Introduction
Pancreatic neuroendocrine tumors (PanNETs) have been established as a molecularly targetable group of cancers with approved inhibitors of mTOR and VEGFR/PDGFR/c-Kit that are available in the treatment paradigm for unresectable or advanced, well-differentiated disease [1, 2]. Comprehensive genomic profiling of 68 non-metastatic and metastatic PanNETs provided amongst the first glimpses of the molecular landscape for this malignancy and identified frequently recurring somatic mutations in MEN1 (44.1%), DAXX (25%), ATRX (17.6%), TSC2 (8.8%), PTEN (7.3%), and PIK3CA (1.4%) [3].
Recently, the Australian Pancreatic Cancer Genome Initiative (APGI) within the framework of the International Cancer Genome Consortium (ICGC) performed whole-genome sequencing of 102 localized, locally advanced, and metastatic PanNETs to further characterize pathogenic mechanisms and uncover novel mutations for therapeutic targeting [4]. In this seminal study, potentially actionable somatic alterations in 4 core pathways were identified: DNA damage repair (including MUTYH (mutational frequency of 6%), CHEK2 (4%), and BRCA2 (1%)), chromatin modification (including MEN1 (41%), SETD2 (5%), and MLL3 (5%)), altered telomere length (including DAXX (22%) and ATRX (10%)), and mTOR signaling (including PTEN (7%), DEPDC5 (2%), EWSR1 fusion (3%), TSC1 (2%), and TSC2 (2%)).
Despite the results of these landmark studies that have provided a framework for defining the most frequently altered genes and molecular pathways in PanNETs, their relationships to clinicopathologic characteristics remain poorly described. In this study, we describe our experience in cataloging alterations identified through multiplatform profiling of a cohort of predominantly locally advanced and metastatic PanNETs. In particular, we focused our analysis on co-occurring alterations of known pathogenicity and correlating molecular and pathway alterations with various clinicopathologic factors. We believe these analyses to be important in further defining molecular signatures and patient subsets of potential clinical relevance to PanNET management.
Results
Patient characteristics
From 12/2014–12/2018, a total of 46 patients diagnosed with predominantly locally advanced and metastatic PanNETs were included in this study (Table 1). The median age was 52 years (range 30–79) and the majority of cases (65%) were low or intermediate grade.
Table 1: Patient characteristics
| Characteristic (n = 46) | Frequency (%)* |
| Age (median) | 52 years (range 30–79) |
| Gender | |
| Female | 24 (52%) |
| Male | 22 (48%) |
| Stagea | |
| IV | 25 (54%) |
| III | 3 (7%) |
| II | 7 (15%) |
| I | 1 (2%) |
| Locally advanced | 10 (22%) |
| Gradeb | |
| Low/intermediate | 30 (65%) |
| High | 16 (35%) |
| Site of biopsy | |
| Liver | 23 (50%) |
| Pancreas | 18 (39.1%) |
| Lymph node | 2 (4.3%) |
| Peritoneum | 1 (2.2%) |
| Spleen | 1 (2.2%) |
| Soft tissue | 1 (2.2%) |
Molecular alterations
The most commonly detected somatic alterations including pathogenic variants and VUS by NGS are shown in Table 2. Mutations in MEN1 (41%), DAXX (28%), TP53 (20%), PTEN (18%), RB1 (13%), and ARID1A (11%) comprised the most frequently altered genes in our NGS cohort of 46 patients with PanNETs. PTEN (100%), mismatch repair (MMR) proteins (100%), TOP1 (64%), TUBB3 (64%), pAKT (58%), ERCC1 (27%), and TLE3 (25%) were among those most frequently expressed in our IHC cohort (Table 2). Analysis by in-situ hybridization did not identify any significant alterations across genes assayed: ALK, HER2, MET, ROS1, and TOP2A.
Table 2: Frequently detected molecular alterations detected by NGS and IHC
| Alteration* | Pathway Association | Frequency (%)+ |
| NGS (n = 46) | ||
| MEN1 | Chromatin remodeling | 19 (41%) |
| DAXX | Chromatin remodeling | 13 (28%) |
| TP53 | Replicative stress | 9 (20%) |
| PTEN | mTOR signaling | 8 (18%) |
| RB1 | Replicative stress | 6 (13%) |
| ARID1A | mTOR signaling | 5 (11%) |
| TSC2 | mTOR signaling | 4 (9%) |
| SETD2 | Chromatin remodeling | 4 (9%) |
| ATRX | Replicative stress | 3 (7%) |
| IHC (n = 28) | ||
| MMR | Microsatellite status/mismatch repair | 25/25 (100%) |
| PTEN | mTOR signaling | 12/12 (100%) |
| TOP1 | Taxane resistance | 7/11 (64%) |
| pAKT | mTOR signaling | 7/12 (58%) |
| TS | 5-FU resistance | 8/26 (31%) |
| ERCC1 | Platinum resistance | 7/26 (27%) |
| TLE3 | Wnt signaling | 3/12 (25%) |
| PD-1 | Immune checkpoint | 2/10 (20%) |
| RRM1 | Gemcitabine resistance | 1/13 (8%) |
| HER2 | HER2 signaling | 0/27 (0%) |
| PD-L1 | Immune checkpoint | 0/13 (0%) |
Correlative analyses
Comparisons of molecular alterations and clinicopathologic variables identified several significant correlations (Table 3). Alterations in MEN1 (next-generation sequencing or NGS) were less associated with having high-grade PanNETs, while alterations in RB1 (NGS), TP53 (NGS), and the replicative stress pathway were associated with having high-grade PanNETs. Alterations in MEN1 (NGS) and age >54 were less associated with metastatic disease at diagnosis. A history of diabetes was significantly associated with having an altered cyclin-dependent kinase (CDK) but less associated with having an altered replicative stress pathway. Several co-occurring alterations reached statistical significance: RB1 (NGS) and TP53 (NGS), MEN1 (NGS) with ERCC1 expression (immunohistochemistry or IHC) PTEN (NGS) and ARID1A (NGS), DAXX (NGS) and ERCC1 (IHC), PTEN (NGS) and TP53 (NGS), and MEN1 (NGS) with DAXX (NGS).
Table 3: Significant correlations between molecular alterations and clinicopathologic variables
Variable 1* | Variable 2* | OR+ | 95% CI | Fisher’s (two-tailed) p-value |
High grade tumor | RB1 (NGS) | Infinite | 2.82–Infinite | 0.0009 |
RB1 (NGS) | TP53 (NGS) | 12.73 | 1.43–175.82 | 0.0095 |
ERCC1 (IHC) | MEN1 (NGS) | 11.68 | 1.07–639.81 | 0.0261 |
High grade tumor | TP53 (NGS) | 10.22 | 1.58–118.17 | 0.0049 |
PTEN (NGS) | ARID1A (NGS) | 9.92 | 0.91–147.03 | 0.0307 |
History of diabetes | Cyclin-dependent kinase pathway | 8.42 | 0.94–95.00 | 0.0292 |
DAXX (NGS) | ERCC1 (IHC) | 8.40 | 0.96–120.72 | 0.0283 |
Replicative stress pathway | High grade tumor | 6.87 | 1.57–35.18 | 0.0043 |
PTEN (NGS) | TP53 (NGS) | 6.23 | 0.87–47.19 | 0.0359 |
MEN1 (NGS) | DAXX (NGS) | 5.74 | 1.23–32.41 | 0.0171 |
Metastatic at diagnosis | MEN1 (NGS) | 0.22 | 0.05–0.90 | 0.0288 |
Metastatic at diagnosis | Age >54 | 0.17 | 0.04–0.72 | 0.0070 |
High grade tumor | MEN1 (NGS) | 0.13 | 0.01–0.72 | 0.0105 |
History of diabetes | Replicative stress pathway | 0 | 0–0.65 | 0.0087 |
Cluster analysis allowed generation of a heatmap to visualize putative correlations across clinicopathologic variables in our PanNET cohort (Figure 1). Notably, NGS alterations in MEN1 and DAXX, both components of the chromatin remodeling pathway, tend to co-occur in non-metastatic, well-differentiated/intermediate grade PanNETS. NGS alterations in TP53 and RB1 tended to co-occur in high-grade tumors. SETD2 was mutually exclusive of these alterations within other low-grade tumors. No other associations across clinicopathologic variables and genomic alterations/IHC expression reached statistical significance beyond those described in Table 3.
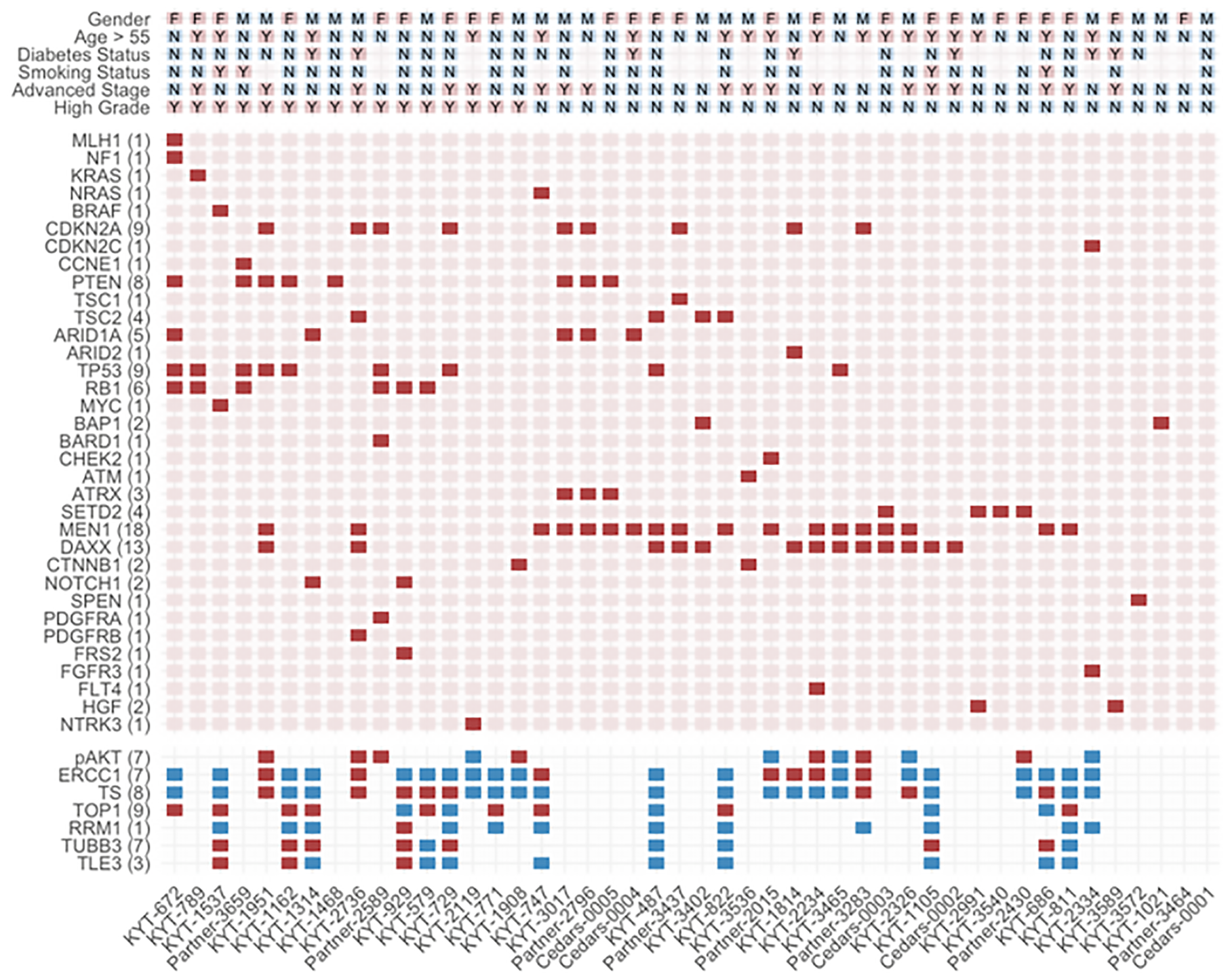
Figure 1: Heatmap illustrating subgroup associations across clinicopathologic variables using hierarchical clustering. Presence of select pathogenic genomic alterations are designated by solid red squares. Positive protein expression is designated by solid red squares while blue squares signify negative expression. Select clinicopathologic variables are designated by M for male gender, F for female gender, and Y or N for yes or no, respectively. Each column represents an individual case. Numbers of cases with genomic alterations or positive protein expression are displayed alongside row names.
Survival analysis
Molecular alterations in the replicative stress pathway were the only variables demonstrating significant correlation with worse OS in our PanNET cohort (Figure 2, hazard ratio (HR) 13.62 [95% confidence interval (CI): 1.51-122.5], p = 0.0198). This finding was primarily driven by alterations in TP53 and RB1. No variables were significantly correlated with progression-free survival (PFS) in our analysis.
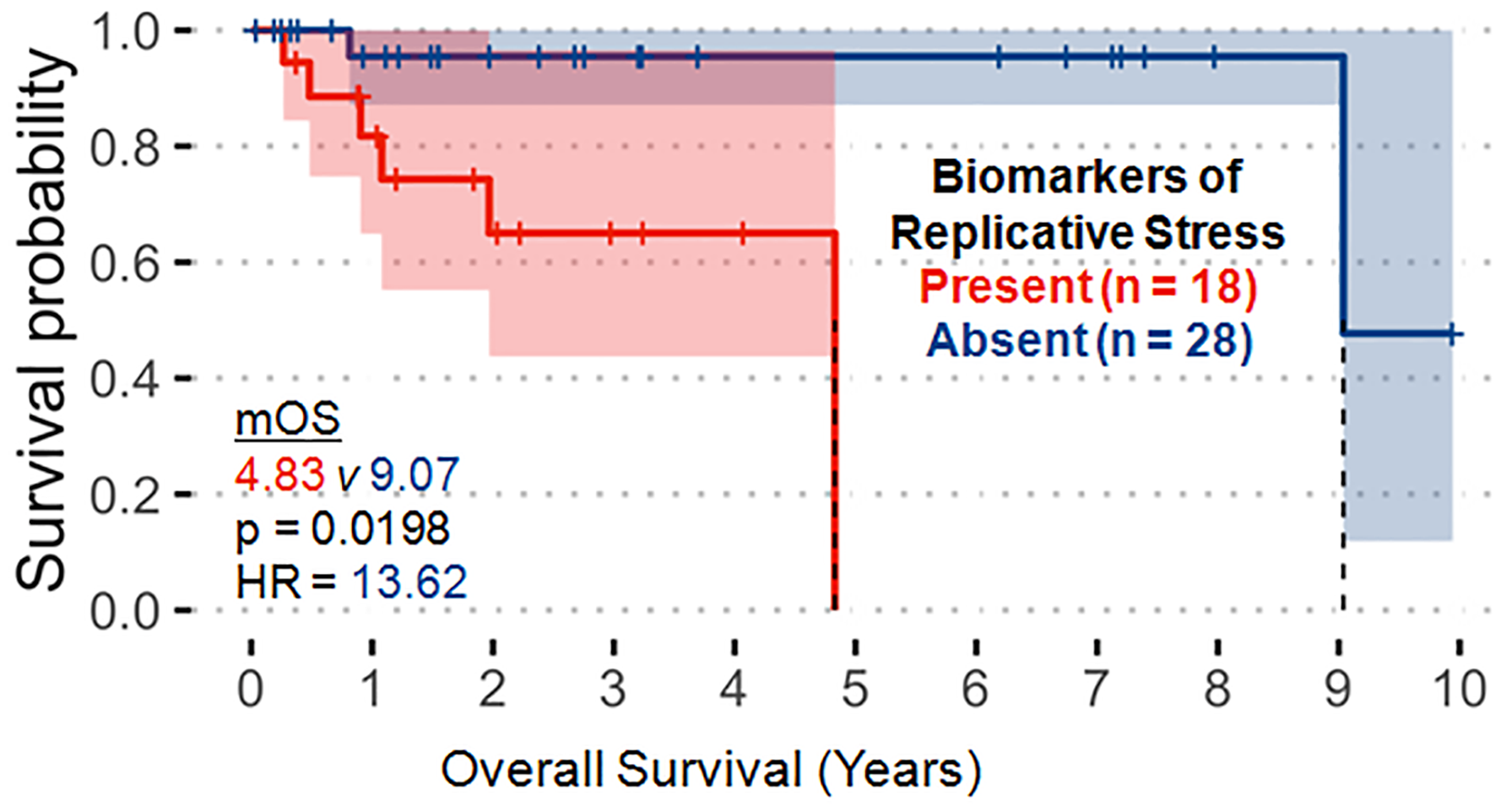
Figure 2: Molecular alterations associated with the replicative stress pathway (p53, RB1, ATRX, ATM, CCNE1, CHEK2, MYC) significantly correlate with decreased overall survival. These observations were primarily driven by TP53 and RB1 alterations.
DISCUSSION
Beyond well-established germline mutations, PanNETs are increasingly recognized as molecularly-driven tumors with somatic mutations affecting chromatin remodeling (MEN1, DAXX, and ATRX) and mTOR signaling (PTEN and TSC1/2) as key oncogenic drivers [5]. Associations between clinicopathologic factors and genomic and proteomic signatures, however, remain poorly described and most large-scale studies have limited their focus on correlating DAXX/ATRX to prognostic variables [3, 4, 6, 7]. To the best of our knowledge, we are among the first to evaluate associations of molecular alterations across a spectrum of clinicopathologic variables and identify co-occurring pathogenic mutations and altered molecular pathways of potential clinical significance in PanNETs.
Alterations in MEN1 and DAXX correlate with prognostic variables
Tumor stage and grade have long been known to be the most important independent predictors of survival in PanNETs [8, 9]. Our correlative analyses identified several significant associations with these 2 prognostic variables (Table 3). Alterations in MEN1 (NGS) were less associated with having high-grade PanNETs and metastatic disease at diagnosis. NGS alterations in MEN1 and DAXX, both components of the chromatin remodeling pathway, tend to co-occur in non-metastatic and well-differentiated/intermediate-grade PanNETs as visualized by hierarchical clustering (Figure 1), and a significant association between MEN1 and DAXX was reached on pairwise comparisons (Table 3). Mutations in ATRX have been shown to be mutually exclusive to DAXX mutations and were excluded from our analysis [4].
Our potential positive prognostic signature with MEN1 and DAXX alterations by NGS are consistent with an earlier NGS study that showed that alterations in MEN1, DAXX/ATRX, or the combination of were significantly associated with prolonged survival, in contrast to tumors lacking such mutations, in a metastatic PanNET cohort [3]. In predominantly non-metastatic and well-differentiated/intermediate grade PanNETs, however, loss of DAXX/ATRX by IHC correlated with higher tumor grade and stage, risk for metastasis, and worse survival [6, 7]. Recently, the ICGC analysis identified that median overall survival (OS) was not significantly different between DAXX/ATRX-mutant and -wild-type tumors, although DAXX/ATRX-mutant World Health Organization (WHO) grade 2 tumors had significantly worse OS compared to wild-type tumors [4]. Further study is needed to resolve this controversy regarding the prognostic value of chromatin remodeling alterations including MEN1, DAXX, and ATRX, which may be tumor stage and grade-specific. SETD2 was mutually exclusive of DAXX and MEN1 alterations within low-grade tumors in our cohort; the prognostic significance of this alteration may need further validation in larger PanNET cohorts.
Replicative stress pathway alterations as primarily driven by TP53 and RB1 correlate with worse overall survival
On our OS analysis, only alterations in the replicative stress pathway were significantly correlated with worse survival (Figure 2). Having an altered replicative stress pathway correlated with high-grade disease as well. Notably, these observations were primarily driven by TP53 (NGS) and RB1 (NGS) alterations, of which both were significantly associated with high-grade disease on pairwise comparison (Table 3) and co-occurred in high-grade tumors (Figure 1). Abnormal expression of TP53 and RB1 have been shown to define poorly differentiated PanNETs with worse disease-specific survival compared to loss of DAXX or ATRX in well-differentiated PanNETs which have much better survival [10].
To sum, alterations in the DNA replicative stress pathway, in particular TP53 and/or RB1 alterations, significantly correlate with a negative prognostic signature. Targeting of DNA damage repair /replicative stress pathways with systemic therapies such as poly(ADP-ribose) polymerase (PARP) inhibitors or checkpoint kinase 1 (CHEK1/CHK1) inhibitors or cell cycle pathways with CDK inhibitors have yet to be as established in PanNETs as in other cancers, but our findings suggest a potentially targetable subset of PanNET patients with DNA replicative stress pathway alterations and poor prognosis that are primarily driven by RB1 and TP53 [11–13]. Future investigation is warranted to test these hypotheses in PanNETs.
Other associations of potential clinical significance
We identified several other significant correlations of potential clinical importance on pairwise comparisons (Table 3): history of diabetes with CDK (positive correlation) or replicative stress pathway alterations (negative correlation), age >54 with non-metastatic disease, ERCC1 expression (IHC) with DAXX or MEN1 alterations (NGS), and PTEN (NGS) with ARID1A or TP53 alterations (NGS). ERCC1 expression has been shown to be putative prognostic and predictive biomarkers across various solid malignancies [14, 15]. In our series, ERCC1 expression positively correlated with favorable prognostic alterations (DAXX and MEN1). In separate studies of PanNET patients, loss of PTEN and ARID1A were associated with poorer prognosis [16, 17]. This appears consistent with our findings where PTEN co-occurred with ARID1A as well as the unfavorable prognostic TP53 alteration.
The rationale behind the associations between a history of diabetes and alterations in the CDK or replicative stress pathway are unclear, but an altered replicative stress pathway portended to a poorer OS in our cohort (Figure 2). Age >54 was significantly less associated with metastatic disease at diagnosis (Table 3). This finding is contradictory to evidence where increasing age has historically portended a worse prognosis in large PanNET cohorts [8, 9]. Instead, this finding is more likely due to the fact that the majority of our metastatic or high-grade PanNET patients were age ≤54 (Figure 1). The tendency for these molecular alterations and clinicopathologic features of various prognostic/predictive significance to co-occur in our cohort are hypothesis generating and provide impetus for subsequent validation in larger cohorts to define combinations of genomic/proteomic signatures of greatest prognostic and/or predictive value in PanNETs.
Lastly, it should be noted that this study was limited by its retrospective design and heterogeneity across patient characteristics and tumor biology. Our decision to include pathogenic variations only for analysis likely further limited our sample size though we believed this decision was justified in conducting our survival and correlative analyses to prognostic variables. Nevertheless, we identified several molecular signatures through multi-omic profiling of potential clinical significance that are hypothesis generating and can inform future investigations of larger and ideally prospective design. Increasing efforts to molecularly profile PanNETs will enhance the possibility to stratify PanNET patients into molecularly-defined treatment subgroups based on prognostic and predictive biomarkers.
Materials and Methods
Study population and tumor samples
Informed consent was obtained from patients with pancreatic cancers through an IRB-approved protocol that facilitated the collection of real-world outcomes for research purposes. Patients diagnosed with histologically-confirmed pancreatic neuroendocrine tumors (PanNETs) were referred to Perthera, Inc. (McLean, VA) between 12/2014–12/2018 by their treating physician at a partnering institution or via the Pancreatic Cancer Action Network’s Know Your Tumor® (KYT) program (Figure 3). Participating patients received personalized treatment recommendations from Perthera’s molecular tumor board based on CLIA/CAP-certified genomic and proteomic testing results that were ordered from commercial testing laboratories. The analysis cohort in this study included histologically confirmed cases of PanNET in which NGS testing results were considered adequately sensitive. Multiplatform profiling data was also available for all but 19 PanNET cases which included IHC protein testing results for chemopredictive markers (e.g. RRM1, ERCC1, TYMS) and other predictive markers (PD-L1, HER2). All PanNET cases were categorized based on the WHO criteria [18]. There were no exclusions to site of biopsy, previous treatment or lines of prior therapy, disease stage, medical comorbidities, or performance status.
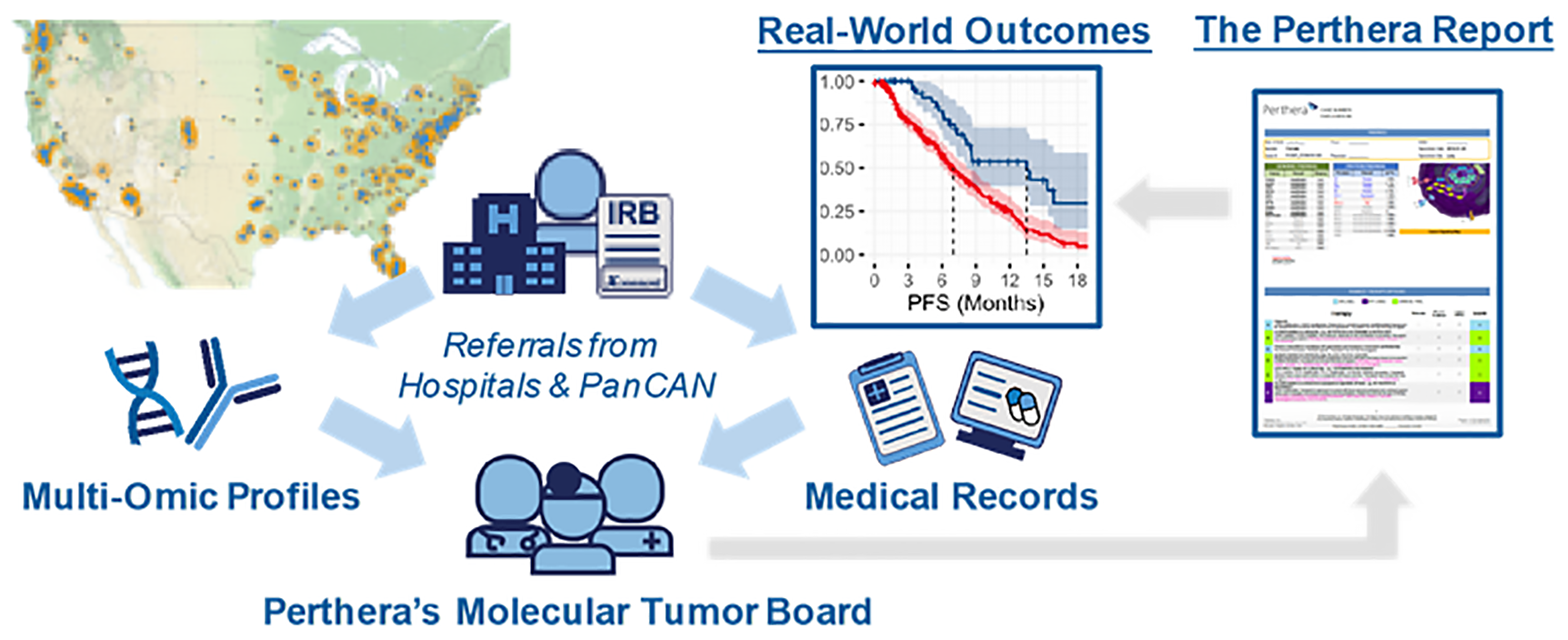
Figure 3: Perthera’s precision oncology process for curating real-world outcomes leverages the expertise of a molecular tumor board for actionability assessment. Perthera’s institutional review board-approved registry includes pancreatic neuroendocrine tumor patients from multiple cancer centers and patients referred via the Pancreatic Cancer Action Network’s Know Your Tumor® (KYT) program. Real-world outcomes were collected after cases were reviewed by Perthera’s Molecular Tumor Board to deliver personalized treatment recommendations based on molecular findings. Next-generation DNA sequencing (NGS) was performed by Foundation Medicine and proteomics/immunohistochemistry (IHC) by Neogenomcs Inc. or Caris Life Sciences, Inc.
Multiplatform profiling
The processing of formalin-fixed paraffin-embedded (FFPE) tissues was performed in accordance to requirements per specific platform as previously described [19, 20]. NGS was performed on submitted tumor samples through FoundationOne (Foundation Medicine, Inc., Cambridge, MA). The FoundationOne NGS assay, which detects all 4 classes of genomic alterations including base substitutions, insertions and deletions, copy number alterations (CNAs), and rearrangements, has been previously described and validated [19]. Analysis by IHC was performed on FFPE tumor tissues through Caris Life Sciences (Phoenix, AZ) and NeoGenomics, Inc. (Fort Myers, FL) using commercially available detection kits and autostainers along with commercially available antibodies as previously described [20–22].
Statistical analysis
Real-world outcomes data including demographics information, clinicopathologic factors, and survival metrics were abstracted from medical records and analyzed alongside molecular testing results from commercial laboratories. Descriptive statistics and other statistical analyses were computed in an R/Bioconductor programming environment. OS and PFS were evaluated for pairwise correlative analyses between individual pathogenic alterations or altered molecular pathways and clinicopathologic variables. Odds ratios (ORs) and 95% CIs were calculated using Cox proportional hazards regression as implemented in the R survival package. Statistical significance was defined by a Fisher’s exact test (two-tailed) p ≤ 0.05.
Pathogenic alterations were grouped into the following molecular pathways: replicative stress (p53, RB1, ATRX, ATM, CCNE1, CHEK2, MYC), chromatin remodeling (ARID1A, ARID2, ASXL1, CHD2, CTCF, DAXX, KMT2D, MEN1, MLL3, PBRM1, SETD2, TET2), cyclin-dependent kinase (CDKN1B, CDKN2A, CDKN2B, CDKN2C, CDK6), WNT signaling (CTNNB1, FAT1, GNAS, RNF43), mTOR signaling (MTOR, PIK3CA, PTEN, FLCN, TSC2), and receptor tyrosine kinase (RTK) signaling (BRAF, ERBB3, EPHA5, FGFR3, FRS2, FLT4, KRAS, NRAS, NTRK3). Only pathogenic alterations were included for correlative and survival analyses, and variants of uncertain significance (VUS) were excluded.
ACKNOWLEDGMENTS
We wish to acknowledge the Pancreatic Cancer Action Network (PanCAN), PanCAN’s Patient Central and Know Your Tumor initiatives, and the patients and their families who were involved in this study.
CONFLICTS OF INTEREST
E.M.B., E.F.P., M.J.P., and G.G. hold ownership interest (including patents) in Perthera. E.F.P., M.J.P., and A.E.H. are consultant/advisory board members for Perthera. No conflicts of interest were reported by J.G., J.R.B., M.G., V.P., and R.T.
FUNDING
None.
References
1. Raymond E, Dahan L, Raoul JL, Bang YJ, Borbath I, Lombard-Bohas C, Valle J, Metrakos P, Smith D, Vinik A, Chen JS, Hörsch D, Hammel P, et al. Sunitinib malate for the treatment of pancreatic neuroendocrine tumors. N Engl J Med. 2011; 364:501–13. https://doi.org/10.1056/NEJMoa1003825. [PubMed].
2. Yao JC, Shah MH, Ito T, Bohas CL, Wolin EM, Van Cutsem E, Hobday TJ, Okusaka T, Capdevila J, de Vries EG, Tomassetti P, Pavel ME, Hoosen S, et al. Everolimus for advanced pancreatic neuroendocrine tumors. N Engl J Med. 2011; 364:514–23. https://doi.org/10.1056/NEJMoa1009290. [PubMed].
3. Jiao Y, Shi C, Edil BH, de Wilde RF, Klimstra DS, Maitra A, Schulick RD, Tang LH, Wolfgang CL, Choti MA, Velculescu VE, Diaz LA, Vogelstein B, et al. DAXX/ATRX, MEN1, and mTOR pathway genes are frequently altered in pancreatic neuroendocrine tumors. Science. 2011; 331:1199–203. https://doi.org/10.1126/science.1200609. [PubMed].
4. Scarpa A, Chang DK, Nones K, Corbo V, Patch AM, Bailey P, Lawlor RT, Johns AL, Miller DK, Mafficini A, Rusev B, Scardoni M, Antonello D, et al. Whole-genome landscape of pancreatic neuroendocrine tumours. Nature. 2017; 543:65–71. https://doi.org/10.1038/nature21063. [PubMed].
5. Mafficini A, Scarpa A. Genomic landscape of pancreatic neuroendocrine tumours: the International Cancer Genome Consortium. J Endocrinol. 2018; 236:R161–R7. https://doi.org/10.1530/JOE-17-0560. [PubMed].
6. Marinoni I, Kurrer AS, Vassella E, Dettmer M, Rudolph T, Banz V, Hunger F, Pasquinelli S, Speel EJ, Perren A. Loss of DAXX and ATRX are associated with chromosome instability and reduced survival of patients with pancreatic neuroendocrine tumors. Gastroenterology. 2014; 146:453–60. https://doi.org/10.1053/j.gastro.2013.10.020. [PubMed].
7. Singhi AD, Liu TC, Roncaioli JL, Cao D, Zeh HJ, Zureikat AH, Tsung A, Marsh JW, Lee KK, Hogg ME, Bahary N, Brand RE, McGrath KM, et al. Alternative lengthening of telomeres and loss of DAXX/ATRX expression predicts metastatic disease and poor survival in patients with pancreatic neuroendocrine tumors. Clin Cancer Res. 2017; 23:600–9. https://doi.org/10.1158/1078-0432.CCR-16-1113. [PubMed].
8. Ellison TA, Wolfgang CL, Shi C, Cameron JL, Murakami P, Mun LJ, Singhi AD, Cornish TC, Olino K, Meriden Z, Choti M, Diaz LA, Pawlik TM, et al. A single institution’s 26-year experience with nonfunctional pancreatic neuroendocrine tumors: a validation of current staging systems and a new prognostic nomogram. Ann Surg. 2014; 259:204–12. https://doi.org/10.1097/SLA.0b013e31828f3174. [PubMed].
9. Rindi G, Falconi M, Klersy C, Albarello L, Boninsegna L, Buchler MW, Capella C, Caplin M, Couvelard A, Doglioni C, Delle Fave G, Fischer L, Fusai G, et al. TNM staging of neoplasms of the endocrine pancreas: results from a large international cohort study. J Natl Cancer Inst. 2012; 104:764–77. https://doi.org/10.1093/jnci/djs208. [PubMed].
10. Tang LH, Basturk O, Sue JJ, Klimstra DS. A practical approach to the classification of WHO grade 3 (G3) well-differentiated neuroendocrine tumor (WD-NET) and poorly differentiated neuroendocrine carcinoma (PD-NEC) of the pancreas. Am J Surg Pathol. 2016; 40:1192–202. https://doi.org/10.1097/PAS.0000000000000662. [PubMed].
11. Brown JS, O’Carrigan B, Jackson SP, Yap TA. Targeting DNA repair in cancer: beyond PARP inhibitors. Cancer Discov. 2017; 7:20–37. https://doi.org/10.1158/2159-8290.CD-16-0860. [PubMed].
12. Grande Pulido E, Teule A, Alonso-Gordoa T, Jiménez-Fonseca P, Benavent M, Capdevila J, Custodio A, Vera R, Munarriz J, La Casta-Muñoa A, Garcia-Carbonero R. A phase II trial of palbociclib in metastatic grade 1/2 pancreatic neuroendocrine tumors: the PALBONET study on behalf of the Spanish Taskforce Group of Neuroendocrine Tumors (GETNE). Ann Oncol. 2017; 28:4290.
13. McNeely S, Conti C, Sheikh T, Patel H, Zabludoff S, Pommier Y, Schwartz G, Tse A. Chk1 inhibition after replicative stress activates a double strand break response mediated by ATM and DNA-dependent protein kinase. Cell Cycle. 2010; 9:995–1004. https://doi.org/10.4161/cc.9.5.10935. [PubMed].
14. Choueiri MB, Shen JP, Gross AM, Huang JK, Ideker T, Fanta P. ERCC1 and TS expression as prognostic and predictive biomarkers in metastatic colon cancer. PLoS One. 2015; 10:e0126898. https://doi.org/10.1371/journal.pone.0126898. [PubMed].
15. Jiang J, Liang X, Zhou X, Huang R, Chu Z, Zhan Q. ERCC1 expression as a prognostic and predictive factor in patients with non-small cell lung cancer: a meta-analysis. Mol Biol Rep. 2012; 39:6933–42. https://doi.org/10.1007/s11033-012-1520-4. [PubMed].
16. Jayakumar R, Lanjewar S, Axiotis CA. Loss of PTEN and Increased pAKT Expression Distinguishes Aggressive Low-grade Neuroendocrine Tumors. Ann Clin Lab Sci. 2018; 48:565–72. [PubMed].
17. Roy S, LaFramboise WA, Liu TC, Cao D, Luvison A, Miller C, Lyons MA, O’Sullivan RJ, Zureikat AH, Hogg ME, Tsung A, Lee KK, Bahary N, et al. Loss of Chromatin-Remodeling Proteins and/or CDKN2A Associates With Metastasis of Pancreatic Neuroendocrine Tumors and Reduced Patient Survival Times. Gastroenterology. 2018; 154:2060–3.e8. https://doi.org/10.1053/j.gastro.2018.02.026. [PubMed].
18. Bosman FT, Carneiro F, Hruban RH, Theise ND. WHO classification of tumours of the digestive system. World Health Organization. 2010. Fourth Edition.
19. Frampton GM, Fichtenholtz A, Otto GA, Wang K, Downing SR, He J, Schnall-Levin M, White J, Sanford EM, An P, Sun J, Juhn F, Brennan K, et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol. 2013; 31:1023–31. https://doi.org/10.1038/nbt.2696. [PubMed].
20. Millis SZ, Ikeda S, Reddy S, Gatalica Z, Kurzrock R. Landscape of phosphatidylinositol-3-kinase pathway alterations across 19 784 diverse solid tumors. JAMA Oncol. 2016; 2:1565–1573. https://doi.org/10.1001/jamaoncol.2016.0891. [PubMed].
21. Herzog TJ, Spetzler D, Xiao N, Burnett K, Maney T, Voss A, Reddy S, Burger R, Krivak T, Powell M, Friedlander M, McGuire W. Impact of molecular profiling on overall survival of patients with advanced ovarian cancer. Oncotarget. 2016; 7:19840–9. https://doi.org/10.18632/oncotarget.7835. [PubMed].
22. Millis SZ, Bryant D, Basu G, Bender R, Vranic S, Gatalica Z, Vogelzang NJ. Molecular profiling of infiltrating urothelial carcinoma of bladder and nonbladder origin. Clin Genitourin Cancer. 2015; 13:e37–e49. https://doi.org/10.1016/j.clgc.2014.07.010. [PubMed].

