Introduction
Pancreatic cancer is a high-risk malignant neoplasm with a 5-year survival rate of less than 10%. The number of new cases and deaths of pancreatic cancer worldwide in 2018 was 458,918 and 432,242, respectively. In 2020, the projected number of incidences and deaths of pancreatic cancer patients in the United States is 57,600 and 47,050, respectively. Pancreatic ductal adenocarcinoma (PDAC) is the most common type of pancreatic cancer, accounting for more than 90% of all pancreatic cancer diagnoses [1, 2]. The poor prognosis of PDAC is due to several challenges, especially the lack of manifestation screening biomarkers, leading to most pancreatic cancer diagnoses being made in the later, more aggressive stages.
New biomarkers and therapeutic targets of PDAC are urgently needed. Currently, carbohydrate antigen (CA) 19-9 is the sole clinically approved serum biomarker for pancreatic cancer [3]. CA-19-9 is only used for disease monitoring due to a lack of specificity and sensitivity for the early diagnosis of asymptomatic cases. PDAC tends to have strong stem cell-like properties, causing pancreatic cancer to be relatively resistant to traditional chemotherapeutic strategies [4]. Mechanistically, mutations in the KRAS gene has been well described in PDAC, but this advanced knowledge has not resulted in therapeutic innovations due to multiple factors [1, 2]. KRAS protein is relatively small and lacks deep binding pockets, making it difficult to target with inhibitory drugs. KRAS is an intracellular protein and cannot be targeted with immunotherapy options such as antibodies or CAR-T cells. Because of these limitations, KRAS has been viewed as a difficult protein to develop targeted therapies [5, 6]. PDAC is presented with a highly immunosuppressive tumor microenvironment, leading to challenges in immunotherapy treatments such as the administration of checkpoint inhibitors [7–9]. Thus, there is a critical need for novel biomarkers relevant to therapeutic outcomes and novel therapeutic targets in PDAC.
Recently, the human lymphocyte antigen-6 (LY6) gene family has received increasing attention for its multi-faceted roles in cancer development, stem cell maintenance, immunomodulation, and association with more aggressive and hard-to-treat cancers [2, 10, 11]. The LY6 gene family is located on chromosomes 6, 8, 11, and 19. The LY6 family members in human chromosome 8 include PSCA, LY6K, SLURP1, LYPD2, SLURP2, LY6D, GML, LY6E, LY6L, LY6H, and GPIHBP1, mapped at 8q24.3 locus [10]. Somatic amplification of 8q has been suggested to be one of the most prevalent copy number gains in cancer [12–14]. LY6D, LY6E, LY6H, and LY6K have increased mRNA expression in tumor tissues of ovarian, colorectal, gastric, breast, lung, bladder, brain, cervical, esophageal, head and neck, and pancreatic cancer compared to adjacent normal tissues. The increased mRNA expression of LY6D, LY6E, LY6H and LY6K is associated with poor outcome in ovarian, colorectal, gastric, breast, lung, bladder or brain and CNS [2]. Increased levels of LY6A/E (Sca-1) has been reported to promote breast tumorigenesis via disruption of TGF-β signaling [15]. However, whether LY6 family members can be a biomarker for PDAC diagnosis, prognosis, and therapy remains elusive.
In this report, we analyzed the mRNA expression profile of 30 LY6 genes located on chromosomes 6, 8, 11 and 19 using the public The Cancer Genome Atlas (TCGA) dataset of 177 PDAC patients and their association with overall survival outcome. We used publicly available online tools Kaplan-Meier Plotter (KM Plotter), cBioPortal, Oncomine and survMisc R package [16–18]. Herein, we demonstrate differential associations of LY6 family members with patient survival outcome to discuss the potential influence of our identified proteins on the tumor microenvironment, immunomodulation, and stem cell maintenance. Although the function of many LY6 family members is yet to be established, several of our identified proteins have known roles in tumorigenesis and may be suitable candidate biomarkers for the diagnosis, prognosis, and treatment of PDAC.
Results
High expression of LY6 mRNA was associated with overall survival outcome in pancreatic ductal carcinoma
We used Kaplan-Meier (KM) plotter web tool (https://kmplot.com) to see if LY6 gene expression was significantly associated with overall survival (OS) in pancreatic ductal adenocarcinoma (PDAC) [18]. We found 21 out of 30 queried genes were significantly associated with OS outcome in PDAC. We found that high mRNA expression of 17 LY6 genes - PSCA, SLURP1, LYPD2, LY6D, GML, Ly6E and LY6L on chromosome 8; LYPD4, PLAUR, LYPD5 on chromosome 19; PATE1, PATE2, PATE3 and CD59 on chromosome 11and LY6G6C, LY6G6D and LY6G6F on chromosome 6 to be significantly associated with poor OS outcome. The hazard ratios for these genes ranged from 1.68 to 2.99 indicating that patients with high mRNA expression of genes had approximately 1.6 to 3 times the risk of death compared to patients with low mRNA expression. We found high mRNA expression of 4 LY6 genes - LY6H on chromosome 8; PINLYP on chromosome 19; and LY6G5C and LY6G5B on chromosome 6 were significantly associated with good OS outcome (Supplementary Table 1 and Supplementary Figure 1).
We explored the association between RNA-seq data from Pancreatic Adenocarcinoma from TCGA with disease free survival (DFS) outcome using R-programming. We found that 16 out of the 30 LY6 genes were significantly associated with DFS outcome in PDAC. We found that high mRNA expression of 12 LY6 genes - PSCA, SLURP1, LYPD2, LY6D, GML and LY6E on chromosome 8; LYPD3, PLAUR, LYPD5 on chromosome 19; CD59 on chromosome 11 and LY6G6C on chromosome 6 to be significantly associated with poor DFS outcome. We found that high mRNA expression of 4 LY6 genes - LY6H and GPIHBP1 on chromosome 8; and LY6G5C and LY6G5B on chromosome 6 were significantly associated with good OS outcome (Figure 1).
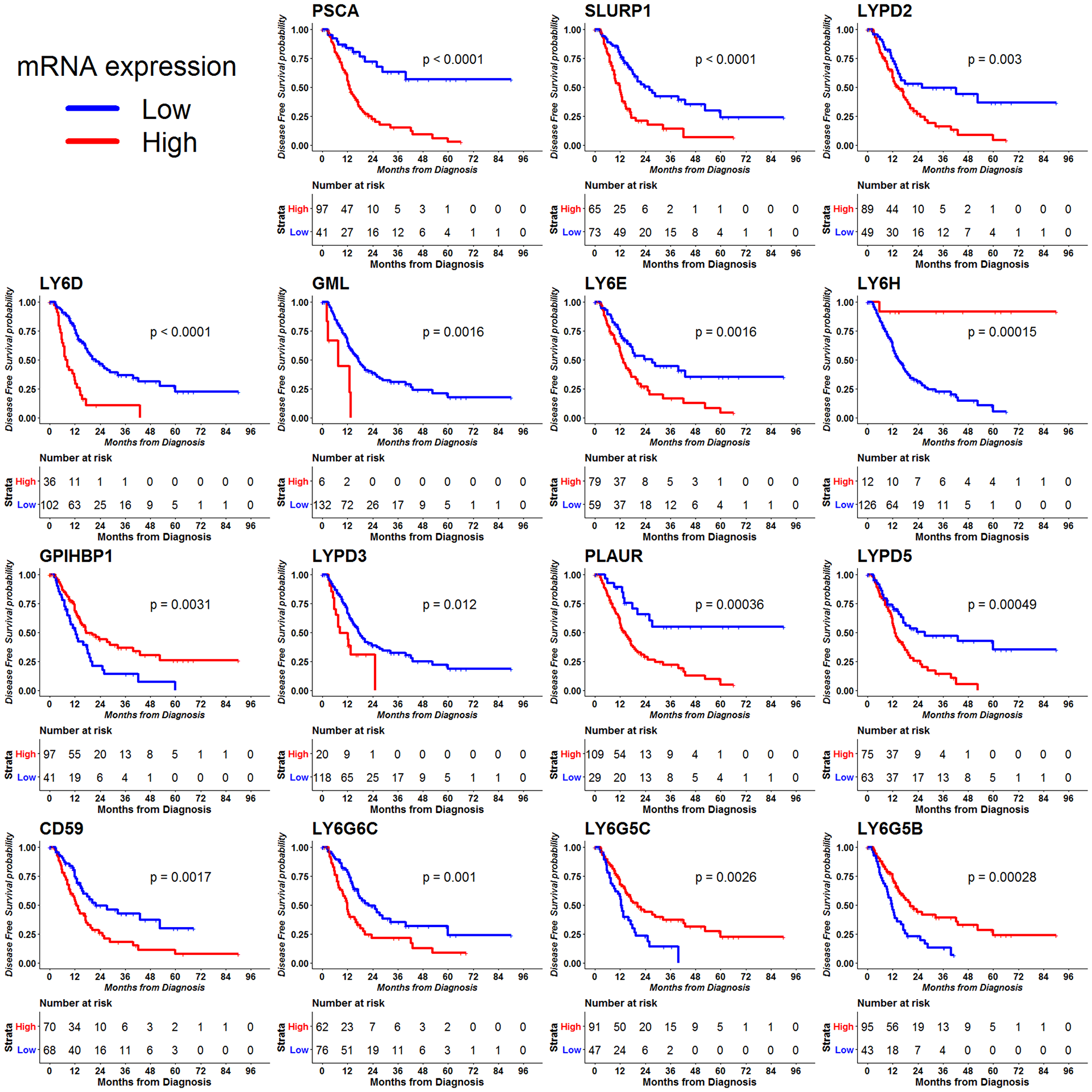
Figure 1: Disease free survival (DFS) outcome for 30 LY6 genes was analyzed using the RNA-seq data using the TCGA data. The clinical and expression data were accessed through the cgdsr package, R programming. The optimal cutoff for mRNA expression was determined using the method implemented in the survMisc R package. High mRNA expression of 15 genes associated with DFS outcome of PDAC. Note: KM plot for GML gene expression has only 6 patients in high expression, however this was included based on p value < 0.05.
We found that 13 LY6 genes were commonly associated with both OS and DFS outcome. We found high expression of 10 LY6 genes - PSCA, SLURP1, LYPD2, LY6D, GML, LY6E on chromosome 8; PLAUR and LYPD5 on chromosome 19; CD59 on chromosome 11 and LY6G6C on chromosome 6 were commonly associated with poor OS and DFS outcomes. We found high expression of 3 LY6 genes LY6H on chromosome 8; LY6G5C and LY6G5B on chromosome 6 were commonly associated with good OS and DFS outcomes (Supplementary Table 1 and Figure 1).
High expression of LY6 mRNA expression and their association with overall survival outcome was dependent on cell type content of the tumors
Tumor microenvironment is composed of various cell types. The enrichment of specific cellular contents in a tumor microenvironment may play an important role as to how the tumor will progress or respond to therapeutic interventions. To estimate if inherent cellular content plays a role in the association of LY6 gene expression with the OS outcome, we used the restricted analysis feature of Kaplan-Meier Plotter tool. This feature allowed us to observe the OS outcome in patient samples with enriched or decreased cellular content of PDAC. Mesenchymal stem cells (MSCs) in tumor microenvironment have shown to be responsible for increased tumor metastasis and immune tolerance of tumors [19–21]. We studied how the LY6 genes affected OS outcome in PDAC patients based on the MSCs status in the tumor samples. We found high mRNA expression of LY6D, SLURP1, CD59, PSCA, PATE2, LY6G6F, LYPD5, LY6E, PATE1, LYPD2, LY6G6D, PATE3, LYPD4, and GML was significantly associated with poor OS outcome only in the MSC enriched patient population. High expression of LY6G6C and GML was associated with poor OS outcome in the MSC enriched population. High expression of LY6G6C and GML was associated with good OS outcome in the MSC decreased population. LY6G5C was associated with good OS outcome only in the MSC enriched population. Ly6G5B was associated with good OS outcome independent of MSC status. Ly6H, PINLYP was associated with good OS outcome only in MSC enriched population. TEX101 was associated with good OS outcome only in MSC decreased population (Table 1 and Supplementary Figure 2).
Table 1: High expression of LY6 genes and its association with overall survival outcome (OS) in TCGA dataset of PDAC patient population based on mesenchymal stem cells (MSC) status, as observed by using KM plotter tool
| Gene | MSC enriched population | MSC decreased population | ||||
|---|---|---|---|---|---|---|
| OS | p value | FDR | OS | p value | FDR | |
| LY6D | Poor | 0.00002 | 1% | NS | 0.11 | 100% |
| SLURP1 | Poor | 0.00002 | 1% | NS | 0.18 | 100% |
| PSCA | Poor | 0.00005 | 1% | NS | 0.33 | 100% |
| LY6G5C | Good | 0.00005 | 2% | NS | 0.16 | 100% |
| CD59 | Poor | 0.00007 | 2% | NS | 0.22 | 100% |
| LY6G6F | Poor | 0.00037 | 10% | NS | 0.39 | 100% |
| PATE1 | Poor | 0.00061 | 20% | NA | NA | NA |
| PATE2 | Poor | 0.00130 | 50% | NS | 0.21 | 100% |
| LY6G6D | Poor | 0.00160 | 10% | NS | 0.17 | 100% |
| LY6E | Poor | 0.00290 | 50% | NS | 0.26 | 100% |
| LY6G5B | Good | 0.00300 | 50% | Good | 0.041 | 50% |
| LY6G6C | Poor | 0.00410 | 20% | Good | 0.046 | 50% |
| LYPD5 | Poor | 0.00480 | 50% | NS | 0.13 | 100% |
| LYPD2 | Poor | 0.00520 | 50% | NS | 0.14 | 100% |
| PATE3 | Poor | 0.00640 | 50% | NS | 0.59 | 100% |
| LY6H | Good | 0.00670 | 50% | NS | 0.091 | 100% |
| PINLYP | Good | 0.00680 | 50% | NS | 0.077 | 100% |
| LYPD4 | Poor | 0.02100 | 50% | NS | 0.15 | 100% |
| LY6L | Poor | 0.02200 | 50% | Good | 0.036 | 50% |
| GML | Poor | 0.03700 | 50% | NS | 0.33 | 100% |
| TEX101 | NS | 0.12000 | 100% | Good | 0.033 | 50% |
Regulatory T cells (Treg cells) play important roles in suppressing immune responses in the tumor microenvironment. They can inhibit cytotoxic T-cell lymphocytes (CTL) from attacking the cancer cells or infiltrating the tumor microenvironment. We observed that high expression of LY6D, SLURP1, PSCA, LY6G6C, LY6E, LYPD5, LY6G6D, PATE2, and PATE1 were significantly associated with poor OS outcome independent of the Treg status. High expression of LY6G5C and LY6G5B were associated good OS outcome independent of Treg status. CD59, LY6G6F, and PATE3 were associated with poor OS outcome only in Treg decreased population. LYPD2 and PLAUR were only associated with poor OS outcome only in Treg enriched population. (Table 2 and Supplementary Figure 3).
Table 2: High expression of LY6 genes and its association with overall survival outcome (OS) in TCGA dataset of PDAC patient population based on T-regulatory cells (Treg) status, as observed by using KM plotter tool
| Gene | Treg enriched population | Treg decreased population | ||||
|---|---|---|---|---|---|---|
| OS | p value | FDR | OS | p value | FDR | |
| LY6D | Poor | 0.00002 | 1% | Poor | 0.00220 | 20% |
| SLURP1 | Poor | 0.00007 | 1% | Poor | 0.00360 | >50% |
| PSCA | Poor | 0.00030 | 2% | Poor | 0.01760 | >50% |
| LY6G6C | Poor | 0.00210 | 20% | Poor | 0.00860 | >50% |
| LYPD2 | Poor | 0.00220 | 20% | NS | 0.09210 | 100% |
| LY6G5C | Good | 0.00230 | 50% | Good | 0.00030 | 5% |
| LY6G5B | Good | 0.00290 | 10% | Good | 0.00790 | >50% |
| PLAUR | Poor | 0.00390 | 50% | NS | 0.20220 | 100% |
| LY6E | Poor | 0.00810 | 50% | Poor | 0.02210 | >50% |
| LYPD5 | Poor | 0.01340 | >50% | Poor | 0.00770 | 50% |
| LY6G6D | Poor | 0.02650 | 100% | Poor | 0.04400 | >50% |
| PATE2 | Poor | 0.02800 | >50% | Poor | 0.00200 | 50% |
| PATE1 | Poor | 0.03690 | >50% | Poor | 0.01290 | >50% |
| LY6G6F | NS | 0.05030 | 100% | Poor | 0.00540 | >50% |
| CD59 | NS | 0.07340 | 100% | Poor | 0.00004 | 1% |
| PATE3 | NS | 0.16470 | 100% | Poor | 0.02020 | >50% |
CD8 positive T-cells are major defense against cancer. Tumors with infiltrated CD8 positive T-cells are termed as inflamed or hot tumors [22]. High expression of PSCA, LY6D, SLURP1, PATE2, LYPD5, PATE1, and LY6G6F was associated with poor OS outcome independent of CD8 positive T-cells status. High expression of LY6G6C and LY6G5B was associated with good OS outcome independent of CD8 positive T-cells status. High expression of PLAUR, LY6E, PATE3, LYPD2, and CD177 was associated with poor OS outcome only in CD8 enriched population. High expression of LY6H was associated with good OS outcome only in CD8 enriched population. High expression of LY6G6C, LY6G6D, CD59, and LYPD4 was associated with poor OS outcome only in CD8 decreased population. High expression of LYPD3 was associated with poor OS outcome in CD8 enriched and with good OS outcome in CD8 decreased population (Table 3 and Supplementary Figure 4).
Table 3: High expression of LY6 genes and its association with overall survival outcome (OS) in TCGA dataset of PDAC patient population based on CD8 positive (CD8+) status, as observed by using KM plotter tool
| Gene | CD8+ enriched population | CD8+ decreased population | ||||
|---|---|---|---|---|---|---|
| OS | p value | FDR | OS | p value | FDR | |
| LY6D | Poor | 0.000074 | 1% | Poor | 0.0016 | 20% |
| SLURP1 | Poor | 0.0012 | 20% | Poor | 0.0017 | 50% |
| PATE2 | Poor | 0.0013 | 20% | Poor | 0.011 | 50% |
| LYPD5 | Poor | 0.0016 | 10% | Poor | 0.035 | 50% |
| LY6G5C | Good | 0.0022 | 10% | Good | 0.0026 | 20% |
| PSCA | Poor | 0.005 | 50% | Poor | 0.0034 | 50% |
| LYPD3 | Poor | 0.0092 | 50% | Good | 0.009 | 50% |
| PATE1 | Poor | 0.0095 | 50% | Poor | 0.022 | 50% |
| PLAUR | Poor | 0.01 | 50% | NS | 0.21 | 100% |
| LY6G5B | Good | 0.011 | 50% | Good | 0.0049 | 50% |
| LY6E | Poor | 0.015 | 50% | NS | 0.082 | 100% |
| LY6H | Good | 0.019 | 50% | NS | 0.078 | 100% |
| LYPD2 | Poor | 0.02 | 50% | NS | 0.097 | 100% |
| CD177 | Poor | 0.023 | 50% | NS | 0.27 | 100% |
| LY6G6F | Poor | 0.038 | 50% | Poor | 0.0064 | 50% |
| LY6L | NS | 0.049 | 50% | NS | 0.13 | 100% |
| LY6G6C | NS | 0.057 | 100% | Poor | 0.015 | 50% |
| LY6G6D | NS | 0.07 | 100% | Poor | 0.014 | 50% |
| CD59 | NS | 0.1 | 100% | Poor | 0.0014 | 20% |
| LYPD4 | NS | 0.21 | 100% | Poor | 0.044 | 50% |
Macrophages play an important role in pancreatic beta cell function, pancreatic tissue homeostasis and pancreatic cancer [23, 24]. High expression of LY6D, CD59, and SLURP1 was associated with poor OS outcome and LY6G5B and LY6G5C was associated with good OS outcome independent of macrophage status. High expression of PATE2 was associated with poor OS outcome in macrophage enriched population. SPACA4, LYPD4, LYPD3, and LY6L was associated with good OS outcome in macrophages enriched population and with poor OS outcome in macrophages decreased population. High expression of LY6H and LY6K was associated with good OS outcome only in macrophages decreased population. High expression of PSCA, TEX101, CD177, LY6G6D, PATE1, LY6E, LY6G6F, LYPD2, PATE3, LYPD5, GML and LY6G6C was associated with poor OS outcome in macrophages decreased population (Table 4 and Supplementary Figure 5).
Table 4: High expression of LY6 genes and its association with overall survival outcome (OS) in TCGA dataset of PDAC patient population based on macrophages status, as observed by using KM plotter tool
| Gene | Macrophages enriched population | Macrophages decreased population | ||||
|---|---|---|---|---|---|---|
| OS | p value | FDR | OS | p value | FDR | |
| LY6D | Poor | 0.00017 | 2% | Poor | 0.00001 | 1% |
| LY6G5B | Good | 0.00090 | 10%% | Good | 0.04400 | 50% |
| CD59 | Poor | 0.00890 | 50% | Poor | 0.00030 | 5% |
| PATE2 | Poor | 0.00970 | 50% | NS | 0.10000 | 100% |
| SLURP1 | Poor | 0.01700 | 50% | Poor | 0.00000 | 1% |
| SPACA4 | Good | 0.01900 | 50% | Poor | 0.02900 | 50% |
| LYPD4 | Good | 0.02100 | 50% | Poor | 0.00350 | 50% |
| LY6G5C | Good | 0.02700 | 50% | Good | 0.00002 | 1% |
| LYPD3 | Good | 0.02900 | 50% | Poor | 0.03500 | 50% |
| LY6L | Good | 0.04100 | 50% | Poor | 0.00330 | 20% |
| PLAUR | NS | 0.06900 | 100% | Poor | 0.00092 | 10% |
| LY6H | NS | 0.07800 | 100% | Good | 0.00070 | 10% |
| PSCA | NS | 0.09500 | 100% | Poor | 0.00000 | 1% |
| TEX101 | NS | 0.09600 | 100% | Poor | 0.00990 | 50% |
| CD177 | NS | 0.12000 | 100% | Poor | 0.00790 | 50% |
| LY6G6D | NS | 0.12000 | 100% | Poor | 0.01400 | 50% |
| PATE1 | NS | 0.13000 | 100% | Poor | 0.01200 | 50% |
| LY6K | NS | 0.14000 | 100% | Good | 0.00470 | 50% |
| LY6E | NS | 0.14000 | 100% | Poor | 0.00014 | 1% |
| LY6G6F | NS | 0.18000 | 100% | Poor | 0.00140 | 20% |
| LYPD2 | NS | 0.27000 | 100% | Poor | 0.00006 | 1% |
| PATE3 | NS | 0.29000 | 100% | Poor | 0.01200 | 50% |
| LYPD5 | NS | 0.35000 | 100% | Poor | 0.00001 | 1% |
| GML | NS | 0.45000 | 100% | Poor | 0.02800 | 50% |
| LY6G6C | NS | 0.48000 | 100% | Poor | 0.00018 | 2% |
Natural killer T-cell (NKT cells) have the characteristics of natural killer (NK) cells and T-cells [25, 26]. High expression of LY6D, SLURP1, LYPD5, PSCA, CD59, PATE2, LY6G6F and LY6E was significantly associated with poor OS outcome independent of NKT status. High expression of LY6G5C, LY6H, and LY6G5B was significantly associated with good OS outcome independent of NKT population. High expression of LYPD2, PLAUR, LY6G6C, and CD177 was significantly associated with poor OS outcome only in the NKT enriched population. High expression of PATE4 was associated with good OS outcome in NKT enriched and poor OS outcome in NKT decreased population. High expression of LYPD4, PATE3, PATE1, and LY6G6D was associated with poor OS outcome only in NKT decreased patient population (Table 5 and Supplementary Figure 6).
Table 5: High expression of LY6 genes and its association with overall survival outcome (OS) in TCGA dataset of PDAC patient population based on NKT cell status, as observed by using KM plotter tool
| Gene | NKT enriched population | NKT decreased population | ||||
|---|---|---|---|---|---|---|
| OS | p value | FDR | OS | p value | FDR | |
| LY6D | Poor | 0.00009 | 2% | Poor | 0.00100 | 20% |
| SLURP1 | Poor | 0.00010 | 2% | Poor | 0.00330 | 50% |
| LY6G5C | Good | 0.00030 | 2% | Good | 0.00490 | 50% |
| LYPD2 | Poor | 0.00035 | 5% | NS | 0.26000 | 100% |
| LYPD5 | Poor | 0.00170 | 20% | Poor | 0.04700 | 50% |
| PSCA | Poor | 0.00410 | 20% | Poor | 0.00260 | 50% |
| PATE4 | Good | 0.00700 | 50% | Poor | 0.01100 | 50% |
| CD59 | Poor | 0.00750 | 50% | Poor | 0.00310 | 20% |
| LY6H | Good | 0.00850 | 50% | Good | 0.00900 | 50% |
| PLAUR | Poor | 0.00990 | 50% | NS | 0.42000 | 100% |
| LY6K | Good | 0.01100 | 50% | NS | 0.40000 | 100% |
| LY6G5B | Good | 0.01400 | 50% | Good | 0.00340 | 50% |
| LY6G6C | Poor | 0.01500 | 50% | NS | 0.17000 | 100% |
| PATE2 | Poor | 0.01600 | 50% | Poor | 0.01100 | 50% |
| CD177 | Poor | 0.01700 | 50% | NS | 0.33000 | 100% |
| LY6G6F | Poor | 0.01700 | 50% | Poor | 0.00660 | 50% |
| LY6E | Poor | 0.03400 | 50% | Poor | 0.00700 | 50% |
| LYPD4 | NS | 0.06300 | 100% | Poor | 0.01900 | 50% |
| PATE3 | Good | 0.06900 | 100% | Poor | 0.03400 | 50% |
| PATE1 | NS | 0.08700 | 100% | Poor | 0.00650 | 50% |
| LY6G6D | NS | 0.29000 | 100% | Poor | 0.00160 | 20% |
CD4 positive T-cells are helper memory T-cells which can activate cytotoxic T-cells, natural killer T-cells, B-cells and macrophages to activate immune responses [27, 28]. High expression of LY6D, LY6E, LY6L, LYPD4, LYPD5, and SLURP1 was significantly associated with poor OS outcome independent of CD4+ memory T-cell status. High expression of LY6G5B was strongly associated with good OS outcome independent of CD4+ memory T-cell status. High expression of PATE1 was associated with good OS outcome in CD4+ enriched and poor OS outcome in CD4+ decreased population. High expression of PATE3, LYPD2, GML, PSCA, PATE2, LY6G6D, CD59, and LY6G6C was associated with poor OS outcome only in the CD4+decraesed population. High expression of LY6H, and LY6G5C was associated with good OS outcome in CD4+ decreased population (Table 6 and Supplementary Figure 7).
Table 6: High expression of LY6 genes and its association with overall survival outcome (OS) in TCGA dataset of PDAC patient population based on CD4+ memory T-cell status, as observed by using KM plotter tool
| Gene | CD4+ enriched population | CD4+ decreased population | ||||
|---|---|---|---|---|---|---|
| OS | p value | FDR | OS | p value | FDR | |
| LY6D | Poor | 0.000048 | 1% | Poor | 0.0001 | 5% |
| LY6E | Poor | 0.0012 | 10% | Poor | 0.0248 | >50% |
| LY6L | Poor | 0.0037 | 20% | Poor | 0.0056 | >50% |
| LYPD4 | Poor | 0.0038 | 20% | Poor | 0.003 | >50% |
| LY6G5B | Good | 0.0062 | >50% | Good | 0.009 | 50% |
| LYPD5 | Poor | 0.0142 | >50% | Poor | 0.0044 | 50% |
| SLURP1 | Poor | 0.0328 | >50% | Poor | 0.000013 | 1% |
| PATE1 | Good | 0.0465 | >50% | Poor | 0.0007 | 10% |
| PATE3 | NS | 0.061 | 100% | Poor | 0.0052 | >50% |
| LYPD2 | NS | 0.0854 | 100% | Poor | 0.0041 | >50% |
| GML | NS | 0.1008 | 100% | Poor | 0.0112 | >50% |
| PSCA | NS | 0.1014 | 100% | Poor | 0.000084 | 2% |
| PATE2 | NS | 0.1275 | 100% | Poor | 0.0004 | 20% |
| LY6G6D | NS | 0.1351 | 100% | Poor | 0.0016 | 10% |
| CD59 | NS | 0.1934 | 100% | Poor | 0.0006 | 10% |
| LY6H | NS | 0.214 | 100% | Good | 0.006 | 50% |
| LY6G5C | NS | 0.277 | 100% | Good | 0.00006 | 2% |
| LY6G6C | NS | 0.4561 | 100% | Poor | 0.0037 | 50% |
Presence of B-cells have been demonstrated to correlate with good prognosis in many different cancers [29–31]. Interestingly, LY6D has been shown to be involved in B-cell differentiation [32, 33]. High expression of LY6D, LY6E, PSCA, SLURP1, LYPD5, and PATE2 was strongly associated with poor outcome independent of B-cell status. High expression of LY6G5B and LY6G5C was associated with good OS outcome independent of B-cell status. High expression of PLAUR, LYPD4, LY6G6C, LYPD2, CD59, LY6G6D, LY6L, GML, PATE3, and PATE1 was associated with poor OS outcome only in B-cell decreased population. High expression of LY6H was associated with good OS outcome only in B-cell decreased population. (Table 7 and Supplementary Figure 8).
Table 7: High expression of LY6 genes and its association with overall survival outcome (OS) in TCGA dataset of PDAC patient population based on B-cell status, as observed by using KM plotter tool
| Gene | B cell enriched population | B cell decreased population | ||||
|---|---|---|---|---|---|---|
| OS | p value | FDR | OS | p value | FDR | |
| LY6D | Poor | 0.00003 | 1% | Poor | 0.0009 | 10% |
| LY6E | Poor | 0.006 | 10% | Poor | 0.0078 | >50% |
| PSCA | Poor | 0.0152 | 100% | Poor | 0.0005 | 5% |
| LY6G5B | Good | 0.0213 | >50% | Good | 0.0017 | 10% |
| SLURP1 | Poor | 0.0242 | >50% | Poor | 0.000009 | 1% |
| LY6G5C | Good | 0.0283 | 100% | Good | 0.0001 | 5% |
| LYPD5 | Poor | 0.0355 | >50% | Poor | 0.003 | 50% |
| PLAUR | NS | 0.0433 | 100% | Poor | 0.0644 | >50% |
| PATE2 | Poor | 0.0488 | 100% | Poor | 0.0014 | 50% |
| LYPD4 | NS | 0.0525 | 20% | Poor | 0.009 | >50% |
| LY6G6C | NS | 0.055 | 100% | Poor | 0.0078 | >50% |
| LYPD2 | NS | 0.0967 | 100% | Poor | 0.0021 | 50% |
| CD59 | NS | 0.1004 | 100% | Poor | 0.0000068 | 1% |
| LY6G6D | NS | 0.1892 | 100% | Poor | 0.0038 | >50% |
| LY6L | NS | 0.2433 | 20% | Poor | 0.043 | >50% |
| LY6H | NS | 0.3599 | 100% | Good | 0.0007 | 20% |
| GML | NS | 0.3734 | 100% | Poor | 0.0385 | >50% |
| PATE3 | NS | 0.4032 | 100% | Poor | 0.0261 | >50% |
| PATE1 | NS | 0.6189 | >50% | Poor | 0.0039 | 50% |
LY6 DNA were amplified in pancreatic ductal carcinoma
LY6 genes have been reported to be upregulated in multiple cancers [11, 34]. To test whether LY6 gene family members are amplified in PDAC, we assessed copy number variation data including DNA amplification and deep deletions from TCGA dataset (n = 177) and Pancreatic Cancer UTSW dataset (n = 109) hosted on cBioPortal tool [35]. We observed that LY6 genes located on the chromosome 8q24.3, PSCA, LY6K, SLURP1, LYPD2, LY6D, GML, LY6E, LY6H, and GPIHBP1 were co-amplified in most 9 to 28% of PDAC cases in TCGA and UTSW datasets. Interestingly, the cluster of LY6 genes which expressed on same genetic location were co-amplified in the same patients. The deep deletions in LY6 genes were only observed in few cases (Figure 2).
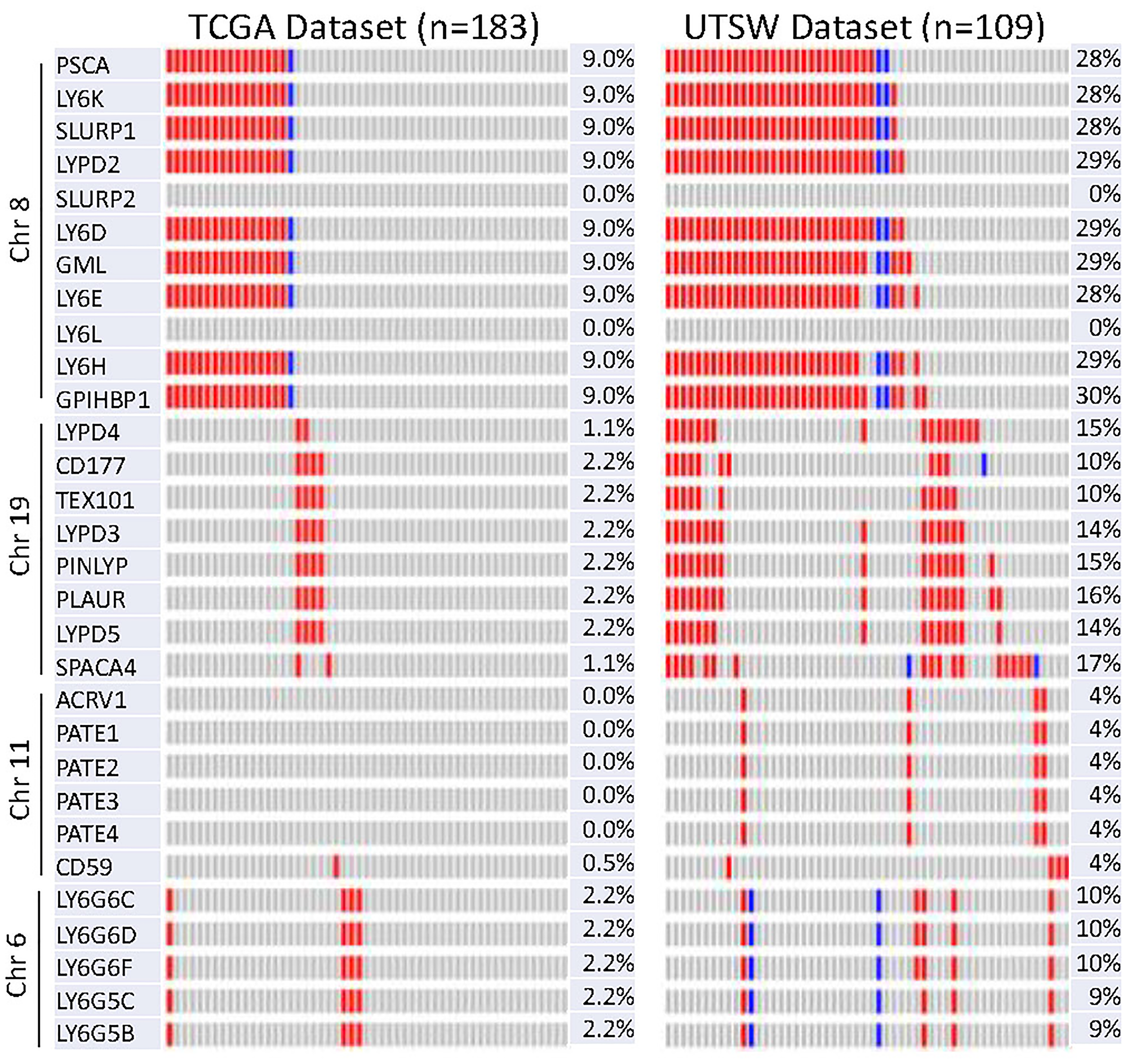
Figure 2: Oncoprint of copy number variation of 30 LY6 genes in TCGA and UTSW dataset using cBioPortal. Each column represents a patient/sample. DNA amplification in red and DNA deep deletion in blue are depicted in the indicated genes.
LY6 mRNA expression is upregulated in cancer compared to normal tissues
The differential expression of LY6 genes between the tumor tissue and the normal tissue will promote the understanding of their potentials as prognosis and therapeutic biomarkers. We evaluated 30 LY6 genes in the Pei dataset of pancreatic tumors versus normal tissues hosted at Oncomine. LY6D, LY6E, PLAUR, PSCA, CD59 and LYPD3 mRNA expression was significantly increased in the PDAC tumor tissues than normal adjacent tissues (p < 0.01) (Figure 3).
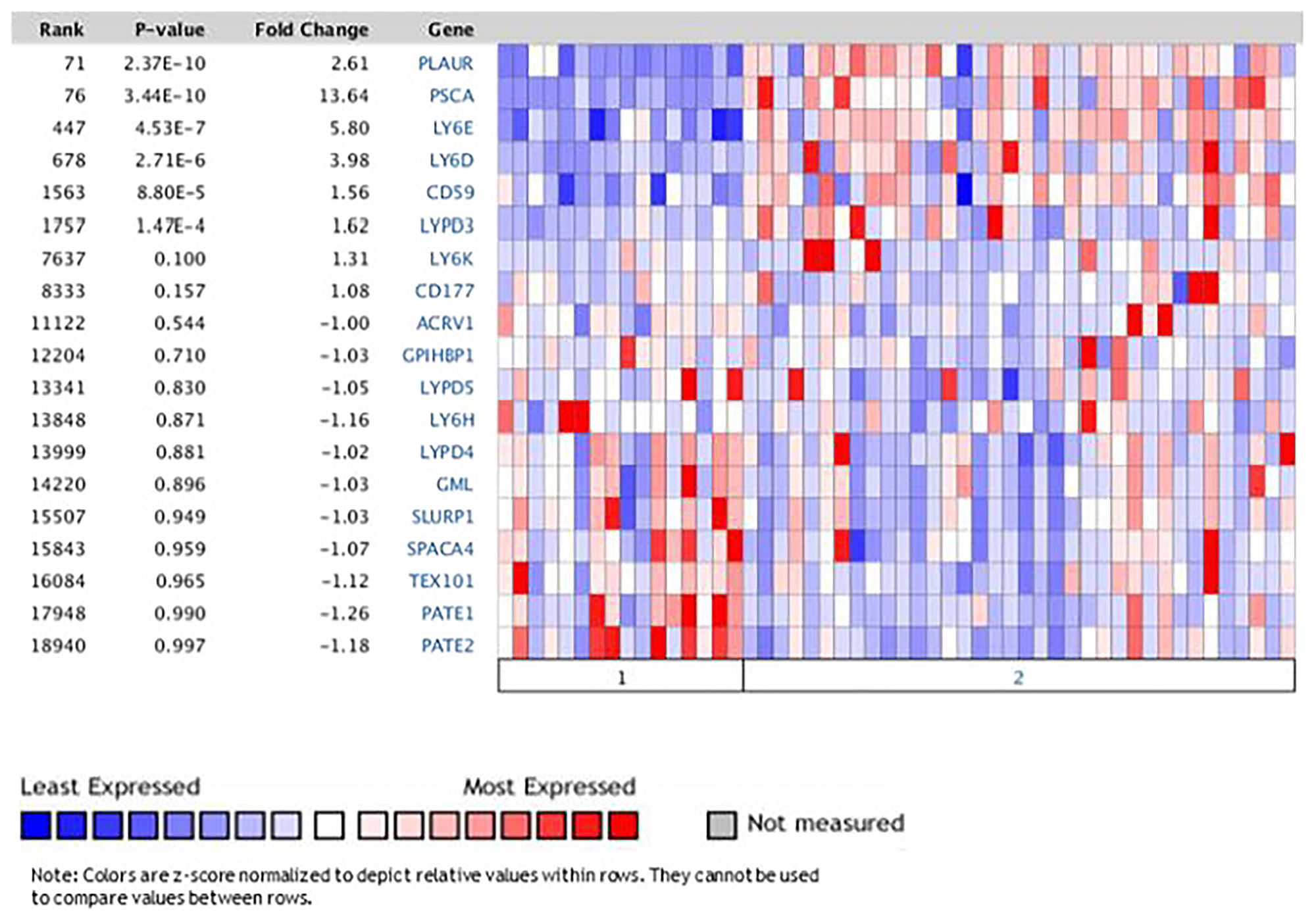
Figure 3: Oncomine tool showed significant increase expression of LY6 genes in pancreatic tumor vs normal tissues in Pei dataset comparing normal vs tumor PDAC tissue. (1) Normal Pancreas (16 cases); (2) Pancreatic Carcinoma (36 cases).
DISCUSSION
The first step towards research and development of pancreatic cancer treatment will be to identify and define the novel tumor-specific biomarkers. Mutations in the oncogenic KRAS gene occur in over 90% of patients and are viewed as driving force of pancreatic cancer [5]. However, a history of detailed knowledge in the KRAS mechanistic pathway has not yet led to a clinical breakthrough in the treatment of PDAC [5, 6]. With a five-year survival rate of less than 10%, there is an urgent need for innovative treatment strategies. Compared to other solid malignancies, challenges in pancreatic cancer include the immensely immunosuppressive tumor microenvironment, in addition to the presence of a dense desmoplastic barrier, which limits the diffusion of therapeutic drugs and the infiltration of immunotherapy-based anti-tumor immune cells [5–8]. An increased understanding of the key molecular pathways unique to pancreatic cancer which contribute to its immunosuppressive and stem cell-like properties is required to develop novel and successful therapeutic strategies against pancreatic cancer. Herein, we analyze the expression of LY6 gene family and its association with OS outcome in clinical samples of PDAC.
LY6 gene expression in the context of tumor microenvironment
Mesenchymal stems cell (MSC) enriched pancreatic tumors
MSC enriched tumors showed the strongest association between high expression of LY6D/SLURP1/PSCA/CD59 and low overall survival outcome in PDAC. PDAC tumors with low MSC did not show significant association with high LY6 gene expression to OS outcome. This observation suggests that LY6 gene expression and its association with OS outcome is specifically relevant in the presence of MSCs in the pancreatic tumor microenvironment. MSCs are instrumental in providing the immunosuppressive tumor microenvironment. They can suppress CD4 and CD8 positive T-cells. MSCs can secrete various growth factors which can regulate gene expression directly on cancer cells [36]. It remains to be seen if the LY6 gene expression is associated with a direct immunosuppressive environment due to presence of MSCs.
Immune cell enriched pancreatic tumors
We observed that high expression of many LY6 genes were associated significantly with lower OS in PDAC population enriched for Treg, CD8, macrophages, NKT, B-cells and CD4+ immune cells. However, a single pattern of association did not emerge for each of the LY6 genes, suggesting the LY6 genes may be differentially regulated. In contrast to other LY6 genes, high expression for CD59 was strongly associated with lower OS in PDAC population decreased for CD8, Treg, macrophages, NKT, B-cells and CD4+ immune cells. This observation suggests that CD59 is associated with tumor environments which present with lower immune cell infiltrates [37]. The overexpression of CD59 in pancreatic cancer has major consequences on the tumor microenvironment and was previously shown to be required for stem cell evasion of complement surveillance, a biological mechanism for eliminating cancer stem cells in epithelial cancer [37].
We observed that increased mRNA of SLURP1 was associated with lower OS outcome in pancreatic cancer. This observation was in agreement with public data from The Human protein Atlas data which showed that high expression of mRNA for SLURP1 is associated with lower OS. This observation, however was in conflict with a previously reported finding that high expression of SLURP1 protein is associated with higher OS outcome in pancreatic cancer [38]. Further studies are required to understand the role of SLURP1 mRNA and protein in pancreatic cancer and its association with OS outcome in PDAC. In vivo evidence indicates that SLURP1 is a major component of maintaining immune privilege through inhibiting leukocytic binding and infiltration in a corneal model, suggesting that SLURP1 can also serve as a potent inhibitor of immune activity [39, 40]. However this has not been tested in tumor models, it is plausible that SLURP1 can contribute to pancreatic cancer’s immunosuppressive tumor microenvironment and suppress anti-tumor immune responses.
LY6 proteins and maintenance of proliferation and stem cell-like properties
LY6D, LY6E, PSCA, and PLAUR, known markers tumorigenesis and cancer cell maintenance, were significantly associated with lower overall survival outcome in our pancreatic cancer analysis [10, 11]. LY6D is a GPI-anchored member of the LY6 family with a recently established association with aggressive cancers and poor patient outcome [10, 11]. Under normal physiological conditions, LY6D is most commonly used as a marker of early B cell lineage; however, in response to genotoxic stressors such as radiation and chemotherapy, LY6D expression is upregulated in numerous cancer types and is suggested to contribute to distant metastasis in breast cancer [41–43]. LY6D serves as a marker of luminal progenitors with bi-lineage capacity and intrinsic castration-resistant properties in prostate cancer [44]. Together, our analysis and previous research suggests that LY6D expression may indicate a more aggressive pancreatic cancer phenotype and it would therefore be beneficial to explore the precise mechanism of action for LY6D to determine its therapeutic potential.
LY6E, a GPI-anchored member of the LY6 family, has recently been implicated as a driver of tumorigenesis and stem cell maintenance through inhibiting expression of the tumor suppressor PTEN and inducing the upregulation of the HIF-1 pathway [45, 46]. Notably, inhibition of LY6E with siRNA was shown to restore PTEN expression, induce G1-S phase cell cycle arrest, and increase apoptosis in gastric cancer, suggesting that LY6E’s inhibition may be enough to cause anti-tumor effects in some cancers [46]. On a clinical level, high LY6E expression correlates with poor overall patient survival in various malignant tumors such as those of gastric, breast, head and neck, lung, bladder, brain, and skin origin [11, 45]. Importantly, in pancreatic cancer specifically, LY6E was suggested to be a marker for cancer cells with stem cell properties and was used in addition to the stem cell markers TACSTD1 and CD44 to establish a sorting technique to obtain clonal colony-forming pancreatic cancer stem cells [47].
GPI-anchored LY6 family member PSCA is most commonly attributed to prostate stem cells and prostate cancer; however, its overexpression in pancreatic cancer and limited expression in normal pancreatic cells is also well recognized [48, 49]. PSCA can be a target of immunotherapy strategies such as anti-PSCA antibody and anti-PSCA CAR-T cell therapy, with a clinical trial currently underway for the latter [6]. However, PSCA’s function in pancreatic cancer is still unknown and requires further research.
Gene copy number amplifications of LY6 proteins on chromosome 8
LY6D, LY6E, LY6H, LY6K, PSCA, SLURP1, LYPD2, and GPIHP1 had co-amplifications of their gene copy numbers as seen on cBioPortal in TCGA dataset While increased DNA copy number does not necessarily indicate increased levels of protein expression, the mRNA expression data for these genes was found to be relatively high in the patients with gene copy number amplifications, suggesting that an amplified copy number for LY6 genes on chromosome 8 increases LY6 gene expression. Understanding what causes both an amplification of LY6 gene copy number and an overexpression of LY6 genes on chromosome 8 may reveal important insight into the molecular phenotype of this subset of pancreatic cancer patients.
The “good” LY6 genes
High expression of LY6G5B, LY6G5C, and LY6G6C was associated with high OS outcome. These LY6 genes are located among the MHC class III proteins and are known as MHC-linked LY6 genes. Although the precise functions of the LY6 family members LY6G5B, LY6G5C, LY6G6C are unknown, they are suggested to be located on filopodia and secreted proteins with binding potential to the cell surface and may be involved in cell signaling. Interestingly, potential ligands for LY6G5C, LY6G6C were found to be present on K562 cells, an undifferentiated megakaryocyte cell line, among a panel of cell lines, indicating a potential role in hematopoietic cell differentiation [50, 51]. It is unclear how these genes are associated with higher OS outcome. It is likely that cell-cell interaction may play important role in LY6 signaling.
Concluding remarks
LY6 family proteins are either attached to the outer cell surface through a GPI-anchor or are secreted into the extracellular matrix, making them relatively accessible for drug inhibition or immunotherapy targeting. In this report, we sought to analyze the mRNA expression of LY6 gene family and its association with overall survival (OS) outcome in pancreatic cancer patients. We focused our exploration on thirty LY6 family genes scattered on various chromosomes [2, 10, 11]. We found that high expression of sixteen LY6 family members significantly associated with lower OS outcome and high expression of four genes significantly associated with higher OS outcome. Future research is required to translate our DNA and mRNA analysis into a proteomic and molecular interaction context. With further research, these findings may lead to potential successful screening markers for PDAC, as well as new, personalized targeted therapy or guidance of current standard chemotherapy regimens based on LY6 gene expression.
Materials and Methods
Overall survival (OS) outcome analysis
The Kaplan-Meier (KM) plotter (https://kmplot.com/analysis/) online web-based tools allows users to observe the association of mRNA expression to over survival outcome with or the context of tumor microenvironment cellular composition [18]. For our analysis, we selected PDAC dataset on 177 patients among the list of pan cancer dataset (TCGA collection) in the gene expression RNA sequencing (RNA-seq) tab on the homepage of KM plotter. This dataset was used to visualize the association of mRNA expression of the LY6 gene family with overall survival (OS) outcome in PDAC. The number of patients in low and high expression of indicated gene is displayed in the Supplementary Table 1. To estimate if inherent cellular content plays a role in the association of LY6 gene expression with the OS outcome, we used the restricted analysis feature of the KM Plotter tool. This feature allowed us to observe the OS outcome in patient samples with enriched or decreased cellular content which included mesenchymal stem cell, CD8+ T cells, macrophages, NK T cells, CD4+ memory T cells, regulatory T cells, and B cells. The number of patients in the various subgroups for low expressing genes were in the range of 43 to 127 and for high expressing genes were 51 to 134. The exact number of patients in each subgroup can be found in Supplementary Figure 1 through 8, which corresponds to the data presented in Supplementary Table 1, Tables 2–7.
Disease free survival (DFS) analysis
RNA-seq normalized gene expression data at z-score level and the corresponding clinical data from the TCGA Pancreatic Adenocarcinoma cohort were downloaded from the cBioPortal (https://www.cbioportal.org/) [16, 17]. The TCGA dataset contained data from 179 patients, of which 139 had disease free survival (DFS) status, including one metastasis sample. Thus, 138 primary tumor samples were used for DFS analysis. DFS analysis was performed using the R statistical programming environment. We used the cgdsr R package (https://CRAN.R-project.org/package=cgdsr) in the R statistical programming software to download and query the data [52]. For each gene, the optimal cutoff into “low” and “high” expression was determined by inbuilt algorithm in the survMisc R package [53]. We used this data to explore the association of the LY6 genes with DFS outcome. The survival analysis and KM plots were done using the survMisc R package. Then the significance of the association of each gene with DFS outcome was reported using a log rank test with p-value < 0.05.
Copy number alteration analysis
We analyzed the copy number alteration data from LY6 genes in TCGA PDAC in the cBioPortal, which included data for 183 patients and UTSW PDAC dataset, which contained data for 109 patients.
Differential gene expression analysis
The Oncomine™ Platform (Thermo Fisher, Ann Arbor, MI) (https://www.oncomine.org [54]) was applied to assess the differential expression of LY6 genes between the pancreatic tumors and normal tissues from the Pei dataset [55]. Pei et al. performed differential gene expression analysis in 16 normal and 36 pancreatic cancer samples using a human genome U133 Plus 2.0 Array measuring 19574 genes, submitted as a public dataset in the NCBI Gene Expression Omnibus public repository GSE16515 [55].
CONFLICTS OF INTEREST
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
The opinions expressed herein are those of the authors and are not necessarily representative of those of the government of the United States, the Uniformed Services University, the Department of Defense, NIH, or any other US federal agency.
FUNDING
This research has been supported by NIH R01CA227694 and Uniformed Services University Start-Up funds by Vice President of Research office, USUHS to Geeta Upadhyay.
References
1. Rawla P, Sunkara T, Gaduputi V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J Oncol. 2019; 10:10–27. https://doi.org/10.14740/wjon1166. [PubMed].
2. Luo L, McGarvey P, Madhavan S, Kumar R, Gusev Y, Upadhyay G. Distinct lymphocyte antigens 6 (Ly6) family members Ly6D, Ly6E, Ly6K and Ly6H drive tumorigenesis and clinical outcome. Oncotarget. 2016; 7:11165–11193. https://doi.org/10.18632/oncotarget.7163. [PubMed].
3. Ballehaninna UK, Chamberlain RS. The clinical utility of serum CA 19-9 in the diagnosis, prognosis and management of pancreatic adenocarcinoma: An evidence based appraisal. J Gastrointest Oncol. 2012; 3:105–119. https://doi.org/10.3978/j.issn.2078-6891.2011.021. [PubMed].
4. Di Carlo C, Brandi J, Cecconi D. Pancreatic cancer stem cells: Perspectives on potential therapeutic approaches of pancreatic ductal adenocarcinoma. World J Stem Cells. 2018; 10:172–182. https://doi.org/10.4252/wjsc.v10.i11.172. [PubMed].
5. Liu P, Wang Y, Li X. Targeting the untargetable KRAS in cancer therapy. Acta Pharm Sin B. 2019; 9:871–879. https://doi.org/10.1016/j.apsb.2019.03.002. [PubMed].
6. McCormick F. KRAS as a Therapeutic Target. Clin Cancer Res. 2015; 21:1797–1801. https://doi.org/10.1158/1078-0432.CCR-14-2662. [PubMed].
7. Martinez-Bosch N, Vinaixa J, Navarro P. Immune Evasion in Pancreatic Cancer: From Mechanisms to Therapy. Cancers (Basel). 2018; 10:6. https://doi.org/10.3390/cancers10010006. [PubMed].
8. Akce M, Zaidi MY, Waller EK, El-Rayes BF, Lesinski GB. The Potential of CAR T Cell Therapy in Pancreatic Cancer. Front Immunol. 2018; 9:2166. https://doi.org/10.3389/fimmu.2018.02166. [PubMed].
9. Looi CK, Chung FF, Leong CO, Wong SF, Rosli R, Mai CW. Therapeutic challenges and current immunomodulatory strategies in targeting the immunosuppressive pancreatic tumor microenvironment. J Exp Clin Cancer Res. 2019; 38:162. https://doi.org/10.1186/s13046-019-1153-8. [PubMed].
10. Loughner CL, Bruford EA, McAndrews MS, Delp EE, Swamynathan S, Swamynathan SK. Organization, evolution and functions of the human and mouse Ly6/uPAR family genes. Hum Genomics. 2016; 10:10. https://doi.org/10.1186/s40246-016-0074-2. [PubMed].
11. Upadhyay G. Emerging Role of Lymphocyte Antigen-6 Family of Genes in Cancer and Immune Cells. Front Immunol. 2019; 10:819. https://doi.org/10.3389/fimmu.2019.00819. [PubMed].
12. Brusselaers N, Ekwall K, Durand-Dubief M. Copy number of 8q24.3 drives HSF1 expression and patient outcome in cancer: an individual patient data meta-analysis. Hum Genomics. 2019; 13:54. https://doi.org/10.1186/s40246-019-0241-3. [PubMed].
13. Ballabio S, Craparotta I, Paracchini L, Mannarino L, Corso S, Pezzotta MG, Vescio M, Fruscio R, Romualdi C, Dainese E, Ceppi L, Calura E, Pileggi S, et al. Multisite analysis of high-grade serous epithelial ovarian cancers identifies genomic regions of focal and recurrent copy number alteration in 3q26.2 and 8q24.3. Int J Cancer. 2019; 145:2670–2681. https://doi.org/10.1002/ijc.32288. [PubMed].
14. Bilal E, Vassallo K, Toppmeyer D, Barnard N, Rye IH, Almendro V, Russnes H, Borresen-Dale AL, Levine AJ, Bhanot G, Ganesan S. Amplified loci on chromosomes 8 and 17 predict early relapse in ER-positive breast cancers. PLoS One. 2012; 7:e38575. https://doi.org/10.1371/journal.pone.0038575. [PubMed].
15. Upadhyay G, Yin Y, Yuan H, Li X, Derynck R, Glazer RI. Stem cell antigen-1 enhances tumorigenicity by disruption of growth differentiation factor-10 (GDF10)-dependent TGF-beta signaling. Proc Natl Acad Sci U S A. 2011; 108:7820–7825. https://doi.org/10.1073/pnas.1103441108. [PubMed].
16. Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012; 2:401–404. https://doi.org/10.1158/2159-8290.CD-12-0095. [PubMed].
17. Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013; 6:pl1. https://doi.org/10.1126/scisignal.2004088. [PubMed].
18. Nagy A, Lanczky A, Menyhart O, Gyorffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep. 2018; 8:9227. https://doi.org/10.1038/s41598-018-27521-y. [PubMed].
19. Chaturvedi P, Gilkes DM, Wong CC, Kshitiz, Luo W, Zhang H, Wei H, Takano N, Schito L, Levchenko A, Semenza GL. Hypoxia-inducible factor-dependent breast cancer-mesenchymal stem cell bidirectional signaling promotes metastasis. J Clin Invest. 2013; 123:189–205. https://doi.org/10.1172/JCI64993. [PubMed].
20. El-Haibi CP, Bell GW, Zhang J, Collmann AY, Wood D, Scherber CM, Csizmadia E, Mariani O, Zhu C, Campagne A, Toner M, Bhatia SN, Irimia D, et al. Critical role for lysyl oxidase in mesenchymal stem cell-driven breast cancer malignancy. Proc Natl Acad Sci U S A. 2012; 109:17460–17465. https://doi.org/10.1073/pnas.1206653109. [PubMed].
21. Patel SA, Meyer JR, Greco SJ, Corcoran KE, Bryan M, Rameshwar P. Mesenchymal stem cells protect breast cancer cells through regulatory T cells: role of mesenchymal stem cell-derived TGF-beta. J Immunol. 2010; 184:5885–5894. https://doi.org/10.4049/jimmunol.0903143. [PubMed].
22. Gajewski TF, Corrales L, Williams J, Horton B, Sivan A, Spranger S. Cancer Immunotherapy Targets Based on Understanding the T Cell-Inflamed Versus Non-T Cell-Inflamed Tumor Microenvironment. Adv Exp Med Biol. 2017; 1036:19–31. https://doi.org/10.1007/978-3-319-67577-0_2. [PubMed].
23. Van Gassen N, Staels W, Van Overmeire E, De Groef S, Sojoodi M, Heremans Y, Leuckx G, Van de Casteele M, Van Ginderachter JA, Heimberg H, De Leu N. Concise Review: Macrophages: Versatile Gatekeepers During Pancreatic beta-Cell Development, Injury, and Regeneration. Stem Cells Transl Med. 2015; 4:555–563. https://doi.org/10.5966/sctm.2014-0272. [PubMed].
24. Cruz AF, Rohban R, Esni F. Macrophages in the pancreas: Villains by circumstances, not necessarily by actions. Immun Inflamm Dis. 2020; 8:807–824. https://doi.org/10.1002/iid3.345. [PubMed].
25. Krijgsman D, Hokland M, Kuppen PJK. The Role of Natural Killer T Cells in Cancer-A Phenotypical and Functional Approach. Front Immunol. 2018; 9:367. https://doi.org/10.3389/fimmu.2018.00367. [PubMed].
26. Terabe M, Berzofsky JA. NKT cells in immunoregulation of tumor immunity: a new immunoregulatory axis. Trends Immunol. 2007; 28:491–496. https://doi.org/10.1016/j.it.2007.05.008. [PubMed].
27. Brightman SE, Naradikian MS, Miller AM, Schoenberger SP. Harnessing neoantigen specific CD4 T cells for cancer immunotherapy. J Leukoc Biol. 2020; 107:625–633. https://doi.org/10.1002/JLB.5RI0220-603RR. [PubMed].
28. Farber DL, Yudanin NA, Restifo NP. Human memory T cells: generation, compartmentalization and homeostasis. Nat Rev Immunol. 2014; 14:24–35. https://doi.org/10.1038/nri3567. [PubMed].
29. Lechner A, Schlosser HA, Thelen M, Wennhold K, Rothschild SI, Gilles R, Quaas A, Siefer OG, Huebbers CU, Cukuroglu E, Goke J, Hillmer A, Gathof B, et al. Tumor-associated B cells and humoral immune response in head and neck squamous cell carcinoma. Oncoimmunology. 2019; 8:1535293. https://doi.org/10.1080/2162402X.2018.1535293. [PubMed].
30. Al-Shibli KI, Donnem T, Al-Saad S, Persson M, Bremnes RM, Busund LT. Prognostic effect of epithelial and stromal lymphocyte infiltration in non-small cell lung cancer. Clin Cancer Res. 2008; 14:5220–5227. https://doi.org/10.1158/1078-0432.CCR-08-0133. [PubMed].
31. Hennequin A, Derangere V, Boidot R, Apetoh L, Vincent J, Orry D, Fraisse J, Causeret S, Martin F, Arnould L, Beltjens F, Ghiringhelli F, Ladoire S. Tumor infiltration by Tbet+ effector T cells and CD20+ B cells is associated with survival in gastric cancer patients. Oncoimmunology. 2016; 5:e1054598. https://doi.org/10.1080/2162402X.2015.1054598. [PubMed].
32. Dress RJ, Dutertre CA, Giladi A, Schlitzer A, Low I, Shadan NB, Tay A, Lum J, Kairi M, Hwang YY, Becht E, Cheng Y, Chevrier M, et al. Plasmacytoid dendritic cells develop from Ly6D(+) lymphoid progenitors distinct from the myeloid lineage. Nat Immunol. 2019; 20:852–864. https://doi.org/10.1038/s41590-019-0420-3. [PubMed].
33. Lee PY, Wang JX, Parisini E, Dascher CC, Nigrovic PA. Ly6 family proteins in neutrophil biology. J Leukoc Biol. 2013; 94:585–594. https://doi.org/10.1189/jlb.0113014. [PubMed].
34. Upadhyay G. Emerging Role of Novel Biomarkers of Ly6 Gene Family in Pan Cancer. Adv Exp Med Biol. 2019; 1164:47–61. https://doi.org/10.1007/978-3-030-22254-3_4. [PubMed].
35. Witkiewicz AK, McMillan EA, Balaji U, Baek G, Lin WC, Mansour J, Mollaee M, Wagner KU, Koduru P, Yopp A, Choti MA, Yeo CJ, McCue P, et al. Whole-exome sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nat Commun. 2015; 6:6744. https://doi.org/10.1038/ncomms7744. [PubMed].
36. Cuiffo BG, Karnoub AE. Mesenchymal stem cells in tumor development: emerging roles and concepts. Cell Adh Migr. 2012; 6:220–230. https://doi.org/10.4161/cam.20875. [PubMed].
37. Chen J, Ding P, Li L, Gu H, Zhang X, Zhang L, Wang N, Gan L, Wang Q, Zhang W, Hu W. CD59 Regulation by SOX2 Is Required for Epithelial Cancer Stem Cells to Evade Complement Surveillance. Stem Cell Reports. 2017; 8:140–151. https://doi.org/10.1016/j.stemcr.2016.11.008. [PubMed].
38. Throm VM, Mannle D, Giese T, Bauer AS, Gaida MM, Kopitz J, Bruckner T, Plaschke K, Grekova SP, Felix K, Hackert T, Giese NA, Strobel O. Endogenous CHRNA7-ligand SLURP1 as a potential tumor suppressor and anti-nicotinic factor in pancreatic cancer. Oncotarget. 2018; 9:11734–11751. https://doi.org/10.18632/oncotarget.24312. [PubMed].
39. Swamynathan S, Buela KA, Kinchington P, Lathrop KL, Misawa H, Hendricks RL, Swamynathan SK. Klf4 regulates the expression of Slurp1, which functions as an immunomodulatory peptide in the mouse cornea. Invest Ophthalmol Vis Sci. 2012; 53:8433–8446. https://doi.org/10.1167/iovs.12-10759. [PubMed].
40. Swamynathan S, Tiwari A, Loughner CL, Gnalian J, Alexander N, Jhanji V, Swamynathan SK. The secreted Ly6/uPAR-related protein-1 suppresses neutrophil binding, chemotaxis, and transmigration through human umbilical vein endothelial cells. Sci Rep. 2019; 9:5898. https://doi.org/10.1038/s41598-019-42437-x. [PubMed].
41. Kurosawa M, Jeyasekharan AD, Surmann EM, Hashimoto N, Venkatraman V, Kurosawa G, Furukawa K, Venkitaraman AR, Kurosawa Y. Expression of LY6D is induced at the surface of MCF10A cells by X-ray irradiation. FEBS J. 2012; 279:4479–4491. https://doi.org/10.1111/febs.12034. [PubMed].
42. Inlay MA, Bhattacharya D, Sahoo D, Serwold T, Seita J, Karsunky H, Plevritis SK, Dill DL, Weissman IL. Ly6d marks the earliest stage of B-cell specification and identifies the branchpoint between B-cell and T-cell development. Genes Dev. 2009; 23:2376–2381. https://doi.org/10.1101/gad.1836009. [PubMed].
43. Mayama A, Takagi K, Suzuki H, Sato A, Onodera Y, Miki Y, Sakurai M, Watanabe T, Sakamoto K, Yoshida R, Ishida T, Sasano H, Suzuki T. OLFM4, LY6D and S100A7 as potent markers for distant metastasis in estrogen receptor-positive breast carcinoma. Cancer Sci. 2018; 109:3350–3359. https://doi.org/10.1111/cas.13770. [PubMed].
44. Barros-Silva JD, Linn DE, Steiner I, Guo G, Ali A, Pakula H, Ashton G, Peset I, Brown M, Clarke NW, Bronson RT, Yuan GC, Orkin SH, et al. Single-Cell Analysis Identifies LY6D as a Marker Linking Castration-Resistant Prostate Luminal Cells to Prostate Progenitors and Cancer. Cell Rep. 2018; 25:3504–3518.e3506. https://doi.org/10.1016/j.celrep.2018.11.069. [PubMed].
45. Yeom CJ, Zeng L, Goto Y, Morinibu A, Zhu Y, Shinomiya K, Kobayashi M, Itasaka S, Yoshimura M, Hur CG, Kakeya H, Hammond EM, Hiraoka M, et al. LY6E: a conductor of malignant tumor growth through modulation of the PTEN/PI3K/Akt/HIF-1 axis. Oncotarget. 2016; 7:65837–65848. https://doi.org/10.18632/oncotarget.11670. [PubMed].
46. Lv Y, Song Y, Ni C, Wang S, Chen Z, Shi X, Jiang Q, Cao C, Zuo Y. Overexpression of Lymphocyte Antigen 6 Complex, Locus E in Gastric Cancer Promotes Cancer Cell Growth and Metastasis. Cell Physiol Biochem. 2018; 45:1219–1229. https://doi.org/10.1159/000487453. [PubMed].
47. Gou S, Liu T, Wang C, Yin T, Li K, Yang M, Zhou J. Establishment of clonal colony-forming assay for propagation of pancreatic cancer cells with stem cell properties. Pancreas. 2007; 34:429–435. https://doi.org/10.1097/MPA.0b013e318033f9f4. [PubMed].
48. Katari UL, Keirnan JM, Worth AC, Hodges SE, Leen AM, Fisher WE, Vera JF. Engineered T cells for pancreatic cancer treatment. HPB (Oxford). 2011; 13:643–650. https://doi.org/10.1111/j.1477-2574.2011.00344.x. [PubMed].
49. Wente MN, Jain A, Kono E, Berberat PO, Giese T, Reber HA, Friess H, Buchler MW, Reiter RE, Hines OJ. Prostate stem cell antigen is a putative target for immunotherapy in pancreatic cancer. Pancreas. 2005; 31:119–125. https://doi.org/10.1097/01.mpa.0000173459.81193.4d. [PubMed].
50. Janeway CA. Immunobiology, 5th edition: The Immune System in Health and Disease. Garland Science: New York, 2001.
51. Mallya M, Campbell RD, Aguado B. Characterization of the five novel Ly-6 superfamily members encoded in the MHC, and detection of cells expressing their potential ligands. Protein Sci. 2006; 15:2244–2256. https://doi.org/10.1110/ps.062242606. [PubMed].
52. Computing RFfS. R: A language and environment for statistical computing: Vienna, Austria, 2013.
53. Dardis C. Miscellaneous Functions for Survival Data, 2018.
54. Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, Barrette T, Pandey A, Chinnaiyan AM. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004; 6:1–6. https://doi.org/10.1016/s1476-5586(04)80047-2. [PubMed].
55. Pei H, Li L, Fridley BL, Jenkins GD, Kalari KR, Lingle W, Petersen G, Lou Z, Wang L. FKBP51 affects cancer cell response to chemotherapy by negatively regulating Akt. Cancer Cell. 2009; 16:259–266. https://doi.org/10.1016/j.ccr.2009.07.016. [PubMed].

