Introduction
Medical oncologists were among the first clinicians to integrate genomics into clinical care. Traditionally, this involved single gene testing or small gene panels in select cancer histotypes with established predictive, prognostic or therapeutic implications [1, 2]. The implementation of comprehensive genomic profiling (CGP) has however been much more complex, with most gene panels comprised of hundreds of genes and applied to populations unselected by cancer type [3]. The rapidity of the evolution of CGP places great responsibility on oncologists to discuss implications of testing, requiring familiarity with ever-expanding genomic and therapeutic knowledgebases [4, 5]. CGP can also reveal secondary germline findings [6–8], with up to 18% of patients having germline variants in cancer susceptibility genes identified [9]. The interpretation and management of germline findings outside a classic familial cancer context creates new challenges for oncologists, patients and their families [10, 11].
Understanding oncologists’ perspectives on their role and needs when communicating results to patients is important to the successful integration of CGP into routine care. Few studies have explored clinicians’ experiences utilising CGP [5, 12–14]. These data demonstrate varied uptake of genomic sequencing, and a lack of confidence amongst oncologists incorporating genomic results into clinical care [3, 15–20]. The uptake of CGP was greatest in high-patient volume and academic centres [15, 20] and amongst clinicians with training in genomics [3]. In an Australian context [3, 15], the Australian Pancreatic Genome Initiative published a practical framework for communicating genomic research results [20], including informed consent, analytical validity, clinical relevance of results and results communication [20]. How this guidance has impacted clinical practice is not clear.
Here we aimed to understand Australian oncologists’ views and experiences in integrating CGP into clinical care, assessing current practice and confidence in performing CGP-related tasks, and potential resource needs.
Results
Survey participants
Between July 2018 and January 2019, a total of 108 oncologists completed the survey (participation rate = 32%) (Table 1). Males predominated (n = 62, 57%, p = 0.15, one-sample proportion test), with most participants having less than 15 years of experience (n = 76, 70%), most were aged under 50 years (n = 83, 77%) and worked within urban-based practices (n = 90, 83%) in New South Wales (n = 68, 63%).
Table 1: Participating oncologists’ demographics
| Characteristic | Frequency | Percentage (%) |
|---|---|---|
| Gender | ||
| Male | 62 | 57.4 |
| Female | 46 | 42.6 |
| Age | ||
| 20–29 years | 1 | 0.9 |
| 30–39 years | 49 | 45.4 |
| 40–49 years | 33 | 30.6 |
| 50–59 years | 11 | 10.2 |
| 60–69 years | 14 | 13.0 |
| Urban or regional | ||
| Urban | 90 | 83.3 |
| Regional | 18 | 16.7 |
| Practicing State | ||
| Australian Capital Territory | 5 | 4.6 |
| New South Wales | 68 | 63.0 |
| Northern Territory | 2 | 1.9 |
| Queensland | 7 | 6.5 |
| South Australia | 9 | 8.3 |
| Tasmania | 1 | 0.9 |
| Victoria | 11 | 10.2 |
| Western Australia | 5 | 4.6 |
| Years worked as an oncologist | ||
| 0–4 years | 32 | 29.6 |
| 5–9 years | 27 | 25.0 |
| 10–14 years | 17 | 15.7 |
| 15–19 years | 7 | 6.5 |
| 20–24 years | 10 | 9.3 |
| 25–29 years | 5 | 4.6 |
| 30–34 years | 8 | 7.4 |
| 35–39 years | 2 | 1.9 |
| >40 years | 0 | 0 |
Experience with genomic testing
Most oncologists (n = 97, 89.6%) had referred patients for CGP, and most had experience referring patients to a Family Cancer Centre (FCC) based on family history (n = 103, 95.4%, Table 2). No significant associations were found between participant characteristics (gender, age, practicing state, or clinical experience) and experience with CGP (multiple regression, p = 0.58, Supplementary Table 1). Approximately two-thirds of oncologists (n = 70, 64.8%) also had experience with patients undergoing direct-to-consumer (DTC) tumor testing (mode: 1–5 patients) through a commercially available sequencing panel. DTC refers to CGP that is instigated and coordinated by the patient and involves them working directly with the service provider.
Table 2: Experience with testing
| Experience referring patients for molecular tumor profiling | Number of patient referrals made | Number (%) of participants |
| Never | 11 (10.2) | |
| 1–5 | 29 (26.9) | |
| 6–10 | 33 (30.6) | |
| 11–20 | 14 (13.0) | |
| >20 | 21 (19.4) | |
| Experience with direct to consumer (DTC) tumor testing | Number of patients having had DTC tumor testing | Number (%) of participants |
| Never | 40 (37.0) | |
| 1–5 | 47 (43.5) | |
| 6–10 | 14 (13.0) | |
| 11–20 | 5 (4.6) | |
| >20 | 2 (1.9) | |
| Experience with DTC germline testing | Any experience with DTC germline testing? | Number (%) of participants |
| No | 96 (88.9) | |
| Yes | 12 (11.1) | |
| Experience with referring patients to a Family Cancer Centre | Number of patient referrals made | Number (%) of participants |
| Never | 5 (4.6) | |
| 1–5 | 10 (9.3) | |
| 6–10 | 12 (11.1) | |
| 11–20 | 18 (16.7) | |
| >20 | 63 (58.3) |
Role of oncologist in ensuring clinical suitability to order tumor profiling
Nearly all oncologists felt it was their responsibility to ensure patients’ clinical suitability to undergo CGP (n = 105, 97%). Most respondents indicated that adequate life expectancy (≥ 12 weeks, n = 78, 74%), organ function (n = 99, 94%), and reasonable Eastern Cooperative Oncology Group (ECOG) performance status (n = 89, 95%) were prerequisites to referring patients. Twenty-three oncologists (21%) indicated they would initiate CGP only to access therapy with reasonable efficacy in the relevant context, in the absence of standard treatment options or where significant comorbidities precluded standard treatment, and where the patient had a full understanding of the potential financial implications.
Obtaining informed consent
Most oncologists (82.4%) believed they had a role in obtaining patient consent for CGP (median VAS 75, IQR: 53–85), but were less comfortable obtaining consent for germline testing (median VAS 44). Oncologists were confident in discussing the process of CGP (median VAS: 75, IQR: 50–86), identifying a ‘therapeutically actionable finding’ (median VAS: 70, IQR: 53–80), and communicating genomic profiling results (median VAS: 70, IQR: 51–80). They were significantly less confident in discussing germline findings (median pairwise difference: –10, IQR: –30–0, p < 0.001, Wilcoxon test with continuity correction).
Pursuit of therapy using CGP results
Most oncologists (60%) would advise against pursuing an unproven targeted therapy based on CGP prior to standard of care treatment, but most (91%) would consider this option in the treatment-refractory setting. Most oncologists (84.3%) still required some evidence to pursue matched therapy outside a trial setting. Relevant evidence included clinical effectiveness in any cancer type (18.7%), where the efficacy was linked to a biomarker (25.3%), especially in the relevant tumor type (56%). Eleven (12.1%) participants reported that case reports/series constituted sufficient evidence. Interestingly, oncologists’ confidence to pursue an unproven targeted therapy increased with years of clinical experience, with every 5 years of clinical experience associated with a mean 4.8 points (95% CI: 2.0–7.5) increase in confidence to pursue an unproven therapy (p = 0.001, Supplementary Table 2). Additional patient, clinician and treatment factors were the absence of standard treatment options (n = 2, 2.2%), the rarity of the cancer and biomarker, and lastly patient preferences (n = 1, 1.1%) (Table 3).
Table 3: Key factors in the use of genomic results to access targeted therapy
| Patient factors | Clinician factors | Treatment factors |
|---|---|---|
| Patient wellness/ECOG | Training/education/experience | Cost |
| Patient preference/interest | Acceptable level of evidence for clinical benefit | Availability/access program/clinical trial |
| Age | Therapeutic Goods Administration approval | Safety/toxicity |
| Financial status | Hospital policy | |
| Rare cancer histology | ||
| Lack of other available therapies | ||
| Informed patient consent |
CGP report
Ninety-eight of 108 participants (90.7%) had experience with CGP reports, and felt it was their role to communicate these results to patients. This includes communication of CGP results initiated by patients (DTC). Most oncologists (n = 103, 95.4%) wanted reports to include genes on the assay and a statement of therapy recommendations (n = 93, 86.1%), and most (80/98, 82%) felt there was sufficient information to understand the results. Areas for suggested improvement included greater explanation and evidence-based ranking of gene prioritisation, clinical significance and treatment recommendations. Some clinicians did not want recommendations to an inaccessible treatment, including international clinical trials, or detailed explanation of the assay. A significant fraction (49, 46%) wanted only those variants that are clinically actionable, while 39 (36.1%) wanted all genetic variants, and 20 (18.5%) only those variants that are therapeutically actionable. Importantly, most participants (n = 99, 91.7%) would value a list of matched trials for Australian patients. Acceptable sources of supportive evidence for treatment recommendations included a molecular tumor board (MTB), published evidence, and genomic database references. Nearly all oncologists (n = 104, 96.3%) wanted the report to include referral recommendations if potential germline variants in cancer predisposition genes were indicated.
Assistance with interpreting CGP results
Most oncologists (n = 68, 62.9%) reported they needed, or wanted assistance with interpreting CGP results. Currently used sources of support included an oncology colleague (80, 74.1%), the local FCC (50, 46.3%), while 33 (30.6%) reported ‘other’, including reporting institution, own research, molecular pathologist or local expert. Most oncologists (n = 77, 71.3%) were significantly less confident in discussing inherited cancer risk; median VAS was 49.5 (IQR: 30.0–63.5). As indicated by very few responses (N = 8, 7.4%) with VAS in the top quintile (≥ 80), only a minority of oncologists were confident in discussing secondary germline findings with patients and their family following tumor profiling.
Resources to assist with communicating results to patients
Participants were asked to identify potential sources of support in communicating the results of CGP to patients and their families (Table 4). The highest priority resources identified included patient information sheets, decision aids, an FCC helpline for patients and clinicians, and a text-based primer of genomics. More experienced oncologists indicated a trend towards lower preference for clinician-based resources (P = 0.04, Supplementary Table 3). p = 0.016). Figure 1 provides a synoptic overview of steps followed by oncologists in the clinical integration of CGP results, synthesizing survey responses.
Table 4: Preferred resources
| Resource | Visual analogue scale Median (IQR) |
|---|---|
| FCC helpline for patients | 69 (50–80) |
| Downloadable patient information sheets on genomic testing and its implications | 80 (70–95) |
| Downloadable patient decision aids | 78 (68–90) |
| FCC helpline for clinicians | 70 (50–80) |
| An online primer on genomics (text) | 65 (50–79) |
| An online primer on genomics (video) | 48 (21–69) |
| Additional suggested resources | |
| Evidence – reference to published literature on genetic variants’ clinical relevance | |
| An app or database of current evidence for therapeutic efficacy per biomarker | |
| Update on variant calling | |
| Case study seminars by an experienced clinician | |
| Structured reports including types of variants and suggested clinical advice | |
| Compendium of relevant clinical and basic science information for clinician | |
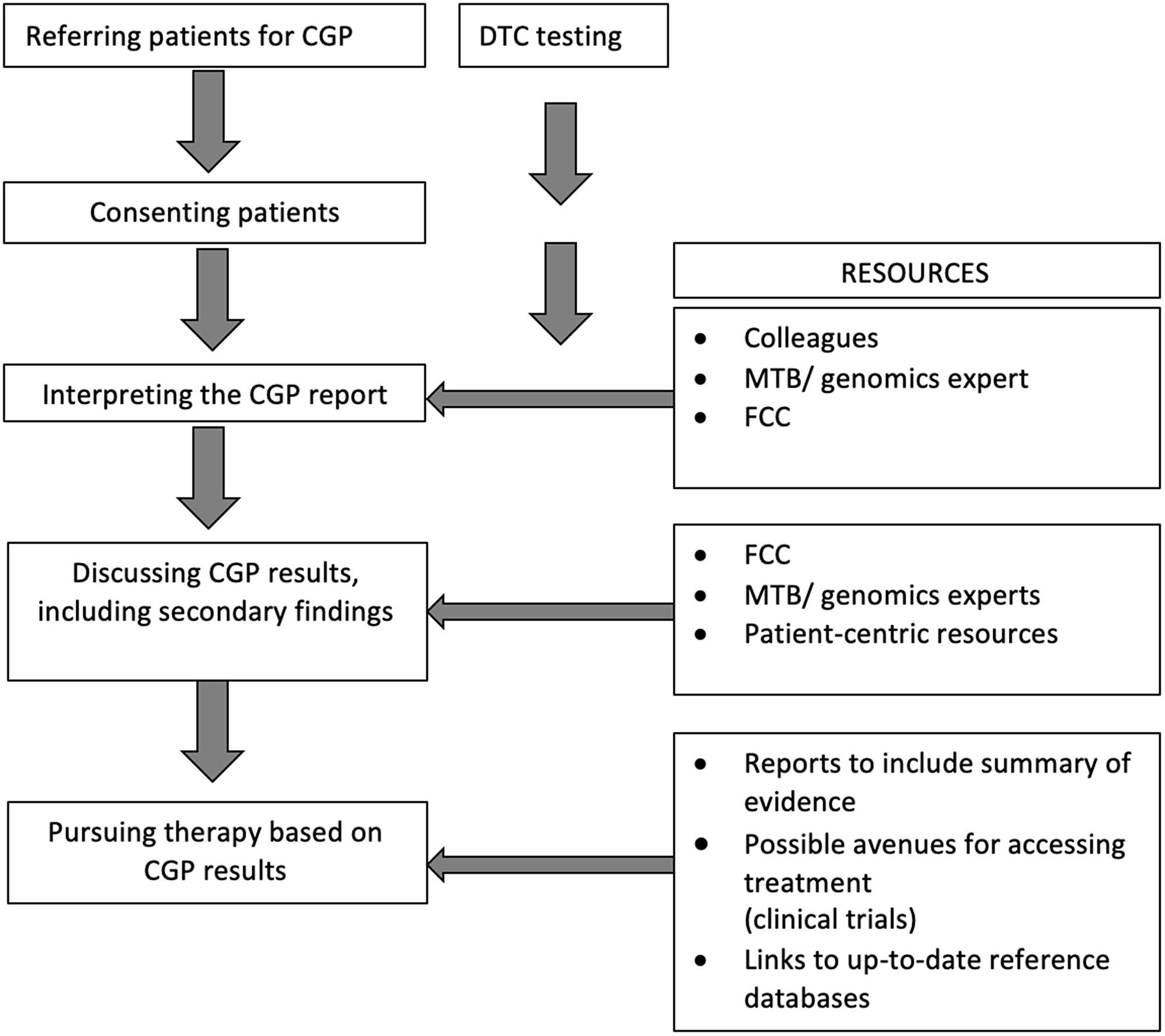
Figure 1: Steps followed by oncologists in the clinical integration of genomic information and resources used or sought. Abbreviations: CGP: comprehensive genomic profiling, DTC: direct to consumer; FCC: Family Cancer Centre; MTB: molecular tumor board.
DISCUSSION
Oncologists clearly identify a role in consenting patients for CGP, and feel comfortable with the process, and the probability of identifying ‘therapeutically actionable findings’ and results. Even when testing was instigated by patients, most oncologists felt it was their role to discuss results, particularly when there were treatment implications. Earlier studies have similarly shown greater use of genomic sequencing in large volume, and academic centres [19]. While our survey did not directly question genomic knowledge, it addressed other components of the genomic confidence scales applied in earlier studies [3, 12], including the ability to discuss CGP results with patients and apply them to treatment recommendations. A study conducted through a single, tertiary, comprehensive cancer care center reported 90% of physicians feeling somewhat/ very confident with their ability to explain genomic concepts to patients and ~75% feeling somewhat/ very confidence with their ability to make treatment recommendations based on genomic information [12]. Comparable levels of variation in ‘genomic confidence’ were seen in our study, with 91% of oncologists believing it was their role to communicate CGP results to patients, but 63% wanting assistance with translating genomic information into treatment recommendations. Our study highlighted the need for greater support for oncologists in interpreting and communicating CGP results effectively. The predominant reason for oncologists rejecting the role to discuss CGP results related to either a lack of expertise to interpret, or uncertainty about the clinical relevance of the finding. Specifically, most participants would like greater guidance as to test validity and the strength of therapeutic recommendations. Clinicians particularly lack confidence in extrapolating treatment decisions based on genomic profiling results outside cancer types in which the evidence for efficacy has been established [21].
Providing a transparent, evidence-based therapy recommendation is key to effectively translating CGP results to the bedside. The confidence of oncologists in applying CGP results to initiate off-label therapies increases with clinical experience, but the majority would pursue investigational agents only in a clinical trial setting. Most oncologists would advise against unproven therapies before standard treatment options had been exhausted. With the rapid evolution of CGP and biomarker-directed therapies, practical guidelines appear to be needed to assist with consistent and effective therapeutic decision-making. Oncologists feel less comfortable dealing with incidental germline findings, despite the high frequency of such results in the context of tumor profiling. This is in keeping with previous studies [22]. Most oncologists confined their role to discussing the potential for secondary findings during consent to CGP. With considerable variation, most participants felt the level of detail on the genomic report was sufficient for interpretation. Many felt the report contained too much detail, wanting only those cancer gene variants with actionability. A list of genes tested during CGP, a statement relating gene variants to recommended therapies, available trials in an Australian context, and a recommendation for referral to the FCC were all valued components of the report. Oncologists expressed a higher preference for patient-centred resources to assist with communicating test results.
Our study has certain limitations. Firstly, this study was performed in the setting of a major national precision oncology trial, the Molecular Screening and Therapeutics (MoST) Program [16]. Despite the breadth of the survey, it is likely to have over-sampled oncologists having referred patients to the Molecular Screening and Therapeutics (MoST) Program for CGP, and therefore not be representative of Australian medical oncologists as a whole. Thus, our findings may over-estimate oncologists’ experience with CGP. If so, the need for support and resources for implementation of CGP in clinical practice is likely even greater. Secondly, in a rapidly evolving field, these findings may change substantially if repeated in 2021. Ongoing surveillance of the oncology landscape is warranted to determine the relevance to clinical experience today.
In conclusion, this study provides insight into how Australian oncologists are coping in the era of precision medicine, and their perceptions of potential barriers to the clinical translation of genomic output. Medical oncologists appear to have accepted CGP as an element of clinical care delivery, and feel responsible for discussing the interpretation and management implications with their patients. Understanding their needs is the foundation for establishing appropriate and effective supports and resources. These considerations will be increasingly important with the rapid evolution and integration of genomic-guided clinical management of cancer patients.
Materials and Methods
Study type
The study employed a cross-sectional survey design. Ethics approval was obtained from St Vincent’s Hospital Human Research Ethics Committee, file number 18/083.
Survey design
The online survey was developed based on discussions within an expert advisory group comprising genomics researchers, molecular pathologists, medical oncologists and psycho-oncologists. The survey comprised 25 items eliciting views/experiences/needs related to utilising CGP results and participant demographic data. A 100-point visual analogue scale score (VAS) was used to assess perceived role and level of confidence in utilising CGP results, with higher values indicating greater confidence. Open-ended questions elicited more detail regarding oncologists’ rationale for views and needs. The survey listed a range of options for participants to indicate their preferred resources to assist with communicating results to patients. The survey was piloted with ten oncologists, and minimal changes were made (Supplementary Data).
Sampling and recruitment
The Medical Oncology Group of Australia membership (n = 333) was sent a single email invitation and weblink in July 2018. Study investigators were blinded to the identity and responses of survey recipients. Between August 2018 and January 2019, individualised follow-up emails were sent to medical oncologists who had referred patients to the Molecular Screening and Therapeutics (MoST) Program [16]. The MoST Program is recruiting adult patients with advanced or metastatic solid cancers. Participants undergo CGP using an evolving range of panels including in-house assays (CCPv2, CCPv2.2, MoSTv1), the Illumina TruSight Tumor 170, and/or the Foundation Medicine panels, with the view to identify actionable findings that can inform novel treatment strategies.
Data analysis
Descriptive statistics were used to summarise cohort characteristics and participants’ responses to survey items. For VAS, medians and IQR are reported given the variables were not normally distributed. Multiple linear regression was used to examine the association between VAS and participant baseline demographics (i.e., practice setting, gender and years of clinical experience). The Type I error rate was set at α = 0.01, accounting for family-wise error from multiple hypothesis testing and modelling. Chi-square testing was employed for survey questions with categorical responses. R environment version 3.5.3 was employed for statistical computing. Open-ended questions were analysed using content analysis [17].
CONFLICTS OF INTEREST
Authors have no conflicts of interest to declare.
FUNDING
We would like to acknowledge study funding provided by the NSW Office of Health and Medical Research, the Australian Department of Health and the NHMRC Project Grant APP1124749 for the Molecular Screening and Therapeutics (MoST) Program. Additionally, several authors were funded by fellowships: M Ballinger - Cancer Institute NSW Career Development Fellowship (CDF171109), PB - NHMRC Senior Principal Research Fellowship (APP1121630); DT – NHMRC Principal Research Fellowship (APP1104364) and M Best - Cancer Institute NSW Post-Doctoral Research Fellowship (2017ECF002).
References
1. Malone ER, Oliva M, Sabatini PJB, Stockley TL, Siu LL. Molecular profiling for precision cancer therapies. Genome Med. 2020; 12:8. https://doi.org/10.1186/s13073-019-0703-1. [PubMed].
2. Agarwal A, Ressler D, Snyder G. The current and future state of companion diagnostics. Pharmgenomics Pers Med. 2015; 8:99–110. https://doi.org/10.2147/PGPM.S49493. [PubMed].
3. de Moor JS, Gray SW, Mitchell SA, Klabunde CN, Freedman AN. Oncologist Confidence in Genomic Testing and Implications for Using Multimarker Tumor Panel Tests in Practice. JCO Precision Oncol. 2020; 4:620–31. https://doi.org/10.1200/po.19.00338. [PubMed].
4. Hughes KS, Ambinder EP, Hess GP, Yu PP, Bernstam EV, Routbort MJ, Clemenceau JR, Hamm JT, Febbo PG, Domchek SM, Chen JL, Warner JL, and OPO Workshop Members. Identifying Health Information Technology Needs of Oncologists to Facilitate the Adoption of Genomic Medicine: Recommendations From the 2016 American Society of Clinical Oncology Omics and Precision Oncology Workshop. J Clin Oncol. 2017; 35:3153–59. https://doi.org/10.1200/JCO.2017.74.1744. [PubMed].
5. Freedman AN, Klabunde CN, Wiant K, Enewold L, Gray SW, Filipski KK, Keating NL, Leonard DGB, Lively T, McNeel TS, Minasian L, Potosky AL, Rivera DR, et al. Use of Next-Generation Sequencing Tests to Guide Cancer Treatment: Results From a Nationally Representative Survey of Oncologists in the United States. JCO Precision Oncol. 2018; 2:1–13. https://doi.org/10.1200/PO.18.00169.
6. Li MM, Datto M, Duncavage EJ, Kulkarni S, Lindeman NI, Roy S, Tsimberidou AM, Vnencak-Jones CL, Wolff DJ, Younes A, Nikiforova MN. Standards and Guidelines for the Interpretation and Reporting of Sequence Variants in Cancer: A Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J Mol Diagn. 2017; 19:4–23. https://doi.org/10.1016/j.jmoldx.2016.10.002. [PubMed].
7. Stjepanovic N, Stockley TL, Bedard PL, McCuaig JM, Aronson M, Holter S, Semotiuk K, Leighl NB, Jang R, Krzyzanowska MK, Oza AM, Gupta A, Elser C, et al. Additional germline findings from a tumor profiling program. BMC Med Genomics. 2018; 11:65. https://doi.org/10.1186/s12920-018-0383-5. [PubMed].
8. Schrader KA, Cheng DT, Joseph V, Prasad M, Walsh M, Zehir A, Ni A, Thomas T, Benayed R, Ashraf A, Lincoln A, Arcila M, Stadler Z, et al. Germline Variants in Targeted Tumor Sequencing Using Matched Normal DNA. JAMA Oncol. 2016; 2:104–11. https://doi.org/10.1001/jamaoncol.2015.5208. [PubMed].
9. Mandelker D, Zhang L, Kemel Y, Stadler ZK, Joseph V, Zehir A, Pradhan N, Arnold A, Walsh MF, Li Y, Balakrishnan AR, Syed A, Prasad M, et al. Mutation Detection in Patients With Advanced Cancer by Universal Sequencing of Cancer-Related Genes in Tumor and Normal DNA vs Guideline-Based Germline Testing. JAMA. 2017; 318:825–35. https://doi.org/10.1001/jama.2017.11137. [PubMed].
10. Douma KF, Meiser B, Kirk J, Mitchell G, Saunders C, Rahman B, Sousa MS, Barlow-Stewart K, Gleeson M, Tucker K. Health professionals' evaluation of delivering treatment-focused genetic testing to women newly diagnosed with breast cancer. Fam Cancer. 2015; 14:265–72. https://doi.org/10.1007/s10689-014-9770-z. [PubMed].
11. Thavaneswaran S, Rath E, Tucker K, Joshua AM, Hess D, Pinese M, Ballinger ML, Thomas DM. Therapeutic implications of germline genetic findings in cancer. Nat Rev Clin Oncol. 2019; 16:386–96. https://doi.org/10.1038/s41571-019-0179-3. [PubMed].
12. Gray SW, Hicks-Courant K, Cronin A, Rollins BJ, Weeks JC. Physicians' attitudes about multiplex tumor genomic testing. J Clin Oncol. 2014; 32:1317–23. https://doi.org/10.1200/JCO.2013.52.4298. [PubMed].
13. Vassy JL, Bates DW, Murray MF. Appropriateness: A Key to Enabling the Use of Genomics in Clinical Practice? Am J Med. 2016; 129:551–53. https://doi.org/10.1016/j.amjmed.2016.02.010. [PubMed].
14. Christensen KD, Vassy JL, Jamal L, Lehmann LS, Slashinski MJ, Perry DL, Robinson JO, Blumenthal-Barby J, Feuerman LZ, Murray MF, Green RC, McGuire AL, and MedSeq Project Team. Are physicians prepared for whole genome sequencing? a qualitative analysis. Clin Genet. 2016; 89:228–34. https://doi.org/10.1111/cge.12626. [PubMed].
15. Chow-White P, Ha D, Laskin J. Knowledge, attitudes, and values among physicians working with clinical genomics: a survey of medical oncologists. Hum Resour Health. 2017; 15:42. https://doi.org/10.1186/s12960-017-0218-z. [PubMed].
16. Thavaneswaran S, Sebastian L, Ballinger M, Best M, Hess D, Lee CK, Sjoquist KM, Hague WE, Butow PN, Simes RJ, Thomas D. Cancer Molecular Screening and Therapeutics (MoST): a framework for multiple, parallel signal-seeking studies of targeted therapies for rare and neglected cancers. Med J Aust. 2018; 209:354–55. https://doi.org/10.5694/mja18.00227. [PubMed].
17. Schreier M, Stamann C, Janssen M, Dahl T, Whittal A. Qualitative Content Analysis: Conceptualizations and Challenges in Research Practice—Introduction to the FQS Special Issue "Qualitative Content Analysis I". Forum Qual Soc Res. 2019; 20. https://doi.org/10.17169/fqs-20.3.3393.
18. Tate JG, Bamford S, Jubb HC, Sondka Z, Beare DM, Bindal N, Boutselakis H, Cole CG, Creatore C, Dawson E, Fish P, Harsha B, Hathaway C, et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res. 2019; 47:D941–47. https://doi.org/10.1093/nar/gky1015. [PubMed].
19. Gray SW, Kim B, Sholl L, Cronin A, Parikh AR, Klabunde CN, Kahn KL, Haggstrom DA, Keating NL. Medical Oncologists' Experiences in Using Genomic Testing for Lung and Colorectal Cancer Care. J Oncol Pract. 2017; 13:e185–96. https://doi.org/10.1200/JOP.2016.016659. [PubMed].
20. Johns AL, Miller DK, Simpson SH, Gill AJ, Kassahn KS, Humphris JL, Samra JS, Tucker K, Andrews L, Chang DK, Waddell N, Pajic M, Pearson JV, et al. Returning individual research results for genome sequences of pancreatic cancer. Genome Med. 2014; 6:42. https://doi.org/10.1186/gm558. [PubMed].
21. Hede K. Genomic testing: a struggle for oncologists. J Natl Cancer Inst. 2014; 106:dju172. https://doi.org/10.1093/jnci/dju172. [PubMed].
22. Gray SW, Park ER, Najita J, Martins Y, Traeger L, Bair E, Gagne J, Garber J, Jänne PA, Lindeman N, Lowenstein C, Oliver N, Sholl L, et al. Oncologists' and cancer patients' views on whole-exome sequencing and incidental findings: results from the CanSeq study. Genet Med. 2016; 18:1011–19. https://doi.org/10.1038/gim.2015.207. [PubMed].

