Introduction
Osteosarcoma (OS), the most common primary tumor of the bone in the pediatric population and second most common in adults (following multiple myeloma), is characterized by a remarkable degree of intratumoral heterogeneity in cell phenotypes, gene expression, and chemoresistance [1–3]. The combination of multi-agent chemotherapy and improved surgical techniques have increased 5-year survival to 70% in patients diagnosed with localized disease.
In contrast, the survival rates for patients with recurrent disease or metastatic disease at diagnosis have barely changed, having remained around 20% for more than five decades [4]. Among the roadblocks that have stymied the advent of new therapies, perhaps the most impenetrable derives from the unique genetic etiology of OS. Whereas 10–15% of OS tumors exhibit relatively specific mutations in Rb or p53, non-targetable mutations in tumor suppressors associated with familial retinoblastoma [5, 6] and Li Fraumeni syndrome [7], most OSs originate as a byproduct of chromothripsis, the result of a catastrophic genomic event of unclear genesis [8–10]. The ensuing chromosomal and mutational diversity leaves oncologists without a singular target to direct their therapeutic focus. Despite their high mutation burden, OS has proven surprisingly recalcitrant to the numerous immunotherapies that have revolutionized the treatment of other mutation-high cancers.
Without OS-specific proteins to target, investigators have increasingly sought to explore whether oncogenic signaling cascades common in other cancers may also play a role in OS. The MAPK, IGF/PI3K/mTOR, and YAP/TAZ pathways, in particular, have generated much interest. Herein, we characterize a potential relationship in OS between the IGF-1R receptor tyrosine kinase (RTK) and two downstream mediators of the Hippo pathway, YAP and TAZ.
The IGF-1R/PI3K/mTOR cascade has been the subject of clinical trials in osteosarcoma with variable success [11–15]. Ligand-mediated receptor phosphorylation is known to initiate a signaling kinase cascade propagated through sequential activation of IRS-1, PI3K, and Akt, ultimately culminating in mTOR activation. Traditionally thought to function exclusively at the cell surface, recent data suggests that ligand-receptor engagement mediates IGF-1R endocytosis. Following its internalization, IGF-1R can recirculate back to the plasma membrane or transit the nuclear membrane where it joins transcription factors to exert complex epigenetic effects [16–18]. Significant controversy exists about the prognostic value of IGF-1R in its ability to predict response to downstream mediators PI3K and mTOR [18, 19].
Yes-associated protein 1 (YAP) and transcriptional coactivator with PDZ-binding motif (WWTR1, TAZ) are mechanoresponsive transcription factors phosphorylated by the active Hippo tumor suppressor pathway. Upon phosphorylation by MST1 and LATS (the mammalian orthologs of Hippo and Warts in Drosophila), YAP and TAZ (often referred to together as YAP/TAZ given their significant functional overlap) undergo binding to 14-3-3, which tags them for nuclear exclusion and degradation. Alternatively, if the Hippo pathway is inactive, YAP/TAZ remain unphosphorylated and translocate into the nucleus [20]. Studies in human mesenchymal stem cells (MSCs) have indicated that stiff microenvironments promote cell spreading, increase nuclear YAP and TAZ, and facilitate cell reprogramming towards an osteogenic lineage; conversely, cell confinement and less stiff microenvironments contribute to commitment towards an adipogenic lineage [21, 22]. Our lab and others have shown that YAP/TAZ localization is also influenced by the architecture and dimensionality of the tumor niche, highlighting the complexity of what cells ‘sense’ as their physical microenvironment [23, 24]. YAP/TAZ-mediated mechanotransduction has also been implicated in tumor chemoresistance and, potentially, worse OS survival [25–27].
In the present study, we evaluate the association between the IGF-1/mTOR and YAP/TAZ pathways, with a major emphasis on the nuclear (i.e., activated) state of phosphorylated IGF-1R (pIGF-1R), non-phosphorylated IGF-1R, and nuclear-to-cytoplasmic ratios of YAP and TAZ. This was accomplished using a human OS tissue microarray (TMA) comprised of 37 post-treatment OS tumor specimens. We performed confocal imaging of each TMA section and quantified nuclear and cytoplasmic protein intensity using semi-automated cell segmentation algorithms. Nuclear staining for pIGF-1R, total IGF-1R, and nuclear to cytoplasmic ratios (N:C ratios) for YAP/TAZ were correlated the patients’ clinical characteristics. Kaplan–Meier analyses were performed, and a proportional hazards model was used to estimate each parameter’s contribution to overall survival. Overall, this research sheds new light on the interrelationship between the IGF-1R/PI3K/mTOR and YAP/TAZ cancer-related pathways.
Results
Patient demographics and biopsy characteristics
All patients received care at the University of Texas MD Anderson Cancer Center between November 1989 and December 2018. Biopsies were taken following neoadjuvant treatment from 37 patients. 67.6% of patients were male, and 37.8% were female (Table 1). Biopsies were taken from the primary tumor (27.0%), recurrent tumor (5.4%), or a metastatic site, most commonly the lung (67.6%). The mean age at diagnosis was 33.7 years, whereas the median age (interquartile range; total range) was 28.75 years (16.6–45.9; 10.4–79.5 years).
Table 1: Patient demographics and biopsy characteristics
| n | Percentage | ||
| Race/Ethnicity | Asian | 2 | 5.4% |
| Black or African American | 4 | 10.8% | |
| White (Hispanic or Latino) | 6 | 16.2% | |
| White (Not Hispanic or Latino) | 25 | 67.6% | |
| Gender | Female | 14 | 37.8% |
| Male | 23 | 62.2% | |
| TMA Biopsy Site | Primary | 10 | 27.0% |
| Metastatic: Other Site | 4 | 10.8% | |
| Metastatic: Lung | 21 | 56.8% | |
| Local Recurrence | 2 | 5.4% |
Correlation of stained markers
YAP, TAZ, IGF-1R, and pIGF-1R demonstrated a wide range of localization and overall signal, which is expected given the high degree of morphological inter-patient and intra-tumoral heterogeneity observed between tumors (Figure 1A and Figure 1B). When average staining intensity of IGF-1R/pIGF-1R and YAP/TAZ N:C ratios were treated as continuous variables, expression of IGF-1R and pIGF-1R were highly correlated (Pearson r correlation coefficient 0.690, p < 0.001). YAP and TAZ similarly correlated with each other (0.822, p < 0.001). Additionally, we identified a likely relationship between pIGF-1R and the YAP/TAZ pathway, as pIGF-1R was highly correlated with YAP and TAZ of 0.522 (p = 0.001). Weaker but statistically significant positive associations could be found between other pairs (Figure 1C).
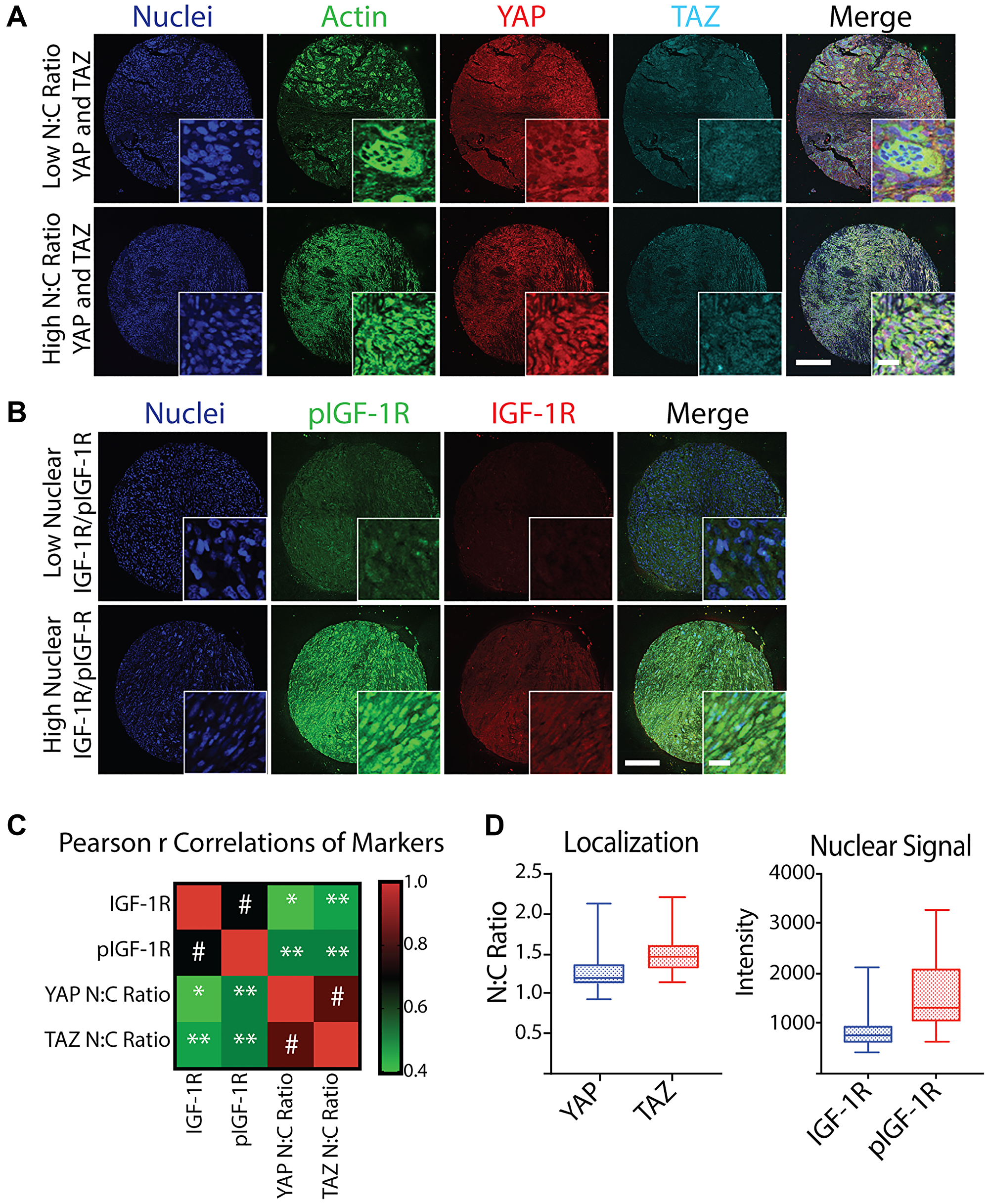
Figure 1: Quantification and correlation of YAP N:C ratio, TAZ N:C ratio, nuclear IGF-1R intensity, and nuclear pIGF-1R intensity from a TMA of post-treatment osteosarcoma biopsies. (A) Representative confocal microscopy images of osteosarcoma taken at post-treatment biopsy in a tissue microarray (TMA) displaying low (top) and high (bottom) N:C ratio of YAP and TAZ co-stained with Hoechst (nuclei, blue), phalloidin (actin, green), scale = 250 μm, inset scale = 20 μm. (B) Representative confocal microscopy images of osteosarcoma TMA displaying low (top) and high (bottom) nuclear IGF-1R and pIGF-1R (bottom) co-stained with Hoechst (nuclei, blue), scale = 250 μm. (C) Heatmap of Spearman r Correlations of YAP N:C ratio, TAZ N:C ratio, mean nuclear IGF-1R intensity, and mean nuclear pIGF-1R intensity for each patient represented in the TMA (n = 37 for IGF-1R/pIGF-1R and n = 36 for YAP/TAZ due to loss of one sample on TMA) with corresponding p values. *p < 0.05, **p < 0.01, #p ≤ 0.001. (D) Box plots of average marker values per patient across all patients represented in the TMA. Shaded boxes with inner line represent the interquartile range and median, where whiskers represent the range of calculated patient values.
Since the YAP/TAZ paralogs must bind TEAD (or other co-factors) and shuttle to the nucleus to exert their epigenetic effects, pathway activation can be measured using either the absolute expression of nuclear YAP/YAZ or the N:C ratios. Our patients’ observed N:C ratio ranged between 0.94 and 2.13 for YAP and 1.15 and 2.19 for TAZ. However, most patient values fell in much smaller interquartile ranges of 1.13 and 1.35 for YAP and 1.33 and 1.60 for TAZ (Figure 1D, left panel). IGF-1R nuclear intensity also had a narrow interquartile range compared to the total range. Conversely, we observed widely varied nuclear expression of pIGF-1R in patient samples (Figure 1D, right panel).
Association of TMA confocal imaging with the histological characterization of disease
Since YAP/TAZ is an important role in directing MSCs lineage commitment toward mature connective tissues, we hypothesized their activity might associate with distinct osteosarcoma subtypes [21, 28]. Each tumor’s predominant histotype and the subtype were first determined by pathologists at MD Anderson Cancer Center that have bone sarcoma expertise. Notably, as OS tumors usually exhibit multi-lineage osteoblastic, chondroblastic, and fibroblastic differentiation, the relevant subclassification was determined by pathologists based upon their review of the entire tumor specimen. Next, we examined whether YAP/TAZ N:C ratios or pIGF-1R/IGF-1R expression levels differed by the OS histopathological subtype. Our findings indicate that YAP N:C ratio was highest among chondroblastic phenotype compared to other histotypes (Figure 2A, right panel). Increased YAP and TAZ N:C ratios were also observed in low- and intermediate-grade OS (Figure 2B, right panel).
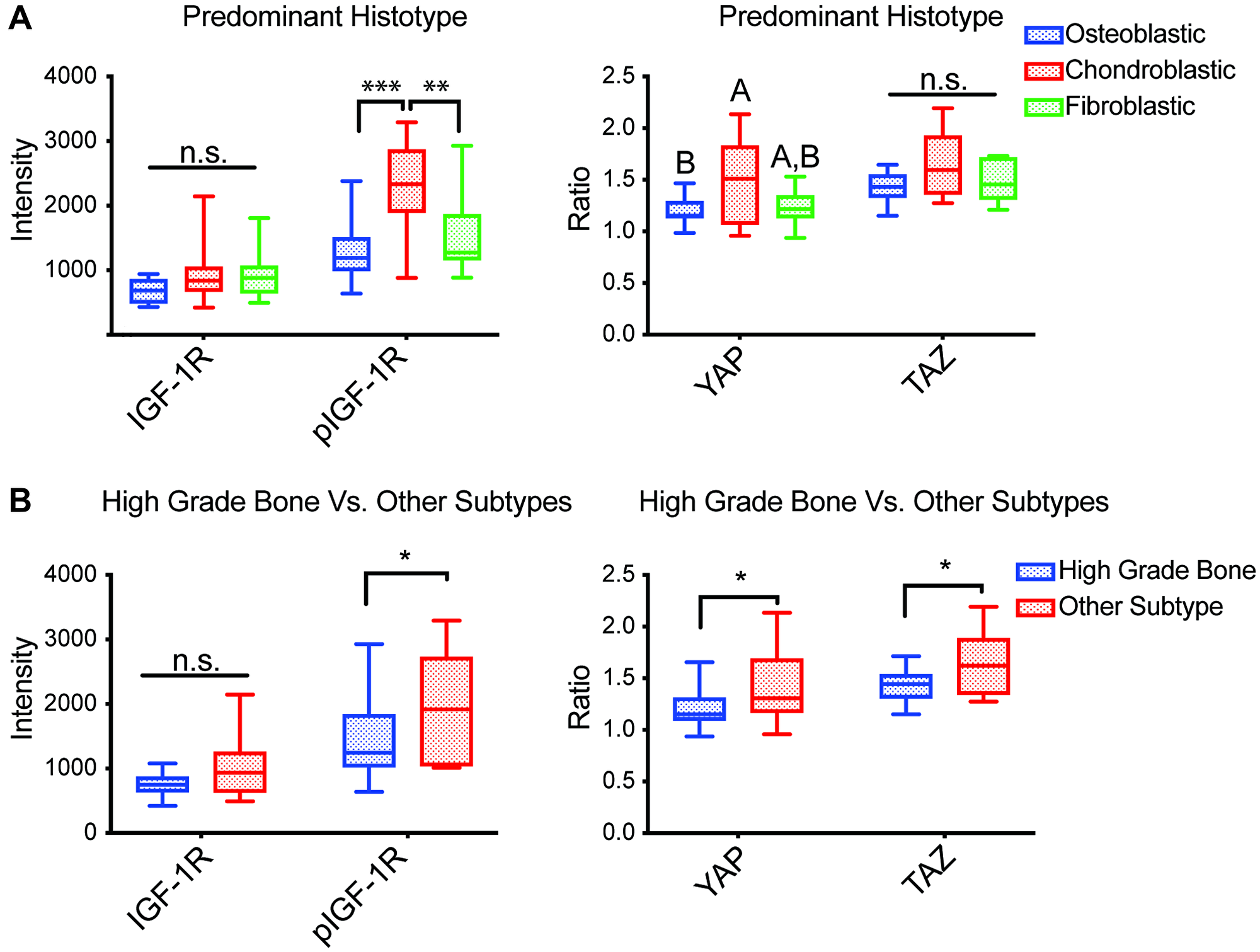
Figure 2: Associations between TMA staining and histopathological aspects of tumors. Average mean nuclear intensity of IGF-1R and pIGF-1R (Y1161) (n = 37) and average N:C ratio of YAP and TAZ (n = 36) in biopsies represented in a TMA subdivided by (A) predominant histotype of constituent cells [fibroblastic n = 10; osteoblastic n = 14; chondroblastic n = 7; other n = 4; not listed = 2], and (B) osteosarcoma subtypes compared to the high grade bone subtype [high-grade bone n = 27; other n = 10]. Box plots with inner line represent the interquartile range and median, where whiskers represent the range of observed values; Abbreviation: n.s.: no significance, differing letters and *p < 0.05, **p < 0.01, ***p < 0.001, ANOVA with post-hoc Sidak test.
Increased nuclear pIGF-1R was also associated with a chondroblastic histopathology and more likely to be seen in osteosarcoma subtypes other than high-grade bone (Figure 2, left panels). Interestingly, although increased pIGF-1R nuclear staining was associated with subtypes other than high-grade bone, IGF-1R nuclear staining intensity did not correlate with osteosarcoma subtype, possibly due to low sample sizes. However, when osteosarcoma subtypes were compared by specific subtype, the only statistical differences among the groups were an increased pIGF-1R nuclear stain and YAP N:C ratio of the myxoid subtype compared to high-grade bone (Supplementary Figure 1). Nuclear IGF-1R intensity and TAZ N:C ratios were not statistically different across the tumors with different predominant histotypes present in the osteosarcoma biopsies.
We also assessed if protein markers correlated with other clinical aspects of disease outcomes, as reported in other studies [29]. Interestingly, the YAP/TAZ N:C ratio and IGF-1R/pIGF-1R nuclear intensity did not appear to correlate with the primary vs. metastatic biopsy (Supplementary Figure 2A). Furthermore, they did not correlate with primary tumor location (Supplementary Figure 2B), the presence or absence of metastatic disease at diagnosis (Supplementary Figure 2C), or the emergence of metastases throughout a patient’s disease course (Supplementary Figure 2D).
YAP/TAZ and pIGF-1R/IGF-1R staining and overall survival
Previous reports have associated high IGF-1R expression with advanced stage, lower overall survival, and distant metastasis. However, less is known about the impact of nuclear or phosphorylated IGF-1R in osteosarcoma [19, 30]. To correlate the nuclear staining and localization of the tumor markers with outcome measures, we found it useful to partition patients into groups with low, medium, and high YAP/TAZ N:C ratio or IGF-1R/pIGF-1R mean nuclear intensity. In our work, we stratified average values by grouping patients within the 1st quartile, interquartile range, or 4th quartile as “low,” “medium,” and “high,” respectively, for the four selected markers over the 37 patients represented in the TMA.
We assessed patients’ overall survival for correlations with patient characteristics, pathology assessments, or tumor marker values generated from confocal imaging of the TMA. Kaplan–Meier assessment yielded no statistical differences between YAP or TAZ N:C ratio or IGF-1R mean nuclear intensity and overall survival (Figure 3A–3C). Conversely, patients with high pIGF-1R mean nuclear intensity had worse overall survival (log-rank test done pairwise, p < 0.05) (Figure 3D).
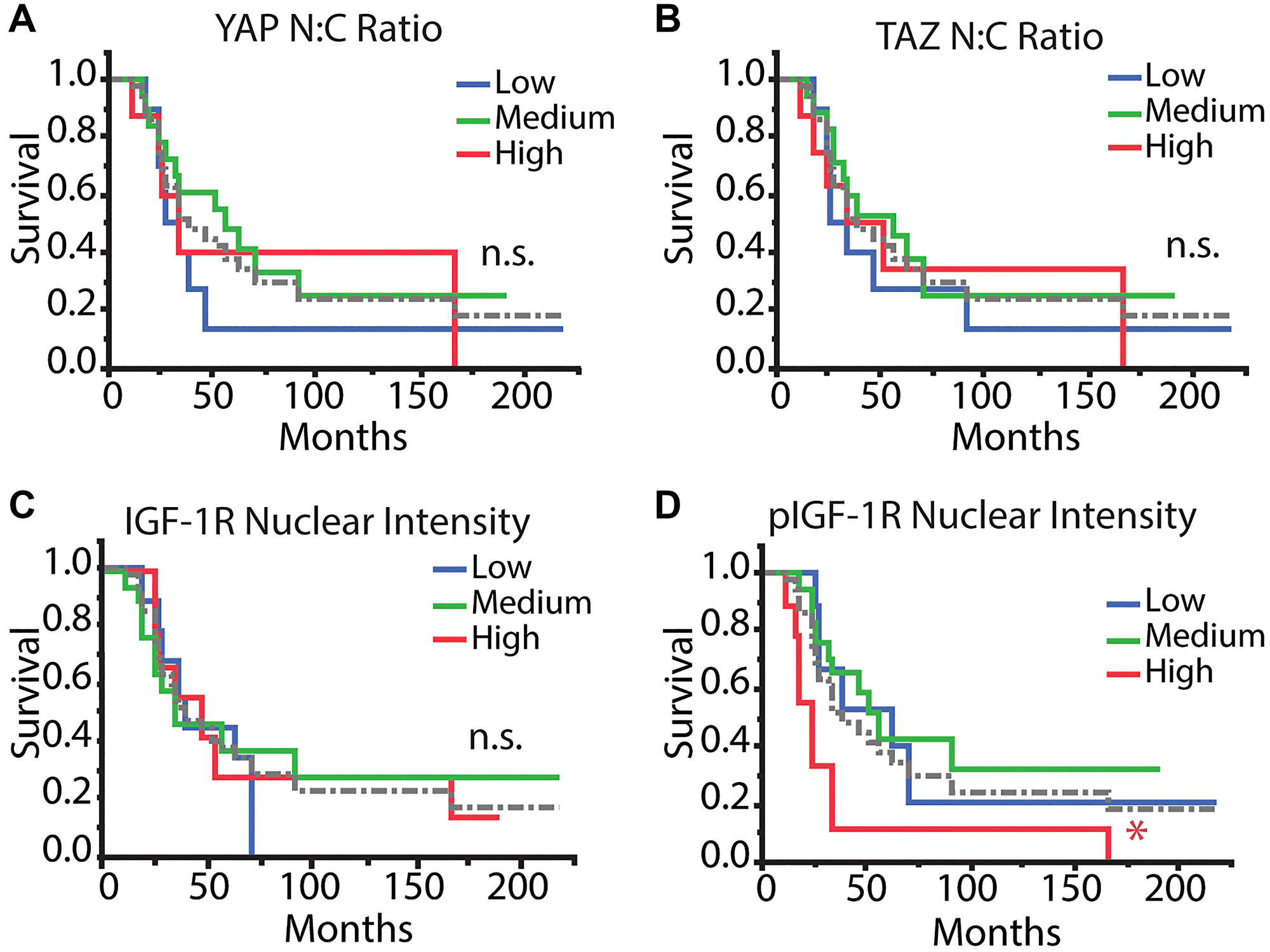
Figure 3: Kaplan–Meier survival analysis of tumor markers. Patients were stratified into low, medium, and high categories (1st quartile [n = 10 IGF-1R/pIGF-1R; n = 9 YAP/TAZ], interquartile range [n = 17 IGF-1R/pIGF-1R; n = 18 YAP/TAZ], 4th quartile [n = 10 IGF-1R/pIGF-1R; n = 9 YAP/TAZ] for each mean value across all patients, respectively) for each parameter and were analyzed with Kaplan–Meier analysis to determine differences in overall survival. Tumor markers included (A) YAP N:C ratio, (B) TAZ N:C ratio, (C) IGF-1R nuclear intensity, and (D) pIGF-1R nuclear intensity. The dotted gray line refers to the average of all patients. Log-rank test, post-hoc: Abbreviation: n.s.: no significance, *p < 0.05, **p < 0.01.
To determine if pIGF-1R’s adverse survival impact was, in part, related to enhanced metastatic potential, we evaluated whether pIGF-1R (and other markers) occurred more commonly in patients who developed metastatic spread. As a first step, we reviewed whether known adverse clinical parameters in our data set were associated with poor patient outcomes. We correlated overall survival with metastasis at diagnosis and the development of clinically observable metastasis throughout the disease, which is a known negative prognostic indicator [31, 32]. As expected, patients diagnosed with metastatic disease at diagnosis or during their disease course had significantly worse overall survival (Figure 4A, 4B, log-rank test **p < 0.01 and *p < 0.05, respectively).
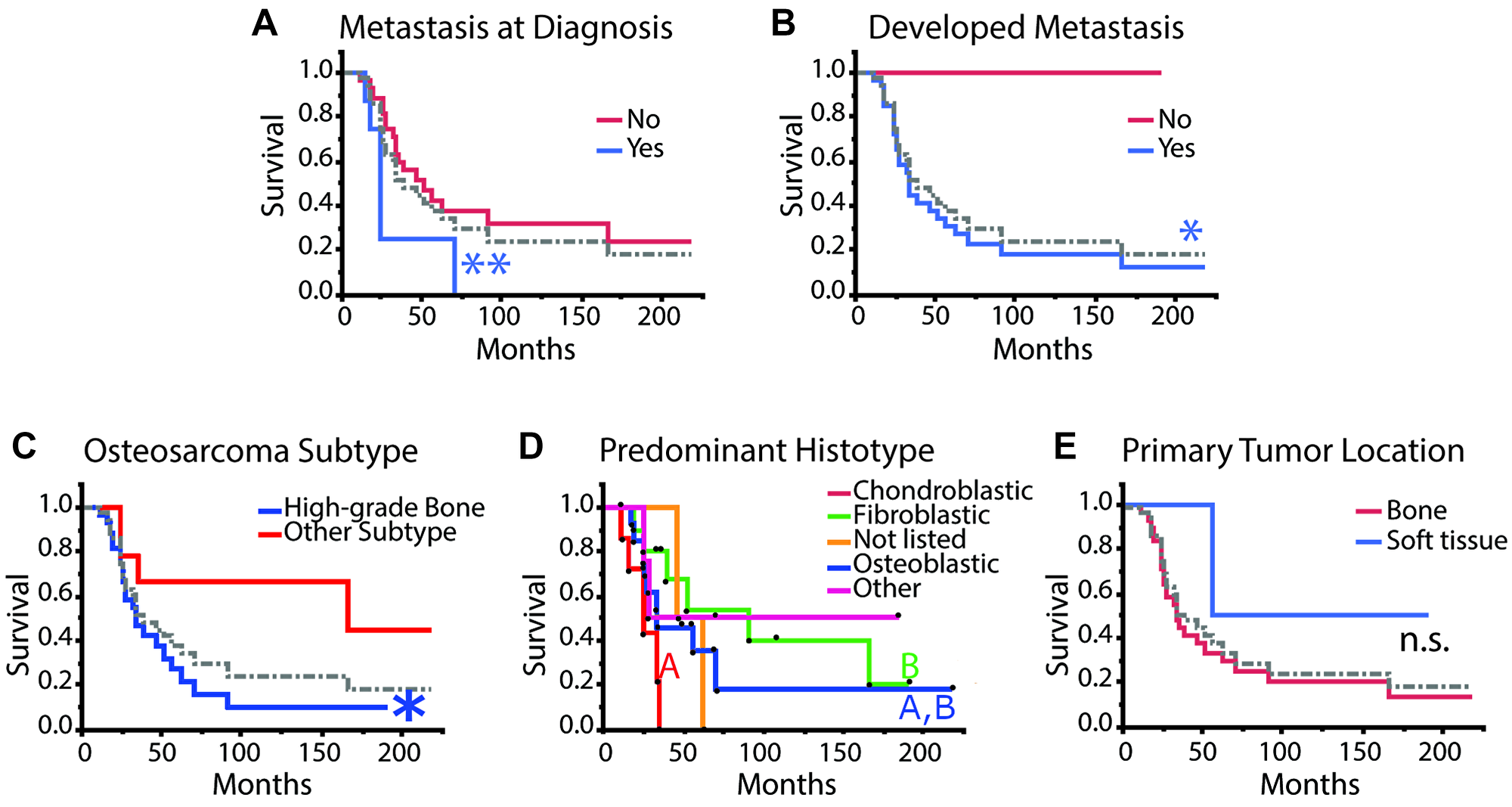
Figure 4: Kaplan–Meier survival analysis for clinical and histopathological aspects of disease. (A) metastasis at diagnosis: yes n = 8; no n = 28. (B) metastasis developed during disease course: yes n = 33; no n = 4. (C) osteosarcoma subtype: high-grade bone n = 27; other n = 10. (D) predominant histotype: excluding “not listed” n = 2 and “other” n = 4 and including “fibroblastic” n = 10; “osteoblastic” n = 14; “chondroblastic” n = 7. (E) primary tumor location: bone n = 33; soft tissue n = 4. Survival curves were compared using the log-rank test, and plots with more than two curves were compared pairwise except for “Predominant Histotype” where only the known histotype curves were compared excluding “not listed” and “other” and including “fibroblastic” n = 10; “osteoblastic” n = 14; “chondroblastic” n = 7. Black dots indicate individual patients that were censored (dot above curve) or died (dot below the curve). The dotted gray line refers to the average of all patients. Log-rank test, post-hoc: Abbreviation: n.s.: no significance, different colored letters and *p < 0.05, **p < 0.01.
Consistent with published reports, the high-grade bone osteosarcoma subtype was associated with decreased overall survival (Figure 4C) [33]. Large studies have previously reported that the predominant histological subtype does not correlate with prognosis or outcomes in high-grade osteosarcoma, while a different group had reported that chondroblastic phenotypes might be associated with worse overall outcomes [34–36]. Interestingly, chondroblastic histotype was associated with worse outcomes than fibroblastic histotype, but neither of these phenotypes was statistically different from the osteoblastic histotype (Figure 4D). Primary tumor location did not seem to affect overall survival, which is consistent with previous studies (Figure 4E) [37]. We note that our study was not powered to detect a survival difference for extraskeletal osteosarcoma, given that this scenario is exceedingly rare.
Given the complexity and potential interactions between variables in our patient sample set, we developed a proportional hazards (Cox) model using a priori selected variables. Tumor markers were fit as categorical variables within the Kaplan–Meier analysis. Possibly due to the small sample size (n = 37), calculated risk ratios were considerable and unreliable (calculated p values for the Wald tests and hazard ratios with 95% CIs are available in Supplementary Table 1). To avoid overfitting our sample data, we employed the Bayesian information criterion (BIC) to select variables suitable for our Cox model that correlated with survival time in those patients who died over the observation period. This approach selected pIGF-1R mean nuclear intensity, YAP N:C ratio, osteosarcoma subtype, and predominant histotype as the variables most potentially predictive of time to death in the osteosarcoma patients represented in the TMA.
After fitting a Cox model to these four variables, Wald test p-values indicated that the effects of pIGF-1R mean nuclear intensity and YAP N:C ratio were significant (p = 0.0023 and p = 0.0224, respectively, Table 2). High pIGF-1R had increased risk ratios of 23.36 (95% CI: 3.83–154.72) and 25.22 (95% CI: 3.67–206.37) compared to medium and low pIGF-1R, respectively. There was no significant difference between medium and low pIGF-1R. The adverse prognostic effect of pIGF-1R upon survival was consistent with the Kaplan–Meier survival analysis. Conversely, it was low YAP N:C ratio compared to a medium N:C ratio that was associated with a higher risk ratio of 7.38 (95% CI: 1.81–35.42; p < 0.01). Lower YAP N:C ratio trended with an increased risk compared to high YAP N:C of 5.03 but was not statistically significant likely due to small sample size (95% CI: 0.58–76.15; p = 0.16). Calculated risk ratios, confidence intervals, and p-values for various pairs of osteosarcoma subtype and predominant histotype can be viewed in Supplementary Table 2, although the overall effect did not appear to be significant based on Wald tests for these variables.
Table 2: Proportional hazards model featuring variables selected with the Bayesian Information Criterion (BIC)
| Effect Wald Tests | |||||
| Source | p-value | ||||
| pIGF-1R Mean Nuclear Intensity | 0.0023* | ||||
| YAP N:C Ratio | 0.0224* | ||||
| Osteosarcoma Subtype | 0.1689 | ||||
| Predominant Histotype | 0.1104 | ||||
| Risk Ratios | |||||
| Risk Ratios for pIGF-1R Mean Nuclear Intensity | |||||
| Level 1 | Level 2 | Risk Ratio | p-value | Lower 95% | Upper 95% |
| High | Low | 25.22 | 0.0009* | 206.37 | 3.67 |
| Medium | Low | 1.08 | 0.9021 | 0.33 | 3.94 |
| High | Medium | 23.36 | 0.0008* | 3.83 | 154.72 |
| Risk Ratios for YAP N:C Ratio | |||||
| Level 1 | Level 2 | Risk Ratio | p-value | Lower 95% | Upper 95% |
| Low | High | 5.03 | 0.1568 | 0.58 | 76.15 |
| Low | Medium | 7.38 | 0.0050* | 1.81 | 35.43 |
| High | Medium | 1.47 | 0.7678 | 0.11 | 16.81 |
The fitted Cox models and the Kaplan–Meier survival curves indicated that YAP and pIGF-1R might be useful prognostic indicators in osteosarcoma. Because there is such a wide range of phenotypic variability in YAP localization and IGF-1R activation among patients, these indicators may be useful in stratifying or selecting patients for YAP/TAZ or IGF-1R targeted therapy.
DISCUSSION
The rarity and heterogeneous intratumoral characteristics of osteosarcoma constitute significant hurdles for the development of prognostic biomarkers [1]. The advent of TMA technology has afforded cancer researchers the ability to examine multiple tissues from multiple patients. TMAs allow researchers to treat each sample simultaneously with stains and antibodies that can reduce variability between samples. The use of tissue microarrays has primarily been in studies focused on immunohistochemistry. Here we describe the use of quantitative image analyses for staining intensity and localization from confocal microscopy to evaluate TMAs, also reported in other studies [38].
In this work, we probed TMAs of core biopsies of osteosarcoma in patients with known outcomes to investigate the relationship between nuclear IGF-1R localization/activation and YAP/TAZ nuclear to cytoplasmic ratios. Usually, the IGF-1R/mTOR pathway facilitates the growth of organs, the proliferation of cells, and the development of many tissue types [39]. However, in osteosarcoma, the IGF-1R/mTOR pathway has been implicated as a driver of anti-apoptotic signals and aberrant growth in many cancers making the pathway an attractive target for potential therapies [40, 41].
The Hippo pathway effectors, YAP and TAZ, have been extensively described as transducers of mechanical inputs such as growth substrate stiffness, actin cytoskeletal tension, and cell shape [21, 22, 42]. Upon stimulation of the Hippo pathway in environments that confine cell growth or with low substrate stiffness, YAP and TAZ are phosphorylated and confined to the cytosol for degradation. This process helps regulate organ size in healthy tissues. Recent preclinical models in our lab and others have shown that increased cell spreading in less stiff 3D microenvironments promote nuclear localization of YAP/TAZ, highlighting a complex regulation of this mechanosensitive pathway [23, 24]. In osteosarcoma, YAP levels are upregulated in about 80% of tumors, and YAP/TAZ signaling is involved in tumor initiation, propagation, and metastasis [43–45]. This data suggests that decreased bone stiffness due to lytic osteosarcoma lesions may increase cell spreading and facilitate nuclear YAP/TAZ localization, thus contributing to pathogenesis.
Given the functions of YAP/TAZ and the IGF-1R/mTOR cascade in stimulating proliferation and growth, it is unsurprising that studies have revealed the potentiation of the mTOR pathway by YAP [46, 47]. However, to our knowledge, our study is the first to demonstrate this relationship in osteosarcoma patient biopsies. In patient samples, there was a wide variation of nuclear (i.e., active) IGF-1R, as evidenced by the distribution of mean intensities of samples stained for pIGF-1R. Kaplan–Meier survival analysis indicated that high nuclear pIGF-1R was associated with worse outcomes. Moreover, a selected Cox model fit to our data confirmed an increased hazard ratio for death for patients with progressively higher levels of nuclear pIGF-1R. Both statistical models corroborated previous studies that implicate IGF-1R/mTOR in aggressive osteosarcoma [19, 30, 48]. Indeed, these results suggest that IGF-1R or downstream signaling proteins may be attractive targets for therapy in osteosarcoma.
When tested in the clinic, IGF/PI3K/mTOR-directed therapies have yielded mixed results. This has led some groups to hypothesize that patient stratification based on IGF-1R expression may be necessary to select appropriate patients for targeted therapy [18, 19]. It is also plausible that single-agent IGF-1R-directed monoclonal antibodies and small molecule inhibitors are not effective enough to fully abrogate downstream mTOR signaling, as demonstrated in Ewing sarcoma preclinical models [49, 50].
Analysis of the TMA revealed consistent expression of YAP/TAZ N:C ratio across patient samples. A potential weakness of our analysis is that the biopsy specimens analyzed in this study were collected post-treatment from variable sites, as indicated in Table 1 and Supplementary Figure 2A. While prior studies assessed the relationship between nuclear or total YAP/TAZ and clinical outcome, we hypothesized that the N:C ratio of mean intensities for cells in each biopsy would serve as a proxy measure of YAP/TAZ activation [26, 43]. This approach allowed us to compare ratios of protein localization across tumor biopsies and determine the extent to which YAP/TAZ function is associated with cancer pathogenesis in multiple tumor types [25]. While univariate Kaplan–Meier survival analysis did not glean any statically significant association between YAP or TAZ N:C ratio and overall survival, a selected Cox model suggested that a low YAP N:C ratio is associated with decreased overall survival but only compared to the medium N:C ratio group not the high N:C ratio group. This result suggests that average values for YAP and TAZ N:C ratios may have prognostic value.
Our research identifies a potentially clinically important relationship between the Hippo pathway and IGF-1R/PI3K/mTOR pathway signaling. Other studies have shown that nuclear YAP and TAZ potentiates the signaling in the canonical IGF-1R/mTOR cascade as a potential mechanistic reason for this relationship [51, 52]. Though single-agent therapies against IGF-1R or mTOR have proved ineffective in osteosarcoma, the ability to co-target these pathways using novel biologically-targeted agents would be expected to yield synergistic anti-cancer activity, as has been observed in preclinical bone sarcoma models [53].
While sole targeting of the IGF/PI3K/mTOR cascade has had limited success in early phase osteosarcoma trials, our study suggests that nuclear pIGF-1R might serve as a prognostic biomarker to identify osteosarcoma patients that have an especially poor prognosis. Of course, prospective studies will be required to determine if patient selection, based on their nuclear pIGF-1R status, correlates with clinical response to IGF/PI3K/mTOR-directed therapies. Presently, most pharmaceutical companies have stopped the development of their IGF1 or IGF-1R-targeted drug candidates. This leaves only PI3K or mTOR as potential targets within this pathway. Given the likely pathway crosstalk between YAP/TAZ and IGF/PI3K/mTOR, one might hypothesize that a dual-targeting approach will have synergistic antineoplastic activity. This remains to be explored now that second-generation inhibitors of YAP1 (NCT04659096) and TEAD have reached the clinic.
Materials and Methods
Study design
To establish our model’s clinical relevance, we used available osteosarcoma TMA samples consisting of two slides representing sequential tumor sections from 37 patients treated at The University of Texas MD Anderson Cancer Center. The TMA slide stained for YAP/TAZ lost one patient sample during preparation. When duplicate patient samples present on the slide, their results were averaged to give the tumor staining scores. The algorithms used to evaluate intensities in TMA samples were applied by a researcher blinded to the patients’ outcomes. Patient outcomes, demographics, and staging were compiled separately and not available to the researchers evaluating images of de-identified patient samples before analysis was complete.
Statistical analyses
All statistics were performed using JMP Pro 14 or GraphPad Prism. Data are displayed as means or medians with standard deviations or as box blots indicating quartile ranges, as indicated in figure captions. Patient sample staining values were compared between groups using paired t-tests or one-way ANOVA with post-hoc Tukey’s HSD (if over two groups) or Sidak correction (2 groups) as described in figure captions with *p < 0.05, **p < 0.01, #p ≤ 0.001.
For Kaplan–Meier curve analyses and initial proportional hazards model, variables were selected a priori. The log-rank test was used to compare groups in Kaplan–Meier curves and done pairwise when three or more groups were present as indicated. To avoid overfitting our proportional hazards model to a small sample size, we selected patient variables for predicting death during the observation period using the Bayesian information criterion (BIC) method in JMP Pro 14. For completeness, we provide results of Cox models that were fit to a variable selected a priori and variables selected by the BIC.
Preparation of tissue microarray
All aspects of our research were performed by recognized ethical guidelines (e.g., Declaration of Helsinki, Belmont Report). A TMA was constructed from archival surgical pathology materials comprising 38 formalin-fixed, paraffin-embedded tissues from 37 patients. Areas of viable tumor were selected by pathologist review of whole slide H&E-stained sections. Selected areas were punched and transferred, in duplicate, to a recipient block using an ATA-100 Advanced Tissue Arrayer (Chemicon International). All human specimens were utilized under an Institutional Review Board-approved research protocol allowing for the retrospective sampling and analysis of existing archival materials collected in the course of normal patient care. Immunofluorescence studies were performed using an antigen retrieval microwave (Biogenex) and manual staining with antibodies and stains described below.
Immunofluorescence staining for confocal microscopy
Samples were then washed three times in 1× PBS on a shaker for 5 min before being blocked with blocking buffer (1 × PBS, 5% normal goat serum, 0.3% Triton™ X-100) for 60 min. Primary antibodies were then diluted 1:100 in antibody dilution buffer (1 × PBS, 1% BSA, 0.3% Triton™ X-100), and samples were incubated in primary antibody overnight at 4°C. Primary antibodies used were targeted to YAP (Santa Cruz, sc-271134), TAZ (Abcam, ab-84927), IGF-1R (Santa Cruz, sc-461), and pIGF-1R (Santa Cruz, sc-101703). Samples were then washed three times with 1 × PBS before incubating with secondary antibody (diluted 1:500, Cell Signaling Technology) and phalloidin conjugated to iFluor 488 (Abcam, diluted 1:1000, only in YAP/TAZ TMA) in antibody dilution buffer for 1.5 hours in the dark at room temperature. After three washes, cells were incubated for 10 min in a 1:5000 dilution of a 1 mg/mL Hoechst 33342 stock solution (ThermoFisher). Samples were then rinsed once with PBS. Coverslips were mounted onto slides using Prolong® Gold Antifade Reagent (Cell Signaling Technology).
Because all samples were on one slide, the same power for each wavelength laser illumination was used for each sample to directly compare patient markers across samples. Individual tumor samples were stained for tumor markers of interest and nuclei and imaged using 20× objective by stitching images in a two-by-two image grid with 10% overlap to create one large image file for each tumor sample.
Quantitative image analysis
All quantitative imaging analysis was done with IMARIS software (V8.6, Bitplane) licensed to Rice University. For YAP and TAZ, all areas staining positive for the Hoechst stain in the blue channel were considered nuclear domains. All areas staining positive for YAP in the red channel but not staining positive for nuclei in the blue channel were considered cytoplasmic domains. Mean nuclear and cytoplasmic intensity values for markers in the domains were identified by the IMARIS “cells” algorithm were then averaged after weighting each distinct identified domain by total area. Because individual cells were not able to be segmented in the highly cellular sections, we weighted each identified nuclear or cytoplasmic domain by area and averaged by the total area of the nuclear or cytoplasmic domain. In cases where the TMA contained two or more samples from the same patient, values were averaged between these two sections for further analysis. With this technique, average nuclear intensity and average cytoplasmic intensity for each tumor sample across the entire image were generated using the following formula:
Where n is the total number of nuclear or cytoplasmic domains identified in each patient sample.
For TMA stained for IGF-1R and pIGF-1R, individual nuclei were identified by distinct areas staining positive for the Hoechst stain in the blue channel. Values for mean nuclear intensities for the entire sample were then calculated by averaging the nuclear intensity of IGF-1R/pIGF-1R signals colocalizing with the blue channel. Each mean nuclear value was weighted by the nuclear area and averaged for each patient in the following formula:
Where n is the total number of nuclear domains identified in all samples from the same patient.
Conclusions
In this study, we demonstrated a correlation between nuclear IGF-1R and the Hippo pathway effectors, YAP and TAZ staining in human osteosarcoma. Further, we show that high nuclear-phosphorylated IGF-1R and low YAP N:C ratio are potentially negative prognostic indicators for overall survival in osteosarcoma. While histotype-specific YAP/TAZ activity must be confirmed in a larger dataset, our results imply a potential need for patient selection in trials directed at the IGF/PI3K/mTOR or Hippo pathways. Given the rarity of osteosarcoma, clinical validation of our results will almost certainly require the active participation of national and international high-volume cancer centers.
Data availability
All data is represented in figures in the main text or supplemental materials. Summary data of average values in each specimen are included in Supplemental Data Set 1. Raw proportional hazards model calculations are included in Supplemental Data Sets 2 and 3. Raw data for segmented images and microscopy images are available upon request to the corresponding author.
ACKNOWLEDGMENTS
We thank the Rice University Shared Equipment Authority and Alloysius Budi Utama for his insight on the use of confocal microscopy and quantitative imaging.
CONFLICTS OF INTEREST
Authors have no conflicts of interest to declare.
FUNDING
Research Support: Investigators at The University of Texas MD Anderson Cancer Center are supported by the National Institutes of Health through Cancer Center Support Grant CA016672. E.R.M. received support from the Baylor College of Medicine Medical Scientist Training Program and NIH grants (F31CA213994). J.A.L. is funded by NIH [R01CA180279]. A.G.M. is funded by NIH [R01CA180279 and P41EB023833]. The Characterized Cell Line Core Facility is funded by NIH [CA016672].
References
1. Isakoff MS, Bielack SS, Meltzer P, Gorlick R. Osteosarcoma: Current Treatment and a Collaborative Pathway to Success. J Clin Oncol. 2015; 33:3029–35. https://doi.org/10.1200/JCO.2014.59.4895. [PubMed].
2. Kansara M, Teng MW, Smyth MJ, Thomas DM. Translational biology of osteosarcoma. Nat Rev Cancer. 2014; 14:722–35. https://doi.org/10.1038/nrc3838. [PubMed].
3. Brown HK, Tellez-Gabriel M, Heymann D. Cancer stem cells in osteosarcoma. Cancer Lett. 2017; 386:189–95. https://doi.org/10.1016/j.canlet.2016.11.019. [PubMed].
4. He JP, Hao Y, Wang XL, Yang XJ, Shao JF, Guo FJ, Feng JX. Review of the molecular pathogenesis of osteosarcoma. Asian Pac J Cancer Prev. 2014; 15:5967–76. https://doi.org/10.7314/apjcp.2014.15.15.5967. [PubMed].
5. Chan H, Pratt CB. A new familial cancer syndrome? A spectrum of malignant and benign tumors including retinoblastoma, carcinoma of the bladder and other genitourinary tumors, thyroid adenoma, and a probable case of multifocal osteosarcoma. J Natl Cancer Inst. 1977; 58:205–7. https://doi.org/10.1093/jnci/58.2.205. [PubMed].
6. Hansen MF, Koufos A, Gallie BL, Phillips RA, Fodstad O, Brøgger A, Gedde-Dahl T, Cavenee WK. Osteosarcoma and retinoblastoma: a shared chromosomal mechanism revealing recessive predisposition. Proc Natl Acad Sci U S A. 1985; 82:6216–20. https://doi.org/10.1073/pnas.82.18.6216. [PubMed].
7. Malkin D, Li FP, Strong LC, Fraumeni JF Jr, Nelson CE, Kim DH, Kassel J, Gryka MA, Bischoff FZ, Tainsky MA, Friend SH. Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science. 1990; 250:1233–38. https://doi.org/10.1126/science.1978757. [PubMed].
8. Forment JV, Kaidi A, Jackson SP. Chromothripsis and cancer: causes and consequences of chromosome shattering. Nat Rev Cancer. 2012; 12:663–70. https://doi.org/10.1038/nrc3352. [PubMed].
9. Zhang CZ, Spektor A, Cornils H, Francis JM, Jackson EK, Liu S, Meyerson M, Pellman D. Chromothripsis from DNA damage in micronuclei. Nature. 2015; 522:179–84. https://doi.org/10.1038/nature14493. [PubMed].
10. Marcozzi A, Pellestor F, Kloosterman WP. The Genomic Characteristics and Origin of Chromothripsis. Methods Mol Biol. 2018; 1769:3–19. https://doi.org/10.1007/978-1-4939-7780-2_1. [PubMed].
11. Chawla SP, Staddon AP, Baker LH, Schuetze SM, Tolcher AW, D’Amato GZ, Blay JY, Mita MM, Sankhala KK, Berk L, Rivera VM, Clackson T, Loewy JW, et al. Phase II study of the mammalian target of rapamycin inhibitor ridaforolimus in patients with advanced bone and soft tissue sarcomas. J Clin Oncol. 2012; 30:78–84. https://doi.org/10.1200/JCO.2011.35.6329. [PubMed].
12. Demetri GD, Chawla SP, Ray-Coquard I, Le Cesne A, Staddon AP, Milhem MM, Penel N, Riedel RF, Bui-Nguyen B, Cranmer LD, Reichardt P, Bompas E, Alcindor T, et al. Results of an international randomized phase III trial of the mammalian target of rapamycin inhibitor ridaforolimus versus placebo to control metastatic sarcomas in patients after benefit from prior chemotherapy. J Clin Oncol. 2013; 31:2485–92. https://doi.org/10.1200/JCO.2012.45.5766. [PubMed].
13. Sclafani F, Kim TY, Cunningham D, Kim TW, Tabernero J, Schmoll HJ, Roh JK, Kim SY, Park YS, Guren TK, Hawkes E, Clarke SJ, Ferry D, et al. A Randomized Phase II/III Study of Dalotuzumab in Combination With Cetuximab and Irinotecan in Chemorefractory, KRAS Wild-Type, Metastatic Colorectal Cancer. J Natl Cancer Inst. 2015; 107:djv258. https://doi.org/10.1093/jnci/djv258. [PubMed].
14. Naing A, LoRusso P, Fu S, Hong DS, Anderson P, Benjamin RS, Ludwig J, Chen HX, Doyle LA, Kurzrock R. Insulin growth factor-receptor (IGF-1R) antibody cixutumumab combined with the mTOR inhibitor temsirolimus in patients with refractory Ewing’s sarcoma family tumors. Clin Cancer Res. 2012; 18:2625–31. https://doi.org/10.1158/1078-0432.CCR-12-0061. [PubMed].
15. Schwartz GK, Tap WD, Qin LX, Livingston MB, Undevia SD, Chmielowski B, Agulnik M, Schuetze SM, Reed DR, Okuno SH, Ludwig JA, Keedy V, Rietschel P, et al. Cixutumumab and temsirolimus for patients with bone and soft-tissue sarcoma: a multicentre, open-label, phase 2 trial. Lancet Oncol. 2013; 14:371–82. https://doi.org/10.1016/S1470-2045(13)70049-4. [PubMed].
16. Aleksic T, Chitnis MM, Perestenko OV, Gao S, Thomas PH, Turner GD, Protheroe AS, Howarth M, Macaulay VM. Type 1 insulin-like growth factor receptor translocates to the nucleus of human tumor cells. Cancer Res. 2010; 70:6412–19. https://doi.org/10.1158/0008-5472.CAN-10-0052. [PubMed].
17. Packham S, Warsito D, Lin Y, Sadi S, Karlsson R, Sehat B, Larsson O. Nuclear translocation of IGF-1R via p150(Glued) and an importin-β/RanBP2-dependent pathway in cancer cells. Oncogene. 2015; 34:2227–38. https://doi.org/10.1038/onc.2014.165. [PubMed].
18. Asmane I, Watkin E, Alberti L, Duc A, Marec-Berard P, Ray-Coquard I, Cassier P, Decouvelaere AV, Ranchère D, Kurtz JE, Bergerat JP, Blay JY. Insulin-like growth factor type 1 receptor (IGF-1R) exclusive nuclear staining: a predictive biomarker for IGF-1R monoclonal antibody (Ab) therapy in sarcomas. Eur J Cancer. 2012; 48:3027–35. https://doi.org/10.1016/j.ejca.2012.05.009. [PubMed].
19. Liang J, Li B, Yuan L, Ye Z. Prognostic value of IGF-1R expression in bone and soft tissue sarcomas: a meta-analysis. Onco Targets Ther. 2015; 8:1949–55. https://doi.org/10.2147/OTT.S88293. [PubMed].
20. Discher DE, Janmey P, Wang YL. Tissue cells feel and respond to the stiffness of their substrate. Science. 2005; 310:1139–43. https://doi.org/10.1126/science.1116995. [PubMed].
21. Dupont S, Morsut L, Aragona M, Enzo E, Giulitti S, Cordenonsi M, Zanconato F, Le Digabel J, Forcato M, Bicciato S, Elvassore N, Piccolo S. Role of YAP/TAZ in mechanotransduction. Nature. 2011; 474:179–83. https://doi.org/10.1038/nature10137. [PubMed].
22. Engler AJ, Sen S, Sweeney HL, Discher DE. Matrix elasticity directs stem cell lineage specification. Cell. 2006; 126:677–89. https://doi.org/10.1016/j.cell.2006.06.044. [PubMed].
23. Caliari SR, Vega SL, Kwon M, Soulas EM, Burdick JA. Dimensionality and spreading influence MSC YAP/TAZ signaling in hydrogel environments. Biomaterials. 2016; 103:314–23. https://doi.org/10.1016/j.biomaterials.2016.06.061. [PubMed].
24. Molina ER, Chim LK, Salazar MC, Mehta SM, Menegaz BA, Lamhamedi-Cherradi SE, Satish T, Mohiuddin S, McCall D, Zaske AM, Cuglievan B, Lazar AJ, Scott DW, et al. Mechanically tunable coaxial electrospun models of YAP/TAZ mechanoresponse and IGF-1R activation in osteosarcoma. Acta Biomater. 2019; 100:38–51. https://doi.org/10.1016/j.actbio.2019.09.029. [PubMed].
25. Ahmed AA, Mohamed AD, Gener M, Li W, Taboada E. YAP and the Hippo pathway in pediatric cancer. Mol Cell Oncol. 2017; 4:e1295127. https://doi.org/10.1080/23723556.2017.1295127. [PubMed].
26. Bouvier C, Macagno N, Nguyen Q, Loundou A, Jiguet-Jiglaire C, Gentet JC, Jouve JL, Rochwerger A, Mattei JC, Bouvard D, Salas S. Prognostic value of the Hippo pathway transcriptional coactivators YAP/TAZ and β1-integrin in conventional osteosarcoma. Oncotarget. 2016; 7:64702–10. https://doi.org/10.18632/oncotarget.11876. [PubMed].
27. Wang DY, Wu YN, Huang JQ, Wang W, Xu M, Jia JP, Han G, Mao BB, Bi WZ. Hippo/YAP signaling pathway is involved in osteosarcoma chemoresistance. Chin J Cancer. 2016; 35:47. https://doi.org/10.1186/s40880-016-0109-z. [PubMed].
28. Hong JH, Hwang ES, McManus MT, Amsterdam A, Tian Y, Kalmukova R, Mueller E, Benjamin T, Spiegelman BM, Sharp PA, Hopkins N, Yaffe MB. TAZ, a transcriptional modulator of mesenchymal stem cell differentiation. Science. 2005; 309:1074–78. https://doi.org/10.1126/science.1110955. [PubMed].
29. Pakos EE, Nearchou AD, Grimer RJ, Koumoullis HD, Abudu A, Bramer JA, Jeys LM, Franchi A, Scoccianti G, Campanacci D, Capanna R, Aparicio J, Tabone MD, et al. Prognostic factors and outcomes for osteosarcoma: an international collaboration. Eur J Cancer. 2009; 45:2367–75. https://doi.org/10.1016/j.ejca.2009.03.005. [PubMed].
30. Wang YH, Han XD, Qiu Y, Xiong J, Yu Y, Wang B, Zhu ZZ, Qian BP, Chen YX, Wang SF, Shi HF, Sun X. Increased expression of insulin-like growth factor-1 receptor is correlated with tumor metastasis and prognosis in patients with osteosarcoma. J Surg Oncol. 2012; 105:235–43. https://doi.org/10.1002/jso.22077. [PubMed].
31. Vasquez L, Tarrillo F, Oscanoa M, Maza I, Geronimo J, Paredes G, Silva JM, Sialer L. Analysis of Prognostic Factors in High-Grade Osteosarcoma of the Extremities in Children: A 15-Year Single-Institution Experience. Front Oncol. 2016; 6:22. https://doi.org/10.3389/fonc.2016.00022. [PubMed].
32. Mialou V, Philip T, Kalifa C, Perol D, Gentet JC, Marec-Berard P, Pacquement H, Chastagner P, Defaschelles AS, Hartmann O. Metastatic osteosarcoma at diagnosis: prognostic factors and long-term outcome--the French pediatric experience. Cancer. 2005; 104:1100–9. https://doi.org/10.1002/cncr.21263. [PubMed].
33. Lin PP, Patel S. Osteosarcoma. Bone Sarcoma. MD Anderson Cancer Care Series. Boston (MA): Springer. 2013; 75–97. https://doi.org/10.1007/978-1-4614-5194-5_5.
34. Hauben EI, Weeden S, Pringle J, Van Marck EA, Hogendoorn PC. Does the histological subtype of high-grade central osteosarcoma influence the response to treatment with chemotherapy and does it affect overall survival? A study on 570 patients of two consecutive trials of the European Osteosarcoma Intergroup. Eur J Cancer. 2002; 38:1218–25. https://doi.org/10.1016/s0959-8049(02)00037-0. [PubMed].
35. Whelan JS, Jinks RC, McTiernan A, Sydes MR, Hook JM, Trani L, Uscinska B, Bramwell V, Lewis IJ, Nooij MA, van Glabbeke M, Grimer RJ, Hogendoorn PC, et al. Survival from high-grade localised extremity osteosarcoma: combined results and prognostic factors from three European Osteosarcoma Intergroup randomised controlled trials. Ann Oncol. 2012; 23:1607–16. https://doi.org/10.1093/annonc/mdr491. [PubMed].
36. Chui MH, Kandel RA, Wong M, Griffin AM, Bell RS, Blackstein ME, Wunder JS, Dickson BC. Histopathologic Features of Prognostic Significance in High-Grade Osteosarcoma. Arch Pathol Lab Med. 2016; 140:1231–42. https://doi.org/10.5858/arpa.2015-0389-OA. [PubMed].
37. Thampi S, Matthay KK, Boscardin WJ, Goldsby R, DuBois SG. Clinical Features and Outcomes Differ between Skeletal and Extraskeletal Osteosarcoma. Sarcoma. 2014; 2014:902620. https://doi.org/10.1155/2014/902620. [PubMed].
38. Camp RL, Chung GG, Rimm DL. Automated subcellular localization and quantification of protein expression in tissue microarrays. Nat Med. 2002; 8:1323–27. https://doi.org/10.1038/nm791. [PubMed].
39. Yuen JS, Macaulay VM. Targeting the type 1 insulin-like growth factor receptor as a treatment for cancer. Expert Opin Ther Targets. 2008; 12:589–603. https://doi.org/10.1517/14728222.12.5.589. [PubMed].
40. Meric-Bernstam F, Gonzalez-Angulo AM. Targeting the mTOR signaling network for cancer therapy. J Clin Oncol. 2009; 27:2278–87. https://doi.org/10.1200/JCO.2008.20.0766. [PubMed].
41. Larsson O, Girnita A, Girnita L. Role of insulin-like growth factor 1 receptor signalling in cancer. Br J Cancer. 2005; 92:2097–101. https://doi.org/10.1038/sj.bjc.6602627. [PubMed].
42. Aragona M, Panciera T, Manfrin A, Giulitti S, Michielin F, Elvassore N, Dupont S, Piccolo S. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell. 2013; 154:1047–59. https://doi.org/10.1016/j.cell.2013.07.042. [PubMed].
43. Chan LH, Wang W, Yeung W, Deng Y, Yuan P, Mak KK. Hedgehog signaling induces osteosarcoma development through Yap1 and H19 overexpression. Oncogene. 2014; 33:4857–66. https://doi.org/10.1038/onc.2013.433. [PubMed].
44. Zhang YH, Li B, Shen L, Shen Y, Chen XD. The role and clinical significance of YES-associated protein 1 in human osteosarcoma. Int J Immunopathol Pharmacol. 2013; 26:157–67. https://doi.org/10.1177/039463201302600115. [PubMed].
45. Shen S, Huang K, Wu Y, Ma Y, Wang J, Qin F, Ma J. A miR-135b-TAZ positive feedback loop promotes epithelial-mesenchymal transition (EMT) and tumorigenesis in osteosarcoma. Cancer Lett. 2017; 407:32–44. https://doi.org/10.1016/j.canlet.2017.08.005. [PubMed].
46. Tumaneng K, Schlegelmilch K, Russell RC, Yimlamai D, Basnet H, Mahadevan N, Fitamant J, Bardeesy N, Camargo FD, Guan KL. YAP mediates crosstalk between the Hippo and PI(3)K–TOR pathways by suppressing PTEN via miR-29. Nat Cell Biol. 2012; 14:1322–29. https://doi.org/10.1038/ncb2615. [PubMed].
47. Goodman CA, Dietz JM, Jacobs BL, McNally RM, You JS, Hornberger TA. Yes-Associated Protein is up-regulated by mechanical overload and is sufficient to induce skeletal muscle hypertrophy. FEBS Lett. 2015; 589:1491–97. https://doi.org/10.1016/j.febslet.2015.04.047. [PubMed].
48. Hu K, Dai HB, Qiu ZL. mTOR signaling in osteosarcoma: Oncogenesis and therapeutic aspects (Review). Oncol Rep. 2016; 36:1219–25. https://doi.org/10.3892/or.2016.4922. [PubMed].
49. Kolb EA, Gorlick R, Houghton PJ, Morton CL, Lock R, Carol H, Reynolds CP, Maris JM, Keir ST, Billups CA, Smith MA. Initial testing (stage 1) of a monoclonal antibody (SCH 717454) against the IGF-1 receptor by the pediatric preclinical testing program. Pediatr Blood Cancer. 2008; 50:1190–97. https://doi.org/10.1002/pbc.21450. [PubMed].
50. Cao Y, Roth M, Piperdi S, Montoya K, Sowers R, Rao P, Geller D, Houghton P, Kolb EA, Gill J, Gorlick R. Insulin-like growth factor 1 receptor and response to anti-IGF1R antibody therapy in osteosarcoma. PLoS One. 2014; 9:e106249. https://doi.org/10.1371/journal.pone.0106249. [PubMed].
51. Xin M, Kim Y, Sutherland LB, Qi X, McAnally J, Schwartz RJ, Richardson JA, Bassel-Duby R, Olson EN. Regulation of insulin-like growth factor signaling by Yap governs cardiomyocyte proliferation and embryonic heart size. Sci Signal. 2011; 4:ra70. https://doi.org/10.1126/scisignal.2002278. [PubMed].
52. Hansen CG, Ng YL, Lam WL, Plouffe SW, Guan KL. The Hippo pathway effectors YAP and TAZ promote cell growth by modulating amino acid signaling to mTORC1. Cell Res. 2015; 25:1299–313. https://doi.org/10.1038/cr.2015.140. [PubMed].
53. Kolb EA, Gorlick R, Maris JM, Keir ST, Morton CL, Wu J, Wozniak AW, Smith MA, Houghton PJ. Combination testing (Stage 2) of the Anti-IGF-1 receptor antibody IMC-A12 with rapamycin by the pediatric preclinical testing program. Pediatr Blood Cancer. 2012; 58:729–35. https://doi.org/10.1002/pbc.23157. [PubMed].
