Introduction
The multi-organ influence of variants in NOD2 and CDKN2A are associated with the risk of cancers occurring in the colon, ovary, breast, lung, larynx, Non-Hodgkin lymphoma and melanoma [1–16]. The NOD2 c.3020insC allele has been shown to occur in 7.3% of the Polish population [1] and could therefore be considered a common genetic risk factor for cancer. In 2005, we investigated 172 bladder and 245 kidney cancer patients [1] and observed nominal evidence for a relationship between the c.3020insC variant and bladder cancer (10.5%; odds ratio OR = 1.5; p = 0.13). We also reported a significantly lower frequency of the c.3020insC allele among renal cancer patients (3.2%; odds ratio OR = 0.4; p = 0.02) [6]. The CDKN2A p.A148T polymorphism is found in 3.5% of the Polish population [2]. In 2006 we determined the frequency of the CDKN2A p.A148T polymorphism in 223 patients with urothelial bladder cancer and 245 patients with kidney cancer [2]. The results of this study revealed the frequency of p. A148T to be similar in bladder or kidney cancer patients compared to controls. To ensure that our original findings were not erroneous and indeed represented the actual frequency found in bladder and kidney cancer patients, we re-evaluated the frequency of the NOD2 c.3020insC allele and the p.A148T polymorphism in CDKN2A among 706 bladder cancer and 410 kidney cancer patients. Due to the size of the patient cohorts we were also able to evaluate the impact of these variants on survival of both the bladder and kidney cancer patients. To our knowledge, this is the first larger-scale study describing the clinical characteristics and survival of bladder and kidney cancer patients that is associated with the NOD2 c.3020insC allele and the CDKN2A p.A148T polymorphism in Poland.
Results
Bladder cancer
NOD2
The NOD2 c.3020insC allele analysis was performed in 706 bladder cancer patients, the characteristics of the study population of bladder cancer are shown in Table 1. The frequency of the c.3020insC allele in the consecutive series of bladder cancer patients was significantly different compared to the control population, the characteristics of the patients with and without the c.c.3020insC allele are presented in Table 2, 63 (8.9%) patients carried the NOD2 variant (OR = 1.4; 95% CI 1.01–1.90; p = 0.04).
Table 1: Characteristics of the study population of bladder cancer (n = 706)
| Gender | |
| Male | 540 |
| Female | 166 |
| Age, mean (range) | 79 (25–91) |
| ≤60 | 222 |
| >61 | 484 |
| Smoking status | |
| Yes | 461 (65%) |
| No | 80 (11%) |
| Missing | 165 |
| Histological features | |
| Noninvasive papillary | 416 (59%) |
| Low grade1 | 250 (35%) |
| High grade2 | 166 (24%) |
| Invasive | 290 (41%) |
| Low | 10 (1%) |
| High | 280 (40%) |
| Stage | |
| Ta | 415 (59%) |
| T1 | 118 (17%) |
| T2 | 84 (12%) |
| T3 | 60 (8%) |
| T4 | 29 (4%) |
| Vital status | |
| Alive | 212 (30%) |
| Dead | 494 (70%) |
Table 2: Clinical characteristics of bladder cancers patients harboring 3020insC allele in NOD2 gene
| Patients with 3020insC allele (63) | Patients with no 3020insC allele (643) | p-value* | Odds ratio (CI) | |
|---|---|---|---|---|
| Age of diagnosis (yr) | ||||
| ≤50 | 3 | 55 | 0.42 | 0.5 0.16–1.76 |
| 51–60 | 23 | 141 | 0.01 | 2.0 1.18–3.53 |
| 61–70 | 14 | 212 | 0.10 | 0.6 0.31–1.07 |
| >71 | 23 | 235 | 1.00 | 1.0 0.58–1.70 |
| Mean | 66 | 79 | ||
| Family history Positive (+) | 3 | 23 | 0.90 | 1.3 0.39–4.62 |
| Smoking | ||||
| no | 10/49 (20) | 70/492 (14) | 0.30 | 1.5 0.73–3.24 |
| ≤20 packyears | 11/49 (23) | 123/492 (25) | 0.80 | 0.9 0.43–1.75 |
| >21 | 28/49 (57) | 299/492 (61) | 0.70 | 0.8 0.47–1.55 |
| Histological features | ||||
| Noninvasive Papillary | ||||
| Low grade | 24/40 (60) | 226/376 (60) | 1.00 | 1.0 0.51–1.94 |
| High grade | 16/40 (40) | 150/376 (40) | 1.00 | 1.0 0.51–1.95 |
| Invasive | ||||
| Low grade | − | 10/267 (4) | − | − |
| High grade | 23/23 (100) | 257/267 (96) | 0.73 | 1.9 0.10–33.7 |
| Stage | ||||
| Ta | 40/63 (63) | 375/643 (58) | 0.50 | 1.2 0.72–2.12 |
| T1 | 12/63 (19) | 106/643 (16) | 0.73 | 1.2 0.61–2.31 |
| T2 | 7/63 (11) | 77/643 (12) | 1.00 | 0.9 0.40–2.09 |
| T3 | 3/63 (5) | 57/643 (9) | 0.34 | 0.5 0.15–1.69 |
| T4 | 1/63 (2) | 28/643 (5) | 0.47 | 0.3 0.04–2.65 |
The study subjects were followed from the date of diagnosis until death or February 2021 (a mean of 21 years). There were 40 deaths (63%) recorded in 63 NOD2 c.3020insC carriers compared with 454 deaths (71%) in 643 non-carriers (HR = 0.94; 95% CI 0.67–1.31; p = 0.7).
The NOD2 c.3020insC allele did not play a significant role in the survival of patients with bladder cancer (Figure 1). The data was stratified for age, smoking status, cancer family history, sex and clinical characteristics. The median survival was 54 months for patients with the NOD2 variant compared to 60 months for non-carriers (Table 3). Among patients with the c.3020insC allele the 10-year survival was 37% compared with 30% for non-carriers. After adjusting for age, smoking status, cancer family history, sex or clinical characteristics, the HR for mortality associated with bladder cancer and the NOD2 variant are presented in Table 4. The HR for patients with positive cancer family history, invasive low grade or T4 couldn’t be calculated, due to low number of patients in those subgroups.
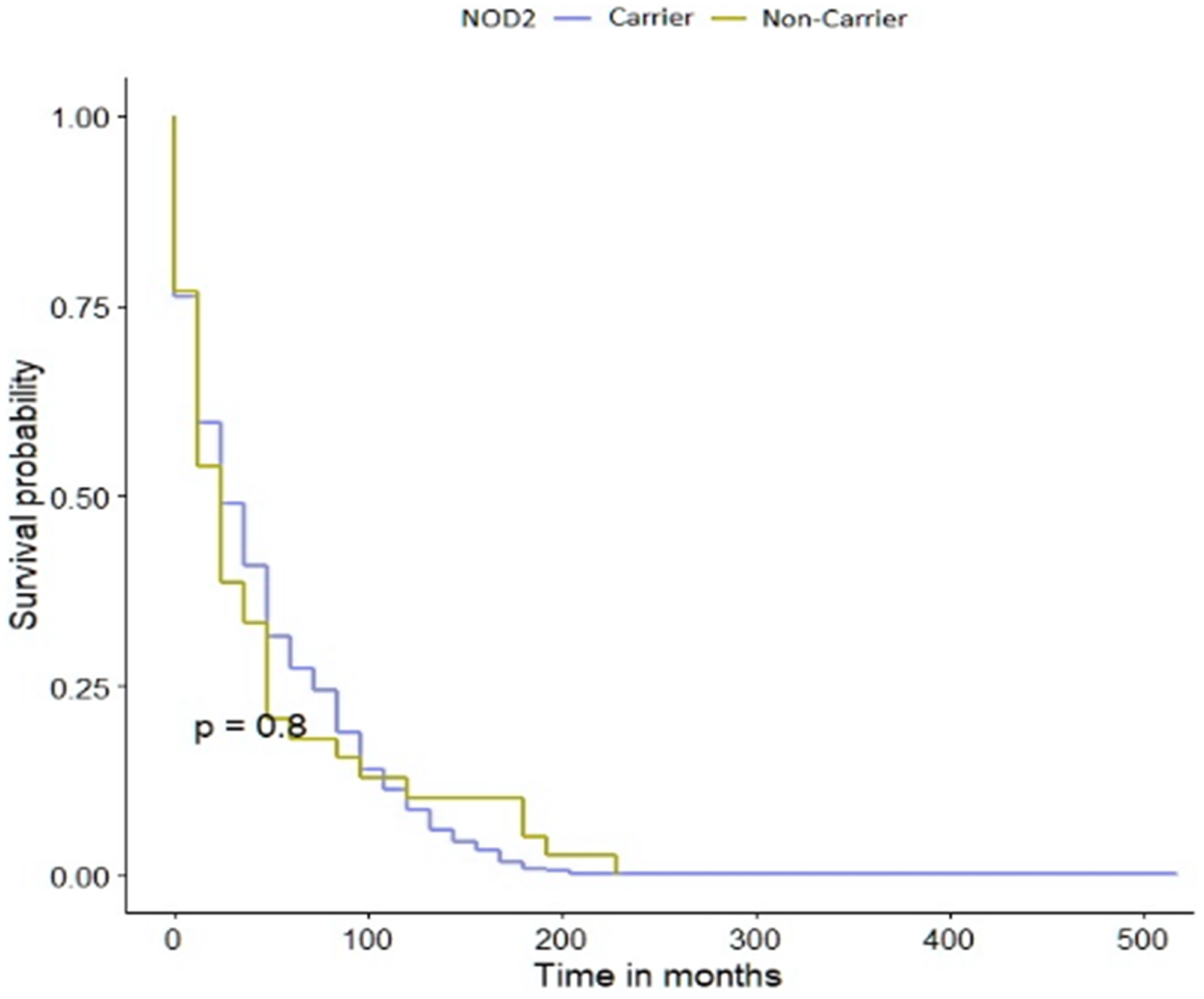
Figure 1: Kaplan-Meier survival curves of bladder cancer patients with NOD2 variant and -negative sub-cohorts.
Table 3: Survival of patients with bladder cancer; by variant of NOD2
| Patients with NOD2 variant (n = 63) | Patients with no variant in NOD2 (n = 643) | |
|---|---|---|
| Median follow-up (mo) | 162 | 156 |
| Proportion of deceased (%) | 63 | 71 |
| Median survival (mo) | 54 | 60 |
| 5-Year survival (%) | 50 | 50 |
| 10-Year survival (%) | 37 | 30 |
| HR | 1.34 | 1.0 |
| 95% CI | 0.89–2.01 | − |
| p-value | 0.2 | − |
Table 4: The HR for mortality associated with bladder cancer and the NOD2 c.3020insC variant
| HR | 95% CI | p-value | |
|---|---|---|---|
| patients younger than 60 years old | 2.14 | 0.89–5.16 | 0.0090 |
| patients older than 60 years old | 1.13 | 0.70–1.84 | 0.6 |
| non-smoking patients | 4.94 | 1.13–21.5 | 0.033 |
| smoking patients | 1.06 | 0.46–2.43 | 0.9 |
| patients with no cancer family history | 1.26 | 0.83–1.91 | 0.3 |
| females | 1.17 | 0.71–1.93 | 0.5 |
| males | 1.69 | 0.74–3.83 | 0.2 |
| patients with noninvasive papillary low grade | 3.11 | 1.38–7.01 | 0.006 |
| patients with noninvasive papillary high grade | 0.88 | 0.39–1.99 | 0.8 |
| patients with invasive high grade | 1.09 | 0.58–2.07 | 0.8 |
| patients with Ta | 1.45 | 0.84–2.51 | 0.2 |
CDKN2A
Of the 706 bladder cancer patients enrolled in the study, 37 (5.2%) carried a p.A148T variant in CDKN2A (OR = 1.5; 95% CI 1.04–2.24; p = 0.04). The characteristics of the study population are shown in Table 1. The characteristics of the patients with and without the polymorphism are presented in Table 5.
Table 5: Clinical characteristics of bladder cancers patients harboring A148T allele in CDKN2A gene
| Patients with 3020insC allele (37) | Patients with no 3020insC allele (669) | p-value* | Odds ratio (CI) | |
|---|---|---|---|---|
| Age of diagnosis (yr) | ||||
| ≤50 | 4 | 54 | 0.78 | 1.4 0.47–4.04 |
| 51–60 | 9 | 155 | 0.87 | 1.1 0.49–2.38 |
| 61–70 | 11 | 215v | 0.90 | 0.9 0.43–1.84 |
| >71 | 13 | 245 | 1.00 | 0.9 0.56–1.87 |
| Mean | 63 | 79 | ||
| Family history Positive (+) | 1 | 25 | 0.74 | 0.7 0.09–5.43 |
| Smoking | ||||
| no | 4/28 (14) | 76/513 (15) | 0.93 | 1.0 0.32–2.84 |
| ≤20 packyears | 7/28 (25) | 166/513 (32) | 0.54 | 0.7 0.29–1.67 |
| >21 | 17/28 (61) | 271/513 (53) | 0.53 | 1.4 0.63–3.00 |
| Histological features | ||||
| Noninvasive Papillary | ||||
| Low grade | 14/25 (56) | 236/391 (60) | 0.80 | 0.8 0.37–1.89 |
| High grade | 11/25 (44) | 155/391 (40) | 0.80 | 1.2 0.53–2.70 |
| Invasive | ||||
| Low grade | − | 10/278 (36) | − | − |
| High grade | 12/12 (100) | 268/278 (96) | 0.50 | 1.0 0.05–17.6 |
| Stage | ||||
| Ta | 25/37 (68) | 390/669 (58) | 0.34 | 1.5 0.73–3.01 |
| T1 | 3/37 (8) | 115/669 (17) | 0.22 | 0.4 0.12–1.40 |
| T2 | 5/37 (13) | 79/669 (12) | 0.95 | 1.1 0.44–3.08 |
| T3 | 1/37 (3) | 59/669 (9) | 0.31 | 0.3 0.04–2.13 |
| T4 | 3/37 (8) | 26/669 (4) | 0.40 | 2.2 0.63–7.57 |
The mean follow-up time was 35 years. There were 23 deaths (62%) in 37 p.A148T carriers versus 471 deaths (70%) in 669 non-carriers (HR = 1.10; 95% CI 0.73–1.68; p = 0.6). The p.A148T allele in CDKN2A gene did not affect the survival of patients with bladder cancer Figure 2. We observed a possible decrease in survival in the subgroup of patients younger than 61 years old carrying the p.A148T variant but this was not statistically significant. The data was stratified for age, smoking status, cancer family history, sex and clinical characteristics. The median survival of patients with CDKN2A polymorphism was the same as for non-carriers, namely 156 months (Table 6). In patients carrying the p.A148T allele the 10-year survival was 27% compared with 31% for non-carriers. After adjusting for age, smoking status, cancer family history, sex or clinical characteristics, the HR for mortality associated with bladder cancer and the CDKN2A variant are presented in Table 7. The HR for patients with positive cancer family history, T1, T3 or invasive low grade disease couldn’t be calculated, due to the low number of patients in those subgroups.
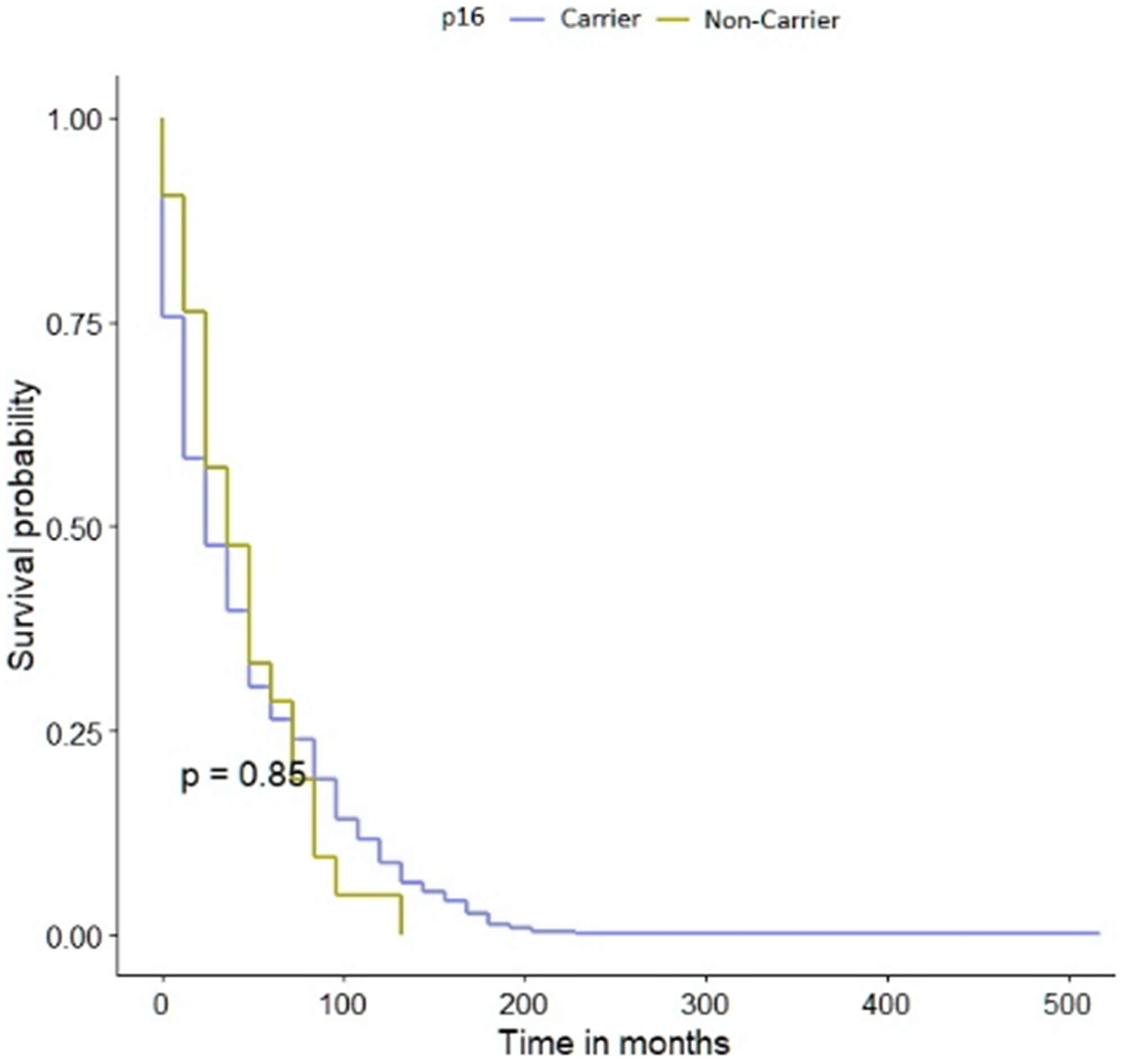
Figure 2: Kaplan-Meier survival curves of bladder cancer patients with CDKN2A polymorphism and -negative sub-cohorts.
Table 6: Survival of patients with bladder cancer; by variant of CDKN2A
| Patients with CDKN2A variant (n = 37) | Patients with no variant in CDKN2A (n = 669) | |
|---|---|---|
| Median follow-up (mo) | 156 | 156 |
| Proportion of deceased (%) | 62 | 70 |
| Median survival (mo) | 72 | 48 |
| 5-Year survival (%) | 54 | 50 |
| 10-Year survival (%) | 27 | 31 |
| HR | 1.38 | 1.0 |
| 95% CI | 0.83–2.29 | − |
| p-value | 0.2 | − |
Table 7: The HR for mortality associated with bladder cancer and the CDKN2A p.A148T variant
| HR | 95% CI | p-value | |
|---|---|---|---|
| patients younger than 60 years old | 2.54 | 0.89–7.26 | 0.082 |
| patients older than 60 years old | 1.10 | 0.59–2.04 | 0.8 |
| non-smoking patients | 0.28 | 0.06–1.36 | 0.12 |
| smoking patients | 1.87 | 0.53–6.65 | 0.3 |
| patients with no cancer family history | 1.31 | 0.79–2.19 | 0.3 |
| females | 1.46 | 0.84–2.53 | 0.2 |
| males | 0.72 | 0.15–3.36 | 0.7 |
| patients with noninvasive papillary low grade | 1.15 | 1.50–2.64 | 0.7 |
| patients with noninvasive papillary high grade | 1.38 | 0.47–4.10 | 0.6 |
| patients with invasive high grade | 1.61 | 0.70–3.69 | 0.3 |
| patients with Ta | 1.21 | 0.63–2.31 | 0.6 |
| patients with T2 | 1.09 | 0.42–2.80 | 0.9 |
| patients with T4 | 5.29 | 0.32–86.9 | 0.2 |
Kidney cancer
NOD2
The NOD2 c.3020insC variant was detected in 7,3% of the kidney cancer patients, a similar frequency as found in the Polish population. The characteristics of the study population are shown in Table 8 both with and without the variant are presented in Table 9. A total of 30 patients carried the NOD2 c.3020insC variant (OR = 1.1; 95% CI 0.74–1.69; p = 0.7) and was found in 20 affected men (5%) and 10 women (2.5%). The NOD2 variant was not observed in familial cases with bladder cancer nor in first- and/or second-degree relatives. Sixteen cancer patients who carried the NOD2 variant had died by February 2021.
Table 8: Characteristics of the study population of kidney cancer (n = 410)
| Sex | |
| Male | 261 |
| Female | 149 |
| Age, mean (range) | 60 (21–81) |
| ≤60 | 166 |
| >61 | 244 |
| Smoking status | |
| Yes | 162 (39%) |
| No | 91 (22%) |
| Missing | 157 |
| Histological features | |
| Clarocellulare | |
| GI* | 59 (14%) |
| GII* | 159 (39%) |
| GIII* | 105 (26%) |
| GIV* | 29 (7%) |
| Chromophobe | |
| GI | 16 (4%) |
| GII | 5 (1%) |
| GIII | 3 (1%) |
| Papillary | |
| GI | 14 (3%) |
| GII | 20 (5%) |
| GIII | − |
| Stage | |
| Ta | 5 (2%) |
| T1 | 278 (68%) |
| T2 | 40 (8%) |
| T3 | 81 (20%) |
| T4 | 6 (2%) |
| Vital status | |
| Alive | 278 (68%) |
| Dead | 132 (32%) |
Table 9: Clinical characteristics of kidney cancers patients harboring 3020insC allele in NOD2 gene
| Patients with 3020insC allele (30) | Patients with no 3020insC allele (380) | p-value* | Odds ratio (CI) | |
|---|---|---|---|---|
| Age of diagnosis (yr) | ||||
| ≤50 | 7 | 50 | 0.16 | 2.0 0.81–4.92 |
| 51-60 | 8 | 101 | 1.00 | 1.0 0.43−2.32 |
| 61-70 | 8 | 167 | 0.08 | 0.4 0.20−1.07 |
| >71 | 7 | 62 | 0.30 | 1.5 0.64−3.79 |
| Mean | 61 | 60 | ||
| Family history Positive (+) | − | 11 | − | − − |
| Smoking | ||||
| no | 4/19 (21) | 87/234 (37) | 0.20 | 0.4 0.14−1.40 |
| ≤20 packyears | 8/19 (42) | 95/234 (41) | 1.00 | 1.0 0.41−2.74 |
| >21 | 7/19 (37) | 52/234 (22) | 0.10 | 2.0 0.76−5.41 |
| Histological features | ||||
| Clarocellulare | ||||
| GI | 2/25 (8) | 57/327 (17) | 0.20 | 0.4 0.09−1.78 |
| GII | 11/25 (44) | 148/327 (45) | 1.00 | 0.9 0.42−2.15 |
| GIII | 8/25 (32) | 97/327 (30) | 0.80 | 1.1 0.46−2.67 |
| GIV | 4/25 (16) | 25/327 (8) | 0.10 | 2.3 0.73−7.22 |
| Chromophobe | ||||
| GI | 4/5 (80) | 12/19 (63) | 0.60 | 2.3 0.21−25.2 |
| GII | − | 5/19 (26) | − | − − |
| GIII | 1/5 (20) | 2/19 (11) | 0.0006 | 34 2.43−474.86 |
| Papillare | ||||
| GI | − | 14/34 (41) | − | − − |
| GII | − | 20/34 (59) | − | − − |
| GIII | − | − | − | − − |
| Stage | ||||
| Ta | − | 5/380 (1) | − | − − |
| T1 | 18/30 (60) | 260/380 (69) | 0.41 | 0.7 0.32−1.48 |
| T2 | 4/30 (13) | 36/380 (9) | 0.50 | 1.5 0.48−4.45 |
| T3 | 8/30 (27) | 73/380 (19) | 0.30 | 1.5 0.65−3.57 |
| T4 | − | 6/380 (2) | − | − − |
CDKN2A
Of the kidney cancer patients enrolled in the study, fourteen (3.4%) carried a CDKN2A p.A148T polymorphism (OR = 0.9; 95% CI 0.55–1.70; p = 0.9). The characteristics of the study population are shown in Table 6 with and without the CDKN2A p.A148T variant are presented in Table 10. Eleven males (0.3%) and three females (0.7%) carried of the p.A148T variant. The p.A148T allele was seen in one family with kidney cancer present in first- and/or second-degree relatives. Seven patients with kidney cancer who carried the CDKN2A variant had died by February 2021.
Table 10: Clinical characteristics of kidney cancers patients harboring A148T allele in CDKN2A gene
| Patients with 3020insC allele (14) | Patients with no 3020insC allele (396) | p-value* | Odds ratio (CI) | |
|---|---|---|---|---|
| Age of diagnosis (yr) | ||||
| ≤50 | 1 | 56 | 0.70 | 0.4 0.05–3.64 |
| 51–60 | 3 | 106 | 0.76 | 0.7 0.20–2.72 |
| 61–70 | 6 | 169 | 1.00 | 1.0 0.34–2.95 |
| >71 | 4 | 65 | 0.26 | 2.0 0.63–6.82 |
| Mean | 66 | 60 | ||
| Family history Positive (+) | 1 | 10 | 0.32 | 2.9 0.35–24.9 |
| Smoking | ||||
| no | 5/9 (56) | 86/244 (35) | 0.28 | 2.3 0.60–8.78 |
| ≤20 packyears | 3/9 (34) | 100/244 (41) | 0.74 | 0.7 0.17–2.94 |
| >21 | 1/9 (10) | 58/244 (24) | 0.68 | 0.4 0.04–3.27 |
| Histological features | ||||
| Clarocellulare | ||||
| GI | 4/13 (31) | 55/339 (16) | 0.24 | 2.3 0.98–7.71 |
| GII | 6/13 (46) | 153/339 (45) | 1.00 | 1.0 0.34–3.16 |
| GIII | 3/13 (23) | 102/339 (30) | 0.76 | 0.7 0.18–2.58 |
| GIV | − | 29/339 (9) | − | − − |
| Chromophobe | ||||
| GI | − | 16/23 (70) | − | − − |
| GII | 1/1 (100) | 4/23 (17) | 0.20 | 13 0.45–374.8 |
| GIII | − | 3/23 (13) | − | − − |
| Papillare | ||||
| GI | − | 14/34 (41) | − | − − |
| GII | − | 20/34 (59) | − | − − |
| GIII | − | − | − | − − |
| Stage | ||||
| Ta | − | 5/396 (1) | − | − − |
| T1 | 10/14 (71) | 268/396 (68) | 1.00 | 1.1 0.36–3.88 |
| T2 | 1/14 (7) | 39/396 (10) | 1.00 | 0.7 0.08–5.53 |
| T3 | 3/14 (22) | 78/396 (20) | 0.74 | 1.1 0.30–4.08 |
| T4 | − | 6/396 (1) | − | − − |
DISCUSSION
We previously reported that the common founder mutation in NOD2 may be more common in patients with bladder cancer and be associated with a reduced susceptibility to kidney cancer but these preliminary findings required further confirmation [1].
The study revealed that the presence of the NOD2 c.3020insC variant and the CDKN2A p.A148T polymorphism did not influence bladder cancer survival. When examining subgroups of these patients, those at the age of 60 or younger, who did not smoke tobacco, and those with histological low grade noninvasive papillary bladder cancer there was nominal evidence that the NOD2 c.3020insC variant effected survival.
When subdividing the patients by age to determine if there was any association with the age of disease onset we were unable to identify any statistically significant association between carriage of either of the two variants and the age of disease diagnosis. There also did not appear to be any relationship between either of the variants and the histological diagnosis of kidney cancer. Notwithstanding, we did identify borderline associations which require larger multi-center studies to determine more precisely whether there is a role of these two polymorphisms in disease risk and or pathology. To our knowledge no such large study examining the clinical characteristics and survival of Polish patients with bladder cancer and the presence of NOD2 and CDKN2A variants has been reported to date.
The strengths of the current study are the number of bladder and kidney cancer patients, which is complemented by detailed participant data. Predictors of disease include sex, age, clinical characteristics, smoking status and cancer family history all of which were controlled for in our statistical analysis. The maximum period of follow-up was 210 months. Xu et al. reported a study of 1953 samples with information about NOD2 from four The Cancer Genome Atlas (TCGA) databases in 2017 and showed that NOD2 could be a predictor of survival of kidney cancer patients [15]. In the present study we were unable to perform a survival analysis for kidney cancer. Liu et al. in a meta-analysis of thirty case-control studies, showed that NOD2 polymorphism was associated with an increased risk of gastric, breast, laryngeal, MALT lymphoma, lung and colorectal cancer [16]. In the review by Kutikhin (2011) the incidence of NOD2 polymorphism in individual populations was compared [17]. In the Polish population, the NOD2 c.3020insC variant has been identified in colorectal, early-onset laryngeal, lung, ductal breast cancer in situ, ovarian cancers [1, 3, 4, 6, 18]. In Poland, the association with the NOD2 polymorphism and rectal [19]; gastric, prostate, thyroid, melanoma [1, 20]; pancreatic [1, 5]; breast [21] and ovarian [22] cancers was not confirmed. Many of the participants enrolled in these previous studies comprised only small number of patient groups, which is likely to have contributed to the variance in the associations observed.
Many studies have investigated the clinical-pathological and prognostic significance of CDKN2A gene in patients with bladder cancer. However due to differences in sample sizes, statistical analysis and the study populations, the results remain inconclusive and evidence-based confirmation from large-scale clinical trials is still lacking. To address these issues we conducted the current study on a large group of Polish patients. The meta-analysis described by Pan et al. examined 12 studies encompassing 975 patients with bladder cancer revealing no correlation between CDKN2A expression and cancer prognosis [23]. In contrast, Gan et al. undertook a meta-analysis of 37 studies (a total of 2246 patients) with bladder cancer which confirmed an association between CDKN2A expression with prognosis and clinical-pathological features in patients with bladder cancer. [24]. They suggested that low CDKN2A expression was correlated with worse prognosis for progression-free survival and recurrence-free survival in Ta–T1 bladder cancer. Sun et al. showed that mutation in CDKN2A plays an important role in kidney cancer metastasis [25]. In 2006 Dębniak et al. evaluated the association between eleven malignancies and the CDKN2A p.A148T polymorphism in the Polish population [2]. No statistically significant relationship was observed for bladder or kidney with CDKN2A p.A148T variant. The results reported herein, concur with these findings and we conclude that this variant is not associated with disease risk or outcome.
In summary, the results of this study indicate that neither the NOD2 c.3020insC variant or the CDKN2A p.A148T polymorphism are associated with the survival of bladder cancer patients regardless of age, cancer family history, smoking status and sex. Thus, the NOD2 c.3020insC or the CDKN2A p.A148T polymorphism cannot be added to the list of genes that are associated with an increased susceptibility to bladder or kidney cancer at this time.
Materials and Methods
Study population
Patients
The unselected case group consisted of 706 urothelial bladder cancer patients (166 women, 540 men, mean age at diagnosis 79 years, range 25–91) and 410 kidney cancer cases (149 women, 261 men, mean age at diagnosis 60 years, range 21–81). All cancers were diagnosed at the Urology Hospital in Szczecin and the Genetic Outpatients Clinic between the years 2000–2018. A total of 1520 bladder cancers and 870 kidney cancers were identified during the study period. Of these, 1419 bladder patients and 835 kidney cancer patients accepted the study invitation. During the interview at the Genetic Outpatients Clinic, the study objectives were explained, informed consent was obtained, family history and smoking status recorded, genetic counseling provided and a blood sample taken for DNA analysis. The pathological diagnosis of bladder and kidney cancer was confirmed by biopsy at one central pathology laboratory in Szczecin, Poland. All cases were unselected for age, sex, clinical characteristics (stage: T, grade, histopathological tumor type), smoking and family history. Clinical data were collected from patient records. If there was no information on the stage, grade and histopathological type of the tumor the case was excluded. Of the 1419 patients with bladder and 835 with kidney cancer, clinical information was missing for 713 bladder and 425 kidney cases and these subjects were excluded from the study. In total, we recorded data from 706 patients with bladder and 410 with kidney cancer. Detailed information on smoking status (pack years) was available for a subgroup of 541 (77%) patients with bladder and 253 (62%) kidney cancer. A family history was taken at the Genetics Outpatients Clinic and a lifestyle questionnaire completed. A total of 26 patients were identified with a family history of at least 1 bladder cancer in their first or second degree relatives and 11 with a family history of at least 1 kidney cancer in their first or second degree relatives. The vital status and date of death of all of the cases were obtained from the Polish Ministry of the Interior and Administration in February 2021. In total, we received information about the deaths of 494 (70%) patients with bladder cancer and 132 (32%) with kidney cancer. All patients and control subjects are of European ancestry and are ethnic Poles. The study was approved by the Ethics Committee of Pomeranian Medical University in Szczecin.
Controls
We used two control groups. The first control group consisted of 2068 unselected, cancer-free individuals. These controls were selected to investigate the potential association between the NOD2 c.3020insC allele and bladder and kidney cancer. The control participants were collected from 1648 adults from Szczecin who submitted blood for paternity testing and 420 lists of adult patients from three family doctors from Szczecin. This control group was described in detail elsewhere [3].
The second control group included 3000 unselected cancer-free individuals to estimate the association between the CDKN2A p.A148T variant and bladder and kidney cancer. This control group consisted of 2000 newborns from 10 hospitals all over Poland and 1000 adults from the Szczecin area not selected for family history. This control group was described in detail elsewhere [10].
The allele frequencies for the NOD2 and CDKN2A variants in our control groups were not dependent on age or sex, and the estimates of the polymorphism frequency in both genes were similar in younger and in older control groups.
Methods
DNA was extracted from peripheral blood for all participants. The c.3020insC and the p.A148T variants were genotyped using a TaqMan assay (Life Technologies, Carlsbad, CA) employing a LightCycler Real-Time PCR 480 System (Roche Life Science, Mannheim, Germany). All mutations were confirmed by Sanger sequencing using a BigDye Terminator v3.1 Cycle Sequencing Kit (Life Technologies), according to the manufacturer’s protocol. In all reaction sets, positive and negative controls (including a reaction without DNA) were used.
Statistical analysis
Survival analysis
In order to perform the analysis, patient survival was followed from the date of bladder or kidney cancer diagnosis to the date of death or February 2021. Death was established by linkage to the Polish Vital Statistics Registry. All subjects in the study were linked to the records of the Vital Statistics Registry using a unique eleven digit identification number (PESEL). Death was all-cause mortality as no specific cause of death was available. The median follow-up was 210 months.
Kaplan-Meier survival curves were constructed for the variant carriers and non-carriers of the two tumour groups and the control participants. The comparison of the survival curves was performed by log-rank test. Multivariable Cox regression analysis was performed on these patients. Covariates included age (≤60; >61 years), gender (females; males), clinical characteristics (stage: T, grade, histopathological type of cancer), smoking status (non-smoking; smoking) and family history of cancer (negative; positive).
Survival analysis was performed first on all subjects, and then on subgroups of individuals divided by: age, gender, clinical characteristics, smoking and family history.
Due to the findings that the NOD2 c.3020insC allele and the CDKN2A p.A148T variant were not statistical significant in patients with renal cancer, we did not perform a survival analysis.
Power calculation
The statistical power for the case-control analysis for bladder cancer is approximately 70% with a two-sided confidence of 0.95, while for the renal cancer group we found the statistical power below 10%. In order to obtain at least 80% statistical power in this group, more than 322 000 subjects would have to be analysed, which is not possible in our region or indeed, Poland.
Odds ratios
The frequencies of the NOD2 c.3020insC allele and the CDKN2A p.A148T variant were compared in bladder and kidney cancer patients against the control group. Odds ratios were generated from two-by-two tables and statistical significance was assessed using the Fisher exact test where appropriate. The odds ratios were used as estimates of relative risk and were adjusted for age, sex, clinical characteristics, smoking and family history by multiple logistic regression.
Ethics approval and consent to participate
The study was approved by Ethics Committee of the Pomeranian Medical University in Szczecin, Poland. All participants gave informed written consent prior blood donating.
Data availability
Our data contain potentially sensitive information therefore we have not included it with our manuscript. Those who would like to request access to data may contact Melissa Sidhu at the Research Ethics Board of Women’s College Hospital by calling (416) 351-3732 x2723 or email [email protected]. The Pomeranian University of Medicine Ethics Committee will grant access to all researchers who meet the criteria for access to confidential data.
ACKNOWLEDGMENTS
We thank: Marcin Słojewski, Cezary Cybulski, Tadeusz Dębniak, Bohdan Górski, Anita Giermakowska, Joanna Trubicka, Artur Lemiński, Michał Soczawa for support in this study.
Ethics statement
The study was conducted in accordance with the principles of the Helsinki Declaration. All patients and controls provided written informed consent.
CONFLICTS OF INTEREST
JL is CEO of Read-Gene S.A. The authors EZ-P, AT-G, TW- declare that they have no conflict of interest relevant to this article.
FUNDING
This study was funded by Pomeranian Medical University, Szczecin. These authors are employees of Pomeranian Medical University, Szczecin: EZP, TD, MS, TW, ATG, CC, JL. The specific roles of these authors are articulated in the ‘author contributions’ section. The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
1. Lubiński J, Huzarski T, Kurzawski G, Suchy J, Masojć B, Mierzejewski M, Lener M, Domagała W, Chosia M, Teodorczyk U, Medrek K, Debniak T, Złowocka E, et al. The 3020insC Allele of NOD2 Predisposes to Cancers of Multiple Organs. Hered Cancer Clin Pract. 2005; 3:59–63. https://doi.org/10.1186/1897-4287-3-2-59. [PubMed].
2. Debniak T, Scott RJ, Huzarski T, Byrski T, Rozmiarek A, Debniak B, Górski B, Cybulski C, Medrek K, Mierzejewski M, Masojc B, Matyjasik J, Złowocka E, et al. CDKN2A common variant and multi-organ cancer risk--a population-based study. Int J Cancer. 2006; 118:3180–82. https://doi.org/10.1002/ijc.21760. [PubMed].
3. Lener MR, Oszutowska D, Castaneda J, Kurzawski G, Suchy J, Nej-Wołosiak K, Byrski T, Huzarski T, Gronwald J, Szymańska A, Szymańska-Pasternak J, Grodzki T, Serwatowski P, et al. Prevalence of the NOD2 3020insC mutation in aggregations of breast and lung cancer. Breast Cancer Res Treat. 2006; 95:141–45. https://doi.org/10.1007/s10549-005-9057-z. [PubMed].
4. Huzarski T, Lener M, Domagała W, Gronwald J, Byrski T, Kurzawski G, Suchy J, Chosia M, Woyton J, Ucinski M, Narod SA, Lubiński J. The 3020insC allele of NOD2 predisposes to early-onset breast cancer. Breast Cancer Res Treat. 2005; 89:91–93. https://doi.org/10.1007/s10549-004-1250-y. [PubMed].
5. Nej K, Bartsch DK, Sina-Frey M, Rieder H, Hahn SA, Lubiński J. The NOD2 3020insC Mutation and The Risk of Familial Pancreatic Cancer? Hered Cancer Clin Pract. 2004; 2:149–50. https://doi.org/10.1186/1897-4287-2-3-149. [PubMed].
6. Kurzawski G, Suchy J, Kładny J, Grabowska E, Mierzejewski M, Jakubowska A, Debniak T, Cybulski C, Kowalska E, Szych Z, Domagała W, Scott RJ, Lubiński J. The NOD2 3020insC mutation and the risk of colorectal cancer. Cancer Res. 2004; 64:1604–6. https://doi.org/10.1158/0008-5472.can-03-3791. [PubMed].
7. Jaworowska E, Masojć B, Tarnowska C, Brzosko M, Fliciński J, Serrano-Fernandez P, Matyjasik J, Amernik K, Scott RJ, Lubiński J. Association between early-onset breast and laryngeal cancers. Breast Cancer Res Treat. 2006; 97:215–19. https://doi.org/10.1007/s10549-005-9116-5. [PubMed].
8. Debniak T, Cybulski C, Górski B, Huzarski T, Byrski T, Gronwald J, Jakubowska A, Kowalska E, Oszurek O, Narod SA, Lubiński J. CDKN2A-positive breast cancers in young women from Poland. Breast Cancer Res Treat. 2007; 103:355–59. https://doi.org/10.1007/s10549-006-9382-x. [PubMed].
9. Debniak T, Scott RJ, Huzarski T, Byrski T, Rozmiarek A, Debniak B, Załuga E, Maleszka R, Kładny J, Górski B, Cybulski C, Gronwald J, Kurzawski G, Lubinski J. CDKN2A common variants and their association with melanoma risk: a population-based study. Cancer Res. 2005; 65:835–39. [PubMed].
10. Debniak T, Górski B, Huzarski T, Byrski T, Cybulski C, Mackiewicz A, Gozdecka-Grodecka S, Gronwald J, Kowalska E, Haus O, Grzybowska E, Stawicka M, Swiec M, et al. A common variant of CDKN2A (p16) predisposes to breast cancer. J Med Genet. 2005; 42:763–65. https://doi.org/10.1136/jmg.2005.031476. [PubMed].
11. Debniak T, Scott R, Masojc B, Serrano-Fernández P, Huzarski T, Byrski T, Debniak B, Górski B, Cybulski C, Medrek K, Kurzawski G, van de Wetering T, Maleszka R, et al. MC1R common variants, CDKN2A and their association with melanoma and breast cancer risk. Int J Cancer. 2006; 119:2597–602. https://doi.org/10.1002/ijc.22210. [PubMed].
12. Lubinski J, Korzen M, Gorski B, Cybulski C, Debniak T, Jakubowska A, Medrek K, Matyjasik J, Huzarski T, Byrski T, Gronwald J, Masojc B, Lener M, et al. Breast cancer susceptibility genes. J BUON. 2007; 12:S23–29. [PubMed].
13. Forrest MS, Skibola CF, Lightfoot TJ, Bracci PM, Willett EV, Smith MT, Holly EA, Roman E. Polymorphisms in innate immunity genes and risk of non-Hodgkin lymphoma. Br J Haematol. 2006; 134:180–83. https://doi.org/10.1111/j.1365-2141.2006.06141.x. [PubMed].
14. Rothman N, Skibola CF, Wang SS, Morgan G, Lan Q, Smith MT, Spinelli JJ, Willett E, De Sanjose S, Cocco P, Berndt SI, Brennan P, Brooks-Wilson A, et al. Genetic variation in TNF and IL10 and risk of non-Hodgkin lymphoma: a report from the InterLymph Consortium. Lancet Oncol. 2006; 7:27–38. https://doi.org/10.1016/S1470-2045(05)70434-4. [PubMed].
15. Xu D, Zhang S, Zhang S, Liu H, Li P, Yu L, Shang H, Hou Y, Tian Y. NOD2 maybe a biomarker for the survival of kidney cancer patients. Oncotarget. 2017; 8:101489–99. https://doi.org/10.18632/oncotarget.21547. [PubMed].
16. Liu J, He C, Xu Q, Xing C, Yuan Y. NOD2 polymorphisms associated with cancer risk: a meta-analysis. PLoS One. 2014; 9:e89340. https://doi.org/10.1371/journal.pone.0089340. [PubMed].
17. Kutikhin AG. Role of NOD1/CARD4 and NOD2/CARD15 gene polymorphisms in cancer etiology. Hum Immunol. 2011; 72:955–68. https://doi.org/10.1016/j.humimm.2011.06.003. [PubMed].
18. Suchy J, Kłujszo-Grabowska E, Kładny J, Cybulski C, Wokołorczyk D, Szymańska-Pasternak J, Kurzawski G, Scott RJ, Lubiński J. Inflammatory response gene polymorphisms and their relationship with colorectal cancer risk. BMC Cancer. 2008; 8:112. https://doi.org/10.1186/1471-2407-8-112. [PubMed].
19. Szeliga J, Sondka Z, Jackowski M, Jarkiewicz-Tretyn J, Tretyn A, Malenczyk M. NOD2/CARD15 polymorphism in patients with rectal cancer. Med Sci Monit. 2008; 14:CR480–84. [PubMed].
20. Debniak T, Kurzawski G, Huzarski T, Byrski T, Gronwald J, Debniak B, Rozmiarek A, Dziuba I, Złowocka E, Suchy J, Górski B, Cybulski C, Mierzejewski M, et al. NOD2 variants and the risk of malignant melanoma. Eur J Cancer Prev. 2005; 14:143–46. https://doi.org/10.1097/00008469-200504000-00010. [PubMed].
21. Janiszewska H, Haus O, Lauda-Swieciak A, Bak A, Mierzwa T, Sir J, Laskowski R. The NOD2 3020insC Mutation in Women with Breast Cancer from the Bydgoszcz Region in Poland. First Results. Hered Cancer Clin Pract. 2006; 4:15–19. https://doi.org/10.1186/1897-4287-4-1-15. [PubMed].
22. Magnowski P, Medrek K, Magnowska M, Stawicka M, Kedzia H, Górski B, Lubiński J, Spaczyński M. The 3020insC NOD2 gene mutation in patients with ovarian cancer. Ginekol Pol. 2008; 79:544–49. [PubMed].
23. Pan JG, Han RF, Wang Y. Significance of p16 gene and p21 gene expression abnormality in the prognosis of bladder carcinoma: a meta-analysis. Chinese Journal of Urology. 2006; 27:400–3.
24. Gan X, Lin X, He R, Lin X, Wang H, Yan L, Zhou H, Qin H, Chen G. Prognostic and Clinicopathological Significance of Downregulated p16 Expression in Patients with Bladder Cancer: A Systematic Review and Meta-Analysis. Dis Markers. 2016; 2016:5259602. https://doi.org/10.1155/2016/5259602. [PubMed].
25. Sun Q, Chen S, Hou Y, Wen X, Teng X, Zhang H, Lai C, Lai M. Mutant CDKN2A regulates P16/p14 expression by alternative splicing in renal cell carcinoma metastasis. Pathol Res Pract. 2021; 223:153453. https://doi.org/10.1016/j.prp.2021.153453. [PubMed].

