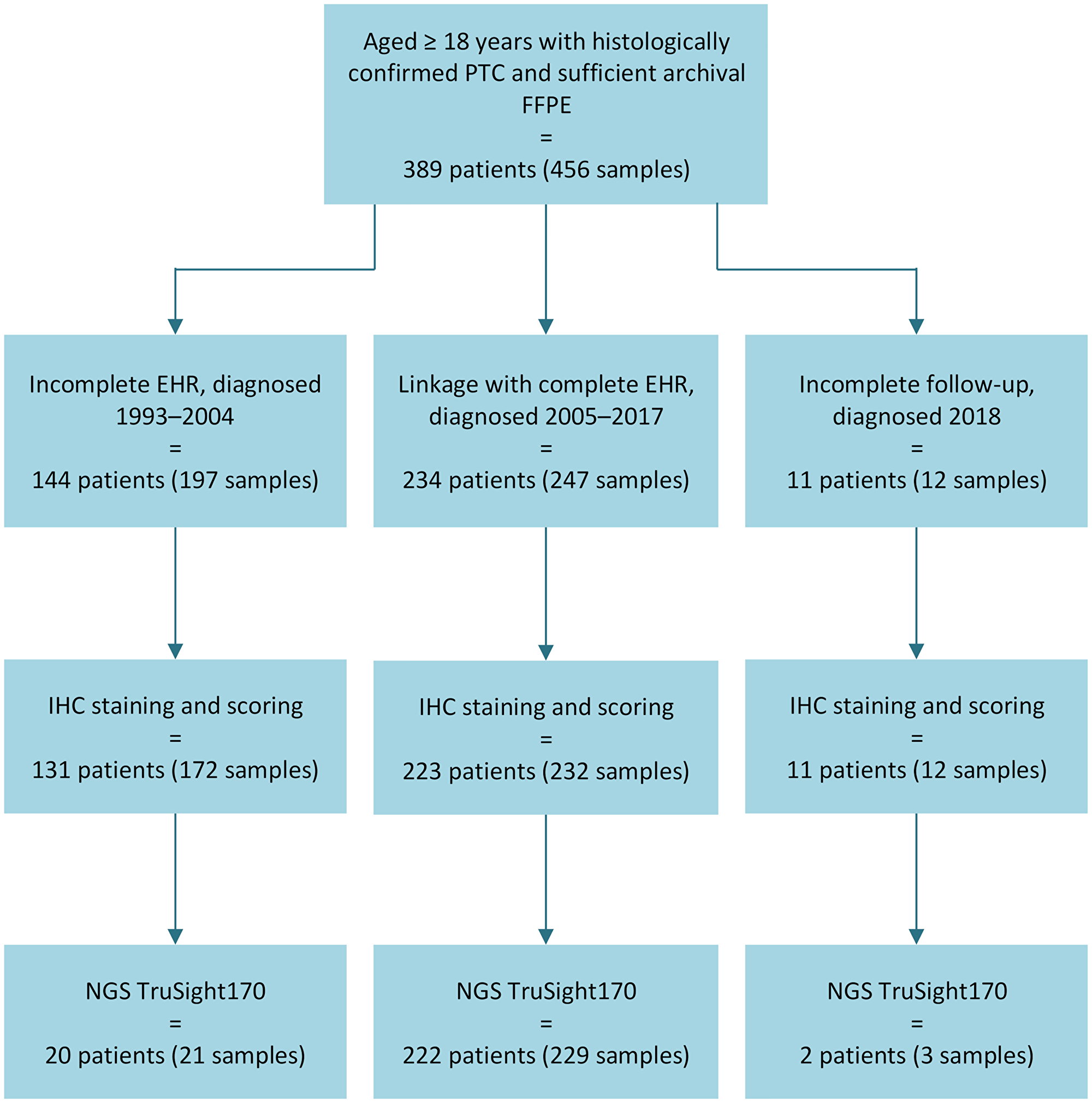
Figure 1: Flowchart depicting patient selection.
Abbreviations: EHR: electronic health record; FFPE: formalin-fixed paraffin-embedded; IHC: immunohistochemistry; NGS: next generation sequencing; PTC: papillary thyroid cancer.
NTRK gene fusion identification
TRK protein expression was determined by pan-TRK IHC, and presence of NTRK gene fusions was confirmed by next generation sequencing (NGS) (Figure 1).
Pan-TRK IHC
Automated IHC was performed on 5 μm histological FFPE sections on a Ventana Discovery autostainer using pan-TRK antibody (clone EPR17341) from Abcam (Cambridge, MA, USA) with DAB detection chemistry as described previously [34]. Positive and negative FFPE samples were used as positive and negative staining run controls. Stained slides were scored by a board-certified pathologist from Dresden University Hospital (KJ) within weeks after staining using the original glass slides. A sample for one patient that could not be scored was excluded (because the tissue was detached from the slide), and a further 39 samples (from 23 patients) that showed no tumor content on the IHC stained slide were also excluded. For each of the remaining 416 samples (from 365 patients), we estimated the percentage of tumor and normal adjacent cells on each slide. Staining was scored in four different categories: ‘0’ for no pan-TRK staining, ‘1’ for weak, ‘2’ for moderate, and ‘3’ for strong. For the normal adjacent area, the percentage of area with the highest score was estimated (e.g., if 20% of the normal adjacent area was scored 1, this implied that 80% was scored 0). The same was undertaken for the tumor area, which was further divided by subcellular compartment (cytoplasmic, membranous, perinuclear, and nuclear). All tumor cells, and not only hot spots, were evaluated.
NGS confirmation
All samples (except one being too scarce) from subcohort members were successfully stained, scored, and underwent NGS confirmation. For members of the study cohort not included in the subcohort (i.e., those without linked EHR data), the samples most strongly stained in the tumor area were selected for NGS confirmation. Next generation sequencing was performed using the TruSight Tumor 170 (TST170) Kit (Illumina, San Diego, CA, USA), which was chosen for its wide coverage across 170 genes associated with solid tumors, detecting single-nucleotide variants/indels, and amplifications from DNA, as well as fusions/splice variants on a subset of genes (including NTRK1/2/3) from RNA. The complete panel of tested genes and content can be found elsewhere [39]. For confirmation of positive samples from pan-TRK IHC and determination of the fusion partner, as well as detection of co-occurring changes in oncogenes, samples nominated for NGS TST170 were analyzed on a Nextseq 500 system at the EN ISO 15189 accredited laboratory Biopticka S.R.O. (Plzeň, Czech Republic). Fusion calling was undertaken using Illumina’s algorithm V2.0.1.8, and correctness of the algorithm was confirmed by concordance with STAR-Fusion [40].
Linkage to EHR and follow-up
We collected information from the hospital EHR database on demographics (age at thyroid cancer diagnosis and sex), comorbidities at cancer diagnosis using the modified Charlson comorbidity diagnosis list [41], CCI, lifestyle variables (body mass index (BMI) and smoking status), cancer treatments (e.g., chemotherapy, radiotherapy), and number of surgeries/hospitalisations (see Supplementary Table 1 for more details). Subcohort members were followed from 1 year before the date of thyroid cancer diagnosis until the end of follow-up, death, or the end of the study.
Statistical analysis
Data analysis was exploratory and descriptive in nature with no pre-specified hypotheses. Frequency of NTRK gene fusion was expressed as a percentage of the total cohort who were confirmed positive for NTRK gene fusion. Characteristics of the subcohort were described using frequency counts and percentages for categorical variables, and medians with interquartile range (IQR) for continuous variables. Descriptions were performed separately for patients positive for NTRK gene fusion and those with wild-type NTRK. Analyses were performed using SAS version 9.4.
Abbreviations
BMI: body mass index; BRAF: proto-oncogene B-Raf; CCI: Charlson Comorbidity Index; EHR: electronic health records; EML: echinoderm microtubule-associated protein-like; ETV: ETS variant transcription factor; KIF: kinesin family; IHC: immunohistochemistry; IQR: inter-quartile range; MAPK: mitogen activated protein kinase; mRNA: messenger ribonucleic acid; NGS: next generation sequencing; NTRK: neurotrophic tyrosine receptor kinase; PTC: papillary thyroid cancer; RAF: rapidly accelerated fibrosarcoma; RAS: rat sarcoma; RB: retinoblastoma tumor suppressor; RBPMS: RNA binding protein with multiple splicing; TRK: tropomyosin receptor kinase.
ACKNOWLEDGMENTS
We thank Susan Bromley of EpiMed Communications Ltd (Abingdon, UK) for medical writing assistance funded by Bayer AG and in accordance with Good Publication Practice. We also thank Dr Petr Martinek at Biopticka SRO for NGS analyses, and Dr Atanas Kamburov, formerly with Bayer AG, for bioinformatics support.
CONFLICTS OF INTEREST
WZ, RSR, AAS, and JHZ are all employees of Bayer. AAS also holds stocks with Bayer AG. BF was an employee of Bayer at the time the study was carried out. KJ was a medical advisor for Provitro AG, Berlin, Germany at the time the study was carried out, and is currently an employee of the Institute of Pathology, Chemnitz Hospital, Chemnitz, Germany. EA has received expert and lecture fees from Bayer outside of this study. REK, MP, NP are employees of the Auria Biobank, University of Turku and Turku University Hospital, which received research funding from Bayer AG for this study; MT was an employee of the Auria Biobank at the time the study was carried out.
ETHICAL STATEMENT
Research use of samples and data was covered by the approvals of Auria Biobank Scientific Steering Committee and Auria Biobank decisions (AB18-6900 and AB18-9957) and permissions of Hospital District of Southwest Finland (T278/2018) and by Statistics Finland (TK-53-448-20).
CONSENT
Auria Biobank stores human biological samples and related data collected for healthcare and/or routine medical examinations for future research based on donor’s consent or legal transfer to biobank according to Finnish Biobank Act (688/2012).
FUNDING
This work was supported by Bayer AG (grant number not applicable).
References
1. Drilon A. TRK inhibitors in TRK fusion-positive cancers. Ann Oncol. 2019 (Suppl 8); 30:viii23–30. https://doi.org/10.1093/annonc/mdz282. [PubMed].
2. Westphalen CB, Krebs MG, Le Tourneau C, Sokol ES, Maund SL, Wilson TR, Jin DX, Newberg JY, Fabrizio D, Veronese L, Thomas M, de Braud F. Genomic context of NTRK1/2/3 fusion-positive tumours from a large real-world population. NPJ Precis Oncol. 2021; 5:69. https://doi.org/10.1038/s41698-021-00206-y. [PubMed].
3. Okamura R, Boichard A, Kato S, Sicklick JK, Bazhenova L, Kurzrock R. Analysis of NTRK Alterations in Pan-Cancer Adult and Pediatric Malignancies: Implications for NTRK-Targeted Therapeutics. JCO Precis Oncol. 2018; 2:1–20. https://doi.org/10.1200/PO.18.00183. [PubMed].
4. Gatalica Z, Xiu J, Swensen J, Vranic S. Molecular characterization of cancers with NTRK gene fusions. Mod Pathol. 2019; 32:147–53. https://doi.org/10.1038/s41379-018-0118-3. [PubMed].
5. Rosen EY, Goldman DA, Hechtman JF, Benayed R, Schram AM, Cocco E, Shifman S, Gong Y, Kundra R, Solomon JP, Bardelli A, Scaltriti M, Drilon A, et al. TRK Fusions Are Enriched in Cancers with Uncommon Histologies and the Absence of Canonical Driver Mutations. Clin Cancer Res. 2020; 26:1624–32. https://doi.org/10.1158/1078-0432.CCR-19-3165. [PubMed].
6. Forsythe A, Zhang W, Phillip Strauss U, Fellous M, Korei M, Keating K. A systematic review and meta-analysis of neurotrophic tyrosine receptor kinase gene fusion frequencies in solid tumors. Ther Adv Med Oncol. 2020; 12:1758835920975613. https://doi.org/10.1177/1758835920975613. [PubMed].
7. Bourgeois JM, Knezevich SR, Mathers JA, Sorensen PH. Molecular detection of the ETV6-NTRK3 gene fusion differentiates congenital fibrosarcoma from other childhood spindle cell tumors. Am J Surg Pathol. 2000; 24:937–46. https://doi.org/10.1097/00000478-200007000-00005. [PubMed].
8. Solomon JP, Linkov I, Rosado A, Mullaney K, Rosen EY, Frosina D, Jungbluth AA, Zehir A, Benayed R, Drilon A, Hyman DM, Ladanyi M, Sireci AN, Hechtman JF. NTRK fusion detection across multiple assays and 33,997 cases: diagnostic implications and pitfalls. Mod Pathol. 2020; 33:38–46. https://doi.org/10.1038/s41379-019-0324-7. [PubMed].
9. Koopman B, Kuijpers CCH, Groen HJM, Timens W, Schuuring E, Willems SM, van Kempen LC. Detection of NTRK Fusions and TRK Expression and Performance of pan-TRK Immunohistochemistry in Routine Diagnostics: Results from a Nationwide Community-Based Cohort. Diagnostics (Basel). 2022; 12:668. https://doi.org/10.3390/diagnostics12030668. [PubMed].
10. Pekova B, Sykorova V, Mastnikova K, Vaclavikova E, Moravcova J, Vlcek P, Lastuvka P, Taudy M, Katra R, Bavor P, Kodetova D, Chovanec M, Drozenova J, et al. NTRK Fusion Genes in Thyroid Carcinomas: Clinicopathological Characteristics and Their Impacts on Prognosis. Cancers (Basel). 2021; 13:1932. https://doi.org/10.3390/cancers13081932. [PubMed].
11. Lange AM, Lo HW. Inhibiting TRK Proteins in Clinical Cancer Therapy. Cancers (Basel). 2018; 10:105. https://doi.org/10.3390/cancers10040105. [PubMed].
12. Amatu A, Sartore-Bianchi A, Siena S. NTRK gene fusions as novel targets of cancer therapy across multiple tumour types. ESMO Open. 2016; 1:e000023. https://doi.org/10.1136/esmoopen-2015-000023. [PubMed].
13. Waguespack SG, Drilon A, Lin JJ, Brose MS, McDermott R, Almubarak M, Bauman J, Casanova M, Krishnamurthy A, Kummar S, Leyvraz S, Oh DY, Park K, et al. Efficacy and safety of larotrectinib in patients with TRK fusion-positive thyroid carcinoma. Eur J Endocrinol. 2022; 186:631–43. https://doi.org/10.1530/EJE-21-1259. [PubMed].
14. Demetri GD, De Braud F, Drilon A, Siena S, Patel MR, Cho BC, Liu SV, Ahn MJ, Chiu CH, Lin JJ, Goto K, Lee J, Bazhenova L, et al. Updated Integrated Analysis of the Efficacy and Safety of Entrectinib in Patients With NTRK Fusion-Positive Solid Tumors. Clin Cancer Res. 2022; 28:1302–12. https://doi.org/10.1158/1078-0432.CCR-21-3597. [PubMed].
15. Cancer Genome Atlas Research Network. Integrated genomic characterization of papillary thyroid carcinoma. Cell. 2014; 159:676–90. https://doi.org/10.1016/j.cell.2014.09.050. [PubMed].
16. Liang J, Cai W, Feng D, Teng H, Mao F, Jiang Y, Hu S, Li X, Zhang Y, Liu B, Sun ZS. Genetic landscape of papillary thyroid carcinoma in the Chinese population. J Pathol. 2018; 244:215–26. https://doi.org/10.1002/path.5005. [PubMed].
17. Lee YC, Chen JY, Huang CJ, Chen HS, Yang AH, Hang JF. Detection of NTRK1/3 Rearrangements in Papillary Thyroid Carcinoma Using Immunohistochemistry, Fluorescent In Situ Hybridization, and Next-Generation Sequencing. Endocr Pathol. 2020; 31:348–58. https://doi.org/10.1007/s12022-020-09648-9. [PubMed].
18. Kong Y, Bu R, Parvathareddy SK, Siraj AK, Siraj N, Al-Sobhi SS, Al-Dayel F, Al-Kuraya KS. NTRK fusion analysis reveals enrichment in Middle Eastern BRAF wild-type PTC. Eur J Endocrinol. 2021; 184:503–11. https://doi.org/10.1530/EJE-20-1345. [PubMed].
19. Prasad ML, Vyas M, Horne MJ, Virk RK, Morotti R, Liu Z, Tallini G, Nikiforova MN, Christison-Lagay ER, Udelsman R, Dinauer CA, Nikiforov YE. NTRK fusion oncogenes in pediatric papillary thyroid carcinoma in northeast United States. Cancer. 2016; 122:1097–107. https://doi.org/10.1002/cncr.29887. [PubMed].
20. Marchiò C, Scaltriti M, Ladanyi M, Iafrate AJ, Bibeau F, Dietel M, Hechtman JF, Troiani T, López-Rios F, Douillard JY, Andrè F, Reis-Filho JS. ESMO recommendations on the standard methods to detect NTRK fusions in daily practice and clinical research. Ann Oncol. 2019; 30:1417–27. https://doi.org/10.1093/annonc/mdz204. [PubMed].
21. Finnish Cancer Registry. Cancer Statistics. https://cancerregistry.fi/statistics/cancer-statistics/ Accessed 31 Jan 31 2023. 2018.
22. Brzeziańska E, Karbownik M, Migdalska-Sek M, Pastuszak-Lewandoska D, Włoch J, Lewiński A. Molecular analysis of the RET and NTRK1 gene rearrangements in papillary thyroid carcinoma in the Polish population. Mutat Res. 2006; 599:26–35. https://doi.org/10.1016/j.mrfmmm.2005.12.013. [PubMed].
23. Bongarzone I, Vigneri P, Mariani L, Collini P, Pilotti S, Pierotti MA. RET/NTRK1 rearrangements in thyroid gland tumors of the papillary carcinoma family: correlation with clinicopathological features. Clin Cancer Res. 1998; 4:223–28. [PubMed].
24. Said S, Schlumberger M, Suarez HG. Oncogenes and anti-oncogenes in human epithelial thyroid tumors. J Endocrinol Invest. 1994; 17:371–19. https://doi.org/10.1007/BF03349004. [PubMed].
25. Kuo CS, Lin CY, Hsu CW, Lee CH, Lin HD. Low frequency of rearrangement of TRK protooncogene in Chinese thyroid tumors. Endocrine. 2000; 13:341–44. https://doi.org/10.1385/ENDO:13:3:341. [PubMed].
26. Chu YH, Dias-Santagata D, Farahani AA, Boyraz B, Faquin WC, Nosé V, Sadow PM. Clinicopathologic and molecular characterization of NTRK-rearranged thyroid carcinoma (NRTC). Mod Pathol. 2020; 33:2186–97. https://doi.org/10.1038/s41379-020-0574-4. [PubMed].
27. Leeman-Neill RJ, Kelly LM, Liu P, Brenner AV, Little MP, Bogdanova TI, Evdokimova VN, Hatch M, Zurnadzy LY, Nikiforova MN, Yue NJ, Zhang M, Mabuchi K, et al. ETV6-NTRK3 is a common chromosomal rearrangement in radiation-associated thyroid cancer. Cancer. 2014; 120:799–807. https://doi.org/10.1002/cncr.28484. [PubMed].
28. Marusyk A, Janiszewska M, Polyak K. Intratumor Heterogeneity: The Rosetta Stone of Therapy Resistance. Cancer Cell. 2020; 37:471–84. https://doi.org/10.1016/j.ccell.2020.03.007. [PubMed].
29. Ramón Y Cajal S, Sesé M, Capdevila C, Aasen T, De Mattos-Arruda L, Diaz-Cano SJ, Hernández-Losa J, Castellví J. Clinical implications of intratumor heterogeneity: challenges and opportunities. J Mol Med (Berl). 2020; 98:161–77. https://doi.org/10.1007/s00109-020-01874-2. [PubMed].
30. Bargas S, Mc Leer A, Mondet J, Chabre O, Laramas M. An impressive response with larotrectinib in a patient with a papillary thyroid carcinoma harboring an SQSTM1-NTRK1 fusion. Eur J Endocrinol. 2022; 186:K5–8. https://doi.org/10.1530/EJE-21-0509. [PubMed].
31. Waguespack SG, Tewari SO, Busaidy NL, Zafereo ME. Larotrectinib Before Initial Radioactive Iodine Therapy in Pediatric TRK Fusion-Positive Papillary Thyroid Carcinoma: Time to Reconsider the Treatment Paradigm for Distantly Metastatic Disease? JCO Precis Oncol. 2022; 6:e2100467. https://doi.org/10.1200/PO.21.00467. [PubMed].
32. Pitoia F. Complete response to larotrectinib treatment in a patient with papillary thyroid cancer harboring an ETV6-NTRK3 gene fusion. Clin Case Rep. 2021; 9:1905–12. https://doi.org/10.1002/ccr3.3900. [PubMed].
33. Hondelink LM, Schrader AMR, Asri Aghmuni G, Solleveld-Westerink N, Cleton-Jansen AM, van Egmond D, Boot A, Ouahoud S, Khalifa MN, Wai Lam S, Morreau H, Bovee JVM, van Wezel T, Cohen D. The sensitivity of pan-TRK immunohistochemistry in solid tumours: A meta-analysis. Eur J Cancer. 2022; 173:229–37. https://doi.org/10.1016/j.ejca.2022.06.030. [PubMed].
34. Hechtman JF, Benayed R, Hyman DM, Drilon A, Zehir A, Frosina D, Arcila ME, Dogan S, Klimstra DS, Ladanyi M, Jungbluth AA. Pan-Trk Immunohistochemistry Is an Efficient and Reliable Screen for the Detection of NTRK Fusions. Am J Surg Pathol. 2017; 41:1547–51. https://doi.org/10.1097/PAS.0000000000000911. [PubMed].
35. Khan M, Khaznadar SS, Routila J, Ventelä S, Schmid E, Gebhart B, Becker ET, Roider HG, Perala M, Schmitz AA, Krahn T, von Ahsen O. Hepatocyte Growth Factor Receptor overexpression predicts reduced survival but its targeting is not effective in unselected HNSCC patients. Head Neck. 2020; 42:625–35. https://doi.org/10.1002/hed.26049. [PubMed].
36. Vizcaya D, Farahmand B, Walter AO, Kneip C, Jöhrens K, Tukiainen M, Schmitz AA. Prognosis of patients with malignant mesothelioma by expression of programmed cell death 1 ligand 1 and mesothelin in a contemporary cohort in Finland. Cancer Treat Res Commun. 2020; 25:100260. https://doi.org/10.1016/j.ctarc.2020.100260. [PubMed].
37. Auria Biobank. https://www.auria.fi/biopankki/en/index.php?lang=en#mika_on_auria_biopankki.
38. Tilastokeskus. Statistics Finland’s free-of-charge statistical databases. https://pxdata.stat.fi:443/PxWeb/sq/e0fc035d-3fd1-4114-b087-870c7fee429b. Accessed 31 January 2023.
40. Haas BJ, Dobin A, Li B, Stransky N, Pochet N, Regev A. Accuracy assessment of fusion transcript detection via read-mapping and de novo fusion transcript assembly-based methods. Genome Biol. 2019; 20:213. https://doi.org/10.1186/s13059-019-1842-9. [PubMed].
41. Quan H, Sundararajan V, Halfon P, Fong A, Burnand B, Luthi JC, Saunders LD, Beck CA, Feasby TE, Ghali WA. Coding algorithms for defining comorbidities in ICD-9-CM and ICD-10 administrative data. Med Care. 2005; 43:1130–39. https://doi.org/10.1097/01.mlr.0000182534.19832.83. [PubMed].

