Introduction
Cancer testis antigens (CTA) are germ cell antigens that are typically minimally expressed in normal tissues, but commonly expressed in tumors. Because of this, they are appealing therapeutic targets with less potential for off target toxicity [1]. Kita-Kyushu lung cancer antigen-1 (KK-LC-1), also known as CT 83 and CXORF61, is a CTA containing 113 amino acids that maps to chromosome Xq22 and is not expressed in normal tissues except for the testis. It has been shown to be expressed in 82% of gastric cancer tumors and 52.9% of breast cancer tumors, with even higher detection rates in triple negative breast cancer [2, 3]. In lung cancer, surgical series have shown KK-LC-1 to be expressed in about one-third of lung cancer tumors [1, 3–5]. In one study with Japanese patients, KK-LC-1 has been shown to be expressed at similar rates in adenocarcinoma and squamous cell carcinoma [5]. KK-LC-1 high expression was shown in a greater proportion of TNM stage II and III tumors compared to TNM stage I tumors [1]. There are no large series studying the prevalence of the disease in patients with stage IV NSCLC.
Cytotoxic T lymphocytes (CTLs) are an emerging modality for immunity against cancers, as antigen specific CTLs can be formulated to specifically target CTA expressing tumor cells. T-cell receptor therapy is designed to modify patient T cells with a selected T-cell receptor targeting a cancer testis antigen which can promote selective targeting of tumor antigens while limiting off target toxicity. One challenge is that not all patients have T cells that will effectively recognize their tumors. TCR-Ts in order to be effective need a high affinity towards the cancer testis antigen and many times the T cells that would be designed to recognize these tumors are eliminated during the negative selection process in the thymus [6–8]. Fukuyama et al. identified the KK-LC-1 CTA by establishing a lung adenocarcinoma cell line and induced a CTL clone from regional lymph node lymphocytes of a patient [9]. More recently, Marcinkowski et al. identified a KK-LC-1 reactive T cell receptor from tumor infiltrating lymphocytes (TIL) in a patient with cervical cancer who had a subsequent complete tumor response to TIL therapy [10]. Paret et al. also showed strong antigen-specific response in HLA-A*02 transgenic mice with KK-LC-1 encoding RNA in triple negative breast cancer cell lines. [11]. In addition, other cancer testis antigens such as MAGE-A4 and NY-ESO-1 have shown promise in lung cancer [6, 12].
The majority of non-small cell lung cancer (NSCLC) is diagnosed at an advanced, incurable stage. Five-year relative survival rate for all stages combined is 21% and only 6% for lung cancer that has spread to distant sites, according to SEER data from 2009–2015 [13]. Lung cancer treatment has made tremendous strides in the past decade with advances in targeted therapies for EGFR, ALK, ROS1, BRAF V600E, MET exon 14 skip mutation, RET, NTRK, and most recently KRAS G12C along with checkpoint inhibitor immunotherapy in other variants of the disease. Targeted therapies have shown a median progression free survival of around 9–18 months with long term results showing 5-year survival of greater than 14% in EGFR patients and 4-year survival in ALK and ROS1 gene rearrangement of 56.6% and 51.1% respectively [14–20]. While immunotherapy has shown significant improvements, standard of care immunotherapy with chemotherapy showed an ORR of 48% and a median PFS of 9 months and OS of 22 months [21].
Because the treatment algorithms have diverged between lung cancer subtypes that are amenable to targeted therapy or immunotherapy, positioning emerging treatments for future clinical trials requires understanding of which molecular subtype of lung cancer would be most appropriate for the new CTA directed immune treatments. Therefore, the aim of this study is to identify the molecular subtypes of lung cancer expressing KK-LC-1 to determine which patients would be most likely to benefit from a clinical trial of T cell receptor therapy (TCR-T) targeting KK-LC-1.
Results
The study population included a total of 9790 tumors with KK-LC-1 expression ranging from 0–266 TPM. Tumors were then categorized into four groups based on quartiles of KK-LC-1 expression (Supplementary Table 1). We observed a relatively equal median age of patients across all four quartiles of KK-LC-1 expression (Supplementary Table 1). Overall, there seemed to be a slightly higher prevalence of males in the lowest quartile (Q1) of KK-LC-1 expression and a higher prevalence of females in the highest quartile (Q4) of KK-LC-1 expression. (Table 1) We see an even greater proportion of adenocarcinoma tumors (1730/2448, 70.6%) in the tumors in Q4 compared to tumors (1294/2448, 52.9%) in Q1 (Supplementary Table 2). Adenocarcinoma tumors were mostly pan-wild type (36.5%), KRAS mutated (35.4%), and EGFR mutated (16.3%) while squamous cell carcinoma tumors were even more predominantly pan-wild type (85.5%) (Supplementary Figure 1).
Table 1: Baseline characteristics of 9790 NSCLC patients
| KK-LC-1 expression quartiles | Total | Female N (%) | Male N (%) | Median age |
|---|---|---|---|---|
| Q1 | 2448 | 1138 (46.5) | 1310 (53.5) | 69 |
| Q2 | 2447 | 1295 (52.9) | 1152 (47.1) | 69 |
| Q3 | 2447 | 1311 (53.6) | 1136 (46.4) | 69 |
| Q4 | 2448 | 1320 (53.9) | 1128 (46.1) | 68 |
| Total | 9790 | 5064 (51.7) | 4726 (48.3) | 69 |
Adenocarcinomas had significantly higher median KK-LC-1 expression than squamous cell carcinomas (3.25 vs. 1.17 transcripts per million (TPM), p < 0.0001). We also saw a significant difference between squamous cell carcinoma and unclear or mixed (1.14 vs. 3.078 TPM, p < 0.001). KK-LC-1 expression between other subtypes were not statistically different (Figure 1).
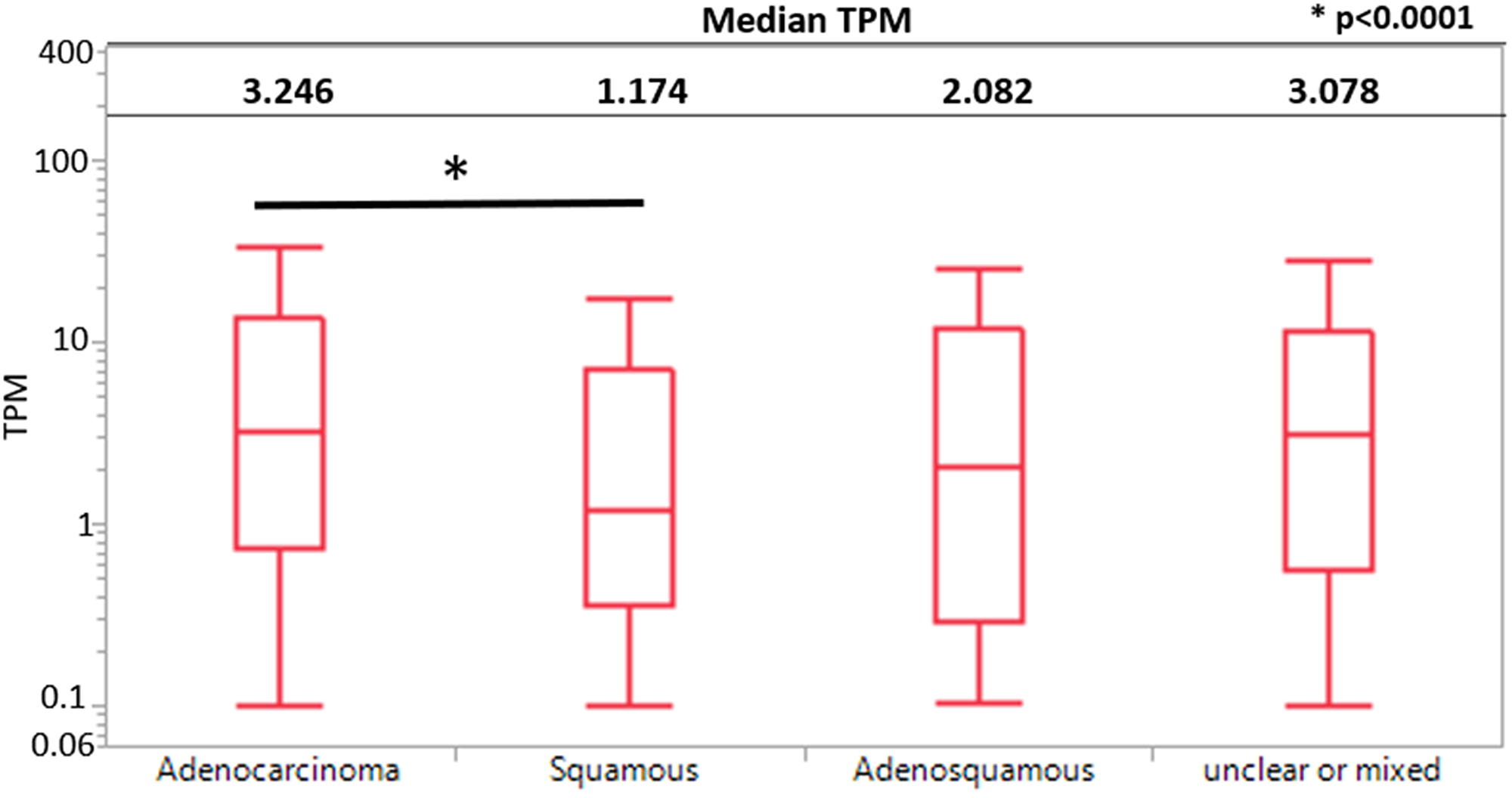
Figure 1: KK-LC-1 expression by histological subtypes between adenocarcinoma, squamous cell carcinoma, adenosquamous carcinoma, and unclear or mixed carcinomas using a boxplot. TPM expression is represented here using a log scale.
Tumors within the highest quartile of KK-LC-1 expression (Q4) had a greater proportion of TMB > 10 mutations per megabase (mt/MB) (44% vs. 28%) compared to Q1. No difference was seen in PD-L1 expression (Figure 2). In adenocarcinoma, Q4 had a higher TMB (9 vs. 5 in Q1).
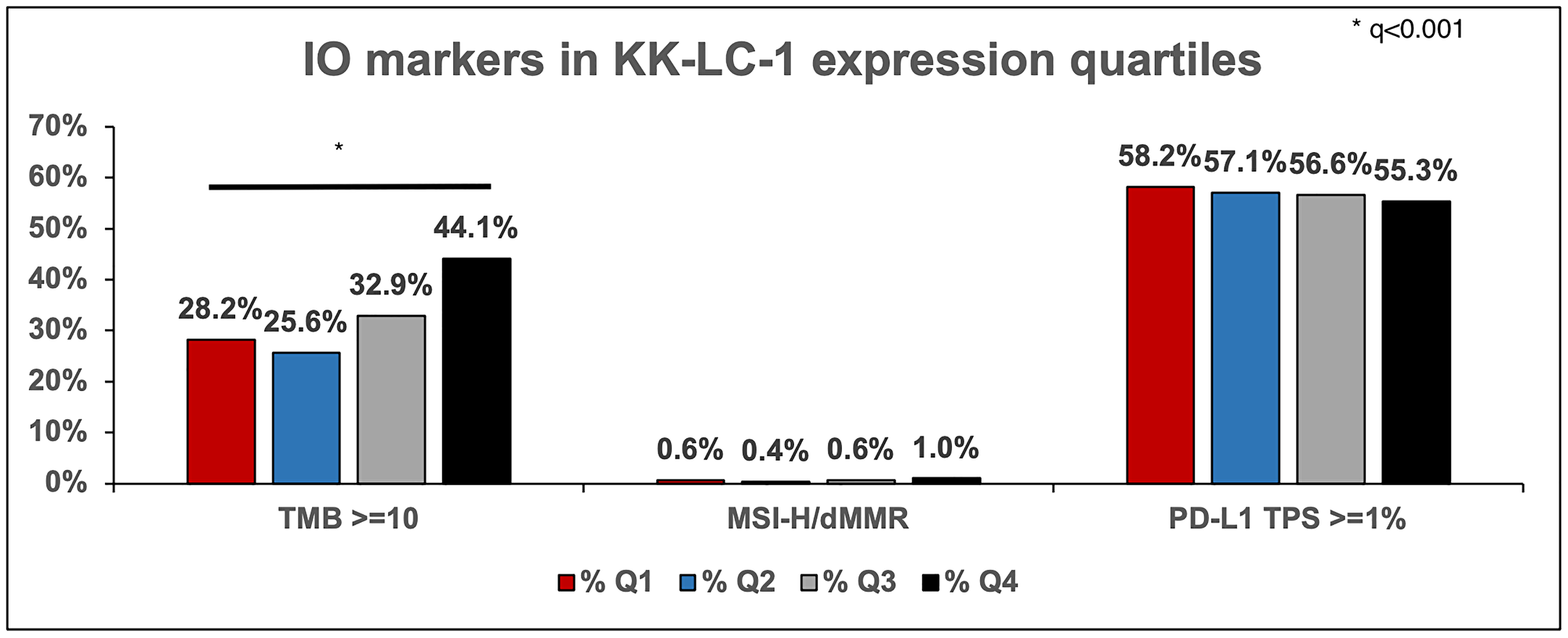
Figure 2: IO markers in tumors in their respective quartile of KK-LC-1 expression that have TMB >= 10, MSI-H/dMMR, and PD-L1 with TPS >= 1%.
There was a statistically higher KRAS mutation prevalence in Q3/Q4 (34.8%/35.0%) than Q1/Q2 (22%/29%) but a lower ALK fusion prevalence in Q3/4 (1.0%/0.5%) compared to Q1/2 (3.3%/2.6%) (Figure 3A, 3B). There is statistically higher expression of KK-LC-1 in pan wild type (3.95 TPM) compared to tumors with EGFR mutation (1.95 TPM, p < 0.0001), ALK fusion (0.6 TPM, p < 0.0001), MET exon-14-skip (1.22 TPM, p < 0.0001), RET fusion (1.42 TPM, p = 0.0001), and ROS1 fusion (1.78 TPM, p = 0.0162) (Figure 4).
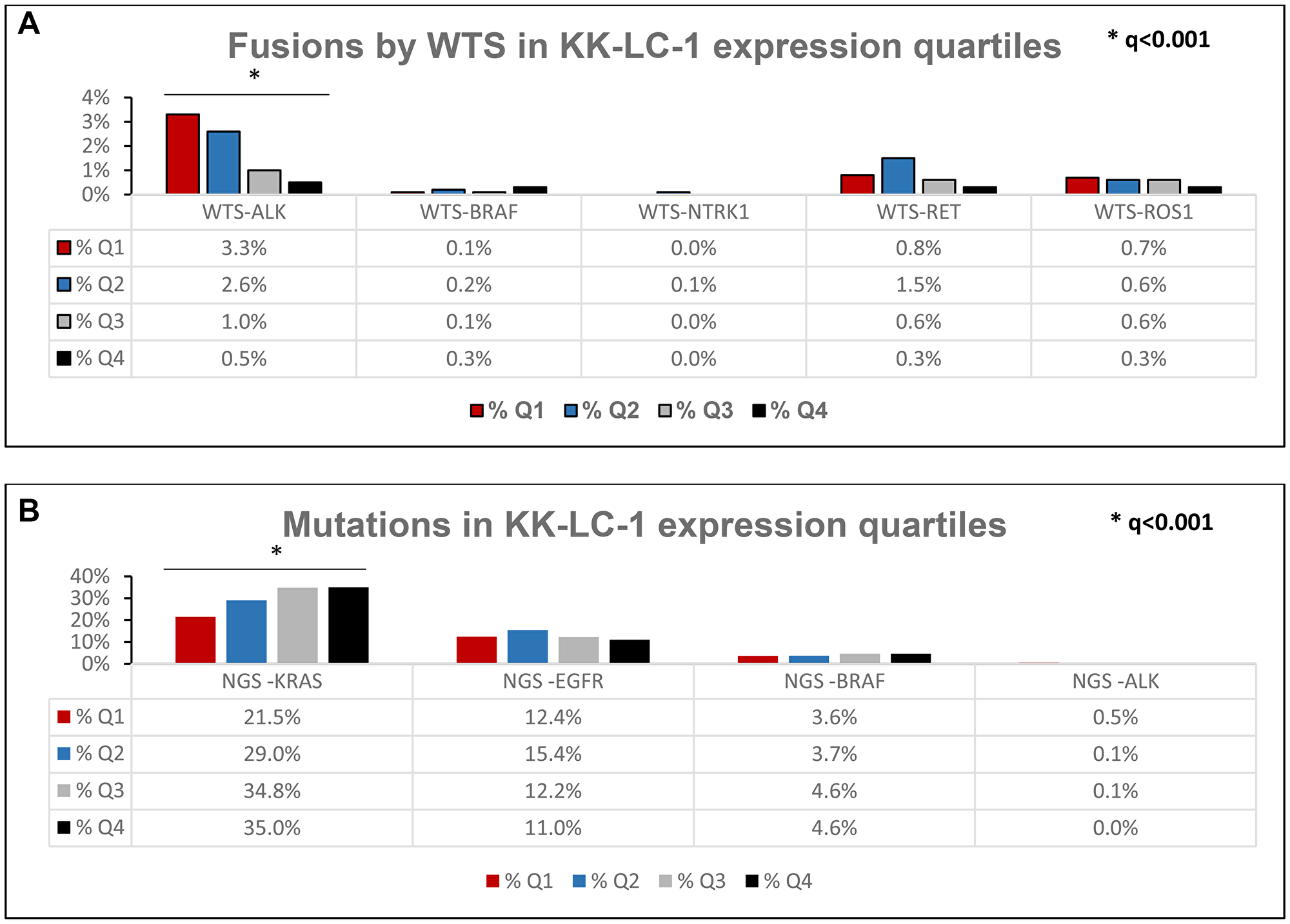
Figure 3: (A) Prevalence of fusions based on KK-LC-1 expression quartiles. (B) Prevalence of oncogenic mutations based on KK-LC-1 quartiles. Further breakdown of KRAS, EGFR, and BRAF mutations are noted in Supplementary Tables 3–5. NGS-ALK mutations refer to pathogenic/likely ALK mutations.
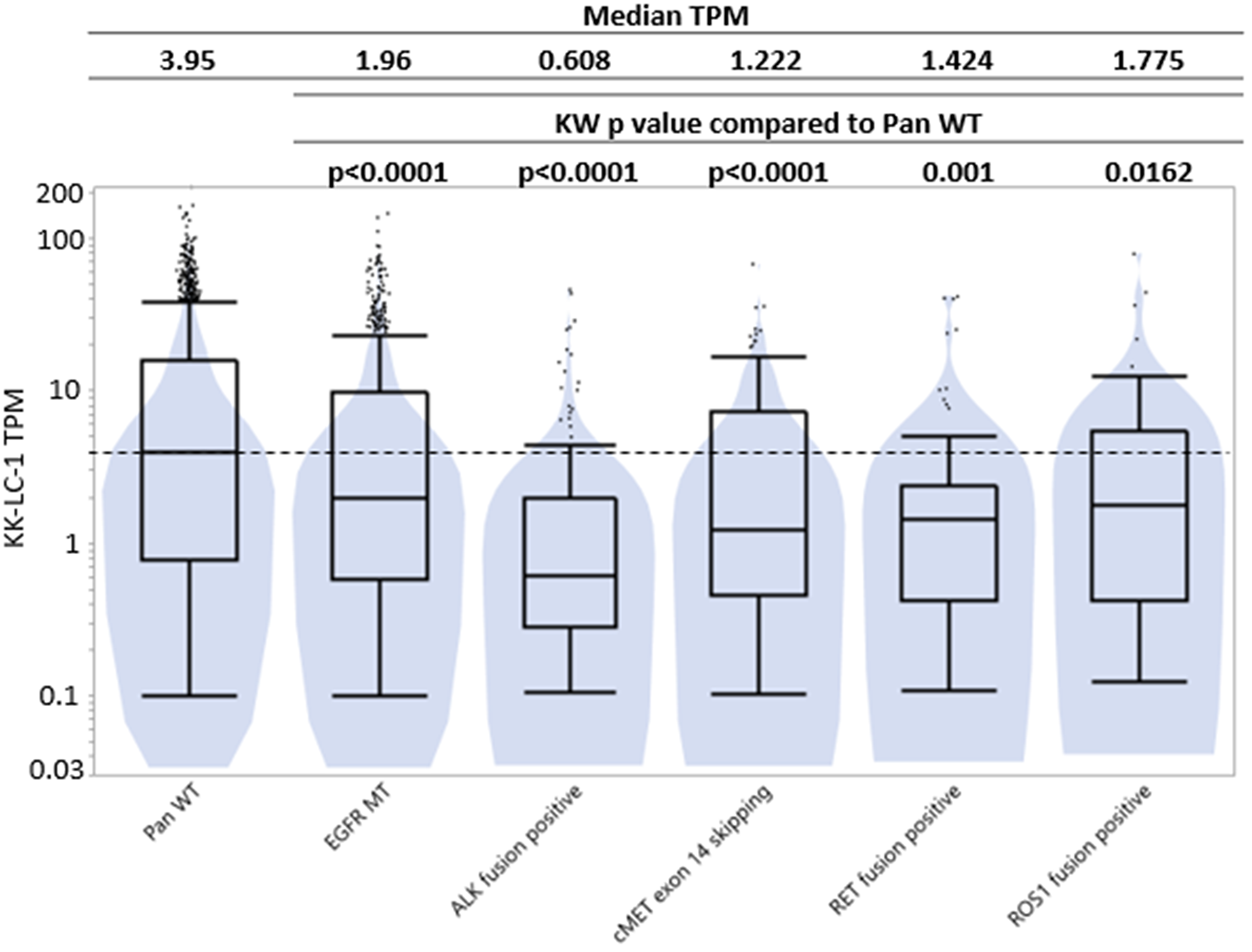
Figure 4: KK-LC-1 expression by transcripts per million (log-scale) based on whether NSCLC tumors were pan-wild type, EGFR mutant, ALK fusion positive, cMET exon 14 skipping, RET fusion positive, and ROS1 fusion positive tumors. TPM expression is represented here using a log scale.
Actionable mutations (ALK, RET, ROS1) are significantly more prevalent in KK-LC-1 low expressors (Q1) where as KRAS mutations are significantly more prevalent in KK-LC-1 high expressors (Q4) (Supplementary Figures 2 and 3).
Specifically, KRAS G12C mutations trended higher in KK-LC-1 Q4 when compared to Q1 (p = 0.0258) (Supplementary Table 3). There was no significance in number of EGFR sensitizing or resistant mutations when comparing KK-LC-1 Q1 vs. Q4. (Supplementary Table 4) In addition, no significance in difference among BRAF mutations was seen across the KK-LC-1 expression quartiles (Supplementary Tables 5 and 6).
When comparing Q4 vs. Q1 in the tumor microenvironment for KK-LC-1, we see a greater proportion of M1 macrophages in Q4 tumors whereas there is a lower proportion of M2 macrophages and CD4+ T cells (Figure 5, Supplementary Figure 4).
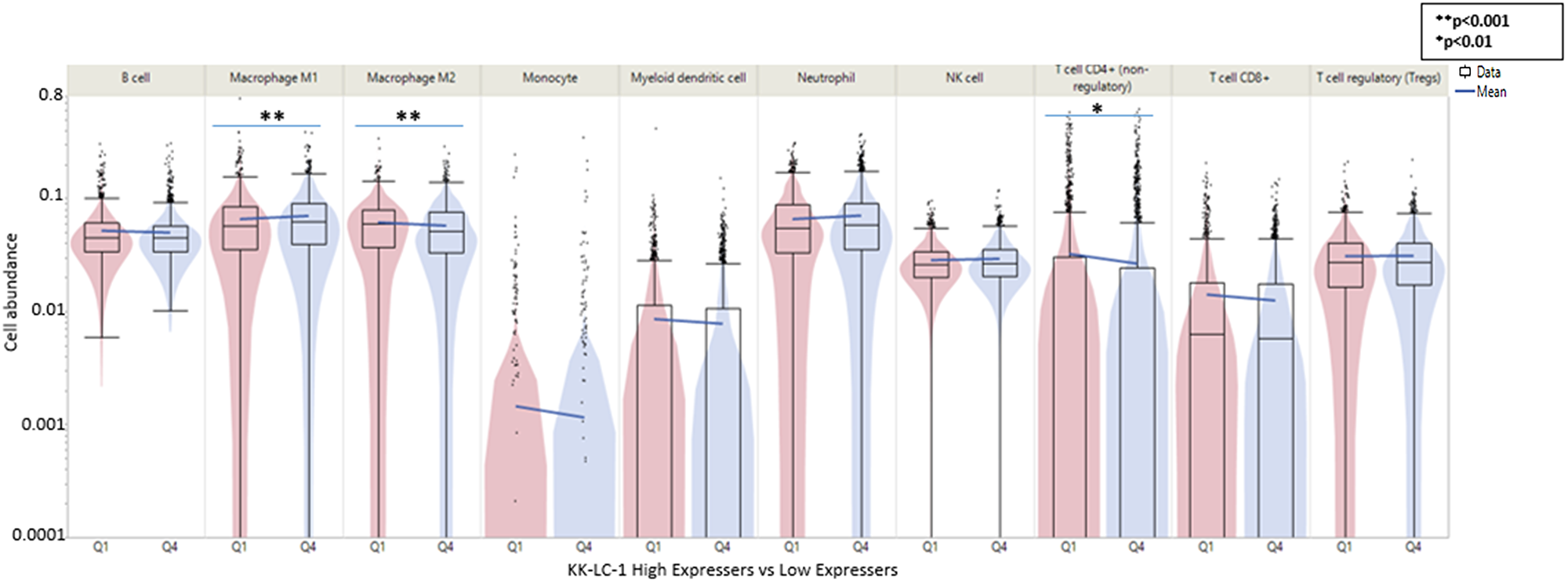
Figure 5: Tumor microenvironment in KK-LC-1 high vs low expressing (Q4 vs. Q1) in adenocarcinomas as measured by immune cell fraction calculated by QuantiSeq. The elements of the tumor microenvironment calculated include B cell, M1 macrophage, M2 macrophage, monocytes, myeloid dendritic cells, neutrophils, NK cells, CD4+ T cells, CD8+ T cells, and regulatory T cells.
DISCUSSION
Models of cancer tumorigenesis have focused cancer cells arising from molecular mutations in mature somatic cells. However, pluripotent stem cells can give rise to either tissue composed of mature somatic tissue or immature embryonic tissue by ectopically induced transcription factors [22]. Both hematopoietic and solid tumor cancer cells have transcriptional activity that mirrors that of pluripotent and multipotent stem cells [23]. These same transcription factors involved in the reprogramming of pluripotent cells have also been shown to be involved in vivo in tissue damage and senescence signaling [24]. Thus, it has been hypothesized that genes involved in early-stage embryogenesis and silenced after might be reactivated as a result of an insult or damage [25]. Targeting CTAs provides an opportunity to help patients based on recognition of this biology.
In determining the optimal patient population for future trials of KK-LC-1 targeted T cells, we see that there is a higher tumor mutation burden in NSCLC expressing high levels of KK-LC-1, as evidenced by higher percentages of tumors with TMB ≥10 in the highest quartile expression of KK-LC-1 and greater median TMB in Q4 versus Q1 (9 vs. 6 in Q1) (Figure 2). Also, a significantly higher percentages of KK-LC-1 expression was observed in pan wild type tumors and those with KRAS mutations compared to actionable oncogenic mutation tumors such as EGFR, BRAF, ALK, NTRK, RET, and ROS1 (Figure 3A, 3B).
Meanwhile, in discordance with previous literature, we find that tumors with adenocarcinoma had significantly higher KK-LC-1 expression levels than squamous cell carcinomas (Supplementary Figure 1) [5]. This may be explained by the propensity for patients with molecular profiling to be of more advanced stage and the use of a more diverse North American patient population than prior surgical series [1, 4, 5, 9].
Tumor associated macrophages (TAMs) are involved in creating an immunosuppressive microenvironment. They affect tumor growth, angiogenesis, immune regulation, metastasis, and chemoresistance. M1 macrophages are part of the classically activated macrophage pathway and are known to secrete proinflammatory cytokines and express markers, notably MHCII, CD68, CD80, iNOS, pSTAT. M2 macrophages meanwhile exert an immunosuppressive phenotype, favoring tissue repair and tumor progression; they secrete anti-inflammatory cytokines such as IL-10, IL-13, and IL-4 along with expressing CD206. TAMs tend to acquire a M2-like phenotype acquired macrophages with macrophage polarization between M1 and M2 macrophages [26]. Previous studies have shown that TAMs in NSCLC have been associated with angiogenesis and lymphangiogenesis with high M2 macrophage ratio and VEGF expression [27].
In evaluating our analysis, tumors with higher KK-LC-1 expression seem to favor a microenvironment enriched for cells representing innate immunity. There was significantly fewer CD4+ cells in Q4 tumors compared to Q1 and while not statistically significant fewer CD8+ and myeloid dendritic cells as well in Q4 tumors. Q4 tumors also trended to have more neutrophils and NK cells. In addition, we saw significantly greater representation of M1 macrophages while a significantly lower fraction of M2 macrophages was observed in highly KK-LC-1 expressing tumors. Previous therapies shown to induce M1 macrophage polarization include agents used to treat lung cancer in paclitaxel and docetaxel [28]. Anti PD-L1 treatment has been shown to remodel the macrophage compartment towards a pro-inflammatory phenotype favoring M1 macrophage expression [29]. We do not have treatment history available on our cohort, further studies looking at pretreated and untreated cohorts may be beneficial in delineating between which patients would be benefit from targeted therapy towards KK-LC-1.
Our findings show that the KK-LC-1 antigen is associated with a higher mutation burden. This suggests that patients with the highest KK-LC-1 expression are likely those with lung adenocarcinoma without targetable mutations aside from KRAS G12C. From our findings, the most likely candidates for a clinical trial using TCR-T therapy directed against KK-LC-1 would be patients who have been treated with checkpoint inhibitors and less likely to have been treated with tyrosine kinase inhibitors.
There are several limitations of our study. First is that the study was performed using de-identified patient samples without the benefit of clinical history. Second, we did not include protein expression measurements which would be important for identification of lung cancer patients most likely to benefit from TCR-T therapy. Finally, although we identified those tumors with higher and lower numbers of transcripts for KK-LC-1, we do not know the threshold of gene expression that would be relevant for targeted T-cells or for any other targeted immunotherapy. Establishing this minimal level will require interventional studies with clinical endpoints. Future directions include investigation of other cancer testis antigens particularly those antigens that are candidates for T-cell based immunotherapy [6, 12].
Materials and Methods
Tumor samples
The study included NSCLC tumor samples submitted to Caris Life Sciences (Phoenix, AZ) for analysis. This study was conducted in accordance with guidelines of the Declaration of Helsinki, Belmont report, and U.S. Common rule. In keeping with 45 CFR 46.101(b)[4], this study was performed utilizing retrospective, deidentified clinical data. Therefore, this study was considered IRB exempt and patient consent was not required.
mRNA expression (WTS)
KK-LC-1 expression data was evaluated using RNA sequencing. Full formalin-fixed paraffin-embedded (FFPE) specimens underwent review by a board-certified pathologist to measure percent tumor content and tumor size; a minimum of 20% of tumor content in the area for microdissection was required to enable enrichment and extraction of tumor-specific RNA. Qiagen RNA FFPE tissue extraction kit was used for extraction, and the RNA quality and quantity were determined using the Agilent TapeStation. Biotinylated RNA baits were hybridized to the synthesized and purified cDNA targets and the bait-target complexes were amplified in a post capture PCR reaction. The Illumina NovaSeq 6500 was used to sequence the whole transcriptome from patients to an average of 60M reads. Raw data was demultiplexed by Illumina Dragen BioIT accelerator, trimmed, counted, PCR-duplicates removed and aligned to human reference genome hg19 by STAR aligner. For transcription counting, transcripts per million molecules was generated using the Salmon expression pipeline. Human All Exon V7 bait panel (Agilent Technologies, Santa Clara, CA). Additionally, immune cell fraction was calculated by QuantiSeq [30] using this transcriptomic data.
Immunohistochemistry (IHC)
Immunohistochemistry (IHC) was performed on FFPE sections of glass slides. Slides were stained using automated staining techniques, per the manufacturer’s instructions, and were optimized and validated per CLIA/CAO and ISO requirements. The primary PD-L1 antibody clone was 22c3 (Dako). Tumor Proportion Score (TPS) was measured, defined as the percentage of viable tumor cells showing partial or complete membrane staining at any intensity. The tumor was considered positive if TPS ≥ 1%.
Next-generation sequencing (NGS)
NGS was performed on genomic DNA isolated from FFPE tumor samples using the NextSeq platform (Illumina, Inc., San Diego, CA). Matched normal tissue was not sequenced. A custom-designed SureSelect XT assay was used to enrich 592 whole-gene targets (Agilent Technologies, Santa Clara, CA). All variants were detected with >99% confidence based on allele frequency and amplicon coverage, with an average sequencing depth of coverage of >500 and an analytic sensitivity of 5%. Prior to molecular testing, tumor enrichment was achieved by harvesting targeted tissue using manual microdissection techniques. Genetic variants were interpreted by molecular geneticists and categorized as “pathogenic,” “presumed pathogenic,” “pathogenic variant”, “variant of unknown significance,” “presumed benign” or “benign” according to the American College of Medical Genetics and Genomics (ACMG) standards. “Pathogenic”, “resumed pathogenic”, and “pathogenic variants” were counted as mutations whereas “benign”, “presumed benign”, and “variants of unknown significance” were excluded. Pan wild type tumors were defined as tumors that did not contain a “pathogenic,” “presumed pathogenic,” or “pathogenic variant” mutation. Specific information on criteria for BRAF and EGFR mutations are provided (Supplementary Tables 7 and 8).
Tumor mutational burden (TMB)
TMB was measured by counting all non-synonymous missense, nonsense, inframe insertion/deletion and frameshift mutations found per tumor that had not been previously described as germline alterations in dbSNP151, Genome Aggregation Database (gnomAD) databases or benign variants identified by Caris geneticists. A cutoff point of ≥10 mutations per MB was used to define high TMB. [31] Caris Life Sciences is a participant in the Friends of Cancer Research TMB Harmonization Project [32].
Microsatellite instability (MSI)
A combination of multiple test platforms was used to determine dMMR/MSI-H status of the tumors profiled, including fragment analysis (FA, Promega, Madison, WI), IHC (MLH1, M1 antibody; MSH2, G2191129 antibody; MSH6, 44 anti-body; and PMS2, EPR3947 antibody [Ventana Medical Systems, Inc., Tucson, AZ, USA]) and NGS (for tumors tested with NextSeq platform, 7,000 target microsatellite loci were examined and compared to the reference genome hg19 from the University of California).
Data and statistical analysis
Prevalence of molecular alterations among KK-LC-1 expression quartiles were analyzed using Chi-square or Fisher Exact tests. KK-LC-1 TPM distribution among molecular subtypes and histological subtypes were analyzed using non-parametric Kruskal-Wallis testing. Similarly, tumor microenvironment cell fractions were analyzed among expression quartiles as described previously. A value of <0.05 was considered a trending difference; p values were further corrected for multiple comparison using Benjamini-Hochberg method to avoid type I error and an adjusted p value (i.e., q value) of <0.05 was considered a significant difference.
Abbreviations
CTAs: Cancer/Testis Antigens; KK-LC-1/CXorf61: Kita-Kyushu Lung Cancer Antigen-1; CT 83: Cancer/Testis Antigen 83; NSCLC: Non-Small Cell Lung Cancer; TCR-T: T-Cell Receptor Therapy; TPM: Transcripts Per Million; EGFR: Epidermal Growth Factor Receptor; ALK: Anaplastic Lymphoma Kinase; KRAS: Kirsten Rat Sarcoma Virus; CTLs: Cytotoxic T Lymphocytes; TIL: Tumor Infiltrating Lymphocytes; MAGE-A4: Melanoma-Associated Antigen A4; NY-ESO-1: New York Esophageal Squamous Cell Carcinoma 1; ROS1: Proto-oncogene tyrosine-protein kinase ROS; BRAF: V-raf murine sarcoma viral oncogene homolog B1; MET: Exon 14 Mesenchymal-to-Epithelial Transition Exon 14; RET: Rearranged During Transfection Proto-oncogene; NTRK: Neurotrophic Tyrosine Receptor Kinase; FFPE: Formalin-Fixed Paraffin-Embedded; IHC: Immunohistochemistry; PD-L1: Programmed Death-Ligand 1; PD-1: Programmed Death-1; TPS: Tumor Proportion Score; NGS: Next-Generation Sequencing; TMB: Tumor Mutational Burden; MSI: Microsatellite Instability; TAMs: Tumor Associated Macrophages; MHC-II: Major Histocompatibility Complex Class II; CD68: Cluster of Differentiation 68; CD80: Cluster of Differentiation 80; iNOS: Inducible Nitric Oxide Synthase; pSTAT: Phosphorylated Signal Transducer and Activator of Transcription; IL-10: Interleukin 10; IL-13: Interleukin 13; IL-4: Interleukin 4; VEGF: Vascular Endothelial Growth Factor; CD4: Cluster of Differentiation 4; CD8: Cluster of Differentiation 8.
CONFLICTS OF INTEREST
Dr. Nieva receives personal fees from Astra Zeneca, Fujirebio and Naveris. He receives research support from Merck and Genentech. Dr. Hsu was a consultant for Targeted Healthcare Communications. Ms. Baca and Dr. Xiu are employees of Caris Life Sciences. Dr. Kim is a consultant or advisory board member for Novartis, Janssen, AstraZeneca, and PierianDx and has received research funding to his institution from AstraZeneca, Bristol-Myers Squibb, Novartis, Genentech, Regeneron, Spectrum, Mirati, Debiopharm, Karyopharm, and Janssen. Dr. Mamdani is a consultant for Zentalis and has been a consultant for AstraZeneca, Caris Life Sciences, and Takeda. Dr. Nagasaka is a consultant for Caris and a speaker for Blueprint. She has also participated in advisory boards for AstraZeneca, Daiichi, Takeda, Novartis, EMD Serono, JNJ, Pfizer, Lilly, Genentech, and has received travel support from AnHeart Therapuetics. Dr. Puri is a consultant/advisor for AstraZeneca and G1 Therapeutics. Dr. Liu is a consultant/serves on advisory board for Amgen, AstraZeneca, Bayer, Beigene, Blueprint, Bristol-Myers Squibb, Daiichi Sankyo, Eisai, Elevation Oncology, Genentech/Roche, Guardant Health, Janssen, Jazz Pharmaceuticals, Lilly, Merck/MSD, Novartis, Regeneron, Takeda, Turning Point Therapeutics. He also has received research funding to his institution from Alkermes, Bayer, Blueprint, Bristol-Myers Squibb, Elevation Oncology, Genentech, Lilly, Merck, Merus, Pfizer, Rain Therapeutics, RAPT, Turning Point Therapeutics. Dr. Korn is an employee of Caris Life Sciences and has stock ownership and stock options from Caris Life Sciences. He also receives consultant fees from Merck. All other authors have no conflicts of interests to disclose.
FUNDING
This work was funded in part by the USC Norris Comprehensive Cancer Center (Core) Grant, P30 CA014089.
References
1. Jin S, Cao S, Li J, Meng Q, Wang C, Yao L, Lang Y, Cao J, Shen J, Pan B, Hu J, Yu Y. Cancer/testis antigens (CTAs) expression in resected lung cancer. Onco Targets Ther. 2018; 11:4491–99. https://doi.org/10.2147/OTT.S159491. [PubMed].
2. Kondo Y, Fukuyama T, Yamamura R, Futawatari N, Ichiki Y, Tanaka Y, Nishi Y, Takahashi Y, Yamazaki H, Kobayashi N, Watanabe M. Detection of KK-LC-1 Protein, a Cancer/Testis Antigen, in Patients with Breast Cancer. Anticancer Res. 2018; 38:5923–28. https://doi.org/10.21873/anticanres.12937. [PubMed].
3. Fukuyama T, Futawatari N, Yamamura R, Yamazaki T, Ichiki Y, Ema A, Ushiku H, Nishi Y, Takahashi Y, Otsuka T, Yamazaki H, Koizumi W, Yasumoto K, Kobayashi N. Expression of KK-LC-1, a cancer/testis antigen, at non-tumour sites of the stomach carrying a tumour. Sci Rep. 2018; 8:6131. https://doi.org/10.1038/s41598-018-24514-9. [PubMed].
4. Baba T, Shiota H, Kuroda K, Shigematsu Y, Ichiki Y, Uramoto H, Hanagiri T, Tanaka F. Cancer/testis antigen expression as a predictor for epidermal growth factor receptor mutation and prognosis in lung adenocarcinoma. Eur J Cardiothorac Surg. 2013; 43:759–64. https://doi.org/10.1093/ejcts/ezs426. [PubMed].
5. Shigematsu Y, Hanagiri T, Shiota H, Kuroda K, Baba T, Mizukami M, So T, Ichiki Y, Yasuda M, So T, Takenoyama M, Yasumoto K. Clinical significance of cancer/testis antigens expression in patients with non-small cell lung cancer. Lung Cancer. 2010; 68:105–10. https://doi.org/10.1016/j.lungcan.2009.05.010. [PubMed].
6. Davari K, Holland T, Prassmayer L, Longinotti G, Ganley KP, Pechilis LJ, Diaconu I, Nambiar PR, Magee MS, Schendel DJ, Sommermeyer D, Ellinger C. Development of a CD8 co-receptor independent T-cell receptor specific for tumor-associated antigen MAGE-A4 for next generation T-cell-based immunotherapy. J Immunother Cancer. 2021; 9:e002035. https://doi.org/10.1136/jitc-2020-002035. [PubMed].
7. Aleksic M, Liddy N, Molloy PE, Pumphrey N, Vuidepot A, Chang KM, Jakobsen BK. Different affinity windows for virus and cancer-specific T-cell receptors: implications for therapeutic strategies. Eur J Immunol. 2012; 42:3174–79. https://doi.org/10.1002/eji.201242606. [PubMed].
8. Palmer E. Negative selection--clearing out the bad apples from the T-cell repertoire. Nat Rev Immunol. 2003; 3:383–91. https://doi.org/10.1038/nri1085. [PubMed].
9. Fukuyama T, Hanagiri T, Takenoyama M, Ichiki Y, Mizukami M, So T, Sugaya M, So T, Sugio K, Yasumoto K. Identification of a new cancer/germline gene, KK-LC-1, encoding an antigen recognized by autologous CTL induced on human lung adenocarcinoma. Cancer Res. 2006; 66:4922–28. https://doi.org/10.1158/0008-5472.CAN-05-3840. [PubMed].
10. Marcinkowski B, Stevanović S, Helman SR, Norberg SM, Serna C, Jin B, Gkitsas N, Kadakia T, Warner A, Davis JL, Rooper L, Hinrichs CS. Cancer targeting by TCR gene-engineered T cells directed against Kita-Kyushu Lung Cancer Antigen-1. J Immunother Cancer. 2019; 7:229. https://doi.org/10.1186/s40425-019-0678-x. [PubMed].
11. Paret C, Simon P, Vormbrock K, Bender C, Kölsch A, Breitkreuz A, Yildiz Ö, Omokoko T, Hubich-Rau S, Hartmann C, Häcker S, Wagner M, Roldan DB, et al. CXorf61 is a target for T cell based immunotherapy of triple-negative breast cancer. Oncotarget. 2015; 6:25356–67. https://doi.org/10.18632/oncotarget.4516. [PubMed].
12. Xia Y, Tian X, Wang J, Qiao D, Liu X, Xiao L, Liang W, Ban D, Chu J, Yu J, Wang R, Tian G, Wang M. Treatment of metastatic non-small cell lung cancer with NY-ESO-1 specific TCR engineered-T cells in a phase I clinical trial: A case report. Oncol Lett. 2018; 16:6998–7007. https://doi.org/10.3892/ol.2018.9534. [PubMed].
13. Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2021. CA Cancer J Clin. 2021; 71:7–33. https://doi.org/10.3322/caac.21654. [PubMed].
14. Peters S, Camidge DR, Shaw AT, Gadgeel S, Ahn JS, Kim DW, Ou SI, Pérol M, Dziadziuszko R, Rosell R, Zeaiter A, Mitry E, Golding S, et al, and ALEX Trial Investigators. Alectinib versus Crizotinib in Untreated ALK-Positive Non-Small-Cell Lung Cancer. N Engl J Med. 2017; 377:829–38. https://doi.org/10.1056/NEJMoa1704795. [PubMed].
15. Ahn MJ, Tsai CM, Shepherd FA, Bazhenova L, Sequist LV, Hida T, Yang JCH, Ramalingam SS, Mitsudomi T, Jänne PA, Mann H, Cantarini M, Goss G. Osimertinib in patients with T790M mutation-positive, advanced non-small cell lung cancer: Long-term follow-up from a pooled analysis of 2 phase 2 studies. Cancer. 2019; 125:892–901. https://doi.org/10.1002/cncr.31891. [PubMed].
16. Hong DS, DuBois SG, Kummar S, Farago AF, Albert CM, Rohrberg KS, van Tilburg CM, Nagasubramanian R, Berlin JD, Federman N, Mascarenhas L, Geoerger B, Dowlati A, et al. Larotrectinib in patients with TRK fusion-positive solid tumours: a pooled analysis of three phase 1/2 clinical trials. Lancet Oncol. 2020; 21:531–40. https://doi.org/10.1016/S1470-2045(19)30856-3. [PubMed].
17. Wolf J, Seto T, Han JY, Reguart N, Garon EB, Groen HJM, Tan DSW, Hida T, de Jonge M, Orlov SV, Smit EF, Souquet PJ, Vansteenkiste J, et al, and GEOMETRY mono-1 Investigators. Capmatinib in MET Exon 14-Mutated or MET-Amplified Non-Small-Cell Lung Cancer. N Engl J Med. 2020; 383:944–57. https://doi.org/10.1056/NEJMoa2002787. [PubMed].
18. Solomon BJ, Kim DW, Wu YL, Nakagawa K, Mekhail T, Felip E, Cappuzzo F, Paolini J, Usari T, Tang Y, Wilner KD, Blackhall F, Mok TS. Final Overall Survival Analysis From a Study Comparing First-Line Crizotinib Versus Chemotherapy in ALK-Mutation-Positive Non-Small-Cell Lung Cancer. J Clin Oncol. 2018; 36:2251–58. https://doi.org/10.1200/JCO.2017.77.4794. [PubMed].
19. Lin JJ, Cardarella S, Lydon CA, Dahlberg SE, Jackman DM, Jänne PA, Johnson BE. Five-Year Survival in EGFR-Mutant Metastatic Lung Adenocarcinoma Treated with EGFR-TKIs. J Thorac Oncol. 2016; 11:556–65. https://doi.org/10.1016/j.jtho.2015.12.103. [PubMed].
20. Shaw AT, Riely GJ, Bang YJ, Kim DW, Camidge DR, Solomon BJ, Varella-Garcia M, Iafrate AJ, Shapiro GI, Usari T, Wang SC, Wilner KD, Clark JW, Ou SI. Crizotinib in ROS1-rearranged advanced non-small-cell lung cancer (NSCLC): updated results, including overall survival, from PROFILE 1001. Ann Oncol. 2019; 30:1121–26. https://doi.org/10.1093/annonc/mdz131. [PubMed].
21. Gadgeel S, Rodríguez-Abreu D, Speranza G, Esteban E, Felip E, Dómine M, Hui R, Hochmair MJ, Clingan P, Powell SF, Cheng SY, Bischoff HG, Peled N, et al. Updated Analysis From KEYNOTE-189: Pembrolizumab or Placebo Plus Pemetrexed and Platinum for Previously Untreated Metastatic Nonsquamous Non-Small-Cell Lung Cancer. J Clin Oncol. 2020; 38:1505–17. https://doi.org/10.1200/JCO.19.03136. [PubMed].
22. Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006; 126:663–76. https://doi.org/10.1016/j.cell.2006.07.024. [PubMed].
23. Palmer NP, Schmid PR, Berger B, Kohane IS. A gene expression profile of stem cell pluripotentiality and differentiation is conserved across diverse solid and hematopoietic cancers. Genome Biol. 2012; 13:R71. https://doi.org/10.1186/gb-2012-13-8-r71. [PubMed].
24. Mosteiro L, Pantoja C, Alcazar N, Marión RM, Chondronasiou D, Rovira M, Fernandez-Marcos PJ, Muñoz-Martin M, Blanco-Aparicio C, Pastor J, Gómez-López G, De Martino A, Blasco MA, et al. Tissue damage and senescence provide critical signals for cellular reprogramming in vivo. Science. 2016; 354:aaf4445. https://doi.org/10.1126/science.aaf4445. [PubMed].
25. Davies P. Exposing cancer’s deep evolutionary roots. Physics World. 2013; 26:37–40. https://doi.org/10.1088/2058-7058/26/07/41.
26. Lin Y, Xu J, Lan H. Tumor-associated macrophages in tumor metastasis: biological roles and clinical therapeutic applications. J Hematol Oncol. 2019; 12:76. https://doi.org/10.1186/s13045-019-0760-3. [PubMed].
27. Hwang I, Kim JW, Ylaya K, Chung EJ, Kitano H, Perry C, Hanaoka J, Fukuoka J, Chung JY, Hewitt SM. Tumor-associated macrophage, angiogenesis and lymphangiogenesis markers predict prognosis of non-small cell lung cancer patients. J Transl Med. 2020; 18:443. https://doi.org/10.1186/s12967-020-02618-z. [PubMed].
28. Malfitano AM, Pisanti S, Napolitano F, Di Somma S, Martinelli R, Portella G. Tumor-Associated Macrophage Status in Cancer Treatment. Cancers (Basel). 2020; 12:1987. https://doi.org/10.3390/cancers12071987. [PubMed].
29. Xiong H, Mittman S, Rodriguez R, Pacheco-Sanchez P, Moskalenko M, Yang Y, Elstrott J, Ritter AT, Müller S, Nickles D, Arenzana TL, Capietto AH, Delamarre L, et al. Coexpression of Inhibitory Receptors Enriches for Activated and Functional CD8+ T Cells in Murine Syngeneic Tumor Models. Cancer Immunol Res. 2019; 7:963–76. https://doi.org/10.1158/2326-6066.CIR-18-0750. [PubMed].
30. Finotello F, Mayer C, Plattner C, Laschober G, Rieder D, Hackl H, Krogsdam A, Loncova Z, Posch W, Wilflingseder D, Sopper S, Ijsselsteijn M, Brouwer TP, et al. Correction to: Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 2019; 11:50. https://doi.org/10.1186/s13073-019-0655-5. [PubMed].
31. Marabelle A, Fakih M, Lopez J, Shah M, Shapira-Frommer R, Nakagawa K, Chung HC, Kindler HL, Lopez-Martin JA, Miller WH Jr, Italiano A, Kao S, Piha-Paul SA, et al. Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. 2020; 21:1353–65. https://doi.org/10.1016/S1470-2045(20)30445-9. [PubMed].
32. Merino DM, McShane LM, Fabrizio D, Funari V, Chen SJ, White JR, Wenz P, Baden J, Barrett JC, Chaudhary R, Chen L, Chen WS, Cheng JH, et al, and TMB Harmonization Consortium. Establishing guidelines to harmonize tumor mutational burden (TMB): in silico assessment of variation in TMB quantification across diagnostic platforms: phase I of the Friends of Cancer Research TMB Harmonization Project. J Immunother Cancer. 2020; 8:e000147. https://doi.org/10.1136/jitc-2019-000147. [PubMed].

