Introduction
Notch1 is a heterodimeric and multifunctional transmembrane receptor that regulates key cellular processes, including cell fate determination, maintenance of stem cells, cell survival, proliferation and apoptosis [1, 2]. Notch1 has an extracellular domain (NECD), transmembrane domain and intracellular domain (NICD) [3, 4]. Ligand mediated activation of Notch1 induces proteolytic cleavage and release of NICD, which translocates to the nucleus and activates transcription of downstream target genes [5].
NOTCH1 is frequently altered in cancer, resulting in aberrant activation or loss of signaling in a tissue and context dependent manner [6]. A comprehensive analysis of multiple head and neck squamous cell carcinoma (HNSCC) datasets demonstrated that 10–15% of HNSCC harbour inactivating NOTCH1 mutations [7–9]. The majority of NOTCH1 mutations occur in the EGF-like ligand binding domain of the NECD, and prevent ligand binding and downstream signaling [3].
The detection of NOTCH1 mutations in dysplastic regions, and reduced expression of NOTCH1 in pre-neoplastic and cancerous skin lesions [10], suggests its potential gate-keeper properties. Some studies have implicated Notch1 signaling in angiogenesis and therapy resistance in HNSCC [11], while in vitro studies have pointed to the role of NOTCH1 in promoting keratinocyte differentiation [12]. Thus, it is important to understand how NOTCH1 contributes to oral tumorigenesis since it regulates multiple cellular processes and is a potential therapeutic target.
We previously performed whole exome sequencing of a panel of HPV-negative keratinocyte lines derived from oral squamous cell carcinomas (OSCCs), and identified NOTCH1 mutations in several of the lines [13]. In the present study we have overexpressed NICD in a patient derived OSCC line with truncating mutations in both NOTCH1 alleles. We provide evidence that the effects of NICD are mediated by negative regulation of serpin peptidase inhibitor, clade E, member 1 (SERPINE1) expression via ETV7. ETV7 is a member of the ETS (E26 transformation specific) family of transcription factors and encodes TEL2, which plays a key role in cell migration and metastasis [14]. Thus, we provide new insights into the mechanism by which NOTCH1 inactivation contributes to OSCC.
Results
NOTCH1 mutations in OSCC lines
Based on whole exome analysis of 15 OSCC and the cell lines derived from them (Supplementary Table 1), we identified a hierarchy of nonsynonymous tumour specific mutations that was representative of mutations found in larger OSCC cohorts [13]. Three of the cell lines, SJG6, SJG17 and SJG41, harboured inactivating NOTCH1 mutations, according to annotation in The Cancer Genome Atlas (Figure 1A, 1B) and were confirmed by Sanger sequencing (Supplementary Table 1). The expression of all 4 NOTCH receptors in the three lines that harbour NOTCH1 mutations was compared with normal oral mucosal keratinocytes (OK) and two OSCC lines that lack NOTCH1 mutations (Supplementary Figure 1A). There was no evidence that NOTCH1 mutations resulted in compensatory upregulation of NOTCH2, 3 or 4; indeed SJG6 had the lowest levels of all 4 receptors.
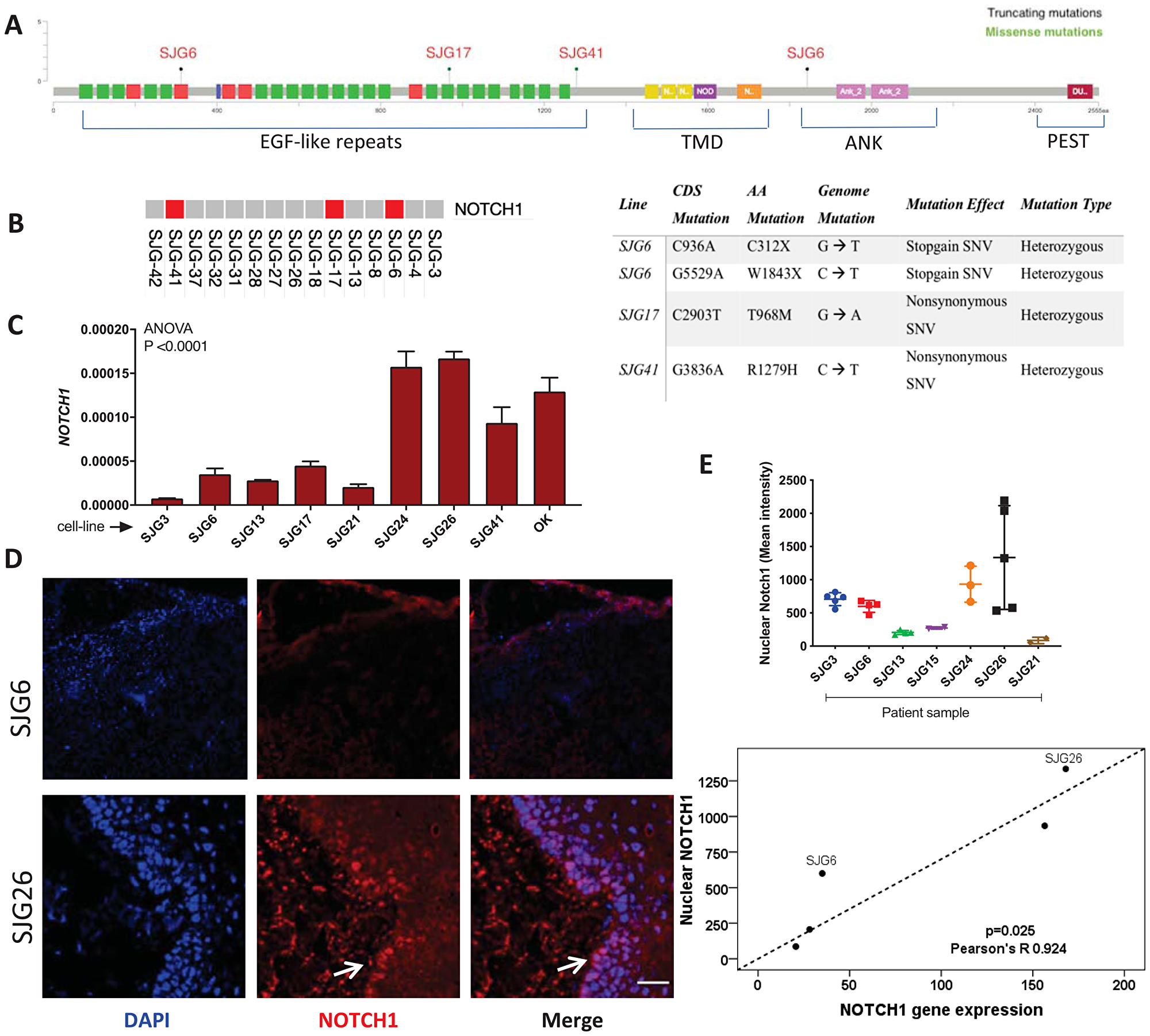
Figure 1: NOTCH1 expression in OSCC cell lines and parental tumours. (A) Schematic showing location of NOTCH1 mutations in SJG lines (red). EGF-like Repeats = epidermal growth factor-like repeats. TMD = transmembrane domain. ANK = ankyrin domain. PEST = proline, glutamic acid, serine, threonine-rich domain. Figure created using cBioPortal [61]. (B) Schematic showing which of the SJG cells lines have mutations (red), and the nature of the mutations (Table). (C) RT-qPCR analysis presenting levels of NOTCH1 mRNA in SJG lines and oral keratinocytes (OK), n = 3. Data represent mean ± SD. (D) Immunostaining of SJG parental tumours for NOTCH1 (red, arrowed) with DAPI counterstain (blue). Scale bars: 100 μm. (E) Quantification of nuclear NOTCH1 mean staining intensity in SJG tumour biopsies (top). Data represent mean ± SD. Correlation between NOTCH1 nuclear staining intensity in parental tumours and NOTCH1 mRNA expression in the corresponding SJG cell lines (bottom). p value was determined by Mann-Whitney test.
To examine the effects of mutations on NOTCH1 expression, we performed real-time PCR of mRNA extracted from cell lines, and immunostaining for NOTCH1 in sections of the original tumours (Figure 1C, 1D). Compared to OK, there was reduced expression of NOTCH1 mRNA in the majority of OSCC lines, including SJG6 and SJG17 (Figure 1C). In those lines for which the original tumour was available (Figure 1D, 1E), there was a positive correlation between NOTCH1 mRNA expression and the mean intensity of nuclear Notch1 protein labelling in the corresponding tumour samples (R = 0.9241, p = 0.025) (Figure 1E, bottom panel). The difference in Notch1 expression between the tumours from which SJG6 and SJG26 were derived was particularly striking (Figure 1C–1E).
Rescue of Notch signaling by NICD overexpression
Since the SJG6 line had inactivating mutations in both NOTCH1 alleles (Figure 1A) and therefore low levels of NOTCH1 mRNA and protein, we chose this line to examine the effects of restoring Notch signaling. A lentiviral system was used to express NICD with an mCherry reporter (SJG6+NICD). A control line was generating by transducing SJG6 with mCherry alone (SJG6 Blank). NICD mRNA levels in SJG6+NICD were approximately 3-fold higher than in SJG6 Blank (Figure 2A) and thus similar to OK (Figure 1C). When SJG6 or OK expressing the highest levels of the NICD mCherry construct were flow sorted they did not proliferate on re-plating (data not shown). This is consistent with the role of NOTCH1 signaling in promoting keratinocyte terminal differentiation [13] and explains the modest level of NICD expression in those SJG6 cells that grew following transduction.
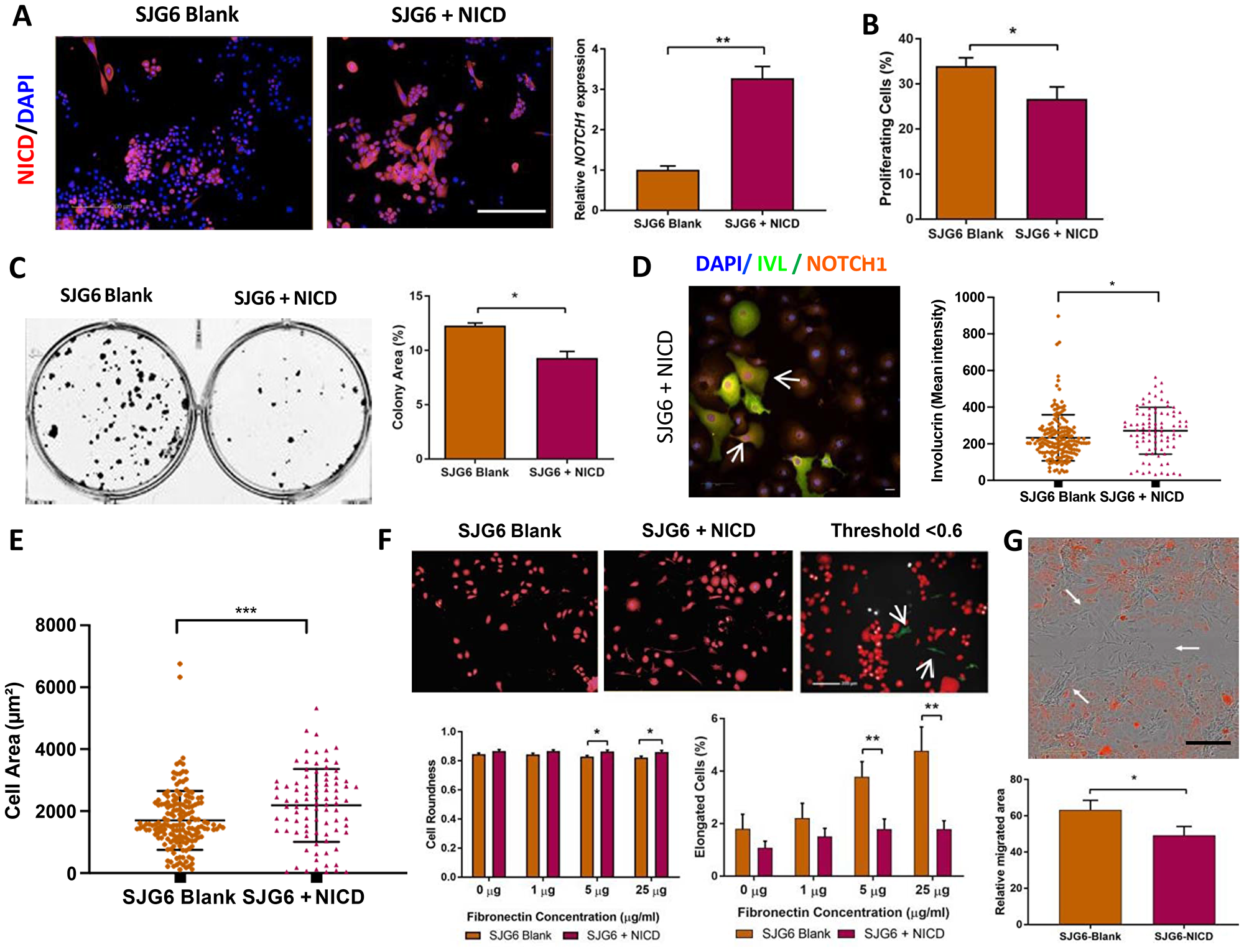
Figure 2: Phenotypic consequences of NICD overexpression. (A) mCherry (red) expression in SJG6 Blank and SJG6+NICD cells with DAPI counterstain (blue). Scale bar: 100 μm (left panels). RT-qPCR (right panel) showing relative expression of NOTCH1 in SJG6 Blank and SJG6+NICD cells, n = 3. **Student’s t-test p < 0.01. (B) Percentage of proliferating (EdU-positive) cells. n = 8. **Student’s t-test p < 0.01. (C) Effect of NICD overexpression on clonal growth of SJG6 cells. Representative dishes and quantitation are shown. **Student’s t-test p < 0.01. (D) Effect of NICD overexpression on differentiation. Representative image of Involucrin (green) and NOTCH1 (orange) immunostaining in SJG6+NICD cells and quantitation of mean staining intensity are shown. Scale bar: 20 μm. *Multiple t-test, p < 0.05. (E) Increase in cell spreading on NICD overexpression. *Multiple t-test, p < 0.05. (F) Quantitation of cell roundness and elongation via automated high content image analysis. In top panel images have overlay mask showing cells with a shape factor of less than 0.6 (red) and elongated cells (green). *Multiple t-test p < 0.05; **p < 0.01. (G) Impact of NICD expression on cell migration. Representative image demonstrating migration of cells (arrowed) into the wound area (top panel) and quantitation of relative migration (bottom panel). Scale bar: 100 μm. *Multiple t-test, p < 0.05. (A–G) Data represent mean ± SD.
To measure the effects of NICD overexpression on cell behaviour, we performed assays of cell proliferation, differentiation, adhesion, migration and shape. We observed a significant reduction in cell proliferation, as measured by EdU incorporation (Figure 2B). This correlated with a reduction in colony formation (Figure 2C), consistent with the increase in keratinocyte proliferation that occurs on NOTCH1 knockdown [15]. NOTCH1 is a known regulator of keratinocyte differentiation [12], and, consistent with this, we detected a higher proportion of Involucrin-positive cells in SJG6+NICD than SJG6 Blank cultures (Figure 2D).
NICD expression also induced changes in cell adhesion, shape and migration [16]. SJG6+NICD cells exhibited a greater spread area 24 h after plating on collagen-coated flasks (Figure 2E). Furthermore we observed a reduced number of elongated cells and a greater number of cells with a rounded morphology in SJG6+NICD compared to SJG6 Blank cells plated on 5 or 25 μg/ml fibronectin (Figure 2F). When scratch wounds were made in confluent sheets of cells, there was faster migration of SJG6 Blank cells into the wound area than SJG6+NICD cells (Figure 2G). We conclude that loss of NOTCH1 enhances OSCC cell proliferation, colony formation and migration, while inhibiting differentiation and inducing changes in cell shape, consistent with previous reports [13, 15].
NICD overexpression alters gene expression in SJG6 cells
To identify potential downstream targets of NOTCH1 we performed comparative gene expression profiling of SJG6 Blank and SJG6+NICD cells using an Affymetrix platform. We compared SJG6+NICD and SJG6 Blank cells from three independent lentiviral transduction experiments. SJG6 Blank cells clustered separately from SJG6+NICD cells (Figure 3A). The maximum fold change in NOTCH1 expression was four-fold (Figure 3B, Supplementary Table 2). Enrichment of KEGG pathways in the gene set that was upregulated in SJG6+NICD cells included the NOTCH pathway itself, and genes involved in cell adhesion (Regulation of actin cytoskeleton, Focal adhesions, ECM-receptor interactions) (Figure 3C and Supplementary Table 3).
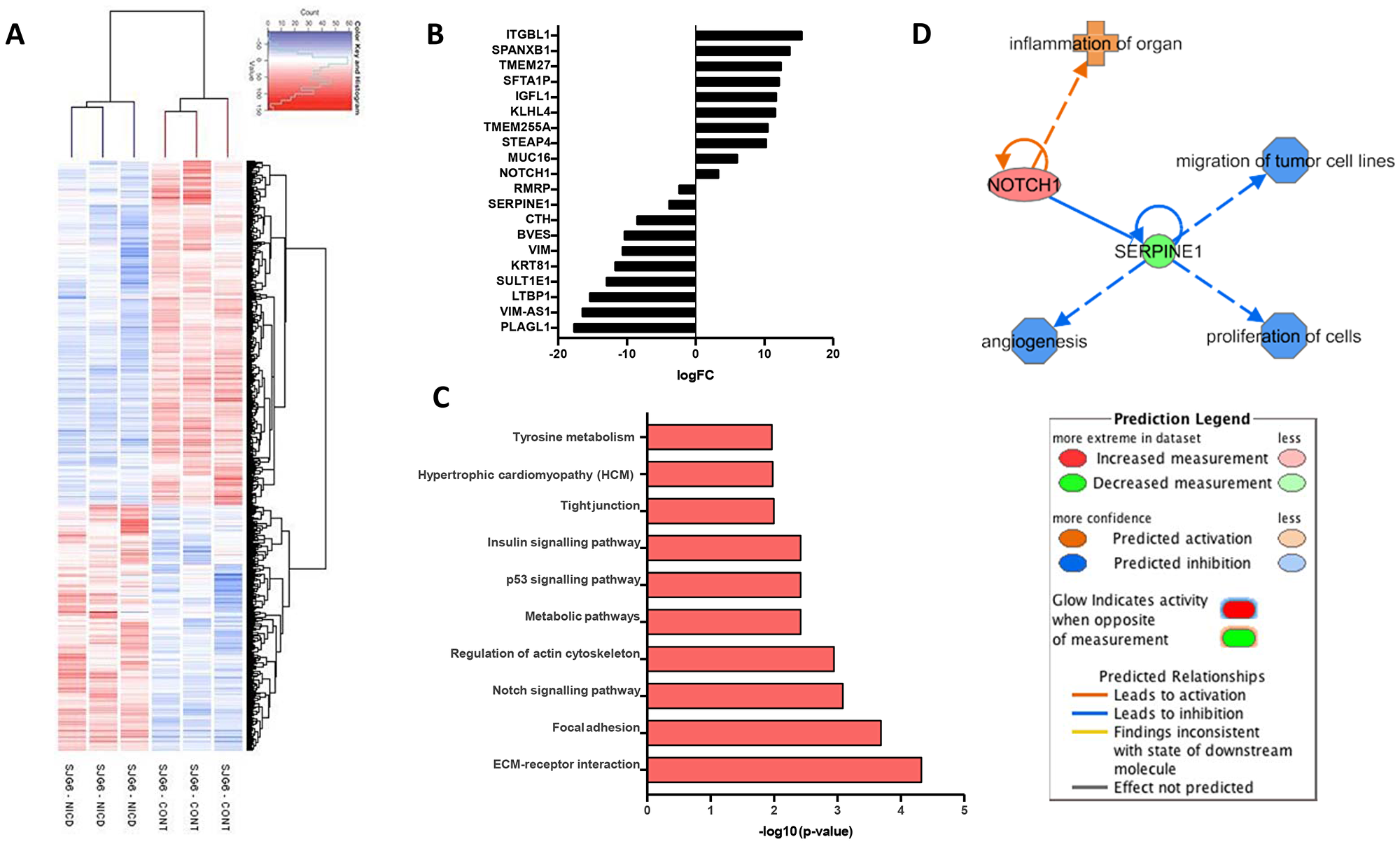
Figure 3: Gene expression profiling of SGJ6 cells transduced with NICD versus control. (A) Hierarchical clustering heatmap. (B) Top 20 up- or down- regulated genes in SJG+NICD cells. (C) Top 10 enriched KEGG pathways in SJG6+NICD cells. (D) Schematic created with IPA software showing SERPINE1 as a common regulator of NOTCH1 mediated cell behaviours.
Of the top 20 genes found to be up- or down- regulated on NICD overexpression (Figure 3B), only STEAP4 (Supplementary Table 3) is known to be a NOTCH target [17]. Of the top down-regulated genes, PLAGL1, CTH and VIM are positively correlated with HNSCC [18–20], while increased expression of RMRP, SULT1 and LTBP1 has been recorded in other cancers [21–23]. Therefore, our data are consistent with a role for NOTCH1 in altering expression of several OSCC genes (Supplementary Table 2). We also observed downregulation of the transcription factor SNAI2, which is known to promote an elongated cell shape, while expression of TWIST1/2 and ZEB1, which promote EMT and metastasis, was reduced [24] (Supplementary Table 2). This is consistent with the changes in cell phenotypes that we observed on NICD expression in SJG6 cells.
We next examined the differentially expressed genes using Gene Set Enrichment Analysis (GSEA) [25, 26] for molecular and biological functions with an activation score of more than 2 (Supplementary Tables 4, 5). Of the 46 GO terms (Supplementary Table 4), 18 predicted that NICD expression is associated with a decrease in cell migration and 5 predicted a decrease in proliferation, consistent with the observed cellular behaviours (Figure 2). Microarray data were mined for genes that could link NOTCH1 to the respective phenotypes, specifically reduced migration and proliferation and increased differentiation. SERPINE1 was found to be common to all pathways (Figure 3D) and observed to be one of the top downregulated genes in SJG6+NICD cells (Figure 3B), suggesting a potential key role in mediating the effects of NOTCH1 signaling in OSCC. SERPINE1 (also known as PAI1) is a plasminogen activator inhibitor involved in the urokinase-type plasminogen activator system (uPA), and its binding to urokinase-type plasminogen activator receptor (uPAR) triggers signals that promote migration, proliferation and cell survival [27, 28].
SERPINE1 knockdown phenocopies NOTCH1 activation in OSCCs
Overexpression of SERPINE1 in OSCC is correlated with increased cell migration, EMT and metastasis [29], while NICD expression has previously been linked to inhibition of SERPINE1 in thyroid cancer [30]. It has been shown previously that downregulation of SERPINE1 in nasopharyngeal carcinoma lines results in decreased cell migration and increased cell death [31]. To investigate whether SERPINE1 mediates any of the phenotypes associated with NOTCH1 mutation in SJG6 cells, we performed transient knockdown of SERPINE1. We predicted that when SERPINE1 expression was silenced, the SJG6 cellular phenotypes would correspond to those observed upon NICD overexpression.
siRNA mediated knockdown of SERPINE1 was confirmed 72 h post-transfection in SJG6-siSERPINE1 (referred as siSERPINE1 cells) compared to SJG6-Scrambled (SCR) by RTq-PCR and Western blotting (Figure 4A). We observed a significantly higher percentage of proliferating cells in SCR compared to siSERPINE1 cells (Figure 4B). siSERPINE1 cells exhibited a significantly greater cell area and increased involucrin labelling intensity than SCR cells (Figure 4C) and the proportion of elongated cells was reduced on 5 and 25 μg/ml fibronectin (Figure 4D). Furthermore, in a scratch wound assay SCR cells had a higher migration rate than siSERPINE1 cells (Figure 4E). These results suggest that upregulation of SERPINE1 on NOTCH1 inactivation could contribute to the changes in OSCC cell adhesion, migration, proliferation and differentiation.
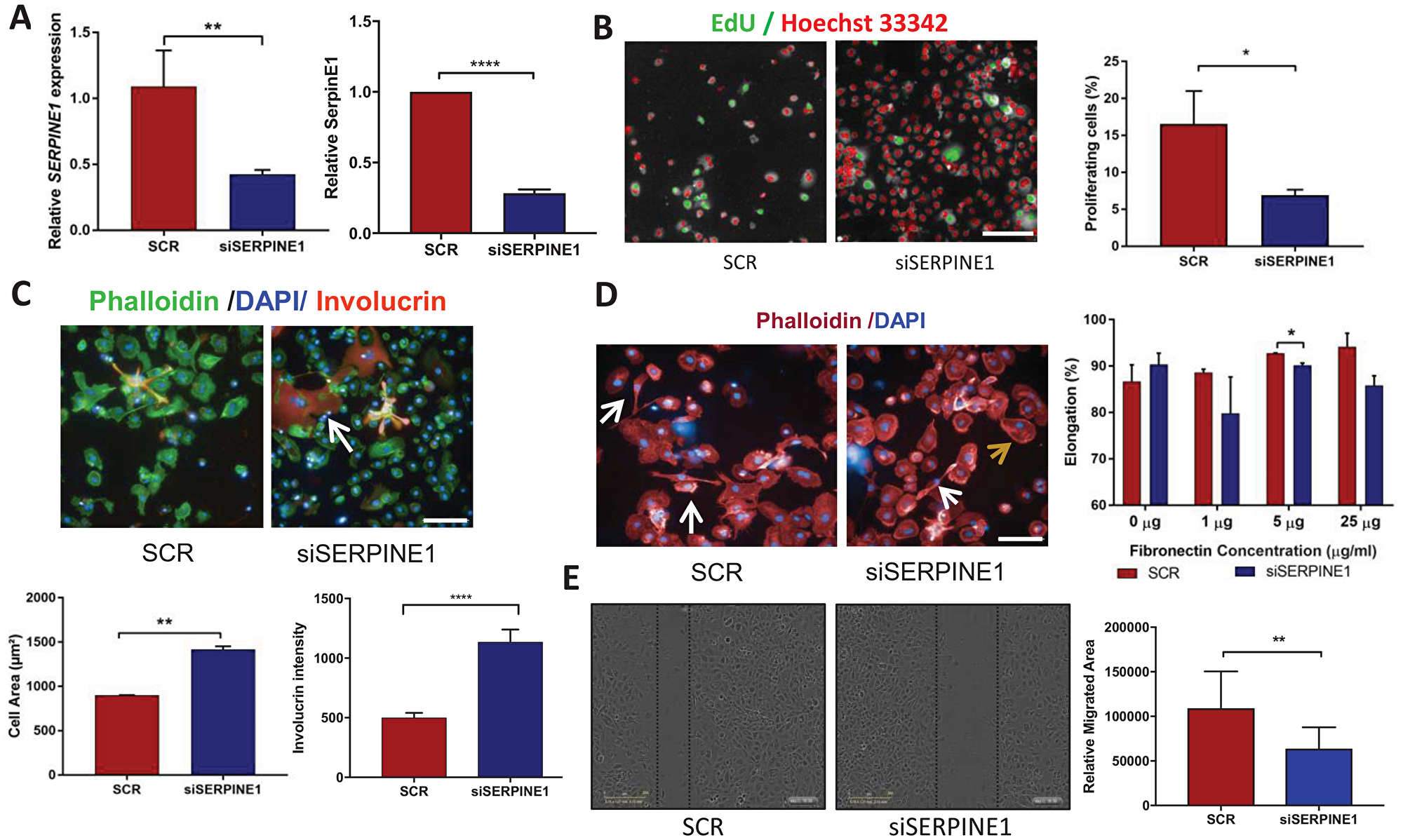
Figure 4: Cell phenotypes resulting from SERPINE1 knockdown. (A) RT-qPCR (left) and Western blot densitometric analysis showing reduced SERPINE1 expression in siSERPINE1 cells compared to SCR controls. Multiple t-test, **p < 0.01; ****p < 0.001. (B) Representative high content images after thresholding for Alexa 488 positive nuclei. EdU positive cells are depicted in green and negative cells in red, together with quantitation of % EdU labelled cells in SCR and siSERPINE1 (right). Multiple t-test, *p < 0.05. (C) Representative images showing Involucrin (orange; arrow) and Phallodin (green) staining with DAPI counterstain (blue). Changes in cell area and Involucrin labelling intensity per cell were quantified by high content imaging. Multiple t-test, **p < 0.01; ****p < 0.001. (D) Representative images of elongated (white arrows) and non-elongated (brown arrow) cells plated on Fibronectin, together with quantification. Multiple t-test, *p < 0.05. (E) Scratch wound assays and quantitation of relative migration of SCR and siSERPINE1 cells. Multiple t-test, **p < 0.01. Scale bars: 100 μm. Data represent mean ± SD (A, E) or SEM (B–D).
NICD represses SERPINE1 expression by regulating ETV7/TEL2
To discover whether NICD suppresses SERPINE1 expression we performed a comparative analysis of the SERPINE1 promoter region using the Eukaryotic Promoter Database ([32], https://epd.epfl.ch/EPDnew_database.php). We identified multiple ETV6/7 transcription factor binding motifs upstream of the transcription start site (Figure 5A). The ETS-domain transcription factor family consists of 29 genes in humans and its members are known to either activate or inhibit gene transcription [33]. Most ETS-transcription factors are aberrantly activated or repressed in tumorigenesis [34], and ETV7, which encodes TEL2, has been shown to downregulate SERPINE1 in nasopharyngeal cancer [31]. Examination of the TCGA HNSCC dataset indicated improved survival (p = 0.0389) in HNSCC patients with high ETV7 expression, whereas high SERPINE1 expression correlated with poor clinical outcome (p = 0.000616) (Figure 5B). Thus, ETV7 and SERPINE1 emerge as potential predictors of HNSCC prognosis.
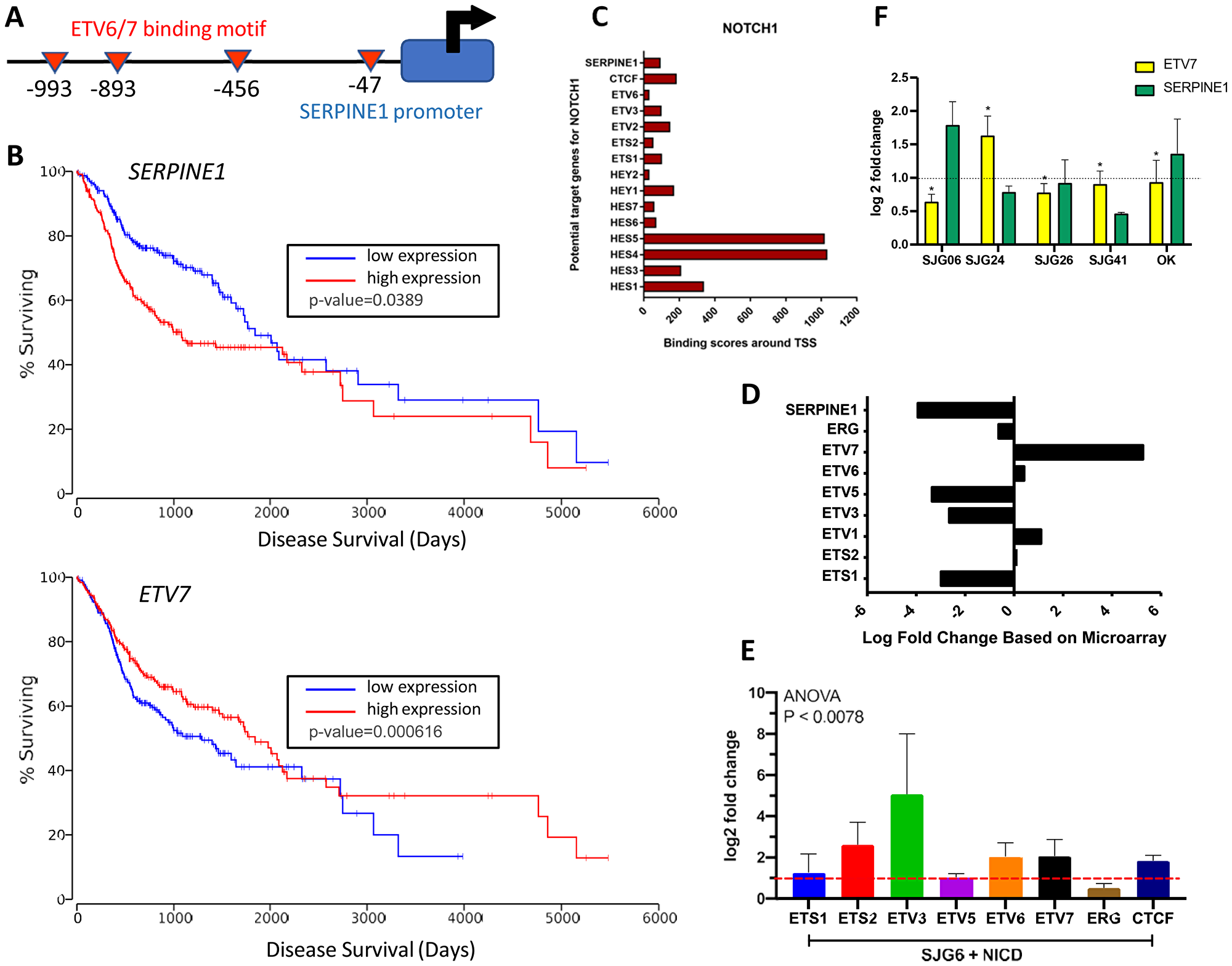
Figure 5: NOTCH1 regulation of SERPINE1 via ETV7. (A) Schematic showing ETV6/7 binding site motifs upstream of SERPINE1 promoter. (B) Kaplan–Meier survival curves in TCGA-HNSCC patient (n = 471) dataset for SERPINE1 and ETV7. Figure reproduced using OncoLnc with cut off of 45.7 [62]. (C) ChIP-Atlas data showing efficiency of NOTCH1 binding to ETS and HES/HEY promoters. (D) Relative up- and down-regulation of ETS-transcription factors in SJG6+NICD versus SJG6 Blank cells in microarray dataset. (E) RT-qPCR in SJG6+NICD versus SJG6 Blank cells (dashed red line) of CTCF and ETS transcription factors upon NICD overexpression. (F) RT-qPCR showing effects of DAPT treatment on SERPINE1 and ETV7 expression in oral keratinocytes (OK) and OSCC lines relative to DMSO control (dashed line). n = 3. 2-Way ANOVA was used for statistical analysis. (E, F) Data represent mean ± SEM.
According to data deposited in ChIP-Atlas, a comprehensive database for visualizing and exploring public ChIP-seq datasets ([35], https://chip-atlas.org/), NOTCH1 is the only NOTCH protein that binds the transcription start site (TSS) of ETS-transcription factors and SERPINE1 with comparable binding efficiency to the gene promoter regions of HES and HEY, two canonical NOTCH1 target genes, (Figure 5C) in T-cell lymphoblastic lymphoma and T-cell Acute Lymphoblastic Leukaemia derived lines [35]. This led us to examine gene expression levels of ETS-transcription factors in the SJG6+NICD versus SJG6 Blank microarray dataset. We observed upregulation of ETV7 (nearly a 6-fold increase) and downregulation of ETV5 (almost 4-fold) in SJG6+NICD compared to SJG6 Blank cells (Figure 5D).
To validate these observations experimentally, we performed qRT-PCR. We observed an upregulation of ETV3 (by 4-fold) and ETV6/7 (2-fold) in SJG6+NICD compared to SJG6 Blank cells (Figure 5E). To further explore NOTCH1 mediated regulation of ETV7 and SERPINE1 expression, NOTCH signaling was inhibited pharmacologically with the γ-secretase inhibitor DAPT. A reduction in ETV7 and upregulation of SERPINE1 were observed in both OK and SJG6 (Figure 5F). These results suggest that NICD suppresses SERPINE1 by upregulation of the ETS-transcription factor ETV7 in OSCC. Nevertheless, the NOTCH pathway is not the sole means by which ETV7 and SERPINE1 are regulated, because in SJG24 cells DAPT treatment led to an increase in ETV7 levels, while in SJG26 and SJG41 cells DAPT did not increase SERPINE1 (Figure 5F).
Relationship between NOTCH1, SERPINE1 and ETV7/TEL2 expression in OSCC
We next compared the expression levels of ETV7 and SERPINE1 in SJG cell lines and the tumours from which they were derived (Figure 6A–6C and Supplementary Figure 1B, 1C). There was very low nuclear ETV7 protein expression in SJG6 (NOTCH1 mutant) compared to SJG26 (NOTCH1 wild type) patient samples, and a significantly lower percentage of ETV7 positive cells (Figure 6B). In addition, ETV7 expression was inversely correlated with SERPINE1, since the SJG6 parental tumour and cell line had higher levels of SERPINE1 than the SJG26 parental tumour and cell line (Figure 6D). Overall, this study suggests a mechanism whereby NICD mediated SERPINE1 downregulation via ETV7 contributes to several of the tumour-suppressive features of NOTCH1 signaling (Figure 6E).
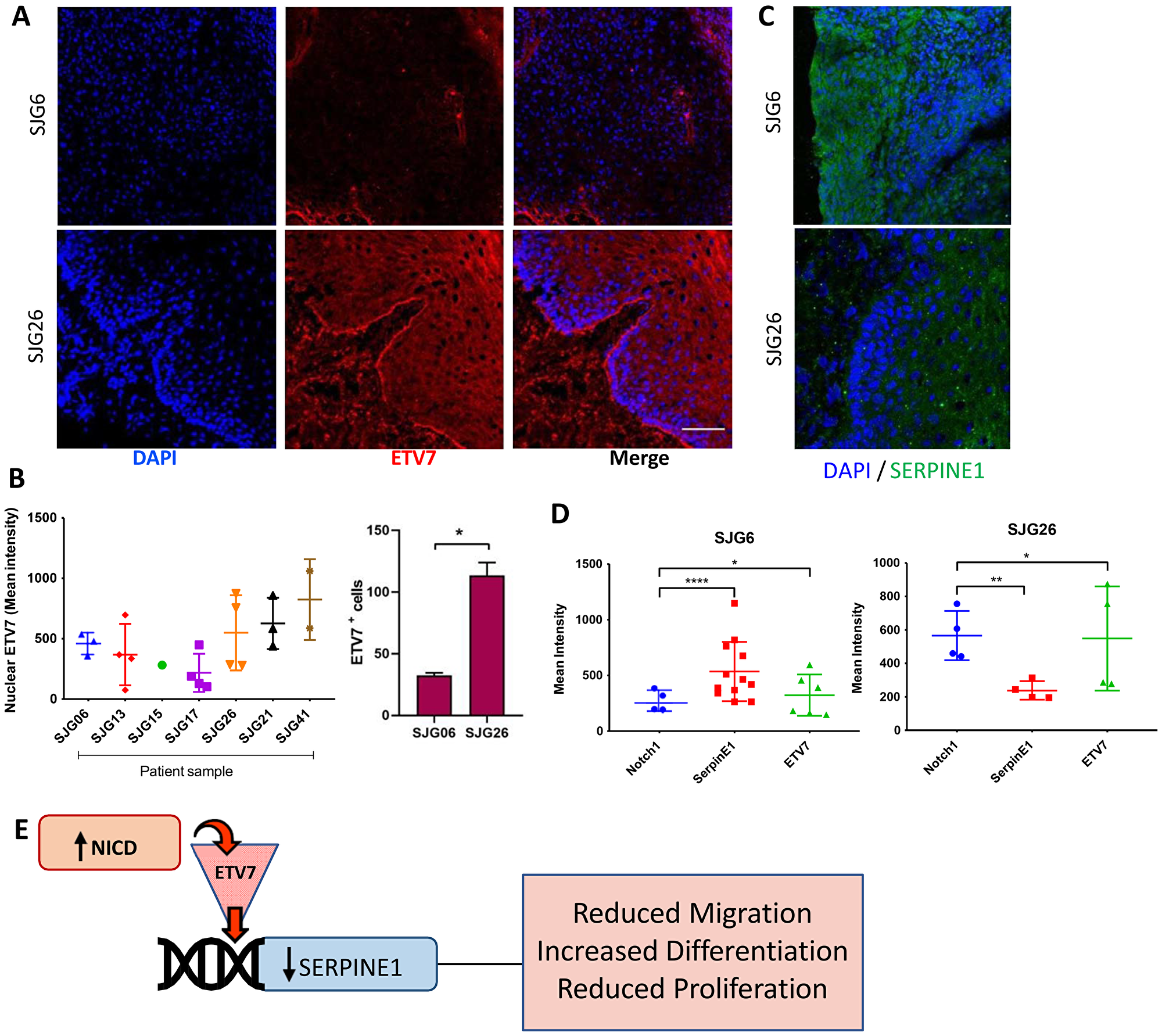
Figure 6: Inverse relationship between ETV7 and SERPINE1 in OSCC. (A) Representative immunostaining for ETV7 (red) in SJG26 and SJG6 tumour biopsies with DAPI counterstain (blue). Scale bar: 200 μm. (B) Quantification of mean nuclear ETV7 staining in OSCC tumour biopsies. Note lower % ETV7 positive cells in NOTCH1 mutant tumour (SJG6) compared to NOTCH1 wild type (SJG26). Multiple t-test, *p < 0.05. (C) Representative immunostaining for SERPINE1 (green) in SJG26 and SJG6 tumour biopsies with DAPI counterstain (blue). Scale bar: 200 μm. (D) Quantification of staining intensities for NOTCH1, ETV7 and SERPINE1 in SJG6 (NOTCH1 mutant) and SJG26 (NOTCH1 WT) tumour biopsies. One sample t-test was used. *p < 0.05; **p < 0.01; ****p < 0.0001. (E) Schematic of the NOTCH1/ETV7/SERPINE1 regulatory axis. (B, D) Data represent mean ± SD.
Discussion
OSCCs are highly heterogeneous, both genetically and in terms of tumour biology, and have a poor clinical outcome [36]. Hence it is critical to understand the role of specific mutations in determining the properties of individual tumours, as this may lead to the development of new treatments. In the present study we have shown that Notch signaling negatively regulates expression of SERPINE1 via ETV7 to control cell behaviour and that SERPINE1 and ETV7 correlate inversely with survival in OSCC. This is consistent with an earlier study demonstrating ETV7-mediated downregulation of SERPINE1 expression and an associated survival benefit in patients with nasopharyngeal carcinoma [31].
The NOTCH1 signaling pathway is highly conserved and regulates diverse cellular processes, including the differentiation of cells in stratified squamous epithelia [37–39]. NOTCH signaling is frequently dysregulated in HNSCCs, often in association with inactivating NOTCH1 mutations [7, 8]. Inhibition of NOTCH1 in oesophageal epithelial cells leads to clonal expansion, representative of field cancerisation [40]. Although mutational inactivation is frequent in HNSCC [7], other context dependent oncogenic or tumour suppressive roles for NOTCH signaling have been observed in various tumours including HNSCCs [41–43]. This would be consistent with our finding that NOTCH1 mutation is not the sole determinant of NOTCH1 expression in OSCC lines (Figure 1).
To elucidate the mechanism by which NOTCH1 inactivation affects cell behaviour in OSCC, we performed comparative gene expression profiling of SJG6 cells with and without NICD overexpression. NICD rescue led to decreased expression of EMT mediators such as ZEB1, TWIST1/2, MMP9 and SNAI2. This is consistent with evidence suggesting NOTCH1 is a mediator of metastasis via the regulation of MMP2 and MMP9 in HNSCC [44]. In addition, NOTCH1 contributes to maintenance of Tumour Necrosis Factor-α (TNF-α)-mediated invasion via transcriptional regulation of SNAI2 and TWIST [45]. Many of the IPA terms in our analysis related NOTCH1 and SERPINE1 to cell adhesion, migration, invasion and cell proliferation. A direct interaction between NOTCH1 and SERPINE1 in the cell behaviour phenotypes studied in IPA was consistently observed. It has been shown previously that downregulation of SERPINE1 depends on NOTCH signaling [46]. Most studies have focused on SERPINE1 upregulation upon wounding, which stimulates epithelial migration [47, 48]. SERPINE1 mediates the response of cell locomotion in ras-transformed human keratinocytes treated with TGFβ1 and EGF, and moreover SERPINE1 upregulation requires the activities of mitogen-activated extracellular kinase (MEK), p21 and pp60 in addition to EGFR [49].
Consistent with the microarray analysis we demonstrated downregulation of SERPINE1 as a result of NICD overexpression. Overexpression of SERPINE1 has been related to high risk of metastatic spread in HNSCC [29]. We demonstrated that the changes in cell behaviour and morphology resulting from knockdown of SERPINE1 resembled those resulting from NICD overexpression, suggesting that SERPINE1 contributes to the effects of NOTCH1 inactivation. This finding supports a study demonstrating SERPINE1 downregulation by NICD in differentiated thyroid tumours [30]. SERPINE1 also protects endothelial cells from FasL-mediated apoptosis [50]. While previous studies have indicated that NICD-induced keratinocyte differentiation is mediated by HES target genes [12, 24], our new results suggest a contribution of ETV7. Although we have not observed cell senescence in our OSCC model, the changes in morphology could be consistent with the senescent phenotype described in OSCC lines [51].
Based on promoter analysis and the ChIP-Atlas database, we identified ETV7/TEL2 as a key factor regulating NICD-mediated SERPINE1 expression. NICD overexpression in SJG6 cells led to upregulation of several ETS-transcription factors, including ETV7, in addition to HES transcription factors. Furthermore ChIP-Atlas data confirmed binding of NOTCH1 to the transcription start sites of both ETS-members and SERPINE1. Negative regulation of SERPINE1 by TEL2 in nasopharyngeal carcinoma has already been described [31]. However, regulation of TEL2 by NICD has not been observed previously. ETS-transcription factors regulate a plethora of cellular process, and play a crucial role in tumorigenesis [52]. Recent studies have highlighted the importance of different combinations of ETS- factors in patient stratification and are emerging as novel therapeutic targets in various cancers [53]. Similarly, we observed ETV7 as a predictor of the patient outcome in HNSCCs.
In summary, our study indicates that the tumour suppressive role of NOTCH1 in OSCC is manifested, at least in part, by ETV7-mediated suppression of SERPINE1. We have identified the NICD/TEL2/SERPINE1 regulatory axis as a stratification and prognostic tool in OSCC that may be valuable for developing new treatments.
Materials and Methods
OSCC biopsies and derivation of OSCC lines
Anonymised biopsies of OSCC or normal oral mucosa were collected with appropriate ethical approval (UK National Research Ethics Service 08/H0306/30), as described previously [13]. The genomic DNA of the 15 SJG cell lines analysed in this study and a further 5 lines for which parental tumour DNA was not available (Supplementary Table 1) was isolated by using the QIAamp DNA Mini Kit (Qiagen). To generate STR profiles of the cells (Supplementary Table 1), PowerPlex assays (Promega) were performed by Source BioScience (Nottingham, UK). The following loci were tested: AMEL, CSF1PO, D13S317, D16S539, D18S51, D21S11, D3S1358, D5S818, D7S820, D8S1179, FGA, Penta D, Penta E, TH01, TPOX, vWA. STR profiles of SJG cells were compared with other cell lines using the STR Similarity Search Tool Cellosaurus 1.1.0 (ExPASy) and found to be distinct.
OSCC lines and a primary oral keratinocyte line (OK) were cultured on a feeder layer of J2 3T3 cells in complete FAD medium as described previously [54]. For some experiments cells were transferred to Keratinocyte Serum Free Medium containing EGF and bovine pituitary extract (KSFM) (Thermo Fisher Scientific).
Plasmid constructs and transfection
To achieve constitutive expression of NICD, 3xFlagNICD1 (Addgene #20183, [55]) was cloned into the multiple cloning site, under the SFFV promoter, of the LeGO-iC2 plasmid (Addgene#27345, [56]) using the Gateway Vector Conversion System (ThermoFisher Scientific). The plasmids were kind gifts from Raphael Kopan and Boris Fehse, respectively. Lentivirus was produced by transfecting HEK293 with second generation packaging plasmids (psPAX2 - Addgene#12260, and pMD2.G-Addgene#12259) using JetPRIME® (Polyplus Transfection) according to the Manufacturer’s guidelines. OSCC cells were subsequently transduced with lentiviruses.
Transient transfection with SMARTpool ON-TARGETplus SERPINE1 (ID 118572, Thermo Fisher) and Silencer® Negative Control No. 1 (Thermo Fisher) siRNAs was carried out using INTERFERin® (Polyplus transfection).
Quantitative real-time RT-PCR (qRT-PCR)
RNA isolation and cDNA synthesis were performed using the RNeasy Mini Kit and QuantiTect Reverse Transcription Kit (QIAGEN). qRT-PCR was performed using TaqMan™ Fast Universal PCR Master Mix, no AmpErase™ UNG (Life Technologies) or SYBR Green Master Mix in a Bio-Rad CFX qPCR Instrument with the TaqMan primers listed in Supplementary Table 6. The following qPCR primers were used. NOTCH1: TCCACCAGTTTGAATGGTCA (forward), AGCTCATCATCTGGGACAGG (reverse); NOTCH2: GATCACCCGAATGGCTATGAAT (forward), GGGGTCACAGTTGTCAATGTT (reverse); NOTCH3, TGGCGACCTCACTTACGACT (forward), CACTGGCAGTTATAGGTGTTGAC (reverse); NOTCH4: TGTGAACGTGATGTCAACGAG (forward), ACAGTCTGGGCCTATGAAACC (reverse). Relative quantification of gene expression was performed using 2-ΔCt or 2-ΔΔCt methods.
Colony formation
72 h after transfection, 500 cells were seeded at single cell density in 6-well plates on a layer of feeders. 10–14 days after seeding, feeders were detached, and keratinocytes were fixed in 10% formalin and stained with a solution of 1% Rhodamine B and 1% Nile blue. Well images were acquired on a Gel Doc™ XR+ (Bio-Rad) and the % of each well covered by colonies was determined using the The ColonyArea plugin tool from ImageJ [57].
Cell migration
0.5 × 106 cells were seeded per well in KSFM on rat tail collagen I (Corning) - coated 6-well plates. After confluence, cells were treated with 4 μg/ml Mitomycin-C for 2 h. A scratch wound was created and cell migration into the wound area was monitored for 20 h using an IncuCyte®ZOOM (Essen BioScience). Relative wound area was measured using ImageJ.
Cell proliferation, differentiation and morphology
To assay proliferation, cells were seeded in KSFM at a concentration of 105 cells/well in collagen-coated 96-well plates. Cell proliferation was determined with a Click-iT Alexa Fluor 488 Imaging Kit (Life Technologies).
For morphology and differentiation assays, cells were seeded at clonal density in 12-well plates coated with collagen for the differentiation assay or fibronectin (0, 1, 5, 25 μg/ml) to measure cell morphology. After 72 h cells were fixed in 4% paraformaldehyde, permeabilised in 0.1% Triton™ X-100 in PBS for 5 min, and stained with DAPI (Thermofisher, D1306) and Phalloidin (detailed in Supplementary Table 7) for 1 h at room temperature or labelled overnight at 4°C with anti-involucrin (SY7 [58]) antibody, followed by 1 h incubation with Alexa Fluor-conjugated Secondary Antibody at room temperature (RT). Details of all the Primary and secondary antibodies used in the study are listed in Supplementary Table 7. Images were acquired using the Operetta® High-Content Imaging System (PerkinElmer). Cell morphology features (cell area, elongation and roundness), % EdU and % Involucrin positive cells were calculated using Harmony Software (PerkinElmer).
Pharmacological inhibition of Notch signaling
The γ-secretase inhibitor N-[N-(3,5-Difluorophenacetyl)-L-alanyl]-S-phenylglycine t-butyl ester (DAPT; Sigma), dissolved in dimethyl sulfoxide (DMSO; Stratech), was used to block Notch signaling. Cells were seeded in 6-well plates at 105 cells per well in 2 ml KSFM supplemented with 10 µM DAPT or DMSO. After 24 h cells were grown in complete FAD medium for 48 h and then harvested.
Gene expression microarrays
cDNA was fragmented and labelled with the Affymetrix GeneChip Labeling Kit, followed by hybridization on Human Gene 1.0 ST Arrays (Affymetrix). The microarrays were scanned on a GeneChip scanner 3000 (Affymetrix) and analysed using the oligo (Carvalho and Irizarry, 2010) and limma [59] packages in R-studio software (https://rstudio.com/). Raw data, in CEL format, were normalized using Robust Multichip Average (RMA) and annotated using the Entrez IDs from the ‘hugene20transcripcluster’ package. A heatmap using the ‘gplots’ package [60] was then created for the differentially expressed genes filtered at a p-value of 0.05 and log-fold change of 2. Go-Term and GSEA were run prior to exporting the gene list and fold change values to Ingenuity Pathway Analysis (IPA) Software. The data are deposited in Gene Expression Omnibus (accession number GSE127770).
Immunofluorescence microscopy
Frozen 6–8 μm sections were fixed in 4% paraformaldehyde and permeabilised in 0.5% Triton™ X-100 in PBS at room temperature or with 1:1 acetone-methanol (for nuclear antigens) at –20°C for 5 min. Sections were blocked with a solution of 10% donkey serum, 0.1% fish skin gelatin, 0.1% Triton X-100, and 0.5% Tween-20 (all from Sigma) in phosphate buffered saline (PBS), and labelled overnight at 4°C with primary antibodies (Supplementary Table 7) diluted in blocking buffer. Sections were washed with PBS and then labelled with Alexa Fluor-conjugated secondary antibodies and DAPI for 1 h at RT, washed with PBS, and mounted with ProLong Gold antifade reagent (Thermo Fisher Scientific). Confocal microscopy was performed with a Nikon A1 Upright Confocal microscope (Tokyo, Japan) using 20× objectives. Image processing was performed with Image J (Fiji) (National Institutes of Health, Bethesda, MD).
Western blotting
30 μg protein were loaded into pre-cast 4–15% Mini-PROTEAN® TGXTM gels (Bio-Rad) alongside Precision Plus Protein™ Dual Colour Standards (Bio-Rad) and run at 150V in Tris/Glycine/SDS Running Buffer (Bio-Rad). The Trans-Blot® Turbo™ Transfer Starter System, Mini PVDF (Bio-Rad) was used to transfer protein to membranes followed by blocking for 1 h, overnight incubation with primary antibodies at 4°C and 1 h incubation with horseradish peroxidase (HRP)-conjugated secondary antibody at RT. ClarityTM ECL Western Blotting Substrate (Bio-Rad) and ChemiDoc Touch (Bio-Rad) were used to image the membranes. Relative protein band intensities were quantified and normalized against Vinculin using densitometric analysis.
Statistical analysis
Graphs were generated using GraphPad Prism7 (La Jolla, CA) and represent Mean ± SEM of 3 independent experiments. One-way analysis of variance parametric test with Bonferroni post-test or Student’s t-test was performed, with p < 0.05 considered significant.
Abbreviations
DAPT: (2S)-N-[(3,5-Difluorophenyl)acetyl]-L-alanyl-2-phenyl]glycine1,1-dimethylethylester; DMSO: Dimethyl Sulfoxide; ECM: Extracellular Matrix; GSEA: Gene Set Enrichment Analysis; HNSCC: Head and Neck Squamous Cell Carcinoma; KEGG: Kyoto Encyclopaedia of Genes and Genomes; NICD: Notch Intracellular Domain; NECD: Notch Extracellular Domain; OSCC: Oral Squamous Cell Carcinoma; OK: Oral Keratinocytes.
ACKNOWLEDGMENTS
We thank all our colleagues for valuable input into the project, in particular Victor Negri, Simon Broad, Kazunori Sunadome and Angela O. Pisco. FMW is currently on secondment as Executive Chair of the Medical Research Council.
CONFLICTS OF INTEREST
The authors state no conflicts of interest.
FUNDING
FMW gratefully acknowledges funding from Cancer Research UK (C219/A23522), the Wellcome Trust (206439/Z/17/Z) and the Medical Research Council (MRC) (MR/PO18823/1). AA-D was funded by the MRC via the King’s College London Doctoral Training Programme. We also acknowledge funding from the Department of Health via the National Institute for Health Research comprehensive Biomedical Research Centre award to Guy’s & St Thomas’ National Health Service Foundation Trust in partnership with King’s College London and King’s College Hospital NHS Foundation Trust.
References
1. Bray SJ. Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol. 2006; 7:678–89. https://doi.org/10.1038/nrm2009. [PubMed].
2. Sherbet GV. Notch signalling in carcinogenesis. Mol Approach to Cancer Manag. Academic Press; 2017. pp. 81–8. https://doi.org/10.1016/B978-0-12-812896-1.00008-8.
3. Kopan R, Ilagan MX. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell. 2009; 137:216–33. https://doi.org/10.1016/j.cell.2009.03.045. [PubMed].
4. Gokulan R, Halagowder D. Expression pattern of Notch intracellular domain (NICD) and Hes-1 in preneoplastic and neoplastic human oral squamous epithelium: their correlation with c-Myc, clinicopathological factors and prognosis in Oral cancer. Med Oncol. 2014; 31:126. https://doi.org/10.1007/s12032-014-0126-1. [PubMed].
5. Borggrefe T, Lauth M, Zwijsen A, Huylebroeck D, Oswald F, Giaimo BD. The Notch intracellular domain integrates signals from Wnt, Hedgehog, TGFβ/BMP and hypoxia pathways. Biochim Biophys Acta. 2016; 1863:303–13. https://doi.org/10.1016/j.bbamcr.2015.11.020. [PubMed].
6. Artavanis-Tsakonas S, Matsuno K, Fortini ME. Notch signaling. Science. 1995; 268:225–32. https://doi.org/10.1126/science.7716513. [PubMed].
7. Agrawal N, Frederick MJ, Pickering CR, Bettegowda C, Chang K, Li RJ, Fakhry C, Xie TX, Zhang J, Wang J, Zhang N, El-Naggar AK, Jasser SA, et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science. 2011; 333:1154–57. https://doi.org/10.1126/science.1206923. [PubMed].
8. Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, Kryukov GV, Lawrence MS, Sougnez C, McKenna A, Shefler E, Ramos AH, Stojanov P, et al. The mutational landscape of head and neck squamous cell carcinoma. Science. 2011; 333:1157–60. https://doi.org/10.1126/science.1208130. [PubMed].
9. Fukusumi T, Califano JA. The NOTCH pathway in head and neck squamous cell carcinoma. J Dent Res. 2018; 97:645–53. https://doi.org/10.1177/0022034518760297. [PubMed].
10. South AP, Purdie KJ, Watt SA, Haldenby S, den Breems N, Dimon M, Arron ST, Kluk MJ, Aster JC, McHugh A, Xue DJ, Dayal JH, Robinson KS, et al. NOTCH1 mutations occur early during cutaneous squamous cell carcinogenesis. J Invest Dermatol. 2014; 134:2630–38. https://doi.org/10.1038/jid.2014.154. [PubMed].
11. Zhao YY, Yu GT, Xiao T, Hu J. The Notch signaling pathway in head and neck squamous cell carcinoma: A meta-analysis. Adv Clin Exp Med. 2017; 26:881–87. https://doi.org/10.17219/acem/64000. [PubMed].
12. Watt FM, Estrach S, Ambler CA. Epidermal Notch signalling: differentiation, cancer and adhesion. Curr Opin Cell Biol. 2008; 20:171–9. https://doi.org/10.1016/j.ceb.2008.01.010. [PubMed].
13. Hayes TF, Benaich N, Goldie SJ, Sipilä K, Ames-Draycott A, Cai W, Yin G, Watt FM. Integrative genomic and functional analysis of human oral squamous cell carcinoma cell lines reveals synergistic effects of FAT1 and CASP8 inactivation. Cancer Lett. 2016; 383:106–14. https://doi.org/10.1016/j.canlet.2016.09.014. [PubMed].
14. Carella C, Potter M, Bonten J, Rehg JE, Neale G, Grosveld GC. The ETS factor TEL2 is a hematopoietic oncoprotein. Blood. 2006; 107:1124–32. https://doi.org/10.1182/blood-2005-03-1196. [PubMed].
15. Negri VA, Logtenberg ME, Renz LM, Oules B, Walko G, Watt FM. Delta-like 1-mediated cis-inhibition of Jagged1/2 signalling inhibits differentiation of human epidermal cells in culture. Sci Rep. 2019; 9:10825. https://doi.org/10.1038/s41598-019-47232-2. [PubMed].
16. Park J, Schwarzbauer JE. Mammary epithelial cell interactions with fibronectin stimulate epithelial-mesenchymal transition. Oncogene. 2014; 33:1649–57. https://doi.org/10.1038/onc.2013.118. [PubMed].
17. Wang C, Zhang CJ, Martin BN, Bulek K, Kang Z, Zhao J, Bian G, Carman JA, Gao J, Dongre A, Xue H, Miller SD, Qian Y, et al. IL-17 induced NOTCH1 activation in oligodendrocyte progenitor cells enhances proliferation and inflammatory gene expression. Nat Commun. 2017; 8:15508. https://doi.org/10.1038/ncomms15508. [PubMed].
18. Marques Filho MF, Walder F, Takahashi HK, Guimarães LL, Tanaka AK, Cervantes O, Straus AH. Glycosphingolipid expression in squamous cell carcinoma of the upper aerodigestive tract. Braz J Otorhinolaryngol. 2006; 72:25–30. https://doi.org/10.1016/S1808-8694(15)30029-X. [PubMed].
19. Giefing M, Zemke N, Brauze D, Kostrzewska-Poczekaj M, Luczak M, Szaumkessel M, Pelinska K, Kiwerska K, Tönnies H, Grenman R, Figlerowicz M, Siebert R, Szyfter K, Jarmuz M. High resolution ArrayCGH and expression profiling identifies PTPRD and PCDH17/PCH68 as tumor suppressor gene candidates in laryngeal squamous cell carcinoma. Genes Chromosomes Cancer. 2011; 50:154–66. https://doi.org/10.1002/gcc.20840. [PubMed].
20. Bayo P, Jou A, Stenzinger A, Shao C, Gross M, Jensen A, Grabe N, Mende CH, Rados PV, Debus J, Weichert W, Plinkert PK, Lichter P, et al. Loss of SOX2 expression induces cell motility via vimentin up-regulation and is an unfavorable risk factor for survival of head and neck squamous cell carcinoma. Mol Oncol. 2015; 9:1704–19. https://doi.org/10.1016/j.molonc.2015.05.006. [PubMed].
21. Xu C, Wang P, Liu Y, Zhang Y, Fan W, Upton MP, Lohavanichbutr P, Houck JR, Doody DR, Futran ND, Zhao LP, Schwartz SM, Chen C, Méndez E. Integrative genomics in combination with RNA interference identifies prognostic and functionally relevant gene targets for oral squamous cell carcinoma. PLoS Genet. 2013; 9:e1003169. https://doi.org/10.1371/journal.pgen.1003169. [PubMed].
22. Cao B, Yang L, Rong W, Feng L, Han N, Zhang K, Cheng S, Wu J, Xiao T, Gao Y. Latent transforming growth factor-beta binding protein-1 in circulating plasma as a novel biomarker for early detection of hepatocellular carcinoma. Int J Clin Exp Pathol. 2015; 8:16046–54. [PubMed].
23. Meng Z, Moroishi T, Guan KL. Mechanisms of Hippo pathway regulation. Genes Dev. 2016; 30:1–17. https://doi.org/10.1101/gad.274027.115. [PubMed].
24. Ambler CA, Watt FM. Adult epidermal Notch activity induces dermal accumulation of T cells and neural crest derivatives through upregulation of jagged 1. Development. 2010; 137:3569–79. https://doi.org/10.1242/dev.050310. [PubMed].
25. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005; 102:15545–50. https://doi.org/10.1073/pnas.0506580102. [PubMed].
26. Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E, Ridderstråle M, Laurila E, Houstis N, Daly MJ, Patterson N, et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003; 34:267–73. https://doi.org/10.1038/ng1180. [PubMed].
27. Santibanez JF. Transforming growth factor-Beta and urokinase-type plasminogen activator: dangerous partners in tumorigenesis-implications in skin cancer. ISRN Dermatol. 2013; 2013:597927. https://doi.org/10.1155/2013/597927. [PubMed].
28. Smith HW, Marshall CJ. Regulation of cell signalling by uPAR. Nat Rev Mol Cell Biol. 2010; 11:23–36. https://doi.org/10.1038/nrm2821. [PubMed].
29. Pavón MA, Arroyo-Solera I, Céspedes MV, Casanova I, León X, Mangues R. uPA/uPAR and SERPINE1 in head and neck cancer: role in tumor resistance, metastasis, prognosis and therapy. Oncotarget. 2016; 7:57351–66. https://doi.org/10.18632/oncotarget.10344. [PubMed].
30. Yu XM, Jaskula-Sztul R, Georgen MR, Aburjania Z, Somnay YR, Leverson G, Sippel RS, Lloyd RV, Johnson BP, Chen H. Notch1 signaling regulates the aggressiveness of differentiated thyroid cancer and inhibits SERPINE1 expression. Clin Cancer Res. 2016; 22:3582–92. https://doi.org/10.1158/1078-0432.CCR-15-1749. [PubMed].
31. Sang Y, Chen MY, Luo D, Zhang RH, Wang L, Li M, Luo R, Qian CN, Shao JY, Zeng YX, Kang T. TEL2 suppresses metastasis by down-regulating SERPINE1 in nasopharyngeal carcinoma. Oncotarget. 2015; 6:29240–53. https://doi.org/10.18632/oncotarget.5074. [PubMed].
32. Dreos R, Ambrosini G, Périer RC, Bucher P. The Eukaryotic Promoter Database: expansion of EPDnew and new promoter analysis tools. Nucleic Acids Res. 2015; 43:D92–96. https://doi.org/10.1093/nar/gku1111. [PubMed].
33. Sharrocks AD. The ETS-domain transcription factor family. Nat Rev Mol Cell Biol. 2001; 2:827–37. https://doi.org/10.1038/35099076. [PubMed].
34. Kar A, Gutierrez-Hartmann A. Molecular mechanisms of ETS transcription factor-mediated tumorigenesis. Crit Rev Biochem Mol Biol. 2013; 48:522–43. https://doi.org/10.3109/10409238.2013.838202. [PubMed].
35. Oki S, Ohta T, Shioi G, Hatanaka H, Ogasawara O, Okuda Y, Kawaji H, Nakaki R, Sese J, Meno C. ChIP-Atlas: a data-mining suite powered by full integration of public ChIP-seq data. EMBO Rep. 2018; 19:e46255. https://doi.org/10.15252/embr.201846255. [PubMed].
36. Leemans CR, Snijders PJ, Brakenhoff RH. The molecular landscape of head and neck cancer. Nat Rev Cancer. 2018; 18:269–82. https://doi.org/10.1038/nrc.2018.11. [PubMed].
37. Lowell S, Jones P, Le Roux I, Dunne J, Watt FM. Stimulation of human epidermal differentiation by delta-notch signalling at the boundaries of stem-cell clusters. Curr Biol. 2000; 10:491–500. https://doi.org/10.1016/S0960-9822(00)00451-6. [PubMed].
38. Rangarajan A, Talora C, Okuyama R, Nicolas M, Mammucari C, Oh H, Aster JC, Krishna S, Metzger D, Chambon P, Miele L, Aguet M, Radtke F, Dotto GP. Notch signaling is a direct determinant of keratinocyte growth arrest and entry into differentiation. EMBO J. 2001; 20:3427–36. https://doi.org/10.1093/emboj/20.13.3427. [PubMed].
39. Nickoloff BJ, Qin JZ, Chaturvedi V, Denning MF, Bonish B, Miele L. Jagged-1 mediated activation of notch signaling induces complete maturation of human keratinocytes through NF-kappaB and PPARgamma. Cell Death Differ. 2002; 9:842–55. https://doi.org/10.1038/sj.cdd.4401036. [PubMed].
40. Alcolea MP, Greulich P, Wabik A, Frede J, Simons BD, Jones PH. Differentiation imbalance in single oesophageal progenitor cells causes clonal immortalization and field change. Nat Cell Biol. 2014; 16:615–22. https://doi.org/10.1038/ncb2963. [PubMed].
41. Lee TV, Sethi MK, Leonardi J, Rana NA, Buettner FF, Haltiwanger RS, Bakker H, Jafar-Nejad H. Negative regulation of Notch signaling by xylose. PLoS Genet. 2013; 9:e1003547. https://doi.org/10.1371/journal.pgen.1003547. [PubMed].
42. Upadhyay P, Nair S, Kaur E, Aich J, Dani P, Sethunath V, Gardi N, Chandrani P, Godbole M, Sonawane K, Prasad R, Kannan S, Agarwal B, et al. Notch pathway activation is essential for maintenance of stem-like cells in early tongue cancer. Oncotarget. 2016; 7:50437–49. https://doi.org/10.18632/oncotarget.10419. [PubMed].
43. Avila JL, Kissil JL. Notch signaling in pancreatic cancer: oncogene or tumor suppressor? Trends Mol Med. 2013; 19:320–27. https://doi.org/10.1016/j.molmed.2013.03.003. [PubMed].
44. Yu CC, Hsieh CR, Hsiao G, Chen PY, Chang ML, Yin HW, Lee TH, Lee CK. Regulated expressions of MMP-2, -9 by diterpenoids from Euphorbia formosana Hayata. Molecules. 2012; 17:2082–90. https://doi.org/10.3390/molecules17022082. [PubMed].
45. Yoshida R, Nagata M, Nakayama H, Niimori-Kita K, Hassan W, Tanaka T, Shinohara M, Ito T. The pathological significance of Notch1 in oral squamous cell carcinoma. Lab Invest. 2013; 93:1068–81. https://doi.org/10.1038/labinvest.2013.95. [PubMed].
46. Münch J, Grivas D, González-Rajal Á, Torregrosa-Carrión R, de la Pompa JL. Notch signalling restricts inflammation and serpine1 expression in the dynamic endocardium of the regenerating zebrafish heart. Development. 2017; 144:1425–40. https://doi.org/10.1242/DEV.143362. [PubMed].
47. Providence KM, Higgins SP, Mullen A, Battista A, Samarakoon R, Higgins CE, Wilkins-Port CE, Higgins PJ. SERPINE1 (PAI-1) is deposited into keratinocyte migration “trails” and required for optimal monolayer wound repair. Arch Dermatol Res. 2008; 300:303–10. https://doi.org/10.1007/s00403-008-0845-2. [PubMed].
48. Simone TM, Higgins CE, Czekay RP, Law BK, Higgins SP, Archambeault J, Kutz SM, Higgins PJ. SERPINE1: a molecular switch in the proliferation-migration dichotomy in wound-”activated” keratinocytes. Adv Wound Care (New Rochelle). 2014; 3:281–90. https://doi.org/10.1089/wound.2013.0512. [PubMed].
49. Freytag J, Wilkins-Port CE, Higgins CE, Higgins SP, Samarakoon R, Higgins PJ. PAI-1 mediates the TGF-β1+EGF-induced “scatter” response in transformed human keratinocytes. J Invest Dermatol. 2010; 130:2179–90. https://doi.org/10.1038/JID.2010.106. [PubMed].
50. Bajou K, Peng H, Laug WE, Maillard C, Noel A, Foidart JM, Martial JA, DeClerck YA. Plasminogen activator inhibitor-1 protects endothelial cells from FasL-mediated apoptosis. Cancer Cell. 2008; 14:324–34. https://doi.org/10.1016/j.ccr.2008.08.012. [PubMed].
51. Pickering CR, Zhang J, Yoo SY, Bengtsson L, Moorthy S, Neskey DM, Zhao M, Ortega Alves MV, Chang K, Drummond J, Cortez E, Xie TX, Zhang D, et al. Integrative genomic characterization of oral squamous cell carcinoma identifies frequent somatic drivers. Cancer Discov. 2013; 3:770–81. https://doi.org/10.1158/2159-8290.CD-12-0537. [PubMed].
52. Findlay VJ, LaRue AC, Turner DP, Watson PM, Watson DK. Understanding the role of ETS-mediated gene regulation in complex biological processes. Adv Cancer Res. 2013; 119:1–61. https://doi.org/10.1016/B978-0-12-407190-2.00001-0. [PubMed].
53. Sizemore GM, Pitarresi JR, Balakrishnan S, Ostrowski MC. The ETS family of oncogenic transcription factors in solid tumours. Nat Rev Cancer. 2017; 17:337–51. https://doi.org/10.1038/nrc.2017.20. [PubMed].
54. Goldie SJ, Mulder KW, Tan DW, Lyons SK, Sims AH, Watt FM. FRMD4A upregulation in human squamous cell carcinoma promotes tumor growth and metastasis and is associated with poor prognosis. Cancer Res. 2012; 72:3424–36. https://doi.org/10.1158/0008-5472.CAN-12-0423. [PubMed].
55. Ong CT, Cheng HT, Chang LW, Ohtsuka T, Kageyama R, Stormo GD, Kopan R. Target selectivity of vertebrate Notch proteins. J Biol Chem. 2006; 281:5106–19. https://doi.org/10.1074/jbc.M506108200. [PubMed].
56. Weber K, Bartsch U, Stocking C, Fehse B. A multicolor panel of novel lentiviral “gene ontology” (LeGO) vectors for functional gene analysis. Mol Ther. 2008; 16:698–706. https://doi.org/10.1038/mt.2008.6. [PubMed].
57. Guzmán C, Bagga M, Kaur A, Westermarck J, Abankwa D. ColonyArea: An ImageJ plugin to automatically quantify colony formation in clonogenic assays. PLoS One. 2014; 9:e92444. https://doi.org/10.1371/journal.pone.0092444. [PubMed].
58. Hudson DL, Weiland KL, Dooley TP, Simon M, Watt FM. Characterisation of eight monoclonal antibodies to involucrin. Hybridoma. 1992; 11:367–79. https://doi.org/10.1089/hyb.1992.11.367. [PubMed].
59. Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015; 43:e47. https://doi.org/10.1093/nar/gkv007. [PubMed].
60. Warnes GR, Bolker B, Bonebakker L, Gentleman R, Liaw WHA, Lumley T, Maechler M, Magnusson A, Moeller S, Schwartz M, Venables B. gplots: Various R programming tools for plotting data. R package version 2.17.0. 2015. p. 2015. https://doi.org/10.1111/j.0022-3646.1997.00569.x.
61. Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012; 2:401–04. https://doi.org/10.1158/2159-8290.CD-12-0095. [PubMed].
62. Anaya J. OncoLnc: linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. PeerJ Comput Sci. 2016; 2:e67. https://doi.org/10.7717/peerj-cs.67.

