Introduction
Acute lymphoblastic leukemia (ALL) and the related malignancy lymphoblastic lymphoma (LBL) dominate pediatric oncology, together representing over one third of all childhood cancer [1–3]. These diseases afflict even more adults in absolute terms, and are more often fatal in them [4, 5]. ALL and LBL can develop in lymphoblasts of either the B or T cell lineage; B-ALL exceeds T-ALL, but T-LBL is more prevalent than B-LBL. Despite treatments for these diseases improving considerably over the past several decades, relapsed ALL is actually the 4th most-frequent pediatric cancer diagnosis and second most lethal childhood malignancy, accounting for ~25% of all deaths [6–8].
Animal studies have been instrumental in investigating the genetics, genomics, drug sensitivities, and other features of ALL and LBL, and several mouse models for these diseases exist [9, 10]. Mammalian systems have advantages such as anatomic, physiologic, and genetic conservation, with mice being the main mammal utilized, but other organisms provide complementary advantages. Zebrafish (Danio rerio) offer flexible genetic tools, powerful live imaging, high fecundity, rapid development, shared oncogenic pathways, affordability, and other favorable traits that have allowed them to quickly gain traction as a cancer model [11–17].
Many groups have exploited these attributes to create zebrafish leukemia models, specifically for acute myeloid leukemia (AML) and T-ALL [18–37]. For T-ALL in particular, zebrafish models have been highly informative, advancing our understanding of T-ALL genetics, pro- and anti-oncogenic interactions between different genes and pathways, tumor heterogeneity, leukemia stem cells, and in screens for new therapeutics [28, 38–53]. However, despite the fact that zebrafish T-ALL models had proven to be fertile grounds for study, B-ALL modeling in D. rerio had not been fruitful, with only one low penetrance and long latency line reported [54]. This was curious because a zebrafish recombination activating gene 2 (rag2) promoter—active in both immature T and B cells—was used to regulate most of these transgenic oncoproteins in the various T-ALL lines, yet D. rerio B-ALL had not been reported in them [55, 56]. Overall, since B-ALL is the more prevalent type in patients, the lack of B-lineage models was particularly unfortunate.
In 2018, the zebrafish ALL field advanced suddenly with reports of B-ALL in two closely-related transgenic lines where T-ALL was already known to occur [57, 58]. In one, a rag2:mMyc (murine Myc) transgene was used [29], with ALL purified as single clones by allo-transplantation. Two of 12 ALL analyzed by RNA sequencing (RNA-seq) exhibited gene expression consistent with B-ALL cells arrested at the pro-B cell stage [57]. In the other, a rag2:hMYC (human MYC) transgene was utilized [32], as well as a transgenic marker, lck:eGFP [59], differentially expressed by B and T cells. Analysis of over one hundred animals demonstrated that many develop B-ALL, others develop T-ALL (as previously known), and still other fish acquire both ALL types concommitantly [58]. A follow-up report by these groups further showed that despite high similarity between the mMyc and hMYC transgenes used, these B-ALL are actually quite different, occurring in distinct B cell lineages and with dissimilar expression patterns [60]. Here, we present new results in the hMYC model, including B- and T-ALL latency and penetrance data in a cohort of over 600 animals, in vivo glucocorticoid and radiation treatment of B-ALL, and expression profiles from single B- and T-ALL cells. We also present new analyses that compare mMyc vs. hMYC B-ALL to reveal key biologic pathways that distinguish them.
Materials and Methods
Zebrafish care
Zebrafish care was provided as previously reported [58]. Animals were housed in an aquatic colony at 28.5°C on a 14:10 hour light:dark circadian cycle and experiments performed according to protocols approved by the University of Oklahoma Health Sciences Center IACUC (12-066 and 15-046). For all procedures, fish were anesthetized with 0.02% tricaine methanesulfonate (MS-222). D. rerio with the cd79a:GFP or cd79b:GFP transgenic markers [61] were bred to rag2:hMYC fish [32] to create the new transgenic lines reported herein.
Fluorescent microscopy
Anesthetized 3–9 month old hMYC; GFP fish were screened for abnormal GFP patterns using a Nikon AZ100 fluorescent microscope. Low exposure (200 ms, 2.8× gain) and high exposure (1.5 s, 3.4× gain) settings were used to obtain images with Nikon DS-Qi1MC camera. Images were processed with Nikon NIS Elements Version 4.13 software.
Fluorescence-Activated Cell Sorting (FACS) and flow cytometric analyses
As previously described [58], cells from whole fish were dissociated using a pestle, and then passed through 35 μm filters. GFPhi, GFPlo, and/or GFP- cells were collected from the lymphoid and precursor gates using a BD-FACSJazz Instrument (Becton Dickinson, San Jose, CA, USA). Flow cytometric analyses were performed using FlowJo software (Ashland, OR, USA).
B- and T-ALL incidence studies
Beginning at ~75 dpf, a cohort of 628 rag2:hMYC;lck:GFP zebrafish was monitored by ~weekly fluorescence microscopy to detect ALL. Animals that developed fluorescent cancers were euthanized and single cell suspensions were prepared as described previously [58]. Cells in the lymphoid and precursor gates were analyzed for GFP intensity using a Beckman-Coulter CytoFLEX™ to determine whether each ALL derived from T, B, or both lymphocyte lineages [58, 62, 63].
In vivo dexamethasone and radiation treatments of D. rerio B-ALL
Zebrafish with B-ALL were treated by continuous immersion in 5 μg/ml dexamethasone (DXM) in normal fish water for 9 days, modified from our prior zebrafish T-ALL DXM protocol [43, 64]. Water and DXM were changed daily, with one or two fish housed in 500 ml. After completing treatment, animals were monitored by weekly fluorescent microscopy to detect relapse. Zebrafish with B-ALL were treated by γ-irradiation (IR) using a Cesium137 irradiator to deliver a total dose of 15 Gy divided in two fractions: 10 Gy on day 0 and a 5 Gy boost on day 5. Animals were imaged by fluorescent microscopy prior to treatment, 2 days post-treatment (day 7), and weekly-to-monthly thereafter to monitor for relapse.
Nanostring™ gene expression analyses
FACS purification of normal lymphocyte and ALL samples from WT and rag2:hMYC fish, RNA extraction, and probe hybridization were performed as described previously [58], with gene identities and probe sequences available in the online supplemental material of that publication. Hybridization data were analyzed using nSolver 3.0 software (Nanostring nCounter Technologies, Seattle, WA, USA). Read counts were log-transformed and converted to z-scores for visualization purposes.
Single-cell qRT-PCR analyses of hMYC B- and T-ALL
ALL cells from double-transgenic hMYC fish were isolated from peritoneal washings as described previously [63]. Individual B- and T-ALL cells were FAC-sorted into 96 well plates, and cells were lysed, RNA extracted, and cDNA synthesized according to the Fluidigm™ single-cell protocol. Twenty-cycle pre-amplification reactions were performed using gene-specific primers (listed in Supplementary Table 1.) according to manufacturer instructions. Unincorporated primers were digested with exonuclease I (NEB M0293L) and samples diluted 5-fold in DNA suspension buffer (TEKnova PN T0221). Single-cell pre-amplified cDNA were then quantified by qRT-PCR in a Fluidigm BioMark HD machine using 48.48 Dynamic Array Chips for Gene Expression following the manufacturer’s protocol. CT values were converted to Log2Ex values for visualization purposes using a limit of detection (LoD) of 28. Additional LoD limits were examined and showed similar results.
RNA-seq and gene expression comparisons of hMYC and mMyc B-ALL
Paired-end RNA-seq reads were trimmed with BBDuk (v.38.22; retrieved from http://sourceforge.net/projects/bbmap) and aligned to the D. rerio GRCz11 genome using STAR (v.2.6.1b; default settings optimized for read-length) [65]. Picard’s MarkDuplicates tool (v.2.18.14; retrieved from http://broadinstitute.github.io/picard) was used to identify potential duplicates. Gene counts were generated with featureCounts (RSubread v.1.32.1) using D. rerio Ensembl annotation (release 92) [66, 67]. Ribosomal and mitochondrial RNA were excluded. Counts were processed and normalized within DESeq2 (v.1.22.1) [68], and pairwise differential expression testing was used to compare hMYC and mMyc B-ALL (adjusted p-value < 0.05, absolute fold-change > 1.5). Additional filtering steps were as described previously [60]. Putative human orthologues were mapped to corresponding D. rerio genes using BEAGLE (update 090718; retrieved from http://chgr.mgh.harvard.edu/genomesrus/index.php). In downstream analyses, a gene count threshold of 100 was used for each up-regulated group (minimum expression in at least 75% of in-group samples). RNA-seq heatmaps depict normalized counts generated via the variance-stabilizing transformation in DESeq2, unless otherwise described. Over-representation analysis was performed with clusterProfiler (v.3.10.0) [69] using putative human orthologues (FDR < 0.05) in the C2, C5, and C7 collections from MSigDB (v.7.0) [70]. The 10 highest-scoring pathways by FDR q-value for each group were chosen for visualization, with pathway names changed for brevity. Highly-redundant pathways were excluded manually, based on gene overlap. Original pathway names (order matching Figure 6B) and related data are listed in Supplementary Table 4.
Results and discussion
B-ALL is early onset and highly penetrant in hMYC zebrafish
We previously analyzed more than 50 unique hMYC B-ALL [58, 60], demonstrating these fish are a robust model, but latency and incidence rates for B- and T-ALL have not been reported in the hMYC line. To determine these, beginning at 3 months of age, we monitored fish by serial fluorescent microscopy to detect GFPlo B-ALL and GFPhi T-ALL, as we described previously (Figure 1A) [58]. Using the lck:GFP marker, GFPlo B-ALL fluoresce dimly, making them difficult to discern by microscopy and more likely to require high disease burdens to be detected. In addition, brightly fluorescent T-ALL can obscure detection of B-ALL. To address these possibilities, we flow cytometrically tested ALL in order to definitively assign them to the correct group.
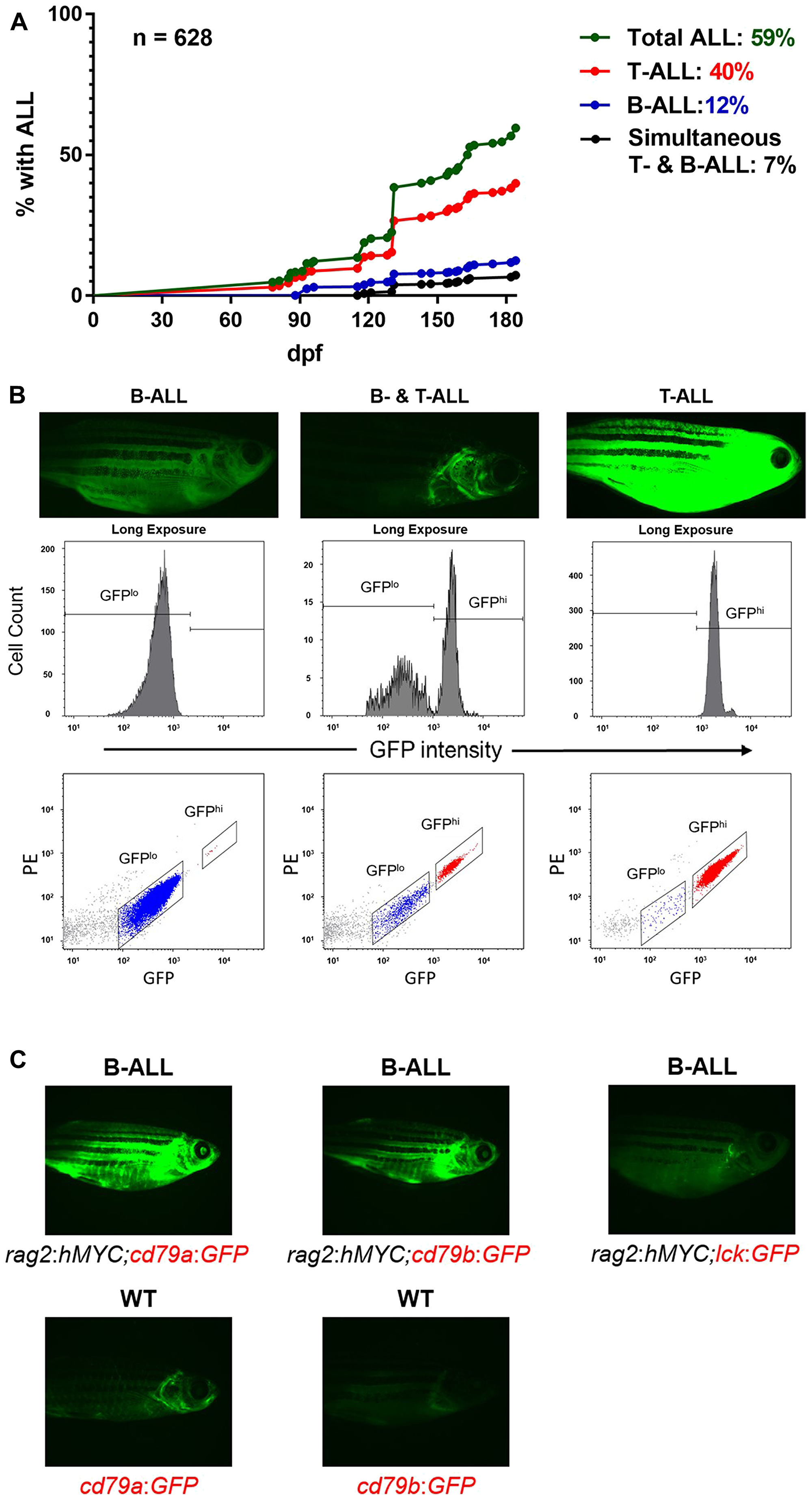
Figure 1: B-ALL in hMYC zebrafish. (A) Curves displaying B- and T-ALL incidence by 184 dpf, as determined by fluorescence microscopy. (B) Fluorescence microscopy images of fish with B-ALL (left), both B- and T-ALL (center), or T-ALL (right). Flow cytometry plots of ALL samples from these fish are shown beneath them. (C) B-ALL appearance in fish with rag2:hMYC and different transgenic markers: cd79a:GFP (left), cd79b:GFP (center), or lck:GFP (right). WT cd79a:GFP or cd79b:GFP fish without B-ALL are shown beneath for comparison. Images are representative of > 50 animals examined for each genotype.
We surveyed over 600 fish until 184 days post fertilization (dpf) and found ~60% developed at least one form of ALL by that time (Figure 1A). Isolated T-ALL was most common (40%), with peak incidence between 4-5 months. B-ALL followed a similar time course, with 12% incidence by 184 dpf, and another 7% of animals developing both T- and B-ALL. To confirm these results, every ALL was analyzed flow cytometrically (Figure 1B). By 300 dpf, every surviving animal had developed at least one type of ALL (64% T-ALL, 23% B-ALL, 13% T- and B-ALL; data not shown). We conclude that ALL is highly penetrant in hMYC fish, with peak onset beginning shortly after 120 dpf.
To simplify detection of B-ALL by fluorescent microscopy and eliminate the need for flow cytometric confirmation, we created double-transgenic fish with rag2:hMYC and either the cd79a:GFP or cd79b:GFP marker (Figure 1C), which utilize promoters from B cell-specific proteins that are components of the surface immunoglobulin signaling complex [61]. In these lines, B-ALL fluoresce brightly, but T-ALL do not fluoresce. These models will be useful for B-ALL studies, particularly in investigations with potential therapeutic agents. The highly-fluorescent B-ALL of these lines can facilitate accurate and high-throughput determinations of both response rates and relapse kinetics, allowing precise regression measurements via fluorescence quantifications, and more sensitive detection of B-ALL relapses when disease burden still remains low.
B-ALL respond to dexamethasone and radiation treatments
The glucocorticoids prednisone and dexamethasone (DXM) are the backbone of ALL therapy in patients, and γ-irradiation (IR) remains an effective adjunct that is also used, although it is reserved for specific cases due to toxicity [71, 72]. To determine whether B-ALL in zebrafish are similarly responsive to these modalities and establish protocols for future therapeutic studies in this model, we devised in vivo regimens for DXM and IR treatment of D. rerio with B-ALL.
For DXM, fish were housed continuously in fish water containing 5 μg/ml DXM for 9 days, modified from our prior DXM studies in zebrafish T-ALL, and monitored by serial fluorescence microscopy [43, 64]. We treated nineteen animals, and every B-ALL showed robust regression, with most becoming undetectable by fluorescent microscopy by the end of treatment (Figure 2A). After completing DXM treatment, we then monitored fish for 9 weeks, with 58% (11/19) recurring by 4 weeks. We conclude that our DXM protocol effectively kills most B-ALL cells, as 100% of animals exhibited brisk responses, but in many cases, sufficient B-ALL cells persist to re-grow the cancer quickly. Based on this, we believe this model can be utilized to investigate the genetic and molecular features of steroid-resistant ALL, a significant clinical problem in ALL patients [73].
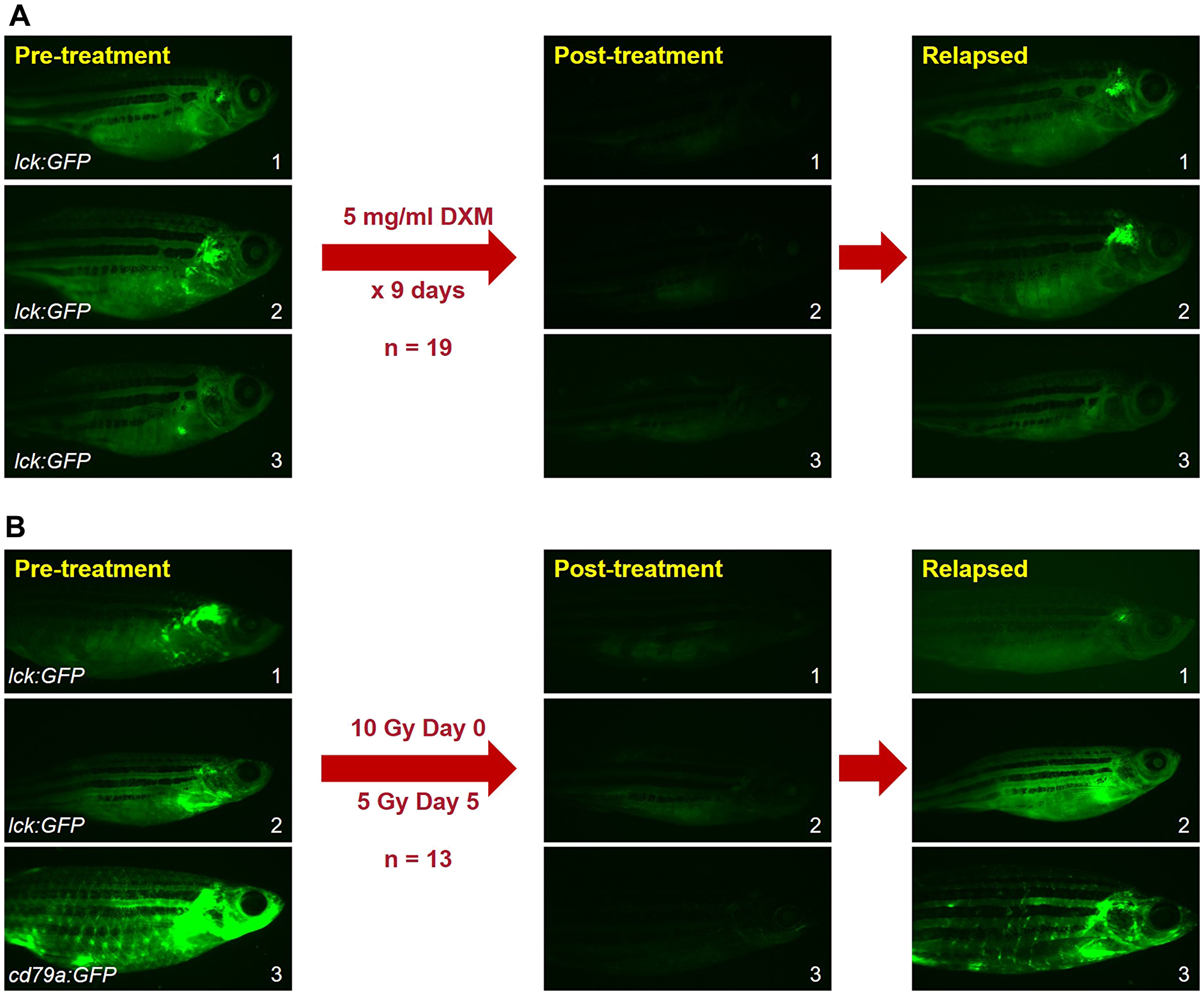
Figure 2: B-ALL regress with DXM or IR treatments. (A) B-ALL in rag2:hMYC fish treated with DXM for 9 days (n = 19). Post-treatment and relapsed images were obtained on days 9 and 28, respectively. Every (19/19) B-ALL regressed after DXM, but 11/19 (58%) B-ALL relapsed by day 28 (19 days after DXM withdrawal). (B) B-ALL in rag2:hMYC fish treated with IR (n = 13). Post-treatment and relapsed images were obtained on days 7 (i. e., 2 days after the 2nd IR dose) and 23, respectively. Every B-ALL (13/13) regressed after IR, but 100% recurred within 3 weeks of the 2nd IR dose. Upon relapse, flow cytometric testing (See Figure 1B) was used to verify that each relapse was GFPlo B-ALL. Images of rag2:hMYC, lck:GFP fish (top 5 animals) have had brightness enhanced to facilitate visualization of dim B-ALL. Images of the rag2:hMYC, cd79a:GFP fish (lowest animal) are unmodified.
IR is toxic in patients, and we also observed this in zebrafish with B-ALL. We initially administered IR treatments of 20 Gray (Gy) as single doses, but many animals died, despite the fact that this dose is well-tolerated by wild-type (WT) fish [59]. To diminish IR toxicity, presumably caused by overwhelming tumor lysis [74], we divided IR into two fractions, giving 10 Gy on day 0 and an additional 5 Gy on day 5. This was better tolerated, and following the second fraction, B-ALL were undetectable by fluorescence microscopy (Figure 2B). However, as seen in many DXM-treated B-ALL, all B-ALL relapsed post-IR, and recurrences were even more rapid (~3 weeks). From these results, we infer that IR (like DXM) kills most B-ALL cells, but surviving cells quickly re-populate the tumor. In addition, the rapid re-growth of B-ALL after IR suggests either more cells persist following IR, or that B-ALL cells that survive IR adopt more aggressive phenotypes. Distinguishing between these possibilities and unravelling the underlying mechanisms responsible will be informative to the clinical scenarios these experiments represent.
Defining B-ALL and B cell gene expression patterns
We discovered many D. rerio B lymphocytes and B-ALL express low levels of lck [58], which fostered new studies not previously done in zebrafish, such as FACS isolation of B-lineage cells (i. e., GFPlo cells from lck:GFP fish) and profiling their gene expression [58, 63]. These findings translate to humans because immature human B cells and many patients’ B-ALL also express low levels of LCK [58, 75, 76]. It is currently not known if these newly-described lcklo/LCKlo B cells represent one, or many, B cell population(s).
To begin to address this, we assessed gene expression in lymphocytes of fish carrying only the lck:GFP marker transgene (henceforth referred to as WT) and double-transgenic rag2:hMYC, lck:GFP fish (henceforth, hMYC). We FACS-purified lymphoid gate cells from both marrow and thymus, and further divided these into GFP-negative (GFP-), GFPlo, and GFPhi fractions [58, 62]. We then used Nanostring™, a multiplexed probe-based hybridization technique, to quantify mRNAs expressed by each population [77, 78]. To categorize distinct cell identities, we selected zebrafish gene homologues whose expression distinguishes mammalian B, T, and other leukocytes for these experiments [58, 79]. We found that GFPhi cells from both thymus and marrow in both genotypes (WT, hMYC) express T-lineage transcripts such as cd2, cd4, cd8a, itk, lat, and T cell receptor (TCR) mRNA (trcd, trcg, trbc2, trac, trbc1; Figure 3A). RNA-seq studies by other groups indicate some D. rerio natural killer (NK) and myeloid cells also express lck [49, 80], but transcripts indicative of these cells were not seen, suggesting they are minor populations in marrow and thymus. In contrast, B cell gene expression (cd79a, cd79b, ighz, btk, cd22, syk, lyn, pax5, blnk, ighm, etc.) was prominent in GFPlo and GFP- fractions from both organs (Figure 3A), particularly in hMYC GFPlo cells of both marrow (Figure 3B) and thymus (Figure 3C), where B-lineage genes were the dominant signature. We draw two inferences from these data: (1) lck- and lcklo lymphocytes both contain B cells, but the lcklo population has greater B cell enrichment and (2) lcklo B cells are more abundant in hMYC fish, as B-lineage genes were more dominant in hMYC GFPlo signatures than in those of WT animals.
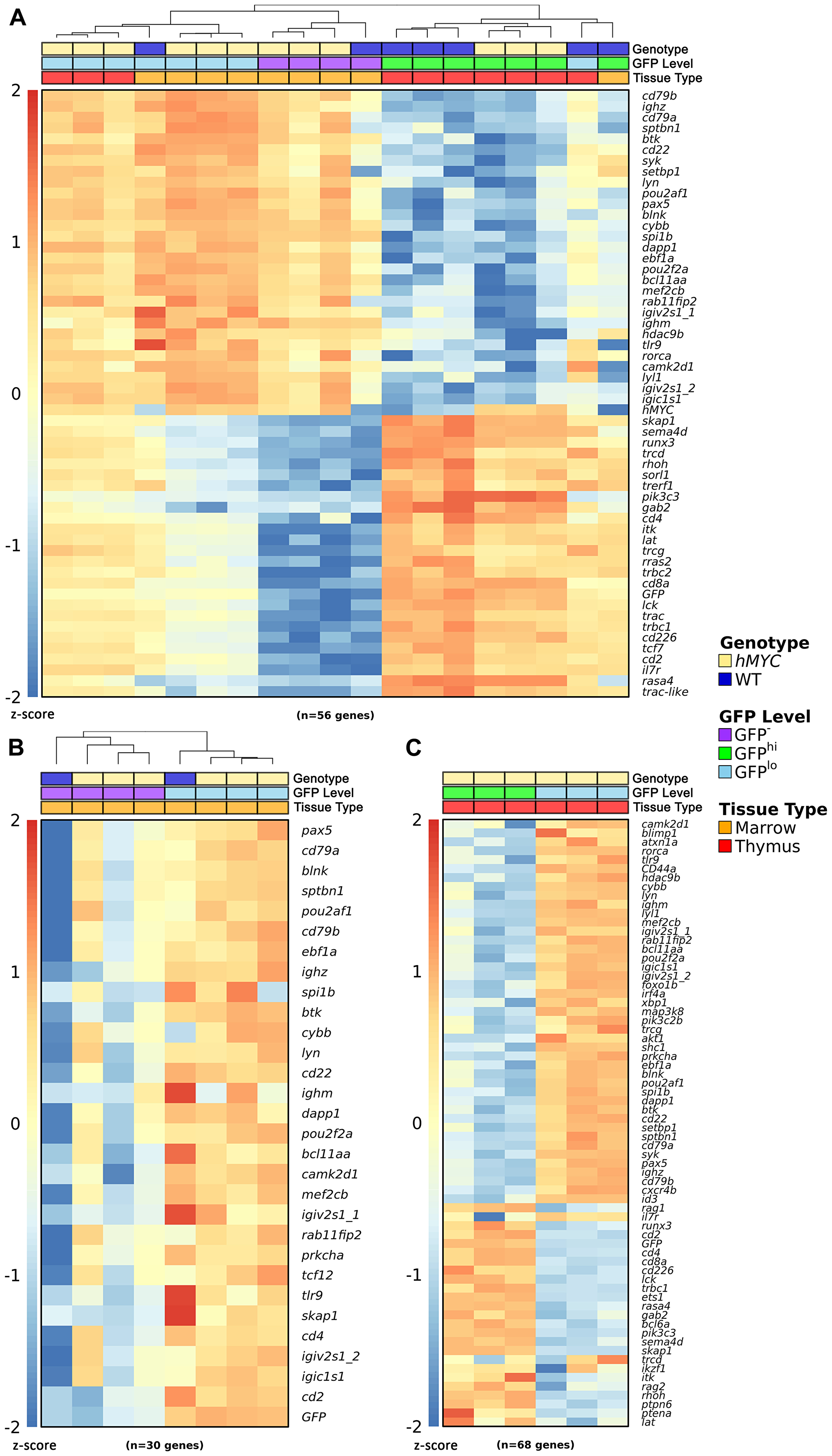
Figure 3: Gene expression in zebrafish lymphocytes. (A) Nanostring™ expression data in GFP-, GFPlo, and GFPhi lymphocytes from WT or rag2:hMYC fish bearing the lck:GFP marker transgene. hMYC GFPlo cells from thymus and marrow express B cell genes, as do WT GFPlo marrow cells. hMYC and WT GFP- marrow cells also express B cell transcripts. WT and hMYC GFPhi thymocytes express T cells genes, as do WT GFPhi marrow cells. WT GFPlo thymocyte fractions contain a mixture of B and T cells (as do hMYC GFPlo thymocytes and WT GFPlo marrow cells), due to the difficulty of obtaining pure GFPlo populations from these tissues [58, 63]. Samples are clustered hierarchically. (B) Nanostring™ expression data for B cell genes in marrow lymphocytes. WT and hMYC GFPlo marrow cells show the strongest B cell signature, but B cell hyperplasia in hMYC fish increases the B cell signature of GFP- hMYC marrow cells compared to WT GFP- marrow cells, which show little-to-no B cell gene expression. Samples are clustered hierarchically. (C) Nanostring™ expression data in hMYC thymocytes. hMYC GFPlo thymocytes express B, but not T, cell genes; hMYC GFPhi thymocytes show the opposite pattern. In panels (A–C), triplicates [hMYC/GFPlo/thymus (tan/lt. blue/red), hMYC/GFPlo/marrow (tan/lt. blue/orange), hMYC/GFP-/marrow (tan/purple/orange), WT/GFPhi/thymus (dk. blue/green/red), and hMYC/GFPhi/thymus (tan/green/red)] represent individual biologic replicates, where each column depicts results from RNA of cells pooled from 10 fish. For each singleton sample [WT/GFPlo/marrow (dk. blue/lt. blue/orange), WT/GFP-/marrow (dk. blue/purple/orange), WT/GFPlo/thymus (dk. blue/lt. blue/red), and WT/GFPhi/marrow (dk. blue/green/orange)], cells were pooled from 30 fish of the indicated genotype for RNA extraction. In panels A–C, read counts were log-transformed and converted to z-scores for visualization purposes. Scale bars are shown at the left of each heatmap.
These results support our conclusions, but using bulk gene expression patterns to compare populations that contain multiple cell types is challenging. Alternatively, single cell analyses can unambiguously define cell identities and precisely quantify different cell types. Such approaches can also reveal heterogeneity in seemingly-uniform populations, which is more powerful than binary categorizations like simple lineage assignments. Recognizing differences between cells of the same lineage, or cells of the same cancer, also has functional relevance, since distinct gene expression patterns correspond to different cellular phenotypes. Lymphocytes are diverse and complex populations, composed of not only B, T, and NK cells, but also innate lymphoid cells (ILC) [81]. Moreover, each of these cell types have multiple additional subtypes, such as the distinct IgM and IgZ B cell lineages in teleost fish [61, 82–84] or the various T lymphocyte subtypes [e. g., cytotoxic (CD8+ CTL), helper (CD4+ TH), TH17, and regulatory (Treg) subclasses] [85–87].
Diversity is not unique to normal lymphocytes; lymphoid cancers also contain heterogeneous populations, as previously shown in D. rerio mMyc-driven T-ALL [15, 40, 45, 48]. To assess tumor heterogeneity in hMYC-induced ALL and begin to define its extent, we analyzed mRNA expression in single cells from B- or T-ALL from hMYC fish (Figure 4). Animals exhibited either dim or bright cancers by fluorescent microscopy (Figure 4A) and by FACS analysis (Figure 4B). Using Fluidigm Biomark™ multiplex quantitative reverse transcriptase polymerase chain reaction (PCR primers are listed in Supplementary Table 1), we analyzed the expression of 17 transcripts: 10 B-lineage, 6 T-lineage, and as a mRNA threshold control, eukaryotic translation elongation factor 1 alpha 1, like 1 (ef1a) in 10 B- and 10 T-ALL individual cells (Figure 4C). All 20 B- and T-ALL cells profiled in accordance with their expected lineage, but expression differences of specific genes in single cells (e. g., pax5, id3, etc.) were evident. In addition, select genes (e. g., foxo1b, skap1) occasionally were mis-expressed by cells of the ‘wrong’ lineage. The significance of these findings remain to be determined, as malignant cells express aberrant markers on occasion [88], but biphenotypic ALL was demonstrated in mMyc fish, so what has previously been termed gene ‘mis-expression’ may actually represent an intriguing finding [57, 60]. Future single-cell studies hold promise as a means to address these and other unanswered questions about the cellular heterogeneity amongst lymphocytes, and within individual cancers, so as to define the precise gene expression differences that underlie their differing cellular phenotypes in vivo.
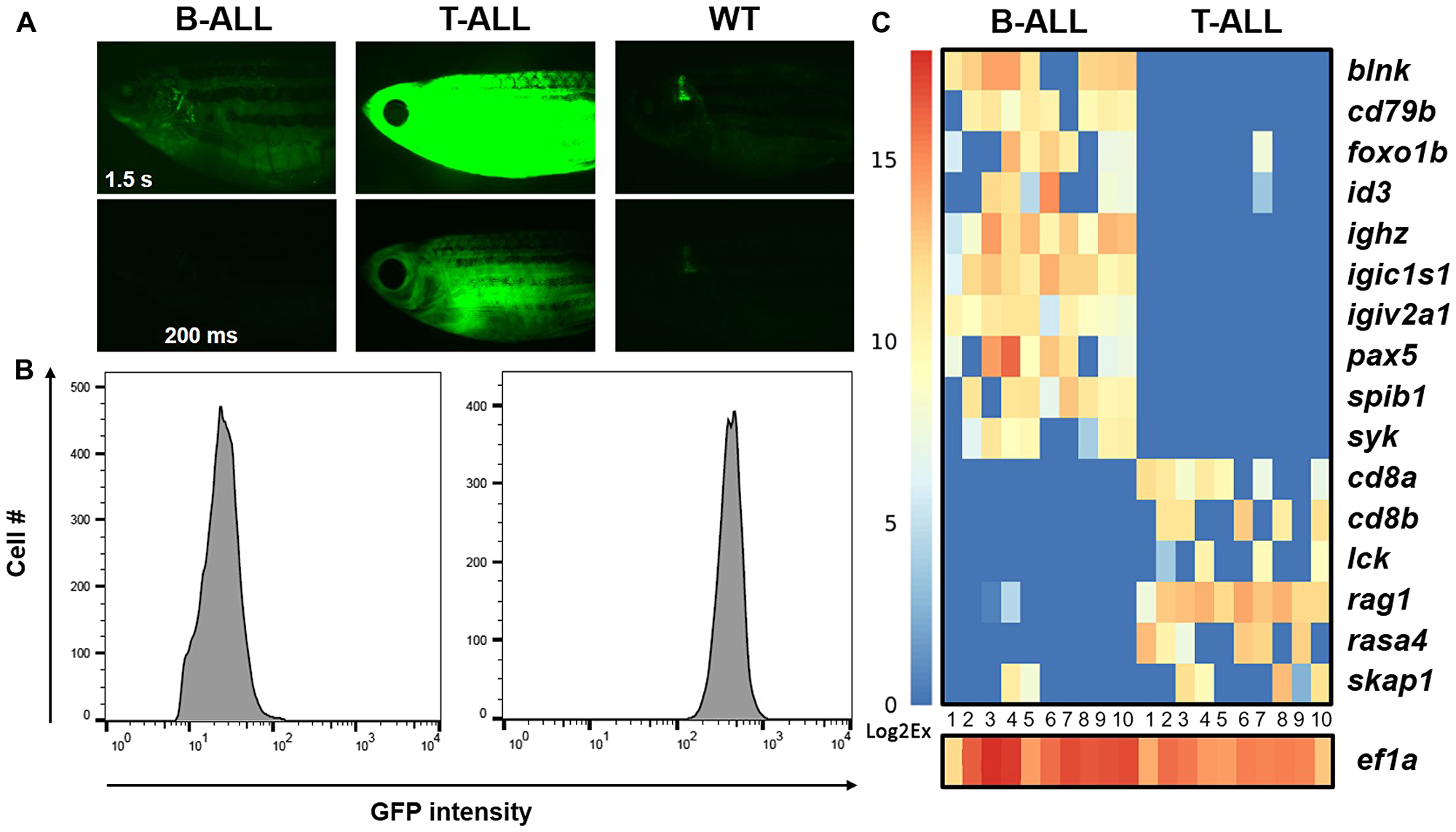
Figure 4: Gene expression in individual hMYC B- and T-ALL cells. (A) rag2:hMYC;lck: GFP fish with dim (left) and bright (center) cancers. B-ALL is only visible with 1.5 s exposure (upper left); T-ALL is visible with either 1.5 s (upper middle) or 200 ms (lower middle) exposures. Control lck:GFP fish images are shown at right, where only the thymus is visible. (B) Flow cytometry plots of the same ALL samples demonstrate GFPlo B-ALL and GFPhi T-ALL specimens. (C) Gene expression in single B- and T-ALL cells of the same cancers (n = 10 cells for each), as determined by Fluidigm™ Biomark qRT-PCR. B-ALL cells express B cell, but not T cell, genes; T-ALL show the opposite pattern. Expression of eef1a1l1 (homologue of human eukaryotic translation elongation factor 1 alpha 1, aka EF1A) was used as a threshold control for the presence of RNA in each well. CT values were converted to Log2Ex for visualization (scale bar at left).
Distinct types of B-ALL driven by hMYC and mMyc
MYC acts oncogenically in many ALL cases, and MYC is hyperactive in several other forms of cancer also [89], so it is not surprising that hMYC and mMyc can induce ALL in several types of lymphocytes [60]. However, the human and mouse MYC oncoproteins in these transgenic fish are highly conserved (435 amino acids, ~92% identical, ~94% similar; Figure 5), and both lines regulate MYC using the same zebrafish rag2 promoter, so it was surprising to discover that B-ALL in hMYC and mMyc fish are quite different. Specifically, hMYC B-ALL arise in ighz-lineage precursor B cells, whereas mMyc B-ALL occur in pro-B cells of the ighm lineage [60]. Unlike mammals which lack IgZ, expression of the zebrafish IgZ isotype does not occur by class switching. Instead, because of the configuration of the D. rerio immunoglobulin heavy chain (IgH) locus, during VDJ recombination zebrafish B cells must delete ighz sequences in order to express IgM [82]. Thus, when B cells commit to the IgM lineage, they lose their ability to express IgZ. As such, IgZ B cells, which are believed to primarily mediate mucosal immunity, can be viewed as an earlier stage in zebrafish B cell development that precedes IgM lineage commitment.

Figure 5: Alignment of human and murine MYC proteins. Transgenic human and mouse MYC are 435 and 439 amino acids, respectively (last four mMyc residues not shown). Proteins show > 91% identity (399 residues, shaded light blue) and > 94% similarity (410 residues; 11 additional similar residues shaded darker blue).
We used RNA-seq to profile gene expression in hMYC ighz+ and mMyc ighm+ B-ALL (Figure 6) [60]. We compared 4 hMYC B-ALL to duplicate transcriptomes from the only 2 mMyc B-ALL thus far reported [57]. As noted, this revealed hMYC B-ALL expressed ighz constant regions while mMyc B-ALL expressed ighm constant region mRNA [60], plus additional differences. Overall, > 400 genes displayed statistically-significant differential expression (Figure 6A and Supplementary Tables 2 and 3), including several pathways of interest. Curiously, in hMYC B-ALL much higher expression levels were seen for 9 of 10 fos and jun family members in the zebrafish genome (green bar in Figure 6A) with the lone exception fosl1b, which was higher in mMyc B-ALL. Also intriguing was the finding that all 6 endogenous D. rerio MYC family members (myca, mycb, mycn, mych, mycla, and myclb; red bar in Figure 6A) were more highly expressed in hMYC-driven B-ALL, as were multiple MYC-binding proteins and several MAX proteins, which heterodimerize with MYC (purple bar in Figure 6A).
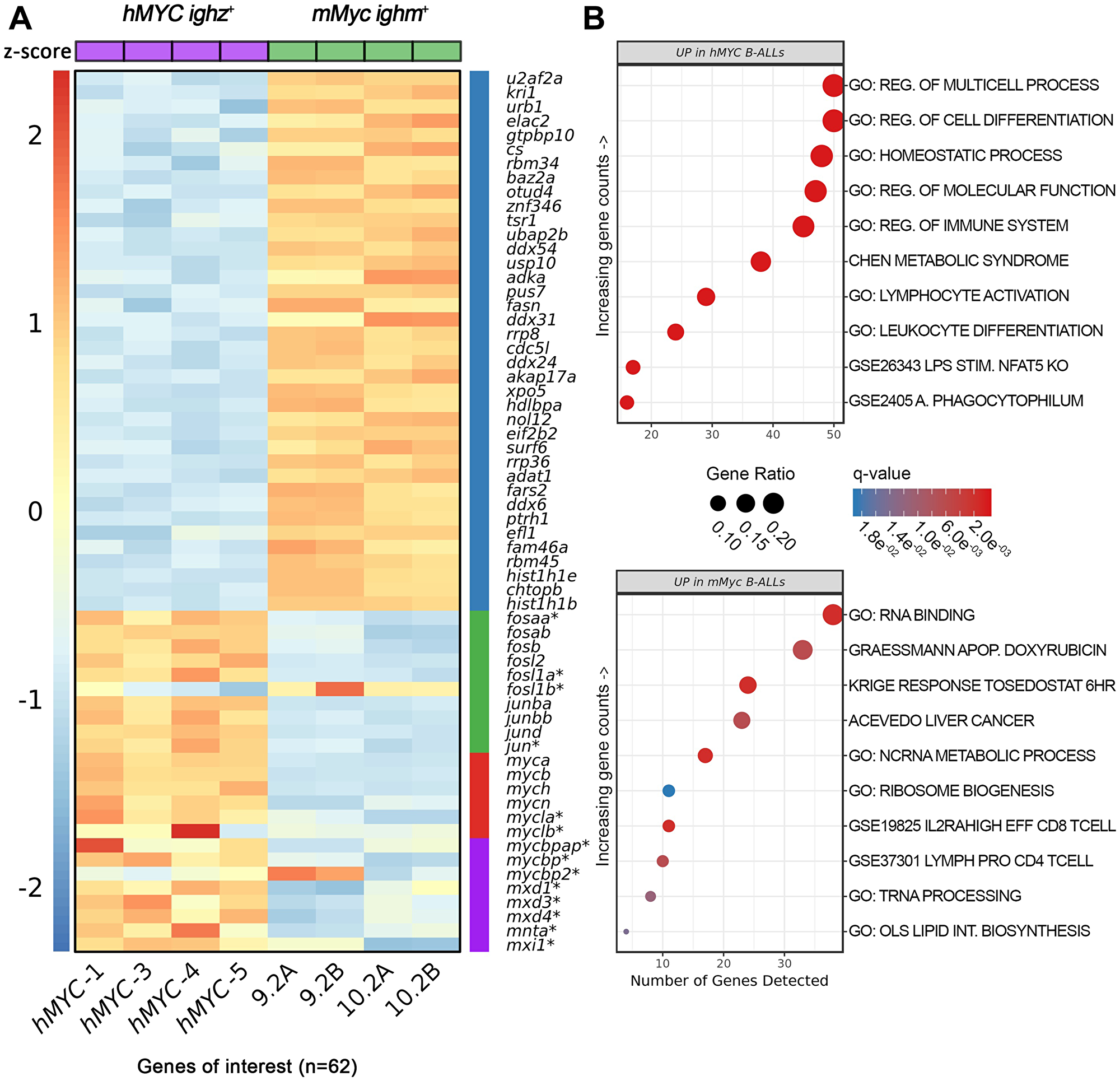
Figure 6: Differentially-expressed genes in hMYC versus mMyc B-ALL. (A) Heatmap depicting 62 differentially-expressed mRNA, including genes of the Gene Ontology (GO) RNA binding pathway (blue bar), D. rerio fos/jun-family members (green bar), endogenous myc-family members (red bar), and other myc-related proteins, including several max heterodimeric partners (purple bar). Samples hMYC-1, -3, -4, and -5 are four distinct hMYC B-ALL; 9.2A/B and 10.2A/B pairs are RNA-seq technical replicates of two mMyc B-ALL [57, 60]. Genes not meeting differential expression testing thresholds or other filtering criteria are marked by asterisks (FDR < 0.05, absolute fold-change ≥ 1.5). Read counts are shown as z-scores (scale bar at left). (B) Top ten biologic pathways up-regulated in hMYC (top) or mMyc B-ALL (bottom; FDR < 0.05). Pathways are ordered according to the number of genes detected in the gene set (x-axis) and colored based on FDR q-value. Data point sizes correspond to the percentage of genes over-expressed in each pathway.
B-ALL from the mMyc model showed dramatic up-regulation of many genes belonging to the Gene Ontology (GO) RNA binding pathway (blue bar in Figure 6A and top entry of Figure 6B lower panel). Several other genes also showed higher expression in mMyc B-ALL (Supplementary Table 3), including D. rerio homo/orthologues of mechanistic target of rapamycin kinase (MTOR) and the MTOR complex 1 member RPTOR, multiple cyclin-dependent kinases and related cell cycle regulators (CDK5, CDK14, CDC5L, CACUL1, TNK2), and the non-receptor tyrosine phosphatases PTPN13 and PTPN21. Together, these results suggest cell growth and division may be regulated differently in mMyc vs. hMYC B-ALL. Other notable orthologues up-regulated in mMyc B-ALL included the MYB-like gene MYSM1, two genes with jumonji lysine demethylase domains (JMJD8, JARID2), and the transcription factor NFAT5.
We postulate these and other differences may explain the apparently disparate oncogenic mechanisms employed by hMYC and mMyc in the B lymphoblasts of these closely-related lines. Pathway analysis of differentially-regulated genes predicted differing activation of several biologic pathways (e.g., cell differentiation, immune system process, lymphocyte activation, RNA binding, etc.; Figure 6B panels and Supplementary Table 4). These markedly different pathway signatures further demonstrate that human and murine MYC are far from synonymous in terms of their oncogenic effects upon zebrafish B lymphoblasts.
Conclusions
Recent discoveries of B-ALL in D. rerio lines previously-known to develop T-ALL were unexpected, but in retrospect, are not surprising. Both B and T lymphoblasts express rag2, so it is predictable that the transgenic promoters used in hMYC and mMyc fish would be active in both lymphocyte lineages. Likewise, MYC is known to be potently oncogenic in multiple types of B cell cancer, so the fact that mammalian MYC proteins can induce zebrafish B-ALL is no more surprising than their already well-documented activities in promoting zebrafish T-ALL. Rather, what is somewhat surprising is that over a decade passed between the first description of rag2:mMyc fish and the recognition that B-ALL occurred in them. This may reflect that B-ALL is less prevalent than T-ALL, but our incidence data in the related rag2:hMYC line (Figure 1A) demonstrate that B-ALL is still quite common—at least in hMYC fish. Irrespective of this, now that B-ALL are described, recognizing them is straightforward, either by differential GFP expression in the lck:GFP background, or with other transgenic markers like cd79a:GFP or cd79b:GFP (Figure 1C). In the case of these latter two lines, T-ALL will of course still develop, but go undetected. In view of this, it is worth noting that other forms of ALL are perhaps yet to be discovered in mMyc and hMYC fish, as alluded to by the discovery of a biphenotypic ALL of mixed B/T-lineage in the aforementioned mMyc study [57]. Going forward, continued efforts to discern the molecular mechanisms by which MYC mediates oncogenic effects in different lymphocyte lineages promise to yield even more insights into how this potent oncogene can drive cancer in zebrafish, and in humans.
ACKNOWLEDGMENTS AND FUNDING
We thank Megan-Malone Perez for zebrafish care, the Laboratory for Molecular Biology and Cytometry Research at OUHSC for use of their Core Facility, which assisted with FACS and Nanostring nCounter™ studies, and the Oklahoma Medical Research Foundation Clinical Genomics Center for assistance with Fluidigm Biomark™ multiplex qRT-PCR and RNA-seq. Research reported in this publication was supported in part by the National Cancer Institute Cancer Center Support Grant P30CA225520 awarded to the University of Oklahoma Stephenson Cancer Center and used the Molecular Biology Shared Resource. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. We also thank Sowmya Iyer, Sara Garcia, and David Langenau, Ph.D., at Massachusetts General Hospital and the Harvard Stem Cell Institute for their prior collaborative RNA-seq analyses. This work was supported by the OUHSC Jimmy Everest Section of Pediatric Hematology-Oncology, and grants from Hyundai Hope On Wheels, the Oklahoma Center for the Advancement of Science and Technology (HRP-067), and an INBRE pilot project award from the National Institute of General Medical Sciences (P20 GM103447). JKF holds the Children’s Hospital Foundation E.L. & Thelma Gaylord Endowed Chair in Pediatric Hematology-Oncology.
CONFLICTS OF INTEREST
Authors declare no conflicts of interests.
References
1. Pui CH, Yang JJ, Hunger SP, Pieters R, Schrappe M, Biondi A, Vora A, Baruchel A, Silverman LB, Schmiegelow K, Escherich G, Horibe K, Benoit YC, et al. Childhood Acute Lymphoblastic Leukemia: Progress Through Collaboration. J Clin Oncol. 2015; 33:2938–48. https://doi.org/10.1200/JCO.2014.59.1636. [PubMed].
2. Pui CH, Evans WE. A 50-year journey to cure childhood acute lymphoblastic leukemia. Semin Hematol. 2013; 50:185–96. https://doi.org/10.1053/j.seminhematol.2013.06.007. [PubMed].
3. Ward E, DeSantis C, Robbins A, Kohler B, Jemal A. Childhood and adolescent cancer statistics, 2014. CA Cancer J Clin. 2014; 64:83–103. https://doi.org/10.3322/caac.21219. [PubMed].
4. Jabbour E, O'Brien S, Konopleva M, Kantarjian H. New insights into the pathophysiology and therapy of adult acute lymphoblastic leukemia. Cancer. 2015; 121:2517–28. https://doi.org/10.1002/cncr.29383. [PubMed].
5. Jabbour E, Pui CH, Kantarjian H. Progress and Innovations in the Management of Adult Acute Lymphoblastic Leukemia. JAMA Oncol. 2018; 4:1413–20. https://doi.org/10.1001/jamaoncol.2018.1915. [PubMed].
6. Curtin SC, Minino AM, Anderson RN. Declines in Cancer Death Rates Among Children and Adolescents in the United States, 1999–2014. NCHS Data Brief. 2016; 257:1–8. [PubMed].
7. Locatelli F, Schrappe M, Bernardo ME, Rutella S. How I treat relapsed childhood acute lymphoblastic leukemia. Blood. 2012; 120:2807–16. https://doi.org/10.1182/blood-2012-02-265884. [PubMed].
8. Woo JS, Alberti MO, Tirado CA. Childhood B-acute lymphoblastic leukemia: a genetic update. Exp Hematol Oncol. 2014; 3:16. https://doi.org/10.1186/2162-3619-3-16. [PubMed].
9. Kohnken R, Porcu P, Mishra A. Overview of the Use of Murine Models in Leukemia and Lymphoma Research. Front Oncol. 2017; 7:22. https://doi.org/10.3389/fonc.2017.00022. [PubMed].
10. Jacoby E, Chien CD, Fry TJ. Murine models of acute leukemia: important tools in current pediatric leukemia research. Front Oncol. 2014; 4:95. https://doi.org/10.3389/fonc.2014.00095. [PubMed].
11. Letrado P, de Miguel I, Lamberto I, Diez-Martinez R, Oyarzabal J. Zebrafish: Speeding Up the Cancer Drug Discovery Process. Cancer Res. 2018; 78:6048–58. https://doi.org/10.1158/0008-5472.CAN-18-1029. [PubMed].
12. Leslie M. Zebrafish larvae could help to personalize cancer treatments. Science. 2017; 357:745. https://doi.org/10.1126/science.357.6353.745. [PubMed].
13. Kirchberger S, Sturtzel C, Pascoal S, Distel M. Quo natas, Danio?-Recent Progress in Modeling Cancer in Zebrafish. Front Oncol. 2017; 7:186. https://doi.org/10.3389/fonc.2017.00186. [PubMed].
14. Langenau DM, Sweet-Cordero A, Wechsler-Reya RJ, Dyer MA. Preclinical Models Provide Scientific Justification and Translational Relevance for Moving Novel Therapeutics into Clinical Trials for Pediatric Cancer. Cancer Res. 2015; 75:5176–86. https://doi.org/10.1158/0008-5472.CAN-15-1308. [PubMed].
15. Blackburn JS, Langenau DM. Zebrafish as a model to assess cancer heterogeneity, progression and relapse. Dis Model Mech. 2014; 7:755–62. https://doi.org/10.1242/dmm.015842. [PubMed].
16. White R, Rose K, Zon L. Zebrafish cancer: the state of the art and the path forward. Nat Rev Cancer. 2013; 13:624–36. https://doi.org/10.1038/nrc3589. [PubMed].
17. Howe K, Clark MD, Torroja CF, Torrance J, Berthelot C, Muffato M, Collins JE, Humphray S, McLaren K, Matthews L, McLaren S, Sealy I, Caccamo M, et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature. 2013; 496:498–503. https://doi.org/10.1038/nature12111. [PubMed].
18. Zhuravleva J, Paggetti J, Martin L, Hammann A, Solary E, Bastie JN, Delva L. MOZ/TIF2-induced acute myeloid leukaemia in transgenic fish. Br J Haematol. 2008; 143:378–82. https://doi.org/10.1111/j.1365-2141.2008.07362.x. [PubMed].
19. Yeh JR, Munson KM, Chao YL, Peterson QP, Macrae CA, Peterson RT. AML1-ETO reprograms hematopoietic cell fate by downregulating scl expression. Development. 2008; 135:401–10. https://doi.org/10.1242/dev.008904. [PubMed].
20. Bolli N, Payne EM, Grabher C, Lee JS, Johnston AB, Falini B, Kanki JP, Look AT. Expression of the cytoplasmic NPM1 mutant (NPMc+) causes the expansion of hematopoietic cells in zebrafish. Blood. 2010; 115:3329–40. https://doi.org/10.1182/blood-2009-02-207225. [PubMed].
21. Dayyani F, Wang J, Yeh JR, Ahn EY, Tobey E, Zhang DE, Bernstein ID, Peterson RT, Sweetser DA. Loss of TLE1 and TLE4 from the del(9q) commonly deleted region in AML cooperates with AML1-ETO to affect myeloid cell proliferation and survival. Blood. 2008; 111:4338–47. https://doi.org/10.1182/blood-2007-07-103291. [PubMed].
22. Shen LJ, Chen FY, Zhang Y, Cao LF, Kuang Y, Zhong M, Wang T, Zhong H. MYCN transgenic zebrafish model with the characterization of acute myeloid leukemia and altered hematopoiesis. PLoS One. 2013; 8:e59070. https://doi.org/10.1371/journal.pone.0059070. [PubMed].
23. Zhang Y, Wang J, Wheat J, Chen X, Jin S, Sadrzadeh H, Fathi AT, Peterson RT, Kung AL, Sweetser DA, Yeh JR. AML1-ETO mediates hematopoietic self-renewal and leukemogenesis through a COX/beta-catenin signaling pathway. Blood. 2013; 121:4906–16. https://doi.org/10.1182/blood-2012-08-447763. [PubMed].
24. Forrester AM, Grabher C, McBride ER, Boyd ER, Vigerstad MH, Edgar A, Kai FB, Da'as SI, Payne E, Look AT, Berman JN. NUP98-HOXA9-transgenic zebrafish develop a myeloproliferative neoplasm and provide new insight into mechanisms of myeloid leukaemogenesis. Br J Haematol. 2011; 155:167–81. https://doi.org/10.1111/j.1365-2141.2011.08810.x. [PubMed].
25. Yeh JR, Munson KM, Elagib KE, Goldfarb AN, Sweetser DA, Peterson RT. Discovering chemical modifiers of oncogene-regulated hematopoietic differentiation. Nat Chem Biol. 2009; 5:236–43. https://doi.org/nchembio.147. [PubMed].
26. Alghisi E, Distel M, Malagola M, Anelli V, Santoriello C, Herwig L, Krudewig A, Henkel CV, Russo D, Mione MC. Targeting oncogene expression to endothelial cells induces proliferation of the myelo-erythroid lineage by repressing the Notch pathway. Leukemia. 2013; 27:2229–41. https://doi.org/10.1038/leu.2013.132. [PubMed].
27. Feng H, Langenau DM, Madge JA, Quinkertz A, Gutierrez A, Neuberg DS, Kanki JP, Look AT. Heat-shock induction of T-cell lymphoma/leukaemia in conditional Cre/lox-regulated transgenic zebrafish. Br J Haematol. 2007; 138:169–75. https://doi.org/10.1111/j.1365-2141.2007.06625.x. [PubMed].
28. Feng H, Stachura DL, White RM, Gutierrez A, Zhang L, Sanda T, Jette CA, Testa JR, Neuberg DS, Langenau DM, Kutok JL, Zon LI, Traver D, et al. T-lymphoblastic lymphoma cells express high levels of BCL2, S1P1, and ICAM1, leading to a blockade of tumor cell intravasation. Cancer Cell. 2010; 18:353–66. https://doi.org/10.1016/j.ccr.2010.09.009. [PubMed].
29. Langenau DM, Traver D, Ferrando AA, Kutok JL, Aster JC, Kanki JP, Lin S, Prochownik E, Trede NS, Zon LI, Look AT. Myc-induced T cell leukemia in transgenic zebrafish. Science. 2003; 299:887–90. https://doi.org/10.1126/science.1080280. [PubMed].
30. Langenau DM, Feng H, Berghmans S, Kanki JP, Kutok JL, Look AT. Cre/lox-regulated transgenic zebrafish model with conditional myc-induced T cell acute lymphoblastic leukemia. Proc Natl Acad Sci U S A. 2005; 102:6068–73. https://doi.org/10.1073/pnas.0408708102. [PubMed].
31. Frazer JK, Meeker ND, Rudner L, Bradley DF, Smith AC, Demarest B, Joshi D, Locke EE, Hutchinson SA, Tripp S, Perkins SL, Trede NS. Heritable T-cell malignancy models established in a zebrafish phenotypic screen. Leukemia. 2009; 23:1825–35. https://doi.org/10.1038/leu.2009.116. [PubMed].
32. Gutierrez A, Grebliunaite R, Feng H, Kozakewich E, Zhu S, Guo F, Payne E, Mansour M, Dahlberg SE, Neuberg DS, den Hertog J, Prochownik EV, Testa JR, et al. Pten mediates Myc oncogene dependence in a conditional zebrafish model of T cell acute lymphoblastic leukemia. J Exp Med. 2011; 208:1595–603. https://doi.org/10.1084/jem.20101691. [PubMed].
33. Reynolds C, Roderick JE, LaBelle JL, Bird G, Mathieu R, Bodaar K, Colon D, Pyati U, Stevenson KE, Qi J, Harris M, Silverman LB, Sallan SE, et al. Repression of BIM mediates survival signaling by MYC and AKT in high-risk T-cell acute lymphoblastic leukemia. Leukemia. 2014; 28:1819–27. https://doi.org/10.1038/leu.2014.78. [PubMed].
34. Anderson NM, Li D, Peng HL, Laroche FJ, Mansour MR, Gjini E, Aioub M, Helman DJ, Roderick JE, Cheng T, Harrold I, Samaha Y, Meng L, et al. The TCA cycle transferase DLST is important for MYC-mediated leukemogenesis. Leukemia. 2016; 30:1365–74. https://doi.org/10.1038/leu.2016.26. [PubMed].
35. Huiting LN, Samaha Y, Zhang GL, Roderick JE, Li B, Anderson NM, Wang YW, Wang L, Laroche F, Choi JW, Liu CT, Kelliher MA, Feng H. UFD1 contributes to MYC-mediated leukemia aggressiveness through suppression of the proapoptotic unfolded protein response. Leukemia. 2018; 32:2339–2351. https://doi.org/10.1038/s41375-018-0141-x. [PubMed].
36. Chen J, Jette C, Kanki JP, Aster JC, Look AT, Griffin JD. NOTCH1-induced T-cell leukemia in transgenic zebrafish. Leukemia. 2007; 21:462–71. https://doi.org/10.1038/sj.leu.2404546. [PubMed].
37. Mansour MR, He S, Li Z, Lobbardi R, Abraham BJ, Hug C, Rahman S, Leon TE, Kuang YY, Zimmerman MW, Blonquist T, Gjini E, Gutierrez A, et al. JDP2: An oncogenic bZIP transcription factor in T cell acute lymphoblastic leukemia. J Exp Med. 2018; 215:1929–45. https://doi.org/10.1084/jem.20170484. [PubMed].
38. Rudner LA, Brown KH, Dobrinski KP, Bradley DF, Garcia MI, Smith AC, Downie JM, Meeker ND, Look AT, Downing JR, Gutierrez A, Mullighan CG, Schiffman JD, et al. Shared acquired genomic changes in zebrafish and human T-ALL. Oncogene. 2011; 30:4289–96. https://doi.org/10.1038/onc.2011.138. [PubMed].
39. Blackburn JS, Liu S, Wilder JL, Dobrinski KP, Lobbardi R, Moore FE, Martinez SA, Chen EY, Lee C, Langenau DM. Clonal evolution enhances leukemia-propagating cell frequency in T cell acute lymphoblastic leukemia through Akt/mTORC1 pathway activation. Cancer Cell. 2014; 25:366–78. https://doi.org/10.1016/j.ccr.2014.01.032. [PubMed].
40. Blackburn JS, Liu S, Raiser DM, Martinez SA, Feng H, Meeker ND, Gentry J, Neuberg D, Look AT, Ramaswamy S, Bernards A, Trede NS, Langenau DM. Notch signaling expands a pre-malignant pool of T-cell acute lymphoblastic leukemia clones without affecting leukemia-propagating cell frequency. Leukemia. 2012; 26:2069–78. https://doi.org/10.1038/leu.2012.116. [PubMed].
41. Gutierrez A, Feng H, Stevenson K, Neuberg DS, Calzada O, Zhou Y, Langenau DM, Look AT. Loss of function tp53 mutations do not accelerate the onset of myc-induced T-cell acute lymphoblastic leukaemia in the zebrafish. Br J Haematol. 2014; 166:84–90. https://doi.org/10.1111/bjh.12851. [PubMed].
42. Lobbardi R, Pinder J, Martinez-Pastor B, Theodorou M, Blackburn JS, Abraham BJ, Namiki Y, Mansour M, Abdelfattah NS, Molodtsov A, Alexe G, Toiber D, de Waard M, et al. TOX Regulates Growth, DNA Repair, and Genomic Instability in T-cell Acute Lymphoblastic Leukemia. Cancer Discov. 2017; 7:1336–53. https://doi.org/10.1158/2159-8290.CD-17-0267. [PubMed].
43. Ridges S, Heaton WL, Joshi D, Choi H, Eiring A, Batchelor L, Choudhry P, Manos EJ, Sofla H, Sanati A, Welborn S, Agarwal A, Spangrude GJ, et al. Zebrafish screen identifies novel compound with selective toxicity against leukemia. Blood. 2012; 119:5621–31. https://doi.org/10.1182/blood-2011-12-398818. [PubMed].
44. Gutierrez A, Pan L, Groen RW, Baleydier F, Kentsis A, Marineau J, Grebliunaite R, Kozakewich E, Reed C, Pflumio F, Poglio S, Uzan B, Clemons P, et al. Phenothiazines induce PP2A-mediated apoptosis in T cell acute lymphoblastic leukemia. J Clin Invest. 2014; 124:644–55. https://doi.org/10.1172/JCI65093. [PubMed].
45. Tang Q, Moore JC, Ignatius MS, Tenente IM, Hayes MN, Garcia EG, Torres Yordan N, Bourque C, He S, Blackburn JS, Look AT, Houvras Y, Langenau DM. Imaging tumour cell heterogeneity following cell transplantation into optically clear immune-deficient zebrafish. Nat Commun. 2016; 7:10358. https://doi.org/10.1038/ncomms10358. [PubMed].
46. Langenau DM, Jette C, Berghmans S, Palomero T, Kanki JP, Kutok JL, Look AT. Suppression of apoptosis by bcl-2 overexpression in lymphoid cells of transgenic zebrafish. Blood. 2005; 105:3278–85. https://doi.org/10.1182/blood-2004-08-3073. [PubMed].
47. Smith AC, Raimondi AR, Salthouse CD, Ignatius MS, Blackburn JS, Mizgirev IV, Storer NY, de Jong JL, Chen AT, Zhou Y, Revskoy S, Zon LI, Langenau DM. High-throughput cell transplantation establishes that tumor-initiating cells are abundant in zebrafish T-cell acute lymphoblastic leukemia. Blood. 2010; 115:3296–303. https://doi.org/10.1182/blood-2009-10-246488. [PubMed].
48. Moore FE, Garcia EG, Lobbardi R, Jain E, Tang Q, Moore JC, Cortes M, Molodtsov A, Kasheta M, Luo CC, Garcia AJ, Mylvaganam R, Yoder JA, et al. Single-cell transcriptional analysis of normal, aberrant, and malignant hematopoiesis in zebrafish. J Exp Med. 2016; 213:979–92. https://doi.org/10.1084/jem.20152013. [PubMed].
49. Tang Q, Iyer S, Lobbardi R, Moore JC, Chen H, Lareau C, Hebert C, Shaw ML, Neftel C, Suva ML, Ceol CJ, Bernards A, Aryee M, et al. Dissecting hematopoietic and renal cell heterogeneity in adult zebrafish at single-cell resolution using RNA sequencing. J Exp Med. 2017; 214:2875–87. https://doi.org/10.1084/jem.20170976. [PubMed].
50. Baeten JT, Waarts MR, Pruitt MM, Chan WC, Andrade J, de Jong JLO. The side population enriches for leukemia propagating cell activity and Wnt pathway expression in zebrafish acute lymphoblastic leukemia. Haematologica. 2019; 104:1388–1395. https://doi.org/10.3324/haematol.2018.206417. [PubMed].
51. Leong WZ, Tan SH, Ngoc PCT, Amanda S, Yam AWY, Liau WS, Gong Z, Lawton LN, Tenen DG, Sanda T. ARID5B as a critical downstream target of the TAL1 complex that activates the oncogenic transcriptional program and promotes T-cell leukemogenesis. Genes Dev. 2017; 31:2343–60. https://doi.org/10.1101/gad.302646.117. [PubMed].
52. Liau WS, Tan SH, Ngoc PCT, Wang CQ, Tergaonkar V, Feng H, Gong Z, Osato M, Look AT, Sanda T. Aberrant activation of the GIMAP enhancer by oncogenic transcription factors in T-cell acute lymphoblastic leukemia. Leukemia. 2017; 31:1798–807. https://doi.org/10.1038/leu.2016.392. [PubMed].
53. Li Z, He S, Look AT. The MCL1-specific inhibitor S63845 acts synergistically with venetoclax/ABT-199 to induce apoptosis in T-cell acute lymphoblastic leukemia cells. Leukemia. 2019; 33:262–6. https://doi.org/10.1038/s41375-018-0201-2. [PubMed].
54. Sabaawy HE, Azuma M, Embree LJ, Tsai HJ, Starost MF, Hickstein DD. TEL-AML1 transgenic zebrafish model of precursor B cell acute lymphoblastic leukemia. Proc Natl Acad Sci U S A. 2006; 103:15166–71. https://doi.org/10.1073/pnas.0603349103. [PubMed].
55. He S, Jing CB, Look AT. Zebrafish models of leukemia. Methods Cell Biol. 2017; 138:563–92. https://doi.org/10.1016/bs.mcb.2016.11.013. [PubMed].
56. Squiban B, Frazer JK. Danio rerio: Small Fish Making a Big Splash in Leukemia. Curr Pathobiol Rep. 2014; 2:61–73. https://doi.org/10.1007/s40139-014-0041-3. [PubMed].
57. Garcia EG, Iyer S, Garcia SP, Loontiens S, Sadreyev RI, Speleman F, Langenau DM. Cell of origin dictates aggression and stem cell number in acute lymphoblastic leukemia. Leukemia. 2018; 32:1860–5. https://doi.org/10.1038/s41375-018-0130-0. [PubMed].
58. Borga C, Park G, Foster C, Burroughs-Garcia J, Marchesin M, Shah R, Hasan A, Ahmed ST, Bresolin S, Batchelor L, Scordino T, Miles RR, Te Kronnie G, et al. Simultaneous B and T cell acute lymphoblastic leukemias in zebrafish driven by transgenic MYC: implications for oncogenesis and lymphopoiesis. Leukemia. 2019; 33:333–47. https://doi.org/10.1038/s41375-018-0226-6. [PubMed].
59. Langenau DM, Ferrando AA, Traver D, Kutok JL, Hezel JP, Kanki JP, Zon LI, Look AT, Trede NS. In vivo tracking of T cell development, ablation, and engraftment in transgenic zebrafish. Proc Natl Acad Sci U S A. 2004; 101:7369–74. https://doi.org/10.1073/pnas.0402248101. [PubMed].
60. Borga C, Foster CA, Iyer S, Garcia SP, Langenau DM, Frazer JK. Molecularly distinct models of zebrafish Myc-induced B cell leukemia. Leukemia. 2019; 33:559–62. https://doi.org/10.1038/s41375-018-0328-1. [PubMed].
61. Liu X, Li YS, Shinton SA, Rhodes J, Tang L, Feng H, Jette CA, Look AT, Hayakawa K, Hardy RR. Zebrafish B Cell Development without a Pre-B Cell Stage, Revealed by CD79 Fluorescence Reporter Transgenes. J Immunol. 2017; 199:1706–15. https://doi.org/10.4049/jimmunol.1700552. [PubMed].
62. Traver D, Paw BH, Poss KD, Penberthy WT, Lin S, Zon LI. Transplantation and in vivo imaging of multilineage engraftment in zebrafish bloodless mutants. Nat Immunol. 2003; 4:1238–46. https://doi.org/10.1038/ni1007. [PubMed].
63. Burroughs-Garcia J, Hasan A, Park G, Borga C, Frazer JK. Isolating Malignant and Non-Malignant B Cells from lck:eGFP Zebrafish. J Vis Exp. 2019; 144:e59191. https://doi.org/10.3791/59191. [PubMed].
64. Christoph S, Deryckere D, Schlegel J, Frazer JK, Batchelor LA, Trakhimets AY, Sather S, Hunter DM, Cummings CT, Liu J, Yang C, Kireev D, Simpson C, et al. UNC569, a novel small-molecule mer inhibitor with efficacy against acute lymphoblastic leukemia in vitro and in vivo. Mol Cancer Ther. 2013; 12:2367–77. https://doi.org/10.1158/1535-7163.MCT-13-0040. [PubMed].
65. Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013; 29:15–21. https://doi.org/10.1093/bioinformatics/bts635. [PubMed].
66. Zerbino DR, Achuthan P, Akanni W, Amode MR, Barrell D, Bhai J, Billis K, Cummins C, Gall A, Giron CG, Gil L, Gordon L, Haggerty L, et al. Ensembl 2018. Nucleic Acids Res. 2018; 46:D754–D61. https://doi.org/10.1093/nar/gkx1098. [PubMed].
67. Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014; 30:923–30. https://doi.org/10.1093/bioinformatics/btt656. [PubMed].
68. Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014; 15:550. https://doi.org/10.1186/s13059-014-0550-8. [PubMed].
69. Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012; 16:284–7. https://doi.org/10.1089/omi.2011.0118. [PubMed].
70. Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP. Molecular signatures database (MSigDB) 3.0. Bioinformatics. 2011; 27:1739–40. https://doi.org/10.1093/bioinformatics/btr260. [PubMed].
71. Childhood Acute Lymphoblastic Leukemia Treatment (PDQ(R)): Health Professional Version. PDQ Cancer Information Summaries. (Bethesda (MD). 2002.
72. Adult Acute Lymphoblastic Leukemia Treatment (PDQ(R)): Health Professional Version. PDQ Cancer Information Summaries. (Bethesda (MD). 2002.
73. Shah DS, Kumar R. Steroid resistance in leukemia. World J Exp Med. 2013; 3:21–5. https://doi.org/10.5493/wjem.v3.i2.21. [PubMed].
74. Hochberg J, Cairo MS. Tumor lysis syndrome: current perspective. Haematologica. 2008; 93:9–13. https://doi.org/10.3324/haematol.12327. [PubMed].
75. Novershtern N, Subramanian A, Lawton LN, Mak RH, Haining WN, McConkey ME, Habib N, Yosef N, Chang CY, Shay T, Frampton GM, Drake AC, Leskov I, et al. Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell. 2011; 144:296–309. https://doi.org/10.1016/j.cell.2011.01.004. [PubMed].
76. Haferlach T, Kohlmann A, Wieczorek L, Basso G, Kronnie GT, Bene MC, De Vos J, Hernandez JM, Hofmann WK, Mills KI, Gilkes A, Chiaretti S, Shurtleff SA, et al. Clinical utility of microarray-based gene expression profiling in the diagnosis and subclassification of leukemia: report from the International Microarray Innovations in Leukemia Study Group. J Clin Oncol. 2010; 28:2529–37. https://doi.org/10.1200/JCO.2009.23.4732. [PubMed].
77. Geiss GK, Bumgarner RE, Birditt B, Dahl T, Dowidar N, Dunaway DL, Fell HP, Ferree S, George RD, Grogan T, James JJ, Maysuria M, Mitton JD, et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol. 2008; 26:317–25. https://doi.org/10.1038/nbt1385. [PubMed].
78. Malkov VA, Serikawa KA, Balantac N, Watters J, Geiss G, Mashadi-Hossein A, Fare T. Multiplexed measurements of gene signatures in different analytes using the Nanostring nCounter Assay System. BMC Res Notes. 2009; 2:80. https://doi.org/10.1186/1756-0500-2-80. [PubMed].
79. Palmer C, Diehn M, Alizadeh AA, Brown PO. Cell-type specific gene expression profiles of leukocytes in human peripheral blood. BMC Genomics. 2006; 7:115. https://doi.org/10.1186/1471-2164-7-115. [PubMed].
80. Carmona SJ, Teichmann SA, Ferreira L, Macaulay IC, Stubbington MJ, Cvejic A, Gfeller D. Single-cell transcriptome analysis of fish immune cells provides insight into the evolution of vertebrate immune cell types. Genome Res. 2017; 27:451–61. https://doi.org/10.1101/gr.207704.116. [PubMed].
81. Hernandez PP, Strzelecka PM, Athanasiadis EI, Hall D, Robalo AF, Collins CM, Boudinot P, Levraud JP, Cvejic A. Single-cell transcriptional analysis reveals ILC-like cells in zebrafish. Sci Immunol. 2018; 3. https://doi.org/10.1126/sciimmunol.aau5265. [PubMed].
82. Fillatreau S, Six A, Magadan S, Castro R, Sunyer JO, Boudinot P. The astonishing diversity of Ig classes and B cell repertoires in teleost fish. Front Immunol. 2013; 4:28. https://doi.org/10.3389/fimmu.2013.00028. [PubMed].
83. Castro R, Jouneau L, Pham HP, Bouchez O, Giudicelli V, Lefranc MP, Quillet E, Benmansour A, Cazals F, Six A, Fillatreau S, Sunyer O, Boudinot P. Teleost fish mount complex clonal IgM and IgT responses in spleen upon systemic viral infection. PLoS Pathog. 2013; 9:e1003098. https://doi.org/10.1371/journal.ppat.1003098. [PubMed].
84. Page DM, Wittamer V, Bertrand JY, Lewis KL, Pratt DN, Delgado N, Schale SE, McGue C, Jacobsen BH, Doty A, Pao Y, Yang H, Chi NC, et al. An evolutionarily conserved program of B-cell development and activation in zebrafish. Blood. 2013; 122:e1–11. https://doi.org/10.1182/blood-2012-12-471029. [PubMed].
85. Zhang H, Fei C, Wu H, Yang M, Liu Q, Wang Q, Zhang Y. Transcriptome profiling reveals Th17-like immune responses induced in zebrafish bath-vaccinated with a live attenuated Vibrio anguillarum. PLoS One. 2013; 8:e73871. https://doi.org/10.1371/journal.pone.0073871. [PubMed].
86. Sugimoto K, Hui SP, Sheng DZ, Nakayama M, Kikuchi K. Zebrafish FOXP3 is required for the maintenance of immune tolerance. Dev Comp Immunol. 2017; 73:156–62. https://doi.org/10.1016/j.dci.2017.03.023. [PubMed].
87. Hui SP, Sheng DZ, Sugimoto K, Gonzalez-Rajal A, Nakagawa S, Hesselson D, Kikuchi K. Zebrafish Regulatory T Cells Mediate Organ-Specific Regenerative Programs. Dev Cell. 2017; 43:659–72.e5. https://doi.org/10.1016/j.devcel.2017.11.010. [PubMed].
88. Cernan M, Szotkowski T, Pikalova Z. Mixed-phenotype acute leukemia: state-of-the-art of the diagnosis, classification and treatment. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2017; 161:234–41. https://doi.org/10.5507/bp.2017.013. [PubMed].
89. Dang CV. MYC on the path to cancer. Cell. 2012; 149:22–35. https://doi.org/10.1016/j.cell.2012.03.003. [PubMed].

