Introduction
Non-invasive circulating tumor DNA (ctDNA) genotyping is a cost-effective alternative to tumor biopsies when these biopsies are associated with significant risk, tumor tissue is insufficient or inaccessible, and/or serial assessment of tumor molecular abnormalities is needed to optimize treatment. ctDNA analysis of epidermal growth factor receptor (EGFR) in non-small cell lung cancer (NSCLC) and v-Ki-ras2 kirsten rat sarcoma viral oncogene homolog (KRAS) in colorectal cancer (CRC) is well established [1, 2]. However, validation studies of the clinical relevance of ctDNA in other tumor types [3] are lacking.
In this systematic review, we summarize the published trials of ctDNA analysis by tumor type and across tumor types, and we discuss the role of ctDNA analysis in selecting patients for enrollment in clinical trials and in guiding targeted therapy. The potential use of plasma genotyping in cancer is illustrated in Figure 1.
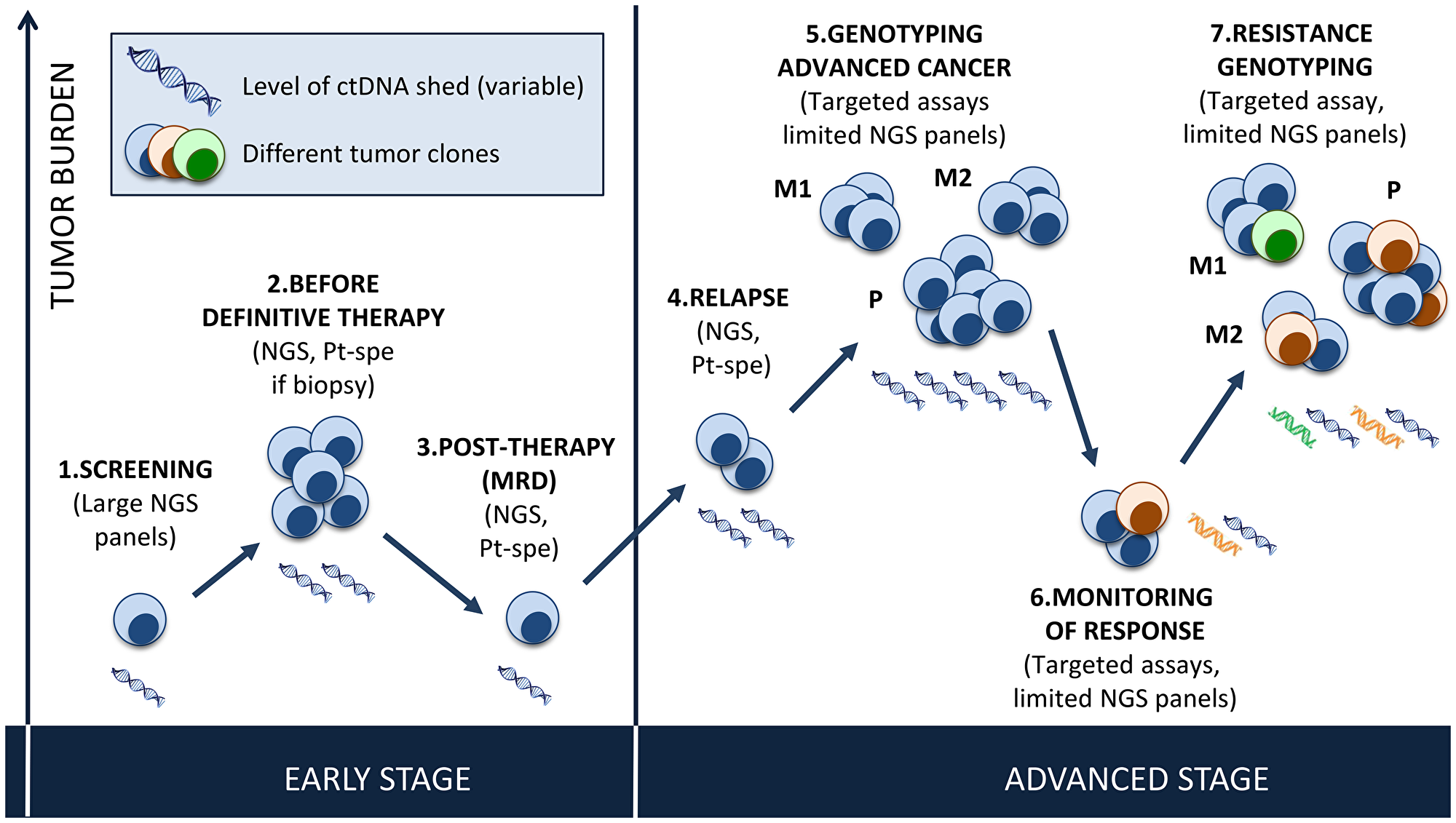
Figure 1: Potential use of plasma genotyping in cancer. Early stage disease: Screening will require the use of large NGS panels, with both high sensitivity and perfect specificity. Before surgery, determination of tumor burden in plasma has the potential to help guide neo-adjuvant or adjuvant therapy and monitor response, using large panels or patient-specific assays based on the molecular profile of the tissue biopsy when available. After surgery, NGS (large gene panels or patient-specific assays) can detect MRD and guide adjuvant therapy (early detection) or detect relapse. Low tumor shed in plasma will be the main limitation to the integration of plasma genotyping in early stage disease. Advanced stage disease: At diagnosis, ctDNA can guide genotype-directed therapy (using targeted assays focusing on a predefined gene of interest (i. e. EGFR in NSCLC) or targeted NGS covering genes of interest). The variations in allelic fractions allow for monitoring of treatment response, which may be helpful for pharmacodynamics analyses in phase I studies. When acquired resistance to targeted therapies occurs, ctDNA can detect specific mechanisms of resistance (targeted assay like for EGFR T790M or targeted NGS), taking into consideration the different clones present within the primary tumor (P) and all metastatic sites (M1, M2), and guide treatment adjustments.
Non-small cell lung cancer
The increasing number of “targetable” genotypes in NSCLC and understanding of tumor resistance to targeted therapies has led to rapid, non-invasive, longitudinal assays to repeatedly assess tumor biology throughout treatment.
ctDNA for NSCLC genotyping in advanced-stage NSCLC
The combination of more targetable genotypes and minimally invasive diagnostic tools (e. g. endobronchial ultrasound) that result in small specimens [4–6] has led to the development of alternative, noninvasive testing methods, such as the U. S. Food and Drug Administration (FDA)-approved targeted ctDNA assay (Cobas) for EGFR genotyping or the CLIA (Clinical Laboratory Improvement Amendments)-certified plasma droplet digital polymerase chain reaction (ddPCR) assay. ddPCR is a highly sensitive (EGFR exon 19 deletion, 82%; EGFR L858R mutation, 74%) and quantitative approach that allows for the longitudinal monitoring of treatment response [7, 8]. Although the specificity of these PCR-based platforms allows for the initiation of EGFR-targeted therapy on the basis of positive plasma testing, negative results must be confirmed by tumor tissue genotyping [9]. While most clinically validated assays are focused on a single predefined gene, next-generation sequencing (NGS) of ctDNA can broadly interrogate the tumor molecular profile across a range of genes and variant types. Hybrid capture-based NGS platforms have already been evaluated in NSCLC [10–12]. Overall, 75% of patients with NSCLC harbor potentially actionable genomic aberrations in ctDNA, although concordance with tissue is suboptimal (specificity, 63.5%) [11, 13–15].
Tumor NGS can help monitor tumor dynamics and detect acquired ALK resistance mutations in crizotinib-resistant patients [14]. Our group has studied various NGS methods and found favorable diagnostic accuracy using a bias-corrected targeted ctDNA NGS (2/3 ALK; 2/3 RET; 2/2 ROS1) [16] or using amplicon-based sequencing (6/7 ALK; 2/2 ROS1) [17]. A larger prospective study is needed to determine the most reliable method for identifying targetable fusions in ctDNA.
At disease progression, ctDNA constitutes a promising alternative to tissue biopsies and is a well-established approach in the EGFR setting. Plasma genotyping is widely used as a screening test for detection of the EGFR T790M resistance mutation, with tumor biopsy needed only if the result is negative [1, 17, 18]. It remains unknown, however, if treatment should be adjusted on the basis of isolated plasma variations. Ongoing trials, such as the (APPLE)-EORTC study [19], will help determine the value of ctDNA analysis in treatment selection.
Clinical trials that validated the use of plasma NGS to guide therapy have demonstrated encouraging results [20–22]. In 323 patients with NSCLC, the addition of ctDNA analysis to tissue NGS analysis increased the identification of driver alterations and resulted in an 85.7% rate of objective response or stable disease [20].
Screening and minimal residual disease in early-stage NSCLC
The National Lung Screening Trial [23] and the Dutch-Belgian Randomized Lung Cancer Screening Trial (NELSON) [24] demonstrated that low-dose computed tomography (CT) screening reduces the mortality rate in lung cancer. Benign lung nodules (false positives) generate invasive procedures. Deep ctDNA sequencing is a more specific and potentially complementary approach to low-dose CT screening in lung cancer but is limited by the low or absent DNA shed of early-stage tumors [25, 26]. Combining ctDNA with other circulating biomarkers such as transcriptomics (ctRNA) could improve sensitivity, while white blood cell sequencing to eliminate “false-positive” variants linked to clonal hematopoiesis may increase specificity [27–29].
ctDNA analysis is also used to detect minimal residual disease (MRD) with plasma genotyping. Tumor molecular profiles from previous biopsies are used to build personalized PCR-based assays with improved sensitivity and specificity. In the TRACERx study, ctDNA from 100 patients was analyzed at the time of diagnosis and followed after definitive treatment. Patient-specific multiplex PCR assays (threshold: ≥2 variants for positivity) demonstrated high sensitivity (93%) and specificity for MRD detection [26, 30].
In another study, ctDNA was analyzed using cancer personalized profiling by deep sequencing (CAPP-seq) in 255 samples from 40 patients with stage I–III lung cancer (stage II–III, n = 33) treated with curative intent and 54 healthy adults. It found that in 94% of patients with stage I–III lung cancer with disease recurrence, ctDNA was detectable in the first post-treatment blood sample and preceded radiographic progression by 5.2 months (median), indicating that detection of post-treatment ctDNA should prompt treatment tailored to the patient’s ctDNA analysis to prevent disease progression [31].
Gastrointestinal tract cancers
Esophageal carcinoma
ctDNA analysis of patients with squamous cell carcinoma (SCC) of the esophagus with the use of a 90-gene panel was associated with sensitivity of 94% and 75%, respectively, when detecting ≥1 or ≥2 mutant genes, suggesting that ctDNA analysis can help monitor treatment effect in these patients [32]. In multivariate analysis, a higher plasma cyclin D1 (CCND1; 11q13) to dopamine receptor D2 (DRD2; 11q22-23) ratio (C/D ratio) was significantly correlated with worse prognosis [33]. In another study, high concordance of multiple somatic mutations was found between plasma ctDNA and tumor (primary or metastases) DNA (83%–100%). The allele frequencies of the mutations increased with tumor burden and preceded radiologic evidence of tumor recurrence by 6 months [34].
In 29 patients with localized esophageal carcinoma treated with chemoradiotherapy, baseline ctDNA levels (NGS-based CAPP-seq) were correlated with metabolic tumor volume and squamous histology. The 2-year overall survival (OS) rates for pretreatment ctDNA-positive vs. ctDNA-negative patients were 47% and 86%, respectively (HR = 6.0; p < 0.05). Compared to patients with undetectable ctDNA, detection of ctDNA post-treatment was predictive of shorter event-free survival (EFS, p < 0.0001) and shorter time to distant metastasis (p < 0.0001) [35].
Gastric cancer
ctDNA analysis is used to detect and monitor HER2 copy numbers in patients with gastric cancer [36]. The preoperative ratio of HER2 to RPPH1 (ddPCR) has been correlated with tumor HER2 status (p < 0.001; sensitivity, 0.73; specificity, 0.93) [37].
In another study, the concordance rate of HER2 amplification detected between formalin-fixed, paraffin-embedded (FFPE) samples vs. ddPCR and immunohistochemical (IHC) analysis/fluorescence in situ hybridization (FISH) was 92%. The concordance rate of FFPE with ctDNA was 62.5%. HER2 positivity by ctDNA analysis was associated with significantly shorter OS compared to HER2 negativity [38]. In late-stage gastric cancer, ctDNA mutations were associated with poor prognosis [39].
In a multiple-parallel cohort, ctDNA-guided plasma-based digital sequencing in patients with metastatic solid tumors, including gastric cancer, identified somatic alterations in 78% of the 76 patients with gastric cancer, and 33% of these 76 patients had targetable alterations. Ten patients with gastric cancer received molecularly matched therapy, which resulted in response and disease control rates of 67% and 100%, respectively [40].
Colorectal cancer
In a prospective study of 106 patient samples of CRC, ctDNA analysis of KRAS and BRAF mutational status was compared to the analysis of tumor tissue [2]. The specificity and sensitivity of ctDNA were both 100% for BRAF V600E mutation and were 98% and 92%, respectively, for KRAS point mutations. Overall, ctDNA was detected in 100% of patients with metastatic CRC, suggesting that ctDNA analysis could replace tumor analysis in these patients [2].
ctDNA has been used to detect MRD in the adjuvant setting in stage II CRC [41, 42]. In a prospective study of 230 patients, [43] the recurrence rate among patients who did not receive adjuvant chemotherapy was 79% (11/14) in patients with detectable ctDNA vs. 9.8% (16/164) in patients with undetectable ctDNA (HR = 18; p < 0.001). ctDNA detection after completion of chemotherapy was associated with shorter recurrence-free survival (HR = 11; p = 0.001) suggesting the presence of MRD [43].
The same investigators analyzed ctDNA in patients with stage III CRC post-operatively, during, and after adjuvant chemotherapy [44]. In 95 patients who received adjuvant chemotherapy, serial ctDNA analysis identified residual metastatic disease not evident on imaging studies [44].
In another longitudinal cohort study, ctDNA was used to monitor tumor burden in 21 CRC [45] patients with localized disease who had ctDNA analysis 3 months post-operatively; disease recurrence within 3 years occurred in all 6 patients with detectable ctDNA at 3 months compared to 27% (4/15) of those with undetectable ctDNA at 3 months (HR = 37.7; p < 0.001). Of 18 patients who had surgical resection of liver metastases, ctDNA detection 3 months post-surgically predicted disease recurrence compared to patients with undetectable ctDNA and metastatic CRC (HR = 4.9; p = 0.007). Therefore, postoperative ctDNA detection identified patients at high risk of relapse [45].
Pancreatic and biliary tract carcinomas
The role of ctDNA is particularly important in patients with pancreatic and biliary tract carcinomas because biopsy samples are often inadequate for molecular profiling. In a prospective analysis in 26 patients with pancreatic (n = 18) or biliary (n = 8) cancer, tumor sequencing using a 54-gene panel failed in 9 patients (35%), and of the remaining 17 patients, 90.3% of the mutations detected in the tumor biopsies were also detected using ctDNA. The diagnostic accuracy of ctDNA sequencing was 97.7% (5 informative genes: sensitivity, 92.3%; specificity, 100%) [46].
In pancreatic cancer, high ctDNA levels of KRAS and/or other mutations have been associated with poor progression-free survival (PFS) and/or OS [47, 48]. However, low concordance between the blood and tissue samples has been reported in patients with pancreatic ductal adenocarcinoma [49].
In a prospective study in advanced pancreatic cancer, ctDNA was analyzed for KRAS mutations using blood samples collected prior to gemcitabine or FOLFIRINOX (5-fluorouracil, oxaliplatin, irinotecan, and leucovorin) treatment and monthly during treatment (median follow-up, 3.7 months). Disease progression was more frequent in patients with ctDNA-identified KRAS mutations at baseline compared to KRAS-mutant-negative patients (9/10 vs. 1/4; p = 0.01). The baseline ctDNA level was associated with PFS (p = 0.014) and OS (p = 0.01). ctDNA level changes were correlated with imaging studies and CA19-9 levels, indicating that ctDNA may be used to monitor disease activity in these patients [47]. In other studies, ctDNA analysis detected mutations in up to 48% of patients, and these mutations were associated with shorter OS. In another study, ctDNA analysis (ddPCR for rare KRAS mutations) detected mutations in 31% of 105 patients with pancreatic ductal adenocarcinoma who underwent pancreatoduodenectomy. Detectable ctDNA KRAS mutations were associated with shorter OS compared to patients without mutations (13.6 months vs. 27.6 months; p < 0.0001) [48]. Other investigators using ctDNA analysis detected KRAS among other mutations (TP53, SMAD4, STK11, PIK3CA, NRAS) using ctDNA in 48% of patients with advanced pancreatic adenocarcinoma. These mutations were strongly correlated with shorter OS compared to OS of patients with undetectable mutations (6.5 vs. 19.0 months; p < 0.001) [50].
ctDNA analysis is a promising prognostic marker in early-stage pancreatic cancer that could also help guide treatment after Whipple surgery [51]. ctDNA analysis of pre- and post-operative plasma samples and tumor tissue (n = 42) using PCR-based SafeSeqS assays to identify KRAS mutation (codons 12, 13, and 61) identified RAS mutations in 90.5% of tumor samples and in 62.2% of 37 pre-operative and 37.1% of 35 post-operative plasma samples. ctDNA detection was an independent factor predicting shorter recurrence-free survival compared to undetectable ctDNA preoperatively (10.3 months vs. not reached, HR = 3.4; p = 0.005) and postoperatively (5.4 months vs. 17.1 months, HR = 5.4; p < 0.0001) [51].
In ctDNA promoter hypermethylation analysis of a selected gene panel, the number of methylated genes was significantly higher in patients with pancreatic adenocarcinoma than in healthy individuals or patients with acute pancreatitis (mean, 8.41 vs. 4.74, respectively; p < 0.001), suggesting that ctDNA promoter hypermethylation may be used for diagnostic purposes in pancreatic adenocarcinoma [52].
ctDNA analysis can also inform the diagnosis, prognosis, and treatment of patients with biliary tract cancer. In patients with metastatic locally advanced/metastatic biliary tract cancer, the concordance rate between plasma ctDNA analysis and tumor tissue analysis (15-gene panel) was 74%; this rate was 92% in intrahepatic tumors. ctDNA variant allele frequency (VAF) was significantly correlated with tumor load and PFS, and the mutational profile changed after chemotherapy in 36% of patients [53].
Hepatocellular carcinoma
ctDNA analysis has been investigated in hepatocellular carcinoma (HCC) [54–56]. In patients who underwent hepatectomy or liver transplantation, disease recurrence and extrahepatic metastases were more frequent in those with mutations detected by ctDNA analysis vs. others (p = 0.01 and 0.04, respectively) and ctDNA was an independent factor predicting invasion of the portal vein (odds ratio = 6.10) [54]. In other studies, hotspot mutations in the TERT, CTNNB1, and TP53 genes detected in the plasma of patients with HCC were associated with vascular invasion and likely predicted a shorter recurrence-free survival time [55].
Breast cancer
The concordance between ctDNA and tissue DNA in breast cancer varies by stage and subtype [57]. In patients with early-stage breast cancer, PIK3CA mutations have been identified presurgically in ctDNA with high sensitivity (93.3%) and specificity (100%) [58]. In a retrospective analysis, the concordance between tissue DNA and ctDNA (digital sequencing of plasma-derived DNA) was robust in PIK3CA mutation and ERBB2 amplification analyses (Cohen’s κ = 0.64 and 0.77, respectively) but poor in TP53 mutation and EGFR amplification analyses (Cohen’s κ = 0.18 and 0.33, respectively) [59, 60]. In another study, TP53 and PIK3CA mutant allele frequencies were associated with response to therapy and PFS [60]. When PCR and targeted exome sequencing were used to detect the hotspot AKT1 E17K mutation in two cohorts of patients with advanced metastatic breast cancer (MBC), the concordance rates between tissue and blood samples were 98% and 97.1% [61].
ctDNA analysis has been used extensively in large trials in hormone-receptor–positive, HER2-negative advanced breast cancer. In the BOLERO-2 study, PIK3CA mutations by ctDNA analysis were associated with efficacy of everolimus [62, 63]. In the BELLE-3 study, patients progressing on or after mTOR inhibition and endocrine therapy [64] with PIK3CA mutations (detected by ctDNA or tissue DNA analysis; concordance, 80%) had significantly longer PFS in the buparlisib plus fulvestrant arm compared to the fulvestrant arm [64, 65].
In patients with HER2-positive breast cancer treated with an anti-HER1/HER2 tyrosine kinase inhibitor, ctDNA analysis for HER2 amplification was associated with disease progression (4/6, 66.7%), whereas TP53 mutations (3/6, 50%) and PI3K/mTOR pathway alterations (3/6, 50.0%) were associated with disease resistance. Dynamic ctDNA analysis identified drug resistance with sensitivity 85.7% and specificity of 55.0%; and had a high concordance rate with CT imaging studies (82.1%) [66].
In patients with non-metastatic triple-negative breast cancer, ddPCR for customized ctDNA analysis detected alterations in 75% of patients at baseline. Lower ctDNA levels were associated with longer OS during neoadjuvant chemotherapy [67]. In this setting, targeted MRD sequencing using serial ctDNA monitoring predicted tumor recurrence 7.9 months (median) before clinical evidence of relapse [68].
Other investigators found that a panel of cell-free methylation markers was a strong predictor of OS in MBC and may have clinical usefulness in risk disease monitoring [69]. Methylated ctDNA from the promoter region of RASSF1A was more sensitive than carcinoembryonic antigen and CA15-3 for monitoring response to neoadjuvant chemotherapy [70]. ctDNA analysis of a 6-gene methylation panel for diagnosis of breast cancer had sensitivity of 79.6% compared to healthy individuals and 82.4% compared to benign disease control (specificity, 72.4% and 78.1%, respectively). This test complemented mammography or ultrasonography [71].
ctDNA for assessment of resistance to aromatase inhibitors as first-line therapy was also prospectively studied in MBC (n = 83) [72]. ESR1 mutations were detected in 56.4% (22/39) of patients who had disease progression 6.7 months (median) prior to clinical progression [72]. On the basis of detection of ESR1 mutations in primary breast cancer using ctDNA analysis at a very low allele frequency, in contrast to a high allele frequency in metastases, it is plausible that in some tumors rare ESR1-mutant clones may be enriched by endocrine therapy [73].
Gynecologic cancers
ctDNA analysis has contributed to diagnosis and monitoring of patients with gynecologic cancers. High ctDNA levels have been associated with poor PFS and OS in patients with resistant epithelial ovarian cancer treated with bevacizumab (n = 144) [74]. In patients with high-grade serous ovarian carcinoma who received standard-of-care therapy, p53 alterations identified using ctDNA analysis at baseline were correlated with volume of disease, and a decrease in TP53 mutant allele fraction ≤60% after 1 cycle of chemotherapy was associated with shorter time to disease progression [75].
In patients with gynecologic cancer (n = 44), ctDNA analysis using ddPCR detected alterations in 93.8% of patients, and detected cancer in 6 patients, 7 months prior to radiologic evidence on CT imaging studies [76]. ctDNA levels were correlated with serum CA-125 levels and CT imaging studies, and they were an independent factor predicting OS [76].
Plasma DNA analysis for detection of chromosomal instability using copy-number alterations in patients with an adnexal mass (n = 68) and in healthy individuals (n = 44) improved detection of malignancy (AUC = 0.89) compared to serum CA-125 (AUC = 0.78) or the RMI (risk of malignancy) index (composite of serum CA 125 level, ultrasound scan result and menopausal status) (AUC = 0.81) [77].
Data suggest that human papillomavirus (HPV) detection by ctDNA analysis is a surrogate marker for HPV-associated cervical cancer and can guide antiviral therapy. In patients with HPV 16/18-associated cervical cancer and tumors >2 cm at diagnosis, HPV ctDNA was identified in 11 of 13 patients and levels were associated with tumor dynamics [78]. In another study, HPV ctDNA was detected in all patients (n = 19) with HPV-positive metastatic cervical cancer but in 0 healthy blood donors (n = 45) [79]. Of 9 patients who received tumor-infiltrating lymphocyte (TIL) immunotherapy, the HPV genotype of the patients’ tumors was identified in serum samples from all patients, and 2 patients with complete response had persistent clearance of HPV ctDNA [79].
Genitourinary cancers
Prostate cancer
In patients with metastatic castration-resistant prostate cancer (mCRPC), ctDNA analysis can identify molecular alterations that are associated with clinical outcomes and can guide therapy. Detection of biallelic BRCA2 gene loss by ctDNA analysis indicated that patients may benefit from therapies targeting defective DNA repair [80]. Others identified alterations in all analyzed patients with mCRPC treated with enzalutamide, including alterations in DNA damage repair and PI3K pathway genes [81]. Aberrations in the androgen receptor (AR) gene by ctDNA analysis have been correlated with resistance to enzalutamide and abiraterone treatment and AR amplification was more common in patients whose disease progressed on enzalutamide compared to abiraterone or other agents (p = 0.02) [82]. Targeted NGS covering all AR coding bases using plasma from patients treated with abiraterone (control, patients’ normal circulating DNA) identified AR copy numbers in 82.5% (80/97) of patients and demonstrated that 45% of tumors had AR gain or T878A or L702H changes before abiraterone treatment, which were associated with shorter PFS and OS [83]. Other investigators also demonstrated that circulating AR copy number gain is a useful biomarker [84]. In patients with CRPC treated with docetaxel followed by enzalutamide (n = 59), patients with AR copy number gain (36%) had higher levels of PSA, alkaline phosphatase, and lactate dehydrogenase (LDH) and shorter PFS (p = 0.0004) and OS (p = 0.0003) compared to those without AR copy number gain. In multivariate analysis, a decrease in PSA ≥50% and AR copy number gain were associated with longer PFS and OS [84].
In a retrospective study, ctDNA genomic profiling using Guardant 360™ demonstrated ≥1 alteration in 94% of 514 men with progressive mCRPC, and had good concordance with the correspondent tumor tissue [85]. Higher numbers of ctDNA alterations were associated with shorter time to treatment failure (HR = 1.05, p = 0.026) in patients treated with chemotherapy or androgen inhibitors [85].
Renal cell carcinoma
Total serum ctDNA levels and CpG island methylation of RASSF1A and VHL were shown to support the diagnosis of renal cell carcinoma (RCC) and VHL methylation indicates clear cell RCC [86]. Higher levels of ctDNA were identified in metastatic RCC or necrotic RCC compared to benign tumors, and they were associated with poorer DFS [86]. Using Guardant 360 ctDNA analysis, 78.6% of 220 patients with RCC had >1 alteration and the most frequent alterations were TP53 (35%), VHL (23%), EGFR (17%), NF1 (16%), and ARID1A (12%) [87]. Higher rates of detection after systemic therapy compared with baseline were noted for NF1 (21% vs. 3%), TP53 (49% vs. 24%), and VHL (29% vs. 18%), indicating clonal evolution of genomic alterations [87].
Bladder cancer
In patients with advanced urothelial carcinoma, high rates (86%–90%) of aberrations using Guardant 360 ctDNA analysis were found; the most common aberrations were TP53 (48%), ARID1A (17%), and PIK3CA (14%) [88, 89]. Using a 62-gene panel (FoundationACT), 73% of patients with metastatic urothelial cancer were found to have ≥1 aberration and the most frequent alterations were TP53 (68%) and TERT-promoter (38%) [90]. In another study, 36% (129/363) of patients with non-muscle-invasive bladder cancer and 11% (44/403) of patients with muscle-invasive bladder cancer who underwent radical cystectomy had ≥1 FGFR3 or PIK3CA mutations and high ctDNA levels were associated with disease progression. ctDNA levels in the urine and plasma were positively correlated and indicated that higher levels of FGFR3- and PIK3CA-mutated DNA can predict disease progression [91].
Melanoma
In patients with metastatic melanoma, identification of BRAF mutations in ctDNA has been associated with higher disease burden and worse prognosis and may precede clinical evidence of disease progression. In patients with unresectable advanced-stage metastatic melanoma, BRAF and NRAS mutations identified in ctDNA analysis at baseline and during treatment with targeted therapy against BRAF or immunotherapy have been associated with larger tumors, increased LDH levels, and brain metastases [92]. Other investigators found that in this setting, the presence of BRAF mutations in ctDNA analysis was associated with a higher number of metastatic sites, higher serum LDH levels or S100 protein concentration, and shorter OS [93].
In one study, 73% of 48 patients with metastatic melanoma had tumor-associated BRAF and NRAS alterations in ctDNA analysis [94]. In another study, patients with the BRAF V600E mutation in ctDNA had shorter PFS and OS compared to those without the mutation (p = 0.02 and p = 0.017, respectively) [95]. Others also demonstrated that lower baseline ctDNA levels were associated with higher rates of response and PFS [94]. In patients with BRAF V600E/V600K-positive tumors enrolled in 4 different studies of dabrafenib or trametinib, ctDNA analysis identified BRAF V600E and BRAF V600K mutations in 76% and 81% of 732 patients, respectively. Patients with undetectable ctDNA BRAF mutations at baseline had higher rates of response, PFS, and OS than those with ctDNA BRAF mutations [96]. Circulating BRAF mutations have been identified in some patients prior to clinical evidence of disease progression [97]. In patients with melanoma who received adoptive transfer of activated autologous TILs, ctDNA analysis for BRAF V600E (n = 48, 388 serum samples) demonstrated a strong correlation between an early peak of circulating V600E mutation and objective response. Patients whose serum had an early ctDNA peak followed by undetectable ctDNA had a higher likelihood of having a complete response in 1–2 years [97]. Others also found a correlation between serial ctDNA analysis of BRAF and NRAS status and tumor response. PFS was longer in patients with an early decrease (1–4 weeks post-treatment) in ctDNA levels than in patients with unchanged or increased ctDNA levels post-treatment (HR = 2.6; p = 0.05) [98]. In the post-surgical setting, ctDNA analysis of BRAF and NRAS mutations predicted OS in 161 patients with high-risk stage II/III melanoma who underwent surgical resection followed by adjuvant bevacizumab for 1 year [99]. The 5-year OS rates were 33% and 65% for patients with detectable and undetectable ctDNA levels, respectively. After adjustment for performance status, patients with detectable ctDNA had shorter OS compared to those with undetectable ctDNA (HR = 2.63; p = 0.003) [99].
An intriguing application of longitudinal ctDNA analysis is the distinction of pseudoprogression from true progression [100]. ctDNA for BRAF and NRAS mutations was analyzed at baseline and at 12 weeks of treatment with PD-1 antibodies with or without ipilimumab [100]. Overall, 23.2% (29/125) of patients had initial disease progression by imaging studies. Thirty-one percent (9/29) of patients had pseudoprogression and 69% (20/29) had true progression. The 9 patients with confirmed pseudoprogression had undetectable ctDNA at baseline or detectable ctDNA at baseline followed by >10-fold decrease (favorable ctDNA profile). Eighteen of 20 patients with true progression had detectable ctDNA at baseline that remained stable or increased (unfavorable ctDNA profile). Among patients with confirmed true progression, the 1-year OS rates were 82% and 39% in those with favorable ctDNA and unfavorable ctDNA profiles, respectively (HR = 4.8; p = 0.02) [100].
Sarcoma
Ewing sarcoma
In patients with localized or metastatic Ewing sarcoma (EWS), copy numbers of the EWSR1 fusion sequence in plasma were associated with tumor volume. Rapid decrease in ctDNA levels of EWSR1 was noted during initial chemotherapy, and increase in ctDNA levels indicated disease recurrence [101]. As a driving EWS-ETS translocation specific to each tumor is identified in up to 95% of patients with Ewing sarcoma, investigators used long-range PCR analysis to identify tumor-specific EWS-ETS breakpoints in plasma DNA [102]. In children with metastatic Ewing sarcoma and primary localized osteosarcoma, detection of ctDNA was associated with inferior outcomes [103]. In patients with localized EWS and detectable ctDNA, the 3-year rates of EFS and OS were lower compared to those with undetectable ctDNA (p = 0.006 and p = .01, respectively). The respective rates in localized osteosarcoma for EFS were 48.6% vs. 82.1% (p = 0.006) and for OS were 79.8% vs. 92.6% (p = 0.01); the risk of death increased proportionately with ctDNA levels [103].
Gastrointestinal stromal tumors
In patients with gastrointestinal stromal tumors (GIST), ctDNA harboring CKIT or PDGFRA was used as a tumor-specific biomarker and the amount of mutant-free circulating DNA was correlated with disease course [104]. In patients with TKI-refractory GIST treated with dovitinib, genotyping of the KIT gene in exon 17 of serum ctDNA using beads, emulsions, amplification, and magnetics assays identified mutations associated with disease resistance [105]. Other investigators suggested that detection of secondary C-KIT mutations in ctDNA may improve the selection of targeted agents [106].
Soft tissue sarcoma
In patients with metastatic soft tissue sarcoma, ctDNA was detected in 36% (4/11) of patients and TP53/PIK3CA mutations in ctDNA analysis were concordant with the primary tumor in 2 of 4 patients [107].
Brain tumors
In brain tumors, ctDNA analysis is used as a non-invasive alternative to tumor biopsies that are associated with significant risk. In patients with glioblastoma multiforme (GBM), ctDNA analysis identified the EGFRvIII deletion in 3 of 13 patients, which was correlated with tumor tissue analysis [108]. This mutation is identified in approximately one third of patients with GBM and is associated with resistance to chemotherapy and radiotherapy. ctDNA levels were correlated with the extent of tumor resection. This test may help select patients for anti-EGFRvIII therapy and monitor response to treatment [108]. In patients with neuroblastoma, serum MYCN amplification (real-time quantitative PCR, sensitivity 86%, specificity 95% compared with tissue analysis) was associated with OS, suggesting that it may help select treatment prior to tumor biopsy, particularly for patients younger than 18 months whose risk assessment and treatment depend on MYCN amplification status [109]. In patients with glioma, the IDH1 R132H mutation was identified in plasma with a specificity of 100% and sensitivity related to the tumor volume and contrast enhancement, suggesting that it may help in the diagnosis of patients not amenable to biopsy [110].
Lymphoma
Classical Hodgkin lymphoma
In patients with classical Hodgkin lymphoma, an XPO1 mutation detected using ctDNA analysis at the end of treatment was associated with a tendency toward shorter PFS compared to patients without the mutation, suggesting that plasma ctDNA may be clinically useful for the noninvasive management of this disease [111].
Diffuse large B-cell lymphoma
In patients with relapsed, refractory de novo, or transformed diffuse large B-cell lymphoma (DLBCL) treated with panobinostat with or without rituximab, ctDNA was detected in ≥1 plasma sample in 96% of the patients and its increase was associated with resistance to treatment [112]. In another study of patients with lymphoma and healthy subjects, the amount of ctDNA at diagnosis was strongly correlated with clinical indices and was independently predictive of patient outcomes [113]. ctDNA genotyping could help distinguish indolent follicular lymphomas from those that transformed into DLBCL and classify transcriptionally defined tumor subtypes, including DLBCL cell of origin [113].
Other investigators used immunoglobulin high-throughput sequencing (Ig-HTS) [114] to analyze circulating leukocytes and ctDNA of patients with DLBCL. At baseline, Ig clonal rearrangement was detected in 82% and 71% of patients in ctDNA and circulating cells, respectively (p = 0.68); at relapse, the respective rates were 100% and 30% (p = 0.001). Interestingly, Ig-HTS detection preceded radiologic evidence of recurrent disease by 88 days with high sensitivity and specificity, indicating that it may be a surrogate marker for monitoring disease after complete remission is achieved [114].
In a retrospective study in patients with DLBCL after first-line treatment, disease progression was evident on imaging studies a median of 3.5 months (range, 0-200 months) after detection on ctDNA analysis of the clonal Ig gene sequence [115].
In contrast to these data showing that ctDNA is associated with prognosis and can be identified prior to radiologic evidence of recurrent DLBCL, in patients with newly diagnosed primary central nervous system lymphoma who had 34-gene panel high-throughput sequencing of primary tumors followed by targeted sequencing of identified somatic mutations on plasma, the correlation between ctDNA concentration and tumor volume was insignificant (R2 coefficient = 0.01) [116]. Overall, 88% of patients with MYD88 L265P in tissue had an identifiable L265P variant in their ctDNA and OS was not significantly correlated with mutations detected in ctDNA [116].
Clinical trials across tumor types
In patients with advanced cancer, circulating nucleic acid biomarker analyses had promising clinically important multipurpose utility awaiting further studies [117]. In an analysis of 105 patients using the Sequenom MassArray System and OncoCarta panel for somatic mutations, the ctDNA concentration was 3 times higher in patients with advanced cancer compared to healthy volunteers. Although the concordance between matched ctDNA and archival tumor tissue was high for selected ‘hot-spot’ mutations (KRAS, BRAF, PIK3CA), some differences were noted between archival tumor and ctDNA. Factors predicting longer OS in multivariate analysis were lower ctDNA concentration, higher albumin levels, and better performance status [117].
We have previously published our experience analyzing ctDNA and archival primary or metastatic tumor tissue (FFPE) from patients with advanced metastatic cancer who were referred to our Phase I program at MD Anderson for participation in clinical trials [118]. We found that the concordance rates between mutations in archival tissue and ctDNA were high in patients with refractory cancer types that progressed on systemic therapy. These rates were 91%, 99%, 83%, and 91% for BRAF, EGFR, KRAS, and PIK3CA mutations, respectively [118]. Patients with >1% of KRAS mutant ctDNA had shorter OS compared to those with ≤1% of KRAS mutant ctDNA (4.8 vs. 7.3 months, p = 0.008). Patients with >1% of mutant ctDNA (BRAF, EGFR, KRAS, or PIK3CA) had shorter OS compared to those with ≤1% of mutant ctDNA (5.5 vs. 9.8 months, p = 0.001) [118].
Other investigators demonstrated that ctDNA and circulating tumor cells are distinct biomarkers, as ctDNA was also detected in patients without any evident circulating tumor cells [119]. ctDNA analysis demonstrated high detection rates (>75%) in advanced breast, bladder, colorectal, gastroesophageal, hepatocellular, head and neck, melanoma, ovarian, and pancreatic cancer. Low detection rates (<50%) were noted in primary brain, renal, prostate, and thyroid cancers. In localized tumor stage, the rate of ctDNA detection was 73% in CRC, 57% in gastroesophageal, and 50% in breast adenocarcinoma. In metastatic CRC (n = 206), ctDNA analysis for KRAS mutation was associated with high sensitivity (87.2%) and specificity (99.2%). ctDNA analysis to assess resistance mechanisms to anti-EGFR treatment in patients with CRC demonstrated that 96% (23/24) of patients developed ≥1 mutation in genes involved in the mitogen-activated protein kinase pathway [119].
In a study of clonal evolution, exome sequencing of ctDNA using serial plasma from 6 patients with breast, ovarian, or lung cancer demonstrated that emergence of resistance was associated with increased mutant allele fractions, and an activating PIK3CA mutation was noted after treatment with paclitaxel [120]. In another study with ctDNA analysis of 23 patients with various tumor types treated with PI3K-AKT-mTOR pathway or MEK inhibitors, clonal response to treatment was noted and some clones changed over time discordantly. Increasing mutational levels were associated with poorer prognosis [121].
Other investigators used NGS to analyze 54 genes and copy number variants in three genes (EGFR, ERBB2 and MET) on ctDNA of patients with various tumor types [122]. Overall, 171 patients with lung (n = 39), breast (n = 39), glioblastoma (n = 33), or other cancers were analyzed. Actionable mutations were noted in 40% (TP53 29.8%, EGFR 17.5%, MET 10.5%, PIK3CA 7%, and NOTCH1 5.8%). Eighteen percent (6/33) of patients with glioblastoma had actionable mutations [122]. The same investigators analyzed 168 patients with diverse cancers [123]. In ctDNA analysis, 58% of patients had ≥1 alteration. Among them, 71.4% had ≥1 alteration potentially targeted by an FDA-approved drug. The concordance rates between ctDNA analysis and tissue analysis were: TP53 70.3%, EGFR 70.3%, PIK3CA 88.1%, and ERBB2, 93.1%. In patients with ≥1 alteration, those with ctDNA ≥5% of total DNA had shorter OS compared to patients with ctDNA <5% (median, 4.03 months vs. NR, p < .001) [123]. With the use of checkpoint inhibitor–based immunotherapy across various histologies, correlation between high alteration number detected in blood-derived ctDNA and favorable response, PFS and OS has been reported [124].
In a meta-analysis of patients with solid tumors (39 studies, 4,052 patients), detection of ctDNA in plasma was associated with shorter OS in multivariable analyses [HR, 2.70; P < .001) compared to patients with undetectable ctDNA [125].
Selected studies by tumor type, the gene (s) used for ctDNA analysis, and outcomes are summarized in Table 1.
Table 1: Selected studies by tumor type, the gene(s) used for ctDNA analysis, and outcomes
| Year, First Author [Ref] | Study Type | Tumor Type | Number of Patients | Biomarker Method | Tested Genes | Outcome |
|---|---|---|---|---|---|---|
| Lung cancer | ||||||
| 2016, Adrian G. Sacher [8] | Prospective | Advanced NSCLC | 180 | ddPCR | KRAS, EGFR | Detection of KRAS and EGFR mutations, lower turnaround time compared to tissue |
| 2016, Jeffrey C. Thompson [10] | Cohort | Advanced NSCLC | 102 | Hybrid capture NGS | 70 cancer-related genes | Detection of potentially actionable variants in 75% of patients; concordance with tissue, 79% |
| 2018, Nicolas Guibert [13] | Cohort, blinded to tissue | Advanced NSCLC | 168 specimens from 46 patients | Amplicon-based NGS | 36 cancer-related genes | Detection of EGFR mutations, rare variants and fusions with high specificity. Early detection of resistance mechanisms in serial samples. |
| 2018, Charu Aggarwal [20] | Prospective | Advanced NSCLC | 323 | Hybrid capture | 73 cancer-related genes | Detection of actionable alterations in 20% of stage IV M1b patients in plasma but not tissue. Complementarity of tissue and plasma |
| 2014, Aaron M. Newman [25] | Cohort | Early-stage lung cancer | 103 | Hybrid capture | 16 cancer-related genes + 8 proteins | Detection of ctDNA in early stages (stage I sensitivity, 50%) |
| 2017, Christopher Abbosh [26] | Cohorts | Early-stage lung cancer | 96 | Patient-specific multiplex PCR | 10–22 SNVs | MRD and subclonal evolution |
| Gastro-intestinal tumors | ||||||
| 2016, Honglei Luo [32] | Cohort | Esophageal, SCC | 8 | Illumina TruSight sequencing | 90 cancer-related genes | Multigene panel has a role in detection and monitoring response to treatment |
| 2010, Hiroki Takeshita [33] | Case-control | Esophageal, SCC | 96 patients, 40 controls | PCR-applied biosystems | CCND1 amplification | Poor prognostic value of CCND1 amplification |
| 2016, Masami Ueda [34] | Cohort | Esophageal, SCC | 13 | HiSeq2000 | 53 cancer-related genes | Multigene panel is associated with a greater accuracy of tumor recurrence compared to imaging methods (post-op) |
| 2015, Katsutoshi Shoda [36] | Case-control | Gastric | 52 patients, 40 controls | PCR-applied biosystems | HER-2 | HER2 amplification can be used for therapeutic monitoring |
| 2017, Katsutoshi Shoda [37] | Case-control | Gastric | 60 patients, 30 controls | PCR-applied biosystems | HER-2 | HER2 amplification can be used for therapeutic monitoring |
| 2015, Hideaki Kinugasa [38] | Cohort | Gastric | 25 | PCR - QX200, Bio-Rad | HER-2 | High concordance in detection of HER-2 between ddPCR and tissue IHC/FISH |
| 2016, Wen-Liang Fang [39] | Cohort | Gastric | 277 | TaqMan qPCR | 68 mutations (8 genes) | High ctDNA levels are associated with peritoneal recurrence and poor prognosis |
| 2017, Jeanne Tie [43] | Prospective, cohort, multicenter | Colon | 230 (1046 plasma samples) | Safe-SeqS PCR | 15 cancer-related genes | ctDNA detection after stage II colon cancer resection provides direct evidence of residual disease and identifies patients at very high risk of recurrence. |
| 2018, Jeanne Tie [44] | Prospective, cohort, multicenter | Colon | 95 | Safe-SeqS PCR | 15 cancer-related genes | ctDNA detection after adjuvant chemotherapy for stage III colon cancer resection can identify patients at very high risk of recurrence. |
| 2017, Lone V. Schøler [45] | Longitudinal cohort | Colon | 45 (371 plasma samples) | QX200 PCR | Somatic structural variants, KRAS | Postoperative ctDNA analysis detects residual disease and identifies patients at very high risk of relapse. Longitudinal surveillance allows early detection of relapse and response to intervention. |
| 2015, Oliver A. Zill [46] | Cohort | Pancreato-biliary carcinomas | 26 | Illumina Hi-Seq 2500 | 54 cancer-related genes | ctDNA sequencing is feasible, accurate, and sensitive in identifying tumor-derived mutations. |
| 2016, Kjersti Tjensvoll [47] | Cohort | Pancreatic | 14 (53 plasma samples) | Mx3000P rtPCR | KRAS | ctDNA can be used as a marker for monitoring treatment efficacy and disease progression. |
| 2016, Naoto Hadano [48] | Cohort | Pancreatic | 105 | TaqMan assay PCR | KRAS | ctDNA can predict poor survival |
| 2017, Daniel Pietrasz [50] | Prospective, cohort | Pancreatic | 135 | Ion AmpliSeq NGS | 112 cancer-related genes | ctDNA is an independent prognostic marker in advanced pancreatic adenocarcinoma |
| 2018, Belinda Lee [51] | Cohort | Pancreatic | 42 | SafeSeqS assays PCR | KRAS | ctDNA analysis is a promising prognostic marker in early-stage pancreatic cancer and guides risk-adaptive treatment strategies. ctDNA detection post-operatively helps to identify disease progression after standard adjuvant chemotherapy. |
| 2016, Stine Dam Henriksen [52] | Prospective case-control | Pancreatic | 95 patients, 183 controls | Methylation-specific PCR | 28 cancer-related genes | ctDNA promoter hypermethylation is a diagnostic biomarker that helps distinguish malignant from benign pancreatic disease. |
| 2018, Andreas Wolfgang Berger [53] | Cohort | Biliary tract cancer | 24 | 1010× depth Sequencing | 15 cancer-related genes | The molecular landscape is represented in ctDNA. |
| 2015, Atsushi Ono [54] | Cohort | Hepatocellular carcinoma | 46 | Illumina Hi-Seq 2500 | ctDNA detection post-surgery reflects tumor progression and disease recurrence. | |
| 2016, Wenjun Liao [55] | Cohort | Hepatocellular carcinoma | 41 | Illumina MiSeq™ | Cancer-related genes TERT, TP53, and CTNNB1 | ctDNA mutation detection is associated with vascular invasion and predicts a shorter recurrence-free survival time. |
| 2016, Ao Huang [56] | Cohort | Hepatocellular carcinoma | 48 | QX200 PCR | Cancer-related genes TERT, TP53, and CTNNB1 | ctDNA analysis can detect intratumoral heterogeneity and may have a promising role in the therapeutic management. |
| Breast cancer | ||||||
| 2017, Heather A. Parsons [57] | Prospective cohort | Triple-negative breast cancer | 26 | HiSeq 2500 Illumina | 33 cancer-related genes | High concordance between ctDNA analysis and tumor tissue analysis, allowing monitoring of the therapeutic effect. |
| 2014, Julia A. Beaver [58] | Prospective cohort | Breast cancer | 29 | ddPCR | PIK3CA mutations | In patients with early-stage breast cancer, mutations can be detected in tumor tissue using ddPCR, and ctDNA can be detected in blood before and after surgery. |
| 2016, Diana H. Liang [60] | Retrospective chart review | Breast cancer | 100 | Illumina Hi-Seq 2500 | TP53, PIK3CA, ERBB2, and EGFR genomic alterations | Robust concordance between tissue and blood for detection of PIK3CA mutation and ERBB2 amplification, but not for TP53 mutation and EGFR amplification. Directional changes of TP53 and PIK3CA mutant allele are associated with response to therapy and PFS. |
| 2016, Marion Rudolph [61] | Cohort | Breast cancer | 600 | HiSeq 2500 Illumina | 306 cancer-related genes | AKT1E17K is the most likely disease driver in selected breast cancer patients and its detection in blood is achievable in advanced-stage disease. |
| 2017, Mary Ellen Moynahan [62] | Prospective | Breast cancer | 724 | QX200 PCR | PIK3CA | Improvement in PFS was maintained using everolimus, irrespective of PIK3CA genotypes (detected by ctDNA), and it was consistent with previous analysis of archival tumor DNA using NGS. |
| 2016, Sarat Chandarlapaty [63] | Prospective | Breast cancer | 541 | QX200 PCR | ESR1 | ESR1 mutations are prevalent in ER-positive metastatic breast cancer treated with aromatase inhibitors. Both Y537S and D538G mutations are associated with aggressive disease biology. |
| 2018, Rosaria Condorelli [65] | Cohort | Breast cancer | 3 | QX200 PCR | RB1 | Somatic RB1 mutations can emerge after exposure to CDK4/6 inhibitors. |
| 2016, Fei Ma [66] | Prospective, cohort | Breast cancer | 18 | HiSeq 2500 Illumina | 368 cancer-related genes | ctDNA analysis provides information regarding resistance to treatment and guides administration of anti-HER2 targeted therapy in the metastatic setting. |
| 2017, Francesca Riva [67] | Cohort | Triple-negative breast cancer | 46 | Illumina MiSeq | TP53 | ctDNA levels decreased quickly during neoadjuvant chemotherapy (NCT) and helped identify minimal residual disease after surgery. Slow decrease of ctDNA levels during NCT was strongly associated with shorter survival. |
| 2015, Isaac Garcia-Murillas [68] | Prospective, cohort | Breast cancer | 55 | HiSeq 2500 Illumina | Cancer-related genes | In patients with early stage breast cancer, ctDNA analysis can identify patients at high risk for relapse and guide adjuvant therapy. |
| 2017, Kala Visvanathan [69] | Cohort | Breast cancer | 141 | cMethDNA assay | 10 cancer-related genes | ctDNA gene methylation is a strong predictor of survival outcomes. |
| 2017, Hiroyo Takahashi [70] | Cohort | Breast cancer | 87 | Methylation-specific PCR | RASSF1A | Met-ctDNA is a more sensitive marker than CEA and CA15-3 and it can be used to monitor clinical tumor response to neoadjuvant chemotherapy. |
| 2016, Ming Shan [71] | Cohort | Breast cancer | 749 | MethyLight | SFN, P16, hMLH1, HOXD13, PCDHGB7 and RASSF1a | Epigenetic markers in serum have potential for diagnosis of breast cancer. |
| 2018, Charlotte Fribbens [72] | Prospective, cohort | Breast cancer | 83 | Enhanced tagged-amplicon sequencing (eTAm-Seq) | ESR1, KRAS, NRAS and HRAS | In patients with progressive disease after first-line aromatase inhibitors, ctDNA analysis demonstrated high levels of genetic heterogeneity and frequent sub-clonal mutations. Sub-clonal KRAS mutations were found at a high frequency. |
| 2016, Peilu Wang [73] | Cohort | Breast cancer | 126 | Bio-Rad QX100 dd PCR | ESR1 | ESR1 mutations were detected at very low allele frequencies in some primary breast cancers, and at high allele frequency in patients with metastatic breast cancer. ESR1-mutant clones are enriched by endocrine therapy. |
| Gynecological cancers | ||||||
| 2014, Karina Dahl Steffensen [74] | Cohort | Ovarian cancer | 144 | QiaSymphony, multiplex qPCR | ctDNA detection | In patients treated with bevacizumab, high ctDNA levels were associated with poor PFS and OS. |
| 2016, Christine A. Parkinson [75] | Retrospective analysis | Ovarian cancer | 40 | ddPCR | TP53 | ctDNA is correlated with volume of disease at the start of treatment. |
| 2015, Elena Pereira [76] | Cohort | Ovarian/ endometrial cancer | 44 | qPCR using TaqMan®, ddPCR | ctDNA detection | ctDNA is an independent predictor of survival in patients with ovarian and endometrial cancers. |
| 2017, Adriaan Vanderstichele [77] | Prospective, cohort | Ovarian cancer | 68 | HiSeq 2500 Illumina | Chromosome instability | ctDNA analysis demonstrated that chromosomal instability can help detect ovarian cancer. |
| 2012, Maura Campitelli [78] | Cohort | Cervical cancer | 16 | RT-qPCR | ctDNA detection | ctDNA analysis demonstrated that the HPV mutational insertion is a highly specific molecular marker and it is detected in most patients with stage 2-4 cervical cancer. |
| 2017, Zhigang Kang [79] | Retrospective analysis | Cervical cancer | 19 | ddPCR | HPV genetic components | HPV genetic insertion in ctDNA represents a promising tumor marker. |
| Urological cancers | ||||||
| 2017, Matti Annala [80] | Prospective, cohort | Prostate cancer | 319 | NimbleGenSeqCap, Illumina | 73 cancer-related genes | Biallelic gene loss detected in ctDNA can help prioritize therapy. |
| 2016, Alexander W. Wyatt [81] | Cohort | Prostate cancer | 65 | Illumina MiSeq, Ion Ampliseq | 19 cancer-related genes | Genomic profiling of ctDNA is feasible in mCRPC patients and provides important insights into enzalutamide response and resistance. |
| 2015, Arun A. Azad [82] | Cohort | Prostate cancer | 62 | PCR-based BEAMing | AR | AR gene aberrations in ctDNA are associated with resistance to enzalutamide and abiraterone in mCRPC. |
| 2015, Alessandro Romanel [83] | Cohort | Prostate cancer | 97 | Ion Torrent Sequencing | AR | Plasma AR sequencing can identify primary resistance to abiraterone. |
| 2016, Samanta Salvi [84] | Cohort | Prostate cancer | 59 | RT-PCR, ddPCR | AR | Detection of circulating AR copy number gain is a non-invasive biomarker for outcome of patients with CRPC treated with enzalutamide. |
| 2017, Sumanta K. Pal [87] | Prospective, cohort | Renal cell carcinoma | 220 | HiSeq 2500 Illumina | 73 cancer-related genes | Higher rates of detection by ctDNA analysis after systemic therapy compared with baseline was noted for NF1, TP53, and VHL, indicating clonal evolution of genomic alterations. |
| 2018, Neeraj Agarwal [89] | Cohort | Urothelial carcinoma | 369 | HiSeq 2500 Illumina | 73 cancer-related genes | ctDNA NGS identified similar genomic alterations with tumor tissue. The genomic landscape was similar between lower tract and upper tract urothelial carcinoma. |
| 2017, Emil Christensen [91] | Cohort | Urothelial carcinoma | 831 | ddPCR | FGFR3 and PIK3CA | ctDNA levels in the urine and plasma were positively correlated and indicated that higher levels of FGFR3- and PIK3CA-mutated DNA can predict disease progression. |
| Skin cancer | ||||||
| 2016, Gregory A. Chang [92] | Cohort | Melanoma | 43 | ddPCR | NRAS, BRAF | ctDNA had a higher sensitivity than LDH to detect disease progression. |
| 2016, Anne C. Knol [93] | Cohort | Melanoma | 38 | RT-PCR | BRAF | ctDNA BRAF mutation is a prognostic factor of OS and it is correlated with tumor burden. |
| 2015, Elin S. Gray [94] | Cohort | Melanoma | 48 | ddPCR | NRAS, BRAF | ctDNA is a biomarker of response to kinase inhibitor therapy and it can be used to monitor resistance to treatment. |
| 2015, Maria Gonzalez-Cao [95] | Cohort | Melanoma | 22 | TaqMan assay PCR | BRAF | Detection and accurate quantification of low- BRAF V600E in ctDNA can predict treatment outcome. |
| 2016, Ademi Santiago-Walker [96] | Cohort | Melanoma | 732 | PCR-based BEAMing | BRAF | BRAF mutation using ctDNA analysis can be detected in >75% of patients and is a prognostic marker. |
| 2016, Max Schreuer [97] | Cohort | Melanoma | 36 | qPCR | BRAF | Quantitative analysis of BRAF mutation in ctDNA is a monitoring tool during treatment with BRAF/MEK inhibitors. |
| 2017, Stephen Q. Wong [98] | Cohort | Melanoma | 52 | Amplicon sequencing, ddPCR | NRAS, BRAF | ctDNA is a powerful complementary modality to functional imaging for real-time monitoring of tumor burden and genomic changes throughout therapy. |
| 2018, R. Jeffrey Lee [99] | Cohort | Melanoma | 161 | QX200 ddPCR | NRAS, BRAF | ctDNA predicts relapse and survival in high-risk resected stage II/III melanoma and can help select patients for adjuvant therapy. |
| Sarcoma | ||||||
| 2016, Manuela Krumbholz [101] | Cohort | Ewing | 20 | AccuPrime Taq DNA PCR | EWSR1-FLI1 fusion | Detection of EWSR1 fusion sequence in plasma is a promising noninvasive biomarker for improved therapeutic monitoring. |
| 2016, Masanori Hayashi [102] | Cohort | Ewing | 3 | ddPCR | EWS-ETS | Tumor specific EWS-ETS translocation breakpoints in plasma DNA is a highly personalized biomarker for relapsed disease. |
| 2018, David S. Shulman [103] | Cohort | Ewing, osteosarcoma | 166 | Illumina HiSeq 2500 | EWSR1, FUS, CIC, CCNB3, TP53, STAG2 | Detectable ctDNA in patients with localized disease is associated with inferior event-free survival and OS at 3 years compared to patients with undetectable ctDNA levels. |
| 2013, Jacqueline Maier [104] | Prospective cohort | Gastrointestinal stromal tumor | 38 | RT-PCR | CKIT, PDGFRA | ctDNA harboring CKIT or PDGFRA was correlated with disease course. |
| 2014, Changhoon Yoo [105] | Cohort | Gastrointestinal stromal tumor | 30 | PCR-based BEAMing | CKIT, PDGFRA, BRAF | Genotyping of the KIT gene in exon 17 of serum ctDNA identified mutations associated with resistance to dovitinib. |
| 2016, Noriko Wada [106] | Cohort | Gastrointestinal stromal tumor | 4 | Sanger sequencing, PCR | C-KIT | Detection of secondary C-KIT mutations in ctDNA is useful to select targeted agents and to predict antitumor effects. |
| 2018, Nicholas C. Eastley [107] | Cohort | Soft tissue sarcoma | 11 | Ion AmpliSeq | 57 cancer-related genes | ctDNA analysis detected TP53/PIK3CA mutations concordant with the primary tumor in 2 of 4 cases. |
| Brain tumors | ||||||
| 2013, Mohamad A. Salkeni [108] | Prospective | Glioblastoma | 13 | Illumina HiSeq | EGFR (vIII deletion) | ctDNA analysis identified the EGFRvIII deletion in 3 of 13 patients, which was correlated with tumor tissue analysis and may help select patients for targeted therapy. ctDNA levels were correlated with the extent of tumor. |
| 2016, Shigeki Yagyu [109] | Retrospective | Neuroblastoma | 151 | Real-time quantitative PCR | MYCN | Serum MYCN amplification (sensitivity 86%, specificity 95% compared with tissue analysis) was associated with OS. It may help select treatment prior to tumor biopsy, particularly for patients < 18 months (risk assessment and treatment depend on MYCN amplification status). |
| 2012, Blandine Boisselier [110] | Prospective | Glioma | 80 patients, 31 controls | Digital PCR, Agilent technologies | IDH1 | The IDH1 R132H mutation was identified in plasma (specificity, 100%; sensitivity related to the tumor volume and contrast enhancement). It may help in the diagnosis of patients not amenable to biopsy. |
| Lymphoma | ||||||
| 2016, Vincent Camus [111] | Cohort | Hodgkin | 94 | TaqMan assay PCR | XPO1 | The XPO1 E571K mutation in ctDNA can be used as a novel biomarker in diagnosis and detection of minimal residual disease. |
| 2016, Sarit E. Assouline [112] | Phase 2 trial | Diffuse large B-cell | 40 | ddPCR | CREBBP, EP300, MLL2, FAS, STAT6, TP53 | Increase in ctDNA levels at 15 days after treatment initiation was associated with resistance to treatment. |
| 2016, Florian Scherer [113] | Case control | Diffuse large B-cell | 92 patients, 24 controls | CAPP-Seq | BCL2, BCL6, MYC, IGH | ctDNA levels at diagnosis were strongly correlated with clinical indices and were independently predictive of patient outcomes. |
| 2015, David M. Kurtz [114] | Prospective cohort | Diffuse large B-cell | 75 | RT-PCR | Immunoglobulin high-throughput sequencing (Ig-HTS) | ctDNA immunoglobulin high-throughput sequencing preceded radiologic evidence of recurrent disease indicating that it may be a surrogate marker for monitoring disease after complete remission. |
| 2015, Mark Roschewski [115] | Retrospective analysis | Diffuse large B-cell | 126 | LymphoSIGHT™ | VDJ | After first-line treatment, disease progression was evident on imaging studies a median of 3.5 months after detection on ctDNA analysis of the clonal Ig gene sequence. |
| Across Tumor Types | ||||||
| 2012, Geraldine Perkins [117] | Cohort from multiple phase 1 trials | Colorectal, breast, melanoma, prostate, ovarian, and other | 105 | Sequenom MassArray, OncoCarta PCR | KRAS, BRAF, PIK3CA | ctDNA analysis has potential clinical multi-purpose utility in patients with advanced cancer. |
| 2015, Filip Janku [118] | Cohort from multiple phase 1 trials | Colorectal, melanoma, non-small cell lung, and other | 157 | PCR-based BEAMing | BRAF, EGFR, KRAS, PIK3CA | Patients with > 1% of mutant ctDNA had shorter median OS compared to patients with ≤ 1%. |
| 2014, Chetan Bettegowda [119] | Cohort | Pancreatic, ovarian, colorectal, bladder, gastroesophageal, breast, melanoma, hepatocellular, head and neck, and other | 640 | BEAMing, PCR-Ligation, Safe-SeqS | 187 cancer-related genes | ctDNA is a broadly applicable, sensitive, and specific biomarker that can be used for clinical and research purposes in patients with various tumor types. |
| 2013, Muhammed Murtaza [120] | Cohort | Breast, ovarian and lung | 6 | HiSeq 2500 Illumina | PIK3CA, RB1, GAS6, EGFR | ctDNA analyses can complement invasive tumor biopsies to identify mutations associated with acquired drug resistance in advanced cancer. |
| 2015, Jean Sebastien Frenel [121] | Cohort | Colorectal, ovarian, breast, bladder, glioblastoma, lung adenocarcinoma, endometrial | 39 | Ion AmpliSeq, ddPCR | Cancer-related genes | Targeted sequencing of ctDNA has potential clinical utility to monitor the effect of targeted therapies. |
| 2016, Maria Schwaederle [122] | Cohort | Lung, breast, glioblastoma, genitourinary, gastrointestinal, of unknown primary, and other | 171 | Illumina Hi-Seq 2500 | 54 cancer-related genes | A large proportion of patients had detectable ctDNA aberration (s), among which the majority are targetable by an approved drug. |
| 2016, Maria Schwaederle [123] | Cohort | Brain, lung, breast | 168 | Illumina Hi-Seq 2500 | 54 cancer-related genes | ctDNA tests provide information complementary to the tissue biopsies and may be useful in determining prognosis and treatment. |
| 2017, Yulian Khagi [124] | Cohort | Skin, lung, breast, glioblastoma, genitourinary, gastrointestinal, and other | 69 | Illumina Hi-Seq 2500 | 54–70 cancer-related genes | Hyper-mutated ctDNA is correlated with response to checkpoint inhibitor-based therapy and investigation of hypermutated ctDNA as a predictive biomarker is warranted. |
Conclusions
ctDNA analysis is a non-invasive, cost-effective test with a potentially significant role in the early detection and diagnosis of tumors. Evolving data from clinical trials indicate the association of ctDNA with tumor burden and the usefulness of ctDNA analysis in assessment of MRD, in understanding mechanisms of resistance to treatment, and in dynamically guiding therapy. The discordance between ctDNA analysis and tumor tissue genomic analysis is attributed, at least in part, to biologic and technical differences in detection of alterations between DNA shed by the tumor in the circulatory system and DNA in tumor tissue. ctDNA is thought to reflect tumor from all sites of disease and is secreted by tumor cells, phagocyte-engulfed tumor cells, and necrotic or apoptotic tumor cells. Selected prospective trials with targeted agents incorporate ctDNA analysis to select targeted therapy. Longitudinal ctDNA analysis starting at the time of diagnosis will enrich our knowledge of the evolution of patients’ tumor biology, will accelerate drug development, and will contribute to the implementation of precision medicine to improve clinical outcomes.
CONFLICTS OF INTEREST
Dr. Rabih Said: has no financial relationship to disclose. Dr. Nicolas Guibert has the following financial relationships to disclose: Consulting fees from Astra Zeneca, Roche, Bristol-Myers-Squibb, and MSD Pharmaceuticals. Dr. Geoffrey Oxnard has the following financial relationships to disclose: Consulting fees from Astra Zeneca, DropWorks, GRAIL, Inivata, Jannsen, Loxo, Sysmex, and Ilumina, and Honoraria from Foundation Medicine, and Guardant. Dr. Apostolia-Maria Tsimberidou has the following financial relationships to disclose: Research Funding (Institution): Immatics, Parker Institute for Cancer Immunotherapy, Tempus, OBI Pharma, EMD Serono, Baxalta, ONYX, Bayer, Boston Biomedical, Placon Therapeutics, and Karus Therapeutics, Tvardi. Consulting or Advisory Role: Roche, Covance, and Genentech.
References
1. Oxnard GR, Thress KS, Alden RS, Lawrance R, Paweletz CP, Cantarini M, Yang JC, Barrett JC, Janne PA. Association Between Plasma Genotyping and Outcomes of Treatment With Osimertinib (AZD9291) in Advanced Non-Small-Cell Lung Cancer. J Clin Oncol. 2016; 34:3375–3382. https://doi.org/10.1200/JCO.2016.66.7162. [PubMed].
2. Thierry AR, Mouliere F, El Messaoudi S, Mollevi C, Lopez-Crapez E, Rolet F, Gillet B, Gongora C, Dechelotte P, Robert B, Del Rio M, Lamy PJ, Bibeau F, et al. Clinical validation of the detection of KRAS and BRAF mutations from circulating tumor DNA. Nat Med. 2014; 20:430–435. https://doi.org/10.1038/nm.3511. [PubMed].
3. Fiala C, Diamandis EP. Utility of circulating tumor DNA in cancer diagnostics with emphasis on early detection. BMC Med. 2018; 16:166. https://doi.org/10.1186/s12916-018-1157-9. [PubMed].
4. Yeow KM, Su IH, Pan KT, Tsay PK, Lui KW, Cheung YC, Chou AS. Risk factors of pneumothorax and bleeding: multivariate analysis of 660 CT-guided coaxial cutting needle lung biopsies. Chest. 2004; 126:748–754. https://doi.org/10.1378/chest.126.3.748. [PubMed].
5. Steinfort DP, Khor YH, Manser RL, Irving LB. Radial probe endobronchial ultrasound for the diagnosis of peripheral lung cancer: systematic review and meta-analysis. Eur Respir J. 2011; 37:902–910. https://doi.org/10.1183/09031936.00075310. [PubMed].
6. Asano F, Eberhardt R, Herth FJ. Virtual bronchoscopic navigation for peripheral pulmonary lesions. Respiration. 2014; 88:430–440. https://doi.org/10.1159/000367900. [PubMed].
7. Oxnard GR, Paweletz CP, Kuang Y, Mach SL, O’Connell A, Messineo MM, Luke JJ, Butaney M, Kirschmeier P, Jackman DM, Janne PA. Noninvasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res. 2014; 20:1698–1705. https://doi.org/10.1158/1078-0432.CCR-13-2482. [PubMed].
8. Sacher AG, Paweletz C, Dahlberg SE, Alden RS, O’Connell A, Feeney N, Mach SL, Janne PA, Oxnard GR. Prospective Validation of Rapid Plasma Genotyping for the Detection of EGFR and KRAS Mutations in Advanced Lung Cancer. JAMA Oncol. 2016; 2:1014–1022. https://doi.org/10.1001/jamaoncol.2016.0173. [PubMed].
9. Luo J, Shen L, Zheng D. Diagnostic value of circulating free DNA for the detection of EGFR mutation status in NSCLC: a systematic review and meta-analysis. Sci Rep. 2014; 4:6269. https://doi.org/10.1038/srep06269. [PubMed].
10. Thompson JC, Yee SS, Troxel AB, Savitch SL, Fan R, Balli D, Lieberman DB, Morrissette JD, Evans TL, Bauml J, Aggarwal C, Kosteva JA, Alley E, et al. Detection of Therapeutically Targetable Driver and Resistance Mutations in Lung Cancer Patients by Next-Generation Sequencing of Cell-Free Circulating Tumor DNA. Clin Cancer Res. 2016; 22:5772–5782. https://doi.org/10.1158/1078-0432.CCR-16-1231. [PubMed].
11. Schwaederle MC, Patel SP, Husain H, Ikeda M, Lanman RB, Banks KC, Talasaz A, Bazhenova L, Kurzrock R. Utility of Genomic Assessment of Blood-Derived Circulating Tumor DNA (ctDNA) in Patients with Advanced Lung Adenocarcinoma. Clin Cancer Res. 2017; 23:5101–5111. https://doi.org/10.1158/1078-0432.CCR-16-2497. [PubMed].
12. Yao Y, Liu J, Li L, Yuan Y, Nan K, Wu X, Zhang Z, Wu Y, Li X, Zhu J, Meng X, Wei L, Chen J, et al. Detection of circulating tumor DNA in patients with advanced non-small cell lung cancer. Oncotarget. 2017; 8:2130–2140. https://doi.org/10.18632/oncotarget.12883. [PubMed].
13. Guibert N, Hu Y, Feeney N, Kuang Y, Plagnol V, Jones G, Howarth K, Beeler JF, Paweletz CP, Oxnard GR. Amplicon-based next-generation sequencing of plasma cell-free DNA for detection of driver and resistance mutations in advanced non-small cell lung cancer. Ann Oncol. 2018; 29:1049–1055. https://doi.org/10.1093/annonc/mdy005. [PubMed].
14. Cui S, Zhang W, Xiong L, Pan F, Niu Y, Chu T, Wang H, Zhao Y, Jiang L. Use of capture-based next-generation sequencing to detect ALK fusion in plasma cell-free DNA of patients with non-small-cell lung cancer. Oncotarget. 2017; 8:2771–2780. https://doi.org/10.18632/oncotarget.13741. [PubMed].
15. Wang Y, Tian PW, Wang WY, Wang K, Zhang Z, Chen BJ, He YQ, Li L, Liu H, Chuai S, Li WM. Noninvasive genotyping and monitoring of anaplastic lymphoma kinase (ALK) rearranged non-small cell lung cancer by capture-based next-generation sequencing. Oncotarget. 2016; 7:65208–65217. https://doi.org/10.18632/oncotarget.11569. [PubMed].
16. Paweletz CP, Sacher AG, Raymond CK, Alden RS, O’Connell A, Mach SL, Kuang Y, Gandhi L, Kirschmeier P, English JM, Lim LP, Janne PA, Oxnard GR. Bias-Corrected Targeted Next-Generation Sequencing for Rapid, Multiplexed Detection of Actionable Alterations in Cell-Free DNA from Advanced Lung Cancer Patients. Clin Cancer Res. 2016; 22:915–922. https://doi.org/10.1158/1078-0432.CCR-15-1627-T. [PubMed].
17. Guibert NM, Paweletz C, Hu Y, Feeney NB, Plagnol V, Poole V, Jones G, Oxnard GR. Early detection of competing resistance mutations using plasma next-generation sequencing (NGS) in patients (pts) with EGFR-mutant NSCLC treated with osimertinib. J Clin Oncol. 2017; 35:1. https://doi.org/10.1200/JCO.2017.35.15_suppl.11529.
18. Remon J, Caramella C, Jovelet C, Lacroix L, Lawson A, Smalley S, Howarth K, Gale D, Green E, Plagnol V, Rosenfeld N, Planchard D, Bluthgen MV, et al. Osimertinib benefit in EGFR-mutant NSCLC patients with T790M-mutation detected by circulating tumour DNA. Ann Oncol. 2017; 28:784–790. https://doi.org/10.1093/annonc/mdx017. [PubMed].
19. Remon J, Menis J, Hasan B, Peric A, De Maio E, Novello S, Reck M, Berghmans T, Wasag B, Besse B, Dziadziuszko R. The APPLE Trial: Feasibility and Activity of AZD9291 (Osimertinib) Treatment on Positive PLasma T790M in EGFR-mutant NSCLC Patients. EORTC 1613. Clin Lung Cancer. 2017; 18:583–588. https://doi.org/10.1016/j.cllc.2017.02.005. [PubMed].
20. Aggarwal C, Thompson JC, Black TA, Katz SI, Fan R, Yee SS, Chien AL, Evans TL, Bauml JM, Alley EW, Ciunci CA, Berman AT, Cohen RB, et al. Clinical Implications of Plasma-Based Genotyping With the Delivery of Personalized Therapy in Metastatic Non-Small Cell Lung Cancer. JAMA Oncol. 2019; 5:173–180. https://doi.org/10.1001/jamaoncol.2018.4305. [PubMed].
21. Li BT, Janku F, Jung B, Hou C, Madwani K, Alden R, Razavi P, Reis-Filho JS, Shen R, Isbell JM, Blocker AW, Eattock N, Gnerre S, et al. Ultra-deep next-generation sequencing of plasma cell-free DNA in patients with advanced lung cancers: results from the Actionable Genome Consortium. Ann Oncol. 2019; 30:597–603. https://doi.org/10.1093/annonc/mdz046. [PubMed].
22. Leighl NB, Page RD, Raymond VM, Daniel DB, Divers SG, Reckamp KL, Villalona-Calero MA, Dix D, Odegaard JI, Lanman RB, Papadimitrakopoulou VA. Clinical Utility of Comprehensive Cell-free DNA Analysis to Identify Genomic Biomarkers in Patients with Newly Diagnosed Metastatic Non-small Cell Lung Cancer. Clin Cancer Res. 2019; 25:4691–4700. https://doi.org/10.1158/1078-0432.CCR-19-0624. [PubMed].
23. Church TR, Black WC, Aberle DR, Berg CD, Clingan KL, Duan F, Fagerstrom RM, Gareen IF, Gierada DS, Jones GC, Mahon I, Marcus PM, Sicks JD, et al. and National Lung Screening Trial Research Team. Results of initial low-dose computed tomographic screening for lung cancer. N Engl J Med. 2013; 368:1980–1991. https://doi.org/10.1056/NEJMoa1209120. [PubMed].
24. Walter JE, Heuvelmans MA, de Jong PA, Vliegenthart R, van Ooijen PMA, Peters RB, Ten Haaf K, Yousaf-Khan U, van der Aalst CM, de Bock GH, Mali W, Groen HJM, de Koning HJ, et al. Occurrence and lung cancer probability of new solid nodules at incidence screening with low-dose CT: analysis of data from the randomised, controlled NELSON trial. Lancet Oncol. 2016; 17:907–916. https://doi.org/10.1016/S1470-2045(16)30069-9. [PubMed].
25. Newman AM, Bratman SV, To J, Wynne JF, Eclov NC, Modlin LA, Liu CL, Neal JW, Wakelee HA, Merritt RE, Shrager JB, Loo BW Jr, Alizadeh AA, et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med. 2014; 20:548–554. https://doi.org/10.1038/nm.3519. [PubMed].
26. Abbosh C, Birkbak NJ, Wilson GA, Jamal-Hanjani M, Constantin T, Salari R, Le Quesne J, Moore DA, Veeriah S, Rosenthal R, Marafioti T, Kirkizlar E, Watkins TBK, et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature. 2017; 545:446–451. https://doi.org/10.1038/nature22364. [PubMed].
27. Oxnard GR, Maddala T, Hubbell E, Aravanis A, Zhang N, Venn O, Valouev A, Shen L, Patel S, Jamshidi A, Jagadeesh K, Gross S, Filippova D, et al. Genome-wide sequencing for early stage lung cancer detection from plasma cell-free DNA (cfDNA): The Circulating Cancer Genome Atlas (CCGA) study. J Clin Oncol. 2018; 36:18.
28. Hu Y, Ulrich BC, Supplee J, Kuang Y, Lizotte PH, Feeney NB, Guibert NM, Awad MM, Wong KK, Janne PA, Paweletz CP, Oxnard GR. False-Positive Plasma Genotyping Due to Clonal Hematopoiesis. Clin Cancer Res. 2018; 24:4437–4443. https://doi.org/10.1158/1078-0432.CCR-18-0143. [PubMed].
29. Paweletz CP, Lau CJ, Oxnard GR. Does Testing Error Underlie Liquid Biopsy Discordance? JCO Precision Oncology. 2019. https://doi.org/10.1200/PO.18.00408.
30. Jamal-Hanjani M, Wilson GA, McGranahan N, Birkbak NJ, Watkins TBK, Veeriah S, Shafi S, Johnson DH, Mitter R, Rosenthal R, Salm M, Horswell S, Escudero M, et al. Tracking the Evolution of Non-Small-Cell Lung Cancer. N Engl J Med. 2017; 376:2109–2121. https://doi.org/10.1056/NEJMoa1616288. [PubMed].
31. Chaudhuri AA, Chabon JJ, Lovejoy AF, Newman AM, Stehr H, Azad TD, Khodadoust MS, Esfahani MS, Liu CL, Zhou L, Scherer F, Kurtz DM, Say C, et al. Early Detection of Molecular Residual Disease in Localized Lung Cancer by Circulating Tumor DNA Profiling. Cancer Discov. 2017; 7:1394–1403. https://doi.org/10.1158/2159-8290.CD-17-0716. [PubMed].
32. Luo H, Li H, Hu Z, Wu H, Liu C, Li Y, Zhang X, Lin P, Hou Q, Ding G, Wang Y, Li S, Wei D, et al. Noninvasive diagnosis and monitoring of mutations by deep sequencing of circulating tumor DNA in esophageal squamous cell carcinoma. Biochem Biophys Res Commun. 2016; 471:596–602. https://doi.org/10.1016/j.bbrc.2016.02.011. [PubMed].
33. Takeshita H, Ichikawa D, Komatsu S, Tsujiura M, Kosuga T, Deguchi K, Konishi H, Morimura R, Shiozaki A, Fujiwara H, Okamoto K, Otsuji E. Prediction of CCND1 amplification using plasma DNA as a prognostic marker in oesophageal squamous cell carcinoma. Br J Cancer. 2010; 102:1378–1383. https://doi.org/10.1038/sj.bjc.6605657. [PubMed].
34. Ueda M, Iguchi T, Masuda T, Nakahara Y, Hirata H, Uchi R, Niida A, Momose K, Sakimura S, Chiba K, Eguchi H, Ito S, Sugimachi K, et al. Somatic mutations in plasma cell-free DNA are diagnostic markers for esophageal squamous cell carcinoma recurrence. Oncotarget. 2016; 7:62280–62291. https://doi.org/10.18632/oncotarget.11409. [PubMed].
35. Azad TD, Chaudhuri A, Newman AM, Stehr H, Schroers-Martin J, Chabon JJ, Fang P, Qiao Y, Liao ZX, Komaki R, Alizadeh AA, Lin SH, Diehn M. Circulating tumor DNA analysis for outcome prediction in localized esophageal cancer. J Clin Oncol. 2017; 35:1. https://doi.org/10.1200/JCO.2017.35.15_suppl.4055.
36. Shoda K, Masuda K, Ichikawa D, Arita T, Miyakami Y, Watanabe M, Konishi H, Imoto I, Otsuji E. HER2 amplification detected in the circulating DNA of patients with gastric cancer: a retrospective pilot study. Gastric Cancer. 2015; 18:698–710. https://doi.org/10.1007/s10120-014-0432-5. [PubMed].
37. Shoda K, Ichikawa D, Fujita Y, Masuda K, Hiramoto H, Hamada J, Arita T, Konishi H, Komatsu S, Shiozaki A, Kakihara N, Okamoto K, Taniguchi H, et al. Monitoring the HER2 copy number status in circulating tumor DNA by droplet digital PCR in patients with gastric cancer. Gastric Cancer. 2017; 20:126–135. https://doi.org/10.1007/s10120-016-0599-z. [PubMed].
38. Kinugasa H, Nouso K, Tanaka T, Miyahara K, Morimoto Y, Dohi C, Matsubara T, Okada H, Yamamoto K. Droplet digital PCR measurement of HER2 in patients with gastric cancer. Br J Cancer. 2015; 112:1652–1655. https://doi.org/10.1038/bjc.2015.129. [PubMed].
39. Fang WL, Lan YT, Huang KH, Liu CA, Hung YP, Lin CH, Jhang FY, Chang SC, Chen MH, Chao Y, Lin WC, Lo SS, Fen-Yau Li A, et al. Clinical significance of circulating plasma DNA in gastric cancer. Int J Cancer. 2016; 138:2974–2983. https://doi.org/10.1002/ijc.30018. [PubMed].
40. Kim ST, Banks KC, Lee SH, Kim K, Park JO, Park SH, Park YS, Lim HY, Kang WK, Lanman RB, Talasaz AA, Park K, Lee J. Prospective Feasibility Study for Using Cell-Free Circulating Tumor DNA–Guided Therapy in Refractory Metastatic Solid Cancers: An Interim Analysis. JCO Precision Oncology. 2017:1–15. https://doi.org/10.1200/PO.16.00059.
41. O’Connor ES, Greenblatt DY, LoConte NK, Gangnon RE, Liou JI, Heise CP, Smith MA. Adjuvant chemotherapy for stage II colon cancer with poor prognostic features. J Clin Oncol. 2011; 29:3381–3388. https://doi.org/10.1200/JCO.2010.34.3426. [PubMed].
42. Andre T, de Gramont A, Vernerey D, Chibaudel B, Bonnetain F, Tijeras-Raballand A, Scriva A, Hickish T, Tabernero J, Van Laethem JL, Banzi M, Maartense E, Shmueli E, et al. Adjuvant Fluorouracil, Leucovorin, and Oxaliplatin in Stage II to III Colon Cancer: Updated 10-Year Survival and Outcomes According to BRAF Mutation and Mismatch Repair Status of the MOSAIC Study. J Clin Oncol. 2015; 33:4176–4187. https://doi.org/10.1200/JCO.2015.63.4238. [PubMed].
43. Tie J, Wang Y, Tomasetti C, Li L, Springer S, Kinde I, Silliman N, Tacey M, Wong HL, Christie M, Kosmider S, Skinner I, Wong R, et al. Circulating tumor DNA analysis detects minimal residual disease and predicts recurrence in patients with stage II colon cancer. Sci Transl Med. 2016; 8:346ra392. https://doi.org/10.1126/scitranslmed.aaf6219. [PubMed].
44. Tie J, Cohen J, Wang Y, Lee M, Wong R, Kosmider S, Ananda S, Cho JH, Faragher I, McKendrick JJ, Corfield C, Lee B, Field KM, et al. Serial circulating tumor DNA (ctDNA) analysis as a prognostic marker and a real-time indicator of adjuvant chemotherapy (CT) efficacy in stage III colon cancer (CC). J Clin Oncol. 2018; 36:3516–3516. https://doi.org/10.1200/JCO.2018.36.15_suppl.3516.
45. Scholer LV, Reinert T, Orntoft MW, Kassentoft CG, Arnadottir SS, Vang S, Nordentoft I, Knudsen M, Lamy P, Andreasen D, Mortensen FV, Knudsen AR, Stribolt K, et al. Clinical Implications of Monitoring Circulating Tumor DNA in Patients with Colorectal Cancer. Clin Cancer Res. 2017; 23:5437–5445. https://doi.org/10.1158/1078-0432.CCR-17-0510. [PubMed].
46. Zill OA, Greene C, Sebisanovic D, Siew LM, Leng J, Vu M, Hendifar AE, Wang Z, Atreya CE, Kelley RK, Van Loon K, Ko AH, Tempero MA, et al. Cell-Free DNA Next-Generation Sequencing in Pancreatobiliary Carcinomas. Cancer Discov. 2015; 5:1040–1048. https://doi.org/10.1158/2159-8290.CD-15-0274. [PubMed].
47. Tjensvoll K, Lapin M, Buhl T, Oltedal S, Steen-Ottosen Berry K, Gilje B, Soreide JA, Javle M, Nordgard O, Smaaland R. Clinical relevance of circulating KRAS mutated DNA in plasma from patients with advanced pancreatic cancer. Mol Oncol. 2016; 10:635–643. https://doi.org/10.1016/j.molonc.2015.11.012. [PubMed].
48. Hadano N, Murakami Y, Uemura K, Hashimoto Y, Kondo N, Nakagawa N, Sueda T, Hiyama E. Prognostic value of circulating tumour DNA in patients undergoing curative resection for pancreatic cancer. Br J Cancer. 2016; 115:59–65. https://doi.org/10.1038/bjc.2016.175. [PubMed].
49. Pishvaian MJ, Joseph Bender R, Matrisian LM, Rahib L, Hendifar A, Hoos WA, Mikhail S, Chung V, Picozzi V, Heartwell C, Mason K, Varieur K, Aberra M, et al. A pilot study evaluating concordance between blood-based and patient-matched tumor molecular testing within pancreatic cancer patients participating in the Know Your Tumor (KYT) initiative. Oncotarget. 2017; 8:83446–83456. https://doi.org/10.18632/oncotarget.13225. [PubMed].
50. Pietrasz D, Pecuchet N, Garlan F, Didelot A, Dubreuil O, Doat S, Imbert-Bismut F, Karoui M, Vaillant JC, Taly V, Laurent-Puig P, Bachet JB. Plasma Circulating Tumor DNA in Pancreatic Cancer Patients Is a Prognostic Marker. Clin Cancer Res. 2017; 23:116–123. https://doi.org/10.1158/1078-0432.CCR-16-0806. [PubMed].
51. Lee B, Lipton LR, Cohen J, Tie J, Javed AA, Li L, Goldstein D, Cooray P, Nagrial A, Burge ME, Tebbutt NC, Nikfarjam M, Harris M, et al. Circulating tumor DNA as a prognostic biomarker in early stage pancreatic cancer. J Clin Oncol. 2018; 36. https://doi.org/10.1200/JCO.2018.36.15_suppl.e16206.
52. Henriksen SD, Madsen PH, Larsen AC, Johansen MB, Drewes AM, Pedersen IS, Krarup H, Thorlacius-Ussing O. Cell-free DNA promoter hypermethylation in plasma as a diagnostic marker for pancreatic adenocarcinoma. Clin Epigenetics. 2016; 8:117. https://doi.org/10.1186/s13148-016-0286-2. [PubMed].
53. Berger AW, Ettrich TJ, Schwerdel D, Dolnik A, Beuter F, Blätte TJ, Schmidt SA, Stanescu N, Steinacker J, Marienfeld R, Kleger A, Bullinger L, Seufferlein T. Genotyping of circulating tumor DNA in biliary tract cancer reveals diagnostic and prognostic information. J Clin Oncol. 2018; 36. https://doi.org/10.1200/JCO.2018.36.15_suppl.e16147.
54. Ono A, Fujimoto A, Yamamoto Y, Akamatsu S, Hiraga N, Imamura M, Kawaoka T, Tsuge M, Abe H, Hayes CN, Miki D, Furuta M, Tsunoda T, et al. Circulating Tumor DNA Analysis for Liver Cancers and Its Usefulness as a Liquid Biopsy. Cell Mol Gastroenterol Hepatol. 2015; 1:516–534. https://doi.org/10.1016/j.jcmgh.2015.06.009. [PubMed].
55. Liao W, Yang H, Xu H, Wang Y, Ge P, Ren J, Xu W, Lu X, Sang X, Zhong S, Zhang H, Mao Y. Noninvasive detection of tumor-associated mutations from circulating cell-free DNA in hepatocellular carcinoma patients by targeted deep sequencing. Oncotarget. 2016; 7:40481–40490. https://doi.org/10.18632/oncotarget.9629. [PubMed].
56. Huang A, Zhang X, Zhou SL, Cao Y, Huang XW, Fan J, Yang XR, Zhou J. Detecting Circulating Tumor DNA in Hepatocellular Carcinoma Patients Using Droplet Digital PCR Is Feasible and Reflects Intratumoral Heterogeneity. J Cancer. 2016; 7:1907–1914. https://doi.org/10.7150/jca.15823. [PubMed].
57. Parsons HA, Beaver JA, Cimino-Mathews A, Ali SM, Axilbund J, Chu D, Connolly RM, Cochran RL, Croessmann S, Clark TA, Gocke CD, Jeter SC, Kennedy MR, et al. Individualized Molecular Analyses Guide Efforts (IMAGE): A Prospective Study of Molecular Profiling of Tissue and Blood in Metastatic Triple-Negative Breast Cancer. Clin Cancer Res. 2017; 23:379–386. https://doi.org/10.1158/1078-0432.CCR-16-1543. [PubMed].
58. Beaver JA, Jelovac D, Balukrishna S, Cochran R, Croessmann S, Zabransky DJ, Wong HY, Toro PV, Cidado J, Blair BG, Chu D, Burns T, Higgins MJ, et al. Detection of cancer DNA in plasma of patients with early-stage breast cancer. Clin Cancer Res. 2014; 20:2643–2650. https://doi.org/10.1158/1078-0432.CCR-13-2933. [PubMed].
59. Lanman RB, Mortimer SA, Zill OA, Sebisanovic D, Lopez R, Blau S, Collisson EA, Divers SG, Hoon DS, Kopetz ES, Lee J, Nikolinakos PG, Baca AM, et al. Analytical and Clinical Validation of a Digital Sequencing Panel for Quantitative, Highly Accurate Evaluation of Cell-Free Circulating Tumor DNA. PLoS One. 2015; 10:e0140712. https://doi.org/10.1371/journal.pone.0140712. [PubMed].
60. Liang DH, Ensor JE, Liu ZB, Patel A, Patel TA, Chang JC, Rodriguez AA. Cell-free DNA as a molecular tool for monitoring disease progression and response to therapy in breast cancer patients. Breast Cancer Res Treat. 2016; 155:139–149. https://doi.org/10.1007/s10549-015-3635-5. [PubMed].
61. Rudolph M, Anzeneder T, Schulz A, Beckmann G, Byrne AT, Jeffers M, Pena C, Politz O, Kochert K, Vonk R, Reischl J. AKT1 (E17K) mutation profiling in breast cancer: prevalence, concurrent oncogenic alterations, and blood-based detection. BMC Cancer. 2016; 16:622. https://doi.org/10.1186/s12885-016-2626-1. [PubMed].
62. Moynahan ME, Chen D, He W, Sung P, Samoila A, You D, Bhatt T, Patel P, Ringeisen F, Hortobagyi GN, Baselga J, Chandarlapaty S. Correlation between PIK3CA mutations in cell-free DNA and everolimus efficacy in HR(+), HER2(-) advanced breast cancer: results from BOLERO-2. Br J Cancer. 2017; 116:726–730. https://doi.org/10.1038/bjc.2017.25. [PubMed].
63. Chandarlapaty S, Chen D, He W, Sung P, Samoila A, You D, Bhatt T, Patel P, Voi M, Gnant M, Hortobagyi G, Baselga J, Moynahan ME. Prevalence of ESR1 Mutations in Cell-Free DNA and Outcomes in Metastatic Breast Cancer: A Secondary Analysis of the BOLERO-2 Clinical Trial. JAMA Oncol. 2016; 2:1310–1315. https://doi.org/10.1001/jamaoncol.2016.1279. [PubMed].
64. Di Leo A, Johnston S, Lee KS, Ciruelos E, Lonning PE, Janni W, O’Regan R, Mouret-Reynier MA, Kalev D, Egle D, Csoszi T, Bordonaro R, Decker T, et al. Buparlisib plus fulvestrant in postmenopausal women with hormone-receptor-positive, HER2-negative, advanced breast cancer progressing on or after mTOR inhibition (BELLE-3): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 2018; 19:87–100. https://doi.org/10.1016/S1470-2045(17)30688-5. [PubMed].
65. Condorelli R, Spring L, O’Shaughnessy J, Lacroix L, Bailleux C, Scott V, Dubois J, Nagy RJ, Lanman RB, Iafrate AJ, Andre F, Bardia A. Polyclonal RB1 mutations and acquired resistance to CDK 4/6 inhibitors in patients with metastatic breast cancer. Ann Oncol. 2018; 29:640–645. https://doi.org/10.1093/annonc/mdx784. [PubMed].
66. Ma F, Zhu W, Guan Y, Yang L, Xia X, Chen S, Li Q, Guan X, Yi Z, Qian H, Yi X, Xu B. ctDNA dynamics: a novel indicator to track resistance in metastatic breast cancer treated with anti-HER2 therapy. Oncotarget. 2016; 7:66020–66031. https://doi.org/10.18632/oncotarget.11791. [PubMed].
67. Riva F, Bidard FC, Houy A, Saliou A, Madic J, Rampanou A, Hego C, Milder M, Cottu P, Sablin MP, Vincent-Salomon A, Lantz O, Stern MH, et al. Patient-Specific Circulating Tumor DNA Detection during Neoadjuvant Chemotherapy in Triple-Negative Breast Cancer. Clin Chem. 2017; 63:691–699. https://doi.org/10.1373/clinchem.2016.262337. [PubMed].
68. Garcia-Murillas I, Schiavon G, Weigelt B, Ng C, Hrebien S, Cutts RJ, Cheang M, Osin P, Nerurkar A, Kozarewa I, Garrido JA, Dowsett M, Reis-Filho JS, et al. Mutation tracking in circulating tumor DNA predicts relapse in early breast cancer. Sci Transl Med. 2015; 7:302ra133. https://doi.org/10.1126/scitranslmed.aab0021. [PubMed].
69. Visvanathan K, Fackler MS, Zhang Z, Lopez-Bujanda ZA, Jeter SC, Sokoll LJ, Garrett-Mayer E, Cope LM, Umbricht CB, Euhus DM, Forero A, Storniolo AM, Nanda R, et al. Monitoring of Serum DNA Methylation as an Early Independent Marker of Response and Survival in Metastatic Breast Cancer: TBCRC 005 Prospective Biomarker Study. J Clin Oncol. 2017; 35:751–758. https://doi.org/10.1200/JCO.2015.66.2080. [PubMed].
70. Takahashi H, Kagara N, Tanei T, Naoi Y, Shimoda M, Shimomura A, Shimazu K, Kim SJ, Noguchi S. Correlation of Methylated Circulating Tumor DNA With Response to Neoadjuvant Chemotherapy in Breast Cancer Patients. Clin Breast Cancer. 2017; 17:61–69.e63. https://doi.org/10.1016/j.clbc.2016.06.006. [PubMed].
71. Shan M, Yin H, Li J, Li X, Wang D, Su Y, Niu M, Zhong Z, Wang J, Zhang X, Kang W, Pang D. Detection of aberrant methylation of a six-gene panel in serum DNA for diagnosis of breast cancer. Oncotarget. 2016; 7:18485–18494. https://doi.org/10.18632/oncotarget.7608. [PubMed].
72. Fribbens C, Garcia Murillas I, Beaney M, Hrebien S, O’Leary B, Kilburn L, Howarth K, Epstein M, Green E, Rosenfeld N, Ring A, Johnston S, Turner N. Tracking evolution of aromatase inhibitor resistance with circulating tumour DNA analysis in metastatic breast cancer. Ann Oncol. 2018; 29:145–153. https://doi.org/10.1093/annonc/mdx483. [PubMed].
73. Wang P, Bahreini A, Gyanchandani R, Lucas PC, Hartmaier RJ, Watters RJ, Jonnalagadda AR, Trejo Bittar HE, Berg A, Hamilton RL, Kurland BF, Weiss KR, Mathew A, et al. Sensitive Detection of Mono- and Polyclonal ESR1 Mutations in Primary Tumors, Metastatic Lesions, and Cell-Free DNA of Breast Cancer Patients. Clin Cancer Res. 2016; 22:1130–1137. https://doi.org/10.1158/1078-0432.CCR-15-1534. [PubMed].
74. Steffensen KD, Madsen CV, Andersen RF, Waldstrom M, Adimi P, Jakobsen A. Prognostic importance of cell-free DNA in chemotherapy resistant ovarian cancer treated with bevacizumab. Eur J Cancer. 2014; 50:2611–2618. https://doi.org/10.1016/j.ejca.2014.06.022. [PubMed].
75. Parkinson CA, Gale D, Piskorz AM, Biggs H, Hodgkin C, Addley H, Freeman S, Moyle P, Sala E, Sayal K, Hosking K, Gounaris I, Jimenez-Linan M, et al. Exploratory Analysis of TP53 Mutations in Circulating Tumour DNA as Biomarkers of Treatment Response for Patients with Relapsed High-Grade Serous Ovarian Carcinoma: A Retrospective Study. PLoS Med. 2016; 13:e1002198. https://doi.org/10.1371/journal.pmed.1002198. [PubMed].
76. Pereira E, Camacho-Vanegas O, Anand S, Sebra R, Catalina Camacho S, Garnar-Wortzel L, Nair N, Moshier E, Wooten M, Uzilov A, Chen R, Prasad-Hayes M, Zakashansky K, et al. Personalized Circulating Tumor DNA Biomarkers Dynamically Predict Treatment Response and Survival In Gynecologic Cancers. PLoS One. 2015; 10:e0145754. https://doi.org/10.1371/journal.pone.0145754. [PubMed].
77. Vanderstichele A, Busschaert P, Smeets D, Landolfo C, Van Nieuwenhuysen E, Leunen K, Neven P, Amant F, Mahner S, Braicu EI, Zeilinger R, Coosemans A, Timmerman D, et al. Chromosomal Instability in Cell-Free DNA as a Highly Specific Biomarker for Detection of Ovarian Cancer in Women with Adnexal Masses. Clin Cancer Res. 2017; 23:2223–2231. https://doi.org/10.1158/1078-0432.CCR-16-1078. [PubMed].
78. Campitelli M, Jeannot E, Peter M, Lappartient E, Saada S, de la Rochefordiere A, Fourchotte V, Alran S, Petrow P, Cottu P, Pierga JY, Lantz O, Couturier J, et al. Human papillomavirus mutational insertion: specific marker of circulating tumor DNA in cervical cancer patients. PLoS One. 2012; 7:e43393. https://doi.org/10.1371/journal.pone.0043393. [PubMed].
79. Kang Z, Stevanovic S, Hinrichs CS, Cao L. Circulating Cell-free DNA for Metastatic Cervical Cancer Detection, Genotyping, and Monitoring. Clin Cancer Res. 2017; 23:6856–6862. https://doi.org/10.1158/1078-0432.CCR-17-1553. [PubMed].
80. Annala M, Struss WJ, Warner EW, Beja K, Vandekerkhove G, Wong A, Khalaf D, Seppala IL, So A, Lo G, Aggarwal R, Small EJ, Nykter M, et al. Treatment Outcomes and Tumor Loss of Heterozygosity in Germline DNA Repair-deficient Prostate Cancer. Eur Urol. 2017; 72:34–42. https://doi.org/10.1016/j.eururo.2017.02.023. [PubMed].
81. Wyatt AW, Azad AA, Volik SV, Annala M, Beja K, McConeghy B, Haegert A, Warner EW, Mo F, Brahmbhatt S, Shukin R, Le Bihan S, Gleave ME, et al. Genomic Alterations in Cell-Free DNA and Enzalutamide Resistance in Castration-Resistant Prostate Cancer. JAMA Oncol. 2016; 2:1598–1606. https://doi.org/10.1001/jamaoncol.2016.0494. [PubMed].
82. Azad AA, Volik SV, Wyatt AW, Haegert A, Le Bihan S, Bell RH, Anderson SA, McConeghy B, Shukin R, Bazov J, Youngren J, Paris P, Thomas G, et al. Androgen Receptor Gene Aberrations in Circulating Cell-Free DNA: Biomarkers of Therapeutic Resistance in Castration-Resistant Prostate Cancer. Clin Cancer Res. 2015; 21:2315–2324. https://doi.org/10.1158/1078-0432.CCR-14-2666. [PubMed].
83. Romanel A, Gasi Tandefelt D, Conteduca V, Jayaram A, Casiraghi N, Wetterskog D, Salvi S, Amadori D, Zafeiriou Z, Rescigno P, Bianchini D, Gurioli G, Casadio V, et al. Plasma AR and abiraterone-resistant prostate cancer. Sci Transl Med. 2015; 7:312re310. https://doi.org/10.1126/scitranslmed.aac9511. [PubMed].
84. Salvi S, Casadio V, Conteduca V, Lolli C, Gurioli G, Martignano F, Schepisi G, Testoni S, Scarpi E, Amadori D, Calistri D, Attard G, De Giorgi U. Circulating AR copy number and outcome to enzalutamide in docetaxel-treated metastatic castration-resistant prostate cancer. Oncotarget. 2016; 7:37839–37845. https://doi.org/10.18632/oncotarget.9341. [PubMed].
85. Sonpavde G, Nagy RJ, Sartor AO, Pond GR, Gourdin TS, Nandagopal L, Ledet EM, Agarwal N, Carroll E, Naik G, Wang J, Bilen MA, Grivas P, et al. Circulating tumor (ct)-DNA alterations in metastatic castration-resistant prostate cancer (mCRPC): Association with outcomes and evolution with therapy. J Clin Oncol. 2017; 35:1. https://doi.org/10.1200/JCO.2017.35.6_suppl.149.
86. de Martino M, Klatte T, Haitel A, Marberger M. Serum cell-free DNA in renal cell carcinoma: a diagnostic and prognostic marker. Cancer. 2012; 118:82–90. https://doi.org/10.1002/cncr.26254. [PubMed].
87. Pal SK, Sonpavde G, Agarwal N, Vogelzang NJ, Srinivas S, Haas NB, Signoretti S, McGregor BA, Jones J, Lanman RB, Banks KC, Choueiri TK. Evolution of Circulating Tumor DNA Profile from First-line to Subsequent Therapy in Metastatic Renal Cell Carcinoma. Eur Urol. 2017; 72:557–564. https://doi.org/10.1016/j.eururo.2017.03.046. [PubMed].
88. Sonpavde G, Nagy RJ, Apolo N, Pal S, Grivas P, Vaishampayan U, Lanman R, Talasaz A. Circulating cell-free DNA profiling of patients with advanced urothelial carcinoma. J Clin Oncol. 2016; 34. https://doi.org/10.1200/jco.2016.34.2_suppl.358.
89. Agarwal N, Pal SK, Hahn AW, Nussenzveig RH, Pond GR, Gupta SV, Wang J, Bilen MA, Naik G, Ghatalia P, Hoimes CJ, Gopalakrishnan D, Barata PC, et al. Characterization of metastatic urothelial carcinoma via comprehensive genomic profiling of circulating tumor DNA. Cancer. 2018; 124:2115–2124. https://doi.org/10.1002/cncr.31314. [PubMed].
90. McGregor BA, Chung J, Bergerot PG, Forcier B, Grivas P, Choueiri TK, Ross JS, Ali SM, Stephens PJ, Miller VA, Gregg JP, MacVicar GR, Bauer TM, Pal SK. Correlation of circulating tumor DNA (ctDNA) ssessment with tissue-based comprehensive genomic profiling (CGP) in metastatic urothelial cancer (MUC). J Clin Oncol. 2018; 36. https://doi.org/10.1200/jco.2018.36.6_suppl.453.
91. Christensen E, Birkenkamp-Demtroder K, Nordentoft I, Hoyer S, van der Keur K, van Kessel K, Zwarthoff E, Agerbaek M, Orntoft TF, Jensen JB, Dyrskjot L. Liquid Biopsy Analysis of FGFR3 and PIK3CA Hotspot Mutations for Disease Surveillance in Bladder Cancer. Eur Urol. 2017; 71:961–969. https://doi.org/10.1016/j.eururo.2016.12.016. [PubMed].
92. Chang GA, Tadepalli JS, Shao Y, Zhang Y, Weiss S, Robinson E, Spittle C, Furtado M, Shelton DN, Karlin-Neumann G, Pavlick A, Osman I, Polsky D. Sensitivity of plasma BRAFmutant and NRASmutant cell-free DNA assays to detect metastatic melanoma in patients with low RECIST scores and non-RECIST disease progression. Mol Oncol. 2016; 10:157–165. https://doi.org/10.1016/j.molonc.2015.09.005. [PubMed].
93. Knol AC, Vallee A, Herbreteau G, Nguyen JM, Varey E, Gaultier A, Theoleyre S, Saint-Jean M, Peuvrel L, Brocard A, Quereux G, Khammari A, Denis MG, et al. Clinical significance of BRAF mutation status in circulating tumor DNA of metastatic melanoma patients at baseline. Exp Dermatol. 2016; 25:783–788. https://doi.org/10.1111/exd.13065. [PubMed].
94. Gray ES, Rizos H, Reid AL, Boyd SC, Pereira MR, Lo J, Tembe V, Freeman J, Lee JH, Scolyer RA, Siew K, Lomma C, Cooper A, et al. Circulating tumor DNA to monitor treatment response and detect acquired resistance in patients with metastatic melanoma. Oncotarget. 2015; 6:42008–42018. https://doi.org/10.18632/oncotarget.5788. [PubMed].
95. Gonzalez-Cao M, Mayo-de-Las-Casas C, Molina-Vila MA, De Mattos-Arruda L, Munoz-Couselo E, Manzano JL, Cortes J, Berros JP, Drozdowskyj A, Sanmamed M, Gonzalez A, Alvarez C, Viteri S, et al. BRAF mutation analysis in circulating free tumor DNA of melanoma patients treated with BRAF inhibitors. Melanoma Res. 2015; 25:486–495. https://doi.org/10.1097/CMR.0000000000000187. [PubMed].
96. Santiago-Walker A, Gagnon R, Mazumdar J, Casey M, Long GV, Schadendorf D, Flaherty K, Kefford R, Hauschild A, Hwu P, Haney P, O’Hagan A, Carver J, et al. Correlation of BRAF Mutation Status in Circulating-Free DNA and Tumor and Association with Clinical Outcome across Four BRAFi and MEKi Clinical Trials. Clin Cancer Res. 2016; 22:567–574. https://doi.org/10.1158/1078-0432.CCR-15-0321. [PubMed].
97. Schreuer M, Meersseman G, Van Den Herrewegen S, Jansen Y, Chevolet I, Bott A, Wilgenhof S, Seremet T, Jacobs B, Buyl R, Maertens G, Neyns B. Quantitative assessment of BRAF V600 mutant circulating cell-free tumor DNA as a tool for therapeutic monitoring in metastatic melanoma patients treated with BRAF/MEK inhibitors. J Transl Med. 2016; 14:95. https://doi.org/10.1186/s12967-016-0852-6. [PubMed].
98. Wong SQ, Raleigh JM, Callahan J, Vergara IA, Ftouni S, Hatzimihalis A, Colebatch AJ, Li J, Semple T, Doig K, Mintoff C, Sinha D, Yeh P, et al. Circulating Tumor DNA Analysis and Functional Imaging Provide Complementary Approaches for Comprehensive Disease Monitoring in Metastatic Melanoma. JCO Precision Oncology. 2017:1–14. https://doi.org/10.1200/po.16.00009.
99. Lee RJ, Gremel G, Marshall A, Myers KA, Fisher N, Dunn JA, Dhomen N, Corrie PG, Middleton MR, Lorigan P, Marais R. Circulating tumor DNA predicts survival in patients with resected high-risk stage II/III melanoma. Ann Oncol. 2018; 29:490–496. https://doi.org/10.1093/annonc/mdx717. [PubMed].
100. Lee JH, Long GV, Menzies AM, Lo S, Guminski A, Whitbourne K, Peranec M, Scolyer R, Kefford RF, Rizos H, Carlino MS. Association Between Circulating Tumor DNA and Pseudoprogression in Patients With Metastatic Melanoma Treated With Anti-Programmed Cell Death 1 Antibodies. JAMA Oncol. 2018; 4:717–721. https://doi.org/10.1001/jamaoncol.2017.5332. [PubMed].
101. Krumbholz M, Hellberg J, Steif B, Bauerle T, Gillmann C, Fritscher T, Agaimy A, Frey B, Juengert J, Wardelmann E, Hartmann W, Juergens H, Dirksen U, et al. Genomic EWSR1 Fusion Sequence as Highly Sensitive and Dynamic Plasma Tumor Marker in Ewing Sarcoma. Clin Cancer Res. 2016; 22:4356–4365. https://doi.org/10.1158/1078-0432.CCR-15-3028. [PubMed].
102. Hayashi M, Chu D, Meyer CF, Llosa NJ, McCarty G, Morris CD, Levin AS, Wolinsky JP, Albert CM, Steppan DA, Park BH, Loeb DM. Highly personalized detection of minimal Ewing sarcoma disease burden from plasma tumor DNA. Cancer. 2016; 122:3015–3023. https://doi.org/10.1002/cncr.30144. [PubMed].
103. Shulman DS, Klega K, Imamovic-Tuco A, Clapp A, Nag A, Thorner AR, Van Allen E, Ha G, Lessnick SL, Gorlick R, Janeway KA, Leavey PJ, Mascarenhas L, et al. Detection of circulating tumour DNA is associated with inferior outcomes in Ewing sarcoma and osteosarcoma: a report from the Children’s Oncology Group. Br J Cancer. 2018; 119:615–621. https://doi.org/10.1038/s41416-018-0212-9. [PubMed].
104. Maier J, Lange T, Kerle I, Specht K, Bruegel M, Wickenhauser C, Jost P, Niederwieser D, Peschel C, Duyster J, von Bubnoff N. Detection of mutant free circulating tumor DNA in the plasma of patients with gastrointestinal stromal tumor harboring activating mutations of CKIT or PDGFRA. Clin Cancer Res. 2013; 19:4854–4867. https://doi.org/10.1158/1078-0432.CCR-13-0765. [PubMed].
105. Yoo C, Ryu MH, Na YS, Ryoo BY, Park SR, Kang YK. Analysis of serum protein biomarkers, circulating tumor DNA, and dovitinib activity in patients with tyrosine kinase inhibitor-refractory gastrointestinal stromal tumors. Ann Oncol. 2014; 25:2272–2277. https://doi.org/10.1093/annonc/mdu386. [PubMed].
106. Wada N, Kurokawa Y, Takahashi T, Hamakawa T, Hirota S, Naka T, Miyazaki Y, Makino T, Yamasaki M, Nakajima K, Takiguchi S, Mori M, Doki Y. Detecting Secondary C-KIT Mutations in the Peripheral Blood of Patients with Imatinib-Resistant Gastrointestinal Stromal Tumor. Oncology. 2016; 90:112–117. https://doi.org/10.1159/000442948. [PubMed].
107. Eastley NC, Ottolini B, Neumann R, Luo JL, Hastings RK, Khan I, Moore DA, Esler CP, Shaw JA, Royle NJ, Ashford RU. Circulating tumour-derived DNA in metastatic soft tissue sarcoma. Oncotarget. 2018; 9:10549–10560. https://doi.org/10.18632/oncotarget.24278. [PubMed].
108. Salkeni MA, Zarzour A, Ansay TY, McPherson CM, Warnick RE, Rixe O. Detection of EGFRvIII mutant DNA in the peripheral blood of brain tumor patients. J Neurooncol. 2013; 115:27–35. https://doi.org/10.1007/s11060-013-1209-0. [PubMed].
109. Yagyu S, Iehara T, Tanaka S, Gotoh T, Misawa-Furihata A, Sugimoto T, London WB, Hogarty MD, Teramukai S, Nakagawara A, Hiyama E, Maris JM, Hosoi H. Serum-Based Quantification of MYCN Gene Amplification in Young Patients with Neuroblastoma: Potential Utility as a Surrogate Biomarker for Neuroblastoma. PLoS One. 2016; 11:e0161039. https://doi.org/10.1371/journal.pone.0161039. [PubMed].
110. Boisselier B, Gallego Perez-Larraya J, Rossetto M, Labussiere M, Ciccarino P, Marie Y, Delattre JY, Sanson M. Detection of IDH1 mutation in the plasma of patients with glioma. Neurology. 2012; 79:1693–1698. https://doi.org/10.1212/WNL.0b013e31826e9b0a. [PubMed].
111. Camus V, Stamatoullas A, Mareschal S, Viailly PJ, Sarafan-Vasseur N, Bohers E, Dubois S, Picquenot JM, Ruminy P, Maingonnat C, Bertrand P, Cornic M, Tallon-Simon V, et al. Detection and prognostic value of recurrent exportin 1 mutations in tumor and cell-free circulating DNA of patients with classical Hodgkin lymphoma. Haematologica. 2016; 101:1094–1101. https://doi.org/10.3324/haematol.2016.145102. [PubMed].
112. Assouline SE, Nielsen TH, Yu S, Alcaide M, Chong L, MacDonald D, Tosikyan A, Kukreti V, Kezouh A, Petrogiannis-Haliotis T, Albuquerque M, Fornika D, Alamouti S, et al. Phase 2 study of panobinostat with or without rituximab in relapsed diffuse large B-cell lymphoma. Blood. 2016; 128:185–194. https://doi.org/10.1182/blood-2016-02-699520. [PubMed].
113. Scherer F, Kurtz DM, Newman AM, Stehr H, Craig AF, Esfahani MS, Lovejoy AF, Chabon JJ, Klass DM, Liu CL, Zhou L, Glover C, Visser BC, et al. Distinct biological subtypes and patterns of genome evolution in lymphoma revealed by circulating tumor DNA. Sci Transl Med. 2016; 8:364ra155. https://doi.org/10.1126/scitranslmed.aai8545. [PubMed].
114. Kurtz DM, Green MR, Bratman SV, Scherer F, Liu CL, Kunder CA, Takahashi K, Glover C, Keane C, Kihira S, Visser B, Callahan J, Kong KA, et al. Noninvasive monitoring of diffuse large B-cell lymphoma by immunoglobulin high-throughput sequencing. Blood. 2015; 125:3679–3687. https://doi.org/10.1182/blood-2015-03-635169. [PubMed].
115. Roschewski M, Dunleavy K, Pittaluga S, Moorhead M, Pepin F, Kong K, Shovlin M, Jaffe ES, Staudt LM, Lai C, Steinberg SM, Chen CC, Zheng J, et al. Circulating tumour DNA and CT monitoring in patients with untreated diffuse large B-cell lymphoma: a correlative biomarker study. Lancet Oncol. 2015; 16:541–549. https://doi.org/10.1016/S1470-2045(15)70106-3. [PubMed].
116. Fontanilles M, Marguet F, Bohers E, Viailly PJ, Bertrand P, Dubois S, Mareschal S, Camus S, Veresezan EL, Ruminy P, Tilly H, Laquerriere A, Picquenot JM, Jardin F. Somatic Mutations Detected in Plasma Cell-Free DNA By Targeted Sequencing: Assessment of Liquid Biopsy in Primary Central Nervous System Lymphoma. Blood. 2015; 126:332. https://doi.org/10.1182/blood.v126.23.332.332.
117. Perkins G, Yap TA, Pope L, Cassidy AM, Dukes JP, Riisnaes R, Massard C, Cassier PA, Miranda S, Clark J, Denholm KA, Thway K, Gonzalez De Castro D, et al. Multi-purpose utility of circulating plasma DNA testing in patients with advanced cancers. PLoS One. 2012; 7:e47020. https://doi.org/10.1371/journal.pone.0047020. [PubMed].
118. Janku F, Angenendt P, Tsimberidou AM, Fu S, Naing A, Falchook GS, Hong DS, Holley VR, Cabrilo G, Wheler JJ, Piha-Paul SA, Zinner RG, Bedikian AY, et al. Actionable mutations in plasma cell-free DNA in patients with advanced cancers referred for experimental targeted therapies. Oncotarget. 2015; 6:12809–12821. https://doi.org/10.18632/oncotarget.3373. [PubMed].
119. Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, Bartlett BR, Wang H, Luber B, Alani RM, Antonarakis ES, Azad NS, Bardelli A, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014; 6:224ra224. https://doi.org/10.1126/scitranslmed.3007094. [PubMed].
120. Murtaza M, Dawson SJ, Tsui DW, Gale D, Forshew T, Piskorz AM, Parkinson C, Chin SF, Kingsbury Z, Wong AS, Marass F, Humphray S, Hadfield J, et al. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature. 2013; 497:108–112. https://doi.org/10.1038/nature12065. [PubMed].
121. Frenel JS, Carreira S, Goodall J, Roda D, Perez-Lopez R, Tunariu N, Riisnaes R, Miranda S, Figueiredo I, Nava-Rodrigues D, Smith A, Leux C, Garcia-Murillas I, et al. Serial Next-Generation Sequencing of Circulating Cell-Free DNA Evaluating Tumor Clone Response To Molecularly Targeted Drug Administration. Clin Cancer Res. 2015; 21:4586–4596. https://doi.org/10.1158/1078-0432.CCR-15-0584. [PubMed].
122. Schwaederle M, Husain H, Fanta PT, Piccioni DE, Kesari S, Schwab RB, Banks KC, Lanman RB, Talasaz A, Parker BA, Kurzrock R. Detection rate of actionable mutations in diverse cancers using a biopsy-free (blood) circulating tumor cell DNA assay. Oncotarget. 2016; 7:9707–9717. https://doi.org/10.18632/oncotarget.7110. [PubMed].
123. Schwaederle M, Husain H, Fanta PT, Piccioni DE, Kesari S, Schwab RB, Patel SP, Harismendy O, Ikeda M, Parker BA, Kurzrock R. Use of Liquid Biopsies in Clinical Oncology: Pilot Experience in 168 Patients. Clin Cancer Res. 2016; 22:5497–5505. https://doi.org/10.1158/1078-0432.CCR-16-0318. [PubMed].
124. Khagi Y, Goodman AM, Daniels GA, Patel SP, Sacco AG, Randall JM, Bazhenova LA, Kurzrock R. Hypermutated Circulating Tumor DNA: Correlation with Response to Checkpoint Inhibitor-Based Immunotherapy. Clin Cancer Res. 2017; 23:5729–5736. https://doi.org/10.1158/1078-0432.CCR-17-1439. [PubMed].
125. Ocana A, Diez-Gonzalez L, Garcia-Olmo DC, Templeton AJ, Vera-Badillo F, Jose Escribano M, Serrano-Heras G, Corrales-Sanchez V, Seruga B, Andres-Pretel F, Pandiella A, Amir E. Circulating DNA and Survival in Solid Tumors. Cancer Epidemiol Biomarkers Prev. 2016; 25:399–406. https://doi.org/10.1158/1055-9965.EPI-15-0893. [PubMed].

