Introduction
Cancer is the second leading cause of morbidity and mortality worldwide. GLOBOCAN (Global cancer statistics) estimated for 2018 a total of 18.1 million people with cancer and 9.6 million deaths from this disease [1]. The most frequent types of cancer differ among populations due to lifestyle and socioeconomic differences. According to a 2019 USA estimate, the three most prevalent types of cancer among men were prostate cancer (3.65 million cases), colorectal cancer (776,000), and melanoma (684,000); among women, these were breast cancer (3.86 million), cervical cancer (807,000), and colorectal cancer (768,000) [2]. In China, 4.3 million new cases and approximately 2.8 million cancer deaths were estimated in 2015. Lung, stomach, liver, esophageal, and colorectal cancer are the leading causes of cancer death among Chinese [3]. Cardiovascular disease (CVD) is the leading cause of morbidity and mortality worldwide, with over 422.7 million estimated cases and 17.92 million deaths in 2015 alone. Coronary artery disease is the most prevalent CVD, followed by stroke [4].
Although cancer and CVD are considered distinct entities, they have similarities and interactions among risk factors (e.g., smoking habits, physical activity, dietary habits, obesity, and diabetes mellitus), supporting a shared biology [5]. Although the rate of cancer death has been decreasing over the years, the risk for CVD in cancer survivors is a health problem that should be monitored by physicians, because individuals in long-term cancer treatment may develop CVD, leading to death or a worsening quality of life [6–8]. CVD in these patients is related to disruption in the organization of the cardiac tissues (such as those caused by chemotherapy, radiotherapy, and other drugs) [6, 9] which can result in several heart problems, hypertension, and cerebrovascular accidents, among other complications. Although chronic inflammation is the link in the pathogenesis of cancer and CVD [10, 11], additional factors may be shared among these diseases, including those that cannot be modified by individuals, such as age, gender, ethnicity, and genetic differences related to the immune response [5].
Cytokines are mediators of the immune response, and some, such as interferon-α (IFN-α), interleukin (IL)-2, and granulocyte macrophage colony stimulating factor (GM-GSF), are currently indicated in cancer treatment. Other cytokines, such as IL-12, -15, and -21, are undergoing clinical trials [12]. Cardiovascular immunotherapy is also being studied for therapeutic use. One strategy is to target the immune system’s ability to regulate arterial inflammation, for instance, neutralizing proinflammatory cytokines such as IL-1β to resolve inflammatory atherosclerosis. Maintaining tissue homeostasis and repairing lesions are also possibilities, which can be carried out through the action of cytokines such as IL-4 and IL-13 [13].
Interleukin-16 (IL-16) is a chemotactic cytokine whose gene (IL16) is located on chromosome 15q26.3. The receptor of IL-16 is a CD4 molecule initially described as inhibiting the interaction between CD4 and HIV-1 [14, 15]. This cytokine is coded as a precursor molecule called pro-IL-16, which is cleaved by caspase-3, generating a smaller secreted molecule [16]. The caspase-3 cleavage site is located between the PDZ2 and PDZ3 domains of the precursor protein and allows processing of pro-IL-16 into its mature form [17]. Both pro-IL-16 and mature IL-16 are biologically active. In lymphocytes, pro-IL-16 acts as a transcriptional repressor in the cell, regulating cell cycle progression through its N-terminal domain [18]. Meanwhile, mature IL-16 is secreted by cells such as CD8+ T and B lymphocytes and is responsible for the processes of chemotaxis, cell growth, and differentiation (e.g., increases IL-2 receptor expression).
IL-16 participates in malignant cell proliferation and transformation, acting on a variety of cells involved in the immune response, alone or in conjunction with other cytokines [18–21]. IL-16 levels were directly correlated with gastrointestinal tumor progression [22] and multiple myeloma [23, 24]; therefore, these malignant tumors are suggested targets for anti-IL-16 therapies [25]. Other studies also demonstrated increases in tissue expression and serum concentrations of IL-16 in malignant ovarian tumors [26] and cutaneous T-cell lymphoma [20, 27]. In cardiovascular diseases, high IL-16 serum concentrations were associated with asymptomatic carotid plaques and reduced numbers of cardiovascular events after surgery, suggesting a protective profile in atherosclerosis and the risk of cardiovascular events [28–30]. However, increased levels of this cytokine were also associated with cardiac fibrosis and myocardial stiffening [31], production of proinflammatory cytokines such as IL-1β and IL-6 [32], as well as the risk for acute myocardial infarction [33], being a potential target for CVD treatments [34].
Single nucleotide polymorphisms (SNPs) are common genetic variations that can alter protein composition or gene expression and thus modify the functioning of immune response components such as cytokines. These modifications may lead to changes in the immune response, cell cycle regulation and metabolism, and DNA repair associated with cancer and CVD susceptibility [35, 36]. SNPs may be located in gene promoter regions (altering gene expression and epigenetic modifications), exons (affecting transcription and translation), introns (modifying splicing and regulation), as well as 5′ and 3′ untranslated regions (UTRs) (affecting translation through microRNA binding) [36]. When associations of SNPs with diseases are confirmed, this knowledge may offer greater understanding of disease prevention, prediction, prognosis, and treatment. Four polymorphisms commonly studied for IL16 are rs4778889 (located in the promoter region), rs11556218 (missense mutation), rs4072111 (missense mutation), and rs1131445 (located in the 3′-UTR).
A better understanding of the connections between cancer and CVD is crucial for identifying and preventing short- and long-term problems in the treatment and prevention of these diseases. In the study of the association of SNPs with diseases, a major analysis tool is meta-analysis. Meta-analysis is the statistical analysis of the evidence from different individual studies, with the aim of integrating them, combining and summarizing their results [37]. Its importance is given by reducing, for example, the standard deviation and the confidence interval, making the result more reliable, in addition to enabling the inclusion of future studies that may be published. The meta-analysis may show an effect that, individually, cannot be observed due to lack of statistical power (limited sample size). Meta-analysis also allows a synthesis of contradictory data even if the statistical power is small [38]. Thus, in this study we performed a systematic review and meta-analysis to evaluate and update information on possible correlations between the polymorphisms rs4778889 T>C, rs11556218 T>G, rs4072111 C>T, and rs1131445 T>C of IL16 and the risk of cancer or CVD.
Results
This systematic review included a total of 6386 cancer subjects and 7395 controls in 19 studies for different types of cancer, as well as 2415 individuals with CVD and 2317 controls in 7 studies. The studied population consisted of Chinese and Iranian individuals. The results are shown in Table 1. SNP rs11556218 of IL16 was statistically associated with the risk of cancer in Chinese (T vs. G; Pooled OR = 1.38; 95% CI 1.23–1.56; random model). Specifically, for gastric cancer in Chinese, the SNP rs4778889 was associated with risk factors for disease development (T vs. C; Pooled OR = 1.18; 95% CI 1.03–1.35; Fixed model). The SNP rs11556218 was also associated with the risk of CVD in Chinese (T vs. G; pooled OR = 1.51; 95% CI 1.07–2.14; random model). Forest plots for all statistically significant comparisons are shown in Supplementary Figure 1. For the selected studies, no statistically significant associations were observed with rs1131445 and rs4072111 and the selected disease groups. For the Iranian population, significance was not found between IL16 polymorphisms and cancer or CVD in this meta-analysis.
Table 1: Meta-analysis results for groups of diseases
| Comparisona | n | Association test | Model | Heterogeneity test | PBEb | Citation | |||
|---|---|---|---|---|---|---|---|---|---|
| OR | CI 95% | τ2 | Phet | I2 | |||||
| Cancer - rs11556218 T>G (total case/control number = 5022/5779) | |||||||||
| T vs G | 14 | 1.38 | (1.23–1.56) | R | 0.04 | <0.01 | 71 | 0.98 | [40, 43, 44, 68, 81–85, 87–89, 93, 98] |
| G/G+T/T vs T/G | 14 | 1.36 | (1.19–1.55) | R | 0.04 | <0.01 | 62 | 0.92 | |
| T/T vs T/G+G/G | 14 | 1.49 | (1.28–1.72) | R | 0.05 | <0.01 | 70 | 0.88 | |
| T/T+T/G vs G/G | 14 | 1.56 | (1.33–1.84) | F | 0.04 | 0.15 | 29 | 0.65 | |
| T/T vs T/G | 14 | 1.43 | (1.24–1.66) | R | 0.05 | <0.01 | 65 | 0.90 | |
| T/T vs G/G | 14 | 1.77 | (1.50–2.10) | F | 0.07 | 0.06 | 41 | 0.93 | |
| T/G vs G/G | 14 | 1.29 | (1.08 – 1.53) | F | 0.00 | 0.72 | 0 | 0.20 | |
| Gastric cancer - rs4778889 T>C (total case/control number = 1046/1310) | |||||||||
| T vs C | 3 | 1.18 | (1.03–1.35) | F | 0.01 | 0.15 | 47 | 0.35 | [68, 81, 93] |
| T/T + T/C vs C/C | 3 | 1.41 | (1.02–1.95) | F | 0.00 | 0.72 | 0 | 0.65 | |
| T/T vs C/C | 3 | 1.48 | (1.06–2.06) | F | 0.00 | 0.61 | 0 | 0.87 | |
| Cardiovascular disease - rs11556218 T>G (total case/control number = 1643/1685) | |||||||||
| T vs G | 5 | 1.51 | (1.07–2.14) | R | 0.17 | <0.01 | 90 | 0.07 | [63, 73, 92, 95, 96] |
| G/G+T/T vs T/G | 5 | 1.87 | (1.08–3.23) | R | 0.43 | <0.01 | 92 | 0.23 | |
| T/T vs T/G+G/G | 5 | 2.00 | (1.11–3.63) | R | 0.52 | <0.01 | 94 | 0.24 | |
| T/T vs T/G | 5 | 2.00 | (1.08–3.71) | R | 0.55 | <0.01 | 94 | 0.24 | |
To avoid a possible bias caused by studies whose genotype frequency distribution in the control groups was not in HWE (Hardy–Weinberg equilibrium), stratified analyses were performed including only studies that met this criterion (P ≥ 0.05), according to the goodness-of-fit test for HWE. These results are shown in Table 2. After these analyses, the IL16 SNP rs11556218 remain associated with the risk of cancer in Chinese (T vs. G; pooled OR = 1.41, 95% CI 1.26–1.59; random model) in the genetic inheritance models described in Table 2. In addition, a subgroup analysis was carried out to separate individuals with gastric and renal cancer from other types of cancer (see Table 2 and Supplementary Figure 2). There was a significant association of rs11556218 with risk for carcinoma and osteosarcoma in all evaluated inheritance models (T vs. G; pooled OR = 1.53; 95% CI 1.34–1.73; random model), while no association was observed with gastric and renal cancer.
Table 2: Meta-analysis results for studies with control groups in the Hardy–Weinberg equilibrium
| Comparisona | n | Association test | Model | Heterogeneity test | PBEb | Citation | |||
|---|---|---|---|---|---|---|---|---|---|
| OR | CI 95% | τ2 | Phet | I2 | |||||
| Cancer - rs11556218 T>G (total case/control number = 4367/5675) | |||||||||
| T vs G | 13 | 1.41 | (1.26–1.59) | R | 0.03 | <0.01 | 68 | 0.99 | [40, 43, 44, 68, 81–84, 87–89, 93, 98] |
| G/G+T/T vs T/G | 13 | 1.41 | (1.23–1.61) | R | 0.04 | <0.01 | 61 | 0.68 | |
| T/T vs T/G+G/G | 13 | 1.54 | (1.33–1.78) | R | 0.05 | <0.01 | 68 | 0.74 | |
| T/T+T/G vs G/G | 13 | 1.66 | (1.39–1.98) | F | 0.04 | 0.21 | 23 | 0.41 | |
| T/T vs T/G | 13 | 1.48 | (1.28–1.71) | R | 0.05 | <0.01 | 64 | 0.69 | |
| T/T vs G/G | 13 | 1.92 | (1.60–2.30) | F | 0.05 | 0.15 | 29 | 0.66 | |
| T/G vs G/G | 13 | 1.31 | (1.09–1.58) | F | 0.00 | 0.62 | 0 | 0.13 | |
| Cancer – Carcinoma and other types - rs11556218 T>G (total case/control number = 3521/4398) | |||||||||
| T vs G | 10 | 1.53 | (1.34–1.73) | R | 0.02 | <0.01 | 62 | 0.63 | [40, 43, 44, 68, 82–84, 88, 89, 98] |
| G/G+T/T vs T/G | 10 | 1.53 | (1.32–1.77) | R | 0.03 | 0.02 | 54 | 0.45 | |
| T/T vs T/G+G/G | 10 | 1.70 | (1.46–1.97) | R | 0.03 | 0.01 | 58 | 0.41 | |
| T/T+T/G vs G/G | 10 | 1.76 | (1.44–2.16) | F | 0.07 | 0.10 | 39 | 0.17 | |
| T/T vs T/G | 10 | 1.63 | (1.40–1.89) | R | 0.03 | 0.02 | 54 | 0.41 | |
| T/T vs G/G | 10 | 2.10 | (1.71–2.58) | F | 0.06 | 0.12 | 35 | 0.06 | |
| T/G vs G/G | 10 | 1.33 | (1.08–1.65) | F | 0.02 | 0.29 | 16 | 0.39 | |
| Cancer – Gastric and renal cancer - rs11556218 T>G (total case/control number = 1154/1515) | |||||||||
| T vs G | 4 | 1.16 | (1.02–1.32) | F | 0.01 | 0.20 | 35 | 0.77 | [68, 81, 85, 87] |
| Cardiovascular disease - rs11556218 T>G (total case/control number = 1405/1507) | |||||||||
| T vs G | 4 | 1.70 | (1.25–2.32) | R | 0.10 | <0.01 | 85 | 0.29 | [63, 92, 95, 96] |
| G/G+T/T vs T/G | 4 | 2.03 | (1.09–3.78) | R | 0.47 | <0.01 | 93 | 0.33 | |
| T/T vs T/G+G/G | 4 | 2.33 | (1.24–4.41) | R | 0.49 | <0.01 | 94 | 0.49 | |
| T/T+T/G vs G/G | 4 | 1.77 | (1.24–2.53) | F | 0.18 | 0.10 | 49 | 0.09 | |
| T/T vs T/G | 4 | 2.25 | (1.12–4.50) | R | 0.59 | <0.01 | 94 | 0.58 | |
| T/T vs G/G | 4 | 2.50 | (1.19–5.25) | R | 0.50 | <0.01 | 72 | 0.39 | |
An association of IL16 rs11556218 with the risk for CVD in Chinese was also found for different genetic inheritance models (T vs. G; pooled OR = 1.70; 95% CI 1.25–2.32; random model). An analysis of the covariates evaluated by CVD studies was carried out with the aim of finding a possible source of heterogeneity for the obtained results. These assessed covariates are described in Supplementary Table 2. In this sense, it was not possible to identify in this analysis which covariables could partially explain the qualitative heterogeneity observed in this CVD analysis.
The statistical power after HWE stratification results, considering a subtle genetic effect (1.25), showed a slight reduction for the association between rs11556218 and cancer (from 99.9% to 93.0% after stratification for HWE) but a substantial reduction for rs11556218 and CVD (90.7% to 81.0%) and for rs4778889 and gastric cancer (72.0% to 54.3%). Forest plots for all statistically significant comparisons after stratification of studies with controls in HWE are shown in Supplementary Figure 2.
The mean minor allele frequencies (MAF) observed among the control groups of the selected studies were 21.5% for rs4778889 (C allele), 21.3% for rs11556218 (G allele), 21.5% for rs4072111 (T allele), and 32.3% for rs1131445 (C allele). These frequencies were similar to those described in the 1000Genomes database for East Asian populations (21.2%, 16.5%, 20.8%, and 31.2%, respectively), and the Fisher test did not indicate a statistically significant difference between the allele frequencies in these populations (P > 0.05).
Egger’s tests did not indicate possible bias in the selection of publications (P ≥ 0.05), and these results are shown in Tables 2 and 3. The funnel plot for visual assessment is available in Supplementary Figure 3. It is noteworthy that the bias in funnel plots should be interpreted carefully for a meta-analysis with a small number of studies (around 10 or less), since tests in a small number of studies have limited statistical power due to the great heterogeneity of these datasets [39]. The quality assessment of the studies selected by the NOS (Table 3) indicated a 95% score of the studies as reasonable or good (greater than 5).
Table 3: Newcastle–ottawa quality assessment scale for selected studies
| Study | Selection | Comparabilitya | Exposure | Total score | |||||
|---|---|---|---|---|---|---|---|---|---|
| Shih et al.[43] | * | * | * | * | ** | * | * | 8 | |
| Wu et al. [44] | * | * | * | * | ** | * | * | 8 | |
| He et al. [81] | * | * | * | * | ** | * | * | * | 9 |
| Li et al. [82] | * | * | * | * | ** | * | * | 8 | |
| Yang et al. [73] | * | * | * | * | * | 5 | |||
| MaiMaiTiMin et al. [83] | * | * | * | * | ** | * | * | 8 | |
| Tang et al. [84] | * | * | * | * | ** | * | * | 8 | |
| Yang et al. [85] | * | * | * | ** | * | * | 7 | ||
| Yao et al. [86] | * | * | * | ** | * | * | 7 | ||
| Kashfi et al. [67] | * | * | ** | * | * | 6 | |||
| Wang and Zhu [87] | * | * | * | * | ** | * | * | 8 | |
| Luo et al. [88] | * | * | * | * | ** | * | * | 8 | |
| Qin et al. [89] | * | * | * | * | ** | * | * | 8 | |
| Hai-Feng et al. [90] | * | * | * | * | ** | * | * | * | 9 |
| Huang et al. [91] | * | * | * | * | ** | * | * | 8 | |
| Liu et al. [63] | * | * | * | ** | * | * | 7 | ||
| Tong et al. [92] | * | * | * | * | ** | * | * | 8 | |
| Zhang and Wang [93] | * | * | * | * | ** | * | * | * | 9 |
| Azimzadeh et al. [72] | * | * | * | * | ** | * | * | 8 | |
| Azimzadeh et al. [94] | * | * | * | * | ** | * | * | 8 | |
| Chen et al. [95]* | * | * | * | * | ** | * | * | 8 | |
| Li et al. [40] | * | * | * | ** | * | * | 7 | ||
| Wu et al. [96] | * | * | ** | * | * | 6 | |||
| Zhu et al. [97] | * | * | * | * | ** | * | * | 8 | |
| Gao et al. [98] | * | * | * | * | * | * | 6 | ||
| Gao et al. [68] | * | * | * | * | ** | * | * | 8 | |
DISCUSSION
This systematic review included a total of 6386 individuals with cancer and 2415 individuals with CVD, which confers high statistical power to verify weaker associations. The IL16 rs11556218 polymorphism was significantly associated with the risk of cancer in Chinese, in different models of genetic inheritance. In addition, to the best of our knowledge, this is the first meta-analysis to show an association between the rs477889 polymorphism and the risk of gastric cancer and the first to demonstrate an association of rs11556218 and the risk of CVD in Chinese.
The association between IL16 rs11556218 and cancer or CVD remained even after careful analysis of studies control groups according to HWE criteria, and perhaps the loss of association observed for rs4778889 and gastric cancer after stratification was due to the significant loss of statistical power when the sample size of the combined studies was reduced. We also see different results when different types of cancer are grouped together. In our meta-analysis, the subgroup of renal and gastric cancer was not associated with rs11556218. These different results in distinct subgroups can be explained by SNPs having different behaviors in different types of cancer [40]. In addition, sample size limitations should be considered in analyzes of smaller number of studies. Finally, selection bias should not be ruled out in the analysis of different studies.
In agreement with our results, two other meta-analyses published in 2014 did not find a significant association between rs4778889 and rs4072111 of IL16 and cancer. However, these same studies observed a statistically significant association between rs11556218 and cancer, an association observed in our study with the addition of new studies published after these meta-analyses results [41–44]. The SNP rs11556218, consisting of a substitution of the T nucleotide by a G, is located in the exon region of the gene and results in a modification of the pro-IL-16 PDZ2 domain (IL-16 isoform with 631 amino acids) and npro-IL-16 (neuronal IL-16 isoform with 1331 amino acids). This modification consists of a substitution of asparagine for lysine in the protein at positions 446 (pro-IL-16) and 1147 (npro-IL-16), as illustrated in Figure 1. This mutation may result in structural changes in the protein, consequently affecting its function in the immune response [45]. This SNP was also associated with prostate cancer in a replication GWAS study in African Americans [46]. It has also been associated with asparaginase-related thrombosis and pancreatitis in the treatment of acute lymphoid leukemia in Caucasians [47], diabetes mellitus [48], autoimmune diseases [49, 50], endometriosis [45, 51], and knee osteoarthritis [52, 53].
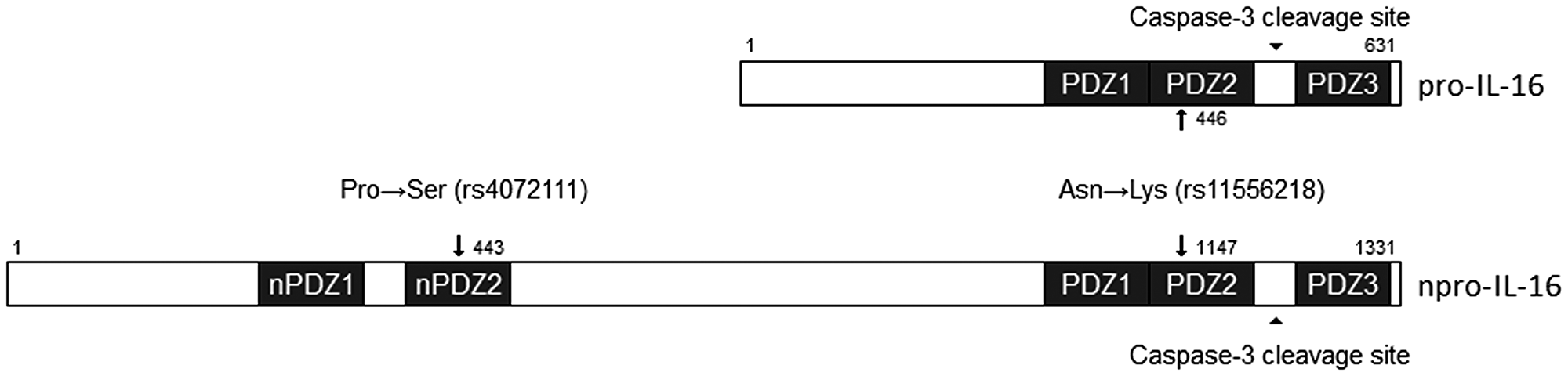
Figure 1: Schematic representation of two SNPs changes in PDZ domains of pro-IL-16 and npro-IL-16. The rs11556218 polymorphism alters the PDZ2 domain at positions 446 and 1147 of pro-IL-16 and n-pro-IL-16, respectively. This polymorphism will cause a substitution of an asparagine to a lysine at the protein, affecting protein recognition by its receptor. The rs4072111 polymorphism is present in the nPDZ2 domain of npro-IL-16 and will cause a substitution of a proline to serine in the protein precursor. Adapted from Bannert et al. [17].
A meta-analysis published in 2019 focusing only on renal cancer association studies also found no association between rs4778889 and rs11556218 and this disease [54]. A possible explanation for this lack of association may be related to the limited statistical power to verify subtle associations, due to the small number of individuals in the evaluated studies (total of 790 individuals). Another meta-analysis, published in 2014, found no association between rs4778889 and cancer, which was confirmed in our study for the general cancer group [55]. However, we observed, for the first time, a statistically significant association of this polymorphism with gastric cancer in Chinese. This SNP in the promoter region of the IL16 gene was related to changes in expression [56]. It was previously associated with precancerous gastric lesions [57], gestational diabetes [58], autoimmune diseases [50, 59], knee osteoarthritis [53], endometriosis [60], Crohn’s disease [61], and allergic contact dermatitis [62].
The SNP rs4072111, consisting of a substitution of the C nucleotide by a T, is located in an intronic region and results in a modification of the nPro-IL-16 PDZ2 domain, with a serine to proline exchange at position 434 [45]. In our study, we found no association of this polymorphism with cancer. Moreover, it was not possible to perform a meta-analysis of the association between rs4072111 and CVD due to the limited number of studies, and no association was observed in the study of Liu et al. [63]. This SNP was previously identified as a marker candidate for aggressive prostate cancer [64], although this was not confirmed for an African American population [46, 65]. This polymorphism was also correlated with better prognosis for chronic lymphoid leukemia in an English population [66], associated with the risk of liver cancer [40], colorectal and gastric cancer [67, 68], autoimmunity [49, 50], endometriosis [51], and knee osteoarthritis [52, 53].
The polymorphism rs1131445 (T>C) is located in the 3′-UTR of the gene, and the mutation predicts disruption of the binding of a microRNA inhibitor (miR135b-mRNA) of the gene sequence, which would result in greater pro-IL-16 expression [69]. In our study, we did not observe any association of this SNP with cancer or CVD. Studies have found that this SNP has been associated with cervical cancer in Chinese and prostate cancer in African Americans [46, 70]. Its mutant allele may also be informative about the time to prostate cancer diagnosis among African Americans [71]. It was also associated with gastric and colorectal cancer in Iranians [67, 72], CVD in Chinese [73], Graves’ disease in Chinese [59], and endometriosis in Iranian women [51].
Important limitations should be considered in our study. First, every systematic review presents some risk of bias, related to the difficulty of obtaining non-indexed studies, and the heterogeneity present in the selected studies. Moreover, the results obtained are limited to the populations surveyed and, therefore, lack information for a meta-analysis on the frequency distributions of these polymorphisms in other populations, although an attempt to broadly describe the literature has been made. Similarly, important covariates must be considered to explain the associations observed in these studies, thus multivariate methods of analysis can be useful, highlighting the need for detailed publication of the profile of the populations studied, although the reliability of these methods depends on 10 or more studies. From the results of this review, further studies may be necessary to clarify the role of these polymorphisms in the immunopathogenesis of cancer and CVD, also helping to clarify the relationship between these diseases.
The SNP rs11556218 of IL16 was significantly associated with an increased risk of cancer or CVD in Chinese. Additionally, the SNP rs4778889 was associated with an increased risk of gastric cancer, while the rs4072111 and rs1131445 SNPs were not associated with cancer or CVD in our study.
Materials and Methods
The methodology of this study was performed according to the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) statement [74]. To identify studies investigating associations between IL16 polymorphisms and cancer or CVD, the MEDLINE, Web of Science and Scopus databases were searched up to June 30, 2020, according to the strategy detailed in Supplementary Table 1. The language of publication was not defined as a search criterion, but only articles in the English language were included. All steps were performed by two pairs of reviewers in duplicate. Article screening procedures are included in Figure 2.
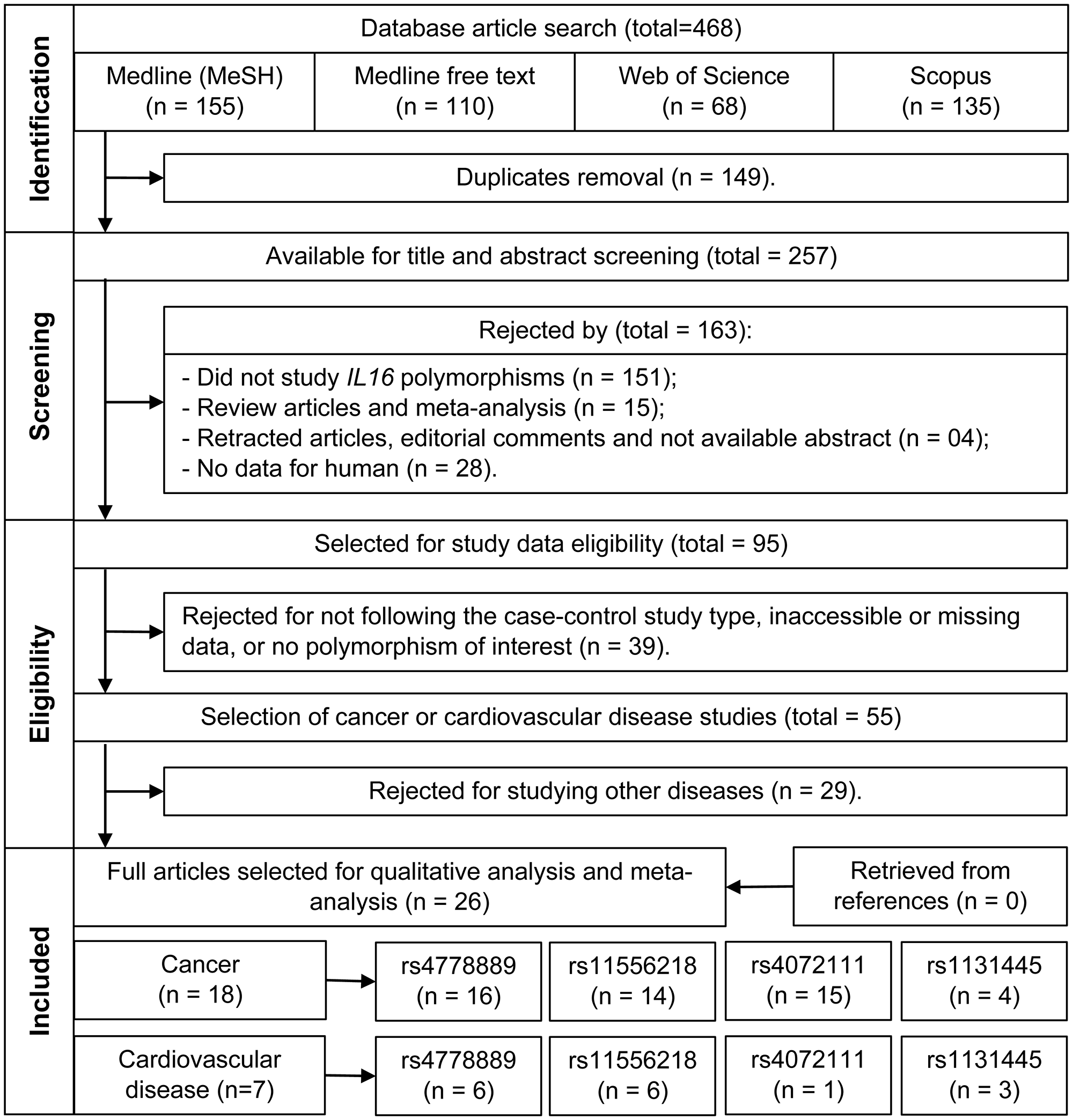
Figure 2: Flowchart of the studies selection.
Articles with available abstracts were selected for title and abstract screening as to their relevance and fitness to the proposed theme. Only original human studies of IL16 polymorphisms were included. In the next step, the 95 selected articles were evaluated by reading the full text, as to their suitability for inclusion in a meta-analysis (case-control study, evaluating at least one of the polymorphisms of interest, availability of genotype frequency data). Thus, 55 articles were obtained that met all these criteria. All doubts regarding the eligibility of a study or not (according to the criteria described above) were discussed among all authors and resolved after consensus. After a consensus was reached between two authors working in pairs, 26 studies were selected for data collection, according to the diseases of interest. Also, the references of these studies were searched for additional studies. The studies and SNPs selected in this review are described in Supplementary Table 3. The 26 selected studies were further reassessed by a third independent pair of authors, composed of authors with more experience in the field of cytokine polymorphisms and association studies, evaluating whether all the studies and results were in accordance with the proposed criteria for this review.
Genotype frequency data were collected from the selected studies. Allele frequencies were obtained by counting the total number of alleles in all genotypes, and the number of individuals used in the meta-analysis was obtained by summing the total number of described genotypes. Distinct groups of cases or controls within the same study were combined into a single group for distinct and unrelated groups of individuals. The cancer group was analyzed by combining all cancer types or individually for each type of cancer (when a comparison was possible). Statistical analyses were performed by using the program R, version 3.5.2, with the meta-analysis package “meta” [75].
The meta-analysis results are represented by the Forest plot. This chart aims to summarize the results of the pooled studies giving them a general trend. It represents the studies with the proportions of events of interest in each study for the case-control groups. The percentage values of weight and effect size (represented by the Odds Ratio, OR) of each study in the final result are presented in two possible models according to observed heterogeneity: fixed and random. Thus, for all comparisons performed, Forest plots were constructed with a heterogeneity test (I2). When the value of I2 was greater than 50% and the test showed statistically significant results, the random OR model was chosen. When I2 was less than 50% with no evidence of significant heterogeneity, the fixed model was chosen. Six comparison groups of alleles and genotypes were used to verify the influence of genetic association with diseases. Considering “M” as the wild-type allele and “m” as the mutated allele, the combinations analyzed were (1) M vs. m alleles; (2) MM vs. Mm genotypes; (3) MM vs. mm and (4) Mm vs. mm; and also genetic inheritance models (5) MM + Mm vs. mm (recessive); (6) MM vs. Mm + mm (dominant); and (7) MM + mm vs. Mm (overdominant).
Quality evaluation of the selected studies was performed according to the Newcastle–Ottawa (NOS) scale by two independent authors. Disagreements were resolved by consensus, and studies with a score higher than 5 were considered as good or reasonable [76]. Possible biases of published meta-analyses were verified by Egger’s test analysis [77], funnel plot visual evaluation, and Begg’s test [78]. All individual study results were expressed as ORs with a 95% confidence interval (CI). The Hardy–Weinberg equilibrium (HWE) of the control group of each study was checked according to the chi-square test for goodness-of-fit (P < 0.05), as recommended for assessing study quality [79]. After this, statistical power analysis was performed using the QUANTO software [80], to evaluate sample size effects on the statistical power. The reference population allele frequencies used for comparison were obtained from the 1000 Genomes Project (phase3 release V3+, available at: https://www.internationalgenome.org/) and the comparison was made using the exact Fisher test in R. For all tests, P < 0.05 was considered to indicate statistical significance.
Abbreviations
GLOBOCAN: Global cancer statistics; CVD: Cardiovascular disease; IFN-α: Interferon-α; IL: Interleukin; GM-GSF: Granulocyte macrophage colony stimulating factor; SNP: Single nucleotide polymorphism; OR: Odds Ratio; CI: Confidence interval; HWE: Hardy–Weinberg equilibrium; NOS: Newcastle–Ottawa Scale; GWAS: Genome-wide association study; PRISMA: Preferred Reporting Items for Systematic Reviews and Meta-Analyses.
ACKNOWLEDGMENTS
We thank Professor Jorge Juarez Vieira Teixeira for his teaching in the area of systematic review in the PRISMA statement and meta-analysis.
CONFLICTS OF INTEREST
Authors have no conflicts of interest to declare.
FUNDING
This study was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Fundação Araucária do Estado do Paraná (#2015/0310), and Laboratory of Immunogenetics at State University of Maringá (#1589/2017-CSD-UEM). The authors declare that there is no conflict of interest regarding the publication of this article.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492. [PubMed].
2. Miller KD, Nogueira L, Mariotto AB, Rowland JH, Yabroff KR, Alfano CM, Jemal A, Kramer JL, Siegel RL. Cancer treatment and survivorship statistics, 2019. CA Cancer J Clin. 2019; 69:363–85. https://doi.org/10.3322/caac.21565. [PubMed].
3. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66:115–32. https://doi.org/10.3322/caac.21338. [PubMed].
4. Roth GA, Johnson C, Abajobir A, Abd-Allah F, Abera SF, Abyu G, Ahmed M, Aksut B, Alam T, Alam K, Alla F, Alvis-Guzman N, Amrock S, et al. Global, Regional, and National Burden of Cardiovascular Diseases for 10 Causes, 1990 to 2015. J Am Coll Cardiol. 2017; 70:1–25. https://doi.org/10.1016/j.jacc.2017.04.052. [PubMed].
5. Koene RJ, Prizment AE, Blaes A, Konety SH. Shared Risk Factors in Cardiovascular Disease and Cancer. Circulation. 2016; 133:1104–14. https://doi.org/10.1161/CIRCULATIONAHA.115.020406. [PubMed].
6. Armenian SH, Lacchetti C, Barac A, Carver J, Constine LS, Denduluri N, Dent S, Douglas PS, Durand JB, Ewer M, Fabian C, Hudson M, Jessup M, et al. Prevention and Monitoring of Cardiac Dysfunction in Survivors of Adult Cancers: American Society of Clinical Oncology Clinical Practice Guideline. J Clin Oncol. 2017; 35:893–911. https://doi.org/10.1200/JCO.2016.70.5400. [PubMed].
7. Armenian SH, Xu L, Ky B, Sun C, Farol LT, Pal SK, Douglas PS, Bhatia S, Chao C. Cardiovascular Disease Among Survivors of Adult-Onset Cancer: A Community-Based Retrospective Cohort Study. J Clin Oncol. 2016; 34:1122–30. https://doi.org/10.1200/JCO.2015.64.0409. [PubMed].
8. Mehta LS, Watson KE, Barac A, Beckie TM, Bittner V, Cruz-Flores S, Dent S, Kondapalli L, Ky B, Okwuosa T, Piña IL, Volgman AS, and American Heart Association Cardiovascular Disease in Women and Special Populations Committee of the Council on Clinical Cardiology; Council on Cardiovascular and Stroke Nursing; and Council on Quality of Care and Outcomes Research. Cardiovascular Disease and Breast Cancer: Where These Entities Intersect: A Scientific Statement From the American Heart Association. Circulation. 2018; 137:e30–66. https://doi.org/10.1161/CIR.0000000000000556. [PubMed].
9. Curigliano G, Cardinale D, Suter T, Plataniotis G, de Azambuja E, Sandri MT, Criscitiello C, Goldhirsch A, Cipolla C, Roila F, and ESMO Guidelines Working Group. Cardiovascular toxicity induced by chemotherapy, targeted agents and radiotherapy: ESMO Clinical Practice Guidelines. Ann Oncol. 2012; 23:vii155–66. https://doi.org/10.1093/annonc/mds293. [PubMed].
10. Libby P. Inflammation in atherosclerosis. Nature. 2002; 420:868–74. https://doi.org/10.1038/nature01323. [PubMed].
11. Coussens LM, Werb Z. Inflammation and cancer. Nature. 2002; 420:860–7. https://doi.org/10.1038/nature01322. [PubMed].
12. Waldmann TA. Cytokines in Cancer Immunotherapy. Cold Spring Harb Perspect Biol. 2018; 10:a028472. https://doi.org/10.1101/cshperspect.a028472. [PubMed].
13. Zupančič E, Fayad ZA, Mulder WJM. Cardiovascular Immunotherapy and the Role of Imaging. Arterioscler Thromb Vasc Biol. 2017; 37:e167–71. https://doi.org/10.1161/ATVBAHA.117.309227. [PubMed].
14. Center DM, Kornfeld H, Cruikshank WW. Interleukin-16. Int J Biochem Cell Biol. 1997; 29:1231–4. https://doi.org/10.1016/s1357-2725(97)00053-8. [PubMed].
15. Center DM, Kornfeld H, Ryan TC, Cruikshank WW. Interleukin 16: implications for CD4 functions and HIV-1 progression. Immunol Today. 2000; 21:273–80.
16. Zhang Y, Center DM, Wu DMH, Cruikshank WW, Yuan J, Andrews DW, Kornfeld H. Processing and Activation of Pro-Interleukin-16 by Caspase-3. J Biol Chem. 1998; 273:1144–9. https://doi.org/10.1074/jbc.273.2.1144. [PubMed].
17. Bannert N, Vollhardt K, Asomuddinov B, Haag M, König H, Norley S, Kurth R. PDZ Domain-mediated Interaction of Interleukin-16 Precursor Proteins with Myosin Phosphatase Targeting Subunits. J Biol Chem. 2003; 278:42190–9. https://doi.org/10.1074/jbc.M306669200. [PubMed].
18. Richmond J, Tuzova M, Cruikshank W, Center D. Regulation of Cellular Processes by Interleukin-16 in Homeostasis and Cancer. J Cell Physiol. 2014; 229:139–47. https://doi.org/10.1002/jcp.24441. [PubMed].
19. Richmond J, Tuzova M, Parks A, Adams N, Martin E, Tawa M, Morrison L, Chaney K, Kupper TS, Curiel-Lewandrowski C, Cruikshank W. Interleukin-16 as a marker of Sézary syndrome onset and stage. J Clin Immunol. 2011; 31:39–50. https://doi.org/10.1007/s10875-010-9464-8. [PubMed].
20. Asadullah K, Haeussler-Quade A, Gellrich S, Hanneken S, Hansen-Hagge TE, Döcke WD, Volk HD, Sterry W. IL-15 and IL-16 overexpression in cutaneous T-cell lymphomas: stage-dependent increase in mycosis fungoides progression. Exp Dermatol. 2000; 9:248–51. https://doi.org/10.1034/j.1600-0625.2000.009004248.x. [PubMed].
21. Sharma V, Sparks JL, Vail JD. Human B-cell lines constitutively express and secrete interleukin-16. Immunology. 2000; 99:266–71. https://doi.org/10.1046/j.1365-2567.2000.00959.x. [PubMed].
22. Kovacs E. The serum levels of IL-12 and IL-16 in cancer patients. Relation to the tumour stage and previous therapy. Biomed Pharmacother. 2001; 55:111–6. [PubMed].
23. Koike M, Sekigawa I, Okada M, Matsumoto M, Iida N, Hashimoto H, Oshimi K. Relationship between CD4(+)/CD8(+) T cell ratio and T cell activation in multiple myeloma: reference to IL-16. Leuk Res. 2002; 26:705–11. https://doi.org/10.1016/s0145-2126(01)00192-8. [PubMed].
24. Alexandrakis MG, Passam FH, Kyriakou DS, Christophoridou AV, Perisinakis K, Hatzivasili A, Foudoulakis A, Castanas E. Serum level of interleukin-16 in multiple myeloma patients and its relationship to disease activity. Am J Hematol. 2004; 75:101–6. https://doi.org/10.1002/ajh.10444. [PubMed].
25. Templin J, Atanackovic D, Hasche D, Radhakrishnan SV, Luetkens T. Oscillating expression of interleukin-16 in multiple myeloma is associated with proliferation, clonogenic growth, and PI3K/NFKB/MAPK activation. Oncotarget. 2017; 8:49253–63. https://doi.org/10.18632/oncotarget.17534. [PubMed].
26. Yellapa A, Bitterman P, Sharma S, Guirguis AS, Bahr JM, Basu S, Abramowicz JS, Barua A. Interleukin 16 expression changes in association with ovarian malignant transformation. Am J Obstet Gynecol. 2014; 210:272.e1–272.e10. https://doi.org/10.1016/j.ajog.2013.12.041. [PubMed].
27. Blaschke V, Reich K, Middel P, Letschert M, Sachse F, Harwix S, Neumann C. Expression of the CD4+ Cell-Specific Chemoattractant Interleukin-16 in Mycosis Fungoides. J Invest Dermatol. 1999; 113:658–63. https://doi.org/10.1046/j.1523-1747.1999.00717.x. [PubMed].
28. Grönberg C, Bengtsson E, Fredrikson GN, Nitulescu M, Asciutto G, Persson A, Andersson L, Nilsson J, Gonçalves I, Björkbacka H. Human Carotid Plaques With High Levels of Interleukin-16 Are Associated With Reduced Risk for Cardiovascular Events. Stroke. 2015; 46:2748–54. https://doi.org/10.1161/STROKEAHA.115.009910. [PubMed].
29. Grönberg C, Asciutto G, Persson A, Fredrikson GN, Nilsson J, Gonçalves I, Björkbacka H. Endarterectomy patients with elevated levels of circulating IL-16 have fewer cardiovascular events during follow-up. Cytokine. 2016; 85:137–9. https://doi.org/10.1016/j.cyto.2016.06.009. [PubMed].
30. McFadden C, Morgan R, Rahangdale S, Green D, Yamasaki H, Center D, Cruikshank W. Preferential migration of T regulatory cells induced by IL-16. J Immunol. 2007; 179:6439–45. https://doi.org/10.4049/jimmunol.179.10.6439. [PubMed].
31. Tamaki S, Mano T, Sakata Y, Ohtani T, Takeda Y, Kamimura D, Omori Y, Tsukamoto Y, Ikeya Y, Kawai M, Kumanogoh A, Hagihara K, Ishii R, et al. Interleukin-16 promotes cardiac fibrosis and myocardial stiffening in heart failure with preserved ejection fraction. PLoS One. 2013; 8:e68893. https://doi.org/10.1371/journal.pone.0068893. [PubMed].
32. Mathy NL, Scheuer W, Lanzendörfer M, Honold K, Ambrosius D, Norley S, Kurth R. Interleukin-16 stimulates the expression and production of pro-inflammatory cytokines by human monocytes. Immunology. 2000; 100:63–9. https://doi.org/10.1046/j.1365-2567.2000.00997.x. [PubMed].
33. Schernthaner C, Paar V, Wernly B, Pistulli R, Rohm I, Jung C, Figulla HR, Yilmaz A, Cadamuro J, Haschke-Becher E, Schulze PC, Hoppe UC, Lichtenauer M, Kretzschmar D. Elevated plasma levels of interleukin-16 in patients with acute myocardial infarction. Medicine (Baltimore). 2017; 96:e8396. https://doi.org/10.1097/MD.0000000000008396. [PubMed].
34. Park SL, Hwang B, Lee SY, Kim WT, Choi YH, Chang YC, Kim WJ, Moon SK. p21WAF1 Is Required for Interleukin-16-Induced Migration and Invasion of Vascular Smooth Muscle Cells via the p38MAPK/Sp-1/MMP-9 Pathway. PLoS One. 2015; 10:e0142153. https://doi.org/10.1371/journal.pone.0142153. [PubMed].
35. Christiansen MK, Larsen SB, Nyegaard M, Neergaard-Petersen S, Ajjan R, Würtz M, Grove EL, Hvas AM, Jensen HK, Kristensen SD. Coronary artery disease-associated genetic variants and biomarkers of inflammation. PLoS One. 2017; 12:e0180365. https://doi.org/10.1371/journal.pone.0180365. [PubMed].
36. Deng N, Zhou H, Fan H, Yuan Y. Single nucleotide polymorphisms and cancer susceptibility. Oncotarget. 2017; 8:110635–49. https://doi.org/10.18632/oncotarget.22372. [PubMed].
37. Haidich AB. Meta-analysis in medical research. Hippokratia. 2010; 14:29–37. https://doi.org/10.5005/jp/books/10519.
38. Lee YH. An overview of meta-analysis for clinicians. Korean J Intern Med. 2018; 33:277–283. https://doi.org/10.3904/kjim.2016.195. [PubMed].
39. Sterne JAC, Sutton AJ, Ioannidis JPA, Terrin N, Jones DR, Lau J, Carpenter J, Rucker G, Harbord RM, Schmid CH, Tetzlaff J, Deeks JJ, Peters J, et al. Recommendations for examining and interpreting funnel plot asymmetry in meta-analyses of randomised controlled trials. BMJ. 2011; 343:d4002. https://doi.org/10.1136/bmj.d4002. [PubMed].
40. Li S, Deng Y, Chen ZP, Huang S, Liao XC, Lin LW, Li H, Peng T, Qin X, Zhao J. Genetic polymorphism of interleukin-16 influences susceptibility to HBV-related hepatocellular carcinoma in a Chinese population. Infect Genet Evol. 2011; 11:2083–8. https://doi.org/10.1016/j.meegid.2011.09.025. [PubMed].
41. Mo CJ, Peng QL, He Y, Wang J, Xie L, Li TJ, Li S, Qin X. Positive association between IL-16 rs11556218 T/G polymorphism and cancer risk: a meta-analysis. Asian Pac J Cancer Prev. 2014; 15:4697–703. https://doi.org/10.7314/apjcp.2014.15.11.4697. [PubMed].
42. Zhao Y, Tao L, Wang B, Nie P, Tang Y, Zhu M. Interleukin-16 Gene Polymorphisms rs4778889, rs4072111, rs11556218, and Cancer Risk in Asian Populations: A Meta-Analysis. Genet Test Mol Biomarkers. 2014; 18:174–82. https://doi.org/10.1089/gtmb.2013.0386. [PubMed].
43. Shih LC, Chang WS, Lee HT, Wang YC, Wang ZH, Chao CY, Yu CC, Lin HY, Shen TC, Kuo CC, Tsai CW, Bau DT. Interaction of Interleukin-16 Genotypes With Betel Quid Chewing Behavior on Oral Cancer in Taiwan. In Vivo. 2020; 34:1759–64. https://doi.org/10.21873/invivo.11969. [PubMed].
44. Wu MF, Wang YC, Shen TC, Chang WS, Li HT, Liao CH, Gong CL, Wang ZH, Tsai CW, Hsia TC, Bau DT. Significant association of interleukin-16 genetic variations to taiwanese lung cancer. In Vivo (Brooklyn). 2020; 34:1117–23. https://doi.org/10.21873/invivo.11883. [PubMed].
45. Matalliotakis M, Zervou MI, Eliopoulos E, Matalliotaki C, Rahmioglu N, Kalogiannidis I, Zondervan K, Spandidos DA, Matalliotakis I, Goulielmos GN. The role of IL-16 gene polymorphisms in endometriosis. Int J Mol Med. 2018; 41:1469–76. https://doi.org/10.3892/ijmm.2018.3368. [PubMed].
46. Batai K, Shah E, Murphy AB, Newsome J, Ruden M, Ahaghotu C, Kittles RA. Fine-Mapping of IL16 Gene and Prostate Cancer Risk in African Americans. Cancer Epidemiol Biomarkers Prev. 2012; 21:2059–68. https://doi.org/10.1158/1055-9965.EPI-12-0707. [PubMed].
47. Abaji R, Gagné V, Xu CJ, Spinella JF, Ceppi F, Laverdière C, Leclerc JM, Sallan SE, Neuberg D, Kutok JL, Silverman LB, Sinnett D, Krajinovic M. Whole-exome sequencing identified genetic risk factors for asparaginase-related complications in childhood ALL patients. Oncotarget. 2017; 8:43752–67. https://doi.org/10.18632/oncotarget.17959. [PubMed].
48. Cheng F, Liu L, Zhang H, Zhu Y, Li X, Li H. Association of IL-16 gene polymorphisms with the risk of developing type 2 diabetes mellitus in the Chinese Han population. Biosci Rep. 2019; 39:BSR20190821. https://doi.org/10.1042/BSR20190821. [PubMed].
49. Farrokhi M, Masoudifar A, Derakhshan A, Saadatmand S, Rouhi-Boroujeni H, Etemadifar M, Rezaei-Zarji S, Javid A, Nobakht R, Deyhimi M, Ekramnia A, Ebrahimi M, Sheikh S, et al. The Association of Interleukin-16 Gene Polymorphisms with IL-16 Serum Levels and Risk of Multiple Sclerosis. Immunol Invest. 2017. [Epub ahead of print]. https://doi.org/10.1080/08820139.2016.1268154. [PubMed].
50. Xue H, Gao L, Wu Y, Fang W, Wang L, Li C, Li Y, Liang W, Zhang L. The IL-16 gene polymorphisms and the risk of the systemic lupus erythematosus. Clin Chim Acta. 2009; 403:223–5. https://doi.org/10.1016/j.cca.2009.03.016. [PubMed].
51. Azimzadeh P, Khorram Khorshid HR, Akhondi MM, Shirazi A. Association of interleukin-16 polymorphisms with disease progression and susceptibility in endometriosis. Int J Immunogenet. 2016; 43:297–302. https://doi.org/10.1111/iji.12281. [PubMed].
52. Liu Z, Ma L, Qiu S, Jia T. Genetic polymorphisms of interleukin-16 are associated with susceptibility to primary knee osteoarthritis. Int J Clin Exp Med. 2015; 8:1401–5. [PubMed].
53. Luo SX, Li S, Zhang XH, Zhang JJ, Long GH, Dong GF, Su W, Deng Y, Liu Y, Zhao JM, Qin X. Genetic polymorphisms of interleukin-16 and risk of knee osteoarthritis. PLoS One. 2015; 10:e0123442. https://doi.org/10.1371/journal.pone.0123442. [PubMed].
54. Zhou T, Li H, Xie WJ, Zhong Z, Zhong H, Lin ZJ. Association of Methylenetetrahydrofolate Reductase, Vitamin D Receptor, and Interleukin-16 Gene Polymorphisms With Renal Cell Carcinoma Risk. Technol Cancer Res Treat. 2019; 18:1533033819859413. https://doi.org/10.1177/1533033819859413. [PubMed].
55. Xu LL, Song ZC, Shang K, Zhao LQ, Zhu ZS. Non-association of IL-16 rs4778889 T/C polymorphism with cancer risk in Asians: a meta-analysis. Asian Pac J Cancer Prev. 2014; 15:803–5. https://doi.org/10.7314/apjcp.2014.15.2.803. [PubMed].
56. Nakayama E, Wasi C, Ajisawa A, Iwamoto A, Shioda T. A new polymorphism in the promoter region of the human interleukin-16 (IL-16) gene. Genes Immun. 2000; 1:293–4. https://doi.org/10.1038/sj.gene.6363672. [PubMed].
57. Wang YM, Li ZX, Tang FB, Zhang Y, Zhou T, Zhang L, Ma JL, You WC, Pan KF. Association of genetic polymorphisms of interleukins with gastric cancer and precancerous gastric lesions in a high-risk Chinese population. Tumour Biol. 2016; 37:2233–42. https://doi.org/10.1007/s13277-015-4022-x. [PubMed].
58. Tarnowski M, Wieczorek A, Dziedziejko V, Safranow K, Ustianowski P, Celewicz Z, Pawlik A. IL16 and IL18 gene polymorphisms in women with gestational diabetes. Ginekol Pol. 2017; 88:249–54. https://doi.org/10.5603/GP.a2017.0047. [PubMed].
59. Gu XJ, Cui B, Zhao ZF, Chen HY, Li XY, Wang S, Ning G, Zhao YJ. Association of the interleukin (IL)-16 gene polymorphisms with Graves’ disease. Clin Immunol. 2008; 127:298–302. https://doi.org/10.1016/j.clim.2008.01.017. [PubMed].
60. Gan XL, Lin YH, Zhang Y, Yu TH, Hu LN. Association of an Interleukin-16 Gene Polymorphism with the Risk and Pain Phenotype of Endometriosis. DNA Cell Biol. 2010; 29:663–7. https://doi.org/10.1089/dna.2010.1049. [PubMed].
61. Glas J, Török HP, Unterhuber H, Radlmayr M, Folwaczny C. The -295T-to-C promoter polymorphism of the IL-16 gene is associated with Crohn’s disease. Clin Immunol. 2003; 106:197–200. https://doi.org/10.1016/s1521-6616(03)00021-4. [PubMed].
62. Reich K, Westphal G, König IR, Mössner R, Krüger U, Ziegler A, Neumann C, Schnuch A. Association of allergic contact dermatitis with a promoter polymorphism in the IL16 gene. J Allergy Clin Immunol. 2003; 112:1191–4. https://doi.org/10.1016/j.jaci.2003.09.012. [PubMed].
63. Liu X, Du J, Zhou Y, Shu Q, Li Y. Interleukin-16 Polymorphism Is Associated with an Increased Risk of Ischemic Stroke. Mediators Inflamm. 2013; 2013:564750. https://doi.org/10.1155/2013/564750. [PubMed].
64. Thomas G, Jacobs KB, Yeager M, Kraft P, Wacholder S, Orr N, Yu K, Chatterjee N, Welch R, Hutchinson A, Crenshaw A, Cancel-Tassin G, Staats BJ, et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat Genet. 2008; 40:310–5. https://doi.org/10.1038/ng.91. [PubMed].
65. Chang BL, Spangler E, Gallagher S, Haiman CA, Henderson B, Isaacs W, Benford ML, Kidd LR, Cooney K, Strom S, Ingles SA, Stern MC, Corral R, et al. Validation of Genome-Wide Prostate Cancer Associations in Men of African Descent. Cancer Epidemiol Biomarkers Prev. 2011; 20:23–32. https://doi.org/10.1158/1055-9965.EPI-10-0698. [PubMed].
66. Sellick GS, Wade R, Richards S, Oscier DG, Catovsky D, Houlston RS. Scan of 977 nonsynonymous SNPs in CLL4 trial patients for the identification of genetic variants influencing prognosis. Blood. 2008; 111:1625–33. https://doi.org/10.1182/blood-2007-08-110130. [PubMed].
67. Kashfi SMH, Behboudi Farahbakhsh F, Nazemalhosseini Mojarad E, Mashayekhi K, Azimzadeh P, Romani S, Derakhshani S, Malekpour H, Asadzadeh Aghdaei H, Zali MR. Interleukin-16 polymorphisms as new promising biomarkers for risk of gastric cancer. Tumour Biol. 2016; 37:2119–26. https://doi.org/10.1007/s13277-015-4013-y. [PubMed].
68. Gao LB, Rao L, Wang YY, Liang WB, Li C, Xue H, Zhou B, Sun H, Li Y, Lv ML, Du XJ, Zhang L. The association of interleukin-16 polymorphisms with IL-16 serum levels and risk of colorectal and gastric cancer. Carcinogenesis. 2009; 30:295–9. https://doi.org/10.1093/carcin/bgn281. [PubMed].
69. Landi D, Gemignani F, Naccarati A, Pardini B, Vodicka P, Vodickova L, Novotny J, Forsti A, Hemminki K, Canzian F, Landi S. Polymorphisms within micro-RNA-binding sites and risk of sporadic colorectal cancer. Carcinogenesis. 2008; 29:579–84. https://doi.org/10.1093/carcin/bgm304. [PubMed].
70. Mi Y, Wang L, Zong L, Pei M, Lu Q, Huang P. Genetic variants in microRNA target sites of 37 selected cancer-related genes and the risk of cervical cancer. PLoS One. 2014; 9:e86061. https://doi.org/10.1371/journal.pone.0086061. [PubMed].
71. Hughes L, Ruth K, Rebbeck TR, Giri VN. Genetic variation in IL-16 miRNA target site and time to prostate cancer diagnosis in African-American men. Prostate Cancer Prostatic Dis. 2013; 16:308–14. https://doi.org/10.1038/pcan.2013.36. [PubMed].
72. Azimzadeh P, Romani S, Mohebbi SR, Mahmoudi T, Vahedi M, Fatemi SR, Zali N, Zali MR. Association of polymorphisms in microRNA-binding sites and colorectal cancer in an Iranian population. Cancer Genet. 2012; 205:501–7. https://doi.org/10.1016/j.cancergen.2012.05.013. [PubMed].
73. Yang L, Liu J, Qi G, Yue W. Interleukin-16 gene polymorphism is associated with acute coronary syndrome in the Chinese Han population. Int J Clin Exp Med. 2018; 11:2624–2629.
74. Moher D, Liberati A, Tetzlaff J, Altman DG, and PRISMA Group. Preferred Reporting Items for Systematic Reviews and Meta-Analyses: The PRISMA Statement. PLoS Med. 2009; 6:e1000097. https://doi.org/10.1371/journal.pmed.1000097. [PubMed].
75. Schwarzer G. An R package for meta-analysis. R News. 2007; 7:40–45.
76. Wells G, Shea B, O’Connell D, Peterson J. The Newcastle-Ottawa Scale (NOS) for assessing the quality of nonrandomised studies in meta-analyses. Ottawa, ON: Ottawa Hospital Research Institute. 2000. http://www.ohri.ca/programs/clinical_epidemiology/oxford.asp.
77. Egger M, Smith GD, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629–34. https://doi.org/10.1136/bmj.315.7109.629. [PubMed].
78. Begg CB, Mazumdar M. Operating characteristics of a rank correlation test for publication bias. Biometrics. 1994; 50:1088–101. https://doi.org/10.2307/2533446. [PubMed].
79. Minelli C, Thompson JR, Abrams KR, Thakkinstian A, Attia J. How should we use information about HWE in the meta-analyses of genetic association studies? Int J Epidemiol. 2008; 37:136–46. https://doi.org/10.1093/ije/dym234. [PubMed].
80. Gauderman WJ. Sample size requirements for matched case-control studies of gene-environment interaction. Stat Med. 2002; 21:35–50. https://doi.org/10.1002/sim.973. [PubMed].
81. He B, Xu T, Pan B, Pan Y, Wang X, Dong J, Sun H, Xu X, Liu X, Wang S. Polymorphisms of TGFBR1, TLR4 are associated with prognosis of gastric cancer in a Chinese population. Cancer Cell Int. 2018; 18:191. https://doi.org/10.1186/s12935-018-0682-0. [PubMed].
82. Li L, Wu C, Zhu X. Role of gene polymorphisms and plasma levels of interleukin-16 in susceptibility to papillary thyroid carcinoma. Int J Clin Exp Med. 2018; 11:2477–2482.
83. MaiMaiTiMin A. AHaTi Y, BaGeDaTi D, AbuDuReYiMu A. Genetic polymorphism of interleukin-16 rs11556218 T/G influences susceptibility to breast cancer in a Chinese population. Int J Clin Exp Pathol. 2016; 9:3931–3936.
84. Tang YJ, Wang JL, Xie KG, Lan CG. Association of interleukin 16 gene polymorphisms and plasma IL16 level with osteosarcoma risk. Sci Rep. 2016; 6:34607. https://doi.org/10.1038/srep34607. [PubMed].
85. Yang SX, Chen F, Zhang JW, Sun ZQ, Chen BP. IL-16 rs4778889 polymorphism contribution to the development of renal cell cancer in a Chinese population. Genet Mol Res. 2016; 15. https://doi.org/10.4238/gmr.15027553. [PubMed].
86. Yao Z, Xia M, Zhu X, Jiang X. ssociation between IL-16 rs4778889 and rs4072111 polymorphisms and ovarian cancer risk in Chinese Han population: a case-control study. Int J Clin Exp Med. 2016; 9:13154–13159.
87. Wang Z, Xu Y, Zhu S. Interleukin-16 rs4778889 polymorphism contributes to the development of renal cell cancer in a Chinese population. Int J Clin Exp Pathol. 2015; 8:15228–33. [PubMed].
88. Luo QS, Wang JL, Deng YY, Huang HD, Fu HD, Li CY, Huang HN. Interleukin-16 Polymorphism Is Associated with an Increased Risk of Glioma. Genet Test Mol Biomarkers. 2014; 18:711–4. https://doi.org/10.1089/gtmb.2014.0170. [PubMed].
89. Qin X, Peng Q, Lao X, Chen Z, Lu Y, Lao X, Mo C, Sui J, Wu J, Zhai L, Yang S, Li S, Zhao J. The association of interleukin-16 gene polymorphisms with IL-16 serum levels and risk of nasopharyngeal carcinoma in a Chinese population. Tumour Biol. 2014; 35:1917–24. https://doi.org/10.1007/s13277-013-1257-2. [PubMed].
90. Hai-Feng T, Wei W, Yuan-Yuan Y, Jun Z, Su-Ping G, Hui-Ming L. Association between Polymorphisms in IL-16 Genes and Coronary Heart Disease risk. Pak J Med Sci. 2013; 29:1033–7. [PubMed].
91. Huang H, Zeng Z, Zhang L, Liu R, Li X, Qiang O, Zhang Q, Chen Y. The association of interleukin-16 gene polymorphisms with susceptibility of coronary artery disease. Clin Biochem. 2013; 46:241–4. https://doi.org/10.1016/j.clinbiochem.2012.11.009. [PubMed].
92. Tong Z, Li Q, Zhang J, Wei Y, Miao G, Yang X. Association between interleukin 6 and interleukin 16 gene polymorphisms and coronary heart disease risk in a Chinese population. J Int Med Res. 2013; 41:1049–56. https://doi.org/10.1177/0300060513483405. [PubMed].
93. Zhang T, Wang H. Variants of interleukin-16 associated with gastric cancer risk. Asian Pac J Cancer Prev. 2013; 14:5269–73. https://doi.org/10.7314/APJCP.2013.14.9.5269. [PubMed].
94. Azimzadeh P, Romani S, Mohebbi SR, Kazemian S, Vahedi M, Almasi S, Fatemi SR, Zali MR. Interleukin-16 (IL-16) gene polymorphisms in Iranian patients with colorectal cancer. J Gastrointestin Liver Dis. 2011; 20:371–6. [PubMed].
95. Chen Y, Huang H, Liu S, Pan LA, Zhou B, Zhang L, Zeng Z. IL-16 rs11556218 gene polymorphism is associated with coronary artery disease in the Chinese Han population. Clin Biochem. 2011; 44:1041–4. https://doi.org/10.1016/j.clinbiochem.2011.06.010. [PubMed].
96. Wu J, Wang Y, Zhang Y, Li L. Association Between Interleukin-16 Polymorphisms and Risk of Coronary Artery Disease. DNA Cell Biol. 2011; 30:305–8. https://doi.org/10.1089/dna.2010.1145. [PubMed].
97. Zhu J, Qin C, Yan F, Wang M, Ding Q, Zhang Z, Yin C. IL-16 polymorphism and risk of renal cell carcinoma: Association in a Chinese population. Int J Urol. 2010; 17:700–7. https://doi.org/10.1111/j.1442-2042.2010.02559.x. [PubMed].
98. Gao LB, Liang WB, Xue H, Rao L, Pan XM, Lv ML, Bai P, Fang WL, Liu J, Liao M, Zhang L. Genetic polymorphism of Interleukin-16 and risk of nasopharyngeal carcinoma. Clin Chim Acta. 2009; 409:132–5. https://doi.org/10.1016/j.cca.2009.09.017. [PubMed].

