Introduction
Many of the first targeted therapeutics for treating patients diagnosed with non-Hodgkin’s lymphoma (NHL) were chimeric or humanized mouse monoclonal antibodies that recognized a member of the CD family of cell surface receptors [1–11]. While the clinical use of these antibodies has improved the outcomes of therapy for many NHL patients over the past two decades, advances in our understanding of the molecular basis of this disease and the discovery of new targets and treatment strategies has markedly expanded the range of options now available for lymphoma therapy. Today the pipelines of many pharmaceutical companies contain a wide range of both small molecule and antibody-based drugs that bind to and/or inhibit many cell surface receptors, proteins and enzymes required for tumor growth and proliferation. These pipelines also include a number of immunotherapies that utilize, augment or activate a patient’s own immune system to mount an effective anti-tumor response.
One family of cell surface receptors that play a central role in immunological surveillance and the induction of anti-tumor immunity are the Major Histocompatibility Complex (MHC) proteins. An interesting therapeutic approach that has potential for treating many types of solid cancer has focused on the isolation and identification of tumor peptide antigens presented by the MHC class I [12–17] and MHC class II [18–26] proteins that trigger the activation of lymphocytes and the induction of an effective anti-tumor immune response. The majority of this work is being driven by a growing interest in utilizing these antigens to develop vaccines for melanoma, lung, cervical, and other cancers whose cells express proteins that contain sequence mutations or structural features not found in normal cells.
Another approach that has shown some success in its application to the treatment of B-cell derived malignancies involves the development of antibodies and other drugs that target the MHC class II proteins directly [27–36]. The characterization of one such antibody, Lym-1, led to the discovery that its target, HLA-DR10 and other HLA-DRs containing a common epitope located on its β-subunit [37–39], is expressed by many B-cell derived lymphomas and leukemias. While the clinical trials conducted with Lym-1 have demonstrated the utility of HLA-DR as a target for NHL therapy [29, 30, 32, 33, 40], a number of patients in the trials developed serious adverse effects commonly encountered in antibody therapies. In an effort to create small molecule targeting agents for use in cancer therapy that exhibit the same avidity and selectivity of antibodies but lack human antibody-mouse antibody (HAMA) responses and other antibody induced adverse effects [41–50], a series of selective high affinity ligands (SHALs) were developed to target the HLA-DR10 epitope recognized by Lym-1. This work led to the development of the family of tridentate SHALs, which include SH7139 and SH7129, that are the first of a new class of NHL small molecule drug and diagnostic candidates that target tumor cells expressing HLA-DR [51, 52].
The preclinical testing of SH7139 has not only shown that this drug exhibits remarkable anti-tumor activity in an aggressive Burkitt lymphoma mouse xenograft model [52], but the pre-IND enabling toxicology and safety studies conducted with SH7139 also suggests the drug should have few if any adverse effects and a safety margin that is uncharacteristically large for an oncology drug. During the course of this testing a biotinylated analog of SH7139, SH7129, was developed for use as a potential companion diagnostic to identify NHL patients whose tumors express the HLA-DRs targeted by the drug. Previous analyses of SH7129 binding to a small number of B-cell derived lymphoma cell lines [51] and human [53] and canine [54] tumor biopsy samples have shown the diagnostic selectively binds to lymphoma cells expressing certain HLA-DRs or their canine DLA-DR orthologs. The specific HLA-DRs SH7129 target include HLA-DR7, HLA-DR9, HLA-DR10, HLA-DR11, HLA-DR12, HLA-DR13, HLA-DR15, and HLA-DR16 [53].
While there have been numerous reports of HLA-DR expression by melanomas [21, 55–58], cervical [59–62], ovarian [63–68] prostate [19, 69–73], liver [74–77], kidney [18, 78], bone [79], breast [80–85], esophageal [86–89], head and neck [90–92], bladder [93–96], colorectal [97–101], lung [102–105], pancreatic [106], larynx [107–109], gastric [110–113], glioma [114–116], and thyroid [117–120] cancers, these MHC class II proteins have not been adequately evaluated as potential targets in the treatment of non-hematological cancers. Although it is not entirely clear why tumors derived from tissues of non-lymphoid origin express HLA-DR, the predominant theory for which there is a great deal of experimental support suggests this expression can be initiated in response to tumor infiltration by lymphocytes, macrophages or dendritic cells [84, 96] and the release of cytokines [84, 121] during the inflammation that often accompanies tumor growth and the progression of the disease. Tumor cell lines treated with IFN-γ, a cytokine released by activated T-cells and NK cells [122, 123], have been observed to upregulate their expression of HLA-DR [121, 124–127]. Analyses of tumors obtained from cervical cancer patients have also shown the tumor cells in biopsy tissues with higher IFN-γ concentrations exhibit higher levels of HLA-DR expression, and these patients survive longer and have a lower risk of disease recurrence [128]. Other studies of hepatocellular carcinoma (HCC) patients have demonstrated a positive correlation between lower plasma concentrations of IFN-γ, advanced tumor stage, and higher rates of HCC recurrence [129].
In the study reported here, the biotinylated analog of SH7139 (SH7129) was used as an immunohistochemical-type stain to examine the frequency with which lymphomas and non-hematological solid cancers express the HLA-DRs targeted by SH7139 and to obtain an estimate of their level of expression by quantifying the amount of SH7129 bound to the cells. The results show that each of the NHL subtypes tested, as well as 18 other types of solid cancer, express the HLA-DRs targeted by SH7139. A number of the non-hematological cancers express higher levels of HLA-DR and bind more SH7129 than the B-cell lymphomas and leukemias.
Results
SH7129 binding to normal tissue
SH7129 binding to normal tissue was evaluated using microarrays containing twenty-four different tissues obtained from three healthy individuals. Following the staining of the microarrays with SH7129, the slides were not counter-stained with hematoxylin. This enables the detection of extremely low levels of SH7129 binding that would normally be obscured by the presence of the counter-stain. Cells expressing the target HLA-DRs that bind SH7129 are stained brown by the horse-radish peroxidase’s conversion of the 3,3′-diaminobenzidine (DAB) substrate to a brown insoluble product. As shown in Figure 1, SH7129 binding was observed to tonsil, thymus, spleen, and bone marrow—all tissues that produce or contain large numbers of antigen presenting cells (APCs). No binding was observed to breast, cerebrum, colon, hypophysis (pituitary), small intestine, ovary, pancreas, salivary gland, skeletal muscle, thyroid, uterine cervix, or peripheral nerve tissue. In each of the three skin samples tested the basal keratinocytes appeared to be stained by SH7129, but examination of the control slides (those stained with hematoxylin and eosin without SH7129) revealed this brown coloration is melanin pigment, not bound SH7129 (Figure 2). Some staining was observed in kidney tissue, but the bound SH7129 was limited to the macrophages, dendritic cells and monocytes located between tubules.

Figure 1: SH7129 binding to normal tissue. Microarrays containing sections from twenty-four normal human tissues (three sections per tissue, each from a different healthy individual) were stained with SH7129 and the bound SHAL was detected using streptavidin conjugated horse-radish peroxidase. Cells binding SH7129 are stained brown. The slides were not counter stained with H&E to enable the detection of very low levels of SH7129 binding. Antigen presenting cells in the bone marrow, thymus, spleen and tonsil bind SH7129 as evidenced by the light staining of cells in those tissues. No binding was observed to breast, cerebrum, colon, hypophysis (pituitary), small intestine, ovary, pancreas, salivary gland, skeletal muscle, thyroid, uterine cervix, peripheral nerve tissue, or skin. The brown coloration in the basal keratinocytes (arrows) in skin is melanin. SH7129 did not bind to lung, esophagus, prostate, cardiac muscle, and parathyroid tissues obtained from two of the three individuals or stomach tissue from one individual. Staining in kidney tissue was limited to the macrophages, dendritic cells and monocytes located between tubules. Liver hepatocytes and zona reticularis cells in the adrenal gland showed very light staining. The images were captured at 40× magnification and the scale bar is the same for all images.
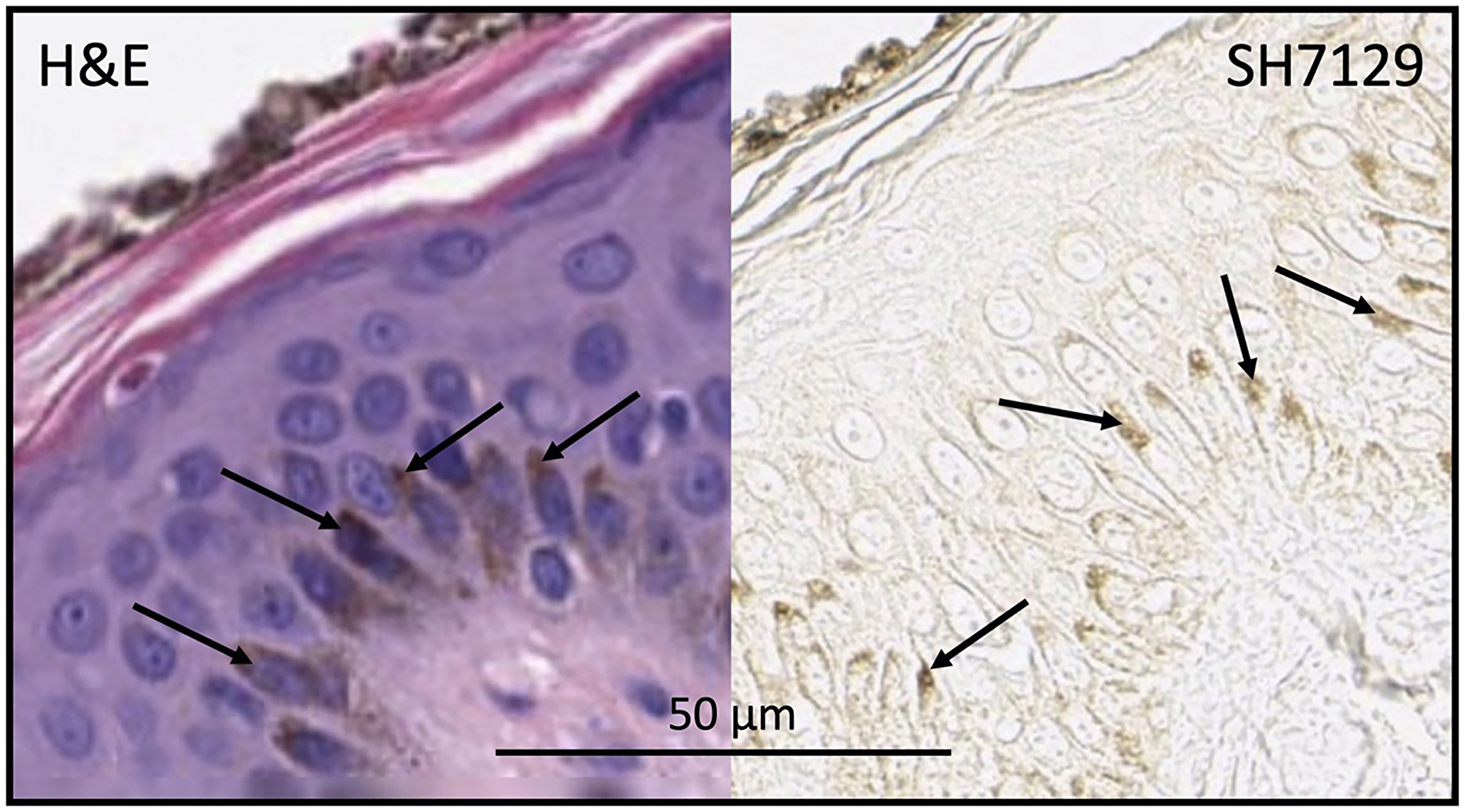
Figure 2: Skin section stained with H&E (without SH7129) or SH7129 (without H&E) showing melanin (arrows) in basal keratinocytes. The apical region of the basal keratinocytes shown in the section stained only with H&E (left panel) has the same brown color as the apical regions of the keratinocytes in the sections stained with SH7129 (right panel). The images were captured at 40× magnification. The scale bar is the same for both images.
Cells in the zona reticularis of adrenal tissue were stained very lightly with SH7129, suggesting these cells may also express very low levels of HLA-DR. HLA-DR expression by adrenal cells has been reported previously [130, 131] and it has been suggested that this expression might be induced during the final maturation step for reticularis cells as they become competent to secrete androgens. It has also been suggested the HLA-DR may trigger the induction of apoptosis in these cells via MHC class II mediated programmed cell death as part of the normal process of adrenal cell turnover [131]. A very low level of SH7129 binding was also observed in cerebellum white matter (Figure 3). Others who have also reported the binding of anti-HLA-DR antibodies to white matter have suggested this binding may be to resting or non-reactive microglia [132], which are cells of the central nervous system that function as macrophages. HLA-DR expression has been shown to increase in the microglia of individuals diagnosed with the neurogenerative diseases Alzheimer’s, Parkinson’s, and multiple sclerosis as well as in the elderly who do not exhibit dementia [132–138]. Very light staining of parietal cells was also observed in stomach sections (Figure 3) obtained from two individuals, while no staining was observed in the section obtained from the third individual. Parietal cells are epithelial cells that have been reported to express HLA-DR in cases of gastritis [139, 140].
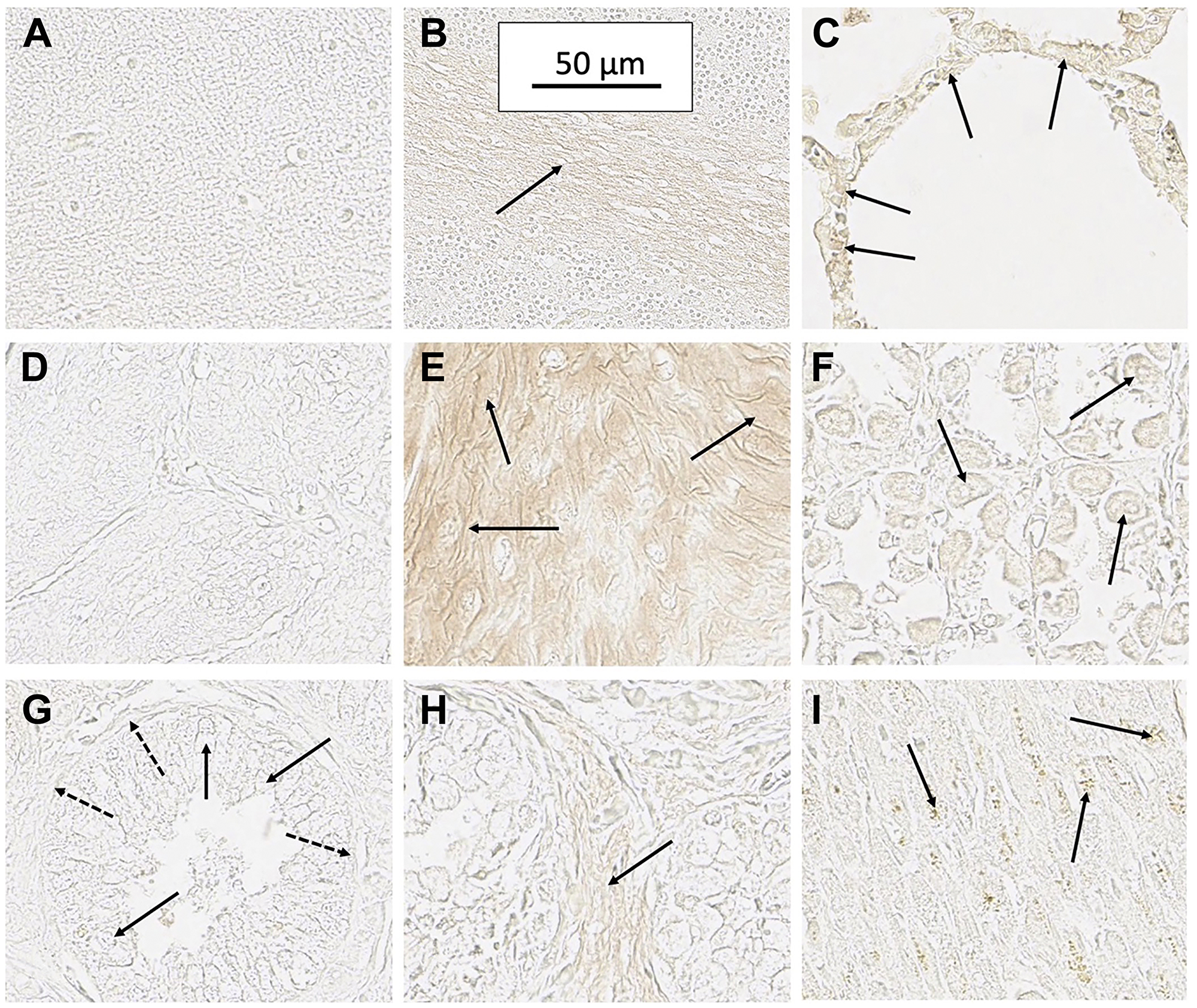
Figure 3: Staining of several normal tissues exhibiting some evidence of SH7129 binding. (A) Cerebellum molecular layer (no binding). (B) Cerebellum white matter (arrow) showing a low level of SH7129 binding. (C) Lung alveolar epithelial cells (arrow) in one section from a normal individual showing SH7129 binding. Lung tissue from two other individuals showed no binding. (D) Esophagus tissue (no binding). (E) Area in tissue section of esophagus from one individual showing binding to squamous epithelial cells (arrows). Esophageal tissue from other two individuals showed no binding. (F) Section of stomach tissue from one individual showing binding to parietal cells (arrows). Stomach tissue from two other individuals showed no binding. (G) Section of normal prostate showing no binding to acinar (solid arrows) and basal cells (dashed arrows). (H) Section of one normal prostate tissue showing low level binding to stroma (arrow). Prostate tissue from two other individuals showed no binding. (I) Section of one normal heart tissue showing very low-level binding to myocytes (arrows). The images were captured at 40× magnification. The scale bar shown in B is the same for all the images.
While SH7129 did not bind to lung, esophagus, prostate, cardiac muscle, and parathyroid tissues obtained from two of the three individuals, the tissue from one individual in each case showed very light staining which is just barely detectable in the captured images. Although these tissues do not normally express HLA-DRs [86, 141–144], the low level binding to the tissues from these individuals may reflect undetected tissue inflammation or very early stage disease. In the lung and esophagus tissue section showing staining, SH7129 binding was localized to the epithelial cells in the alveolar ducts of the lung and the squamous epithelium of the esophagus (Figure 3), which are cell types that have been shown to express HLA-DRs during inflammatory lung and esophagus injury or disease [86, 145–152]. The prostate case showed a very light staining of the stroma, while no binding was observed to the acinar or basal cells (Figure 3). In certain cases of chronic immuno-mediated inflammation, such as benign prostatic hyperplasia, prostate stromal cells have been reported to express HLA-DR and function as antigen presenting cells [153, 154]. In the cardiac tissue section from the one individual showing extremely light staining, the binding appeared to be associated with some (not all) of the myocytes (Figure 3)—an observation others have reported to occur in association with transplant tissue [144, 155, 156], myocarditis [157–159] and other types of cardiovascular disease [157, 160, 161]. Thus, the very low level of SH7129 staining of these normal tissues appears consistent with observations in previous studies that have shown HLA-DR expression in healthy lung, esophagus, prostate, cardiac tissue, cerebellum and stomach is confined to non-lymphoid cells that begin functioning as antigen presenting cells in response to injury or disease.
The only other tissue to which SH7129 binding was observed to bind at a very low level was the liver (Figure 1). Hepatocytes in normal liver do not express HLA-DR [162–164], but epithelial cells surrounding portal tracts have occasionally been observed to express HLA-DR in tissue obtained from healthy individuals and much more frequently in cases of disease [165, 166]. Hepatocytes expressing HLA-DR have only been observed in patients with immune mediated liver disorders [164]. Since SH7129 binding was observed in the liver sections from all three individuals and appears to be localized specifically to hepatocytes, it is highly unlikely the tissues were obtained from three individuals that all have a liver disorder. A more likely possibility is that the staining may reflect a low level of SH7129 binding to something other than HLA-DR. Recent studies have shown that SH7139 inhibits OATP1B1 and OATP1B3 [unpublished results], transporters that have only been found in the liver (normal hepatocytes) [167, 168] or certain cancers [168–170]. The abundance of these transporters (3.18 pmoles OATP1B1 and 2.73 pmoles OATP1B3 per 106 hepatocytes [171]) exceed the number of HLA-DR molecules expressed by the Burkitt lymphoma cell line Raji (2.66 pmoles/106 Raji cells [172, 173]) and some of the highest expressing ovarian cancer cells (0.5 pmoles/106 cells [67]) by 2 to 10-fold, respectively. While the binding affinity of SH7129 has not been determined for either transporter, the IC50 for OATP1B1 (0.29 μM) and OATP1B3 (0.15 μM) inhibition by SH7139 indicate SH7129’s affinity for the transporters will be lower than its affinity for HLA-DR (KD = 23 pM [174]). However, SH7129 would be expected to bind to both transporters under the staining conditions used.
HLA-DR expression by non-Hodgkin’s lymphoma
Using the same SH7129 staining protocol, tumor biopsy sections obtained from patients diagnosed with seven subtypes of NHL were screened for the expression of HLA-DRs targeted by SH7139. Digital images of the tumor cells in each section were obtained from the stained and control (no SH7129 treatment) slides, the images were inverted, and the amount of bound SH7129 was estimated by processing the captured images of each tumor and quantifying the amount of brown oxidized DAB product generated by the horse-radish peroxidase using NIH ImageJ version 1.42 software.
SH7129 staining of biopsy tissues containing cells expressing HLA-DR show binding to HLA-DR proteins located on the surface of the tumor cells, in the cytoplasm, and near the nucleus where the endoplasmic reticulum is located in agreement with previously reported results in cultured Burkitt lymphoma (Raji) cells [51, 175]. Connective tissue is not stained. As shown in Table 1, a significant number of the tested tumors in each of the types of NHL examined were found to bind SH7129. Tumor biopsies obtained from all twenty-four of the anaplastic large cell lymphoma (ALCL) cases examined expressed the targeted HLA-DR and bound SH7129. Nearly every MALT lymphoma (75 of the 80 cases) biopsy sample examined also bound the diagnostic. At the other end of the spectrum, only 28% of the mantle cell and 34% of the follicular lymphomas were observed to express the target and bind SH7129.
Table 1: Solid cancers tested for SH7129 binding as an indicator of their expression of the HLA-DRs targeted by SH7139
| Cancer Type | Number of Cases | Percent Expressing Target |
|---|---|---|
| Non-Hodgkin’s Lymphoma | ||
| DLBC Lymphoma | 75 | 57.3 |
| Follicular Lymphoma | 118 | 33.9 |
| Anaplastic LC Lymphoma | 24 | 100 |
| MALT Lymphoma | 80 | 93.8 |
| Mantle Cell Lymphoma | 71 | 28.2 |
| Burkitt Lymphoma | 25 | 32.0 |
| Small Lymphocytic Lymphoma | 78 | 60.2 |
| Other Cancers | ||
| Ovarian | 64 | 100 |
| Lung | 85 | 98.8 |
| Cervical | 66 | 98.4 |
| Pancreatic | 89 | 65.1 |
| Gastric | 90 | 91.1 |
| Esophageal | 99 | 2 |
| Breast, Medullary Carcinoma | 77 | 66.2 |
| Breast, Invasive Ductal | 90 | 4.4 |
| Kidney | 69 | 79.7 |
| Prostate | 72 | 95.8 |
| Thyroid | 80 | 65 |
| Liver | 75 | 89.3 |
| Colorectal | 92 | 94.5 |
| Bone | 78 | 69.2 |
| Bladder | 60 | 61.7 |
| Plasma Cell Myeloma | 14 | 92.9 |
| Larynx | 70 | 48.6 |
| Melanoma | 122 | 46.7 |
| Head and Neck | 40 | 5 |
Within a typical lymphoma biopsy section, the cell to cell variation in SH7129 binding, as measured by image analysis of the horse-radish peroxidase generated oxidation product of DAB deposited in individual tumor cells, was less than 3-fold (Table 2). The level of HLA-DR expression and SH7129 binding by the cells in different patient’s tumors within the same type of NHL, however, differed by as much as 10 to 100-fold (Figures 4 and 5). Statistical analyses of SH7129 bound by the seven types of NHL indicate these NHL tumors fall into three groups. Biopsy tissues obtained from patients diagnosed with anaplastic large cell lymphoma (ALCL) exhibited the highest level of SH7129 binding. Diffuse large B-cell lymphomas (DLBCL), small lymphocytic lymphomas (SLL), and mucosa-associated lymphoid tissue (MALT) lymphomas bound intermediate amounts of SH7129. Follicular lymphomas (FL), Burkitt’s lymphomas (BL) and mantle cell lymphomas (MCL) bound the least SH7129 of the types of NHL analyzed.
Table 2: Cell to cell variation in SH7129 binding in a typical NHL tumor
| Lymphoma Type | Biopsy Sample ID | Level of SH7129 Binding | Cell Range SH7129 Bound |
|---|---|---|---|
| Diffuse Large B-Cell | LM482E3 | High | 2.98 |
| Follicular | T203B6 | Moderate | 1.69 |
| Burkitt’s | LM482D8 | Moderate | 2.42 |
| MALT | LY804E7 | High | 1.94 |
| Anaplastic Large Cell | LM242D5 | High | 2.75 |
| Mantle Cell | MC2B9 | Moderate | 1.78 |
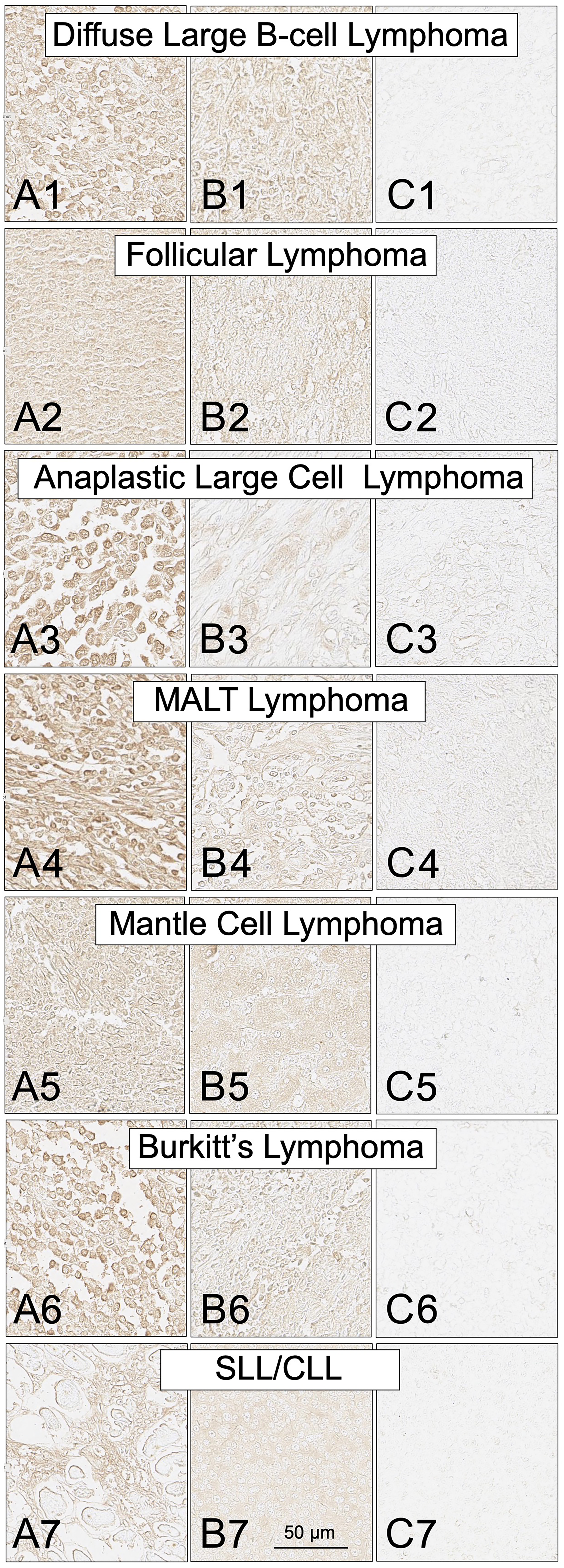
Figure 4: SH7129 staining of tumor biopsy sections from seven subtypes of non-Hodgkin’s lymphoma. Sections from three different tumors are shown for each subtype: high level of SH7129 binding (A), moderate level of SH7129 binding (B), and no SH7129 binding (C). Diffuse large B-cell lymphoma: (A1) Tissue sample ODCTLYMLY02E3. (B1) Tissue sample LM482C8. (C1) Tissue sample LM482C5. Follicular Lymphoma: (A2) Tissue sample T203B6. (B2) Tissue sample 6050459. (C2) Tissue sample 1051864. Anaplastic large cell lymphoma: (A3) Tissue sample LM242D5. (B3) Tissue sample LM242D4. (C3) Tissue sample LM242B1. Mucosa-associated lymphoid tissue lymphoma: (A4) Tissue sample LY804E7. (B4) Tissue sample LY804A6. (C4) Tissue sample LY804G5. Mantle cell lymphoma: (A5) Tissue sample M685565. (B5) Tissue sample 15726/09. (C5) Tissue sample F721798. Burkitt lymphoma: (A6) Tissue sample LM482E3. (B6) Tissue sample BL1A8. (C6) Tissue sample BL1A1. Small lymphocytic lymphoma/chronic lymphocytic leukemia: (A7) Tissue sample F683134. (B7) Tissue sample 9543432. (C7) Tissue sample M672708. The images were captured at 40× magnification. The scale bar is the same for all images.
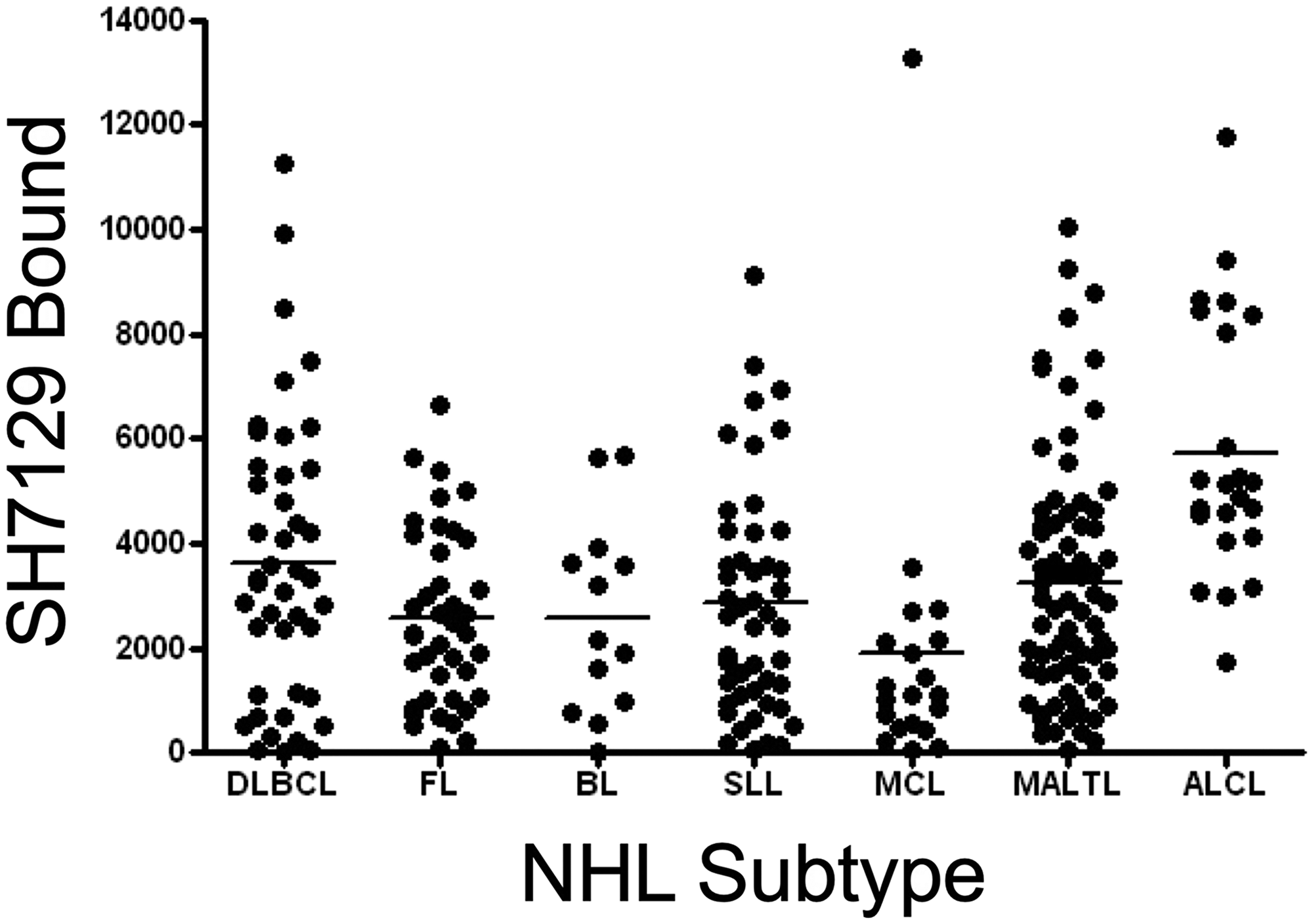
Figure 5: Tumor to tumor variability in SH7129 binding to seven types of NHL. Tumor microarrays containing biopsy tissue sections from tumors of patients diagnosed with diffuse large B-cell lymphoma (DLBCL), follicular lymphoma (FL), Burkitt lymphoma (BL), small lymphocytic lymphoma (SLL), mantle cell lymphoma (MCL), mucosa-associated lymphoid tissue lymphoma (MALTL), and anaplastic large cell lymphoma (ALCL) were stained with SH7129 and the biotin in the bound SHAL was detected using SAHRP and the substrate DAB. The amount of bound SH7129 was determined by densitometric analysis of each tumor section. Not all biopsies contained tumor cells expressing the HLA-DR target to which SH7129 binds. Only those tumors exhibiting SH7129 binding are included in the plot. The value plotted is the amount of bound SH7129 (integrated staining density) per 384-pixel area of tumor tissue. Each point corresponds to a tumor from a different patient. A significant number of tumors representing each subtype of NHL were observed to bind SH7129. The range in SH7129 binding to the tumors within each NHL subtype varied by as much as 100-fold.
Identification of other solid cancers that also express HLA-DRs targeted by SH7139
Biopsy sections from eighteen additional solid cancers that have been reported by others to express MHC class II proteins were also examined for expression of the HLA-DRs targeted by SH7139 using the same staining protocol. While many (33–100%) of the tumors analyzed in sixteen of these cancers were found to bind SH7129 (Table 1), only two of the ninety-nine esophageal (2%) and two of the forty head and neck tumors (5%) (Figure 6) showed detectable SH7129 binding. The highest percentage of cases showing binding was observed for ovarian, lung, cervical, gastric, prostate, myeloma and colorectal cancers. Ovarian, colorectal, prostate and cervical cancers (Figure 7) exhibited the highest levels of SH7129 binding of all the solid tumors examined (Figure 8). In marked contrast to the other types of cancer, which had cases representing the full spectrum (low to high) of HLA-DR expression and SH7129 binding (Figures 9–11), all of the ovarian and all but one of the cervical cancer biopsy sections examined bound moderate to high levels of SH7129. The amount of SH7129 bound by the two esophageal and two head and neck tumors were amongst the lowest of all the cancers tested.
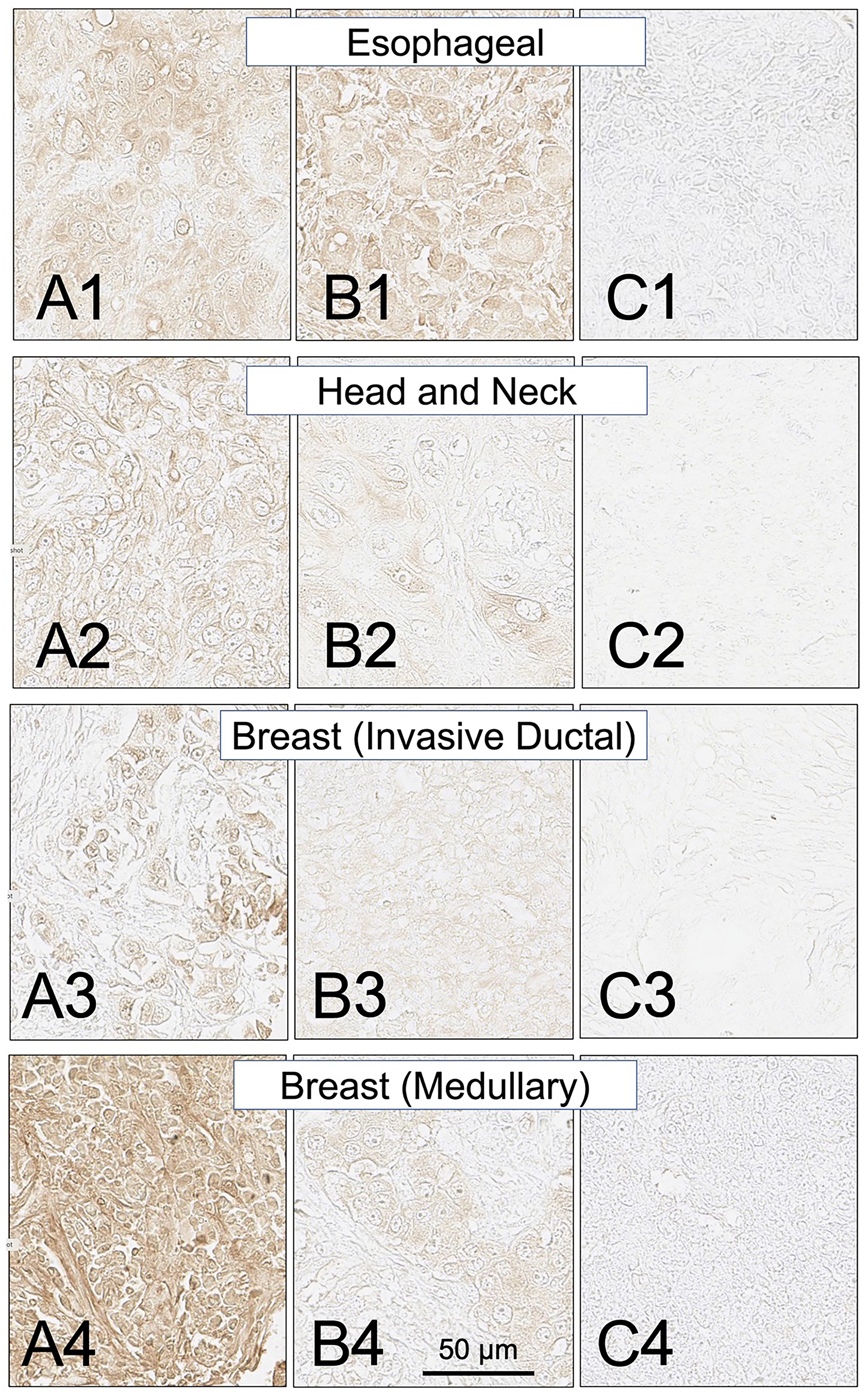
Figure 6: SH7129 stained sections of representative esophageal, head and neck, and breast (invasive ductal and medullary carcinoma) cancers expressing HLA-DRs targeted by SH7139. Sections from three different tumors are shown for each cancer: high level of SH7129 binding (A), moderate level of SH7129 binding (B), and no SH7129 binding (C). Esophageal cancer: (A1) Tissue sample ESC1021H1, adenocarcinoma. (B1) Tissue sample EC1021A12, squamous cell carcinoma. (C1) Tissue sample ESC1021B2, squamous cell carcinoma. Head and Neck: (A2) Tissue sample HN483C3, squamous cell carcinoma. (B2) Tissue sample HN483A4, squamous cell carcinoma. (C2) Tissue sample HN483E5, squamous cell carcinoma. Invasive Ductal breast cancer: (A3) Tissue sample BC08118A9. (B3) Tissue sample BC08118A5. (C3) Tissue sample BC08118H3. Medullary breast cancer: (A4) Tissue sample BR807A1. (B4) Tissue sample BR807A2. (C4) Tissue sample BR807B8. The images were captured at 40× magnification. The scale bar is the same for all images.
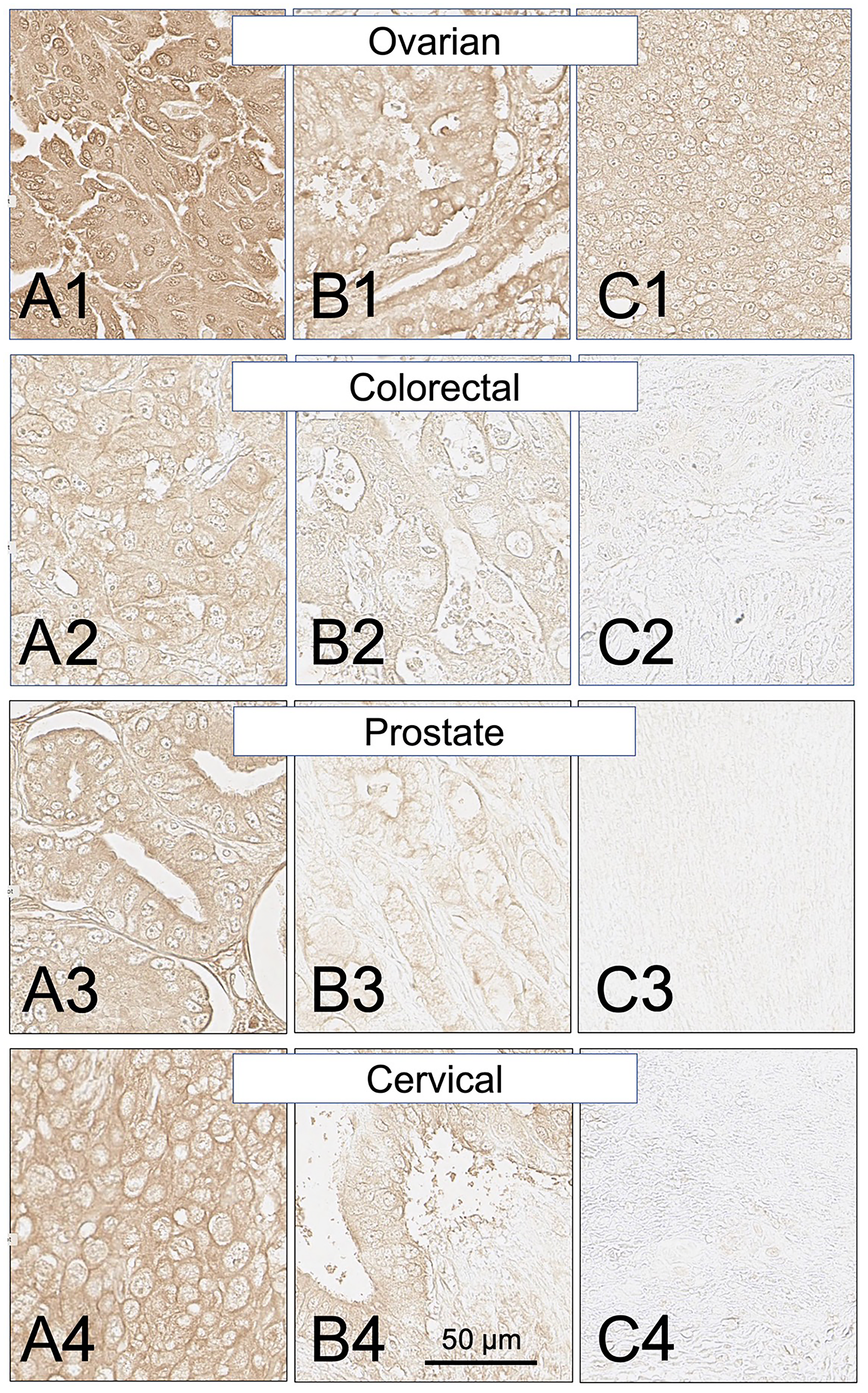
Figure 7: SH7129 stained sections of representative ovarian, colorectal, prostate, and cervical cancers expressing HLA-DRs targeted by SH7139. Three examples are shown for each cancer: high level of SH7129 binding (A), moderate level of SH7129 binding (B), and no SH7129 binding (C). Since all of the tested ovarian cancer biopsy sections bound SH7129, the ovarian cancer example shown in (C) is a third tumor showing SH7129 binding. Ovarian cancer: (A1) Tissue sample OVC1501E4, endometrioid adenocarcinoma. (B1) Tissue sample OVC1501D1, mucinous cystadenocarcinoma. (C1) Tissue sample OVC1501G4, serous cystadenocarcinoma. Colorectal cancer: (A2) Tissue sample ODCTDGCOL04D1, adenocarcinoma. (B2) Tissue sample ODCTDGCOL04B3, adenocarcinoma. (C2) Tissue sample ODCTDGCOL04F8, adenocarcinoma. Prostate cancer: (A3) Tissue sample PR803CA10, adenocarcinoma. (B3) Tissue sample PR803CE1. (C3) Tissue sample PR803CH2. Cervical cancer: (A4) Tissue sample CXC1501C6, squamous cell carcinoma. (B4) Tissue sample CXC1501G3, adenocarcinoma. (C4) Tissue sample CXC1501F6, squamous cell carcinoma. The images were captured at 40× magnification. The scale bar is the same for all images.
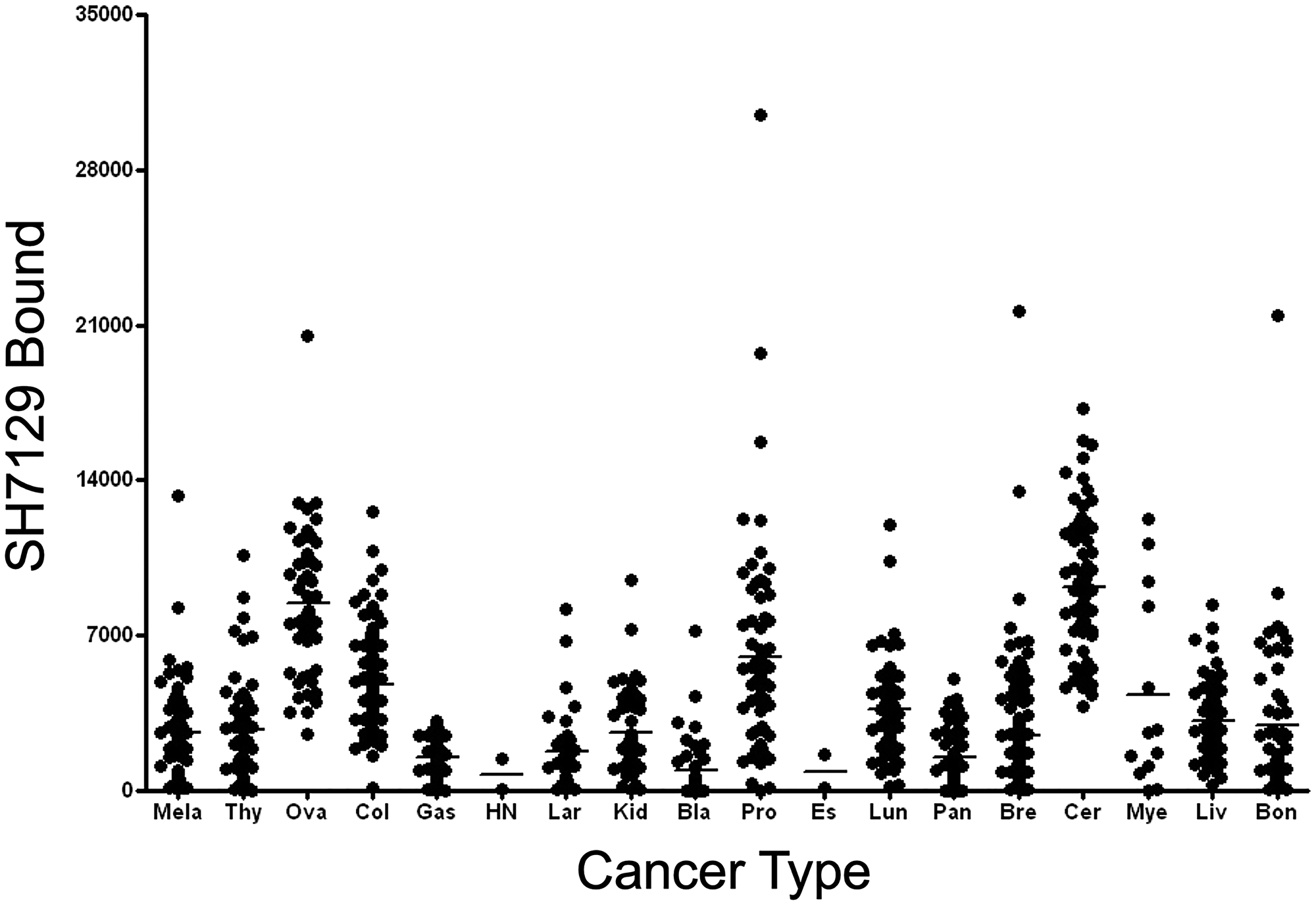
Figure 8: Tumor to tumor variability in SH7129 binding to other cancers. Tumor microarrays containing biopsy sections from tumors of patients diagnosed with one of eighteen different types of solid cancers were stained with SH7129 and the biotin in the bound SHAL was detected using SAHRP and the substrate DAB. The amount of bound SH7129 was determined by densitometric analysis of each tumor section. Not all biopsies contained tumor cells expressing the HLA-DR target to which SH7129 binds. Only those tumors exhibiting SH7129 binding are included in the plot. The value plotted is the amount of bound SH7129 (integrated stain density) per 384-pixel area of tumor tissue. Each point corresponds to a tumor from a different patient. A significant number of tumors representing 16 of the 18 cancers analyzed were observed to bind SH7129. Only two of the head and neck and two esophageal cancers bound SH7129. While the range in SH7129 binding to different cervical, ovarian, breast or myeloma tumors expressing the HLA-DR target is small (5 to 16-fold), SH7129 binding to individual tumors in many of the other non-hematological cancers vary by more than 100-fold. The solid cancers tested include: melanoma (Mela), thyroid (Thy), ovarian (Ova), colorectal (Col), gastric (Gas), head and neck (HN), laryngeal (Lar), kidney (Kid), bladder (Bla), prostate (Pro), esophageal (Es), lung (Lun), pancreatic (Pan), breast (Bre), cervical (Cer), plasma cell myeloma (Mye), liver (Liv), and bone (Bon).
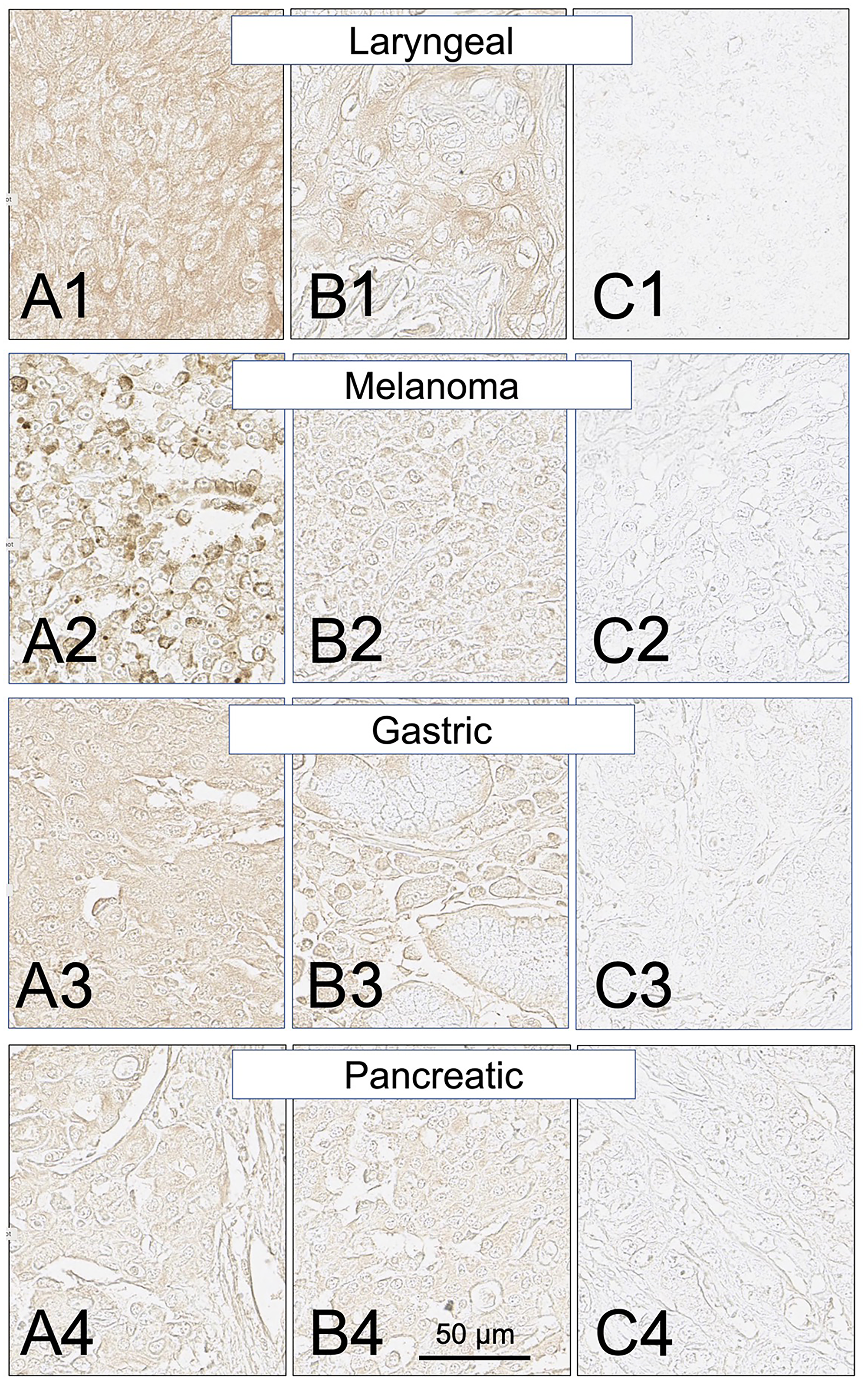
Figure 9: SH7129 stained sections of representative laryngeal, melanoma, gastric, and pancreatic cancers expressing HLA-DRs targeted by SH7139. Sections from three different tumors are shown for each cancer: high level of SH7129 binding (A), moderate level of SH7129 binding (B), and no SH7129 binding (C). Laryngeal cancer: (A1) Tissue sample LP801A7, squamous cell carcinoma. (B1) Tissue sample LP801C6, basaloid squamous cell carcinoma. (C1) Tissue sample LP801G1, squamous cell carcinoma. Melanoma: (A2) Tissue sample Me1004EA10. (B2) Tissue sample Me1004EG6. (C2) Tissue sample Me1004 EB1. Gastric cancer: (A3) Tissue sample ODCTDGSTM01E4, adenocarcinoma. (B3) Tissue sample ODCTDGSTM01J4, ring cell carcinoma. (C3) Tissue sample ODCTDGSTM01I9, adenocarcinoma. Pancreatic cancer: (A4) Tissue sample PA961CF1, duct adenocarcinoma. (B4) Tissue sample PA961CH2, carcinoid. (C4) Tissue sample PA961CF10, adenocarcinoma. The images were captured at 40× magnification. The scale bar is the same for all images.
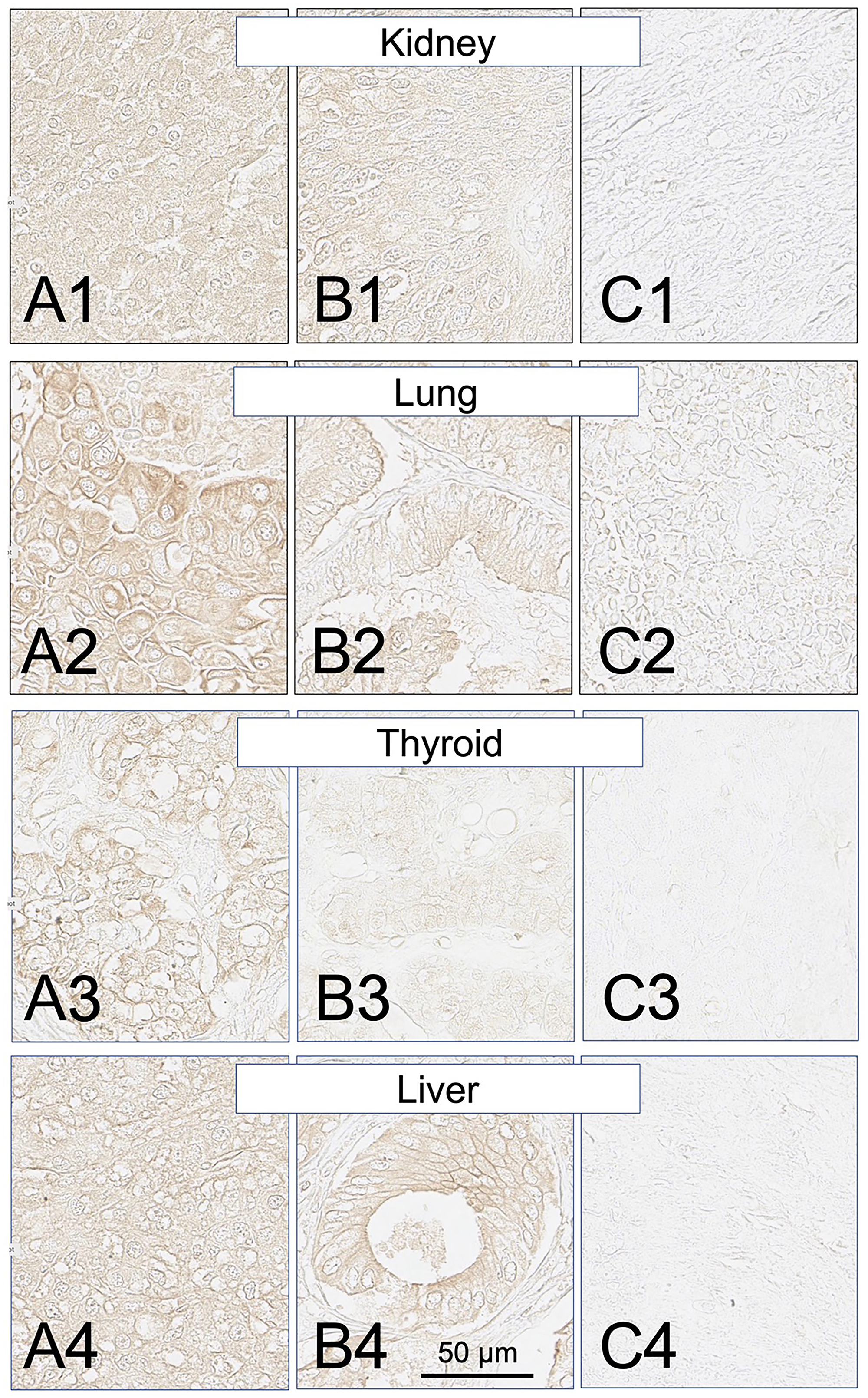
Figure 10: SH7129 stained sections of representative kidney, lung, thyroid and liver cancers expressing HLA-DRs targeted by SH7139. Sections from three different tumors are shown for each cancer: high level of SH7129 binding (A), moderate level of SH7129 binding (B), and no SH7129 binding (C). Kidney cancer: (A1) Tissue sample BC07014B1, clear cell carcinoma. (B1) Tissue sample BC07014AH2, transitional cell carcinoma. (C1) Tissue sample BC07014AB7, clear cell carcinoma. Lung cancer: (A2) Tissue sample LUC1021E1, squamous cell carcinoma. (B2) Tissue sample LUC1021C1, adenocarcinoma. (C2) Tissue sample LUC1021E3, squamous cell carcinoma. Thyroid cancer: (A3) Tissue sample TH802C1, papillary carcinoma. (B3) Tissue sample TH802E6, follicular adenoma. (C3) Tissue sample TH802G7, embryonic adenoma. Liver cancer: (A4) Tissue sample LVC1501A9, hepatocellular carcinoma. (B4) Tissue sample LVC1501A8, bile duct carcinoma. (C4) Tissue sample LVC1501F7, hepatocellular carcinoma. The images were captured at 40× magnification. The scale bar is the same for all images.
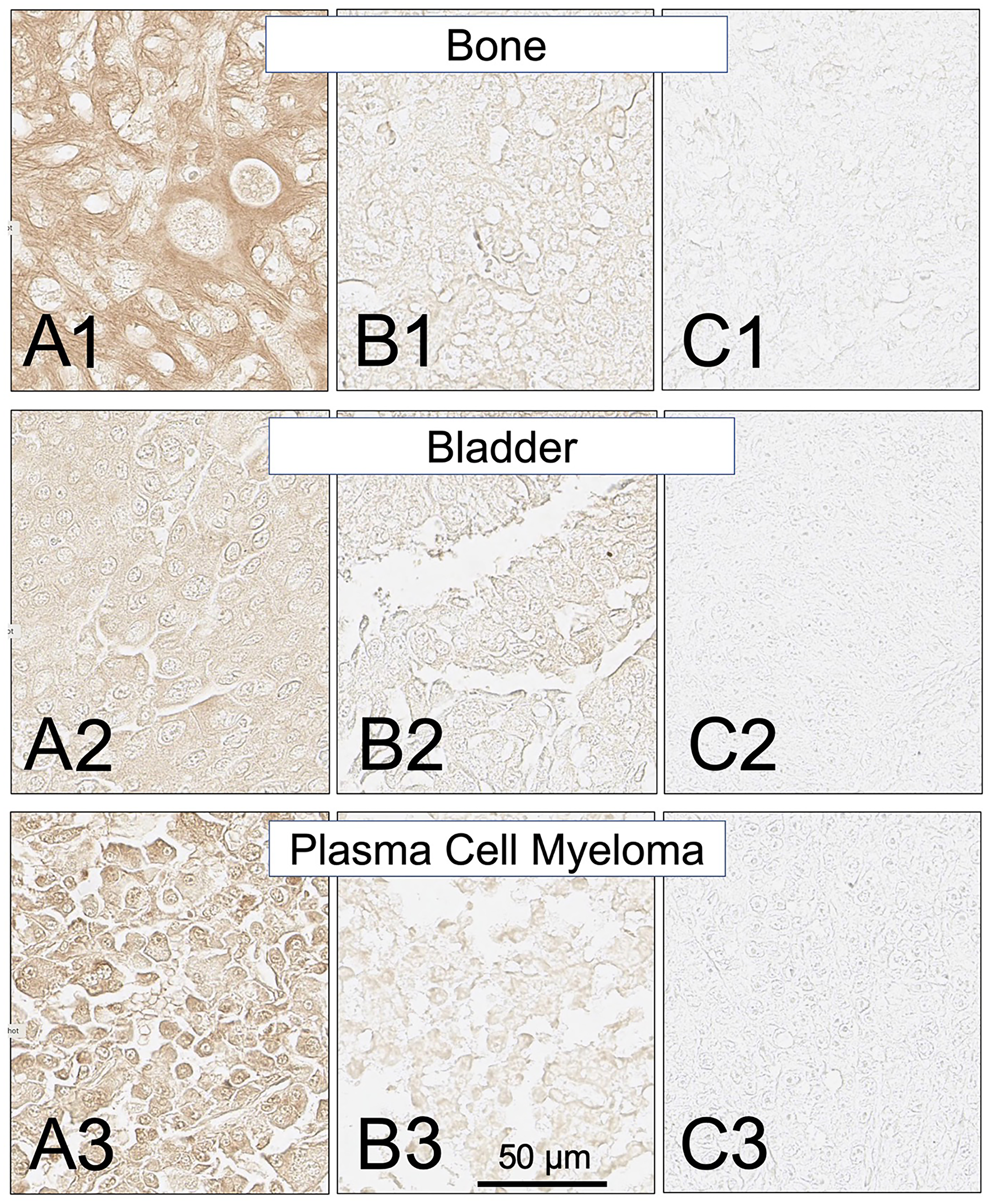
Figure 11: SH7129 stained sections of representative bone, bladder and plasma cell myeloma cancers expressing HLA-DRs targeted by SH7139. Sections from three different tumors are shown for each cancer: high level of SH7129 binding (A), moderate level of SH7129 binding (B), and no SH7129 binding (C). Bone cancer: (A1) Tissue sample OS802F5, chondrosarcoma. (B1) Tissue sample OS802B8, osteosarcoma. (C1) Tissue sample OS802C6, osteosarcoma. Bladder cancer: (A2) Tissue sample BC12011BD3, urothelial carcinoma. (B2) Tissue sample BC12011BE4, urothelial carcinoma (C2) Tissue sample BC12011E6, urothelial carcinoma. Plasma cell myeloma: (A3) Tissue sample T291B3. (B3) Tissue sample LM482A6. (C3) Tissue sample LM482A1. The images were captured at 40× magnification. The scale bar is the same for all images.
Similar to the results obtained in the analysis of the different subtypes of NHL, a broad range in the level of SH7129 binding/HLA-DR target expression was observed between individual tumors for many of these solid cancers. The gastric and pancreatic cancer cases (Figures 8 and 9) showed the least variability in SH7129 binding. Within the ovarian, prostate, melanoma, breast and bone cancers the tumors from a small number of outlier cases expressed extremely high levels of the HLA-DR target (Figure 8). The amount of SH7129 bound by these outliers was nearly twice the amount bound by the lymphoma outliers (Figure 5).
Variation in SH7129 binding by tumor type or grade within the non-hematological cancers
SH7129 binding to the different types of nine of the non-lymphoid solid cancers analyzed (Figure 12) was also compared to determine if a particular type might express more or less of the HLA-DRs. With one exception, the level of target HLA-DR expression by the different types of lung, liver, ovarian, laryngeal, gastric, breast, and bone cancers were not found to be statistically different. Within the cervical cancers analyzed, the squamous cell carcinomas (SC) bound more SH7129 than the adenocarcinomas (A) (p = 0.006).
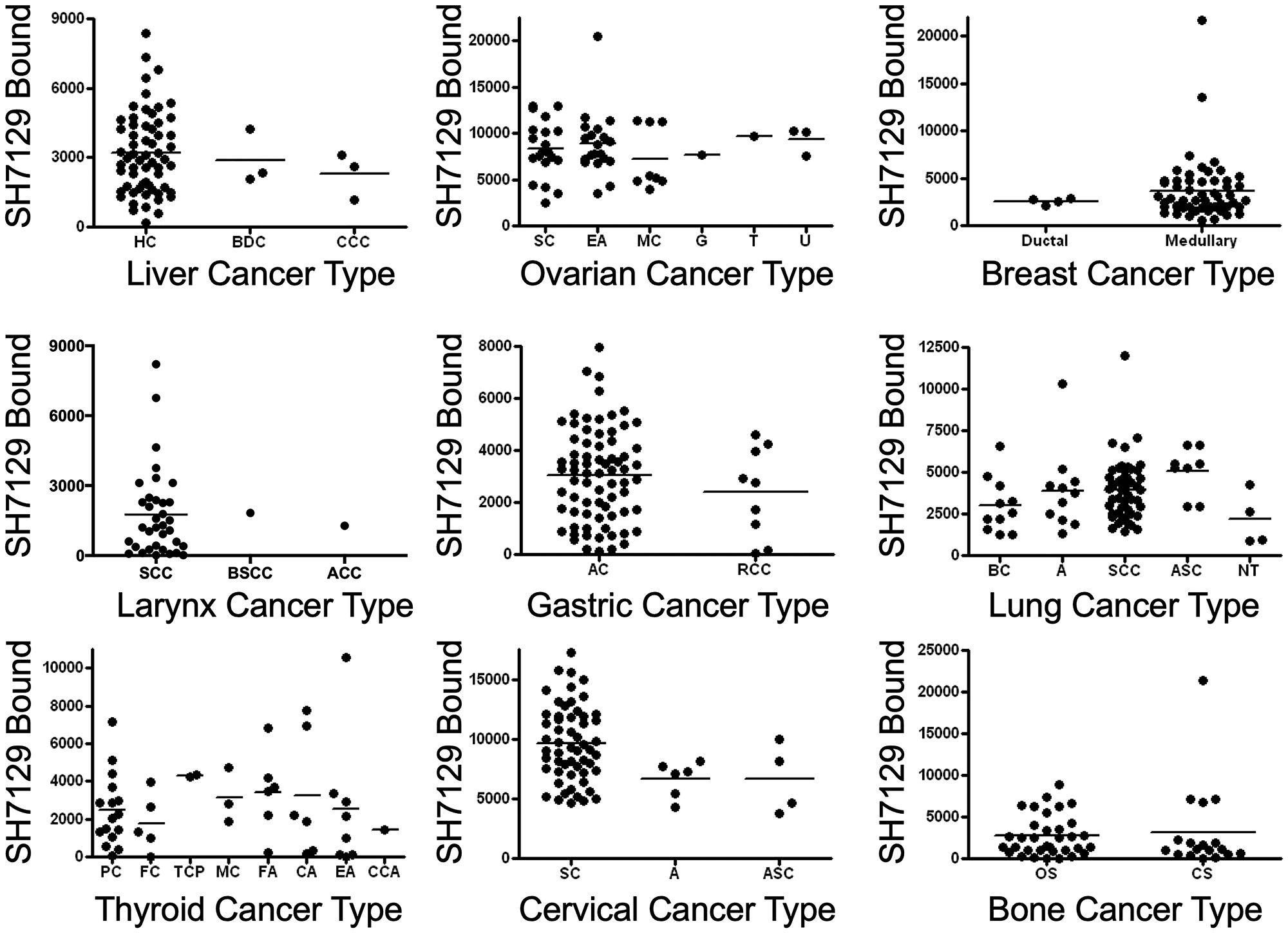
Figure 12: SH7129 binding to different types of nine non-lymphoid solid cancers. SH7129 binding data shown in Figure 8 were sorted by type for nine of the cancers and the binding to the different tumors within each type were plotted for comparison. SH7129 binding to the different types of lung, liver, ovarian, laryngeal, gastric, breast, and bone cancers were not found to be significantly different. In a number of cases there were two few cases to provide a meaningful comparison. A statistically significant difference was only observed for two types of cervical cancer; the squamous cell carcinomas (SC) bound more SH7129 than the adenocarcinoma (A) type (p = 0.006). Liver cancers: hepatocellular carcinoma (HC), bile duct carcinoma (BDC), and clear cell carcinoma (CCC). Ovarian cancers: serous cystadenocarcinoma (SC), endometrioid (EA), mucinous cystadenocarcinoma (MC), granulosa cell tumor (G), thecoma (T), and undifferentiated adenocarcinoma (U). Breast cancers: ductal and medullary. Larynx cancer: squamous cell carcinoma (SCC), basaloid squamous cell carcinoma (BSCC), and acinic cell carcinoma (ACC). Gastric cancers: adenocarcinoma (AC) and ring cell carcinoma (RCC). Lung cancers: bronchioloalveolar carcinoma (BC), adenocarcinoma (A), squamous cell carcinoma (SCC), adenosquamous carcinoma (ASC), and neuroendocrine tumor (NT). Thyroid cancers: papillary carcinomas (PC), follicular papillary carcinoma (FC), tall cell papillary carcinoma (TCP), medullary carcinoma (MC), follicular adenoma (FA), colloid adenoma (CA), embryonic adenoma (EA) and clear cell adenoma (CCA). Cervical cancers: squamous cell carcinoma (SC), adenocarcinoma (A), and adenosquamous carcinoma (ASC). Bone cancers: osteosarcoma (OS) and chondrosarcoma (CS).
A comparison of the amount of SH7129 bound to tumors classified by grade yielded a similar result (Figure 13). No difference in SH7129 binding was observed as a function of tumor grade in liver, ovarian, gastric, prostate, laryngeal, lung, cervical or pancreatic cancers. The comparison suggested what appears to be a significantly higher level of SH7129 binding to grade 3 compared to grade 2 kidney cancers (p = 0.0350), but this result is based on the analysis of only two grade 3 cases. While this difference may prove to be real, such a conclusion cannot be confirmed until a much larger number of cases are examined. It is important to point out that the same is true for many of the cancer types and grades analyzed. The results obtained from the comparisons involving only a few cases per cancer type or grade may not reflect the true variation that is present in the larger population.
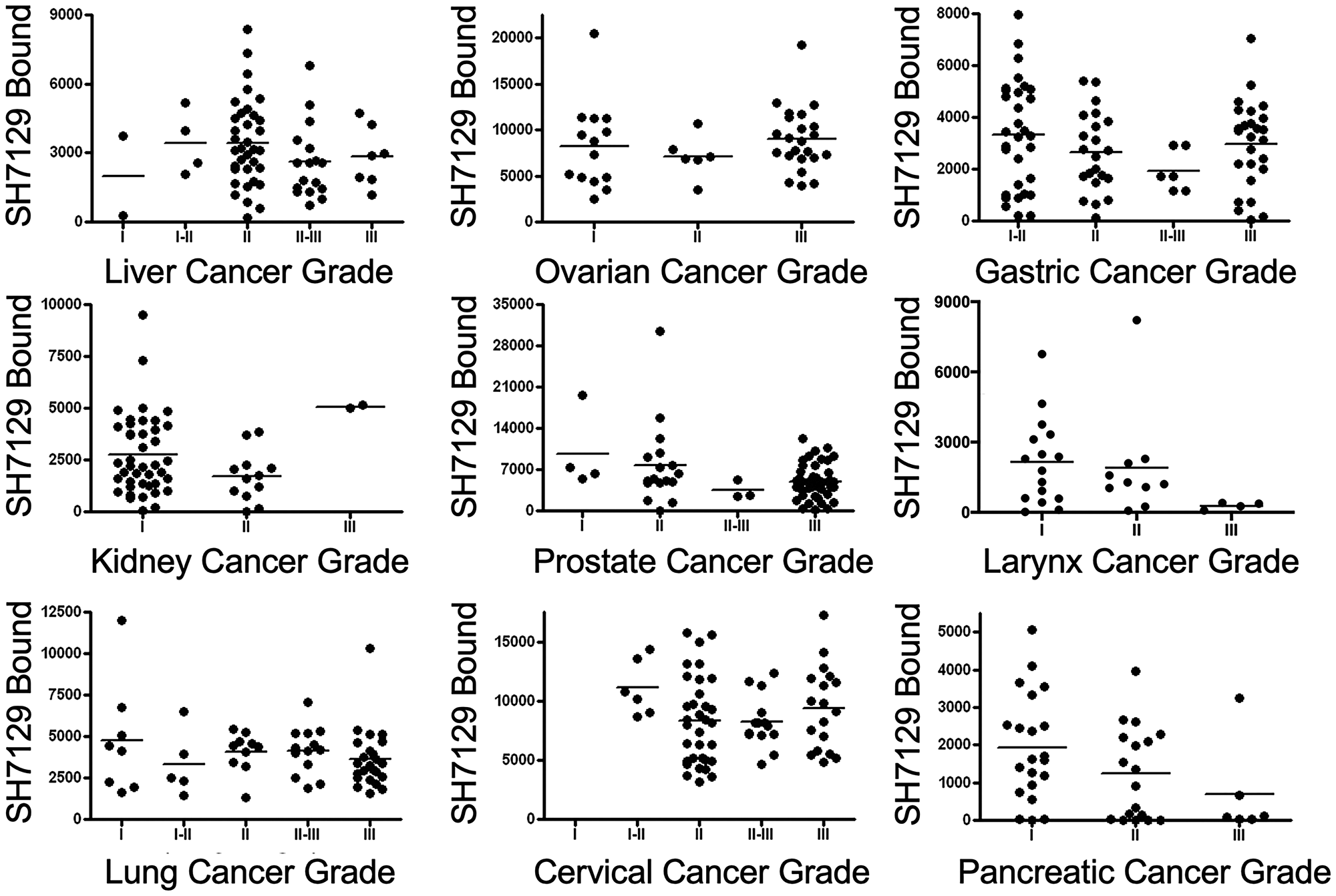
Figure 13: Comparison of SH7129 binding to nine cancers by grade. SH7129 binding data shown in Figure 8 were sorted by grade for nine of the cancers for which there was grade information, and the binding to the different tumors within each type were plotted for comparison. Statistical analyses of the data indicate there is no correlation between the amount of SH7129 bound and tumor grade in liver, ovarian, gastric, prostate, laryngeal, lung, cervical, or pancreatic cancers. The comparison suggested what appears to be a significantly higher level of SH7129 binding to grade III compared to grade II kidney cancers (p = 0.0350), but this result is based on the analysis of only two grade III cases.
DISCUSSION
The SH7129 used to stain the tumor biopsy tissue sections in this study is a biotinylated analog of SH7139 that was synthesized for use in cell binding studies and as a potential companion diagnostic for prescreening patients to identify those with tumors that express the HLA-DRs SH7139 targets. The HLA-DR targeting domains of SH7139 and SH7129 are identical and the two molecules differ only in the effector (a DOTA chelating group or a biotin, respectively) conjugated to the free amine at the non-functional end of the scaffold that links the recognition elements together. The replacement of the DOTA by biotin has been reported in earlier studies to have no impact on either the selectivity of the targeting of HLA-DR [51] or its biological activity [51–53].
With the exception of a single section of lung, esophagus, prostate, stomach and heart tissue derived from a single tissue donor, SH7129 binding to normal tissues was very weak or not observed. Expression of the MHC class II protein HLA-DR is typically restricted to antigen presenting cells in a healthy individual [37, 176]. These include the B-lymphocytes, monocytes, macrophages and dendritic cells found in blood, tonsils, spleen, lymph nodes, thymus, and bone marrow, Kupffer cells in liver, microglia in brain, Langerhans cells in skin, other lymphocytes and macrophages that are frequently found infiltrating numerous organs (e.g., the kidney, breast, and lung), as well as certain epithelial or endothelial cells that provide the first line of an organ’s defense against infection or injury. HLA-DRs are expressed at relatively low levels in resting lymphocytes (approximately 1–2 × 105 molecules per cell) [172].
Expression of HLA-DR in normal tissues that lack APCs does not typically occur except following injury or the onset of disease. Low levels of HLA-DR expression have been observed in cases of primary biliary cirrhosis [177], gastritis [139, 140], cutaneous graft-vs-host disease [178], prostatitis [179], and hyperthyroidism [142]. HLA-DR expression has also been associated with inflammations involving the lung [180], bowel [181, 182], pancreas [183], myocardial tissue [184], and skin [178] and with B-cell derived lymphomas, certain T-cell lymphomas, leukemias, myelomas, and a number of other solid cancers. Although HLA-DR is not normally expressed in skin [178, 185], it has been observed in keratinocytes in skin biopsies taken from individuals with psoriasis [186] and many other diseases caused by an overactive immune response that triggers inflammation, including lupus erythematosus, vitiligo, lymphocytic vasculitis, lichen sclerosus, morphea, lichen planus, erythema nodosum, granulomatous dermatoses, allergic dermatitis, various infectious dermatoses, and Sweet’s syndrome [187, 188].
As expected, normal lymphoid tissues containing APCs bound SH7129. The extremely low levels of SH7129 binding to zona reticularis (adrenal) and white matter (cerebellum) appear related to HLA-DR expression by adrenal cells destined for turnover [131] and microglia or fibrous astrocytes that take on the role of antigen presenting cells in the white matter of the cerebellum of the elderly [132–138] or following brain injury or the onset of neurological disease [132–134, 136, 189, 190]. In each of the isolated cases of a single normal tissue section showing HLA-DR expression, as well as the observed staining of hepatocytes and cells in the white matter of the cerebellum, the level of SH7129 binding to the expressed HLA-DR was only a small fraction of that observed for resting B-cell lymphocytes [54] and was so low that it could only be detected in tissue sections that were not counterstained. The staining of hepatocytes, which do not express HLA-DRs, may be explained by SH7129’s binding to the abundant OATP1B1 and OATP1B3 transporters present in liver.
Based on the results of a series of toxicology and safety studies conducted with SH7139 in dogs and rats at doses of the drug up to 4,000 and 12,000 times, respectively, of the anticipated therapeutic dose (data not shown), the low-level of SHAL binding to these normal tissues does not appear to have an adverse impact on their function. No organ or tissue in the treated animals showed any macroscopic or microscopic indication of pathology or toxicity, B-cell lymphocytes expressing low levels of HLA-DR were not adversely affected, the serum chemistry of the treated animals showed no evidence of liver or renal damage, there were no observed abnormalities in electrocardiography parameters (heart rate, RR interval, PR interval, QRS duration, QT interval or QTc interval), and the Functional Observational Battery performed to assess the central nervous system for pharmacological effects showed no indication of an adverse association with SH7139 exposure.
Consistent with previous reports of HLA-DR expression by a number of different lymphomas and leukemias [30, 37, 191–193], a significant fraction of each of the seven NHL subtypes tested in this study were found to express HLA-DRs that bind SH7129. In addition to providing additional confirmation that HLA-DR is expressed by most, if not all, B-cell lymphoma subtypes, the current results also show the majority of the tumors examined (56% of all cases tested) expressed HLA-DRs recognized by SH7129. Binding was observed in each of the anaplastic large cell lymphoma cases tested. Only about a quarter of the mantle cell lymphomas and one third of the follicular lymphomas were found to bind SH7129. Although the median level of binding was lower than that observed for the other NHL subtypes, one mantle cell lymphoma bound more SH7129 than any other NHL subtype.
Similar results were obtained for sixteen of the eighteen other solid cancers examined. Cervical, ovarian, prostate and colorectal cancers bound considerably more SH7129 than any of the lymphomas and many of the other solid cancers. SH7129 binding comparable to the levels found in NHL were observed in melanoma, kidney, breast, bladder, pancreatic, lung, bone, prostate, larynx, liver, gastric, and thyroid cancers. SH7129 binding was rarely detected in esophageal and head and neck cancers. As observed for the different types of NHL, a wide range in tumor to tumor variability in SH7129 binding was also observed for the majority of the non-hematological cancers expressing HLA-DR.
Although there were too few cases of the non-hematological cancer types represented in the tumor biopsy sets analyzed to be able to draw definitive conclusions regarding cancer type differences in SH7129 binding, the available data suggest differences in HLA-DR target expression and SH7129 binding amongst the different types of liver, larynx, thyroid, ovarian, gastric, cervical, lung, or bone cancers are likely to be small if they differ at all. Only the medullary carcinomas (MC) and invasive ductal carcinomas (IDC) of breast cancer were observed to differ in their expression of HLA-DRs recognized by SH7129. While the amount of SH7129 binding appears to be similar, based on the analysis of a very limited number of invasive ductal cases, the frequency with which the two types of breast cancer expressed the HLA-DR target was quite different. Approximately 4% of the IDC tumors were found to express the target HLA-DRs and bind detectable levels of SH7129. Two thirds of the MC tumors, in contrast, bound SH7129. The larger percentage of medullary carcinomas binding SH7129 in this study are consistent with reports by others who also show a much higher incidence of HLA-DR expression by medullary breast cancers, which typically have heavier and more frequent lymphocytic infiltration [81, 82], compared to invasive ductal carcinomas [194–197].
While SH7139 was originally developed as a therapeutic and imaging agent for B-cell derived lymphomas, the current results suggest SH7139 should also be considered as a viable alternative for treating many other types of cancer. Since the HLA-DR target is unique, SHALs can be used in combination therapies with most other drugs. Because cervical, ovarian and colorectal cancer express much higher levels of the HLA-DRs that bind SH7129, these cancers would be expected to respond even more favorably to SH7139 therapy than NHL. The cell surface HLA-DR molecules to which SH7139 binds are rapidly internalized [198], and tumor cells with significantly larger numbers of HLA-DR molecules on its surface would be expected to more rapidly accumulate SH7139 and achieve much higher intracellular concentrations of the drug.
One unexpected finding in our analysis of SH7129 binding to the non-hematological cancers was that a significant number of pancreatic and liver cancer cases bind the diagnostic, indicating they also express HLA-DRs targeted by SH7139. Although the median level of SH7129 binding was low compared to other cancers, it was similar to some types of NHL, such as the mantle cell lymphomas. Patients diagnosed with pancreatic, liver, and lung cancers currently have the poorest prognoses, with five-year survival rates of only 9%, 19%, and 20% respectively [199]. Therapies targeting the HLA-DRs expressed by pancreatic, liver and lung cancers, either alone or in combination with other drugs, could markedly improve the outcomes for patients diagnosed with these malignancies.
HLA-DR expression by tumor cells, as in normal antigen presenting cells, is often accompanied by the presentation of peptide antigens to T-cell lymphocytes [19, 67, 73, 200–202]. Among these antigens are peptides derived from proteins that bear mutations not found in normal cells [21, 22, 203–207], contain a unique conformational epitope or post-translational modification [208, 209], or whose expression is specific to the tumor cell [26, 210–212] or the normal cell from which the tumor developed [213–218]. One consequence of HLA-DR’s presentation of these abnormal or ‘non-self’ antigenic peptides to T-cells is the induction or enhancement of an immune response that targets the tumor producing the proteins from which the peptides originated [22, 200, 219]. Such responses have been reported to lead to systemic immunity [21] and better prognoses for patients with lymphoma [220], melanoma [221], colorectal [98, 100, 101, 222], gastric [111], ovarian [63] and certain types of laryngeal [107], and breast [81, 223, 224] cancers expressing HLA-DR than others diagnosed with same cancers that do not express HLA-DR. Tumors that do not express MHC class II proteins or have a deletion of the H2-DM gene which prevents the loading of antigenic peptides onto HLA-DRs containing CLIP [225] also appear to be better at escaping immunosurveillance [226–230] which has been suggested to lead to poorer tumor containment [228, 231–233], the outcome of which provides a worse prognosis for the patient [84, 100, 222, 224, 233–238]. While some exceptions have been reported [88, 239–241], this has led to the suggestion that one approach for improving cancer therapy might be to force all cancer cells to become antigen presenting cells [219, 226, 242]. Should such an approach be implemented as a means to trigger the patient’s immune system to mount a more effective antitumor response, it would also sensitize those tumors that do not normally express HLA-DR to SH7139 and other HLA-DR targeted therapeutics.
The correlation between HLA-DR expression and more positive outcomes for patients has not, however, been observed for all types of cancer. HLA-DR expression by acute myelogenous leukemias [241], myelomas [240] and diffuse large B-cell lymphomas [237] have been reported to be highly predictive of poor patient survival. These reports, combined with the fact that a large fraction of so many different types of cancer express HLA-DRs, suggest the presence of tumor associated HLA-DRs may in some cases confer a benefit to the growth or survival of cancer cells. At least two mechanisms that help cancer cells expressing HLA-DR avoid detection by the immune system have been described in support of this idea. Increased expression of HLA-DR by tumors in the absence of the co-stimulatory receptors CD80 or CD86, as one example, has been shown to suppress T-cell activation and tumor infiltration by lymphocytes. In the absence of these receptors the tumor not only evades detection, but its cells can more easily kill those lymphocytes that do invade tumor tissue [243]. Expression of the transcription regulator CD74, which always accompanies expression of HLA-DR, and its binding to the chemokine migration inhibitory factor (MIF) has also been shown to improve tumor cell survival by suppressing CD-44 mediated-apoptosis [244] and modulating other pathways that involve immune regulation and cell survival [245].
CD74, which is also called “Ii”, has a second function as a chaperone protein that is co-expressed along with HLA-DR to facilitate its proper folding, protect its antigen binding site from premature loading with peptides that will be presented to T-cells, and to direct the HLA-DR proteins to the late endosomal lysosomal antigen-processing compartments containing the antigenic proteins and peptides they will eventually present to T-cells [246]. In the endoplasmic reticulum, the α and β-subunits of HLA-DR bind to trimers of CD74 to form large nonameric complexes [247]. Following the transport of these complexes across the trans-Golgi network into late endosomal lysosomal antigen-processing compartments containing the antigenic proteins and peptides, the CD74 in the complex is processed by a series of proteolytic cleavages leaving the class II-associated Ii chain peptide (CLIP) bound inside the peptide binding site of HLA-DR. Upon removal of CLIP by HLA-DM, the antigenic peptides then bind inside the HLA-DR peptide binding site. In addition to those HLA-DR: CD74 complexes trafficked to the endosome, some of the complexes are also transported directly to the plasma membrane. Many of these complexes are under populated by HLA-DRs, leaving CD74 molecules on the surface of the cell that can bind to MIF and function as a transcription factor to promote tumor cell survival [245, 248].
The observation that HLA-DR expression by some cancers provides a benefit to the patient and its expression by other cancers provides a benefit to the tumor demonstrates the complexity of the cellular mechanisms that contribute to tumor cell survival and the immune system’s response to cells considered non-self. In the absence of other contributing factors, HLA-DR expression by tumors can stimulate an immune response targeting the tumor in those cases where the peptide antigens are immunogenic when presented to T-cells [21, 200, 219]. In cases where a sufficiently strong anti-tumor response cannot be induced by the antigens presented by HLA-DR, the co-expression of receptors such as CD74 that promote tumor cell survival may dominate the result. In other cases the outcome may be dictated by unrelated immunosurveillance evasion processes that come into play, such as increased production of T-reg lymphocytes [249], the secretion of immunosuppressive molecules (programmed death-ligand 1, TGF-β, indoleamine 2,3-dioxygenase, IL-10 or Fc receptor-like 6) that induce pathways that limit or repress the activity of cytotoxic T-cell lymphocytes or NK cells [250, 251], blocking of inhibitory checkpoints for immune activity [252], blockage of death receptor signaling [253, 254], downregulation or loss of MHC class I molecules [255–258], defects or alterations in MHC class I antigen processing [259] or interferon signal transduction, or the loss of gene function [260].
The SH7129 staining protocol used in this study provided a simple method to screen biopsy sections to identify tumors expressing the HLA-DRs SH7139 targets and to also obtain an estimate of the amount of SH7139 one might expect to bind to different tumors. In contrast to the Lym-1 antibody, which could also be used to detect the expression of a subset of HLA-DRs (including HLA-DR10), SH7129 works well for staining formalin fixed tissues. During our analysis of the samples, however, several limitations of the current method were identified that will need to be addressed as the diagnostic and staining protocol are developed further. One relates to the observation that the detection of SH7129 binding using horse-radish peroxidase and the current DAB substrate cannot be used to quantify HLA-DR target expression and SH7129 binding to tumor cells that are pigmented or contain significant amounts of melanin. The image analysis approach we used to quantify SH7129 binding to the tumor cells could not distinguish between the brown insoluble product formed by horse-radish peroxidase oxidation of the DAB substrate and the similarly colored melanin pigment present in a number of the melanoma biopsies (Figure 14A and 14B) or the melanin present in the basal keratinocytes found in normal skin tissue. Melanin does not fluoresce under visible or ultraviolet light excitation. One solution might be to replace SH7129 with a fluorochrome-tagged analog of SH7139, provided the presence of the fluorescent dye does not influence HLA-DR binding or alter the reagent’s selectivity.
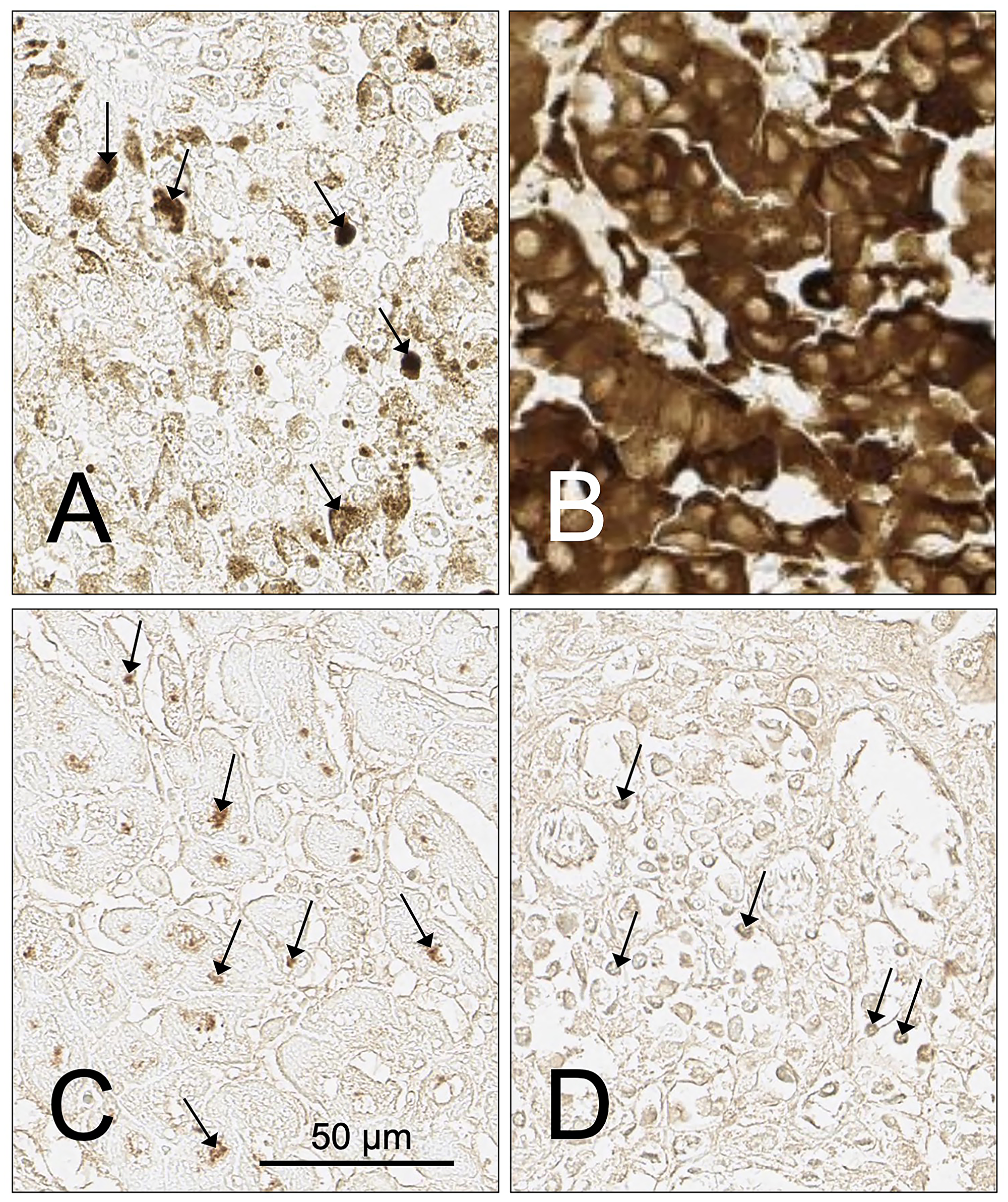
Figure 14: Examples of tumor cases with infiltrating lymphocytes or cells containing pigment that impact or prevent the assessment of SH7129 binding using DAB as a substrate. (A) Tissue sample ME481AB6, unstained moderately pigmented melanoma showing small pigment particles (arrows) similar in color to the insoluble product produced by SAHRP oxidation of DAB. (B) Tissue sample ME481AA8, unstained heavily pigmented melanoma section. (C) Tissue sample M672708, SLL/CLL tumor section showing SH7129 stained lymphocytes and macrophages (arrows). (D) Tissue sample BR807C3, breast medullary carcinoma section showing SH7129 stained infiltrating lymphocytes and macrophages (arrows). The images were captured at 40× magnification. The scale bar is the same for all images.
Other factors, such as the presence of large numbers of tumor infiltrating lymphocytes (TILs), macrophages or dendritic cells that also express HLA-DR (Figure 14C and 14D), could make it difficult to accurately quantify SH7129 binding by the tumor cells. If the numbers of TILs in the section are small, the impact will be negligible. But if large numbers of these cells are present in the tissue, the extent of SH7129 binding to the tumor cells can only be determined in areas free of the antigen presenting cells. The lack of structural uniformity within the tissue being imaged can also present quantification issues. While all cores in a tumor microarray are usually cut to the same thickness, differences in thickness can contribute to variation in the amount of cellular material that is stained and the amount of diagnostic bound. This is particularly important when the target protein, such as HLA-DR, is not only found bound to the membrane but is also abundant in the cytoplasm. Local variation in the optical properties of unstained cells and supporting or connective tissue also contribute background absorption and attenuation of the light transmitted through the tissue that adds to the signal provided by the oxidized product of the DAB substrate. Tumor to tumor differences in density of cells packed within the section, disruptions in the integrity of the core (tears or missing tissue) or the presence of vacuoles, capillaries, or necrotic regions can change the amount of tumor tissue that is available to bind the diagnostic. The use of sequentially cut cores for the stained and unstained control slides should help minimize these contributions to variability and reduce errors in the quantification of SH7129 binding to tumor cells.
Materials and Methods
Chemicals and reagents
The chemicals and reagents used in this study were obtained from the following sources: Key Organics, Camelford, UK (4-[4-[(4-chlorophenyl) methyl] piperazin-1-yl]-3-nitrobenzoic acid and 3-[2-[3-chloro-5-(trifluoromethyl) pyridin-2-yl] oxyanilino]-3-oxopropanoic acid), Sigma-Aldrich, St. Louis, MO (3,3′-diaminobenzidine tetrahydrochloride, hydroxybenzotriazole, trifluoroacetic acid, anhydrous dimethylformamide, 2-(1H-benzotriazol-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate, Wang trityl chloride resin, phosphate buffered saline, citrate buffer, hematoxylin, xylene, ethanol), Macrocyclics, Plano, TX, USA (DOTA NHS ester), TCI, Shanghai, China (Dabsyl chloride), GL Biochem, Shanghai, China (Fmoc-D-Lys (Boc)-OH, Fmoc-D-Lys (Dde)-OH), Fmoc-L-Val-OH), Biomatrik, Zhejiang, China (Fmoc-8-amino-3,6-dioxaoctanoic acid), Electron Microscopy Sciences, Hatfield, PA, USA (Permount, 30% hydrogen peroxide), VWR Scientific, Radnor, PA, USA (acetonitrile, biotin N-hydroxysuccinimide ester, N, N-diisopropylethylamine) Roche Molecular Diagnostics, Pleasanton, CA, USA (Ventana Endogenous Biotin Blocking kit), Leica Biosystems Inc, Buffalo Grove, IL, USA (Leica BOND RX Automated Slide Stainer reagents) and Vector Labs, Burlingame, CA, USA (streptavidin conjugated horse-radish peroxidase).
Synthesis of the biotin analog of SH7139
SH7129 was synthesized and purified by AmbioPharm Inc., (North Augusta, SC, USA) using a modification of the procedure described previously [51, 52]. Briefly, the SHAL was synthesized using solid phase chemistry by the stepwise attachment to a Wang resin of Fmoc-D-Lys#1(Boc)-OH, Fmoc-AEEA-OH#1 (Fmoc-8-amino-3,6-dioxaoctanoic acid), Fmoc-D-Lys#2(Dde)-OH, Fmoc-AEEA-OH#2, Fmoc-L-Val-OH, and Dabsyl chloride using standard Fmoc (N-9-fluorenylmethoxycarbonyl) chemistry with HBTU (2-(1H-benzotriazol-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate)/HOBt (Hydroxybenzotriazole)/DIPEA (N, N-Diisopropylethylamine) as the coupling reagents. The side chain amino group of D-Lys#2-(Dde)-OH was deprotected with 4% hydrazine in dimethylformamide (DMF) and then coupled to Fmoc-D-Lys#3-(Dde)-OH using the same coupling procedure. Fmoc-AEEA-OH#3 was next coupled to deprotected D-Lys#3(Dde) and 4-(4-(4-chlorobenzyl) piperazine)-3-nitrobenzenecarboxylic acid (Cb ligand) was then linked to the deprotected AEEA-OH#3 using the same coupling procedure. The third ligand Ct (3-(2-((3-chloro-5-(trifluoromethyl)-2-pyridinyl) oxy)-anilino)-3-oxopropanoic acid) was then attached to the deprotected ε-amine of D-Lys#3. The assembled free amine form of the SHAL was cleaved from the resin, deprotected and subsequently precipitated as a crude solid. The crude product was purified by standard RP-HPLC methods and isolated by lyophilization. Biotin was attached to the free amine on the terminal lysine by dissolving the SHAL in anhydrous DMF, N, N-Diisopropylethylamine (DIEA) and adding solid biotin N-hydroxysuccinimide ester (biotinyl-OSu). The mixture was nutated for 15 min, and the reaction was monitored by analytical HPLC. Upon completion, the reaction solution was diluted with a small volume of water/acetonitrile (50/50) containing 1% trifluoroacetic acid (TFA) and purified by HPLC. The purified SH7129 was lyophilized and then analyzed by LC/MS and NMR spectroscopy to determine its purity (96.2%) and confirm its molecular mass (2165.6 Da) and structure, respectively.
Tissue and tumor microarrays
The normal human tissue microarrays (FDA808-1 and FDA808-2) containing fixed and paraffin embedded sections of twenty-four different tissues obtained from three individuals and tumor microarrays (TMAs) containing fixed and paraffin embedded tumor biopsy sections obtained from patients diagnosed with different non-Hodgkin’s lymphoma subtypes (LM241, LM242, LM482, LY804, LY1001b, NHL801, NHL802, NHL803, OD-CT-LyMly02) and other solid cancers (BC07014a, BC08118, BC041115d, BC12011b, BM483, BR807, CXC1501, ESC1021, HN483, LP801, LUC1021, LVC1501, Me481a, Me1004e, OD-CT-DgCol04, OD-CT-DgStm01, OD-CT-LyMly02, OS802, OVC1501, PA961c, PR803c, T111, T291, TH802) were obtained from U. S. Biomax (Rockville, MD, USA). An additional set of diffuse large B-cell lymphoma, mantle cell lymphoma, follicular lymphoma and SLL/CLL TMAs were obtained from Dr. John G. Gribben, Barts Cancer Institute, Queen Mary University of London, Charterhouse Square, London, UK.
SH7129 staining protocol
SH7129 was prepared as a stock solution by dissolving 10 mg of the dry compound in 1 ml dimethyl sulfoxide. The formalin fixed normal tissue and tumor microarrays were stained using a Leica BOND RX Automated Slide Stainer (Leica Biosystems Inc., Buffalo Grove, IL, USA) to maximize slide-to-slide uniformity in staining and processing. The fixed slides were deparaffinized using the Leica dewax solution, rehydrated with an alcohol series (100%, 95%, 70% and 30% for 4 min each) followed by antigen retrieval in citrate buffer at pH 6 and 90°C for 20 min. After performing a 5 min hydrogen peroxide block, the slides were washed three times with BOND Wash Solution, endogenous biotin was blocked using the Ventana Endogenous Biotin Blocking kit (Roche Molecular Diagnostics, Pleasanton, CA, USA), the slides were washed three additional times with BOND Wash Solution, and then stained with SH7129 (100 μg/ml in PBS, 1% DMSO) for 30 min. Following three washes with BOND Washing Solution, the slides were treated with Streptavidin-horse radish peroxidase (SAHRP) for 30 min, washed 3 times with BOND Wash Solution and once with deionized water, treated with Mixed DAB (3,3-diaminobenzidine) Refine for 10 min, and then washed four times with deionized water, once with BOND Wash Solution and a final deionized water wash as per the BOND Polymer Refine IHC protocol (Histowiz Inc., Brooklyn, NY, USA). The SH7129 stained tumor microarray slides were not counterstained with hematoxylin. The slides were then dehydrated by immersion in an alcohol series (30%, 70%, 95% and 100% for 4 min each), cleared with xylene and mounted with Permount.
Analysis of SH7129 binding to NHL and other solid tumors
Whole slide images of the SH7129 stained and control (duplicate slide cut from same core treated with PBS instead of SH7129) normal tissue and tumor microarrays were captured at 40× magnification using an Aperio AT2 Digital Pathology Scanner (Leica Biosystems, Buffalo Grove, IL, USA). SH7129 binding or lack of binding to the cells was confirmed by visual inspection. Cells expressing the HLA-DRs that bind SH7129 showed stain associated with both the membranes and cytoplasm. Cell-to-cell variation in SH7129 binding was determined by performing densitometric analyses of fifty (50) individual tumor cells from a representative moderate to high SH7129 binding tumor for six of the NHL subtypes using NIH ImageJ version 1.42 software [262]. An effort was made during the analysis to include cells representing the full range of SH7129 binding. Individual cell data were not obtained for the small lymphocytic lymphomas due to our inability to accurately define the individual cell boundaries in these tumors. To estimate the tumor-to-tumor variation in SH7129 binding, additional lower magnification digital images containing the array of cores for the two slides (SH7129 stained and control without SH7129) were captured at 10× magnification, the images were inverted, and the amount of bound SH7129 was determined by densitometric analysis of each tumor section using the same NIH ImageJ software. Integrated density data were collected from a 384-pixel area of each core and from ten blank (background) 384-pixel areas distributed across the slide near or between the cores. Core sections containing voids or tears (missing tissue), lacking a corresponding core in the control slide, or obtained from pigmented tumors were not analyzed. In cases where there were duplicate or triplicate cores for each biopsy on the slides, the data obtained from the analyses of the replicates were averaged. The amount of bound SH7129 (per 384-pixel area) was then calculated for each biopsy sample as follows:
Bound SH7129 = (IntDenSH7129 – IntDenSH7129Bkg) – (IntDenNoSH7129 – IntDenNoSH7129Bkg)
where IntDenSH7129 is the integrated density of the biopsy section stained with SH7129, IntDenSH7129Bkg is the mean of the integrated densities of the ten blank regions of the SH7129 stained slide, IntDenNoSH7129 is the integrated density of the biopsy section that was processed for staining without SH7129, and IntDenNoSH7129Bkg is the mean of the integrated density of the ten blank regions of the control slide processed for staining without SH7129.
The data were analyzed and plotted using GraphPad Prism version 8.1.2. Statistical analyses of two groups of data were performed using an unpaired t-test. Analyses of three or more groups of data in which the standard deviations of the groups were similar were analyzed using a one-way ANOVA test followed by Tukey’s multiple comparison test. Data sets containing three or more groups in which the standard deviations of the groups being compared were different were analyzed using both a Brown-Forsythe and Welch’s ANOVA test followed by Dunnett’s T3 multiple comparisons test. In both cases in which three or more groups were compared, the assumption was made that the data fit a Gaussian distribution based on the observation that the amount of SH7129 bound per biopsy case did roughly fit a Gaussian distribution.
Conclusions
The tumor biopsy binding studies conducted with SH7129 have shown the HLA-DRs targeted by SH7139 are expressed by many different types of cancer. A significant fraction of the cases in each of the seven subtypes of B-cell lymphomas were found to express these HLA-DRs, as indicated by the binding of SH7129, and the range of expression varied by as much as 10 to 100-fold. Plasma cell myelomas and seventeen other types of non-hematological cancers have also been found to express the HLA-DRs recognized by these SHALs, with some at levels much higher than many types of NHL. Cervical, ovarian, prostate and colorectal cancers bound the most SH7129, followed by non-Hodgkin’s lymphomas, plasma cell myelomas, lung, liver, bone, kidney, thyroid, melanoma, breast, laryngeal, gastric, pancreatic and bladder cancers. Only a few cases of head and neck and esophageal cancers bound SH7129. Comparisons of SH7129 binding to different types of nine non-hematological cancers only revealed a significant difference for two types of cervical and two types of breast cancer. SH7129 binding did not correlate well with tumor grade in the nine cancers for which data was available. The results obtained in this study also provide additional evidence of SH7129’s utility as a diagnostic to identify tumors expressing the HLA-DRs targeted by SH7139 and to estimate the magnitude of their expression. SH7129 should prove useful in future clinical trials as a surrogate biomarker detection reagent for screening formalin-fixed paraffin embedded biopsy sections to identify patients with HLA-DR expressing tumors targeted by SH7139, Lym-1 antibody in Lym-1 CAR T-cell therapy clinical studies [261], and SHALs administered in combination with other drugs to individualize therapy and optimize treatment protocols for each patient. The observation of moderate to high levels of HLA-DRs expression by so many of the tumors tested suggest there are patients diagnosed with many cancers, in addition to NHL, that may benefit from the development of oncology drugs that target HLA-DR.
Abbreviations
AEEA: 8-amino-3,6-dioxaoctanoic acid; ALCL: anaplastic large cell lymphoma; ANOVA: analysis of variance; BL: Burkitt lymphoma; Cb: 4-[4-[(4-chlorophenyl)methyl]piperazin-1-yl]-3-nitrobenzoic acid; CD: cluster of differentiation; Ct: 3-[2-[3-chloro-5-(trifluoromethyl)pyridin-2-yl]oxyanilino]-3-oxopropanoic acid; Da: Daltons; DAB: 3,3′-diaminobenzidine; Dde: N-(1-(4,4-dimethyl-2,6-dioxocyclohexylidene)ethyl); DIEA: Diisopropylethylamine; DIPEA: N,N-Diisopropylethylamine; DLBCL: diffuse large B-cell lymphoma; DMF: dimethylformamide; DOTA: 1,4,7,10-tetraazacyclododecane-1,4,7,10-tetraacetic acid; Dv: 4-(Dimethylamino)azobenzene-4'-sulfonyl-L-valine; FL: follicular lymphoma; Fmoc: fluorenylmethoxycarbonyl; HAMA: human antibody-mouse antibody; HBTU: 2-(1H-benzotriazol-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate; H&E: hematoxylin and eosin; HED: human equivalent dose; HLA-DR: human leukocyte antigen-DR type; HOBT: Hydroxybenzotriazole; HPLC: high performance liquid chromatography; IDC: invasive ductal carcinoma of the breast; IHC: immunohistochemistry; IND: investigational new drug; LC: liquid chromatography; LC/MS: liquid chromatography-mass spectrometry; MALT: mucosa-associated lymphoid tissue; MC: medullary carcinoma of the breast; MCL: mantle cell lymphoma; MHC: major histocompatibility complex; NHL: non-Hodgkin’s lymphoma; NMR: nuclear magnetic resonance; OATP: organic anion transporter polypeptide; OSu: N-hydroxysuccinimide ester; PBS: phosphate buffered saline; SAHRP: streptavidin horse-radish peroxidase; SHAL: Selective high affinity ligand; SLL: small lymphocytic lymphoma; TFA: trifluoroacetic acid; TILs: tumor infiltrating lymphocytes; TMAs: tumor microarrays.
ACKNOWLEDGMENTS AND FUNDING
Funding for this research was provided by the National Cancer Institute (SBIR Award R44CA159844) and by SHAL Technologies Inc.
CONFLICTS OF INTEREST
R. Balhorn and M. Cosman Balhorn are co-founders and employees of SHAL Technologies Inc.
References
1. Caron PC, Jurcic JG, Scott AM, Finn RD, Divgi CR, Graham MC, Jureidini IM, Sgouros G, Tyson D, Old LJ. A phase 1B trial of humanized monoclonal antibody M195 (anti-CD33) in myeloid leukemia: specific targeting without immunogenicity. Blood. 1994; 83:1760–1768. https://doi.org/10.1182/blood.V83.7.1760.1760. [PubMed].
2. Cartron G, Watier H, Golay J, Solal-Celigny P. From the bench to the bedside: ways to improve rituximab efficacy. Blood. 2004; 104:2635–2642. https://doi.org/10.1182/blood-2004-03-1110. [PubMed].
3. Kaminski MS, Tuck M, Estes J, Kolstad A, Ross CW, Zasadny K, Regan D, Kison P, Fisher S, Kroll S, Wahl RL. 131I-tositumomab therapy as initial treatment for follicular lymphoma. N Engl J Med. 2005; 352:441–449. https://doi.org/10.1056/NEJMoa041511. [PubMed].
4. Leonard JP, Coleman M, Ketas JC, Chadburn A, Furman R, Schuster MW, Feldman EJ, Ashe M, Schuster SJ, Wegener WA, Hansen HJ, Ziccardi H, Eschenberg M, et al. Epratuzumab, a humanized anti-CD22 antibody, in aggressive non-Hodgkin’s lymphoma: phase I/II clinical trial results. Clin Cancer Res. 2004; 10:5327–5334. https://doi.org/10.1158/1078-0432.CCR-04-0294. [PubMed].
5. Lim SH, Beers SA, French RR, Johnson PW, Glennie MJ, Cragg MS. Anti-CD20 monoclonal antibodies: historical and future perspectives. Haematologica. 2010; 95:135–143. https://doi.org/10.3324/haematol.2008.001628. [PubMed].
6. Pfreundschuh M, Schubert J, Ziepert M, Schmits R, Mohren M, Lengfelder E, Reiser M, Nickenig C, Clemens M, Peter N, Bokemeyer C, Eimermacher H, Ho A, et al, and German High-Grade Non-Hodgkin Lymphoma Study Group (DSHNHL). Six versus eight cycles of bi-weekly CHOP-14 with or without rituximab in elderly patients with aggressive CD20+ B-cell lymphomas: a randomised controlled trial (RICOVER-60). Lancet Oncol. 2008; 9:105–116. https://doi.org/10.1016/S1470-2045(08)70002-0. [PubMed].
7. Plosker GL, Figgitt DP. Rituximab: a review of its use in non-Hodgkin’s lymphoma and chronic lymphocytic leukaemia. Drugs. 2003; 63:803–43. https://doi.org/10.2165/00003495-200363080-00005. [PubMed].
8. Press OW, Unger JM, Braziel RM, Maloney DG, Miller TP, Leblanc M, Fisher RI, and Southwest Oncology Group. Phase II trial of CHOP chemotherapy followed by tositumomab/iodine I-131 tositumomab for previously untreated follicular non-Hodgkin’s lymphoma: five-year follow-up of Southwest Oncology Group Protocol S9911. J Clin Oncol. 2006; 24:4143–4149. https://doi.org/10.1200/JCO.2006.05.8198. [PubMed].
9. Rieger K, Von Grunhagen U, Fietz T, Thiel E, Knauf W. Efficacy and tolerability of alemtuzumab (CAMPATH-1H) in the salvage treatment of B-cell chronic lymphocytic leukemia–change of regimen needed? Leuk Lymphoma. 2004; 45:345–349. https://doi.org/10.1080/10428190310001598017. [PubMed].
10. Topp MS, Gokbuget N, Zugmaier G, Klappers P, Stelljes M, Neumann S, Viardot A, Marks R, Diedrich H, Faul C, Reichle A, Horst HA, Bruggemann M, et al. Phase II trial of the anti-CD19 bispecific T cell-engager blinatumomab shows hematologic and molecular remissions in patients with relapsed or refractory B-precursor acute lymphoblastic leukemia. J Clin Oncol. 2014; 32:4134–4140. https://doi.org/10.1200/JCO.2014.56.3247. [PubMed].
11. van de Donk NW, Janmaat ML, Mutis T, Lammerts van Bueren JJ, Ahmadi T, Sasser AK, Lokhorst HM, Parren PW. Monoclonal antibodies targeting CD38 in hematological malignancies and beyond. Immunol Rev. 2016; 270:95–112. https://doi.org/10.1111/imr.12389. [PubMed].
12. Brennick CA, George MM, Corwin WL, Srivastava PK, Ebrahimi-Nik H. Neoepitopes as cancer immunotherapy targets: key challenges and opportunities. Immunotherapy. 2017; 9:361–371. https://doi.org/10.2217/imt-2016-0146. [PubMed].
13. Kissick HT, Sanda MG, Dunn LK, Arredouani MS. Immunization with a peptide containing MHC class I and II epitopes derived from the tumor antigen SIM2 induces an effective CD4 and CD8 T-cell response. PLoS One. 2014; 9:e93231. https://doi.org/10.1371/journal.pone.0093231. [PubMed].
14. Nishimura MI, Avichezer D, Custer MC, Lee CS, Chen C, Parkhurst MR, Diamond RA, Robbins PF, Schwartzentruber DJ, Rosenberg SA. MHC class I-restricted recognition of a melanoma antigen by a human CD4+ tumor infiltrating lymphocyte. Cancer Res. 1999; 59:6230–6238. [PubMed].
15. Rozanov DV, Rozanov ND, Chiotti KE, Reddy A, Wilmarth PA, David LL, Cha SW, Woo S, Pevzner P, Bafna V, Burrows GG, Rantala JK, Levin T, et al. MHC class I loaded ligands from breast cancer cell lines: A potential HLA-I-typed antigen collection. J Proteomics. 2018; 176:13–23. https://doi.org/10.1016/j.jprot.2018.01.004. [PubMed].
16. Shingler WH, Chikoti P, Kingsman SM, Harrop R. Identification and functional validation of MHC class I epitopes in the tumor-associated antigen 5T4. Int Immunol. 2008; 20:1057–1066. https://doi.org/10.1093/intimm/dxn063. [PubMed].
17. Sotiriadou R, Perez SA, Gritzapis AD, Sotiropoulou PA, Echner H, Heinzel S, Mamalaki A, Pawelec G, Voelter W, Baxevanis CN, Papamichail M. Peptide HER2(776-788) represents a naturally processed broad MHC class II-restricted T cell epitope. Br J Cancer. 2001; 85:1527–1534. https://doi.org/10.1054/bjoc.2001.2089. [PubMed].
18. Dengjel J, Nastke MD, Gouttefangeas C, Gitsioudis G, Schoor O, Altenberend F, Muller M, Kramer B, Missiou A, Sauter M, Hennenlotter J, Wernet D, Stenzl A, et al. Unexpected abundance of HLA class II presented peptides in primary renal cell carcinomas. Clin Cancer Res. 2006; 12:4163–4170. https://doi.org/10.1158/1078-0432.CCR-05-2470. [PubMed].
19. Doonan BP, Haque A. Prostate Cancer Immunotherapy: Exploiting the HLA Class II Pathway in Vaccine Design. J Clin Cell Immunol. 2015; 6:351. https://doi.org/10.4172/2155-9899.1000351. [PubMed].
20. Hohn H, Pilch H, Gunzel S, Neukirch C, Hilmes C, Kaufmann A, Seliger B, Maeurer MJ. CD4+ tumor-infiltrating lymphocytes in cervical cancer recognize HLA-DR-restricted peptides provided by human papillomavirus-E7. J Immunol. 1999; 163:5715–22. [PubMed].
21. Novellino L, Renkvist N, Rini F, Mazzocchi A, Rivoltini L, Greco A, Deho P, Squarcina P, Robbins PF, Parmiani G, Castelli C. Identification of a mutated receptor-like protein tyrosine phosphatase kappa as a novel, class II HLA-restricted melanoma antigen. J Immunol. 2003; 170:6363–6370. https://doi.org/10.4049/jimmunol.170.12.6363. [PubMed].
22. Pieper R, Christian RE, Gonzales MI, Nishimura MI, Gupta G, Settlage RE, Shabanowitz J, Rosenberg SA, Hunt DF, Topalian SL. Biochemical identification of a mutated human melanoma antigen recognized by CD4(+) T cells. J Exp Med. 1999; 189:757–766. https://doi.org/10.1084/jem.189.5.757. [PubMed].
23. Schroers R, Shen L, Rollins L, Xiao Z, Sonderstrup G, Slawin K, Huang XF, Chen SY. Identification of MHC class II-restricted T-cell epitopes in prostate-specific membrane antigen. Clin Cancer Res. 2003; 9:3260–3271. [PubMed].
24. Shimato S, Maier LM, Maier R, Bruce JN, Anderson RC, Anderson DE. Profound tumor-specific Th2 bias in patients with malignant glioma. BMC Cancer. 2012; 12:561. https://doi.org/10.1186/1471-2407-12-561. [PubMed].
25. Topalian SL, Rivoltini L, Mancini M, Markus NR, Robbins PF, Kawakami Y, Rosenberg SA. Human CD4+ T cells specifically recognize a shared melanoma-associated antigen encoded by the tyrosinase gene. Proc Natl Acad Sci U S A. 1994; 91:9461–9465. https://doi.org/10.1073/pnas.91.20.9461. [PubMed].
26. Zeng G, Wang X, Robbins PF, Rosenberg SA, Wang RF. CD4(+) T cell recognition of MHC class II-restricted epitopes from NY-ESO-1 presented by a prevalent HLA DP4 allele: association with NY-ESO-1 antibody production. Proc Natl Acad Sci U S A. 2001; 98:3964–3969. https://doi.org/10.1073/pnas.061507398. [PubMed].
27. Cardillo TM, Govindan SV, Zalath MB, Rossi DL, Wang Y, Chang CH, Goldenberg DM. IMMU-140, a Novel SN-38 Antibody-Drug Conjugate Targeting HLA-DR, Mediates Dual Cytotoxic Effects in Hematologic Cancers and Malignant Melanoma. Mol Cancer Ther. 2018; 17:150–160. https://doi.org/10.1158/1535-7163.MCT-17-0354. [PubMed].
28. DeNardo SJ, DeNardo GL, O’Grady LF, Levy NB, Mills SL, Macey DJ, McGahan JP, Miller CH, Epstein AL. Pilot studies of radioimmunotherapy of B cell lymphoma and leukemia using I-131 Lym-1 monoclonal antibody. Antibody Immunoconjugates Radiopharm. 1988; 1:17–33.
29. DeNardo GL, Lamborn KR, Goldstein DS, Kroger LA, DeNardo SJ. Increased survival associated with radiolabeled Lym-1 therapy for non-Hodgkin’s lymphoma and chronic lymphocytic leukemia. Cancer. 1997; 80:2706–2711. https://doi.org/10.1002/(SICI)1097-0142(19971215)80:12+<2706::AID-CNCR49>3.0.CO;2-D. [PubMed].
30. DeNardo GL, DeNardo SJ, Lamborn KR, Goldstein DS, Levy NB, Lewis JP, O’Grady LF, Raventos A, Kroger LA, Macey DJ, McGahan JP, Mills SL, Shen S. Low-dose, fractionated radioimmunotherapy for B-cell malignancies using 131I-Lym-1 antibody. Cancer Biother Radiopharm. 1998; 13:239–254. https://doi.org/10.1089/cbr.1998.13.239. [PubMed].
31. DeNardo GL, DeNardo SJ, Goldstein DS, Kroger LA, Lamborn KR, Levy NB, McGahan JP, Salako Q, Shen S, Lewis JP. Maximum-tolerated dose, toxicity, and efficacy of (131)I-Lym-1 antibody for fractionated radioimmunotherapy of non-Hodgkin’s lymphoma. J Clin Oncol. 1998; 16:3246–3256. https://doi.org/10.1200/JCO.1998.16.10.3246. [PubMed].
32. Lamborn KR, DeNardo GL, DeNardo SJ, Goldstein DS, Shen S, Larkin EC, Kroger LA. Treatment-related parameters predicting efficacy of Lym-1 radioimmunotherapy in patients with B-lymphocytic malignancies. Clin Cancer Res. 1997; 3:1253–1260. [PubMed].
33. O’Donnell RT, DeNardo GL, Kukis DL, Lamborn KR, Shen S, Yuan A, Goldstein DS, Carr CE, Mirick GR, DeNardo SJ. A clinical trial of radioimmunotherapy with 67Cu-2IT-BAT-Lym-1 for non-Hodgkin’s lymphoma. J Nucl Med. 1999; 40:2014–2020. [PubMed].
34. Stein R, Qu Z, Chen S, Solis D, Hansen HJ, Goldenberg DM. Characterization of a humanized IgG4 anti-HLA-DR monoclonal antibody that lacks effector cell functions but retains direct antilymphoma activity and increases the potency of rituximab. Blood. 2006; 108:2736–2744. https://doi.org/10.1182/blood-2006-04-017921. [PubMed].
35. Stein R, Gupta P, Chen X, Cardillo TM, Furman RR, Chen S, Chang CH, Goldenberg DM. Therapy of B-cell malignancies by anti-HLA-DR humanized monoclonal antibody, IMMU-114, is mediated through hyperactivation of ERK and JNK MAP kinase signaling pathways. Blood. 2010; 115:5180–5190. https://doi.org/10.1182/blood-2009-06-228288. [PubMed].
36. Stephens DM, Starodub AN, Byrd JC, Horne H, Wegener WA, Goldenberg DM, Christian BA. Subcutaneous injections of IMMU-114 (anti-HLA-DR IgG4 monoclonal antibody): Initial results of a Phase I First-in-Man Study in hematologic malignancies. Blood. 2015; 126:2740. https://doi.org/10.1182/blood.V126.23.2740.2740.
37. Epstein AL, Marder RJ, Winter JN, Stathopoulos E, Chen FM, Parker JW, Taylor CR. Two new monoclonal antibodies, Lym-1 and Lym-2, reactive with human B-lymphocytes and derived tumors, with immunodiagnostic and immunotherapeutic potential. Cancer Res. 1987; 47:830–840. [PubMed].
38. Raftery MJ, Seron D, Koffman G, Hartley B, Janossy G, Cameron JS. The relevance of induced class II HLA antigens and macrophage infiltration in early renal allograft biopsies. Transplantation. 1989; 48:238–243. https://doi.org/10.1097/00007890-198908000-00011. [PubMed].
39. Rose LM, Deng CT, Scott SL, Xiong CY, Lamborn KR, Gumerlock PH, DeNardo GL, Meares CF. Critical Lym-1 binding residues on polymorphic HLA-DR molecules. Mol Immunol. 1999; 36:789–797. https://doi.org/10.1016/S0161-5890(99)00083-8. [PubMed].
40. Denardo GL, Denardo SJ, Kukis DL, O’Donnell RT, Shen S, Goldstein DS, Kroger LA, Salako Q, Denardo DA, Mirick GR, Mausner LF, Srivastava SC, Meares CF. Maximum tolerated dose of 67Cu-2IT-BAT-LYM-1 for fractionated radioimmunotherapy of non-Hodgkin’s lymphoma: a pilot study. Anticancer Res. 1998; 18:2779–2788. [PubMed].
41. Alig SK, Dreyling M, Seppi B, Aulinger B, Witkowski L, Rieger CT. Severe cytokine release syndrome after the first dose of Brentuximab Vedotin in a patient with relapsed systemic anaplastic large cell lymphoma (sALCL): a case report and review of literature. Eur J Haematol. 2015; 94:554–557. https://doi.org/10.1111/ejh.12396. [PubMed].
42. Casanova Estruch B. Safety profile and practical considerations of monoclonal antibody treatment. Neurologia. 2013; 28:169–178. https://doi.org/10.1016/j.nrl.2011.02.004. [PubMed].
43. de Vos S, Forero-Torres A, Ansell SM, Kahl B, Cheson BD, Bartlett NL, Furman RR, Winter JN, Kaplan H, Timmerman J, Whiting NC, Drachman JG, Advani R. A phase II study of dacetuzumab (SGN-40) in patients with relapsed diffuse large B-cell lymphoma (DLBCL) and correlative analyses of patient-specific factors. J Hematol Oncol. 2014; 7:44. https://doi.org/10.1186/1756-8722-7-44. [PubMed].
44. Freeman CL, Morschhauser F, Sehn L, Dixon M, Houghton R, Lamy T, Fingerle-Rowson G, Wassner-Fritsch E, Gribben JG, Hallek M, Salles G, Cartron G. Cytokine release in patients with CLL treated with obinutuzumab and possible relationship with infusion-related reactions. Blood. 2015; 126:2646–2649. https://doi.org/10.1182/blood-2015-09-670802. [PubMed].
45. Gruber R, van Haarlem LJ, Warnaar SO, Holz E, Riethmuller G. The human antimouse immunoglobulin response and the anti-idiotypic network have no influence on clinical outcome in patients with minimal residual colorectal cancer treated with monoclonal antibody CO17-1A. Cancer Res. 2000; 60:1921–1926. [PubMed].
46. Hansel TT, Kropshofer H, Singer T, Mitchell JA, George AJ. The safety and side effects of monoclonal antibodies. Nat Rev Drug Discov. 2010; 9:325–338. https://doi.org/10.1038/nrd3003. [PubMed].
47. Mirick GR, Bradt BM, Denardo SJ, Denardo GL. A review of human anti-globulin antibody (HAGA, HAMA, HACA, HAHA) responses to monoclonal antibodies. Not four letter words. Q J Nucl Med Mol Imaging. 2004; 48:251–257. [PubMed].
48. Rotz SJ, Leino D, Szabo S, Mangino JL, Turpin BK, Pressey JG. Severe cytokine release syndrome in a patient receiving PD-1-directed therapy. Pediatr Blood Cancer. 2017; 64:e26642. https://doi.org/10.1002/pbc.26642. [PubMed].
49. Wing MG, Moreau T, Greenwood J, Smith RM, Hale G, Isaacs J, Waldmann H, Lachmann PJ, Compston A. Mechanism of first-dose cytokine-release syndrome by CAMPATH 1-H: involvement of CD16 (FcgammaRIII) and CD11a/CD18 (LFA-1) on NK cells. J Clin Invest. 1996; 98:2819–2826. https://doi.org/10.1172/JCI119110. [PubMed].
50. Winkler U, Jensen M, Manzke O, Schulz H, Diehl V, Engert A. Cytokine-release syndrome in patients with B-cell chronic lymphocytic leukemia and high lymphocyte counts after treatment with an anti-CD20 monoclonal antibody (rituximab, IDEC-C2B8). Blood. 1999; 94:2217–2224. https://doi.org/10.1182/blood.V94.7.2217.419k02_2217_2224. [PubMed].
51. DeNardo GL, Natarajan A, Hok S, Mirick G, DeNardo SJ, Corzett M, Sysko V, Lehmann J, Beckett L, Balhorn R. Nanomolecular HLA-DR10 antibody mimics: A potent system for molecular targeted therapy and imaging. Cancer Biother Radiopharm. 2008; 23:783–796. https://doi.org/10.1089/cbr.2008.0589. [PubMed].
52. DeNardo GL, Mirick GR, Hok S, DeNardo SJ, Beckett LA, Adamson GN, Balhorn RL. Molecular specific and cell selective cytotoxicity induced by a novel synthetic HLA-DR antibody mimic for lymphoma and leukemia. Int J Oncol. 2009; 34:511–516. [PubMed].
53. Balhorn R, Balhorn MC, Balakrishnan K, Rebhun RB. The small molecule antibody mimic SH7139 targets a family of HLA-DRs expressed by B-cell lymphomas and other solid cancers. J Drug Target. 2020. https://doi.org/10.1080/1061186X.2020.1787418. [PubMed] [Epub ahead of print]
54. Balhorn RL, Skorupski KA, Hok S, Balhorn MC, Guerrero T, Rebhun RB. A selective high affinity ligand (SHAL) designed to bind to an over-expressed human antigen on non-Hodgkin’s lymphoma also binds to canine B-cell lymphomas. Vet Immunol Immunopathol. 2010; 137:235–242. https://doi.org/10.1016/j.vetimm.2010.05.014. [PubMed].
55. Deffrennes V, Vedrenne J, Stolzenberg MC, Piskurich J, Barbieri G, Ting JP, Charron D, Alcaide-Loridan C. Constitutive expression of MHC class II genes in melanoma cell lines results from the transcription of class II transactivator abnormally initiated from its B cell-specific promoter. J Immunol. 2001; 167:98–106. https://doi.org/10.4049/jimmunol.167.1.98. [PubMed].
56. Johnson DB, Estrada MV, Salgado R, Sanchez V, Doxie DB, Opalenik SR, Vilgelm AE, Feld E, Johnson AS, Greenplate AR, Sanders ME, Lovly CM, Frederick DT, et al. Melanoma-specific MHC-II expression represents a tumour-autonomous phenotype and predicts response to anti-PD-1/PD-L1 therapy. Nat Commun. 2016; 7:10582. https://doi.org/10.1038/ncomms10582. [PubMed].
57. van Duinen SG, Ruiter DJ, Broecker EB, van der Velde EA, Sorg C, Welvaart K, Ferrone S. Level of HLA antigens in locoregional metastases and clinical course of the disease in patients with melanoma. Cancer Res. 1988; 48:1019–1025. [PubMed].
58. van Essen TH, Bronkhorst IH, Maat W, Verduyn W, Roelen DL, Luyten GP, Jager MJ. A comparison of HLA genotype with inflammation in uveal melanoma. Invest Ophthalmol Vis Sci. 2012; 53:2640–2646. https://doi.org/10.1167/iovs.11-8901. [PubMed].
59. Glew SS, Duggan-Keen M, Cabrera T, Stern PL. HLA class II antigen expression in human papillomavirus-associated cervical cancer. Cancer Res. 1992; 52:4009–4016. [PubMed].
60. Hilders CG, Houbiers JG, Krul EJ, Fleuren GJ. The expression of histocompatibility-related leukocyte antigens in the pathway to cervical carcinoma. Am J Clin Pathol. 1994; 101:5–12. https://doi.org/10.1093/ajcp/101.1.5. [PubMed].
61. Hohn H, Pilch H, Gunzel S, Neukirch C, Freitag K, Necker A, Maeurer MJ. Human papillomavirus type 33 E7 peptides presented by HLA-DR*0402 to tumor-infiltrating T cells in cervical cancer. J Virol. 2000; 74:6632–6636. https://doi.org/10.1128/JVI.74.14.6632-6636.2000. [PubMed].
62. Zehbe I, Hohn H, Pilch H, Neukirch C, Freitag K, Maeurer MJ. Differential MHC class II component expression in HPV-positive cervical cancer cells: implication for immune surveillance. Int J Cancer. 2005; 117:807–815. https://doi.org/10.1002/ijc.21226. [PubMed].
63. Callahan MJ, Nagymanyoki Z, Bonome T, Johnson ME, Litkouhi B, Sullivan EH, Hirsch MS, Matulonis UA, Liu J, Birrer MJ, Berkowitz RS, Mok SC. Increased HLA-DMB expression in the tumor epithelium is associated with increased CTL infiltration and improved prognosis in advanced-stage serous ovarian cancer. Clin Cancer Res. 2008; 14:7667–73. https://doi.org/10.1158/1078-0432.CCR-08-0479. [PubMed].
64. Kubler K, Arndt PF, Wardelmann E, Krebs D, Kuhn W, van der Ven K. HLA-class II haplotype associations with ovarian cancer. Int J Cancer. 2006; 119:2980–2985. https://doi.org/10.1002/ijc.22266. [PubMed].
65. Kubler K, Arndt PF, Wardelmann E, Landwehr C, Krebs D, Kuhn W, van der Ven K. Genetic alterations of HLA-class II in ovarian cancer. Int J Cancer. 2008; 123:1350–1356. https://doi.org/10.1002/ijc.23624. [PubMed].
66. Rangel LB, Agarwal R, Sherman-Baust CA, Mello-Coelho V, Pizer ES, Ji H, Taub DD, Morin PJ. Anomalous expression of the HLA-DR alpha and beta chains in ovarian and other cancers. Cancer Biol Ther. 2004; 3:1021–7. https://doi.org/10.4161/cbt.3.10.1142. [PubMed].
67. Schuster H, Peper JK, Bosmuller HC, Rohle K, Backert L, Bilich T, Ney B, Loffler MW, Kowalewski DJ, Trautwein N, Rabsteyn A, Engler T, Braun S, et al. The immunopeptidomic landscape of ovarian carcinomas. Proc Natl Acad Sci U S A. 2017; 114:E9942–51. https://doi.org/10.1073/pnas.1707658114. [PubMed].
68. Tamiolakis D, Kotini A, Venizelos J, Jivannakis T, Simopoulos C, Papadopoulos N. Prognostic significance of HLA-DR antigen in serous ovarian tumors. Clin Exp Med. 2003; 3:113–118. https://doi.org/10.1007/s10238-003-0013-0. [PubMed].
69. Bigotti G, Coli A, Castagnola D. Distribution of Langerhans cells and HLA class II molecules in prostatic carcinomas of different histopathological grade. Prostate. 1991; 19:73–87. https://doi.org/10.1002/pros.2990190108. [PubMed].
70. Carlson CB, Mowery P, Owen RM, Dykhuizen EC, Kiessling LL. Selective tumor cell targeting using low-affinity, multivalent interactions. ACS Chem Biol. 2007; 2:119–127. https://doi.org/10.1021/cb6003788. [PubMed].
71. Doonan BP, Haque A. HLA Class II Antigen Presentation in Prostate Cancer Cells: A Novel Approach to Prostate Tumor Immunotherapy. Open Cancer Immunol J. 2010; 3:1–7. https://doi.org/10.2174/1876401001003010001. [PubMed].
72. Masucci GV, Andersson E, Villabona L, Helgadottir H, Bergfeldt K, Cavallo F, Forni G, Ferrone S, Choudhury A, Seliger B, Kiessling R. Survival of the fittest or best adapted: HLA-dependent tumor development. J Nucleic Acids Investig. 2010; 1:41–46.
73. Younger AR, Amria S, Jeffrey WA, Mahdy AE, Goldstein OG, Norris JS, Haque A. HLA class II antigen presentation by prostate cancer cells. Prostate Cancer Prostatic Dis. 2008; 11:334–341. https://doi.org/10.1038/sj.pcan.4501021. [PubMed].
74. Matoba K, Iizuka N, Gondo T, Ishihara T, Yamada-Okabe H, Tamesa T, Takemoto N, Hashimoto K, Sakamoto K, Miyamoto T, Uchimura S, Hamamoto Y, Oka M. Tumor HLA-DR expression linked to early intrahepatic recurrence of hepatocellular carcinoma. Int J Cancer. 2005; 115:231–240. https://doi.org/10.1002/ijc.20860. [PubMed].
75. Paterson AC, Sciot R, Kew MC, Callea F, Dusheiko GM, Desmet VJ. HLA expression in human hepatocellular carcinoma. Br J Cancer. 1988; 57:369–373. https://doi.org/10.1038/bjc.1988.84. [PubMed].
76. Sung CH, Hu CP, Hsu HC, Ng AK, Chou CK, Ting LP, Su TS, Han SH, Chang CM. Expression of class I and class II major histocompatibility antigens on human hepatocellular carcinoma. J Clin Invest. 1989; 83:421–429. https://doi.org/10.1172/JCI113900. [PubMed].
77. Wadee AA, Paterson A, Coplan KA, Reddy SG. HLA expression in hepatocellular carcinoma cell lines. Clin Exp Immunol. 1994; 97:328–333. https://doi.org/10.1111/j.1365-2249.1994.tb06089.x. [PubMed].
78. Gastl G, Ebert T, Finstad CL, Sheinfeld J, Gomahr A, Aulitzky W, Bander NH. Major histocompatibility complex class I and class II expression in renal cell carcinoma and modulation by interferon gamma. J Urol. 1996; 155:361–7.
79. Trieb K, Lechleitner T, Lang S, Windhager R, Kotz R, Dirnhofer S. Evaluation of HLA-DR expression and T-lymphocyte infiltration in osteosarcoma. Pathol Res Pract. 1998; 194:679–684. https://doi.org/10.1016/S0344-0338(98)80126-X. [PubMed].
80. da Silva GB, Silva TG, Duarte RA, Neto NL, Carrara HH, Donadi EA, Goncalves MA, Soares EG, Soares CP. Expression of the Classical and Nonclassical HLA Molecules in Breast Cancer. Int J Breast Cancer. 2013; 2013:250435. https://doi.org/10.1155/2013/250435. [PubMed].
81. Feinmesser M, Sulkes A, Morgenstern S, Sulkes J, Stern S, Okon E. HLA-DR and beta 2 microglobulin expression in medullary and atypical medullary carcinoma of the breast: histopathologically similar but biologically distinct entities. J Clin Pathol. 2000; 53:286–291. https://doi.org/10.1136/jcp.53.4.286. [PubMed].
82. Lazzaro B, Anderson AE, Kajdacsy-Balla A, Hessner MJ. Antigenic characterization of medullary carcinoma of the breast: HLA-DR expression in lymph node positive cases. Appl Immunohistochem Mol Morphol. 2001; 9:234–241. https://doi.org/10.1097/00129039-200109000-00007. [PubMed].
83. Mostafa AA, Codner D, Hirasawa K, Komatsu Y, Young MN, Steimle V, Drover S. Activation of ERalpha signaling differentially modulates IFN-gamma induced HLA-class II expression in breast cancer cells. PLoS One. 2014; 9:e87377. https://doi.org/10.1371/journal.pone.0087377. [PubMed].
84. Park IA, Hwang SH, Song IH, Heo SH, Kim YA, Bang WS, Park HS, Lee M, Gong G, Lee HJ. Expression of the MHC class II in triple-negative breast cancer is associated with tumor-infiltrating lymphocytes and interferon signaling. PLoS One. 2017; 12:e0182786. https://doi.org/10.1371/journal.pone.0182786. [PubMed].
85. Scanlan MJ, Jager D. Challenges to the development of antigen-specific breast cancer vaccines. Breast Cancer Res. 2001; 3:95–98. https://doi.org/10.1186/bcr278. [PubMed].
86. Dunne MR, Michielsen AJ, O’Sullivan KE, Cathcart MC, Feighery R, Doyle B, Watson JA, O’Farrell NJ, Ravi N, Kay E, Reynolds JV, Ryan EJ, O’Sullivan J. HLA-DR expression in tumor epithelium is an independent prognostic indicator in esophageal adenocarcinoma patients. Cancer Immunol Immunother. 2017; 66:841–850. https://doi.org/10.1007/s00262-017-1983-1. [PubMed].
87. Kuwano H, Sumiyoshi K, Sonoda K, Kitamura K, Toh Y, Nakashima H, Sugimachi K. Pathogenesis of esophageal squamous cell carcinoma with lymphoid stroma. Hepatogastroenterology. 2001; 48:458–461. [PubMed].
88. Sadanaga N, Kuwano H, Watanabe M, Maekawa S, Mori M, Sugimachi K. Local immune response to tumor invasion in esophageal squamous cell carcinoma. The expression of human leukocyte antigen-DR and lymphocyte infiltration. Cancer. 1994; 74:586–591. https://doi.org/10.1002/1097-0142(19940715)74:2<586::AID-CNCR2820740209>3.0.CO;2-4. [PubMed].
89. Uesato M, Kono T, Shiratori T, Akutsu Y, Hoshino I, Murakami K, Horibe D, Maruyama T, Semba Y, Urahama R, Ogura Y, Oide T, Tanizawa T, Matsubara H. Lymphoepithelioma-like esophageal carcinoma with macroscopic reduction. World J Gastrointest Endosc. 2014; 6:385–389. https://doi.org/10.4253/wjge.v6.i8.385. [PubMed].
90. Houck JR, Sexton FM, Zajdel G. HLA class I and class II antigen expression on squamous cell carcinoma of the head and neck. Arch Otolaryngol Head Neck Surg. 1990; 116:1181–1185. https://doi.org/10.1001/archotol.1990.01870100075016. [PubMed].
91. Mattijssen V, De Mulder PH, Schalkwijk L, Manni JJ, Van ’t Hof-Grootenboer B, Ruiter DJ. HLA antigen expression in routinely processed head and neck squamous cell carcinoma primary lesions of different sites. Int J Cancer Suppl. 1991; 6:95–100. https://doi.org/10.1002/ijc.2910470719. [PubMed].
92. Russell S, Angell T, Lechner M, Liebertz D, Correa A, Sinha U, Kokot N, Epstein A. Immune cell infiltration patterns and survival in head and neck squamous cell carcinoma. Head Neck Oncol. 2013; 5:24. [PubMed].
93. Ioachim E, Charchanti A, Stavropoulos NE, Skopelitou A, Athanassiou ED, Agnantis NJ. Immunohistochemical expression of retinoblastoma gene product (Rb), p53 protein, MDM2, c-erbB-2, HLA-DR and proliferation indices in human urinary bladder carcinoma. Histol Histopathol. 2000; 15:721–727. https://doi.org/10.14670/HH-15.721. [PubMed].
94. Ioachim E, Stavropoulos NE, Charalabopoulos K, Agnantis NJ. Immunohistochemical prognostic markers in bladder cancer. J BUON. 2002; 7:207–216. [PubMed].
95. Ioachim-Velogianni E, Stavropoulos NE, Kitsiou E, Stefanaki S, Agnantis NJ. HLA-DR antigen expression and lymphocyte subsets in transitional cell carcinoma of the urinary bladder. An immunohistological study on frozen sections. J Pathol. 1994; 174:183–189. https://doi.org/10.1002/path.1711740307. [PubMed].
96. Prescott S, James K, Busuttil A, Hargreave TB, Chisholm GD, Smyth JF. HLA-DR expression by high grade superficial bladder cancer treated with BCG. Br J Urol. 1989; 63:264–269. https://doi.org/10.1111/j.1464-410X.1989.tb05187.x. [PubMed].
97. Kunihiro M, Tanaka S, Haruma K, Yoshihara M, Sumii K, Kajiyama G, Shimamoto F. Combined expression of HLA-DR antigen and proliferating cell nuclear antigen correlate with colorectal cancer prognosis. Oncology. 1998; 55:326–33. https://doi.org/10.1159/000011870. [PubMed].
98. Lovig T, Andersen SN, Thorstensen L, Diep CB, Meling GI, Lothe RA, Rognum TO. Strong HLA-DR expression in microsatellite stable carcinomas of the large bowel is associated with good prognosis. Br J Cancer. 2002; 87:756–762. https://doi.org/10.1038/sj.bjc.6600507. [PubMed].
99. Matsushita K, Takenouchi T, Kobayashi S, Hayashi H, Okuyama K, Ochiai T, Mikata A, Isono K. HLA-DR antigen expression in colorectal carcinomas: influence of expression by IFN-gamma in situ and its association with tumour progression. Br J Cancer. 1996; 73:644–648. https://doi.org/10.1038/bjc.1996.112. [PubMed].
100. Matsushita K, Takenouchi T, Shimada H, Tomonaga T, Hayashi H, Shioya A, Komatsu A, Matsubara H, Ochiai T. Strong HLA-DR antigen expression on cancer cells relates to better prognosis of colorectal cancer patients: Possible involvement of c-myc suppression by interferon-gamma in situ. Cancer Sci. 2006; 97:57–63. https://doi.org/10.1111/j.1349-7006.2006.00137.x. [PubMed].
101. Sconocchia G, Eppenberger-Castori S, Zlobec I, Karamitopoulou E, Arriga R, Coppola A, Caratelli S, Spagnoli GC, Lauro D, Lugli A, Han J, Iezzi G, Ferrone C, et al. HLA class II antigen expression in colorectal carcinoma tumors as a favorable prognostic marker. Neoplasia. 2014; 16:31–42. https://doi.org/10.1593/neo.131568. [PubMed].
102. Foukas PG, Tsilivakos V, Zacharatos P, Mariatos G, Moschos S, Syrianou A, Asimacopoulos PJ, Bramis J, Fotiadis C, Kittas C, Gorgoulis VG. Expression of HLA-DR is reduced in tumor infiltrating immune cells (TIICs) and regional lymph nodes of non-small-cell lung carcinomas. A putative mechanism of tumor-induced immunosuppression? Anticancer Res. 2001; 21:2609–2615. [PubMed].
103. Kamma H, Yazawa T, Ogata T, Horiguchi H, Iijima T. Expression of MHC class II antigens in human lung cancer cells. Virchows Arch B Cell Pathol Incl Mol Pathol. 1991; 60:407–412. https://doi.org/10.1007/BF02899573. [PubMed].
104. Perea F, Bernal M, Sanchez-Palencia A, Carretero J, Torres C, Bayarri C, Gomez-Morales M, Garrido F, Ruiz-Cabello F. The absence of HLA class I expression in non-small cell lung cancer correlates with the tumor tissue structure and the pattern of T cell infiltration. Int J Cancer. 2017; 140:888–899. https://doi.org/10.1002/ijc.30489. [PubMed].
105. Redondo M, Ruiz-Cabello F, Concha A, Hortas ML, Serrano A, Morell M, Garrido F. Differential expression of MHC class II genes in lung tumour cell lines. Eur J Immunogenet. 1998; 25:385–391. https://doi.org/10.1046/j.1365-2370.1998.00116.x. [PubMed].
106. Pandha H, Rigg A, John J, Lemoine N. Loss of expression of antigen-presenting molecules in human pancreatic cancer and pancreatic cancer cell lines. Clin Exp Immunol. 2007; 148:127–135. https://doi.org/10.1111/j.1365-2249.2006.03289.x. [PubMed].
107. Esteban F, Ruiz-Cabello F, Concha A, Perez-Ayala M, Sanchez-Rozas JA, Garrido F. HLA-DR expression is associated with excellent prognosis in squamous cell carcinoma of the larynx. Clin Exp Metastasis. 1990; 8:319–328. https://doi.org/10.1007/BF01810678. [PubMed].
108. Sikorska B, Danilewicz M, Wagrowska-Danilewicz M. Quantitative analysis of HLA DR expression and lymphocytic infiltrate in laryngeal cancer including clinical and morphological correlations. Gen Diagn Pathol. 1998; 143:297–303. [PubMed].
109. Sikorska B, Danilewicz M, Wagrowska-Danilewicz M. HLA-DR expression is a significant prognostic factor in laryngeal cancer. A morphometric study. APMIS. 1999; 107:383–388. https://doi.org/10.1111/j.1699-0463.1999.tb01569.x. [PubMed].
110. Ishigami S, Aikou T, Natsugoe S, Hokita S, Iwashige H, Tokushige M, Sonoda S. Prognostic value of HLA-DR expression and dendritic cell infiltration in gastric cancer. Oncology. 1998; 55:65–9. https://doi.org/10.1159/000011837. [PubMed].
111. Ma XC, Hattori T, Kushima R, Terata N, Kodama M. Expression of HLA-class II antigen in gastric carcinomas. Its relationship to histopathological grade, lymphocyte infiltration and five-year survival rate. Acta Oncol. 1994; 33:187–190. https://doi.org/10.3109/02841869409098403. [PubMed].
112. Murata S, Eguchi Y, Terata N, Tani T, Kodama M. Expression of HLA-DR and urokinase-type plasminogen activator in stage IV gastric cancer. Gastric Cancer. 1998; 1:71–77. https://doi.org/10.1007/s101200050057. [PubMed].
113. Sakai K, Takiguchi M, Mori S, Kobori O, Morioka Y, Inoko H, Sekiguchi M, Kano K. Expression and function of class II antigens on gastric carcinoma cells and gastric epithelia: differential expression of DR, DQ, and DP antigens. J Natl Cancer Inst. 1987; 79:923–932. [PubMed].
114. Fan X, Liang J, Wu Z, Shan X, Qiao H, Jiang T. Expression of HLA-DR genes in gliomas: correlation with clinicopathological features and prognosis. Chinese Neurosurgical Journal. 2017; 3:27. https://doi.org/10.1186/s41016-017-0090-7.
115. Mullins CS, Walter A, Schmitt M, Classen CF, Linnebacher M. Tumor antigen and MHC expression in glioma cells for immunotherapeutic interventions. World J Immunol. 2013; 3:62–67. https://doi.org/10.5411/wji.v3.i3.62.
116. Rossi ML, Jones NR, Karr GF, Esiri MM, Havas L, Coakham HB. HLA-Dr expression by tumor cells compared with survival in high grade astrocytomas. Tumori. 1991; 77:122–125. https://doi.org/10.1177/030089169107700206. [PubMed].
117. Fisfalen ME, Franklin WA, DeGroot LJ, Cajulis RS, Soltani K, Ryan M, Jones N. Expression of HLA ABC and DR antigens in thyroid neoplasia and correlation with mononuclear leukocyte infiltration. J Endocrinol Invest. 1990; 13:41–48. https://doi.org/10.1007/BF03348579. [PubMed].
118. Jo YS, Lee JC, Li S, Choi YS, Bai YS, Kim YJ, Lee IS, Rha SY, Ro HK, Kim JM, Shong M. Significance of the expression of major histocompatibility complex class II antigen, HLA-DR and -DQ, with recurrence of papillary thyroid cancer. Int J Cancer. 2008; 122:785–790. https://doi.org/10.1002/ijc.23167. [PubMed].
119. Kawai K, Resetkova E, Enomoto T, Fornasier V, Volpe R. Is human leukocyte antigen-DR and intercellular adhesion molecule-1 expression on human thyrocytes constitutive in papillary thyroid cancer? Comparative studies in human thyroid xenografts in severe combined immunodeficient and nude mice. J Clin Endocrinol Metab. 1998; 83:157–164. https://doi.org/10.1210/jcem.83.1.4489. [PubMed].
120. Lahat N, Rahat MA, Sadeh O, Kinarty A, Kraiem Z. Regulation of HLA-DR and costimulatory B7 molecules in human thyroid carcinoma cells: differential binding of transcription factors to the HLA-DRalpha promoter. Thyroid. 1998; 8:361–369. https://doi.org/10.1089/thy.1998.8.361. [PubMed].
121. Carlo-Stella C, Guidetti A, Di Nicola M, Lavazza C, Cleris L, Sia D, Longoni P, Milanesi M, Magni M, Nagy Z, Corradini P, Carbone A, Formelli F, Gianni AM. IFN-gamma enhances the antimyeloma activity of the fully human anti-human leukocyte antigen-DR monoclonal antibody 1D09C3. Cancer Res. 2007; 67:3269–3275. https://doi.org/10.1158/0008-5472.CAN-06-3744. [PubMed].
122. Farrar MA, Schreiber RD. The molecular cell biology of interferon-gamma and its receptor. Annu Rev Immunol. 1993; 11:571–611. https://doi.org/10.1146/annurev.iy.11.040193.003035. [PubMed].
123. Schoenborn JR, Wilson CB. Regulation of interferon-gamma during innate and adaptive immune responses. Adv Immunol. 2007; 96:41–101. https://doi.org/10.1016/S0065-2776(07)96002-2. [PubMed].
124. Basham TY, Merigan TC. Recombinant interferon-gamma increases HLA-DR synthesis and expression. J Immunol. 1983; 130:1492–1494. [PubMed].
125. Shaw AR, Chan JK, Reid S, Seehafer J. HLA-DR synthesis induction and expression in HLA-DR-negative carcinoma cell lines of diverse origins by interferon-gamma but not by interferon-beta. J Natl Cancer Inst. 1985; 74:1261–1268. [PubMed].
126. Yi Q, Dabadghao S, Osterborg A, Bergenbrant S, Holm G. Myeloma bone marrow plasma cells: evidence for their capacity as antigen-presenting cells. Blood. 1997; 90:1960–1967. https://doi.org/10.1182/blood.V90.5.1960. [PubMed].
127. Lahat N, Sobel E, Kraiem Z. Control of HLA-DR antigen expression by gamma-interferon: separate signal transduction mechanisms in malignant and nonmalignant human thyroid cells. Cancer Res. 1993; 53:3943–3947. [PubMed].
128. Tartour E, Gey A, Sastre-Garau X, Lombard Surin I, Mosseri V, Fridman WH. Prognostic value of intratumoral interferon gamma messenger RNA expression in invasive cervical carcinomas. J Natl Cancer Inst. 1998; 90:287–294. https://doi.org/10.1093/jnci/90.4.287. [PubMed].
129. Lee IC, Huang YH, Chau GY, Huo TI, Su CW, Wu JC, Lin HC. Serum interferon gamma level predicts recurrence in hepatocellular carcinoma patients after curative treatments. Int J Cancer. 2013; 133:2895–2902. https://doi.org/10.1002/ijc.28311. [PubMed].
130. Khoury EL, Greenspan JS, Greenspan FS. Adrenocortical cells of the zona reticularis normally express HLA-DR antigenic determinants. Am J Pathol. 1987; 127:580–591. [PubMed].
131. Marx C, Bornstein SR, Wolkersdorfer GW, Peter M, Sippell WG, Scherbaum WA. Relevance of major histocompatibility complex class II expression as a hallmark for the cellular differentiation in the human adrenal cortex. J Clin Endocrinol Metab. 1997; 82:3136–3140. https://doi.org/10.1210/jcem.82.9.4194. [PubMed].
132. Styren SD, Civin WH, Rogers J. Molecular, cellular, and pathologic characterization of HLA-DR immunoreactivity in normal elderly and Alzheimer’s disease brain. Exp Neurol. 1990; 110:93–104. https://doi.org/10.1016/0014-4886(90)90054-V. [PubMed].
133. Hopperton KE, Mohammad D, Trépanier MO, Giuliano V, Bazinet RP. Markers of microglia in post-mortem brain samples from patients with Alzheimer’s disease: a systematic review. Mol Psychiatry. 2018; 23:177–198. https://doi.org/10.1038/mp.2017.246. [PubMed].
134. Itagaki S, McGeer PL, Akiyama H, Zhu S, Selkoe D. Relationship of microglia and astrocytes to amyloid deposits of Alzheimer disease. J Neuroimmunol. 1989; 24:173–182. https://doi.org/10.1016/0165-5728(89)90115-X. [PubMed].
135. Martin R, Howell MD, Jaraquemada D, Flerlage M, Richert J, Brostoff S, Long EO, McFarlin DE, McFarland HF. A myelin basic protein peptide is recognized by cytotoxic T cells in the context of four HLA-DR types associated with multiple sclerosis. J Exp Med. 1991; 173:19–24. https://doi.org/10.1084/jem.173.1.19. [PubMed].
136. McGeer PL, Itagaki S, Tago H, McGeer EG. Reactive microglia in patients with senile dementia of the Alzheimer type are positive for the histocompatibility glycoprotein HLA-DR. Neurosci Lett. 1987; 79:195–200. https://doi.org/10.1016/0304-3940(87)90696-3. [PubMed].
137. Ulvestad E, Williams K, Bø L, Trapp B, Antel J, Mørk S. HLA class II molecules (HLA-DR, -DP, -DQ) on cells in the human CNS studied in situ and in vitro. Immunology. 1994; 82:535–541. [PubMed].
138. Weissert R, Kuhle J, de Graaf KL, Wienhold W, Herrmann MM, Müller C, Forsthuber TG, Wiesmüller KH, Melms A. High immunogenicity of intracellular myelin oligodendrocyte glycoprotein epitopes. J Immunol. 2002; 169:548–556. https://doi.org/10.4049/jimmunol.169.1.548. [PubMed].
139. Archimandritis A, Sougioultzis S, Foukas PG, Tzivras M, Davaris P, Moutsopoulos HM. Expression of HLA-DR, costimulatory molecules B7-1, B7-2, intercellular adhesion molecule-1 (ICAM-1) and Fas ligand (FasL) on gastric epithelial cells in Helicobacter pylori gastritis; influence of H. pylori eradication. Clin Exp Immunol. 2000; 119:464–471. https://doi.org/10.1046/j.1365-2249.2000.01164.x. [PubMed].
140. Spencer J, Finn T, Isaacson PG. Expression of HLA-DR antigens on epithelium associated with lymphoid tissue in the human gastrointestinal tract. Gut. 1986; 27:153–157. https://doi.org/10.1136/gut.27.2.153. [PubMed].
141. Arbustini E, Morbini P, Diegoli M, Grasso M, Fasani R, Vitulo P, Fiocca R, Cremaschi P, Volpato G, Martinelli L, Vigano M, Samloff IM, Solicia E. Coexpression of aspartic proteinases and human leukocyte antigen-DR in human transplanted lung. Am J Pathol. 1994; 145:310–321. [PubMed].
142. Bjerneroth G, Juhlin C, Gudmundsson S, Rastad J, Akerstrom G, Klareskog L. Major histocompatibility complex class II expression and parathyroid autoantibodies in primary hyperparathyroidism. Surgery. 1998; 124:503–9. https://doi.org/10.1016/S0039-6060(98)70096-0. [PubMed].
143. Kramer G, Steiner GE, Sokol P, Mallone R, Amann G, Marberger M. Loss of CD38 correlates with simultaneous up-regulation of human leukocyte antigen-DR in benign prostatic glands, but not in fetal or androgen-ablated glands, and is strongly related to gland atrophy. BJU Int. 2003; 91:409–16. https://doi.org/10.1046/j.1464-410X.2003.04091.x. [PubMed].
144. Rose ML, Coles MI, Griffin RJ, Pomerance A, Yacoub MH. Expression of class I and class II major histocompatibility antigens in normal and transplanted human heart. Transplantation. 1986; 41:776–780. https://doi.org/10.1097/00007890-198606000-00021. [PubMed].
145. Beninati W, Derdak S, Dixon PF, Grider DJ, Strollo DC, Hensley RE, Lucey DR. Pulmonary eosinophils express HLA-DR in chronic eosinophilic pneumonia. J Allergy Clin Immunol. 1993; 92:442–9. [PubMed].
146. Campbell DA, du Bois RM, Butcher RG, Poulter LW. The density of HLA-DR antigen expression on alveolar macrophages is increased in pulmonary sarcoidosis. Clin Exp Immunol. 1986; 65:165–171. [PubMed].
147. de Blic J, Peuchmaur M, Carnot F, Danel C, Deruesne M, Reynaud P, Scheinmann P, Brousse N. Rejection in lung transplantation–an immunohistochemical study of transbronchial biopsies. Transplantation. 1992; 54:639–644. https://doi.org/10.1097/00007890-199210000-00014. [PubMed].
148. Glanville AR, Tazelaar HD, Theodore J, Imoto E, Rouse RV, Baldwin JC, Robin ED. The distribution of MHC class I and II antigens on bronchial epithelium. Am Rev Respir Dis. 1989; 139:330–334. https://doi.org/10.1164/ajrccm/139.2.330. [PubMed].
149. Horie Y, Chiba M, Iizuka M, Masamune O, Class II. HLA-DR, -DP, and -DO) antigens on intestinal epithelia in ulcerative colitis, Crohn’s disease, colorectal cancer and normal small intestine. Gastroenterol Jpn. 1990; 25:575–584. https://doi.org/10.1007/BF02779357. [PubMed].
150. Mulder DJ, Pooni A, Mak N, Hurlbut DJ, Basta S, Justinich CJ. Antigen presentation and MHC class II expression by human esophageal epithelial cells: role in eosinophilic esophagitis. Am J Pathol. 2011; 178:744–753. https://doi.org/10.1016/j.ajpath.2010.10.027. [PubMed].
151. Oberhuber G, Puspok A, Peck-Radosavlevic M, Kutilek M, Lamprecht A, Chott A, Vogelsang H, Stolte M. Aberrant esophageal HLA-DR expression in a high percentage of patients with Crohn’s disease. Am J Surg Pathol. 1999; 23:970–976. https://doi.org/10.1097/00000478-199908000-00016. [PubMed].
152. Yousem SA, Curley JM, Dauber J, Paradis I, Rabinowich H, Zeevi A, Duquesnoy R, Dowling R, Zenati M, Hardesty R, Griffith BP. HLA-class II antigen expression in human heart-lung allografts. Transplantation. 1990; 49:991–995. https://doi.org/10.1097/00007890-199005000-00030. [PubMed].
153. Penna G, Fibbi B, Amuchastegui S, Cossetti C, Aquilano F, Laverny G, Gacci M, Crescioli C, Maggi M, Adorini L. Human benign prostatic hyperplasia stromal cells as inducers and targets of chronic immuno-mediated inflammation. J Immunol. 2009; 182:4056–4064. https://doi.org/10.4049/jimmunol.0801875. [PubMed].
154. Theyer G, Kramer G, Assmann I, Sherwood E, Preinfalk W, Marberger M, Zechner O, Steiner GE. Phenotypic characterization of infiltrating leukocytes in benign prostatic hyperplasia. Lab Invest. 1992; 66:96–107. [PubMed].
155. Milton AD, Fabre JW. Massive induction of donor-type class I and class II major histocompatibility complex antigens in rejecting cardiac allografts in the rat. J Exp Med. 1985; 161:98–112. https://doi.org/10.1084/jem.161.1.98. [PubMed].
156. Steinhoff G, Wonigeit K, Haverich A, Schäfers HJ, Kemnitz J. Class 1 MHC antigen induction on myocytes in cardiac allograft rejection. Eur Heart J. 1987; 8:25–28. https://doi.org/10.1093/eurheartj/8.suppl_J.25.
157. Caforio AL, Stewart JT, Bonifacio E, Burke M, Davies MJ, McKenna WJ, Bottazzo GF. Inappropriate major histocompatibility complex expression on cardiac tissue in dilated cardiomyopathy. Relevance for autoimmunity? J Autoimmun. 1990; 3:187–200. [PubMed].
158. Seko Y, Tsuchimochi H, Nakamura T, Okumura K, Naito S, Imataka K, Fujii J, Takaku F, Yazaki Y. Expression of major histocompatibility complex class I antigen in murine ventricular myocytes infected with Coxsackievirus B3. Circ Res. 1990; 67:360–367. https://doi.org/10.1161/01.RES.67.2.360. [PubMed].
159. Wojnicz R, Nowalany-Kozielska E, Wodniecki J, Szczurek-Katański K, Nozyński J, Zembala M, Rozek MM. Immunohistological diagnosis of myocarditis. Potential role of sarcolemmal induction of the MHC and ICAM-1 in the detection of autoimmune mediated myocyte injury. Eur Heart J. 1998; 19:1564–1572. https://doi.org/10.1053/euhj.1998.1085. [PubMed].
160. Grundtman C, Hollan I, Forre OT, Saatvedt K, Mikkelsen K, Lundberg IE. Cardiovascular disease in patients with inflammatory rheumatic disease is associated with up-regulation of markers of inflammation in cardiac microvessels and cardiomyocytes. Arthritis Rheum. 2010; 62:667–673. https://doi.org/10.1002/art.27264. [PubMed].
161. Ansari AA, Wang YC, Danner DJ, Gravanis MB, Mayne A, Neckelmann N, Sell KW, Herskowitz A. Abnormal expression of histocompatibility and mitochondrial antigens by cardiac tissue from patients with myocarditis and dilated cardiomyopathy. Am J Pathol. 1991; 139:337–354. [PubMed].
162. Barbatis C, Kelly P, Greveson J, Heryet A, McGee JO. Immunocytochemical analysis of HLA class II (DR) antigens in liver disease in man. J Clin Pathol. 1987; 40:879–884. https://doi.org/10.1136/jcp.40.8.879. [PubMed].
163. Gehring AJ, Sun D, Kennedy PT, Nolte-’t Hoen E, Lim SG, Wasser S, Selden C, Maini MK, Davis DM, Nassal M, Bertoletti A. The level of viral antigen presented by hepatocytes influences CD8 T-cell function. J Virol. 2007; 81:2940–2949. https://doi.org/10.1128/JVI.02415-06. [PubMed].
164. Senaldi G, Lobo-Yeo A, Mowat AP, Mieli-Vergani G, Vergani D. Class I and class II major histocompatibility complex antigens on hepatocytes: importance of the method of detection and expression in histologically normal and diseased livers. J Clin Pathol. 1991; 44:107–114. https://doi.org/10.1136/jcp.44.2.107. [PubMed].
165. Terada T, Nakanuma Y, Hoso M, Obata H. Expression of HLA-DR antigen on hepatic vascular endothelial cells in idiopathic portal hypertension. Clin Exp Immunol. 1991; 84:303–307. [PubMed].
166. Terada T, Nakanuma Y, Obata H. HLA-DR expression on the microvasculature of portal tracts in idiopathic portal hypertension. Immunohistochemical characteristics and relation to portal phlebosclerosis. Arch Pathol Lab Med. 1991; 115:993–997. [PubMed].
167. Obaidat A, Roth M, Hagenbuch B. The expression and function of organic anion transporting polypeptides in normal tissues and in cancer. Annu Rev Pharmacol Toxicol. 2012; 52:135–151. https://doi.org/10.1146/annurev-pharmtox-010510-100556. [PubMed].
168. Thakkar N, Lockhart AC, Lee W. Role of Organic Anion-Transporting Polypeptides (OATPs) in Cancer Therapy. AAPS J. 2015; 17:535–545. https://doi.org/10.1208/s12248-015-9740-x. [PubMed].
169. Brenner S, Riha J, Giessrigl B, Thalhammer T, Grusch M, Krupitza G, Stieger B, Jager W. The effect of organic anion-transporting polypeptides 1B1, 1B3 and 2B1 on the antitumor activity of flavopiridol in breast cancer cells. Int J Oncol. 2015; 46:324–332. https://doi.org/10.3892/ijo.2014.2731. [PubMed].
170. Svoboda M, Wlcek K, Taferner B, Hering S, Stieger B, Tong D, Zeillinger R, Thalhammer T, Jäger W. Expression of organic anion-transporting polypeptides 1B1 and 1B3 in ovarian cancer cells: relevance for paclitaxel transport. Biomed Pharmacother. 2011; 65:417–426. https://doi.org/10.1016/j.biopha.2011.04.031. [PubMed].
171. Burt HJ, Riedmaier AE, Harwood MD, Crewe HK, Gill KL, Neuhoff S. Abundance of Hepatic Transporters in Caucasians: A Meta-Analysis. Drug Metab Dispos. 2016; 44:1550–1561. https://doi.org/10.1124/dmd.116.071183. [PubMed].
172. Roucard C, Garban F, Mooney NA, Charron DJ, Ericson ML. Conformation of human leukocyte antigen class II molecules. Evidence for superdimers and empty molecules on human antigen presenting cells. J Biol Chem. 1996; 271:13993–14000. https://doi.org/10.1074/jbc.271.24.13993. [PubMed].
173. West J, Perkins J, Hok S, Balhorn R, Lightstone FC, Cosman M, DeNardo SJ, DeNardo GL. Direct antilymphoma activity of novel, first-generation “antibody mimics” that bind HLA-DR10-positive non-Hodgkin’s lymphoma cells. Cancer Biother Radiopharm. 2006; 21:645–654. https://doi.org/10.1089/cbr.2006.21.645. [PubMed].
174. Balhorn R, Balhorn MC. Selective high affinity ligands: A new class of targeting agents for cancer imaging and therapy. In: Baum RP, ed. Therapeutic Nuclear Medicine, in Series Medical Radiology/Radiation Oncology (Series Editors LW Brady, H-P Heilmann, M Molls, and C Nieder). (Berlin/Heidelberg: Springer-Verlag). 2014; pp. 139–50.
175. Balhorn R, Hok S, DeNardo S, Natarajan A, Mirick G, Corzett M, Denardo G. Hexa-arginine enhanced uptake and residualization of selective high affinity ligands by Raji lymphoma cells. Mol Cancer. 2009; 8:25. https://doi.org/10.1186/1476-4598-8-25. [PubMed].
176. Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson A, Kampf C, Sjostedt E, Asplund A, Olsson I, Edlund K, Lundberg E, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015; 347:1260419. https://doi.org/10.1126/science.1260419. [PubMed].
177. Ballardini G, Mirakian R, Bianchi FB, Pisi E, Doniach D, Bottazzo GF. Aberrant expression of HLA-DR antigens on bileduct epithelium in primary biliary cirrhosis: relevance to pathogenesis. Lancet. 1984; 2:1009–1013. https://doi.org/10.1016/S0140-6736(84)91108-5. [PubMed].
178. Volc-Platzer B, Majdic O, Knapp W, Wolff K, Hinterberger W, Lechner K, Stingl G. Evidence of HLA-DR antigen biosynthesis by human keratinocytes in disease. J Exp Med. 1984; 159:1784–1789. https://doi.org/10.1084/jem.159.6.1784. [PubMed].
179. Blumenfeld W, Ye JQ, Dahiya R, Griffiss JM, Narayan P. HLA expression by benign and malignant prostatic epithelium: augmentation by interferon-gamma. J Urol. 1993; 150:1289–92. https://doi.org/10.1016/s0022-5347(17)35762-2. [PubMed].
180. Kreisel D, Richardson SB, Li W, Lin X, Kornfeld CG, Sugimoto S, Hsieh CS, Gelman AE, Krupnick AS. Cutting edge: MHC class II expression by pulmonary nonhematopoietic cells plays a critical role in controlling local inflammatory responses. J Immunol. 2010; 185:3809–3813. https://doi.org/10.4049/jimmunol.1000971. [PubMed].
181. Mayer L, Eisenhardt D, Salomon P, Bauer W, Plous R, Piccinini L. Expression of class II molecules on intestinal epithelial cells in humans. Differences between normal and inflammatory bowel disease. Gastroenterology. 1991; 100:3–12. https://doi.org/10.1016/0016-5085(91)90575-6. [PubMed].
182. Qin OY, El-Youssef M, Yen-Lieberman B, Sapatnekar W, Youngman KR, Kusugami K, Fiocchi C. Expression of HLA-DR antigens in inflammatory bowel disease mucosa: Role of intestinal lamina propria mononuclear cell-derived interferon γ. Dig Dis Sci. 1988; 33:1528–1536. https://doi.org/10.1007/BF01535942. [PubMed].
183. Bedossa P, Bacci J, Lemaigre G, Martin E. Lymphocyte subsets and HLA-DR expression in normal pancreas and chronic pancreatitis. Pancreas. 1990; 5:415–420. https://doi.org/10.1097/00006676-199007000-00007. [PubMed].
184. Vallbracht KB, Schwimmbeck PL, Seeberg B, Kuhl U, Schultheiss HP. Endothelial dysfunction of peripheral arteries in patients with immunohistologically confirmed myocardial inflammation correlates with endothelial expression of human leukocyte antigens and adhesion molecules in myocardial biopsies. J Am Coll Cardiol. 2002; 40:515–20. https://doi.org/10.1016/s0735-1097(02)01990-3. [PubMed].
185. Davis AL, McKenzie JL, Hart DN. HLA-DR-positive leucocyte subpopulations in human skin include dendritic cells, macrophages, and CD7-negative T cells. Immunology. 1988; 65:573–581. [PubMed].
186. Gottlieb AB, Lifshitz B, Fu SM, Staiano-Coico L, Wang CY, Carter DM. Expression of HLA-DR molecules by keratinocytes, and presence of Langerhans cells in the dermal infiltrate of active psoriatic plaques. J Exp Med. 1986; 164:1013–1028. https://doi.org/10.1084/jem.164.4.1013. [PubMed].
187. Aubock J, Romani N, Grubauer G, Fritsch P. HLA-DR expression on keratinocytes is a common feature of diseased skin. Br J Dermatol. 1986; 114:465–472. https://doi.org/10.1111/j.1365-2133.1986.tb02851.x. [PubMed].
188. Tjernlund U. Beta-2-microglobulin and HLA-DR(Ia)-antigens in normal and pathological skin. Acta Derm Venereol Suppl (Stockh). 1981; 94:1–40. [PubMed].
189. Lee SC, Moore GR, Golenwsky G, Raine CS. Multiple sclerosis: a role for astroglia in active demyelination suggested by class II MHC expression and ultrastructural study. J Neuropathol Exp Neurol. 1990; 49:122–136. https://doi.org/10.1097/00005072-199003000-00005. [PubMed].
190. Ransohoff RM, Estes ML. Astrocyte expression of major histocompatibility complex gene products in multiple sclerosis brain tissue obtained by stereotactic biopsy. Arch Neurol. 1991; 48:1244–1246. https://doi.org/10.1001/archneur.1991.00530240048017. [PubMed].
191. DeNardo SJ, DeNardo GL, O’Grady LF, Hu E, Sytsma VM, Mills SL, Levy NB, Macey DJ, Miller CH, Epstein AL. Treatment of B cell malignancies with 131I Lym-1 monoclonal antibodies. Int J Cancer Suppl. 1988; 3:96–101. https://doi.org/10.1002/ijc.2910410819. [PubMed].
192. DeNardo GL, Lewis JP, DeNardo SJ, O’Grady LF. Effect of Lym-1 radioimmunoconjugate on refractory chronic lymphocytic leukemia. Cancer. 1994; 73:1425–1432. https://doi.org/10.1002/1097-0142(19940301)73:5<1425::AID-CNCR2820730517>3.0.CO;2-D. [PubMed].
193. Tobin E, Denardo G, Zhang N, Epstein AL, Liu C, Denardo S. Combination immunotherapy with anti-CD20 and anti-HLA-DR monoclonal antibodies induces synergistic anti-lymphoma effects in human lymphoma cell lines. Leuk Lymphoma. 2007; 48:944–956. https://doi.org/10.1080/10428190701272272. [PubMed].
194. Kuroda H, Tamaru J, Sakamoto G, Ohnisi K, Itoyama S. Immunophenotype of lymphocytic infiltration in medullary carcinoma of the breast. Virchows Arch. 2005; 446:10–14. https://doi.org/10.1007/s00428-004-1143-9. [PubMed].
195. Malyuchik SS, Kiyamova RG. Medullary breast carcinoma. Exp Oncol. 2008; 30:96–101. [PubMed].
196. Moore OS Jr, Foote FW Jr. The relatively favorable prognosis of medullary carcinoma of the breast. Cancer. 1949; 2:635–642. https://doi.org/10.1002/1097-0142(194907)2:4<635::AID-CNCR2820020411>3.0.CO;2-Q. [PubMed].
197. Park I, Kim J, Kim M, Bae SY, Lee SK, Kil WH, Lee JE, Nam SJ. Comparison of the characteristics of medullary breast carcinoma and invasive ductal carcinoma. J Breast Cancer. 2013; 16:417–425. https://doi.org/10.4048/jbc.2013.16.4.417. [PubMed].
198. Roche PA, Teletski CL, Stang E, Bakke O, Long EO. Cell surface HLA-DR-invariant chain complexes are targeted to endosomes by rapid internalization. Proc Natl Acad Sci U S A. 1993; 90:8581–8585. https://doi.org/10.1073/pnas.90.18.8581. [PubMed].
199. Cancer Facts & Figures 2019, https://www.cancer.org/research/cancer-facts-statistics/all-cancer-facts-figures/cancer-facts-figures-2019.html. (Accessed on August 21, 2019).
200. Mortara L, Castellani P, Meazza R, Tosi G, De Lerma Barbaro A, Procopio FA, Comes A, Zardi L, Ferrini S, Accolla RS. CIITA-induced MHC class II expression in mammary adenocarcinoma leads to a Th1 polarization of the tumor microenvironment, tumor rejection, and specific antitumor memory. Clin Cancer Res. 2006; 12:3435–3443. https://doi.org/10.1158/1078-0432.CCR-06-0165. [PubMed].
201. Qi L, Rojas JM, Ostrand-Rosenberg S. Tumor Cells Present MHC Class II-Restricted Nuclear and Mitochondrial Antigens and Are the Predominant Antigen Presenting Cells In Vivo. J Immunol. 2000; 165:5451–5461. https://doi.org/10.4049/jimmunol.165.10.5451. [PubMed].
202. Sundstrom JB, Jollow KC, Braud V, Villinger F, McMichael AJ, Lawrencec EC, Ades EW, Ansari AA. Magnitude of alloresponses to MHC class I/II expressing human cardiac myocytes is limited by their intrinsic ability to process and present antigenic peptides. Clin Dev Immunol. 2003; 10:213–226. https://doi.org/10.1080/10446670310001642410. [PubMed].
203. Cai A, Keskin DB, DeLuca DS, Alonso A, Zhang W, Zhang GL, Hammond NN, Nardi V, Stone RM, Neuberg D, Sidney J, Brusic V, Wu CJ. Mutated BCR-ABL generates immunogenic T-cell epitopes in CML patients. Clin Cancer Res. 2012; 18:5761–5772. https://doi.org/10.1158/1078-0432.CCR-12-1182. [PubMed].
204. Makita M, Azuma T, Hamaguchi H, Niiya H, Kojima K, Fujita S, Tanimoto M, Harada M, Yasukawa M. Leukemia-associated fusion proteins, dek-can and bcr-abl, represent immunogenic HLA-DR-restricted epitopes recognized by fusion peptide-specific CD4+ T lymphocytes. Leukemia. 2002; 16:2400–2407. https://doi.org/10.1038/sj.leu.2402742. [PubMed].
205. Robbins PF, El-Gamil M, Li YF, Kawakami Y, Loftus D, Appella E, Rosenberg SA. A mutated beta-catenin gene encodes a melanoma-specific antigen recognized by tumor infiltrating lymphocytes. J Exp Med. 1996; 183:1185–1192. https://doi.org/10.1084/jem.183.3.1185. [PubMed].
206. Wang RF, Wang X, Atwood AC, Topalian SL, Rosenberg SA. Cloning genes encoding MHC class II-restricted antigens: mutated CDC27 as a tumor antigen. Science. 1999; 284:1351–1354. https://doi.org/10.1126/science.284.5418.1351. [PubMed].
207. Wölfel T, Hauer M, Schneider J, Serrano M, Wölfel C, Klehmann-Hieb E, De Plaen E, Hankeln T, Meyer zum Büschenfelde KH, Beach D. A p16INK4a-insensitive CDK4 mutant targeted by cytolytic T lymphocytes in a human melanoma. Science. 1995; 269:1281–1284. https://doi.org/10.1126/science.7652577. [PubMed].
208. Doyle HA, Zhou J, Wolff MJ, Harvey BP, Roman RM, Gee RJ, Koski RA, Mamula MJ. Isoaspartyl post-translational modification triggers anti-tumor T and B lymphocyte immunity. J Biol Chem. 2006; 281:32676–32683. https://doi.org/10.1074/jbc.M604847200. [PubMed].
209. Golgher D, Korangy F, Gao B, Gorski K, Jaffee E, Edidin M, Pardoll DM, Elliott T. An immunodominant MHC class II-restricted tumor antigen is conformation dependent and binds to the endoplasmic reticulum chaperone, calreticulin. J Immunol. 2001; 167:147–155. https://doi.org/10.4049/jimmunol.167.1.147. [PubMed].
210. Consogno G, Manici S, Facchinetti V, Bachi A, Hammer J, Conti-Fine BM, Rugarli C, Traversari C, Protti MP. Identification of immunodominant regions among promiscuous HLA-DR-restricted CD4+ T-cell epitopes on the tumor antigen MAGE-3. Blood. 2003; 101:1038–1044. https://doi.org/10.1182/blood-2002-03-0933. [PubMed].
211. Slager EH, van der Minne CE, Krüse M, Krueger DD, Griffioen M, Osanto S. Identification of multiple HLA-DR-restricted epitopes of the tumor-associated antigen CAMEL by CD4+ Th1/Th2 lymphocytes. J Immunol. 2004; 172:5095–5102. https://doi.org/10.4049/jimmunol.172.8.5095. [PubMed].
212. Wen W, Zhang L, Peng J, Chen J, Hao J, Li X, Qian X, Zeng P, Zhang Y, Yin Y. Identification of promiscuous HLA-DR-restricted CD4+ T-cell epitopes on the cancer-testis antigen HCA587. Cancer Sci. 2011; 102:1455–1461. https://doi.org/10.1111/j.1349-7006.2011.01986.x. [PubMed].
213. Coulie PG, Brichard V, Van Pel A, Wölfel T, Schneider J, Traversari C, Mattei S, De Plaen E, Lurquin C, Szikora JP, Renauld JC, Boon T. A new gene coding for a differentiation antigen recognized by autologous cytolytic T lymphocytes on HLA-A2 melanomas. J Exp Med. 1994; 180:35–42. https://doi.org/10.1084/jem.180.1.35. [PubMed].
214. Kang X, Kawakami Y, el-Gamil M, Wang R, Sakaguchi K, Yannelli JR, Appella E, Rosenberg SA, Robbins PF. Identification of a tyrosinase epitope recognized by HLA-A24-restricted, tumor-infiltrating lymphocytes. J Immunol. 1995; 155:1343–1348. [PubMed].
215. Kawakami Y, Eliyahu S, Sakaguchi K, Robbins PF, Rivoltini L, Yannelli JR, Appella E, Rosenberg SA. Identification of the immunodominant peptides of the MART-1 human melanoma antigen recognized by the majority of HLA-A2-restricted tumor infiltrating lymphocytes. J Exp Med. 1994; 180:347–352. https://doi.org/10.1084/jem.180.1.347. [PubMed].
216. Kawakami Y, Eliyahu S, Jennings C, Sakaguchi K, Kang X, Southwood S, Robbins PF, Sette A, Appella E, Rosenberg SA. Recognition of multiple epitopes in the human melanoma antigen gp100 by tumor-infiltrating T lymphocytes associated with in vivo tumor regression. J Immunol. 1995; 154:3961–3968. [PubMed].
217. Wang RF, Appella E, Kawakami Y, Kang X, Rosenberg SA. Identification of TRP-2 as a human tumor antigen recognized by cytotoxic T lymphocytes. J Exp Med. 1996; 184:2207–2216. https://doi.org/10.1084/jem.184.6.2207. [PubMed].
218. Wölfel T, Van Pel A, Brichard V, Schneider J, Seliger B, Meyer zum Büschenfelde KH, Boon T. Two tyrosinase nonapeptides recognized on HLA-A2 melanomas by autologous cytolytic T lymphocytes. Eur J Immunol. 1994; 24:759–764. https://doi.org/10.1002/eji.1830240340. [PubMed].
219. de Charette M, Marabelle A, Houot R. Turning tumour cells into antigen presenting cells: The next step to improve cancer immunotherapy? Eur J Cancer. 2016; 68:134–147. https://doi.org/10.1016/j.ejca.2016.09.010. [PubMed].
220. Rimsza LM, Farinha P, Fuchs DA, Masoudi H, Connors JM, Gascoyne RD. HLA-DR protein status predicts survival in patients with diffuse large B-cell lymphoma treated on the MACOP-B chemotherapy regimen. Leuk Lymphoma. 2007; 48:542–546. https://doi.org/10.1080/10428190601078605. [PubMed].
221. Johnson DB, Nixon MJ, Wang Y, Wang DY, Castellanos E, Estrada MV, Ericsson-Gonzalez PI, Cote CH, Salgado R, Sanchez V, Dean PT, Opalenik SR, Schreeder DM, et al. Tumor-specific MHC-II expression drives a unique pattern of resistance to immunotherapy via LAG-3/FCRL6 engagement. JCI Insight. 2018; 3:e120360. https://doi.org/10.1172/jci.insight.120360. [PubMed].
222. Andersen SN, Rognum TO, Lund E, Meling GI, Hauge S. Strong HLA-DR expression in large bowel carcinomas is associated with good prognosis. Br J Cancer. 1993; 68:80–85. https://doi.org/10.1038/bjc.1993.290. [PubMed].
223. Forero A, Li Y, Chen D, Grizzle WE, Updike KL, Merz ND, Downs-Kelly E, Burwell TC, Vaklavas C, Buchsbaum DJ, Myers RM, LoBuglio AF, Varley KE. Expression of the MHC Class II Pathway in Triple-Negative Breast Cancer Tumor Cells Is Associated with a Good Prognosis and Infiltrating Lymphocytes. Cancer Immunol Res. 2016; 4:390–399. https://doi.org/10.1158/2326-6066.CIR-15-0243. [PubMed].
224. Sheen-Chen SM, Chou FF, Eng HL, Chen WJ. An evaluation of the prognostic significance of HLA-DR expression in axillary-node-negative breast cancer. Surgery. 1994; 116:510–515. [PubMed].
225. Brooks AG, Campbell PL, Reynolds P, Gautam AM, McCluskey J. Antigen presentation and assembly by mouse I-Ak class II molecules in human APC containing deleted or mutated HLA DM genes. J Immunol. 1994; 153:5382–5392. [PubMed].
226. Humphreys RE, Hillman GG, von Hofe E, Xu M. Forcing tumor cells to present their own tumor antigens to the immune system: a necessary design for an efficient tumor immunotherapy. Cell Mol Immunol. 2004; 1:180–185. [PubMed].
227. List AF, Spier CM, Miller TP, Grogan TM. Deficient tumor-infiltrating T-lymphocyte response in malignant lymphoma: relationship to HLA expression and host immunocompetence. Leukemia. 1993; 7:398–403. [PubMed].
228. Momburg F, Herrmann B, Moldenhauer G, Moller P. B-cell lymphomas of high-grade malignancy frequently lack HLA-DR, -DP and -DQ antigens and associated invariant chain. Int J Cancer. 1987; 40:598–603. https://doi.org/10.1002/ijc.2910400504. [PubMed].
229. Papadimitriou L, Morianos I, Michailidou V, Dionyssopoulou E, Vassiliadis S, Athanassakis I. Characterization of intracellular HLA-DR, DM and DO profile in K562 and HL-60 leukemic cells. Mol Immunol. 2008; 45:3965–3973. https://doi.org/10.1016/j.molimm.2008.06.017. [PubMed].
230. Yazawa T, Ito T, Kamma H, Suzuki T, Okudela K, Hayashi H, Horiguchi H, Ogata T, Mitsui H, Ikeda M, Kitamura H. Complicated mechanisms of class II transactivator transcription deficiency in small cell lung cancer and neuroblastoma. Am J Pathol. 2002; 161:291–300. https://doi.org/10.1016/S0002-9440(10)64181-8. [PubMed].
231. Ansell SM, Stenson M, Habermann TM, Jelinek DF, Witzig TE. Cd4+ T-cell immune response to large B-cell non-Hodgkin’s lymphoma predicts patient outcome. J Clin Oncol. 2001; 19:720–726. https://doi.org/10.1200/JCO.2001.19.3.720. [PubMed].
232. Ramaswamy S, Ross KN, Lander ES, Golub TR. A molecular signature of metastasis in primary solid tumors. Nat Genet. 2003; 33:49–54. https://doi.org/10.1038/ng1060. [PubMed].
233. Riemersma SA, Jordanova ES, Schop RF, Philippo K, Looijenga LH, Schuuring E, Kluin PM. Extensive genetic alterations of the HLA region, including homozygous deletions of HLA class II genes in B-cell lymphomas arising in immune-privileged sites. Blood. 2000; 96:3569–3577. https://doi.org/10.1182/blood.V96.10.3569. [PubMed].
234. Hilton DA, West KP. An evaluation of the prognostic significance of HLA-DR expression in gastric carcinoma. Cancer. 1990; 66:1154–1157. https://doi.org/10.1002/1097-0142(19900915)66:6<1154::AID-CNCR2820660613>3.0.CO;2-3. [PubMed].
235. Miller TP, Lippman SM, Spier CM, Slymen DJ, Grogan TM. HLA-DR (Ia) immune phenotype predicts outcome for patients with diffuse large cell lymphoma. J Clin Invest. 1988; 82:370–372. https://doi.org/10.1172/JCI113598. [PubMed].
236. Rimsza LM, Roberts RA, Miller TP, Unger JM, LeBlanc M, Braziel RM, Weisenberger DD, Chan WC, Muller-Hermelink HK, Jaffe ES, Gascoyne RD, Campo E, Fuchs DA, et al. Loss of MHC class II gene and protein expression in diffuse large B-cell lymphoma is related to decreased tumor immunosurveillance and poor patient survival regardless of other prognostic factors: a follow-up study from the Leukemia and Lymphoma Molecular Profiling Project. Blood. 2004; 103:4251–8. https://doi.org/10.1182/blood-2003-07-2365. [PubMed].
237. Rosenwald A, Wright G, Chan WC, Connors JM, Campo E, Fisher RI, Gascoyne RD, Muller-Hermelink HK, Smeland EB, Giltnane JM, Hurt EM, Zhao H, Averett L, et al, and Lymphoma/Leukemia Molecular Profiling Project. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med. 2002; 346:1937–1947. https://doi.org/10.1056/NEJMoa012914. [PubMed].
238. Rybski JA, Spier CM, Miller TP, Lippman SM, McGee DL, Grogan TM. Prediction of Outcome in Diffuse Large Cell Lymphoma by the Major Histocompatibility Complex Class II (HLA-DR, DP, DQ) and Class I (HLA-A, B, C) Phenotype. Leuk Lymphoma. 1991; 6:31–38. https://doi.org/10.3109/10428199109064876. [PubMed].
239. Diao J, Xia T, Zhao H, Liu J, Li B, Zhang Z. Overexpression of HLA-DR is associated with prognosis of glioma patients. Int J Clin Exp Pathol. 2015; 8:5485–5490. [PubMed].
240. Wang H, Zhou X, Zhu JW, Ye JN, Guo HF, Sun C. Association of CD117 and HLA-DR expression with shorter overall survival and/or progression-free survival in patients with multiple myeloma treated with bortezomib and thalidomide combination treatment without transplantation. Oncol Lett. 2018; 16:5655–5666. https://doi.org/10.3892/ol.2018.9365. [PubMed].
241. Xiao S, Zhu H. Correlation study on CD7, CD34, CD56 and HLA-DR expressions and its prognosis among patients with acute myeloid leukemia. Biomed Res. 2017; 28:6786–6790.
242. Ostrand-Rosenberg S. Tumor immunotherapy: the tumor cell as an antigen-presenting cell. Curr Opin Immunol. 1994; 6:722–7. https://doi.org/10.1016/0952-7915(94)90075-2. [PubMed].
243. Byrne SN, Halliday GM. High levels of Fas ligand and MHC class II in the absence of CD80 or CD86 expression and a decreased CD4+ T cell Infiltration, enables murine skin tumours to progress. Cancer Immunol Immunother. 2003; 52:396–402. https://doi.org/10.1007/s00262-003-0380-0. [PubMed].
244. Maharshak N, Cohen S, Lantner F, Hart G, Leng L, Bucala R, Shachar I. CD74 is a survival receptor on colon epithelial cells. World J Gastroenterol. 2010; 16:3258–3266. https://doi.org/10.3748/wjg.v16.i26.3258. [PubMed].
245. Gil-Yarom N, Radomir L, Sever L, Kramer MP, Lewinsky H, Bornstein C, Blecher-Gonen R, Barnett-Itzhaki Z, Mirkin V, Friedlander G, Shvidel L, Herishanu Y, Lolis EJ, et al. CD74 is a novel transcription regulator. Proc Natl Acad Sci U S A. 2017; 114:562–567. https://doi.org/10.1073/pnas.1612195114. [PubMed].
246. Roche PA, Furuta K. The ins and outs of MHC class II-mediated antigen processing and presentation. Nat Rev Immunol. 2015; 15:203–216. https://doi.org/10.1038/nri3818. [PubMed].
247. Roche PA, Marks MS, Cresswell P. Formation of a nine-subunit complex by HLA class II glycoproteins and the invariant chain. Nature. 1991; 354:392–394. https://doi.org/10.1038/354392a0. [PubMed].
248. Lindner R. Invariant Chain Complexes and Clusters as Platforms for MIF Signaling. Cells. 2017; 6:6. https://doi.org/10.3390/cells6010006. [PubMed].
249. Sakaguchi S, Ono M, Setoguchi R, Yagi H, Hori S, Fehervari Z, Shimizu J, Takahashi T, Nomura T. Foxp3+ CD25+ CD4+ natural regulatory T cells in dominant self-tolerance and autoimmune disease. Immunol Rev. 2006; 212:8–27. https://doi.org/10.1111/j.0105-2896.2006.00427.x. [PubMed].
250. Campoli M, Ferrone S. Tumor escape mechanisms: potential role of soluble HLA antigens and NK cells activating ligands. Tissue Antigens. 2008; 72:321–334. https://doi.org/10.1111/j.1399-0039.2008.01106.x. [PubMed].
251. Waldhauer I, Steinle A. NK cells and cancer immunosurveillance. Oncogene. 2008; 27:5932–5943. https://doi.org/10.1038/onc.2008.267. [PubMed].
252. Korman AJ, Peggs KS, Allison JP. Checkpoint blockade in cancer immunotherapy. Adv Immunol. 2006; 90:297–339. https://doi.org/10.1016/S0065-2776(06)90008-X. [PubMed].
253. Griffith TS, Chin WA, Jackson GC, Lynch DH, Kubin MZ. Intracellular regulation of TRAIL-induced apoptosis in human melanoma cells. J Immunol. 1998; 161:2833–2840. [PubMed].
254. Ullenhag GJ, Mukherjee A, Watson NF, Al-Attar AH, Scholefield JH, Durrant LG. Overexpression of FLIPL is an independent marker of poor prognosis in colorectal cancer patients. Clin Cancer Res. 2007; 13:5070–5075. https://doi.org/10.1158/1078-0432.CCR-06-2547. [PubMed].
255. Korkolopoulou P, Kaklamanis L, Pezzella F, Harris AL, Gatter KC. Loss of antigen-presenting molecules (MHC class I and TAP-1) in lung cancer. Br J Cancer. 1996; 73:148–153. https://doi.org/10.1038/bjc.1996.28. [PubMed].
256. Meissner M, Reichert TE, Kunkel M, Gooding W, Whiteside TL, Ferrone S, Seliger B. Defects in the human leukocyte antigen class I antigen processing machinery in head and neck squamous cell carcinoma: association with clinical outcome. Clin Cancer Res. 2005; 11:2552–2560. https://doi.org/10.1158/1078-0432.CCR-04-2146. [PubMed].
257. Nie Y, Yang G, Song Y, Zhao X, So C, Liao J, Wang LD, Yang CS. DNA hypermethylation is a mechanism for loss of expression of the HLA class I genes in human esophageal squamous cell carcinomas. Carcinogenesis. 2001; 22:1615–1623. https://doi.org/10.1093/carcin/22.10.1615. [PubMed].
258. Sanda MG, Restifo NP, Walsh JC, Kawakami Y, Nelson WG, Pardoll DM, Simons JW. Molecular characterization of defective antigen processing in human prostate cancer. J Natl Cancer Inst. 1995; 87:280–285. https://doi.org/10.1093/jnci/87.4.280. [PubMed].
259. Garrido F, Cabrera T, Aptsiauri N. “Hard” and “soft” lesions underlying the HLA class I alterations in cancer cells: implications for immunotherapy. Int J Cancer. 2010; 127:249–256. https://doi.org/10.1002/ijc.25270. [PubMed].
260. Norell H, Carlsten M, Ohlum T, Malmberg KJ, Masucci G, Schedvins K, Altermann W, Handke D, Atkins D, Seliger B, Kiessling R. Frequent loss of HLA-A2 expression in metastasizing ovarian carcinomas associated with genomic haplotype loss and HLA-A2-restricted HER-2/neu-specific immunity. Cancer Res. 2006; 66:6387–6394. https://doi.org/10.1158/0008-5472.CAN-06-0029. [PubMed].
261. Zheng L, Ren L, Kouhi A, Khawli LA, Hu P, Kaslow HR, Epstein AL. A Humanized Lym-1 CAR with Novel DAP10/DAP12 Signaling Domains Demonstrates Reduced Tonic Signaling and Increased Antitumor Activity in B-Cell Lymphoma Models. Clin Cancer Res. 2020; 26:3694–3706. https://doi.org/10.1158/1078-0432.CCR-19-3417. [PubMed].
262. Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012; 9:671–675. https://doi.org/10.1038/nmeth.2089. [PubMed].

