Introduction
Lung cancer is the leading cause of cancer-related deaths across the globe with more than 1.7 million deaths annually [1]. In India, lung cancer contributes to 8.1% of all cancer-related deaths [1]. Non-small cell lung cancer (NSCLC), more common type of lung cancer, accounts for 85% of all lung cancers comprise of two major histological subtypes, adenocarcinoma and squamous cell carcinoma [2]. The adenocarcinoma of the lung arises mostly in patients with no previous significant tobacco exposure, while the squamous subtype is found almost exclusively in former or current smokers [3] with relatively higher overall mutational load [4]. Despite distinct histological and biological characteristics, the two NSCLC subtypes are largely treated with the same chemotherapeutic agents except for pemetrexed to treat non-squamous NSCLC [5]. Significant advances in the molecularly targeted therapies have been made to treat lung adenocarcinoma patients harboring mutant EGFR, ALK, RET, ROS1, BRAF, MET, NTRK-1 & 2, ERBB2, and FGFR3 [6–8]. However, no approved targeted therapy regimens are available for lung squamous patients in spite of distinct genetic alterations identified in the tumor type, including alterations in TP53, PIK3CA, CDKN2A, MLL2, PTEN, KEAP1, NFE2L2,DDR2, FGFR1, PDGFRA, SOX2, and CCND1 [9–15]. Moreover, most of the reported studies describe Caucasian, Chinese, Korean and Japanese population, with sparse information on the molecular profile of lung squamous patients of Indian origin that accounts for about 30% of Indian lung cancer disease [16].
In this study, we sought to describe the first genetic landscape of alterations underlying 430 Indian lung squamous genomes and uncover the prevalence of known targetable somatic alterations using next generation sequencing followed by validation using mass spectrometry.
Results
Genomic landscape of alterations in lung squamous carcinoma samples
We performed whole exome sequencing of 20 lung squamous tumors – pathologically confirmed to have tumor content above 40%, at an average on-target coverage of 50–70X, followed by mass spectrometry-based genotyping of 430 tumors (Table 1 and Supplementary Table 1) using a customized panel of 53 hotspot mutations across 17 genes (Supplementary Table 2). The sequencing quality, tumor purity and characteristic features of exome analysis for all the samples are detailed in Supplementary Tables 3 and 4. The exome analysis revealed a non-synonymous mean somatic mutation rate of 11.6 mutations/Mb and median of 11 mutations/Mb with enrichment of C>T transitions as opposed to putative C>A transversions indicative of a smoking signature [17, 18] . However, these C>T transitions did not show concordance (cosine similarity > 0.9) with any of the 30 signatures defined in COSMIC. A total of 9261 alterations including 4181 missense mutations, 3658 indels, 837 splice-site mutations and 585 nonsense and nonstop alterations were observed. Consistent with the literature, TP53 was identified to be the most commonly mutated gene (Figure 1) at a frequency of 65%. Similarly, mutations in CDKN2A (20%) and PTEN (10%) were observed with co-occurring TP53 alteration, as reported earlier [9, 12, 15]. Additionally, alterations in tumor suppressor genes including NFE2L2 (20%, 4 cases), KMT2C (15%, 3 cases), LRP1B (15%, 3 cases), FAT1 (15%, 3 cases), NF1 (15%, 3 cases) and PREX2 (5%, 1 case) were observed at comparable frequencies, as reported earlier [9, 11] (Figure 1). Interestingly, alterations in KMT2D were altered at a higher frequency of 40%. The complete list of the somatic substitutions obtained from exome analysis is detailed in Supplementary Table 5.
Table 1: Demographic characteristics of lung squamous patients
| Total number of lung squamous patients | 430 |
|---|---|
| Age | |
| < 65 years | 283 |
| > 65 years | 147 |
| Mean | 59.5 (21–85 years) |
| Gender | |
| Male | 364 (84%) |
| Female | 66 (16%) |
| Habits | |
| Smokers | 309 (72%) |
| Non-smokers | 53 (12%) |
| Not available | 68 |
| Stage | |
| Stage IV | 208 (48%) |
| Stage III | 83 (19%) |
| Stage II | 25 (6%) |
| NA | 114 |
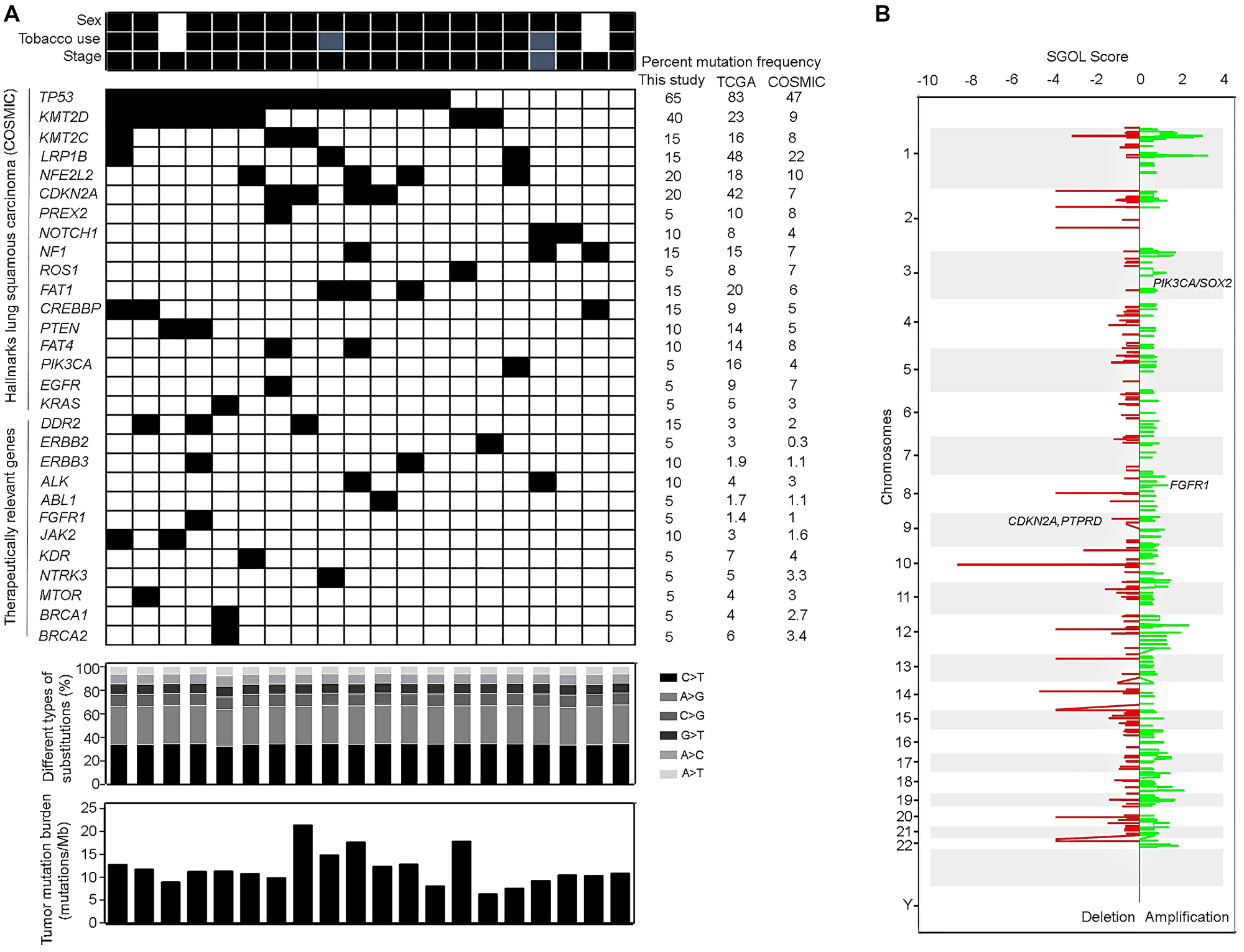
Figure 1: Somatic mutations and copy number variations observed in lung squamous carcinoma patients. (A) The spectrum of mutations obtained after whole exome sequencing analysis of discovery set of 20 Indian origin lung squamous carcinoma patients is represented in the form of heatmap. Black solid box indicates the patient samples positive for the mutation in the specified genes and white box indicates the wildtype status of the particular gene. The clinical characteristics of the patients including the sex, tobacco use and stage are mentioned above the heatmap. Males, tobacco users and stage IV tumors are indicated by black solid boxes while females, non-tobacco users are indicated by white boxes. Grey boxes indicate information not available. Alterations in genes known to be hallmarks of lung squamous carcinoma (based on COSMIC) alongwith therapeutically relevant genes observed in our dataset are depicted. The mutation frequencies of the genes observed in this study (n = 20) are compared with those already reported in COSMIC (n > 1000) and TCGA (n = 587) databases. Additional characteristics of the whole exome data including the distribution of different types of transitions and transversions (according to different shades as specified in the color code) and the tumor mutation burden (mutations/Mb) are shown in the bar graphs below the heatmap. (B) Somatic copy number changes obtained from whole exome sequencing data based on CODEX2 pipeline. The score for segment gain or loss (horizontal axis) is plotted for each chromosome (vertical axis) and represents copy number gain (positive SGOL score, green color) or loss (negative SGOL score, red color). The representative hallmark cancer associated genes are mentioned in the respective amplified/deleted regions.
Inferred copy number analysis based on the exome data identified previously described copy number amplifications harboring genes including FGFR1 (10%), SOX2/ PIK3CA (5%), MYC (10%), CDK6 (10%) and MET (5%) (Figure 1B) [9, 17, 19, 20]. FGFR1 amplification is therapeutically relevant hallmark alteration in lung squamous cancer [20]. Thus, based on the availability and quality of genomic DNA samples, we selected a subset of 100 patients to validate copy number status at FGFR1 amplification in these patients using real-time PCR. We observed FGFR1 amplified in 14% of Indian Lung squamous patients (Supplementary Figure 1).
Similarly, activating kinase domain mutations of EGFR, including the in-frame 15bp deletion in exon 19, EGFR E746_A750 del and point mutation in exon 21, EGFR L858R were observed at a cumulative frequency of 5.8% (Figure 2) across 25/430 tumors (Supplementary Table 6). Interestingly, the activating mutations in the exon 18 of EGFR were not observed in any of the tumors, as reported earlier [9, 14]. In overall, EGFR activating mutations (5.8%) were observed at a significantly higher frequency than the Caucasians (0.2%, p = 0.0005, n = 487) [9] , Korean population (0.1%, p = 0.0405, n = 104) [12] and Chinese populations (3.7%, p = 0.285, n = 271) [11, 21]. The activating KRAS mutations affecting the codon 12, observed at a frequency of 1.1% in our cohort, were mutually exclusive to EGFR mutations (p = 0.37, EGFR = 25, KRAS = 5, both EGFR and KRAS = 0, neither = 400). The canonical PIK3CA mutations, characteristic to lung squamous tumors, were observed at a frequency of 5.5% (E542K: 2.8%, E545K: 2.5% and H1047R: 0.2%, Supplementary Table 6), comparable to as reported among the Caucasian population [9, 14]. Interestingly, of the 49 samples harboring either PIK3CA or EGFR alterations, only one sample showed co-occurring EGFR L858R and PIK3CA H1047R mutation. However, the mutual exclusivity of mutations among these two genes was not found to be significant (p = 0.575, EGFR = 25, PIK3CA = 23, both EGFR and PIK3CA = 1 neither 381). Furthermore, activating mutations at a lower frequency were observed in other genes including FGFR2 (0.5%), AKT1 (0.2%), NRAS (0.2%) and ERBB3 (0.2%) (Supplementary Table 6). Somatic mutations probed in other oncogenes CTNNB1, DDR2, ERBB2 and FGFR3 were not observed in any of the 430 tumor samples.
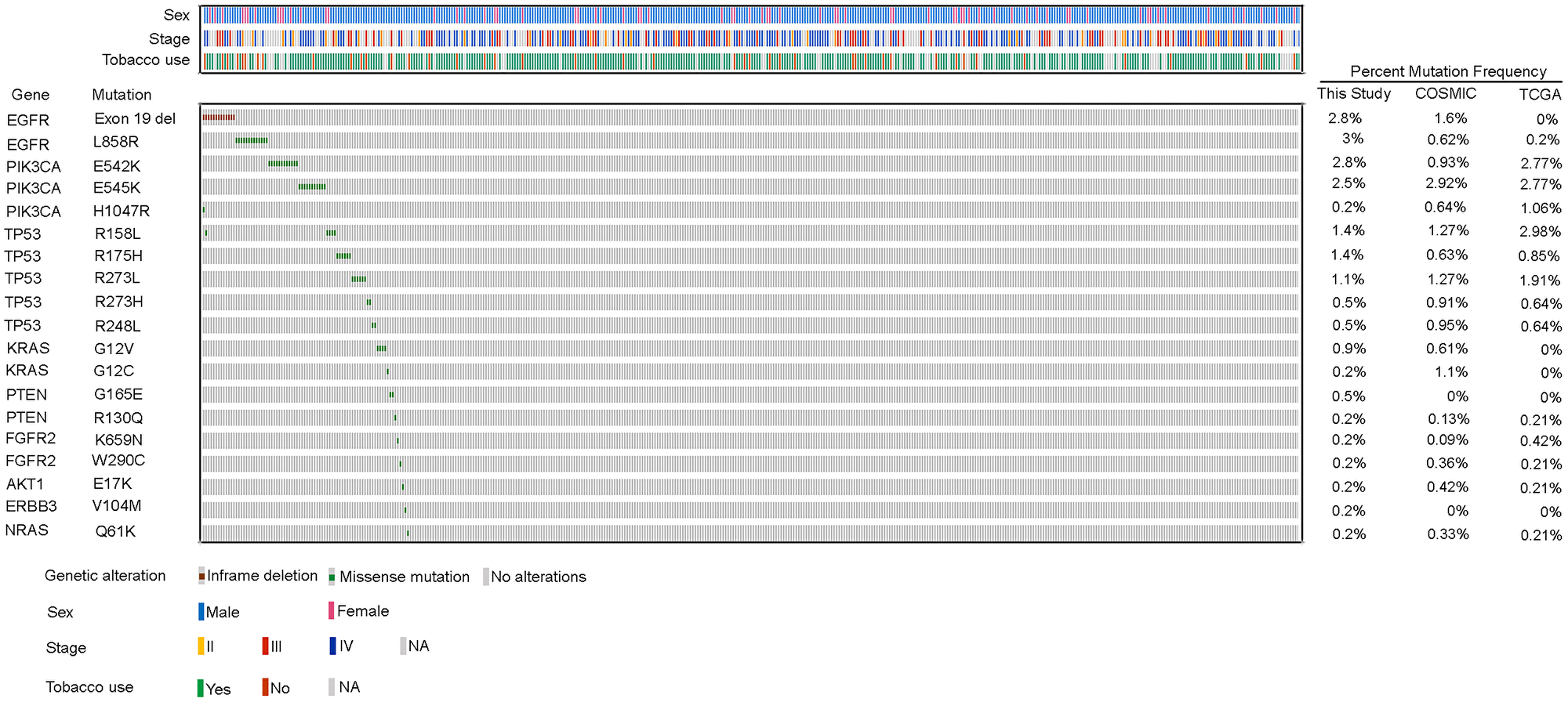

Figure 2: Recurrent genetic alterations in Indian lung squamous carcinoma patients. Heatmap representation of genetic alterations identified by mass spectrometry based genotyping in 430 lung squamous patients. The clinical characteristics of the patients including the sex, tobacco use and stage are mentioned above the heatmap and the respective color codes are mentioned below the heatmap. Only alterations observed at a frequency > 0 are depicted in the heatmap. Alterations which were a part of panel, but not observed in any of the samples are not included. Comparison of frequencies of each alteration between our study and those reported in COSMIC and TCGA databases is shown on the right side of the heatmap.
Clinical correlation with genetic alterations in lung squamous cancer
Our study did not reveal any significant association between clinical features such as age, gender, tumor stage, smoking, tobacco usages with mutations across any of the 17 genes probed in the genotyping experiment, other than fourfold higher EGFR mutations (p = 0.001), and twofold higher mutations (p = 0.02) among non-smokers (Table 2 and Supplementary Table 7), consistent with literature [21]. Next, we analysed whether these mutations are associated with the disease prognosis. Of all the mutations analyzed, the median overall survival of KMT2D mutated patients was found to be significantly lower than among patients with KMT2D wildtype (151.5 days, 284 days, p = 0.032) (Figure 3A), as reported earlier [11, 12]. Similarly, EGFR and PIK3CA mutations were associated with poor prognosis with a median overall survival of 185 and 165 days compared to 219 and 220 days for patients with wildtype, respectively (Figure 3B and 3C), irrespective of their smoking status. Of note, the association of EGFR and PIK3CA mutations and the lower overall survival of lung squamous carcinoma patients however did not attain statistical significance, likely due to lower incidence of the alterations. Additionally, 13.5% of lung squamous patients harboring druggable oncogenic mutations (including KRAS G12C) showed lower median overall survival (165 days) compared to patients harboring other mutations, (221 days) (Figure 4A). And, of most significance, 19% patients harboring a mutation in at least one gene, known to be associated with lung squamous cancer, as inferred by mass-spectrometry based genotyping resulted in significantly shorter median overall survival compared to lung squamous patients with no mutations (167 days vs 225 days, Figure 4B), wherein no patient received any targeted therapy.
Table 2: Corelation of clinicopathological features and gene alterations in Indian lung squamous patients
| Clinico- pathological features | Variable | Number (%), along the column | Gene alterations from mass-array based genotyping, Number (%) along the row | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EGFR | P value | PIK3CA | P value | Oncogenic mutations | P value | Any mutation of panel | P value | ||||||||||
| Mutant | Wildtype | Mutant | Wildtype | Mutant | Wildtype | Mutant | Wildtype | ||||||||||
| Age | < 65 years | 283 | 21 (7.4%) | 262 (92.6%) | 0.051 | 16 (5.6%) | 267 (94.4%) | 1 | 43 (15%) | 240 (85%) | 0.18 | 59 (21%) | 224 (79%) | 0.153 | |||
| > 65 years | 147 | 4 (2.7%) | 143 (97.2%) | 8 (5.4%) | 139 (94.6%) | 15 (10%) | 132 (90%) | 22 (15%) | 125 (85%) | ||||||||
| Gender | Females | 66 | 7 (10.6%) | 59 (89.4%) | 0.084 | 6 (9%) | 60 (91%) | 0.236 | 13 (19.6%) | 53 (80.4%) | 0.118 | 14 (21%) | 52 (79%) | 0.608 | |||
| Males | 364 | 18 (5%) | 346 (95%) | 18 (5%) | 346 (95%) | 45 (12.3%) | 319 (87.7%) | 67 (18.4%) | 297 (81.6%) | ||||||||
| Tumor stage | II | 25 | 1 (4%) | 24 (96%) | 0.828 | 2 (8%) | 23 (92%) | 0.32 | 4 (16%) | 21 (84%) | 0.651 | 6 (24%) | 19 (76%) | 0.373 | |||
| III | 83 | 5 (6%) | 78 (94%) | 2 (2.4%) | 81 (97.6%) | 8 (9.6%) | 75 (90.4%) | 13 (15.6%) | 70 (84.4%) | ||||||||
| IV | 208 | 9 (4.3%) | 199 (95.7%) | 12 (5.7%) | 196 (94.3%) | 23 (11%) | 185 (89%) | 30 (14.4%) | 178 (85.6%) | ||||||||
| Information not available | 114 | 10 | 104 | 8 | 106 | 23 | 91 | 32 | 82 | ||||||||
| Smoking status | Non-smoker | 53 | 8 (15%) | 45 (85%) | 0.002 | 4 (7.5%) | 49 (92.5%) | 0.512 | 12 (22.6%) | 41 (77.4%) | 0.022 | 16 (30%) | 37 (70%) | 0.022 | |||
| Smoker | 309 | 11 (3.5%) | 298 (96.5%) | 16 (5%) | 293 (95%) | 33 (10.6%) | 276 (89.4%) | 51 (16.5%) | 258 (83.5%) | ||||||||
| Information not available | 68 | 6 | 62 | 4 | 64 | 13 | 55 | 14 | 54 | ||||||||
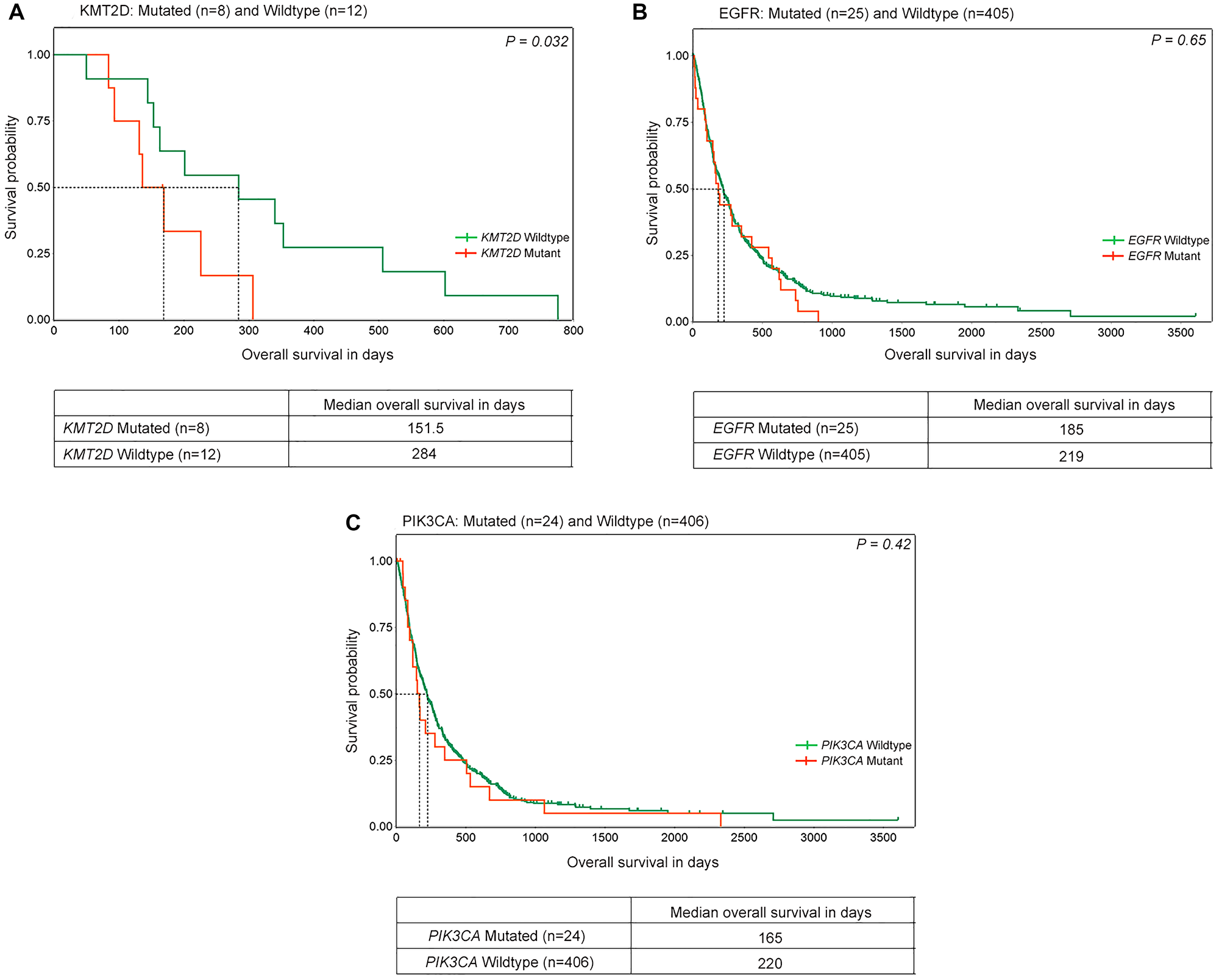
Figure 3: Overall survival in lung squamous tumors with distinct gene alterations. Kaplan-Meier plots of overall survival (in days) in lung squamous tumors with mutations in (A) KMT2D (B) EGFR and (C) PIK3CA are shown. The orange and green lines denote mutated and wild-type patients respectively. The dotted lines indicate median overall survival of the respective groups. P-value is indicated on the top right corner of the plots. The number of patients and median survival in each group is mentioned in the table below.
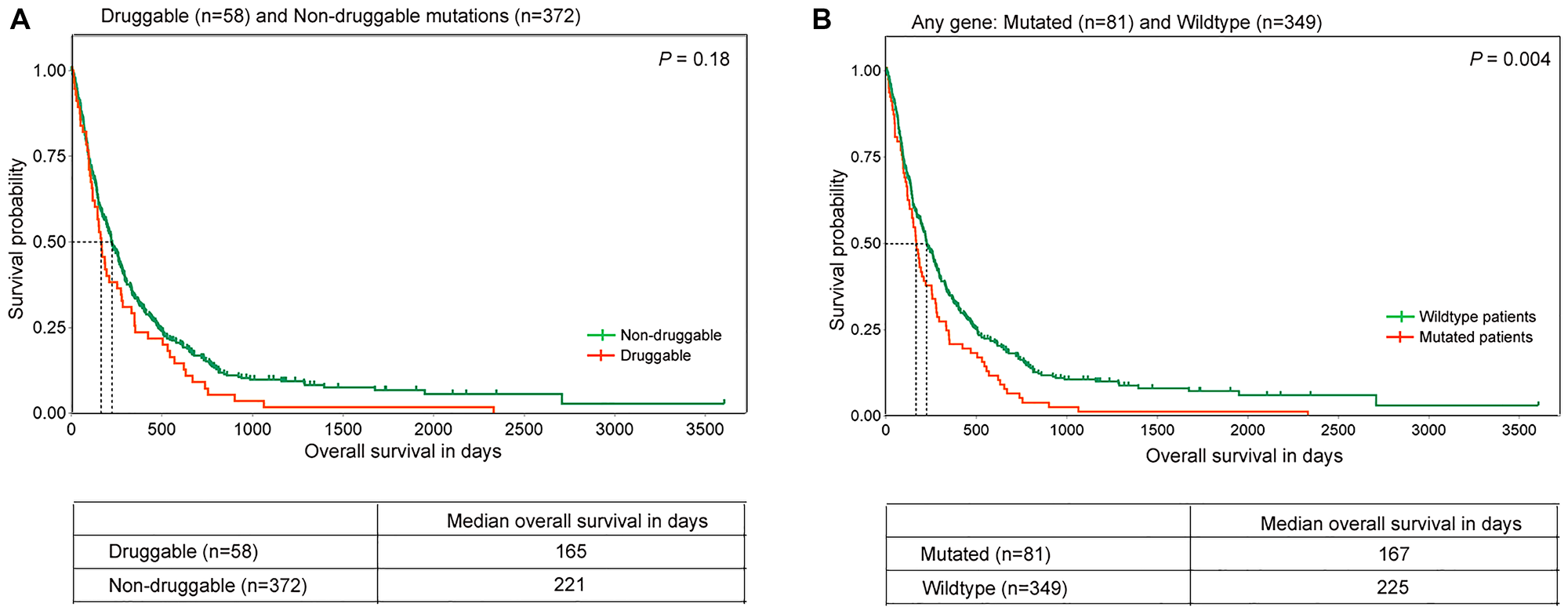
Figure 4: Overall survival in lung squamous tumors with distinct gene alterations. Kaplan-Meier plots of overall survival (in days) in lung squamous tumors with mutations in (A) Druggable genes and (B) any gene are shown. The orange and green lines denote mutated and wild-type patients respectively. The dotted lines indicate median overall survival of the respective groups. P-value is indicated on the top right corner of the plots. The number of patients and median survival in each group is mentioned in the table below.
DISCUSSION
We earlier reported a distinct frequency of 23% EGFR mutations and 18% KRAS mutations in lung adenocarcinoma patients of Indian ethnicity compared to 10–15% vs. 27–62% EGFR mutations and 25–50% vs. 5–15% KRAS mutations among the Caucasians and East Asians populations [6, 22–24]. These studies underscore the somatic mutation variability in lung cancer across populations. Large scale genomic analyses have identified alterations that underlie squamous cell lung cancers [10, 11, 25]. However, most of the lung squamous studies described so far includes mainly the Caucasian, Chinese, Korean and Japanese population, while the molecular profiling of lung squamous patients of Indian origin remains sparsely explored. Here, we describe the first comprehensive landscape of genomic alterations prevalent in Indian lung squamous patients using a combination of next generation sequencing and genotyping technologies. Our study is marked by a deficiency of a lower average on-target coverage of 50–70X of orphan FFPE tumors. However, several lines of distinct features underlie this study attributing to unique etiology and specific population, which has been previously described for early stage tongue tumors among patients of Indian origin [26].
To begin with, we observed a relatively higher non-synonymous mean somatic mutation rate of 11.6 mutations/Mb compared to 8.1 mutations/ Mb among Caucasian population, 8.7 mutations/Mb among Korean population and 7.1 mutations/Mb among Chinese population, which is also considerably higher than the mutation rate observed in various non-tobacco associated cancer types [9, 12]. Interestingly, our data suggests an enrichment of C>T transitions as opposed to putative C>A transversions indicative of a smoking signature [17, 18] consistent with our previous report in tongue squamous tumors [26] and gingiva-buccal tumors as reported by the ICGC-India [27] with tobacco chewing habit. Furthermore, our lung squamous data is largely consistent with the “squamousness” characteristic alterations as described underlying all squamous tumors arising across different anatomical sites [28] — such as TP53, PIK3CA, CDKN2A, and SOX2. However, we observed a significant exception with the absence of alterations at the CCND1 loci among the squamous cell lung carcinoma patients from India compared to 7% frequency among Caucasian patients [29]. Of other significant alterations known to occur in lung squamous, we observed 65% TP53 alterations in our study as reported across different ethnic groups [9–14]. Also, alterations in other tumor suppressor genes, including CDKN2A (20%), NFE2L2 (20%), FAT1 (15%), KMT2C (15%), LRP1B (15%), PTEN (10%) and PREX2 (5%) were comparable to earlier reports [9–12, 14, 17].
Among the therapeutically relevant alterations, frequent oncogenic alterations were found in the PI3K-AKT pathway at a cumulative frequency of 10.7%, as reported in other studies [9, 14]: including 5% amplification at SOX2/PIK3CA; 5.5% PIK3CA mutations; and, AKT1 E17K mutation. Interestingly, the BASALT-1 phase II trial emphasised the prognostic impact of the PI3K pathway in lung squamous cancer, suggesting PIK3CA alterations in lung squamous as a good prognostic marker [30]. Secondly, we found 14% lung squamous cancer patients harboring FGFR1 amplification, comparable to as reported in other population as a promising therapeutic target [19, 20]. While a correlation between the FGFR1 copy number and protein expression remains to be established, this study underlines the relevance of the clinical trials testing the fibroblast growth factor receptor inhibitors in clinical use for the treatment of lung squamous cancer [31, 32]. Despite the dismal survival benefit of the LUME-Lung 1 trial in unselected advanced lung squamous cancer patients trial with combination of FGFR inhibitor with chemotherapeutic agent, the findings of this study along with preclinical studies [33, 34], suggest that pre-selection of ~14% lung squamous patients harboring FGFR1 alterations are more likely to benefit from the treatment; Thirdly, significant alterations in KMT2D were observed at a frequency of 40% compared to 10% and 24%, compared to the Caucasian and East Asian population [12, 17],based on our whole exome data that necessitate validation in a larger cohort of squamous carcinoma samples. And, lastly but most significantly, EGFR-tyrosine kinase inhibitor sensitive alterations were observed at a frequency of 5.8% in our cohort, significantly higher than as reported in the TCGA and studies from the East Asian populations [9–12]. As the LUX-Lung 8 trial underline benefit from the anti-EGFR tyrosine kinase inhibitor among lung squamous patients associated with ERBB alterations, the occurrence of 5.8% EGFR mutations among Indian lung squamous cell carcinoma patients emphasizes the potential significance and relevance of the outcome of this trial in the Indian context [35]. Taken together, 13.5% of lung squamous tumors harbored one or more mutually exclusive therapeutically relevant oncogenic mutations (including KRAS G12C). This is consistent with earlier reports suggesting a 10–11% frequency of potentially targetable alterations in lung squamous carcinoma [36, 37].
Our survival analysis demonstrate that lung squamous patients harboring EGFR or PIK3CA mutations have a shorter median overall survival compared to patients with no mutations; 13.5% patients harboring potentially actionable oncogenic mutations similarly have a lower median overall survival (165 days) compared to patients harboring other mutations (221 days), as reported earlier [36]; and, 19% lung squamous patients harboring a mutation in at least one gene resulted in statistically significant shorter median overall survival compared to lung squamous patients with (167 days vs 225 days). These mutations could thus help inform designing a panel of specific and actionable mutations to select patients likely to benefit from personalized treatment and follow up diagnosis based on liquid biopsy for disease progression and recurrence as shown for lung adenocarcinoma [38].
In overall, we present a striking variation of genetic heterogeneity among lung squamous cell carcinoma patients of Indian descent. The findings from this study extend the scope of the ongoing umbrella clinical trials such as the Lung-MAP master protocol that aims to evaluate multiple targeted therapeutic strategies in lung squamous cell carcinoma patients and the AACR Project GENIE database collaborative project [29, 39]. A systematic exploration of these target genes in lung squamous cell carcinoma patients and variability across ethnicity could further extend our insights into the etiology of lung squamous cancer.
Materials and Methods
Patient cohort
Tumors from 430 lung squamous patients, registered at Tata Memorial Hospital, Mumbai, India during the year 2011–2016 were obtained retrospectively from the tumor tissue repository. All the tumor tissues were stored as formalin fixed paraffin embedded (FFPE) blocks as per the standard protocol of Tata Memorial Hospital, Mumbai, India. The sample cohort and the study protocols were approved by Institutional Review Board and Ethics Committee of Tata Memorial Centre-ACTREC.
Patient details and sample information
The 430 lung squamous patient cohort comprise of 84% males and 16% females with a mean age of diagnosis at 59.5 years (range 21–84 years), 71% cases with a history of tobacco use (including former/current tobacco chewers and smokers) and tumors with stages IV (42%), III (18%) and II (7%). The medical and histopathological records including immunohistochemical staining status of p63, CK7, p40, TTF1 and Napsin A of all the patients were reviewed by oncopathologists to ensure the diagnosis of lung squamous carcinoma. Tumors diagnosed as adenosquamous tumors based on TTF1 expression were excluded from this study. The adequacy of tumor content in all the tissues was assessed by pathologists and was also inferred from whole exome sequencing dataset. All the samples had a minimum of 40% malignant cells. The complete clinical and histopathological characteristics of all the patients including age, sex, tumor stage, smoking status/tobacco usage, nature of biopsy material and immunohistochemical staining status of various markers are detailed in Table 1 and Supplementary Table 1 respectively.
DNA extraction
Genomic DNA extraction from FFPE tumor blocks was performed according to the standard protocol of QIAmp DNA FFPE Tissue Kit. For whole exome sequencing, DNA concentration and quality was assessed using Qubit 3.0 fluorometer and Tape Station respectively and for MassArray genotyping, the DNA concentration was measured using Nanodrop 2000c spectrophotometer (Thermo Fischer Scientific Inc).
Whole exome sequencing
Whole exome sequencing was performed on lung squamous carcinoma DNA samples by Genotypic Technology Pvt Ltd, Bengaluru, India. A minimum of 100ng input DNA (based on Qubit quantification) was utilized for library preparation. Exome capture was performed using the SureSelect Human All Exon V6 (target size 60Mb). Library preparation and exome capture was performed following the manufacturer’s instructions. Exome sequencing was performed on Illumina platform according to standard protocol using 150bp paired end reads to obtain an average coverage of 100X across all the samples.
Exome sequencing analysis for identification of somatic mutations
The raw data was analyzed using the in-house developed pipeline as described earlier [6, 26, 40]. Variants called by the tumor only mode of both GATK and Mutect2 pipelines were combined for further analysis. As described earlier [6, 41], FFPE artefact correction was applied by removing C>T and G>A variants occurring at an allele fraction of < 5%. GATK-Funcotator (https://gatk.broadinstitute.org/hc/en-us/articles/360046786432-Funcotator) was used to annotate cancer associated variants based on which the analysis was restricted only to variants in the coding region. Germline variants were depleted based on databases including dbSNP (dbSNP_151) [42], ExAC (v0.3.1) [43], TMC-SNPdb [44], gnomAD (gnomAD v2.1.1) [45] and Genome Asia 100K while the variants either present in COSMIC (v90) [46] or none of the four databases (the novel variants) were retained. Lastly, the deleterious nature of these variants was determined using functional prediction tool based analysis on somatic non-synonymous variants using seven different tools which are part of dbNSFP v4.0a [47]. Variants called deleterious by at least four out of seven tools were considered for further analysis. The total number of non-synonymous somatic mutations within the coding region were extracted for calculation of tumor mutation burden across each tumor. The percentage of tumor cells in all the samples were computed from the exome sequencing data using the tool AITAC [48] (https://github.com/BDanalysis/aitac). Signature analysis on the exome sequencing data was performed using the R/Bioconductor package MutationalPattens [49] (http://bioconductor.org/packages/MutationalPatterns).
Copy number analysis from exome data
Copy number variations in the whole exome data were assessed using CODEX2, a normalization based CNV detection method which works with or without matched normal samples [50]. We employed fractional mode of CODEX2 for segmentation of our data and categorized an event to be a high gain (copy number > 3.3), gain (copy number 2.3–3.3), diploid (copy number 1.7–2.3), one copy-deletion (copy number 0.7–1.7) and homozygous deletion (copy number < 0.7).
Mass spectrometry-based genotyping
A custom panel of 53 frequently occurring hallmark mutations across 17 cancer associated genes were designed. 200ng input DNA at a final concentration of 20ng/ul from 430 Lung squamous samples was submitted to Imperial Life Sciences Pvt Ltd, New Delhi, India for validation by Mass spectrometry-based genotyping using the iPlex Pro chemistry. Using the assay design software, these 53 mutations were pooled into four wells. Accordingly, the PCR amplification and single base pair extension primers for iPLEX reaction were designed as per manufacturer’s instructions. The mutation calls were analyzed using the Typer 4.0 software and the spectra were also manually revived.
FGFR1 amplification using quantitative real-time PCR
Real-time primers designed to specifically amplify FGFR1 and GAPDH from genomic DNA were used for real time PCR. The specificity of primers was tested by using melt cure analysis during real time PCR. The real time PCR was performed using 20 ng of genomic DNA per 6ul of reaction volume on Light cycler 480 (Roche, Mannheim, Germany). The FGFR1 amplification for the tumor samples was calculated by normalizing the Ct values to the Ct values housekeeping control GAPDH and samples with normalized Ct < 1 were considered to harbor FGFR1 amplification.
Availability of data
The FastQ files containing the raw sequence data for all the samples have been uploaded on ArrayExpress (http://www.ebi.ac.uk/arrayexpress/) hosted by the European Bioinformatics Institute under the accession number E-MTAB-8801 titled ‘Whole exome sequencing of Lung Squamous Carcinoma Patients of Indian Origin’.
Statistical analysis
For mutation mutual exclusivity analysis, the mutant and wildtype status were defined based on the genotyping experiment and the statistical significance of the exclusivity was computed using CoMET method [51]. For correlation of clinico-pathological features and gene alterations and comparisons of mutational frequencies between different ethnic groups, Fisher exact test was used to calculate significance. In both the analysis, P value < 0.05 was considered significant. The Kaplan-Meier survival data analysis and clinicopathological correlation analysis was performed in R as described previously [6, 40] using R survival packages (http://cran.r-project.org/package=survival). The end point was taken as date of death wherever available, else the date of last contact was used for censoring. Tumors with mutations in genes including EGFR, PIK3CA, KRAS, FGFR2, AKT1, NRAS and ERBB3 were grouped as therapeutically relevant alterations.
Ethics approval and consent to participate
The sample cohort and the study protocols were approved by Institutional Review Board and Ethics Committee of Tata Memorial Centre-ACTREC.
ACKNOWLEDGMENTS
All the members of Dutt laboratory for critically reviewing the manuscript. Ms. Trupti Togar for help in collating the samples. Gentotypic Technologies Pvt Ltd and Imperial Life Sciences Pvt Ltd for exome sequencing and genotyping services. A.J., S.H. and S.D are supported by senior research fellowship at ACTREC. The funders had no role in study design, data collection and analysis, decision to publish or preparation of manuscript.
CONFLICTS OF INTEREST
Authors have no conflicts of interest to declare.
FUNDING
The project was funded by an extramural grant from SERB-DST (EMR/2016/007218) to A.D.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492. [PubMed].
2. Herbst RS, Morgensztern D, Boshoff C. The biology and management of non-small cell lung cancer. Nature. 2018; 553:446–54. https://doi.org/10.1038/nature25183. [PubMed].
3. Okamoto T, Takada K, Sato S, Toyokawa G, Tagawa T, Shoji F, Nakanishi R, Oki E, Koike T, Nagahashi M, Ichikawa H, Shimada Y, Watanabe S, et al. Clinical and Genetic Implications of Mutation Burden in Squamous Cell Carcinoma of the Lung. Ann Surg Oncol. 2018; 25:1564–71. https://doi.org/10.1245/s10434-018-6401-1. [PubMed].
4. Gandara DR, Hammerman PS, Sos ML, Lara PN Jr, Hirsch FR. Squamous cell lung cancer: from tumor genomics to cancer therapeutics. Clin Cancer Res. 2015; 21:2236–43. https://doi.org/10.1158/1078-0432.CCR-14-3039. [PubMed].
5. Selvaggi G, Scagliotti GV. Histologic subtype in NSCLC: does it matter? Oncology. 2009; 23:1133–40. [PubMed].
6. Chandrani P, Prabhash K, Prasad R, Sethunath V, Ranjan M, Iyer P, Aich J, Dhamne H, Iyer DN, Upadhyay P, Mohanty B, Chandna P, Kumar R, et al. Drug-sensitive FGFR3 mutations in lung adenocarcinoma. Ann Oncol. 2017; 28:597–603. https://doi.org/10.1093/annonc/mdw636. [PubMed].
7. Tsao AS, Scagliotti GV, Bunn PA Jr, Carbone DP, Warren GW, Bai C, de Koning HJ, Yousaf-Khan AU, McWilliams A, Tsao MS, Adusumilli PS, Rami-Porta R, Asamura H, et al. Scientific Advances in Lung Cancer 2015. J Thorac Oncol. 2016; 11:613–38. https://doi.org/10.1016/j.jtho.2016.03.012. [PubMed].
8. Kris MG, Johnson BE, Berry LD, Kwiatkowski DJ, Iafrate AJ, Wistuba II, Varella-Garcia M, Franklin WA, Aronson SL, Su PF, Shyr Y, Camidge DR, Sequist LV, et al. Using multiplexed assays of oncogenic drivers in lung cancers to select targeted drugs. JAMA. 2014; 311:1998–2006. https://doi.org/10.1001/jama.2014.3741. [PubMed].
9. Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature. 2012; 489:519–25. https://doi.org/10.1038/nature11404. [PubMed].
10. Ding Y, Zhang L, Guo L, Wu C, Zhou J, Zhou Y, Ma J, Li X, Ji P, Wang M, Zhu W, Shi C, Li S, et al. Comparative study on the mutational profile of adenocarcinoma and squamous cell carcinoma predominant histologic subtypes in Chinese non-small cell lung cancer patients. Thorac Cancer. 2020; 11:103–12. https://doi.org/10.1111/1759-7714.13208. [PubMed].
11. Zhang XC, Wang J, Shao GG, Wang Q, Qu X, Wang B, Moy C, Fan Y, Albertyn Z, Huang X, Zhang J, Qiu Y, Platero S, et al. Comprehensive genomic and immunological characterization of Chinese non-small cell lung cancer patients. Nat Commun. 2019; 10:1772. https://doi.org/10.1038/s41467-019-09762-1. [PubMed].
12. Kim Y, Hammerman PS, Kim J, Yoon JA, Lee Y, Sun JM, Wilkerson MD, Pedamallu CS, Cibulskis K, Yoo YK, Lawrence MS, Stojanov P, Carter SL, et al. Integrative and comparative genomic analysis of lung squamous cell carcinomas in East Asian patients. J Clin Oncol. 2014; 32:121–8. https://doi.org/10.1200/JCO.2013.50.8556. [PubMed].
13. Kenmotsu H, Serizawa M, Koh Y, Isaka M, Takahashi T, Taira T, Ono A, Maniwa T, Takahashi S, Mori K, Endo M, Abe M, Hayashi I, et al. Prospective genetic profiling of squamous cell lung cancer and adenosquamous carcinoma in Japanese patients by multitarget assays. BMC Cancer. 2014; 14:786. https://doi.org/10.1186/1471-2407-14-786. [PubMed].
14. Campbell JD, Alexandrov A, Kim J, Wala J, Berger AH, Pedamallu CS, Shukla SA, Guo G, Brooks AN, Murray BA, Imielinski M, Hu X, Ling S, et al. Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat Genet. 2016; 48:607–16. https://doi.org/10.1038/ng.3564. [PubMed].
15. Stewart PA, Welsh EA, Slebos RJC, Fang B, Izumi V, Chambers M, Zhang G, Cen L, Pettersson F, Zhang Y, Chen Z, Cheng CH, Thapa R, et al. Proteogenomic landscape of squamous cell lung cancer. Nat Commun. 2019; 10:3578. https://doi.org/10.1038/s41467-019-11452-x. [PubMed].
16. Mohan A, Garg A, Gupta A, Sahu S, Choudhari C, Vashistha V, Ansari A, Pandey R, Bhalla AS, Madan K, Hadda V, Iyer H, Jain D, et al. Clinical profile of lung cancer in North India: A 10-year analysis of 1862 patients from a tertiary care center. Lung India. 2020; 37:190–7. https://doi.org/10.4103/lungindia.lungindia_333_19.
17. Choi M, Kadara H, Zhang J, Parra ER, Rodriguez-Canales J, Gaffney SG, Zhao Z, Behrens C, Fujimoto J, Chow C, Kim K, Kalhor N, Moran C, et al. Mutation profiles in early-stage lung squamous cell carcinoma with clinical follow-up and correlation with markers of immune function. Ann Oncol. 2017; 28:83–9. https://doi.org/10.1093/annonc/mdw437. [PubMed].
18. Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, Carter SL, Stewart C, Mermel CH, Roberts SA, Kiezun A, Hammerman PS, McKenna A, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013; 499:214–8. https://doi.org/10.1038/nature12213. [PubMed].
19. Ramos AH, Dutt A, Mermel C, Perner S, Cho J, Lafargue CJ, Johnson LA, Stiedl AC, Tanaka KE, Bass AJ, Barretina J, Weir BA, Beroukhim R, et al. Amplification of chromosomal segment 4q12 in non-small cell lung cancer. Cancer Biol Ther. 2009; 8:2042–50. https://doi.org/10.4161/cbt.8.21.9764. [PubMed].
20. Weiss J, Sos ML, Seidel D, Peifer M, Zander T, Heuckmann JM, Ullrich RT, Menon R, Maier S, Soltermann A, Moch H, Wagener P, Fischer F, et al. Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Sci Transl Med. 2010; 2:62ra93. https://doi.org/10.1126/scitranslmed.3001451. [PubMed].
21. Tao D, Han X, Zhang N, Lin D, Wu D, Zhu X, Song W, Shi Y. Genetic alteration profiling of patients with resected squamous cell lung carcinomas. Oncotarget. 2016; 7:36590–601. https://doi.org/10.18632/oncotarget.9096. [PubMed].
22. Choughule A, Sharma R, Trivedi V, Thavamani A, Noronha V, Joshi A, Desai S, Chandrani P, Sundaram P, Utture S, Jambhekar N, Gupta S, Aich J, et al. Coexistence of KRAS mutation with mutant but not wild-type EGFR predicts response to tyrosine-kinase inhibitors in human lung cancer. Br J Cancer. 2014; 111:2203–4. https://doi.org/10.1038/bjc.2014.401. [PubMed].
23. Chougule A, Prabhash K, Noronha V, Joshi A, Thavamani A, Chandrani P, Upadhyay P, Utture S, Desai S, Jambhekar N, Dutt A. Frequency of EGFR mutations in 907 lung adenocarcioma patients of Indian ethnicity. PLoS One. 2013; 8:e76164. https://doi.org/10.1371/journal.pone.0076164. [PubMed].
24. Noronha V, Prabhash K, Thavamani A, Chougule A, Purandare N, Joshi A, Sharma R, Desai S, Jambekar N, Dutt A, Mulherkar R. EGFR mutations in Indian lung cancer patients: clinical correlation and outcome to EGFR targeted therapy. PLoS One. 2013; 8:e61561. https://doi.org/10.1371/journal.pone.0061561. [PubMed].
25. Friedlaender A, Banna G, Malapelle U, Pisapia P, Addeo A. Next Generation Sequencing and Genetic Alterations in Squamous Cell Lung Carcinoma: Where Are We Today? Front Oncol. 2019; 9:166. https://doi.org/10.3389/fonc.2019.00166. [PubMed].
26. Upadhyay P, Gardi N, Desai S, Chandrani P, Joshi A, Dharavath B, Arora P, Bal M, Nair S, Dutt A. Genomic characterization of tobacco/nut chewing HPV-negative early stage tongue tumors identify MMP10 asa candidate to predict metastases. Oral Oncol. 2017; 73:56–64. https://doi.org/10.1016/j.oraloncology.2017.08.003. [PubMed].
27. India Project Team of the International Cancer Genome Consortium. Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups. Nat Commun. 2013; 4:2873. https://doi.org/10.1038/ncomms3873. [PubMed].
28. Schwaederle M, Elkin SK, Tomson BN, Carter JL, Kurzrock R. Squamousness: Next-generation sequencing reveals shared molecular features across squamous tumor types. Cell Cycle. 2015; 14:2355–61. https://doi.org/10.1080/15384101.2015.1053669. [PubMed].
29. AACR Project GENIE Consortium. AACR Project GENIE: Powering Precision Medicine through an International Consortium. Cancer Discov. 2017; 7:818–31. https://doi.org/10.1158/2159-8290.CD-17-0151. [PubMed].
30. Vansteenkiste JF, Canon JL, De Braud F, Grossi F, De Pas T, Gray JE, Su WC, Felip E, Yoshioka H, Gridelli C, Dy GK, Thongprasert S, Reck M, et al. Safety and Efficacy of Buparlisib (BKM120) in Patients with PI3K Pathway-Activated Non-Small Cell Lung Cancer: Results from the Phase II BASALT-1 Study. J Thorac Oncol. 2015; 10:1319–27. https://doi.org/10.1097/JTO.0000000000000607. [PubMed].
31. Tiseo M, Gelsomino F, Alfieri R, Cavazzoni A, Bozzetti C, De Giorgi AM, Petronini PG, Ardizzoni A. FGFR as potential target in the treatment of squamous non small cell lung cancer. Cancer Treat Rev. 2015; 41:527–39. https://doi.org/10.1016/j.ctrv.2015.04.011.
32. Paik PK, Shen R, Berger MF, Ferry D, Soria JC, Mathewson A, Rooney C, Smith NR, Cullberg M, Kilgour E, Landers D, Frewer P, Brooks N, et al. A Phase Ib Open-Label Multicenter Study of AZD4547 in Patients with Advanced Squamous Cell Lung Cancers. Clin Cancer Res. 2017; 23:5366–73. https://doi.org/10.1158/1078-0432.CCR-17-0645. [PubMed].
33. Reck M, Kaiser R, Mellemgaard A, Douillard JY, Orlov S, Krzakowski M, von Pawel J, Gottfried M, Bondarenko I, Liao M, Gann CN, Barrueco J, Gaschler-Markefski B, et al. Docetaxel plus nintedanib versus docetaxel plus placebo in patients with previously treated non-small-cell lung cancer (LUME-Lung 1): a phase 3, double-blind, randomised controlled trial. Lancet Oncol. 2014; 15:143–55. https://doi.org/10.1016/S1470-2045(13)70586-2.
34. Dutt A, Ramos AH, Hammerman PS, Mermel C, Cho J, Sharifnia T, Chande A, Tanaka KE, Stransky N, Greulich H, Gray NS, Meyerson M. Inhibitor-sensitive FGFR1 amplification in human non-small cell lung cancer. PLoS One. 2011; 6:e20351. https://doi.org/10.1371/journal.pone.0020351. [PubMed].
35. Goss GD, Felip E, Cobo M, Lu S, Syrigos K, Lee KH, Goker E, Georgoulias V, Li W, Guclu S, Isla D, Min YJ, Morabito A, et al. Association of ERBB Mutations With Clinical Outcomes of Afatinib- or Erlotinib-Treated Patients With Lung Squamous Cell Carcinoma: Secondary Analysis of the LUX-Lung 8 Randomized Clinical Trial. JAMA Oncol. 2018; 4:1189–97. https://doi.org/10.1001/jamaoncol.2018.0775. [PubMed].
36. Lindquist KE, Karlsson A, Leveen P, Brunnstrom H, Reutersward C, Holm K, Jonsson M, Annersten K, Rosengren F, Jirstrom K, Kosieradzki J, Ek L, Borg A, et al. Clinical framework for next generation sequencing based analysis of treatment predictive mutations and multiplexed gene fusion detection in non-small cell lung cancer. Oncotarget. 2017; 8:34796–810. https://doi.org/10.18632/oncotarget.16276. [PubMed].
37. Lam VK, Tran HT, Banks KC, Lanman RB, Rinsurongkawong W, Peled N, Lewis J, Lee JJ, Roth J, Roarty EB, Swisher S, Talasaz A, Futreal PA, et al. Targeted Tissue and Cell-Free Tumor DNA Sequencing of Advanced Lung Squamous-Cell Carcinoma Reveals Clinically Significant Prevalence of Actionable Alterations. Clin Lung Cancer. 2019; 20:30–6.e3. https://doi.org/10.1016/j.cllc.2018.08.020. [PubMed].
38. Balaji SA, Shanmugam A, Chougule A, Sridharan S, Prabhash K, Arya A, Chaubey A, Hariharan A, Kolekar P, Sen M, Ravichandran A, Katragadda S, Sankaran S, et al. Analysis of solid tumor mutation profiles in liquid biopsy. Cancer Med. 2018; 7:5439–47. https://doi.org/10.1002/cam4.1791. [PubMed].
39. Herbst RS, Gandara DR, Hirsch FR, Redman MW, LeBlanc M, Mack PC, Schwartz LH, Vokes E, Ramalingam SS, Bradley JD, Sparks D, Zhou Y, Miwa C, et al. Lung Master Protocol (Lung-MAP)-A Biomarker-Driven Protocol for Accelerating Development of Therapies for Squamous Cell Lung Cancer: SWOG S1400. Clin Cancer Res. 2015; 21:1514–24. https://doi.org/10.1158/1078-0432.CCR-13-3473.
40. Iyer P, Shrikhande SV, Ranjan M, Joshi A, Gardi N, Prasad R, Dharavath B, Thorat R, Salunkhe S, Sahoo B, Chandrani P, Kore H, Mohanty B, et al. ERBB2 and KRAS alterations mediate response to EGFR inhibitors in early stage gallbladder cancer. Int J Cancer. 2019; 144:2008–19. https://doi.org/10.1002/ijc.31916. [PubMed].
41. Bhagwate AV, Liu Y, Winham SJ, McDonough SJ, Stallings-Mann ML, Heinzen EP, Davila JI, Vierkant RA, Hoskin TL, Frost M, Carter JM, Radisky DC, Cunningham JM, et al. Bioinformatics and DNA-extraction strategies to reliably detect genetic variants from FFPE breast tissue samples. BMC Genomics. 2019; 20:689. https://doi.org/10.1186/s12864-019-6056-8. [PubMed].
42. Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, Sirotkin K. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 2001; 29:308–11. https://doi.org/10.1093/nar/29.1.308. [PubMed].
43. Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, O’Donnell-Luria AH, Ware JS, Hill AJ, Cummings BB, Tukiainen T, Birnbaum DP, Kosmicki JA, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016; 536:285–91. https://doi.org/10.1038/nature19057. [PubMed].
44. Upadhyay P, Gardi N, Desai S, Sahoo B, Singh A, Togar T, Iyer P, Prasad R, Chandrani P, Gupta S, Dutt A. TMC-SNPdb: an Indian germline variant database derived from whole exome sequences. Database (Oxford). 2016; 2016:baw104. https://doi.org/10.1093/database/baw104. [PubMed].
45. Wang Q, Pierce-Hoffman E, Cummings BB, Alföldi J, Francioli LC, Gauthier LD, Hill AJ, O’Donnell-Luria AH, Karczewski KJ, MacArthur DG, and Genome Aggregation Database Production Team, and Genome Aggregation Database Consortium. Landscape of multi-nucleotide variants in 125,748 human exomes and 15,708 genomes. Nat Commun. 2020; 11:2539. https://doi.org/10.1038/s41467-019-12438-5. [PubMed].
46. Tate JG, Bamford S, Jubb HC, Sondka Z, Beare DM, Bindal N, Boutselakis H, Cole CG, Creatore C, Dawson E, Fish P, Harsha B, Hathaway C, et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res. 2019; 47:D941–D7. https://doi.org/10.1093/nar/gky1015. [PubMed].
47. Liu X, Wu C, Li C, Boerwinkle E. dbNSFP v3.0: A One-Stop Database of Functional Predictions and Annotations for Human Nonsynonymous and Splice-Site SNVs. Hum Mutat. 2016; 37:235–41. https://doi.org/10.1002/humu.22932. [PubMed].
48. Yuan X, Li Z, Zhao H, Bai J, Zhang J. Accurate Inference of Tumor Purity and Absolute Copy Numbers From High-Throughput Sequencing Data. Front Genet. 2020; 11:458. https://doi.org/10.3389/fgene.2020.00458. [PubMed].
49. Blokzijl F, Janssen R, van Boxtel R, Cuppen E. MutationalPatterns: comprehensive genome-wide analysis of mutational processes. Genome Med. 2018; 10:33. https://doi.org/10.1186/s13073-018-0539-0. [PubMed].
50. Jiang Y, Wang R, Urrutia E, Anastopoulos IN, Nathanson KL, Zhang NR. CODEX2: full-spectrum copy number variation detection by high-throughput DNA sequencing. Genome Biol. 2018; 19:202. https://doi.org/10.1186/s13059-018-1578-y.
51. Leiserson MD, Wu HT, Vandin F, Raphael BJ. CoMEt: a statistical approach to identify combinations of mutually exclusive alterations in cancer. Genome Biol. 2015; 16:160. https://doi.org/10.1186/s13059-015-0700-7. [PubMed].

