Introduction
Target therapy has revolutionized the oncological approach to cancer patients carrying druggable molecular alterations [1]. New generation targeted therapies are increasingly effective and with fewer side effects, so no patient should be left behind without an appropriated treatment. A correct target treatment implies the identification of specific biomarkers alterations, as reported in dedicated guidelines [2–4].
Some of the biomarkers for target therapy are frequent events in particular tumor types (eg RAS mutations in colorectal cancer or BRAF mutations in melanoma). In these cases, accurate identification of the molecular alteration with dedicated methods is feasible and the cost and time of analysis to find a positive patient is acceptable [3, 4].
On the other hand, several biomarkers are rare or extremely rare events, while remaining valid to select cancer patients for very effective treatments. In practical terms, rare alterations may be defined as those present in less than 5% of patients. Within these alterations, important examples are ALK1 and ROS1 fusions, present in 3-5% and 1-2% of lung tumors, respectively, as well as in many other tumor types at lower prevalence rates [5–7]. The detection of rare mutations with a mono-marker test implies long time frames and high costs to identify a positive/druggable patient. Consider that the cost per positive test (CPT) is inversely related to the prevalence of the genomic alteration, as reported in the equation in Figure 1A.
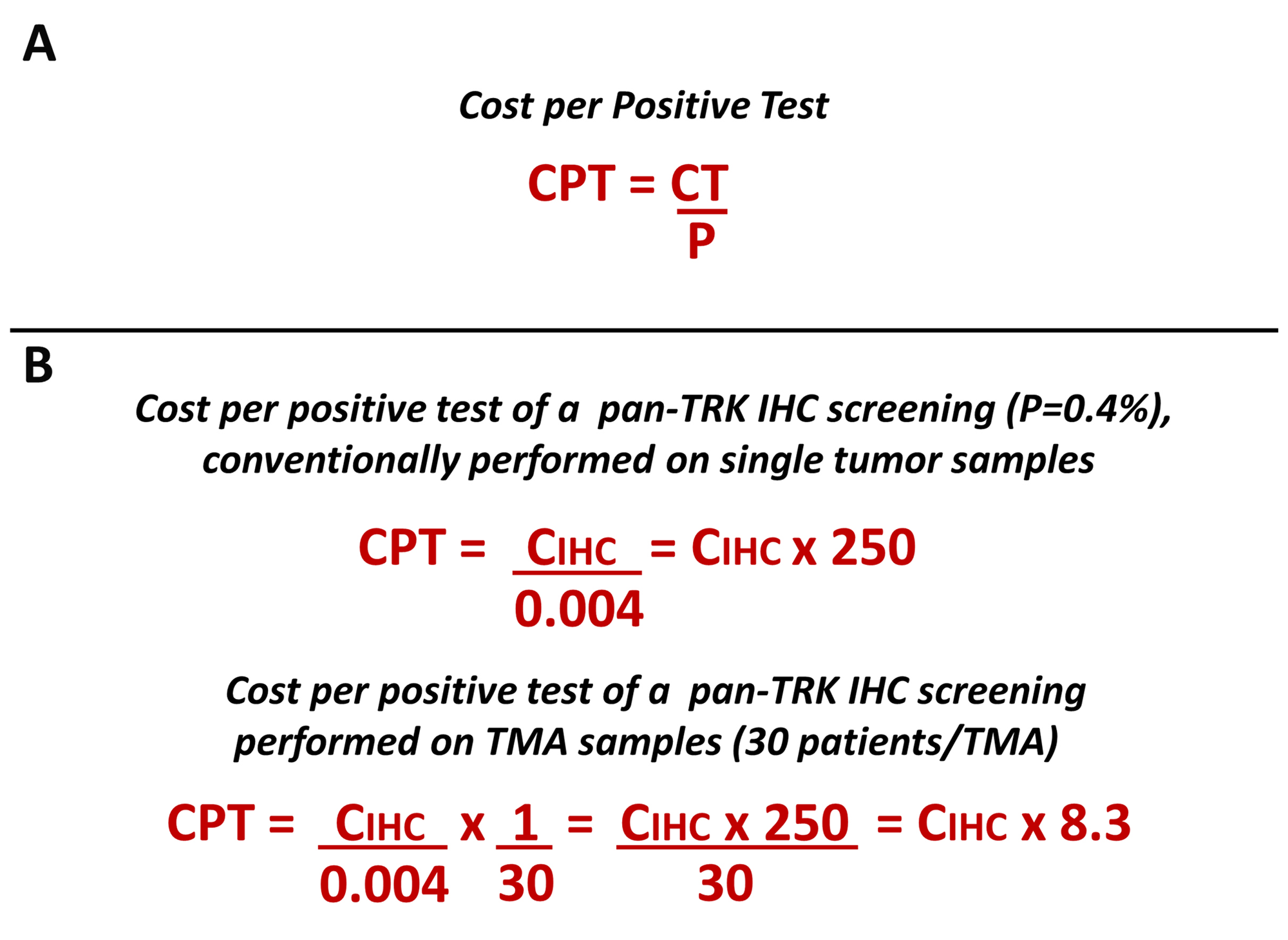
Figure 1: (A) The figure reports the equation to calculate the cost per positive test. CPT, cost per positive test; CT, cost per single test; P, prevalence of biomarker alteration. (B) The equation has been applied to calculate the cost of the pan-TRK IHC assay (CIHC) as an example of a test for the detection of a rare mutation (see the results section for furter details).
Moreover, a mass of data produced by next generation sequencing in the last years indicate that some biomarkers are no longer restricted to specific tumor types, leading to histology agnostic treatments [8, 9]. This new therapeutic vision requires the analysis of molecular targets in many different tumor types if not in all, as in the case of the rare alterations affecting mismatch repair and NTRK genes [10, 11].
Two approaches are possible to meet these new diagnostic needs: 1) screening with methods applicable on a large scale as IHC followed by orthogonal tests (FISH, RT-PCR, Next generation sequencing) to confirm the alterations identified; 2) a direct and extended approach to all tumors through massive parallel sequencing. However, even a simple screening test, if extended to all currently needed biomarkers in clinical practice, and to all neoplastic forms, is not practical as it would have unacceptable timing and costs.
On the other hand, a large-scale NGS approach with large gene panels is desirable but at the moment, the costs and the low diffusion of the technology make it not realistically feasible [12].
Driven by these management difficulties, we have developed a diagnostic strategy based on large scale IHC screening of rare molecular alterations on tissue microarrays (TMAs) and Tissue Slice Arrays (TSAs) (see further text) to select cancer patients for histology-agnostic therapies. The approach has been finely tuned in order to meet the diagnostic needs of a standard pathology laboratory.
Results
A diagnostic strategy for the detection of rare molecular targets to select cancer patients for histology-agnostic treatments has been devised, as described in detail in the Material and Methods section. The innovative workflow provides that malignant tumor samples are identified by histological examination and subdivided into large and small samples based on size, regardless of the type of malignancy, for further processing. Very small samples, with inadequate amount of tissue for molecular diagnosis are excluded.
In a retrospective series, corresponding to a standard six months routine activity in our department (Table 1), malignant tumors (total n. 1865) accounted for about 15% of cases submitted for histopathological diagnosis and 311 malignant samples were collected each month, of which 146 (47%) were large samples, having at least one fragment above 0.5 cm2, and the remaining 165 (53%) were small samples.
Table 1: Retrospective series of different malignant tumors divided according to the size of embedded samples
| Malignant Tumor | Samples | Total | |
|---|---|---|---|
| small | large | ||
| Bladder | 111 | 18 | 129 |
| Prostate | 87 | 90 | 177 |
| Kidney | 0 | 27 | 27 |
| Testis | 0 | 9 | 9 |
| Penis | 3 | 0 | 3 |
| Gynecopathological | 87 | 114 | 201 |
| Gastrointestinal and hepatic | 78 | 81 | 159 |
| Pancreas | 3 | 15 | 18 |
| Lung | 77 | 55 | 132 |
| Thyroid | 0 | 22 | 22 |
| Salivary | 0 | 10 | 10 |
| Lymph Node | 18 | 60 | 78 |
| Oral cavity | 27 | 9 | 36 |
| Breast | 345 | 366 | 711 |
| Skin | 153 | 0 | 153 |
| Total (%) | 989 (53) | 876 (47) | 1865 (100) |
| Total per month | 165 | 146 | 311 |
Large samples were selected for the construction of TMA blocks using 2 (2mm) cores per tumor. The reliability of IHC staining for the expression of driver mutations performed on 2 (2mm) TMA cores versus the original tissue sections was investigated on a series of 60 selected blocks from lung neoplastic lesions including 23 blocks carrying ALK gene fusion, 12 with ROS1 gene fusion, 3 with NTRK1 gene fusion, and a series of 12 blocks from selected colorectal carcinomas with absence of mismatch repair protein expression in 7 cases. Two (2 mm) cores were representative of the tumor immunophenotypic profiling in 100% of cases with concordant data in the two cores of each patient (data not shown).
It is advisable not to process tissues samples below 0.5 cm2 for TMA construction due to the paucity of biological material available. In these cases, in order to reduce the time and costs for the screening of rare molecular targets, we decided to proceed with the preparation of TSAs, an innovative approach that allows to maintain the integrity of tissue blocks, while ensuring the possibility of simultaneously examining multiple samples (Figure 2), as detailed in Materials and Methods.
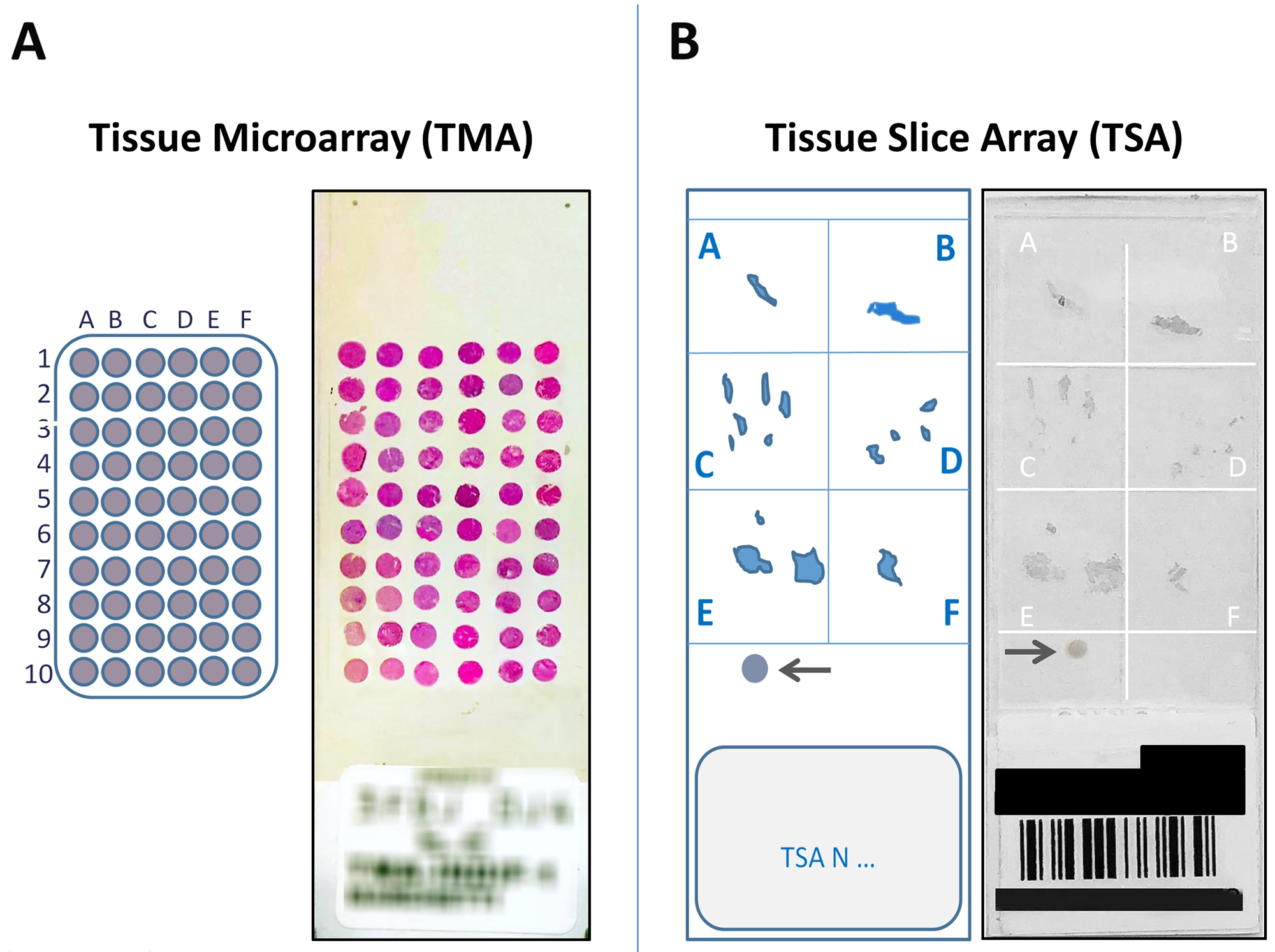
Figure 2: (A) The figure shows a 5 μm section from a 60 cores (30 tumors) Tissue MicroArray (TMA), obtained from large samples, stained with Hematoxylin–eosin, corresponding scheme on the left. (B) An example of a Tissue Slice Array (TSA), obtained from small samples, immunostained with the VENTANA pan-TRK (EPR17341) assay, corresponding scheme on the left. The arrows indicate an internal positive control.
Five micrometers TMA sections and TSAs were screened by IHC to evaluate mismatch repair and TRK A, B, C proteins expression. In case of neoplastic forms for which routine clinical molecular testing is planned (i.e non- small cell lung cancer, colorectal carcinoma, melanoma etc.), the search for rare biomarker alterations was conducted after the analysis of conventional predictive tests. Positive cases were subjected to confirmation orthogonal tests (FISH, NGS) for the detection of microsatellite instability and NTRK gene fusions.
Large-scale analysis of MSI and NTRK fusions allowed to detect 48 (2.6%) MSI cases and 6 (0.3%) NTRK fusion cases in a series of 1865 malignant tumor samples examined, as reported in Table 2. In particular, IHC screening of mismatch repair proteins revealed an absence of nuclear staining within tumor cells in 53 (2.8%) cases of which 48 (91%) showed MSI by molecular analysis. The IHC screening test for TRK (A,B,C) proteins was found to be positive in 8 (0.4%) tumors. In 6 (75%) of these 8 tumors a NTRK gene fusion was shown by FISH. Two (0.3%) of 711 patients with ductal infiltrating breast carcinomas were found to be positive for NTRK1 gene fusions (Figure 3) as well as a patient (0.8%) in a series of 132 cases with lung adenocarcinoma, 2 (1.3%) of 159 patients with gastrointestinal tumors, specifically a colorectal carcinoma and a gastrointestinal stromal tumor (GIST). In a series of 22 thyroid malignant tumors, a papillary thyroid carcinoma (4.5%) was found to be positive for a NTRK3 fusion.
Table 2: Incidence of MMR deficiency, micro satellite instability and Neurothropic Receptor Tyrosine Kinase fusions in solid tumors
| Malignant Tumor | Microsatellite Analysis | NTRK analysis | total | ||||
|---|---|---|---|---|---|---|---|
| dMMR | pMMR | MSI | MSS | Positive | Negative | ||
| Bladder | 1 (0.8) | 128 (99.2) | 1 (0.8) | 128 (99.2) | 0 | 129 (100) | 129 |
| Prostate | 1 (0.6) | 176 (99.4) | 1 (0.6) | 176 (99.4) | 0 | 177 (100) | 177 |
| Kidney | 1 (3.7) | 26 (96.3) | 1 (0.7) | 26 (96.3) | 0 | 27 (100) | 27 |
| Testis | 0 | 9 (100) | 0 | 9 (100) | 0 | 9 (100) | 9 |
| Penis | 0 | 3 (100) | 0 | 3 (100) | 0 | 3 (100) | 3 |
| Gynecopathological | 12 (6) | 189 (94) | 10 (5) | 191 (95) | 0 | 201 (100) | 201 |
| Gastrointestinal and hepatic | 23 (14.5) | 136 (85.5) | 21 (13.2) | 138 (86.8) | 2 (1.3) | 157 (98.7) | 159 |
| Pancreas | 1 (5.6) | 17 (94.4) | 1 (5.6) | 17 (94.4) | 0 | 18 (100) | 18 |
| Lung | 1 (0.8) | 131 (99.2) | 1 (0.8) | 131 (99.2) | 1 (0.8) | 131 (99.3) | 132 |
| Thyroid | 0 | 22 (100) | 0 | 22 (100) | 1 (4.5) | 21 (95.5) | 22 |
| Salivary | 0 | 10 (100) | 0 | 10 (100) | 0 | 10 (100) | 10 |
| Lymph Node | 0 | 78 (100) | 0 | 78 (100) | 0 | 78 (100) | 78 |
| Oral cavity | 0 | 36 (100) | 0 | 36 (100) | 0 | 36 (100) | 36 |
| Breast | 8 (1.1) | 703 (98.9) | 7 (1) | 704 (99) | 2 (0.3) | 709 (99.7) | 711 |
| Skin | 5 (3.3) | 148 (96.7) | 5 (3.3) | 148 (96.7) | 0 | 153 (100) | 153 |
| Total | 53 (2.8) | 1812 (97.2) | 48 (2.6) | 1817 (97.4) | 6 (0.3) | 1859 (99.7) | 1865 (100) |
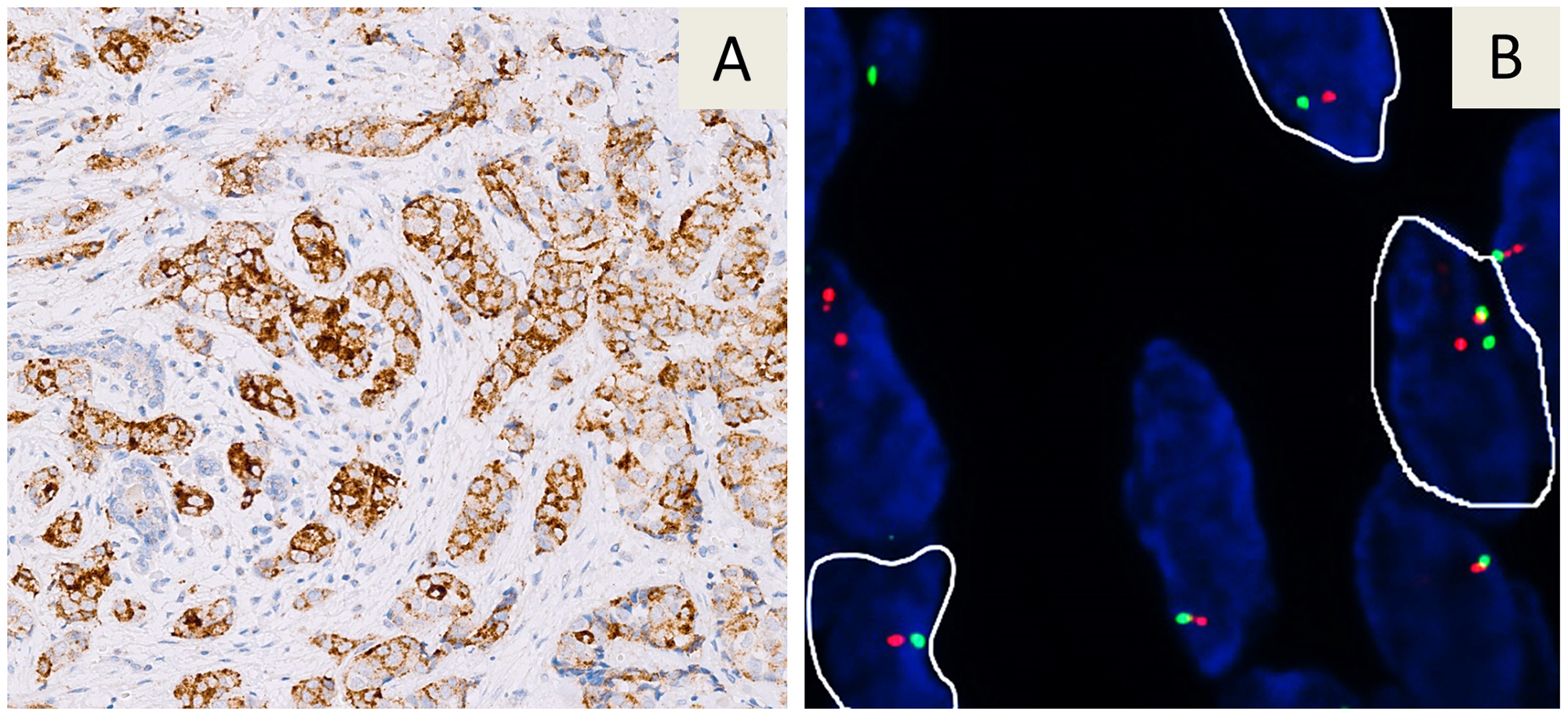
Figure 3: Immunohistochemical staining with the VENTANA pan-TRK (EPR17341) assay in a breast carcinoma. A strong cytoplasmic immunoreaction is evident (A). The tumor showed a NTRK1 gene fusion by FISH analysis with a Break Apart probe. Nuclei highlighted with white borders show split signals (B).
In 4 of these 6 cases, tissue was available for genomic assessment of the fusion patterns by NGS and/or multiplexing PCR/hybridization (Table 3). For two small samples included in the TSA series, NGS analysis could not be conducted due to the paucity of the starting material available. The TSA approach in these two cases has therefore allowed by IHC+FISH the identification of rare fusions otherwise undetectable by direct NGS analysis.
Table 3: TRK protein expression and NTRK gene fusions detected by immunohistochemistry, Fluorescence in situ hybridization and next generation sequencing in different tumor types
| Tumor type | Histotype | NTRK Analysis | ||
|---|---|---|---|---|
| IHC | FISH | NGS | ||
| Gastrointestinal | Colorectal | cytoplasmic | NTRK1 | NTRK1/TPM3 |
| GIST | cytoplasmic | NTRK1 | NTRK1/MPRIP | |
| Breast | Infiltrating ductal | cytoplasmic | NTRK1 | NTRK1/LMNA |
| Infiltrating ductal | perinuclear | NTRK1 | NA | |
| Lung | Adenocarcinoma | cytoplasmic | NTRK1 | NTRK1/CD74 |
| Thyroid | Papillary carcinoma | nuclear | NTRK3 | NA |
The diagnostic strategy described has enabled IHC analysis of up to 30 patients simultaneously per TMA and 6 patients simultaneously per TSA with a significant saving in terms of analytical costs and time. According to the equation reported in Figure 1B, it has been calculated that the analytical cost of IHC screening per TRK-positive patient on TMA samples was about 8 times the cost per single test and on TSA samples was about 42 times compared to 250 times if the IHC test had been carried on single samples. Using both TMA and TSA strategies an average 20 times saving was obtained.
The high throughput of the described diagnostic strategy can be further enhanced by multiplex IHC on TMA sections/TSAs. In order to verify the feasibility of this approach, a section of a TMA with a positive TRKA sample and a positive ALK1 sample was subjected to multiplex IHC which simultaneously showed the different biomarkers in different colours (Figure 4).
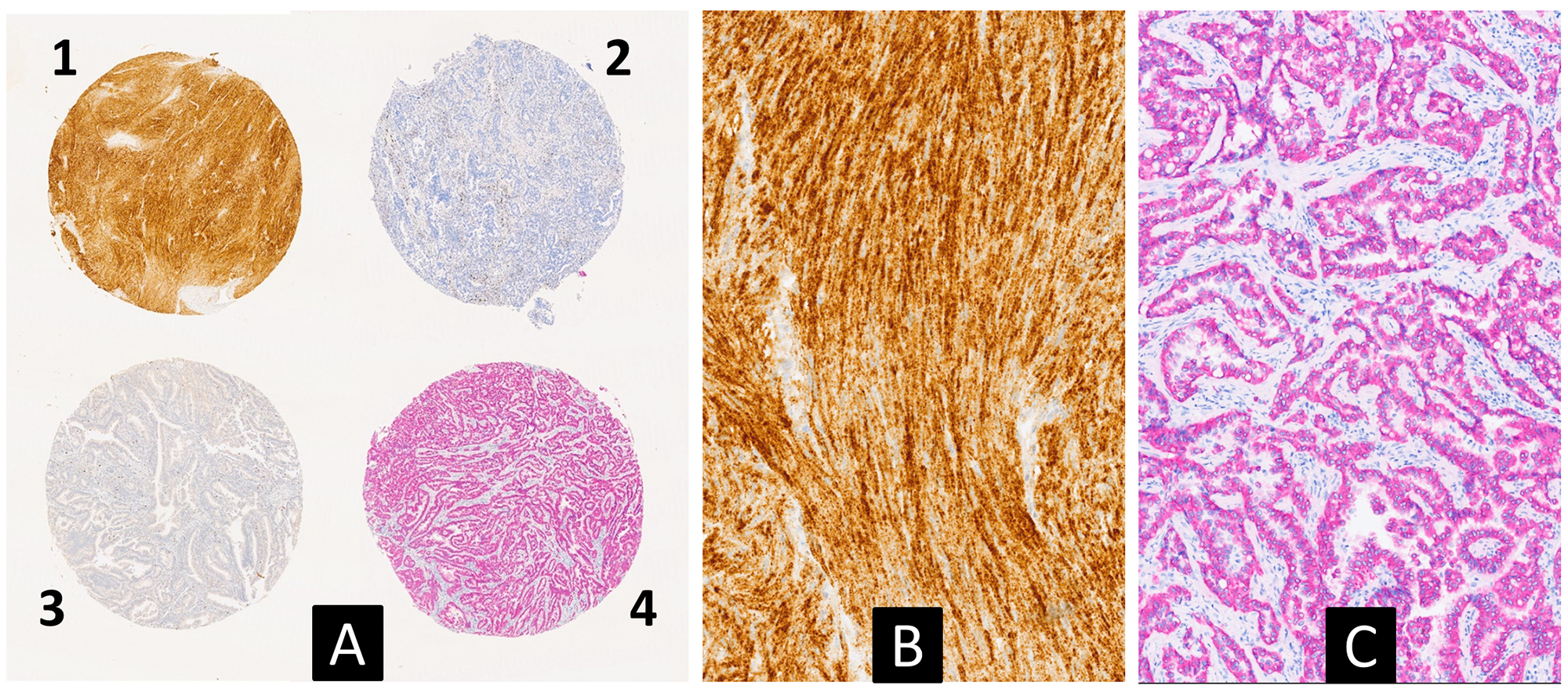
Figure 4: Multiplex immunohistochemical analysis for simultaneous detection of TRK (brown) and ALK1 (red) proteins on a multi-tumor TMA slide (A) (2.5×). Case 1 corresponds to a gastrointestinal stromal tumor, carring a NTRK1 fusion, with a strong TRKA cytoplasmic expression (see also Table 3). Case 4 corresponds to a lung adenocarcinoma with an EML4-ALK fusion showing a cytoplasmic expression of the ALK1 protein. (B and C) are higher magnification pictures of cases 1 and 4, respectively (10×).
DISCUSSION
Among the driver mutations which can be targeted with specific treatments in different forms of human malignancies, there are uncommon genomic alterations affecting a minority of cancer patients [13, 14]. The assessment of rare but clinically important molecular alterations represents a challenge that, due to problems concerning diagnostic costs and time, can only be tackled efficiently with multiple simultaneous analyses. Two potential approaches are currently available in this regard. A first possibility is to analyze single samples for multiple alterations, this is possible through a series of multimarker techniques including massive parallel sequencing [15, 16], a second possibility is to simultaneously evaluate multiple samples with mono-(oligo-)marker screening technologies While the analysis of multiple alterations on a single sample does not alter a conventional prospective approach, the analysis of multiple samples per single marker requires a retrospective approach repeated prospectively at time intervals compatible with clinical needs. In the present study we have shown the feasibility of an innovative diagnostic strategy for the simultaneous analysis of rare mutations in multiple samples, based on the use of TMAs and TSAs. This has allowed to identify a series of patients affected by uncommon tumor types which can efficiently be treated with specific target therapies.
In particular, the method has been applied to the screening of patients affected by malignant tumors, irrespective of the histological types, for the detection of microsatellite instability and NTRK genetic fusions. Both of these two genetic alterations nowadays allow histology agnostic treatments [17].
It has been shown that metastatic or unresectable MSI-H patients or mismatch repair deficient (dMMR) metastatic or unresectable patients or dMMR patients affected by solid tumors that have progressed following previous therapies and who have no satisfactory alternative treatment options could be successfully treated with pembrolizumab. This has led the approval of pembrolizumab (Keytruda®, Merck & Co) by FDA in 2017, for a solid tumor treatment regardless the tumor primary site of origin and histology [18–20].
More recently, FDA and EMA have also approved the oral agent larotrectinib (VITRAKVI®, Loxo Oncology Inc. and Bayer) for treating metastatic or unresectable solid tumors of any type with a NTRK gene fusion. Moreover, FDA has granted a priority review designation to Entrectinib (RXDX-101, Roche Genentech) for patients with NTRK fusion–positive locally advanced or metastatic solid tumors [21–26].
Overall, in a series of 1865 cases of malignant tumors analyzed in this study during a trial period of 6 months, 48 cases with microsatellite instability in 9 different tumor forms and 6 cases of fusions in the NTRK gene family in 4 tumor forms were detected.
The number of cases identified does not yet allow a systematic statistical study of the incidence of these rare mutations in all types of solid tumors. This will be one of the aims of a multicenter study under the aegis of the Italian Society of Pathology (SIAPEC) that foresees the diffusion of this diagnostic approach to a series of reference centers of pathology in Italy that could allow to select patients on a large scale and to have rapid information on incidences. Theoretically, on the basis of the data obtained in our pilot study, considering at minimum to have available tissue in 60% of all cancer patients, we have estimated that a widespread diffusion of the strategy in Pathology Centers could allow us to identify more than 6.000 cases/year in Italy (26.000 cases/year in USA) with microsatellite alterations and more than 1000 cases/year in Italy (4500 cases/year in USA) with fusions of NTRK genes with consequent possible treatment of patients with effective drugs.
Other potential applications of this pathological strategy in oncology concern the IHC screening of several other genomic alterations including ALK1 and ROS1 fusions and BRAF-(V600E) mutations which may be drivers of various neoplastic forms, regardless of the tumor type, and could be treated in a near future with drugs already available for treatment of lung adenocarcinomas (ALK1 and ROS1) and melanoma and lung carcinoma (BRAF) patients [2, 3, 27].
This diagnostic approach can potentially be applied to other clinical fields, even non-neoplastic, whenever it is necessary to identify a rare marker for which a rapid tissue screening method is available, such as IHC or FISH.
TMAs were first introduced in 1998, by J. Kononen and collaborators and since then used in research to study and validate cancer biomarkers in various patient cohorts [28–30]. We have generated and tested in a pathology department a diagnostic screening strategy based on the use of TMA samples. In addition we have developed a complementary diagnostic approach based on TSA which can be used to test multiple samples when the tissue in each case is very limited. By using both these approaches we could screen for rare mutations more than 95% of the tumors received in our Pathology Department. Based on this experience we have developed a new pathological workflow designed to be easily transferable in routine diagnostics, as reported in Table 4.
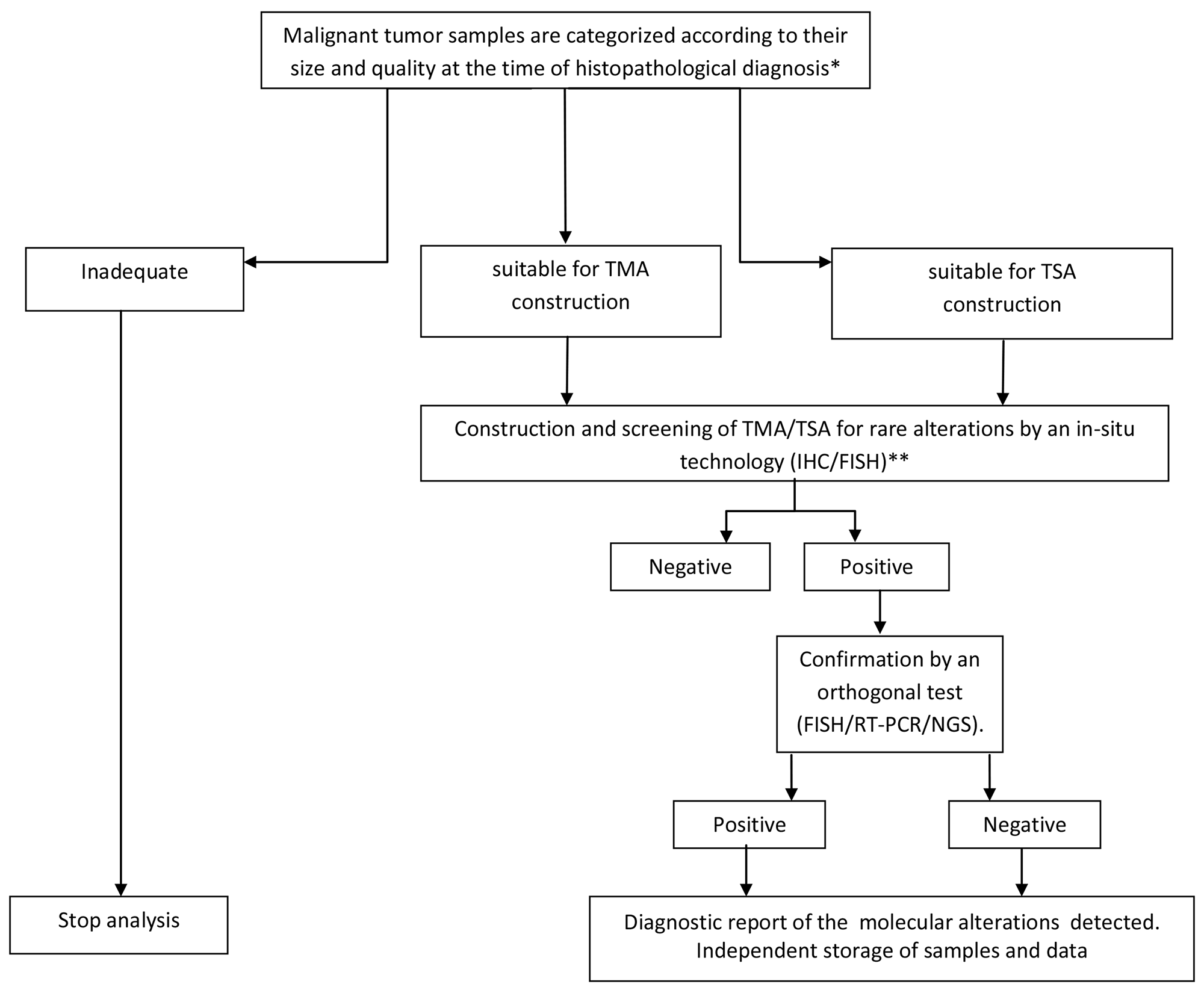
Table 4: A pathological workflow for screening of rare genomic alterations on TMA/TSA samples in routine diagnostics. *Tissue categories are reported in the management software of the pathology department. At fifteen-days/1 month intervals (depending on the number of samples available) the recorded information are used for sample selection and construction of TMA and TSA, with the tissues arranged by organ pathology (See Material and Methods).
**In the case of neoplastic forms for which routine clinical molecular testing is planned (i.e non-small cell lung cancer, colorectal carcinoma, melanoma etc.), TMA and TSA construction must be performed after the analysis of conventional predictive tests.
Abbreviations: IHC, immunohistochemistry; FISH, fluorescence in situ hybridization; RT-PCR, real-time PCR; NGS, next generation sequencing
The main limitation in the immediate application of this diagnostic strategy in all pathology departments are the limited diffusion of the TMA technology in diagnostic centers and the need of personnel with specific skills. However, the technology is not expensive and learning times are short. The specific steps that need to be implemented for this new activity can be easily addressed in most pathology departments. The selection of cases and areas of tissue at the time of reporting is conducted directly by the pathologist, the preparation of TMAs and IHC screening reactions at fortnightly intervals, require the work of a biomedical laboratory technician for about 5 days/month. Reading the IHC results on TMA/TSA samples by an experienced pathologist takes no more than 3 hours/month. TMA and TSA data can be managed by administrative staff within the management software of a pathological anatomy department, making minor changes to the software. We suggest an independent storage of TMAs at appropriate temperature (18–25°C) and controlled humidity. A diagnostic TMA bank within a pathological anatomy can have so many potential applications that the time and economic commitment necessary for its realization is in our opinion amply repaid. Based on this trial experience we have recently decided to adopt the TMA/TSA strategy in clinical practice. The approach was found to be easily adaptable to the routine workflow of our pathological department.
As reported in detail in the result section, the proposed diagnostic strategy can allow a substantial saving in terms of costs and time that makes feasible a diagnostic activity otherwise difficult to manage. The use of multiple IHC could further increase the throughput of the strategy. For example purposes only, we report for the first time, to the best of our knowledge, the possibility of simultaneously identifying different cancer biomarkers in different tumor forms (different cores) on the same TMA section. Considering the current progress of multiplexing technologies it is possible to hypothesize a multicolor analysis of several markers on the same TMA/TSA.
In the medium to long term future it can be hypothesized that multi-parametric molecular analyses such as massive parallel sequencing will be used for an accurate genomic characterization of all malignant tumors in order to treat patients with specific therapies [31]. An approach like the one presented here, essentially based on IHC screening of multiple samples may now be easily activated to implement these new therapeutic strategies. Furthermore, the described pathological approach has the advantage over genomic technologies of directly identifying the druggable targets, i.e the altered proteins, that can sometimes be the consequence of a post-transcriptional regulation which cannot be assessed by a genomic approach. In addition the TSA strategy described can allow the detection of rare alterations in cases with very limited amount of biologic material, insufficient for large scale genomic analysis. This could make the proposed diagnostic strategy a valid complement to massive parallel sequencing even when the latter method will eventually become the standard method of analysis, not to leave behind potential responders.
Materials and Methods
Tissue samples
Tissue samples for this study were obtained from the archives of the Pathology Department - SS Annunziata Hospital, University of Chieti, the number based on the average volume for a six month period of routine activity. Table 1 shows the number of malignant tumors examined during a standard six month period in our Center, calculated on the average of the last five years. A numerically corresponding series of the different malignant tumors consecutively collected at our institution was used for this study. Informed consent was obtained from all patients and that the study was conducted in accordance with the precepts of the Helsinki Declaration.
Malignant tumor samples were subjected to histopathological revision and further processed in a dedicated workflow. Samples were categorized according to their size in “large” and “small”. Large samples were defined as those containing at least one fragment ≥ 0,5 cm2 of vital tumor tissue. From each large sample the most representative tumor areas were selected for TMA construction. All the other cases were indicated as small samples or, if very small, inadequate for additional processing.
TMA and TSA construction
From each large sample, two (2mm) cores, captured in the selected areas, were deposited in a recipient 6×10 (60 cores) TMA block for simultaneous analysis of 30 patients, using the semi-automated Galileo CK3500 Micro Arrayer platform (Integrated Systems, Engineering srl, Milano). Internal specific control samples were added at the time of slide preparation.
For small samples, an innovative technique based on the preparation of Tissue Slice Array (TSA) was set up. The method consists of cutting 5 μm tissue slices from different original blocks and arranging up to 6 of them in an array on a single slide previously prepared with an alphabetized grid for patient identification (Figure 2).
Immunohistochemical screening
TMA and TSA samples were screened by IHC for the expression of mismatch repair and TRK proteins in order to indirectly assess a potential MSI status or fusions in the NTRK gene family, respectively. The expression of mismatch repair proteins was evaluated on TSAs and 4 micron sections obtained from TMAs by IHC with anti-MLH1 (MutL Protein Homolog1, ES05 clone, Dako, Glostrup, Denmark), MSH2 (mutS protein homolog 2, FE11 clone, Dako, Glostrup, Denmark), PMS2 (post-meiotic segregation increate 2,EP51 clone, Dako, Glostrup, Denmark) and MSH6 (mutS protein homolog 6, EP49 clone, Dako, Glostrup, Denmark) antibodies on a DAKO Omnis platform, following the manufacturer’s protocol. Non neoplastic tissues, stromal cells and infiltrating lymphocytes were used as internal positive controls. Normal expression was defined as nuclear staining within tumor cells, while negative protein expression (suggesting a potential MSI) was defined as complete absence of nuclear staining within tumor cells with concurrent internal positive controls.
The expression of TRK proteins was assessed by the VENTANA pan-TRK (EPR17341) assay, based on a monoclonal primary antibody directed against the C-terminal region of the TRK proteins A, B and C, on the BenchMark XT Immunostainer platform, using the OptiView DAB IHC Detection Kit (Ventana, Tucson, Arizona, USA). A sample was considered as potentially rearranged if it showed a moderate/strong cytoplasmic, perinuclear and/or nuclear immunoreaction, as reported in literature. Sequential double staining of TRK and ALK1 proteins with two horseradish peroxidase chromogenic substrates (diaminobenzidine/Magenta), was performed using the pan-TRK (EPR17341) assay and the 5A4 clone (Novocastra, Leica Biosystems, Newcastle, Upon Tyne, UK) with the EnVision FLEX HRP Magenta, High pH (Dako Omnis).
Confirmation by orthogonal methods
Cases found to be positive by IHC were tested with an orthogonal method. For the detection of MSI, fragment analysis by capillary gel electrophoresis was used. FISH, next generation sequencing (NGS) and multiplex PCR/hybridization were used for the detection of fusions in the NTRK gene family.
FISH analysis
FISH analysis was performed on unstained 4- to 5-micron, formalin-fixed, paraffin-embedded tumor tissue sections using a commercial NTRK1-NTRK2-NTRK3 Break Apart FISH probe (Empire Genomics, Buffalo NY, USA) after pretreatment and denaturation steps, according to the manufacturers’ protocols.
FISH analysis was performed with a slide scanning system under a 60X oil immersion objective with a fluorescence microscope (Olympus BX61; Olympus Corporation, Tokyo, Japan). From 50 up to 150 cancer cells per patient were scored, and signals were evaluated using the FISH imaging and capturing software SoloTouch (Bioview Duet; BioView, Rehovat, Israel). Tumor samples were considered NTRK FISH positive if more than 15% of the tumor cells showed split red and green signals (signals separated by one or more signal diameters). Otherwise, the samples were considered FISH negative.
MSI analysis
After DNA extraction from FFPE samples, MSI testing was performed using the MSI Analysis System, Version 1.2 (Promega, Madison, WI, USA), a multiplex PCR assay that included primers fluorescently labeled for two pentanucleotide repeat markers, PentaC and PentaD and five mononucleotide repeat markers, NR21 (KIT), NR24 (ZNF2), BAT25 (SLC7A8), BAT26 (MSH2), and MONO27 (MAP4K3). After PCR, amplicons were detected by capillary electrophoresis on the ABI 310 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) and the results were analyzed using the GeneMapper software version 4.1 (Applied Biosystems). MSI status was defined as MSI-High, MSI-Low, or MS-Stable, depending on the number of mononucleotide markers with instability and corresponding to two or more (≥30%), one (<30% but >0%), or zero markers, respectively.
NGS analysis for NTRK rearrangements
TRK-positive cases were analyzed for NTRK1, 2 and 3 gene rearrangements by next generation sequencing using the Archer® FusionPlex® (ArcherDX, Boulder, CO, USA) panel on the Illumina Miseq System (Illumina, San Diego, Ca, USA). To this aim, RNA was extracted from formalin-fixed, paraffin-embedded (FFPE) tumor blocks by the QIAmp FFPE tissue kit (Qiagen, Antwerp, Belgium). After cDNA synthesis, a library of DNA fragments was constructed for targeted Anchored Multiplex PCR (AMP). AMPTM chemistry utilizes open-ended targeted amplification to identify gene fusions in a single sequencing assay, even without prior knowledge of fusion partners or breakpoints.
Multiplex PCR/hybridization
The IntelliplexTM RET/NTRK rearrangement test (PlexBio, Taipei, Taiwan) was used on a multiplexing technology based on two strategies, SelectAmp and πCode (PlexBio, Taipei, Taiwan). The first one uses the Locked Nucleic Acid (LNA) to block the PCR amplification of the wild-type sequence, dramatically increasing the sensitivity and the specificity. πCode strategy is based on MicroDisc, manufactured to generate up to 16,000 distinct circular image patterns for multiplexing applications. Each πCode has a distinct circular image pattern, which corresponds to a specific capture agent conjugated to the surface of the disc. All capture agent tagged πCode are pooled, enabling capturing and detection of specific analytes in one well reaction.
CONFLICTS OF INTEREST
The authors have no conflicts of interest.
References
1. Herbst RS, Morgensztern D, Boshoff C. The biology and management of non-small cell lung cancer. Nature. 2018; 553:446–54. https://doi.org/10.1038/nature25183. [PubMed].
2. Lindeman NI, Cagle PT, Aisner DL, Arcila ME, Beasley MB, Bernicker EH, Colasacco C, Dacic S, Hirsch FR, Kerr K, Kwiatkowski DJ, Ladanyi M, Nowak JA, et al. Updated molecular testing guideline for the selection of lung cancer patients for treatment with targeted tyrosine kinase inhibitors: guideline from the college of american pathologists, the international association for the study of lung cancer, and the association for molecular pathology. Arch Pathol Lab Med. 2018; 142:321–46. https://doi.org/10.5858/arpa.2017-0388-CP. [PubMed].
3. Coit DG, Thompson JA, Albertini MR, Barker C, Carson WE, Contreras C, Daniels GA, DiMaio D, Fields RC, Fleming MD, Freeman M, Galan A, Gastman B, et al. Cutaneous melanoma, version 2.2019, NCCN clinical practice guidelines in oncology. J Natl Compr Canc Netw. 2019; 17:367–402. https://doi.org/10.6004/jnccn.2019.0018. [PubMed].
4. Sepulveda AR, Hamilton SR, Allegra CJ, Grody W, Cushman-Vokoun AM, Funkhouser WK, Kopetz SE, Lieu C, Lindor NM, Minsky BD, Monzon FA, Sargent DJ, Singh VM, et al. Molecular biomarkers for the evaluation of colorectal cancer: guideline from the american society for clinical pathology, college of american pathologists, association for molecular pathology, and the american society of clinical oncology. J Clin Oncol. 2017; 35:1453–86. https://doi.org/10.1200/JCO.2016.71.9807. [PubMed].
5. Marchetti A, Di Lorito A, Pace MV, Iezzi M, Felicioni L, D’Antuono T, Filice G, Guetti L, Mucilli F, Buttitta F. ALK protein analysis by IHC staining after recent regulatory changes: a comparison of two widely used approaches, revision of the literature, and a new testing algorithm. J Thorac Oncol. 2016; 11:487–95. https://doi.org/10.1016/j.jtho.2015.12.111. [PubMed].
6. Marchetti A, Barberis M, Di Lorito A, Pace MV, Di Lisio C, Felicioni L, Guerini-Rocco E, Vingiani A, D’Antuono T, Liberatore M, Filice G, De Luca G, De Marinis F, et al. ROS1 gene fusion in advanced lung cancer in women: a systematic analysis, review of the literature, and diagnostic algorithm. JCO Precis Oncol. 2017; 1:1–9. https://doi.org/10.1200/PO.16.00010.
7. Ettinger DS, Aisner DL, Wood DE, Akerley W, Bauman J, Chang JY, Chirieac LR, D'Amico TA, Dilling TJ, Dobelbower M, Govindan R, Gubens MA, Hennon M, et al. NCCN clinical practice guidelines in oncology (NCCN guidelines): non-small cell lung cancer. version 6.2017. Natl Compr Canc Netw. Available at https://www.nccn.org. Accessed June 12, 2017.
8. Blumenthal GM, Pazdur R. Approvals in 2018: a histology-agnostic new molecular entity, novel end points and real-time review. Nat Rev Clin Oncol. 2019; 16:139–41. https://doi.org/10.1038/s41571-019-0170-z. [PubMed].
9. Hierro C, Matos I, Martin-Liberal J, Ochoa de Olza M, Garralda E. Agnostic-Histology Approval of New Drugs in Oncology: Are We Already There? Clin Cancer Res. 2019; 25:3210–19. https://doi.org/10.1158/1078-0432.CCR-18-3694. [PubMed].
10. Cocco E, Scaltriti M, Drilon A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat Rev Clin Oncol. 2018; 15:731–47. https://doi.org/10.1038/s41571-018-0113-0. [PubMed].
11. Wakai T, Prasoon P, Hirose Y, Shimada Y, Ichikawa H, Nagahashi M. Next-generation sequencing-based clinical sequencing: toward precision medicine in solid tumors. Int J Clin Oncol. 2019; 24:115–22. https://doi.org/10.1007/s10147-018-1375-3. [PubMed].
12. Marino P, Touzani R, Perrier L, Rouleau E, Kossi DS, Zhaomin Z, Charrier N, Goardon N, Preudhomme C, Durand-Zaleski I, Borget I, Baffert S and NGSEco Group. Cost of cancer diagnosis using next-generation sequencing targeted gene panels in routine practice: a nationwide french study. Eur J Hum Genet. 2018; 26:314–23. https://doi.org/10.1038/s41431-017-0081-3. [PubMed].
13. Kummar S, Lassen UN. TRK Inhibition: A New Tumor-Agnostic Treatment Strategy. Target Oncol. 2018; 13:545–56. https://doi.org/10.1007/s11523-018-0590-1. [PubMed].
14. Chang L, Chang M, Chang HM, Chang F. Microsatellite instability: a predictive biomarker for cancer immunotherapy. Appl Immunohistochem Mol Morphol. 2018; 26:e15–21. [PubMed].
15. Dietel M, Bubendorf L, Dingemans AM, Dooms C, Elmberger G, García RC, Kerr KM, Lim E, López-Ríos F, Thunnissen E, Van Schil PE, von Laffert M. Diagnostic procedures for non-small-cell lung cancer (NSCLC): recommendations of the european expert group. Thorax. 2016; 71:177–84. https://doi.org/10.1136/thoraxjnl-2014-206677. [PubMed].
16. Yohe S, Thyagarajan B. Review of clinical next-generation sequencing. Arch Pathol Lab Med. 2017; 141:1544–57. https://doi.org/10.5858/arpa.2016-0501-RA. [PubMed].
17. Yan L, Zhang W. Precision medicine becomes reality-tumor type-agnostic therapy. Cancer Commun (Lond). 2018; 38:6. https://doi.org/10.1186/s40880-018-0274-3. [PubMed].
18. Cortes-Ciriano I, Lee S, Park WY, Kim TM, Park PJ. A molecular portrait of microsatellite instability across multiple cancers. Nat Commun. 2017; 8:15180. https://doi.org/10.1038/ncomms15180. [PubMed].
19. Palmieri G, Colombino M, Cossu A, Marchetti A, Botti G, Ascierto PA. Genetic instability and increased mutational load: which diagnostic tool best direct patients with cancer to immunotherapy? J Transl Med. 2017; 15:17. https://doi.org/10.1186/s12967-017-1119-6. [PubMed].
20. Dudley JC, Lin MT, Le DT Eshleman JR. Microsatellite instability as a biomarker for PD1 blockade. Clin Cancer Res. 2016; 22:813–20. https://doi.org/10.1158/1078-0432.CCR-15-1678. [PubMed].
21. Gatalica Z, Xiu J, Swensen J, Vranic S. Molecular characterization of cancers with NTRK gene fusions. Mod Pathol. 2019; 32:147–53. https://doi.org/10.1038/s41379-018-0118-3. [PubMed].
22. Drilon A, Laetsch TW, Kummar S, DuBois SG, Lassen UN, Demetri GD, Nathenson M, Doebele RC, Farago AF, Pappo AS, Turpin B, Dowlati A, Brose MS, et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N Engl J Med. 2018; 378:731–39. https://doi.org/10.1056/NEJMoa1714448. [PubMed].
23. Hechtman JF, Benayed R, Hyman DM, Drilon A, Zehir A, Frosina D, Arcila ME, Dogan S, Klimstra DS, Ladanyi M, Jungbluth AA. Pan-Trk immunohistochemistry is an efficient and reliable screen for the detection of NTRK fusions. Am J Surg Pathol. 2017; 41:1547–51. https://doi.org/10.1097/PAS.0000000000000911. [PubMed].
24. Penault-Llorca F, Rudzinski ER, Sepulveda AR. Testing algorithm for identification of patients with TRK fusion cancer. J Clin Pathol. 2019; 72:460–67. https://doi.org/10.1136/jclinpath-2018-205679. [PubMed].
25. Hsiao SJ, Zehir A, Sireci AN, Aisner DL. Detection of tumor NTRK gene fusions to identify patients who may benefit from tyrosine kinase (TRK) inhibitor therapy. J Mol Diagn. 2019; 21:553–71. https://doi.org/10.1016/j.jmoldx.2019.03.008. [PubMed].
26. Medicines European Agency. First ‘histology-independent’ treatment for solid tumours with a specific gene mutation. Press release. Available at https://www.ema.europa.eu/en/news/first-histology-independent-treatment-solid-tumours-specific-gene-mutation.
27. Marchetti A, Ardizzoni A, Papotti M, Crinò L, Rossi G, Gridelli C, Barberis M, Maiorano E, Normanno N, Taddei GL, Scagliotti G, Clemente C, Pinto C. Recommendations for the analysis of ALK gene rearrangements in non-small-cell lung cancer: a consensus of the italian association of medical oncology and the italian society of pathology and cytopathology. J Thorac Oncol. 2013; 8:352–58. https://doi.org/10.1097/JTO.0b013e31827d5280. [PubMed].
28. Kononen J, Bubendorf L, Kallioniemi A, Bärlund M, Schraml P, Leighton S, Torhorst J, Mihatsch MJ, Sauter G, Kallioniemi OP. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med. 1998; 4:844–47. https://doi.org/10.1038/nm0798-844. [PubMed].
29. Vogel U. Overview on techniques to construct tissue arrays with special emphasis on tissue microarrays. Microarrays (Basel). 2014; 3:103–36. https://doi.org/10.3390/microarrays3020103. [PubMed].
30. Behling F, Schittenhelm J. Tissue microarrays - translational biomarker research in the fast lane. Expert Rev Mol Diagn. 2018; 18:833–35. https://doi.org/10.1080/14737159.2018.1522252. [PubMed].
31. Sicklick JK, Kato S, Okamura R, Schwaederle M, Hahn ME, Williams CB, De P, Krie A, Piccioni DE, Miller VA, Ross JS, Benson A, Webster J, et al. Molecular profiling of cancer patients enables personalized combination therapy: the I-PREDICT study. Nat Med. 2019; 25:744–50. https://doi.org/10.1038/s41591-019-0407-5. [PubMed].

