Introduction
In the last century, carcinoma of the lung has progressed from an uncommon and obscure disease to the most common cancer in the world, and the most common cause of death from cancer. In 2018, new cases of lung cancer in the US were estimated at 121,680 for men and at 112,350 for women, for a total of 234,030 cases, representing the equivalent of 641 cases of lung cancer diagnosed each day [1, 2]. Many gene mutations have previously been identified in several solid tumors [3, 4]. For lung cancer, in particular, the most common mutations found are in the epidermal growth factor receptor (EGFR) gene and the K-Raf kinase gene family (KRAS) [5]. In addition to these somatic mutations, which are the most frequent, other mutations in several genes have been discovered, including BRAF, c-MET, DDR2, HER2, MAP2K1, NRAS, PIK3CA, and RET mutations [6, 7]. The main feature that characterizes these “niche” mutations is their low prevalence in the population of patients with lung cancer when compared with KRAS and EGFR. In the present study, we identified these “niche mutations” on the basis of a population prevalence that was equal to or less than 10% based on literature data [8–10]. In particular, according to recently published data concerning 14 healthcare institutions in the US, the prevalence for these mutations was as follows: KRAS 25%, EGFR 23%, HER2 2.7%, BRAF 2.6%, PIK3CA 0.8%, NRAS 0.7%, c-MET 0.7%, and MAP2K1 0.3% [8]. Moreover, the prevalence of the RET mutation in adenocarcinoma was estimated to be 1.7% [9], and the prevalence of DDR2 mutation in lung cancer was 2.2% [3, 10]. Overall, the KRAS mutation was considered to be the most lethal mutation, exhibiting the worst prognosis [3–5]. Nevertheless, the potential aggressiveness of niche mutations remains unclear [6, 8].
With regard to treatment, discoveries of gene mutations have allowed the development of targeted therapies, which are considered more effective for survival than chemotherapy in patients with advanced mutated disease [11].
The purpose of this observational retrospective cohort study was to assess differences in overall survival (OS) rates in patients with histologically diagnosed lung adenocarcinoma harboring either the most frequently detected or niche somatic mutations, to highlight new aspects to consider for prognosis and treatment. Considering the potential aggressiveness of niche mutations in this context, the technological advances of next-generation sequencing (NGS), which is currently used in clinical practice, represents a precise approach to identifying a large panel of mutations in oncologic patients [12].
Results
Description of patients
Data were gathered from 252 patients. The clinical and demographic characteristics of the studied patients are reported in Table 1. The average age was 67.3 ± 9.9 years, 63.5% of patients were male, and 82.5% of them were smokers. The majority of patients had been diagnosed with stage IV (63.1%) or III (19.0%) cancer according to the TNM (8th Edition). Patients were divided into three groups as follows: 42 (16.7%) patients in the niche mutations group, 155 (61.5%) in the KRAS group, and 55 (21.8%) in the EGFR group. We found differences in distributions of gender and smoking habits among the three cohorts. Male patients and smokers were more common in the KRAS group (74.8% males and 90.3% smokers) than in the niche mutations group (61.9% and 81.0%) or the EGFR mutation group (32.7% and 61.8%) (p = 0.000 and p = 0.000). Moreover, the cohorts received different treatments because the frequency of chemotherapy and target therapy was not similar between the groups. In the KRAS groups, chemotherapy was performed on 79.4% of the patients, whereas chemotherapy was performed on 69.0% and 38.2% of patients in the niche mutations and EGFR mutations groups, respectively (p = 0.000). However, target therapy was more common in the EGFR mutation group (52.7%) than in the KRAS mutation (0.0%) and niche mutations (7.1%) groups (p = 0.000). The three cohorts were similar in terms of the remaining characteristics (i. e., age (p = 0.376), TNM stage (p = 0.078), surgery (p = 0.940), radiotherapy (p = 0.462), and immunotherapy (p = 0.409). The most common first-line treatment was chemotherapy for the KRAS and niche mutations cohorts (53.5% and 42.9% of patients, respectively), whereas in the EGFR cohort, target therapy was more common (29.1% of patients). Eight (3.2%) patients did not undergo any treatment after diagnosis. The characteristics of patients harboring every single-niche mutation are reported in Supplementary Table 1.
Table 1: Patients characteristics at primary diagnosis and treatments after diagnosis
| KRAS (n = 155) | EGFR (n = 55) | Niche mutations (n = 42) | All patients (n = 252) | p-value | ||
|---|---|---|---|---|---|---|
| Patients characteristics at primary diagnosis | ||||||
| Age – Years | mean ± SD | 67.4 ± 9.4 | 65.9 ± 12.1 | 68.8 ± 8.4 | 67.3 ± 9.9 | 0.376 |
| Gender – M | n (%) | 116 (74.8%) | 18 (32.7%) | 26 (61.9%) | 160 (63.5%) | 0.000 |
| Smoker – Yes | n (%) | 140 (90.3%) | 34 (61.8%) | 34 (81.0%) | 208 (82.5%) | 0.000 |
| Stage - I | n (%) | 9 (5.8%) | 8 (14.5%) | 1 (2.4%) | 18 (7.1%) | 0.078 |
| Stage – II | n (%) | 18 (11.6%) | 5 (9.1%) | 4 (9.5%) | 27 (10.7%) | |
| Stage – III | n (%) | 27 (17.4%) | 7 (12.7%) | 14 (33.3%) | 48 (19.0%) | |
| Stage - IV | n (%) | 101 (65.2%) | 35 (63.6%) | 23 (54.8%) | 159 (63.1%) | |
| Treatments after diagnosis | ||||||
| Surgery | n (%) | 42 (27.1%) | 16 (29.1%) | 12 (28.6%) | 70 (27.8%) | 0.940 |
| RT | n (%) | 75 (48.4%) | 27 (49.1%) | 25 (59.5%) | 127 (50.4%) | 0.462 |
| CT | n (%) | 123 (79.4%) | 21 (38.2%) | 29 (69.0%) | 173 (68.7%) | 0.000 |
| IMT | n (%) | 36 (23.2%) | 8 (14.5%) | 8 (19.0%) | 52 (20.6%) | 0.409 |
| TT | n (%) | 0 (0.0%) | 29 (52.7%) | 3 (7.1%) | 32 (12.7%) | 0.000 |
| First line treatment | ||||||
| Surgery | n (%) | 35 (22.6%) | 12 (21.8%) | 9 (21.4%) | 56 (22.2%) | 0.000 |
| RT | n (%) | 27 (17.4%) | 8 (14.5%) | 7 (16.7%) | 42 (16.7%) | |
| CT | n (%) | 83 (53.5%) | 12 (21.8%) | 18 (42.9%) | 113 (44.8%) | |
| IMT | n (%) | 6 (3.9%) | 6 (10.9%) | 3 (7.1%) | 15 (6.0%) | |
| TT | n (%) | 0 (0.0%) | 16 (29.1%) | 2 (4.8%) | 18 (7.1%) | |
| No treatment | n (%) | 4 (2.6%) | 1 (1.8%) | 3 (7.1%) | 8 (3.2%) | |
Description of mutations
The observed distribution of single mutations is reported in Table 2. The most frequent mutation was found in the KRAS gene (found in 63.5% of the patients), whereas EGFR gene mutations were observed in 22.6% of the patients. In the niche mutations group, BRAF gene mutations were observed in 6.7% of the patients, whereas HER2 mutations were observed in 4.4% of the patients, PIK3CA gene mutations were detected in 4.0% of the patients and NRAS in 1.6%. No c-MET, DDR2, MAP2K1, or RET mutations were identified in any patient included in the study. Seven patients (2.8%) showed concomitant mutations in two genes and were all classified in the niche mutations cohort. The specific codons, exons, and amino acid alterations identified are reported in detail in Table 2.
Table 2: Mutations, codons, exons and amino acid alterations identified in the study population
| Frequent mutations | Niche mutations | Overall | ||||||
|---|---|---|---|---|---|---|---|---|
| n | % | n | % | n | % | |||
| KRAS single mutation | 155 | 61.5% | BRAF single mutation | 16 | 6.3% | KRAS | 160 | 63.5% |
| Codon 12 | 136 | 54.0% | Codon 469 | 4 | 1.6% | EGFR | 57 | 22.6% |
| G12C | 70 | 27.8% | Codon 600 | 6 | 2.4% | BRAF | 17 | 6.7% |
| G12V | 27 | 10.7% | Codon 466 | 3 | 1.2% | HER2 | 11 | 4.4% |
| G12D | 22 | 8.7% | HER2 single mutation | 10 | 4.0% | PIK3CA | 10 | 4.0% |
| G12A | 15 | 6.0% | Codon 775 & 776 ins YVMA | 7 | 2.8% | NRAS | 4 | 1.6% |
| Codon 13 | 13 | 5.2% | PIK3CA single mutation | 6 | 2.4% | Concomitant mutations | 7 | 2.8% |
| G13C | 8 | 3.2% | Codon 542 | 1 | 0.4% | |||
| Codon 61 | 6 | 2.4% | Codon 545 | 1 | 0.4% | |||
| EGFR single mutation | 55 | 21.8% | NRAS single mutation | 3 | 1.2% | |||
| Exon 19 | 16 | 6.3% | Codon 61 | 1 | 0.4% | |||
| Codon del 746 & 750 | 14 | 5.6% | Concomitant mutations | 7 | 2.8% | |||
| Exon 21 | 17 | 6.7% | KRAS & PIK3CA | 2 | 0.8% | |||
| Codon 858 | 13 | 5.2% | EGFR & PIK3CA | 2 | 0.8% | |||
| Exon 18 | 4 | 1.6% | KRAS & BRAF | 1 | 0.4% | |||
| Exon 20 | 3 | 1.2% | KRAS & NRAS | 1 | 0.4% | |||
| Exons 20 & 21 | 5 | 2.0% | KRAS & HER2 | 1 | 0.4% | |||
| Exons 19 & 20 | 4 | 1.6% | ||||||
| Exons 18 & 20 | 2 | 0.8% | ||||||
| Exons 19 & 21 | 1 | 0.4% | ||||||
Mortality rates
Over an average follow-up period of 1.3 (range 0.1 to 4.8) years, there were 168 (66.7%) deaths, 167 of which were cancer-related. In the niche mutations cohort, 29 (69.0%) patients died, in the KRAS gene cohorts 109 (70.3%) died, and in the EGFR gene cohort 30 (54.5%) died.
MRs for each gene mutation and cohort considered are reported in Figure 1. Patients with KRAS mutations had an MR of 0.59 (95% CI = 0.49–0.71) deaths per year, whereas the MRs were 0.31 (0.21–0.44) for EGFR gene patients, 0.54 (0.29–0.98) for BRAF gene patients, 0.42 (0.15–0.91) for HER2 gene patients, 0.69 (0.30–1.35) for PIK3CA gene patients, and 0.60 (0.12–0.74) for NRAS gene patients. The subjects with two mutations presented an MR of 0.70 (0.26–1.53). The mortality was higher in the KRAS gene cohort (MR = 0.58, 95% CI = 0.48–0.70) and in the niche mutations cohort (MR = 0.55, 95% CI = 0.37–0.78) than in the EGFR gene cohort (MR = 0.31, 95% CI = 0.21–0.44).
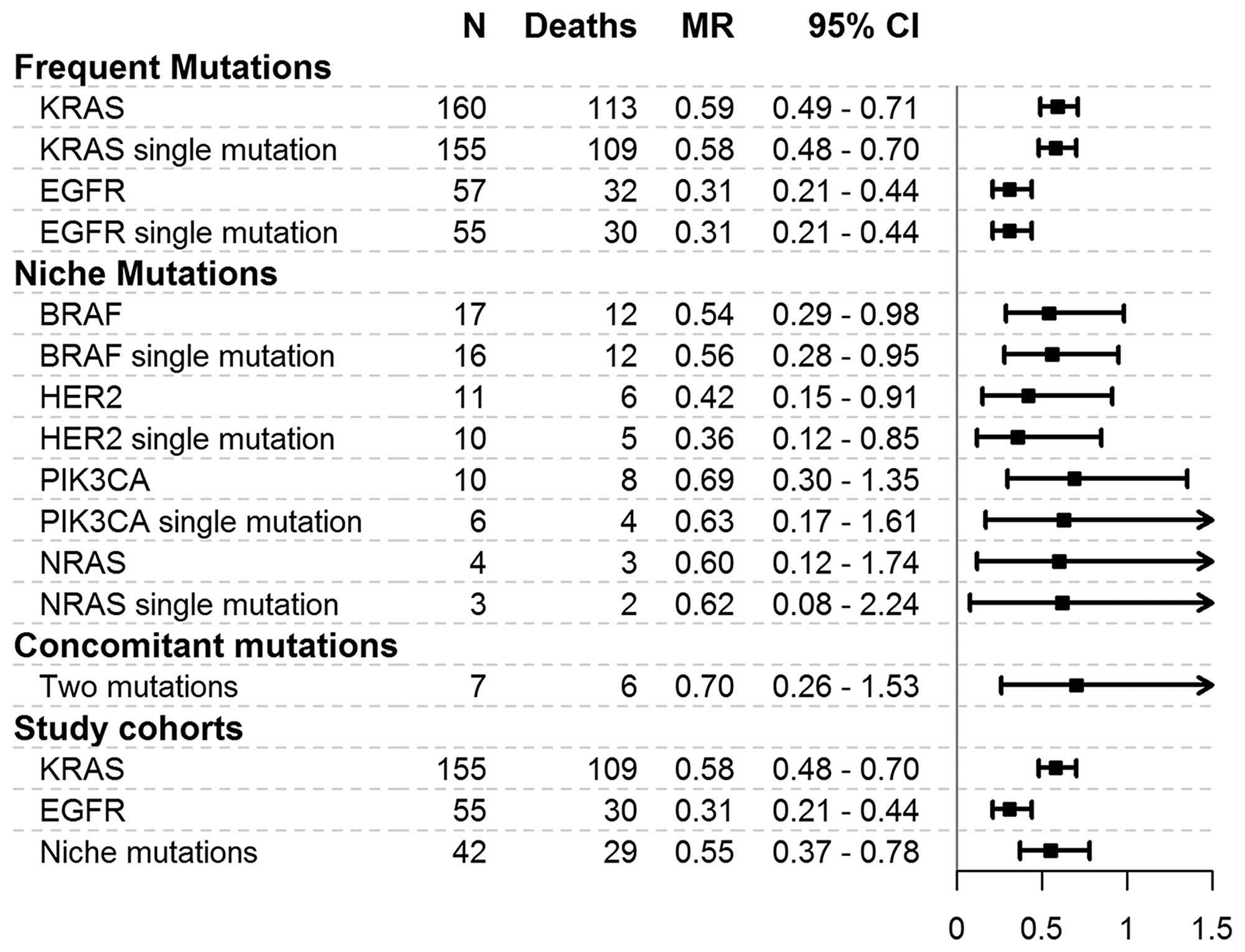
Figure 1: Mortality rates for mutations and cohorts considered in the study. MR = mortality rate, expressed as the number of deaths per person-year. 95% CI = 95% confidence interval. N = number of patients. The plot reports mortality rates and 95% confidence intervals.
Differences in overall survival
The survival curves for the three cohorts are shown in Figure 2. The median OS was measured as 1.5 years (95% CI = 1.2–1.8) for the whole population, 1.5 years (95% CI = 0.8–2.3) for the niche mutations cohort, 1.3 (95% CI = 1.0–1.6) years for the KRAS gene cohort, and 2.2 years (95% CI not estimable) for the EGFR gene cohort. The cumulative survival probability at 1 year from diagnosis was measured as 56.5% (95% CI = 49.1–65.0%) for the KRAS gene mutation cohort, as 60.1% (46.6–77.6%) for the niche mutations cohort, and as 69.2% (57.7–83.1%) for the EGFR gene mutations cohort, whereas at 2 years these values were measured as 28.4% (21.2–38.0%), 38.2% (24.9–58.6%), and 58.2% (45.8–73.8%), respectively.
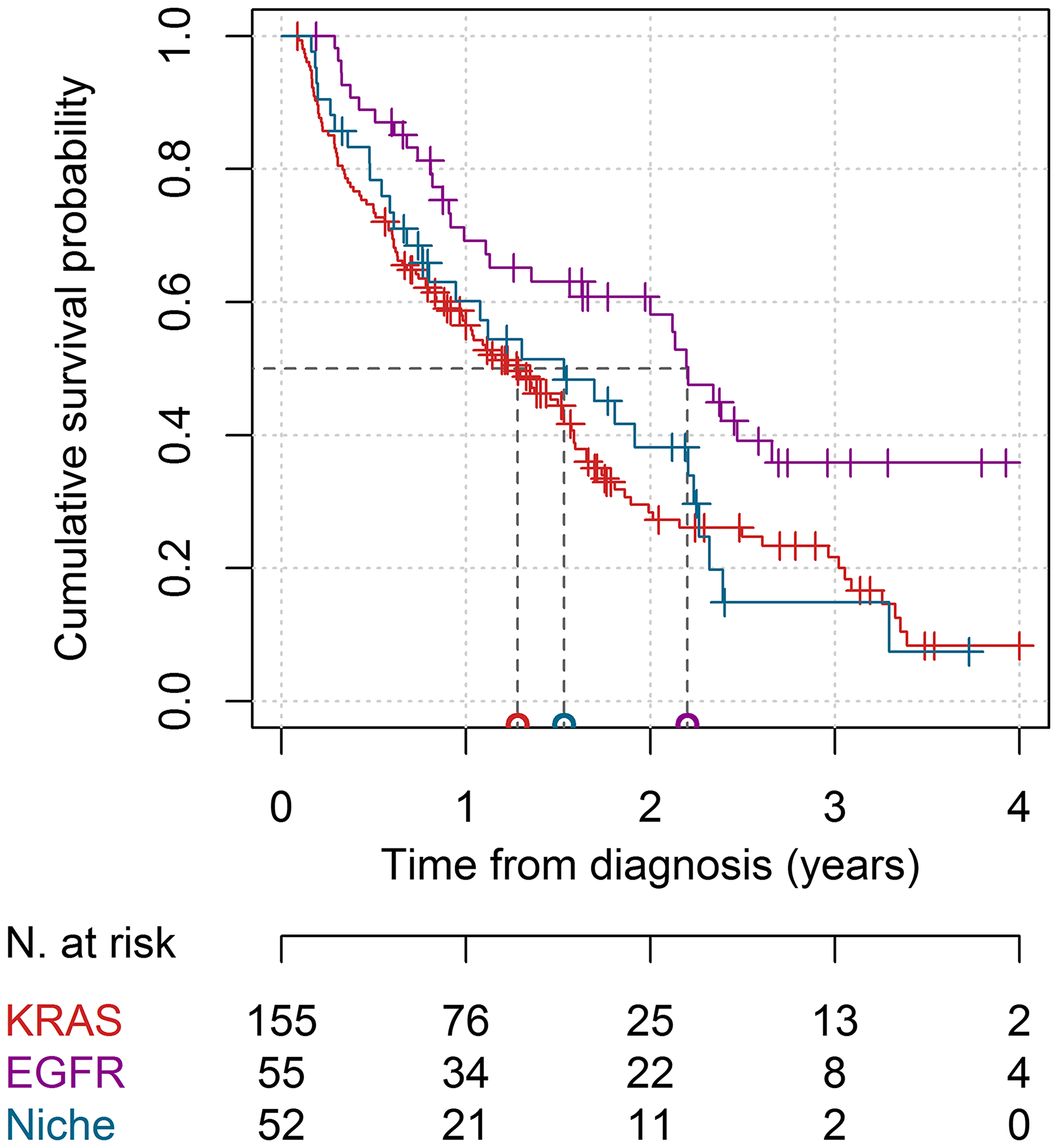
Figure 2: Survival curves in the KRAS, EGFR, and niche mutations cohorts. The red line represents the KRAS cohort, the purple line represents the EGFR cohort and the blue line represents the niche mutations cohort. The median survival times for the three cohorts are reported on the horizontal axis. The number of patients at risk at 0, 1, 2, 3, and 4 years from baseline is reported. Thick marks represent subjects lost to follow-up.
The differences in OS observed between the three cohorts of interest are reported in Figure 3 from the unadjusted and multivariable regression analyses. According to the multivariable analysis that adjusted for age, sex, smoking habits, stage, and treatments, we observed that 1) patients with niche mutations had a higher risk of death than patients with EGFR gene mutations (HR = 2.3; 95% CI = 1.2–4.4; p = 0.009); 2) patients with KRAS gene mutations had a higher risk of death than patients with EGFR gene mutations (HR = 2.5; 95% CI = 1.4–4.5; p = 0.003); 3) patients with niche mutations presented a risk of death similar to patients with KRAS gene mutations (HR = 0.9; 95% CI = 0.6–1.5; p = 0.797). These results did not change when patients with concomitant mutations were excluded or when patients with stage I or II diagnoses were excluded (Figure 3). The associations between all the recorded independent variables and the risk of death, as assessed by the Cox multivariable model, are reported in Supplementary Table 2. The proportional hazards assumption was verified for all HRs of interest (data not shown).
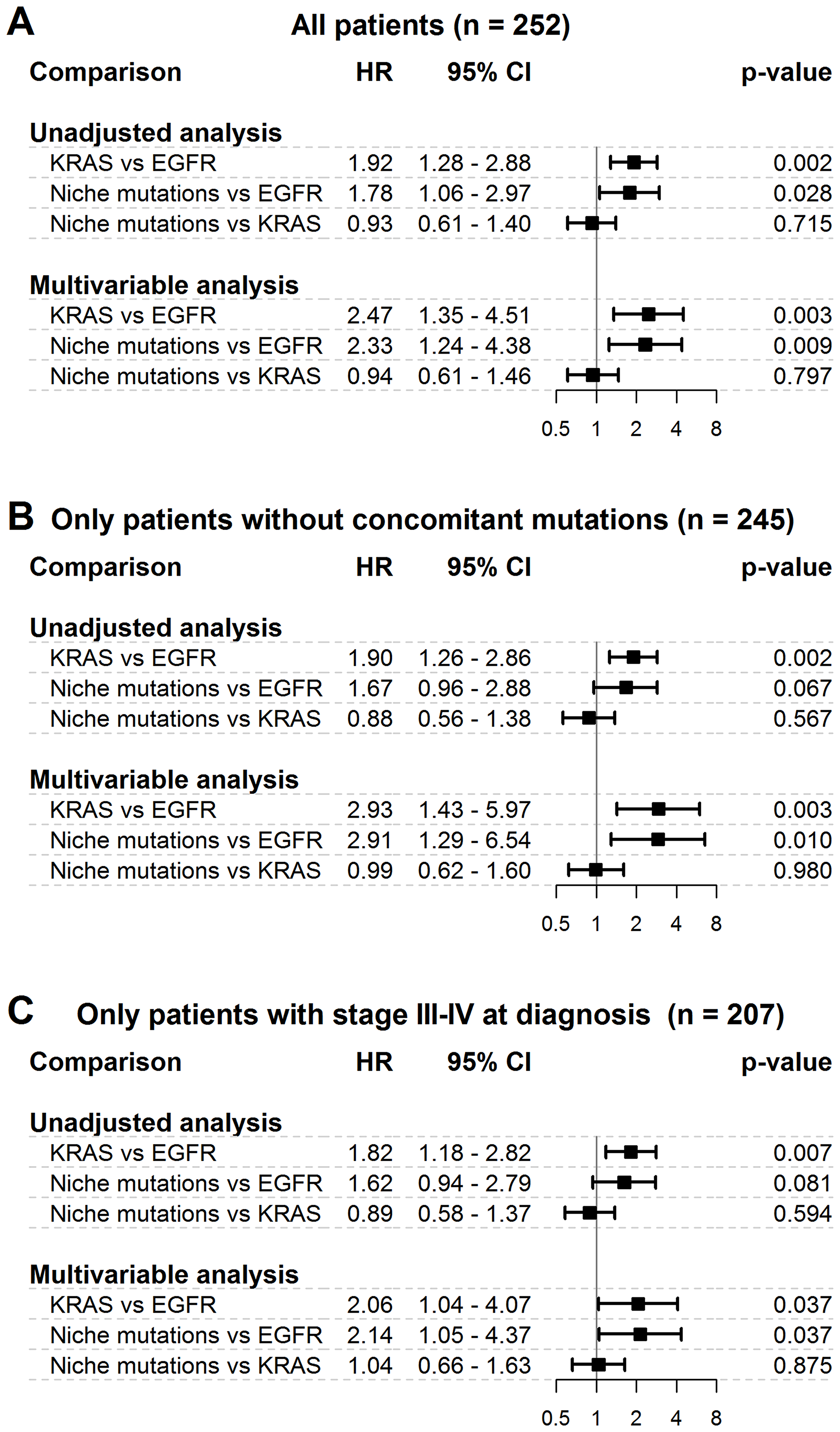
Figure 3: Differences in overall survival among niche mutations cohort and frequent mutations cohorts. HR = hazard ratio; 95% CI = 95% confidence interval. (A) = main analysis - all patients. (B) = secondary analysis – only patients without concomitant mutations. (C) = secondary analysis – only patients with stage III–IV at diagnosis. Results from both unadjusted and multivariable Cox regression analysis are reported. The multivariable model was adjusted for age, sex, smoking habits, stage and treatments. The plots report hazard ratios and 95% confidence intervals.
DISCUSSION
Our study reports a retrospective analysis on OS rates observed in patients with lung adenocarcinoma harboring somatic mutations either in EGFR and KRAS genes, or in less frequently studied genes, such as BRAF, PIK3CA, NRAS, and HER2 (niche mutations). In our work, we analyzed and described the clinical characteristics of the three cohorts of patients. From our data, male subjects and smokers were more common in the KRAS group than in the niche mutations and EGFR groups. Furthermore, target therapies were more common in the EGFR group, whereas chemotherapy was more common in the KRAS and niche mutations cohorts. This is justified by the fact that no specific target treatment has yet been developed for KRAS and niche mutations.
Regarding treatments given after diagnosis, we observed that our cohorts of patients were treated as previously described for general mutations [15, 16]. This is likely because niche mutations are poorly understood, especially in terms of biological characteristics and interactions. However, we believe that the percentage of patients harboring niche mutations suggests further study, in particular with respect to treatment strategies.
In our study, we underlined the importance of niche mutations in terms of OS relative to the most frequent general somatic mutations and for KRAS in particular, which is considered to be the most frequent and aggressive mutation in lung adenocarcinoma. EGFR and KRAS gene mutations are already recorded in the molecular report that an oncologist needs to provide “personalized” therapy [17–20]. In addition to the mutations for which target therapies have been generally successful, for example, for the EGFR gene, there are still mutations without effective therapies, as for the KRAS and niche mutations. The scientific community has agreed on the fact that KRAS is one of the most aggressive mutations in terms of prognosis and survival because it exhibits a high mortality rate and there is no existing targeted treatment available [17, 18]. Our data described a similar OS for KRAS and niche mutations. This aspect is very interesting and needs to be further considered, although little has been published on niche mutations in lung cancer, probably because they are only observed in a small proportion of the population.
We first described a large cohort study analyzing niche mutations in adenocarcinoma of the lung and their prognostic role as compared with the more frequently diagnosed mutations. In the current scientific literature, many studies have described the occurrence of general somatic mutations in NSCLC, although some niche mutations, such as c-MET and RET in adenocarcinoma of the lung have been described in a few short reports [19–22], or in a larger population focusing on a single-niche mutation of PIK3CA [23–25]. We took into consideration different rare mutations observed in BRAF, PIK3CA, HER2, and NRAS detected with a total frequency equal to 16.7% of patients with lung adenocarcinoma harboring somatic mutations, which is not a negligible prevalence if transposed to the entire population. Moreover, niche mutations presented a high risk of death, accounting for 55% of deaths in one person-year.
The most interesting and novel results that came out of this research were that 1) niche mutations showed very similar rates of survival as those of patients with KRAS gene mutation; 2) patients harboring niche or KRAS gene mutations showed a higher risk of mortality than those with EGFR gene mutations.
We know that mutations in the KRAS gene are the most aggressive, and for this reason, we highlighted the importance of evaluating these niche mutations through our results. The life expectancy of these patients was low and seemed to be very similar to that of patients with well-established aggressive general mutations. These patients may benefit from the use of specific inhibitors such as dabrafenib/trametinib, which are used for BRAF v600E mutations [26].
Limitations
Finally, the present study has some limitations that should be discussed. First, the observational and retrospective nature of this study presents a risk of bias; however, adjustments made to the multivariable analysis protected against confounding bias due to the observed patient characteristics. Nevertheless, other unobserved factors may have biased our results. Second, even if we observed statistically significant differences in the OS between niche mutations and general mutations in a large set of consecutive patients enrolled over 5 years, another limitation relates to the low absolute number of patients with niche mutations that were present. Because of this, it was not possible properly to assess differences in OS between the single-niche mutations. Moreover, because c-MET, DDR2, MAP2K1, and RET gene mutations were not observed in this study, we cannot draw conclusions about the OS of patients harboring these niche mutations.
Taken together, these limitations highlight the need for an extended study to confirm our findings and to reduce further the uncertainty regarding the prognostic role of niche mutations.
Materials and Methods
This retrospective cohort study was carried out in accordance with the STrengthening the Reporting of OBservational studies in Epidemiology (STROBE) statement [13].
Study population
All patients with lung adenocarcinoma identified at the Modena University Hospital between January 1, 2013 and December 31, 2017, and for whom a molecular analysis of gene mutations was available were eligible for inclusion. Inclusion criteria were histologically diagnosed primary or metastatic adenocarcinoma of the lung, age at primary diagnosis between 18 and 90 years, presence of at least one of BRAF, c-MET, DDR2, EGFR, HER2, KRAS, MAP2K1, NRAS, PIK3CA, and RET gene mutations. Exclusion criteria were patients with wild-type genes, patients with an unknown tumor, squamous cell carcinoma of the lung, large-cell carcinoma of the lung, small-cell lung cancer, and neuroendocrine tumors of the lung.
All patients received standard treatments according to the current guidelines based on clinical stage and other relevant clinical factors.
Data collection
Data were retrospectively collected based on electronic hospital patient records. The following information was gathered: date of primary diagnosis; age at diagnosis (years); sex (M, F); clinical stage at diagnosis (TNM 8th Edition) [14]; smoking status at diagnosis (yes, no); treatments performed after diagnosis (surgery, chemotherapy, radiotherapy, immunotherapy, target therapy); first-line treatment; presence of BRAF, c-MET, DDR2, EGFR, HER2, KRAS, MAP2K1, NRAS, PIK3CA, or RET gene mutations and related codons; date and cause of death; date of last follow-up.
The somatic mutations that were considered were those detectable by our genotyping platform.
Molecular analysis
Molecular analysis of gene mutations was performed for patients with advanced lung cancer or for early-stage surgical patients after recurrence.
DNA was extracted from 10 μm thick representative sections cut from formalin-fixed and paraffin-embedded blocks of each tumor sample containing at least 50% tumor cells. Extraction was performed with the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany), and the DNA was quantified with Xpose NGS (Trinean NV, Gentbrugge, Belgium). Mutations were detected using the high-throughput genotyping platform Sequenom MassARRAY® System (Sequenom, San Diego, CA, USA) and the Myriapod® Lung Status kit (Diatech Pharmacogenetics, Italy) following the manufacturers’ protocols. The molecular array allowed for the identification of the mutations of the BRAF, c-MET, DDR2, EGFR, HER2, KRAS, MAP2K1, NRAS, PIK3CA, and RET genes. In brief, 25 ng DNA of each specific primitive tumor was amplified through a multiplex polymerase chain reaction and then incorporated nucleotides were inactivated using shrimp alkaline phosphatase. A single-base extension reaction was performed using extension primers that hybridized immediately adjacent to the mutations and a custom mixture of nucleotides. Salts were removed using a cation exchange resin. Multiplexed reactions were spotted in SpectroCHIP II arrays, and DNA fragments were resolved with matrix-assisted laser desorption/ionization–time-of-flight (MALDI–TOF) using a compact mass spectrometer (Sequenom, San Diego, CA, USA) with a detection limit of 5%. The data were evaluated using MassARRAY Typer analyzer software 4.0, which allowed the identification of mutated alleles by comparing the ratio of the wild-type peak of all suspected mutants.
Statistical analysis
Descriptive characteristics of the patients at the time of diagnosis were recorded. Continuous variables were recorded as mean ± standard deviation (SD) values, and categorical variables were recorded as absolute and percentage frequencies. Comparisons of group characteristics were drawn using analysis of variance and chi-square tests. OS was defined as the time span from diagnosis to death (all-cause mortality was considered). The OS of patients who were still alive at the end of the follow-up period were treated as censored survival times. The main aim of the analysis was to compare the OS of three cohorts of patients harboring: 1) niche mutations (BRAF, c-MET, DDR2, HER2, MAP2K1, NRAS, PIK3CA, or RET); 2) KRAS mutations; 3) EGFR mutations. The mortality rates (MRs) were calculated as the number of deaths per person-year. The OS curves were calculated using the Kaplan–Meier method. The pairwise differences in OS between the three cohorts of interest were assessed by means of a Cox proportional hazards regression model. The Cox model results were reported as hazard ratios (HRs) with 95% confidence interval (95% CI) from unadjusted analysis (i. e., with the cohort as the only independent variable) and from a multivariable analysis that adjusted for the confounding effects attributable to age (years); sex (M vs F); smoking habits (yes vs no); clinical stage (III, IV vs I–II) at diagnosis; and treatments after diagnosis (surgery, chemotherapy, radiotherapy, immunotherapy, and target therapy – all yes vs no). The proportional hazards assumption was validated by a graphical assessment of scaled Schoenfeld residuals. All statistical analyses were performed with R 3.4.3 statistical software (The R Foundation for Statistical Computing, Wien) at the p < 0.05 significance level.
Conclusions
We found evidence of a higher risk of death for patients harboring niche mutations relative to patients with EGFR gene mutations, whereas the risk for patients with niche mutations was found to be comparable to that of patients harboring KRAS gene mutations. We may summarize this message in two points: 1. As soon as the molecular biologist is able to identify a niche mutation, it is necessary to highlight this in the patient report to set a targeted treatment that offers a better solution than chemotherapy for the patient; 2. The NGS technique, which is able to select a myriad of genes, is more useful for the oncologist who is conscious of the importance of considering niche mutations.
The correct selection of mutations will be helpful in terms of the greater efficacy of treatment in association with better prognosis and a higher quality of life for oncologic patients.
Abbreviations
NSCLC: non-small cell lung cancer; EGFR: epidermal growth factor receptor; BRAF: B-Raf kinase family; KRAS: K-Raf kinase family; c-MET: tyrosine-protein kinase Met; PIK3CA: phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha; NRAS: neuroblastoma RAS viral oncogene homolog; MAP2K1: Mitogen-Activated Protein Kinase Kinase; HER2: human epidermal growth factor receptor 2; DDR2: discoidin domain receptor tyrosine kinase 2; RET: REarranged during Transfection; OS: overall survival; MRs: Mortality rates; NGS: the next-generation sequencing; STROBE: STrenghening the Reporting of Observational studies in Epidemiology.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
FUNDING
The Project has been supported in part by funds from the Division of Thoracic Surgery of the University Hospital of Modena and from the Laboratories of the Institute of Pathology of the University of Modena and Reggio Emilia, Modena, Italy.
References
1. de Groot PM, Wu CC, Carter BW, Munden RF. The epidemiology of lung cancer. Transl Lung Cancer Res. 2018; 7:220–33. https://doi.org/10.21037/tlcr.2018.05.06. [PubMed].
2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018; 68:7–30. https://doi.org/10.3322/caac.21442. [PubMed].
3. Zito Marino F, Bianco R, Accardo M, Ronchi A, Cozzolino I, Morgillo F, Rossi G, Franco R. Molecular heterogeneity in lung cancer: from mechanisms of origin to clinical implications. Int J Med Sci. 2019; 16:981–89. https://doi.org/10.7150/ijms.34739. [PubMed].
4. Davidson MR, Gazdar AF, Clarke BE. The pivotal role of pathology in the management of lung cancer. J Thorac Dis. 2013 (Suppl 5); 5:S463–78. https://doi.org/10.3978/j.issn.2072-1439.2013.08.43. [PubMed].
5. Raquel-Cunha A, Cardoso-Carneiro D, Reis RM, Martinho O. Current Status of Raf Kinase Inhibitor Protein (RKIP) in Lung Cancer: Behind RTK Signaling. Cells. 2019; 8. https://doi.org/10.3390/cells8050442. [PubMed].
6. Carper MB, Claudio PP. Clinical potential of gene mutations in lung cancer. Clin Transl Med. 2015; 4:33. https://doi.org/10.1186/s40169-015-0074-1. [PubMed].
7. Rothschild SI. Targeted Therapies in Non-Small Cell Lung Cancer-Beyond EGFR and ALK. Cancers (Basel). 2015; 7:930–49. https://doi.org/10.3390/cancers7020816. [PubMed].
8. Sholl LM, Aisner DL, Varella-Garcia M, Berry LD, Dias-Santagata D, Wistuba II, Chen H, Fujimoto J, Kugler K, Franklin WA, Iafrate AJ, Ladanyi M, Kris MG, et al, and LCMC Investigators. Multi-institutional Oncogenic Driver Mutation Analysis in Lung Adenocarcinoma: The Lung Cancer Mutation Consortium Experience. J Thorac Oncol. 2015; 10:768–77. https://doi.org/10.1097/JTO.0000000000000516. [PubMed].
9. Wang R, Hu H, Pan Y, Li Y, Ye T, Li C, Luo X, Wang L, Li H, Zhang Y, Li F, Lu Y, Lu Q, et al. RET fusions define a unique molecular and clinicopathologic subtype of non-small-cell lung cancer. J Clin Oncol. 2012; 30:4352–59. https://doi.org/10.1200/JCO.2012.44.1477. [PubMed].
10. Fathi Z, Mousavi SA, Roudi R, Ghazi F. Distribution of KRAS, DDR2, and TP53 gene mutations in lung cancer: an analysis of Iranian patients. PLoS One. 2018; 13:e0200633. https://doi.org/10.1371/journal.pone.0200633. [PubMed].
11. Linee guida AIOM. Neoplasie del polmone. Available online: https://www.aiom.it/wp-content/uploads/2019/10/2019_LG_AIOM_Polmone-1.pdf (Last accessed on October 2019).
12. Coco S, Truini A, Vanni I, Dal Bello MG, Alama A, Rijavec E, Genova C, Barletta G, Sini C, Burrafato G, Biello F, Boccardo F, Grossi F. Next generation sequencing in non-small cell lung cancer: new avenues toward the personalized medicine. Curr Drug Targets. 2015; 16:47–59. https://doi.org/10.2174/1389450116666141210094640. [PubMed].
13. von Elm E, Altman DG, Egger M, Pocock SJ, Gøtzsche PC, Vandenbroucke JP; STROBE Initiative. The Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement: guidelines for reporting observational studies. J Clin Epidemiol. 2008; 61:344–9. https://doi.org/10.1016/j.jclinepi.2007.11.008. [PubMed].
14. Detterbeck FC, Boffa DJ, Kim AW, Tanoue LT. The Eighth Edition Lung Cancer Stage Classification. Chest. 2017; 151:193–203. https://doi.org/10.1016/j.chest.2016.10.010. [PubMed].
15. Schirrmacher V. From chemotherapy to biological therapy: A review of novel concepts to reduce the side effects of systemic cancer treatment (Review). Int J Oncol. 2019; 54:407–19. https://doi.org/10.3892/ijo.2018.4661. [PubMed].
16. Masters GA, Temin S, Azzoli CG, Giaccone G, Baker S Jr, Brahmer JR, Ellis PM, Gajra A, Rackear N, Schiller JH, Smith TJ, Strawn JR, Trent D, Johnson DH, and American Society of Clinical Oncology Clinical Practice. Systemic Therapy for Stage IV Non-Small-Cell Lung Cancer: American Society of Clinical Oncology Clinical Practice Guideline Update. J Clin Oncol. 2015; 33:3488–515. https://doi.org/10.1200/JCO.2015.62.1342. [PubMed].
17. Nussinov R, Jang H, Tsai CJ, Cheng F. Review: Precision medicine and driver mutations: Computational methods, functional assays and conformational principles for interpreting cancer drivers. PLoS Comput Biol. 2019; 15:e1006658. https://doi.org/10.1371/journal.pcbi.1006658. [PubMed].
18. Loomans-Kropp HA, Umar A. Cancer prevention and screening: the next step in the era of precision medicine. NPJ Precis Oncol. 2019; 3:3. https://doi.org/10.1038/s41698-018-0075-9. [PubMed].
19. Ahmadzada T, Kao S, Reid G, Boyer M, Mahar A, Cooper WA. An Update on Predictive Biomarkers for Treatment Selection in Non-Small Cell Lung Cancer. J Clin Med. 2018; 7. https://doi.org/10.3390/jcm7060153. [PubMed].
20. Sokolenko AP, Imyanitov EN. Molecular Diagnostics in Clinical Oncology. Front Mol Biosci. 2018; 5:76. https://doi.org/10.3389/fmolb.2018.00076. [PubMed].
21. Gautschi O. SC11.04 Rare Mutations in Lung Cancer. J Thorac Oncol. 2017; 12:S104. https://doi.org/10.1016/j.jtho.2016.11.089.
22. Ferreira CG, Zalis M, Reis M, Schluckebier L, Montella T. PUB070 Rare Actionable Mutations in a Lung Adenocarcinoma Cohort in Brazil. J Thoracis Oncology. 12:S2388–89. https://doi.org/10.1016/j.jtho.2017.09.1933.
23. Song Z, Yu X, Zhang Y. Mutation and prognostic analyses of PIK3CA in patients with completely resected lung adenocarcinoma. Cancer Med. 2016; 5:2694–700. https://doi.org/10.1002/cam4.852. [PubMed].
24. Chaft JE, Arcila ME, Paik PK, Lau C, Riely GJ, Pietanza MC, Zakowski MF, Rusch V, Sima CS, Ladanyi M, Kris MG. Coexistence of PIK3CA and other oncogene mutations in lung adenocarcinoma-rationale for comprehensive mutation profiling. Mol Cancer Ther. 2012; 11:485–91. https://doi.org/10.1158/1535-7163.MCT-11-0692. [PubMed].
25. Wu SG, Chang YL, Yu CJ, Yang PC, Shih JY. The Role of PIK3CA Mutations among Lung Adenocarcinoma Patients with Primary and Acquired Resistance to EGFR Tyrosine Kinase Inhibition. Sci Rep. 2016; 6:35249. https://doi.org/10.1038/srep35249. [PubMed].
26. Baik CS, Myall NJ, Wakelee HA. Targeting BRAF-Mutant Non-Small Cell Lung Cancer: From Molecular Profiling to Rationally Designed Therapy. Oncologist. 2017; 22:786–96. https://doi.org/10.1634/theoncologist.2016-0458. [PubMed].


