Introduction
Breast cancer is the first cancer in terms of incidence among women, with 1.7 million new cases in 2012 worldwide. It is also the second cause of death from cancer in western countries with 40,000 deaths per year [1, 2]. In the last 20 years, breast cancer mortality has continuously decreased as a result of mass screening programs and early diagnosis, but also as a consequence of improved treatment for both localized and metastatic disease [3, 4].
In the era of personalized cancer medicine, advances in genomics are essential assets. In this review, we will address current knowledge in genomics applied to the treatment of breast cancers.
HISTOLOGIC AND MOLECULAR CLASSIFICATIONS
There are schematically three main histologic types of breast cancer (Figure 1): i) estrogen-dependent breast cancers expressing the estradiol receptor (ER) and treated with a panel of drugs that target the estradiol receptor pathway [5]; ii) breast cancers overexpressing the human epidermal growth factor receptor 2 (HER2) oncoprotein and treated with anti-HER2-based chemotherapies, the first anti-HER2 being a therapeutic monoclonal antibody, trastuzumab [6]; and iii) “triple negative” breast cancers which lack the expression of the estradiol receptor, the progesterone receptor, and HER2. There are still no targeted therapies for triple-negative breast cancers, which have a high metastatic potential, and consequently a bad prognosis [7].
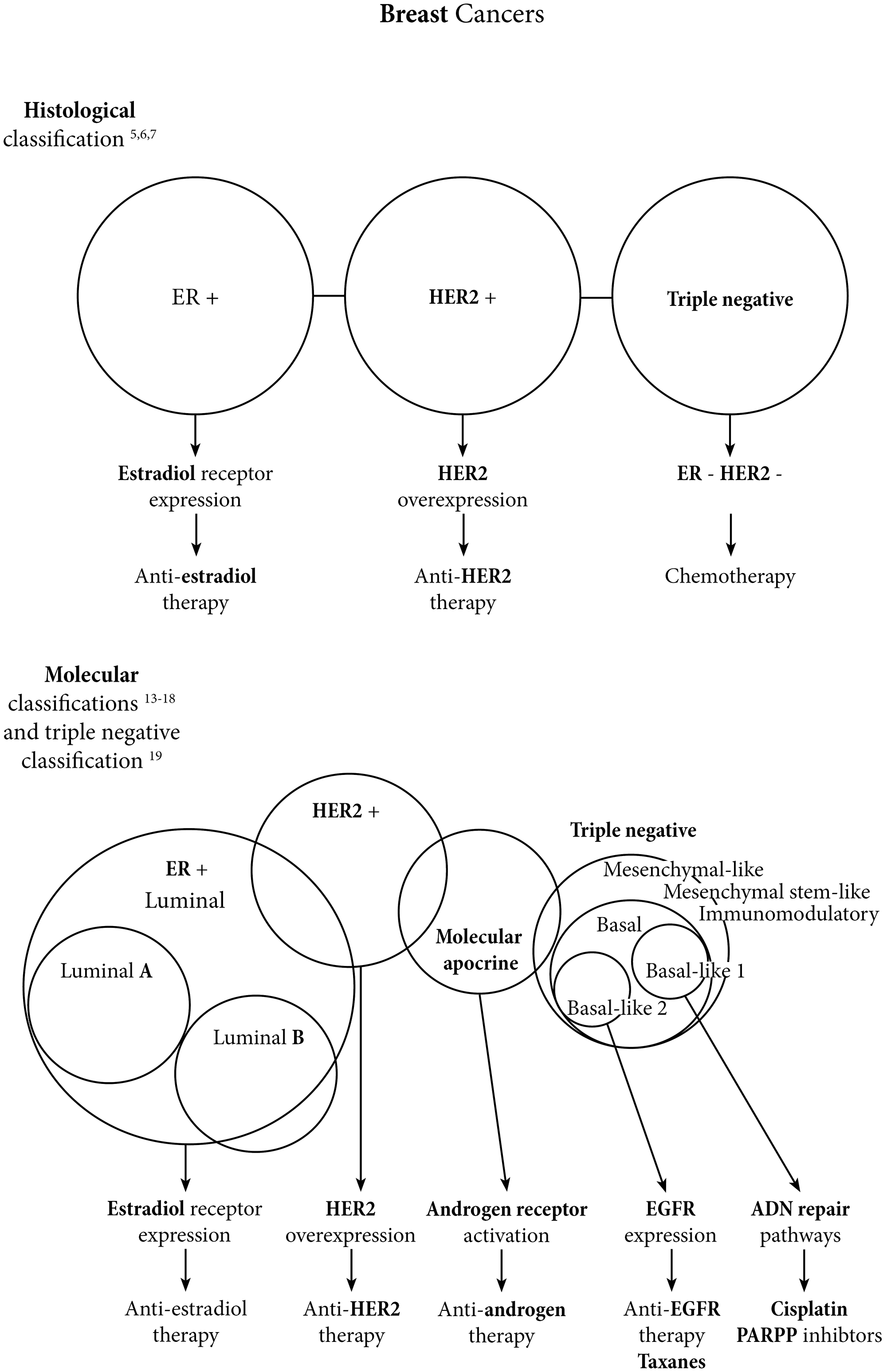
Figure 1: Breast cancers landscape evolution from histologic to molecular classifications.
Since 2000, several molecular subtypes have been identified, including the aforementioned histological subtypes (Figure 1). Peru et al. identified six molecular sub-types: Luminal A, Luminal B, Luminal C, HER2-enriched, Basal-like and Normal-like [8–10]. The luminal A subtype has the best prognosis and the lowest proliferative potential, including highly estrogen-dependent breast cancers. Luminal B subtype is also estrogen-dependent, but to a lesser extent, and corresponds to a sub-group of more aggressive cancers with a higher proliferative potential [11]. For the basal-like subtype, 50 to 75% are triple-negative breast cancers. Their aggressiveness is frequently the result of the loss of the retinoblastoma protein 1 (pRb1), of a high rate of tumor protein 53 (TP53) gene mutations and also of mutations in genes involved in DNA-repair mechanisms, such as breast cancer type 1 susceptibility (BRCA1) gene [12].
This early molecular classification was later revised, with the following subtypes: Luminal A and B, HER2-positive/ER-positive, HER2-positive/ER-negative, basal p53-altered and basal p53-normal (Figure 1). In this more recent version, 7% of cancers are of unknown sub-type because of the heterogenic expression of markers [13, 14]. Since 2005, among ER-negative cancers, several teams have identified a “molecular apocrine” subtype, characterized by an activation of the androgen receptor (AR) pathway and by the expression of AR target genes in 50% of cases, or by HER2 overexpression [15–17]. In 2012, a new molecular classification was established which includes Luminal, molecular Apocrine and Basal-like subtypes (Figure 1) [18]. Different immunohistochemistry markers have been proposed for in situ characterization of molecular apocrine breast cancers, including androgen receptor (AR) and gross cystic disease fluid protein 15 (GCDFP15) [17].
In 2011, Lehman et al. proposed a molecular classification of triple-negative breast cancers which we will discuss later [19].
MOLECULAR SIGNATURES AND ESTROGEN-DEPENDENT BREAST CANCERS
For estrogen-dependent breast cancers, the contribution of genomics has been small. However, molecular signatures have substantially modified clinical practice, enabling the therapeutic decision for adjuvant chemotherapy to be redefined for localized breast cancers [20]. Three commercially available tests are currently used: MammaPrint® (Agilent, The Netherlands) [21, 22], OncotypeDx® [23] and PAM50 (Prediction of Microarray using 50 classifier genes plus 5 reference genes) (Prosigna® kit) (see Table 1) [24, 25]. These tests quantify the expression levels of a limited panel of genes in the primary tumor. Most of them were developed for formalin-fixed paraffin-embedded tissue samples for implementation in daily practice.
Table 1: Gene panel tests used for therapeutic decision of localized breast cancers (Adapted from [28])
| Signature | Number of genes | Clinical application | Risk category | References |
|---|---|---|---|---|
| MammaPrint | 70 | N−, ER+ or ER− Estimates relapse risk | Low and high | [22] |
| OncotypeDX | 21 | ER+, HER2−, N− Estimates chemotherapy benefit and relapse risk during hormonotherapy | Low, intermediate and high | [23] |
| EndoPredict | 11 | ER+, HER2−, N− or N+ Predicts local and metastatic relapse during hormonotherapy | Low and high | [74] |
| Prosigna (PAM50) | 50 | ER+/N− and N+ treated by hormonotherapy Predicts 10-year metastasis-free survival | Low, intermediate and high | [25] |
| Breast Cancer Index | 5 and 2 genes ratio | ER+, N− Estimates metastatic risk Predicts late metastatic risk and efficacy of prolonged hormonotherapy | Low and high | [75] |
| Rotterdam | 76 | ER+, N− Predicts relapse under treatment with tamoxifen | Low and high | [76] |
| BluePrint | 80 | Discriminates sub-types with different level of sensitivity to adjuvant treatment | Not applicable | [77] |
These molecular signatures are used to classify patients according to their risk of metastatic relapse, to guide the decision for adjuvant chemotherapy when conventional criteria are insufficient [20]. This is particularly true for ER-positive, HER2-negative breast cancers without lymph node involvement (N0): in this sub-group of patients, adjuvant chemotherapy significantly reduces the risk of metastatic relapse only for high-risk patients [26–28]. The OncotypeDx® signature comprises three risk categories, raising the question of how to treat «intermediate-risk» patients. Recently, the TAILORx study clearly demonstrated the absence of benefit from adjuvant chemotherapy in this subgroup of intermediate-risk patients [29]. A meta-analysis of 147 articles concluded that molecular signatures for breast cancer enable 10% of patients at high clinical risk of relapse to be reclassified as low-risk patients, thus reducing the use of chemotherapy, with a favorable cost-efficiency ratio and improved quality of life for non-treated patients [30].
For estrogen-dependent breast cancers, the other contributions of genomics remain in the research field. For example, recent studies have identified the presence of mutations of the estrogen receptor 1 (ESR1) gene, such as the D538G or Y537S/C/N mutations, associated with resistance to anti-estrogens. These mutations change the conformation of the ligand binding site, thereby reducing the affinity of tamoxifen for the estrogen receptor [31, 32]. The systematic screening for these mutations is not currently recommended. Further studies are required to demonstrate their possible usefulness in guiding hormone-therapy prescription in daily practice.
GENOMICS APPLIED TO THE TREATMENT OF HER2-OVEREXPRESSING BREAST CANCERS
It is recommended to determine HER2 status in the primary tumor or in metastatic samples using a standardized immunostaining method. For doubtful cases, in situ hybridization methods are currently used to determine the HER2 gene copy number [33, 34]. In 2013, an international consensus clearly defined the criteria for HER2 protein overexpression and for HER2 gene amplification [35].
However, these methods entail certain limitations, typically for micro-invasive foci in a primary tumor or a micro-metastatic axillary lymph node [36].
Digital droplet PCR (ddPCR), easier to implement than in situ hybridization methods, seems to be a reliable alternative for the evaluation of the HER2 copy number in breast or gastric cancers [37–39]. A major limitation could be intra-tumor heterogeneity. Indeed, primary breast cancers are heterogeneous [28], and this is also the case for HER2 status [40]. Combining laser-microdissection with ddPCR overcomes this limitation, by enabling a precise assessment of the HER2 copy number within a cancer sample. In a recent study, we validated the use of laser microdissection combined with ddPCR to assess HER2 copy number in micro-invasive breast cancers with at least 50 invasive cancer cells. We then applied this methodology to a 45-year-old patient with extensive in situ breast cancer, and no associated micro-invasion except a micro-metastasis found only on one section of the sentinel axillary lymph node. We first laser-microdissected the micro-metastatic foci, and then used ddPCR to demonstrate that HER2 was amplified. This led us to optimize the adjuvant treatment for our patient, and she received trastuzumab-based adjuvant chemotherapy [37].
For HER2-overexpressing breast cancers, another issue remains to be deciphered: is there any clinical benefit to be drawn from the systematic determination of HER2 amplification level within the tumor? Singer et al. demonstrated a correlation between the level of HER2 amplification and the response to anti-HER2 treatments in the neoadjuvant setting [41]. Laser-microdissection of cancer cells from a metastatic tumor biopsy combined with ddPCR could be used to overcome signal dilution by enrichment with tumor cells. So, in a skin metastasis of a HER2-expressing breast cancer, we have demonstrated that HER2 copy number evaluated by ddPCR passed from 6 to 34, without and with prior laser microdissection respectively (Figure 2).
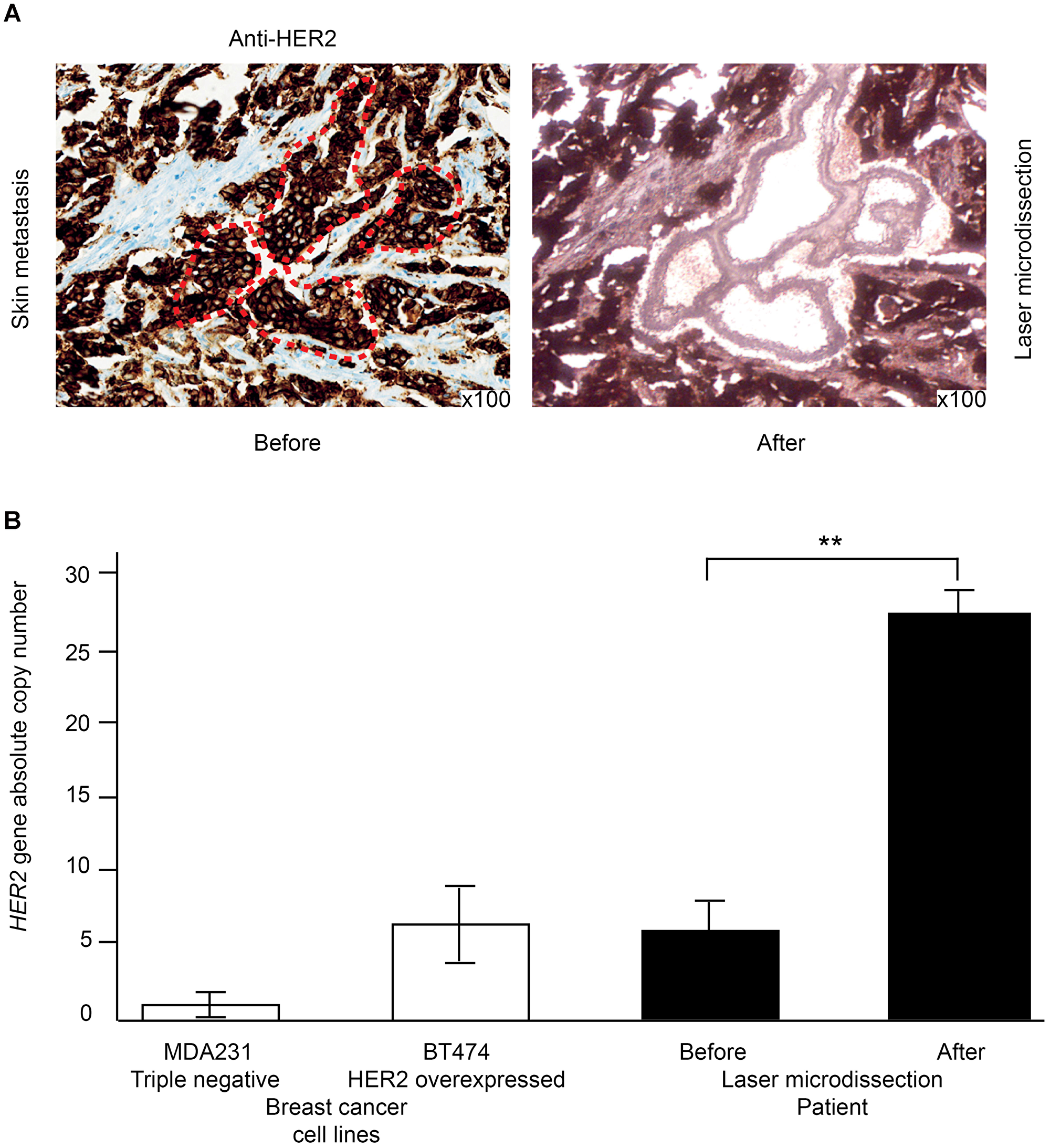
Figure 2: Laser-microdissection of cancer cells combined with ddPCR to precisely assess HER2 amplification level on a skin metastasis of recurrent HER2-overexpressing breast cancer. (A) HER2 is typically overexpressed using immunohistochemistry (left panel). The right panel shows the laser-microdissected HER2-overexpressing cancer cells. (B) the HER2 copy number is much higher in the laser-microdissected cells than in the whole tumor. MDA231 triple-negative breast cancer cell lines serve as a negative control while the BT474 HER2-overexpressing cancer cell line serves as a positive control.
GENOMICS APPLIED TO THE TREATMENT OF TRIPLE-NEGATIVE BREAST CANCERS
Triple-negative breast cancers remain a therapeutic challenge. Lehmann et al. established a molecular classification with six molecular subtypes to personalize the treatment of these breast cancer sub-types: Basal-like 1 (BL1), Basal-like 2 (BL2), Immunomodulatory (IM), Mesenchymal-like (M), Mesenchymal stem-like (MSL), and Luminal androgen receptor (LAR) (Figure 1) [19]. In the neoadjuvant setting, Masuda et al. demonstrated significant associations between these molecular subtypes and pathological complete response under chemotherapy. BL1 subtype is associated with greater chemo-sensitivity, whereas BL2 and LAR subtypes are more chemo-resistant subtypes [42].
Unfortunately, this classification still has few clinical applications, even in clinical trials dedicated to triple-negative breast cancers. For example, the BL2 subtype, characterized by an activation of the epidermal growth factor pathway, could respond to anti-epidermal growth factor receptor (EGFR) therapies. A phase II study assessed the benefit of cetuximab, a monoclonal anti-EGFR antibody, in the treatment of 173 women with metastatic triple-negative breast cancers. The results were disappointing, with a non-significant survival gain of 2.2 months [43]. For anti-EGFR-based treatments, a study could be dedicated exclusively to BL2 subtypes, ideally with high EGFR copy numbers and a quadruple wild-type status of KRAS, NRAS, BRAF, and PIK3CA genes, as for colon cancers [43, 44].
The same arguments can be applied to the LAR subtype, characterized by androgen receptor signaling pathway activation, and corresponding to certain molecular apocrine sub-types [16]. The LAR subtype accounts for 11% of triple-negative breast cancers [19]. In two phase II studies using anti-androgens for the treatment of patients with metastatic triple-negative breast cancers, disease stabilization at 6 months was observed for less than 20% of patients [45, 46]. It should be noted that the patients were not pre-selected according to LAR subtype.
In 2012 and 2013, we conducted a pilot study among five women with metastatic triple-negative breast cancers. For the five patients, we performed transcriptomic analyses on metastasis biopsies, and classified their respective metastatic cancers according to Lehmann’s classification. We also established individual xenograft models from the same metastasis biopsies. For each patient xenograft model, we tested a panel of drugs or drug combinations, guided by transcriptomic data. One patient was classified BL2, with EGF pathway activation and no mutation of the EGF pathway genes. In the corresponding xenograft, the most effective regimen was a combination of paclitaxel and cetuximab. This regimen was offered to the patient as a third-line resort treatment with almost complete metabolic response [47]. On the basis of transcriptomic analyses and chemosensitivity data obtained from the different xenografts, we personalized the resort treatment for the four other women in our study. In all cases, despite the fact that this resort treatment was third-line or fourth-line, the time-to-progression was longer than that observed with previous lines of treatment [48].
GENOMICS AND TUMOR HETEROGENEITY
Tumor heterogeneity is probably insufficiently taken into account in daily clinical practice, particularly for the treatment of metastatic disease. Most molecular analyses are performed on primary tumors, even in metastatic stages. However, metastatic clones can be a minority in the primary tumors they are deriving from [49, 50]. For HER2 status assessment, we have previously noted the benefit of combining molecular and tissue analyses, particularly with the contribution of laser-microdissection to overcome the limitation of tumor heterogeneity [37–39, 51, 52].
Molecular analyses on metastases are rare, mainly because of difficulties in obtaining these samples, and despite the fact that radiology-guided biopsies have considerably reduced this limitation [53]. For breast cancer, eleven studies have been dedicated to whole-genome analyses of metastatic biopsies (Table 2). In addition, most of these studies only included small numbers of patients, and the genome analyses were generally performed on tumors that were not laser-microdissected. One large study included 216 metastatic samples, some of them paired with samples from the corresponding primary breast cancers. They showed that metastatic clones are enriched with certain molecular abnormalities compared to the primary tumors [54].
Table 2: Genomics studies on breast cancer metastasis samples
| Number of samples | References |
|---|---|
| 8 | Weigelt, Proc Natl Acad Sci U S A. 2003 [78] |
| 14 | Wang, Genes Chromosomes Cancer 2009 [79] |
| 30 | Desouki, J Cancer Res Clin Oncol 2011 [80] |
| 14 | Craig, Mol Cancer Ther 2013 [81] |
| 15 | Lee, Oncotarget 2015 [82] |
| 62 | Onstenk, Cancer Lett 2015 [83] |
| 13 | McBryan, Clin Cancer Res 2015 [84] |
| 55 | Lang, Breast Cancer Res Treat 2015 [85] |
| 80 | Kimbung, Clin Cancer Res 2016 [86] |
| 88 | Fumagalli, Ann Oncol 2016 [87] |
| 216 | Lefebvre, PLoS Med 2016 [54] |
Our research team is conducting a program on brain metastases. As part of this program, we performed transcriptomic analyses on laser-microdissected metastatic lymph-nodes of 28 women with HER2-overexpressing or triple negative metastatic breast cancers. Supervised analyses compared the transcriptomic profiles of women who developed brain metastases with those who did not. We identified CDKN2A/p16 as a gene associated with the risk of brain metastases and decreased survival [55].
GENOMICS AND CONSTITUTIONAL GENETICS OF BREAST CANCER
In the last ten years, a panel of genes has been proposed for the diagnosis of hereditary familial cancers. In the context of hereditary predisposition for breast and/or ovarian cancers, the United States National Comprehensive Cancer Network recommends a panel of nineteen genes and proposes corresponding clinical screening tests (Table 3) [56]. In France, the Genetics and Cancer Group, supported by the French National Cancer Institute, recently updated their recommendation to test a panel of 13 genes accompanied by prevention and screening measures for patients and their families (Tables 4 and 5) [57, 58]. With twenty-eight platforms covering the French territory and dedicated to molecular biology, patients at risk for hereditary cancer can have the benefit of recent technologies applied to constitutional genetics. These platforms have implemented high-throughput sequencing tools like the Next Generation Sequencing (NGS) systems, to identify mutational hot-spots in a panel of high penetrance genes. Several types of NGS sequencers are currently used for routine care and also for research purposes, such as Illumina, Applied Biosystems SOLiD System, 454 Life Sciences (Roche), Helicos HeliScope, Complete Genomics, Pacific Biosciences PacBio and Life Technologies Ion Torrent [59].
Table 3: National Comprehensive Cancer Network guidelines for breast and ovarian cancer management based on genetic and familial high-risk assessment (Adapted from [56])
| Gene | Breast cancer risk management | Ovarian cancer risk management | Other cancer risk management |
|---|---|---|---|
| ATM | Increased risk Annual mammography and breast MRI starting at 40 years RRS: based on FH | Potential increase in risk with insufficient evidence to recommend RRS | Insufficient evidence for pancreas or prostate cancers |
| BRCA1 | Increased risk 25–29 years, annual breast MRI or mammogram 30–75 years, annual mammogram and breast MRI >75 years, based on IR RRS: based on IR and FH | Increased risk RRS: based on individual risk and FH between 35-40 years | Prostate, uterine (possible) |
| BRCA2 | Increased risk 25–29 years, annual breast MRI or mammogram 30–75 years, annual mammogram and breast MRI >75 years, based on IR RRS: based on IR and FH | Increased risk RRS: based on IR and FH between 40-45 years | Pancreas, prostate, melanoma |
| PALB2 | Increased risk Annual mammography and MRI starting at age 30 years RRS: based on FH | Insufficient evidence | Insufficient evidence |
| TP53 | Increased risk 20–29 years, annual breast MRI 30–75 years, annual breast MRI and mammogram >75 years, based on IR RRS: based on IR and FH | No increased risk | Neurological cancers, colon, skin cancers |
| CDH1 | Increased risk for lobular cancer Annual mammogram and breast MRI starting at age 30 years RRS: based on FH | No increased risk | Diffuse gastric cancer |
| PTEN | Increased risk Annual mammogram with breast MRI starting at age 30–35 years or 5-10 years before earliest breast cancer in family >75 years, based on IR RRS: based on IR and FH | No increased risk | Endometrial cancer, thyroid, colon, renal cancer, skin cancers |
| BRIP1 | Insufficient evidence | Increased risk Consider RRS at 45–50 years | Not available |
| CHEK2 | Increased risk Annual mammogram and breast MRI starting at age 40 years RRS: based on FH | No increased risk | Colon cancer |
| NBN | Increased risk Annual mammogram and breast MRI starting at age 40 years RRS: based on FH | Insufficient evidence | Insufficient evidence |
| NF1 | Increased risk Annual mammogram from age 40 years and consider breast MRI from 30–50 years RRS: based on FH | No increased risk | Malignant peripheral nerve sheath tumors, GIST, others |
| STK11 | Increased risk Annual mammogram and breast MRI starting at age 25 years RRS: based on FH | Increased risk of non-epithelial cancers Annual pelvic examination and PAP smear | Colon, stomach, pancreas, cervix, uterine, testis, lung |
| RAD51C | Insufficient evidence | Increased risk Consider RRS at 45–50 years | Not available |
| RAD51D | Insufficient evidence | Increased risk Consider RRS at 45–50 years | Not available |
| MLH1 | Insufficient evidence | Increased risk | Colon, uterine, others |
| MSH2 | manage based on FH | ||
| MSH6 | |||
| PMS2 | |||
| EPCAM |
Table 4: Recommendations of the Cancer and Genetics Group and the French National Institute of Cancer concerning gene panel analyses in the context of a hereditary predisposition to breast and ovarian cancers (Adapted from [57])
| Gene | Cytogenetic location | Penetrance | Protein functions | Cumulate risk of breast cancer | References |
|---|---|---|---|---|---|
| BRCA1 | 17q21.31 | High | Repair of DNA double-strand breaks using homologous recombination, cell cycle control, maintaining of genome integrity | 46–87% lifetime risk | [88, 89] |
| BRCA2 | 13q13.1 | High | Repair of DNA double-strand breaks using homologous recombination | 38–84% lifetime risk | [88, 90] |
| PALB2 | 16p12.2 | Moderate | Partner of BRCA2 and regulator of its stability and its nuclear localization | 35% at 70 years | [91, 92] |
| TP53 | 17p13.1 | High | Transcription Factor, cellular cycle, apoptosis, senescence, DNA Repair | 80% life-time risk (premenopausal) | [93, 94] |
| CDH1 | 16q22.1 | Moderate | E-cadherin, cellular adhesion molecule | 39–52% before 40 years (lobular cancer) | [95, 96] |
| PTEN | 10q23.31 | High | Tumor phosphatase suppressor inhibiting PI3K and MAPK pathways | 25–50% lifetime risk | [97, 98] |
| RAD51C | 17q22 | Moderate | Repair of DNA using homologous recombination in interaction with BRCA1/2 | Not known | [99–102] |
| RAD51D | 17q12 | Moderate | Repair of DNA using homologous recombination and maintaining of telomere | Not known | [103–105] |
| MLH1 | 3p22.2 | High | Mismatch repair system | 5–18% | [106–111] |
| MSH2 | 2p21-p16 | ||||
| MSH6 | 2p16.3 | Moderate | Not known | ||
| PMS2 | 7p22.1 | ||||
| EPCAM | 2p21 | Partner of MSH2 |
Table 5: Screening or prevention recommendations for persons carrying mutations of genes analyzed in the Cancer and Genetics Group panel (Adapted from [58])
| Gene | Breast surveillance | Risk reduction surgery | Gynecologic surveillance | |
|---|---|---|---|---|
| Breast | Pelvis | |||
| BRCA1BRCA2 | 30 - 65 year, annual MRI + mammogram (± Ultrasound) After 65 years, mammogram ± Ultrasound (Recommendations HAS 2014* and INCa 2017**) | Prophylactic mastectomy (Recommendations HAS 2014 * and INCa 2017**) | Prophylactic annexectomy (discussed from the age of 40 years and according to mutations and FH of OC) ** | Before RRS: standard surveillance and no efficacious ovarian screening available * |
| PALB2CDH1 | No specific gynecological guidelines If FH of OC: MDC | Standard surveillance | ||
| PTEN | Standard surveillance If gynecologic lesions of CD: MDC | |||
| TP53 | Starting at 20 years, annual MRI + Ultrasound (no systematic mammogram) | Standard surveillance | ||
| RAD51C RAD51D | No specific breast surveillance To be adapted to FH of BC according to guidelines HAS 2014* | Not indicated | Prophylactic annexectomy (discussed from the age of 45 years and according to FH of OC) | Before RRS: standard surveillance and no efficacious ovarian screening to be proposed ** |
| MLH1MSH2MSH6PMS2EPCAM | Ovarian and/or uterine RRS To be discussed in MDC according to Lynch syndrome guidelines | Uterine surveillance according to Lynch syndrome guidelines | ||
In sporadic breast cancers, NGS sequencing enables four subtypes to be discriminated on the basis of different genetic and epigenetic modifications, with three genes (PIK3CA, TP53, and GATA3) that are modified in more than 10% of patients. Basal-like breast cancers typically harbor mutations in the TP53, RB1, and BRCA1 genes, together with MYC amplifications [60].
Because of the potential therapeutic applications, the identification of inactivating mutations in the BRCA1 gene, a tumor suppressor gene, is important. BRCA1, because of its critical role in DNA repair mechanisms through homologous recombination, is one of the most important genes associated with hereditary breast cancer [61]. More than 75% of BRCA1-mutated breast cancers have a triple-negative phenotype and are classified as basal-like [8]. Constitutional BRCA1 mutations are of high penetrance, occur in 10% of breast cancer patients and in 20% of young women with triple-negative breast cancers [62]. BRCA1 sporadic mutations are also found in 1% of breast cancers, and the promoter can be hypermethylated in 11 to 14% of cases, resulting in BRCA1 gene inactivation [63–65].
A deficit in homologous recombination viaBRCA1 inactivation has provided the rationale to concomitantly inhibit other DNA repair pathways, particularly the (ADP-ribose) polymerase (PARP) enzyme pathway. Olaparib, a PARP inhibitor, showed considerable benefit in patients with metastatic BRCA1-mutated breast cancers [66, 67]. We, therefore, need to look for BRCA1 germline mutations in patients at risk for hereditary breast cancer. For sporadic breast cancer, particularly for the triple negative sub-type, we also need to identify BRCA1-inactivating mutations in tumors.
In this domain of constitutional genetics, many questions remain unanswered, particularly the translational value of identifying mutations of unknown significance in genes of low to moderate penetrance. The contributions of Genome-Wide Association Studies (GWAS) have not been very great. To date, more than sixty GWAS have been conducted on breast cancer samples. A meta-analysis of these GWAS identified 84 loci of interest possibly associated with an increased risk of breast cancer [68, 69]. Numerous low penetrance variants have been identified, without validating their functional significance. One of these variants concerns the oncogene FGFR2, the FGFR2 protein being overexpressed in 5% of breast cancers. This variant corresponds to a single nucleotide polymorphism (SNP) that affects the binding site of FGFR2, thus activating the downstream signaling pathway in a ligand-independent manner [70]. It is necessary to address the potential benefit of targeting FGFR2 for therapeutic purpose. Another SNP, in the 8q24 region, participates in regulating MYC oncogene transcription which is distant from this SNP by more than 300 kb [71]. Most GWAS studies suggest that mutations in low penetrance genes could partially explain genetic predisposition to breast cancer, even though their functional significance remains unclear [72].
Conclusions
The contribution of genomics applied to the treatment of breast cancer remains moderate. In practice, it is limited to informing adjuvant treatment decisions for early-stage diseases and to HER2-overexpressing breast cancers whatever the stage. However, breast cancers are heterogeneous and complex. Treatments need to be adjusted according to molecular subtypes and guided by the underlying genetic events. Several programs are ongoing to map the complex genetics of breast cancer, using multi-omic approaches such as the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) [73], which will help to take tumor heterogeneity into account more efficiently. Also, the increasingly widespread utilization of NGS will help to decipher the individual molecular complexity of breast cancers. This will rapidly increase the contribution of genomics in shaping breast cancer treatment in the next few years, especially in the present era of personalized treatments.
Abbreviations
ER: estradiol receptor; HER2: human epidermal growth factor receptor 2; Rb1: retinoblastoma protein 1; TP53: tumor protein 53 gene; BRCA1: breast cancer type 1 susceptibility gene; AR: androgen receptor; GCDFP15: gross cystic disease fluid protein 15; PAM50: Prediction of Microarray using 50 classifier genes plus 5 reference genes; ESR1: estrogen receptor 1 gene; ddPCR: digital droplet polymerase chain reaction; BL1: basal-like 1; BL2: basal-like 2; IM: immunomodulatory; M: mesenchymal-like; MSL: mesenchymal stem-like; LAR: luminal androgen receptor; EGFR: epidermal growth factor receptor; NGS: next generation sequencing; PARP: ADP-ribose polymerase; GWAS: genome-wide association studies; SNP: single nucleotide polymorphism; METABRIC: Molecular Taxonomy of Breast Cancer International Consortium; N: nodes; TNM: tumor, node, metastases; RT-PCR: Reverse Transcription-Polymerase Chain Reaction; MRI: Magnetic Resonance Imaging; RRS: Risk reduction surgery; FH: Family History; IR: Individual Risk; BC: Breast Cancers; OC: Ovarian Cancers; CD: Cowden disease; MDC: Multidisciplinary Committee; HAS: French Health Authority; INCa: French National Cancer Institute.
ACKNOWLEDGMENTS
We would like to thank Mrs. A. Swaine for revising the English language.
CONFLICTS OF INTEREST
No authors have any conflicts of interest to report.
FUNDING
None.
References
1. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015; 136:E359–86. https://doi.org/10.1002/ijc.29210. [PubMed].
2. Tang Y, Wang Y, Kiani MF, Wang B. Classification, Treatment Strategy, and Associated Drug Resistance in Breast Cancer. Clin Breast Cancer. 2016; 16:335–43. https://doi.org/10.1016/j.clbc.2016.05.012. [PubMed].
3. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015; 65:5–29. https://doi.org/10.3322/caac.21254. [PubMed].
4. Sundquist M, Brudin L, Tejler G. Improved survival in metastatic breast cancer 1985-2016. Breast. 2017; 31:46–50. https://doi.org/10.1016/j.breast.2016.10.005. [PubMed].
5. Osborne CK, Schiff R. Mechanisms of endocrine resistance in breast cancer. Annu Rev Med. 2011; 62:233–47. https://doi.org/10.1146/annurev-med-070909-182917. [PubMed].
6. Higgins MJ, Baselga J. Targeted therapies for breast cancer. J Clin Invest. 2011; 121:3797–803. https://doi.org/10.1172/JCI57152. [PubMed].
7. Foulkes WD, Smith IE, Reis-Filho JS. Triple-negative breast cancer. N Engl J Med. 2010; 363:1938–48. https://doi.org/10.1056/NEJMra1001389. [PubMed].
8. Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, et al. Molecular portraits of human breast tumours. Nature. 2000; 406:747–52. https://doi.org/10.1038/35021093. [PubMed].
9. Sørlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A. 2001; 98:10869–74. https://doi.org/10.1073/pnas.191367098. [PubMed].
10. Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, Demeter J, Perou CM, Lønning PE, et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci U S A. 2003; 100:8418–23. https://doi.org/10.1073/pnas.0932692100. [PubMed].
11. Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thürlimann B, Senn HJ, Albain KS, André F, Bergh J, Bonnefoi H, Bretel-Morales D, Burstein H, et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol. 2013; 24:2206–23. https://doi.org/10.1093/annonc/mdt303. [PubMed].
12. Perou CM. Molecular stratification of triple-negative breast cancers. Oncologist. 2011; 16:61–70. https://doi.org/10.1634/theoncologist.2011-S1-61. [PubMed].
13. Green AR, Powe DG, Rakha EA, Soria D, Lemetre C, Nolan CC, Barros FF, Macmillan RD, Garibaldi JM, Ball GR, Ellis IO. Identification of key clinical phenotypes of breast cancer using a reduced panel of protein biomarkers. Br J Cancer. 2013; 109:1886–94. https://doi.org/10.1038/bjc.2013.528. [PubMed].
14. Rakha EA, Soria D, Green AR, Lemetre C, Powe DG, Nolan CC, Garibaldi JM, Ball G, Ellis IO. Nottingham Prognostic Index Plus (NPI+): a modern clinical decision making tool in breast cancer. Br J Cancer. 2014; 110:1688–97. https://doi.org/10.1038/bjc.2014.120. [PubMed].
15. McNamara KM, Moore NL, Hickey TE, Sasano H, Tilley WD. Complexities of androgen receptor signalling in breast cancer. Endocr Relat Cancer. 2014; 21:T161–81. https://doi.org/10.1530/ERC-14-0243. [PubMed].
16. Farmer P, Bonnefoi H, Becette V, Tubiana-Hulin M, Fumoleau P, Larsimont D, Macgrogan G, Bergh J, Cameron D, Goldstein D, Duss S, Nicoulaz AL, Brisken C, et al. Identification of molecular apocrine breast tumours by microarray analysis. Oncogene. 2005; 24:4660–71. https://doi.org/10.1038/sj.onc.1208561. [PubMed].
17. Lehmann-Che J, Hamy AS, Porcher R, Barritault M, Bouhidel F, Habuellelah H, Leman-Detours S, de Roquancourt A, Cahen-Doidy L, Bourstyn E, de Cremoux P, de Bazelaire C, Albiter M, et al. Molecular apocrine breast cancers are aggressive estrogen receptor negative tumors overexpressing either HER2 or GCDFP15. Breast Cancer Res. 2013; 15:R37. https://doi.org/10.1186/bcr3421. [PubMed].
18. Guedj M, Marisa L, de Reynies A, Orsetti B, Schiappa R, Bibeau F, MacGrogan G, Lerebours F, Finetti P, Longy M, Bertheau P, Bertrand F, Bonnet F, et al. A refined molecular taxonomy of breast cancer. Oncogene. 2012; 31:1196–206. https://doi.org/10.1038/onc.2011.301. [PubMed].
19. Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest. 2011; 121:2750–67. https://doi.org/10.1172/JCI45014. [PubMed].
20. van ’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002; 415:530–36. https://doi.org/10.1038/415530a. [PubMed].
21. Ng CK, Schultheis AM, Bidard FC, Weigelt B, Reis-Filho JS. Breast cancer genomics from microarrays to massively parallel sequencing: paradigms and new insights. J Natl Cancer Inst. 2015; 107. https://doi.org/10.1093/jnci/djv015. [PubMed].
22. van de Vijver MJ, He YD, van’t Veer LJ, Dai H, Hart AA, Voskuil DW, Schreiber GJ, Peterse JL, Roberts C, Marton MJ, Parrish M, Atsma D, Witteveen A, et al. A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med. 2002; 347:1999–2009. https://doi.org/10.1056/NEJMoa021967. [PubMed].
23. Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004; 351:2817–26. https://doi.org/10.1056/NEJMoa041588. [PubMed].
24. Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009; 27:1160–67. https://doi.org/10.1200/JCO.2008.18.1370. [PubMed].
25. Dowsett M, Sestak I, Lopez-Knowles E, Sidhu K, Dunbier AK, Cowens JW, Ferree S, Storhoff J, Schaper C, Cuzick J. Comparison of PAM50 risk of recurrence score with oncotype DX and IHC4 for predicting risk of distant recurrence after endocrine therapy. J Clin Oncol. 2013; 31:2783–90. https://doi.org/10.1200/JCO.2012.46.1558. [PubMed].
26. Harris LN, Ismaila N, McShane LM, Andre F, Collyar DE, Gonzalez-Angulo AM, Hammond EH, Kuderer NM, Liu MC, Mennel RG, Van Poznak C, Bast RC, Hayes DF. Use of Biomarkers to Guide Decisions on Adjuvant Systemic Therapy for Women With Early-Stage Invasive Breast Cancer: American Society of Clinical Oncology Clinical Practice Guideline. J Clin Oncol. 2016; 34:1134–50. https://doi.org/10.1200/JCO.2015.65.2289. [PubMed].
27. Sotiriou C, Neo SY, McShane LM, Korn EL, Long PM, Jazaeri A, Martiat P, Fox SB, Harris AL, Liu ET. Breast cancer classification and prognosis based on gene expression profiles from a population-based study. Proc Natl Acad Sci U S A. 2003; 100:10393–98. https://doi.org/10.1073/pnas.1732912100. [PubMed].
28. Rakha EA, Green AR. Molecular classification of breast cancer: what the pathologist needs to know. Pathology. 2017; 49:111–19. https://doi.org/10.1016/j.pathol.2016.10.012. [PubMed].
29. Sparano JA, Gray RJ, Makower DF, Pritchard KI, Albain KS, Hayes DF, Geyer CE Jr, Dees EC, Goetz MP, Olson JA Jr, Lively T, Badve SS, Saphner TJ, et al. Adjuvant Chemotherapy Guided by a 21-Gene Expression Assay in Breast Cancer. N Engl J Med. 2018; 379:111–21. https://doi.org/10.1056/NEJMoa1804710. [PubMed].
30. Blok EJ, Bastiaannet E, van den Hout WB, Liefers GJ, Smit VT, Kroep JR, van de Velde CJ. Systematic review of the clinical and economic value of gene expression profiles for invasive early breast cancer available in Europe. Cancer Treat Rev. 2018; 62:74–90. https://doi.org/10.1016/j.ctrv.2017.10.012. [PubMed].
31. Li S, Shen D, Shao J, Crowder R, Liu W, Prat A, He X, Liu S, Hoog J, Lu C, Ding L, Griffith OL, Miller C, et al. Endocrine-therapy-resistant ESR1 variants revealed by genomic characterization of breast-cancer-derived xenografts. Cell Rep. 2013; 4:1116–30. https://doi.org/10.1016/j.celrep.2013.08.022. [PubMed].
32. Robinson DR, Wu YM, Vats P, Su F, Lonigro RJ, Cao X, Kalyana-Sundaram S, Wang R, Ning Y, Hodges L, Gursky A, Siddiqui J, Tomlins SA, et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat Genet. 2013; 45:1446–51. https://doi.org/10.1038/ng.2823. [PubMed].
33. Rakha EA, Reis-Filho JS, Ellis IO. Combinatorial biomarker expression in breast cancer. Breast Cancer Res Treat. 2010; 120:293–308. https://doi.org/10.1007/s10549-010-0746-x. [PubMed].
34. Lal S, McCart Reed AE, de Luca XM, Simpson PT. Molecular signatures in breast cancer. Methods. 2017; 131:135–46. https://doi.org/10.1016/j.ymeth.2017.06.032. [PubMed].
35. Wolff AC, Hammond ME, Hicks DG, Dowsett M, McShane LM, Allison KH, Allred DC, Bartlett JM, Bilous M, Fitzgibbons P, Hanna W, Jenkins RB, Mangu PB, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol. 2013; 31:3997–4013. https://doi.org/10.1200/JCO.2013.50.9984. [PubMed].
36. American Joint Committee on Cancer (AJCC) Cancer Staging Manual. New York: Springer; 2017.
37. Elbouchtaoui M, Tengher I, Miquel C, Brugière C, Benbara A, Zelek L, Ziol M, Bouhidel F, Janin A, Bousquet G, Leboeuf C. Micromolecular methods for diagnosis and therapeutic strategy: a case study. Oncotarget. 2018; 9:22862–69. https://doi.org/10.18632/oncotarget.25161. [PubMed].
38. Belgrader P, Tanner SC, Regan JF, Koehler R, Hindson BJ, Brown AS. Droplet digital PCR measurement of HER2 copy number alteration in formalin-fixed paraffin-embedded breast carcinoma tissue. Clin Chem. 2013; 59:991–94. https://doi.org/10.1373/clinchem.2012.197855. [PubMed].
39. Zhu Y, Lu D, Lira ME, Xu Q, Du Y, Xiong J, Mao M, Chung HC, Zheng G. Droplet digital polymerase chain reaction detection of HER2 amplification in formalin fixed paraffin embedded breast and gastric carcinoma samples. Exp Mol Pathol. 2016; 100:287–93. https://doi.org/10.1016/j.yexmp.2015.11.027. [PubMed].
40. Lehmann-Che J, Amira-Bouhidel F, Turpin E, Antoine M, Soliman H, Legres L, Bocquet C, Bernoud R, Flandre E, Varna M, de Roquancourt A, Plassa LF, Giacchetti S, et al. Immunohistochemical and molecular analyses of HER2 status in breast cancers are highly concordant and complementary approaches. Br J Cancer. 2011; 104:1739–46. https://doi.org/10.1038/bjc.2011.135. [PubMed].
41. Singer CF, Tan YY, Fitzal F, Steger GG, Egle D, Reiner A, Rudas M, Moinfar F, Gruber C, Petru E, Bartsch R, Tendl KA, Fuchs D, et al. Pathological Complete Response to Neoadjuvant Trastuzumab Is Dependent on HER2/CEP17 Ratio in HER2-Amplified Early Breast Cancer. Clin Cancer Res. 2017; 23:3676–83. https://doi.org/10.1158/1078-0432.CCR-16-2373. [PubMed].
42. Masuda H, Baggerly KA, Wang Y, Zhang Y, Gonzalez-Angulo AM, Meric-Bernstam F, Valero V, Lehmann BD, Pietenpol JA, Hortobagyi GN, Symmans WF, Ueno NT. Differential response to neoadjuvant chemotherapy among 7 triple-negative breast cancer molecular subtypes. Clin Cancer Res. 2013; 19:5533–40. https://doi.org/10.1158/1078-0432.CCR-13-0799. [PubMed].
43. De Roock W, Claes B, Bernasconi D, De Schutter J, Biesmans B, Fountzilas G, Kalogeras KT, Kotoula V, Papamichael D, Laurent-Puig P, Penault-Llorca F, Rougier P, Vincenzi B, et al. Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol. 2010; 11:753–62. https://doi.org/10.1016/S1470-2045(10)70130-3. [PubMed].
44. Park HS, Jang MH, Kim EJ, Kim HJ, Lee HJ, Kim YJ, Kim JH, Kang E, Kim SW, Kim IA, Park SY. High EGFR gene copy number predicts poor outcome in triple-negative breast cancer. Mod Pathol. 2014; 27:1212–22. https://doi.org/10.1038/modpathol.2013.251. [PubMed].
45. Gucalp A, Tolaney S, Isakoff SJ, Ingle JN, Liu MC, Carey LA, Blackwell K, Rugo H, Nabell L, Forero A, Stearns V, Doane AS, Danso M, et al. Phase II trial of bicalutamide in patients with androgen receptor-positive, estrogen receptor-negative metastatic Breast Cancer. Clin Cancer Res. 2013; 19:5505–12. https://doi.org/10.1158/1078-0432.CCR-12-3327. [PubMed].
46. Bonnefoi H, Grellety T, Tredan O, Saghatchian M, Dalenc F, Mailliez A, L’Haridon T, Cottu P, Abadie-Lacourtoisie S, You B, Mousseau M, Dauba J, Del Piano F, et al. A phase II trial of abiraterone acetate plus prednisone in patients with triple-negative androgen receptor positive locally advanced or metastatic breast cancer (UCBG 12-1). Ann Oncol. 2016; 27:812–18. https://doi.org/10.1093/annonc/mdw067. [PubMed].
47. Bousquet G, Feugeas JP, Ferreira I, Vercellino L, Jourdan N, Bertheau P, de Bazelaire C, Barranger E, Janin A. Individual xenograft as a personalized therapeutic resort for women with metastatic triple-negative breast carcinoma. Breast Cancer Res. 2014; 16:401. https://doi.org/10.1186/bcr3615. [PubMed].
48. Nguyen TT, El Bouchtaoui M, Feugeas JP, Vercillino L, de Bazelaire C, Janin A, Bousquet G. Individual xenografts for personalized treatment of women with metastatic triple negative breast cancer. Journal of Translational Genetics and Genomics. 2018; 2:19. https://doi.org/10.20517/jtgg.2018.20.
49. Bousquet G, El Bouchtaoui M, Leboeuf C, Battistella M, Varna M, Ferreira I, Plassa LF, Hamdan D, Bertheau P, Feugeas JP, Damotte D, Janin A. Tracking sub-clonal TP53 mutated tumor cells in human metastatic renal cell carcinoma. Oncotarget. 2015; 6:19279–89. https://doi.org/10.18632/oncotarget.4220. [PubMed].
50. Haffner MC, Mosbruger T, Esopi DM, Fedor H, Heaphy CM, Walker DA, Adejola N, Gürel M, Hicks J, Meeker AK, Halushka MK, Simons JW, Isaacs WB, et al. Tracking the clonal origin of lethal prostate cancer. J Clin Invest. 2013; 123:4918–22. https://doi.org/10.1172/JCI70354. [PubMed].
51. Otsuji K, Sasaki T, Tanaka A, Kunita A, Ikemura M, Matsusaka K, Tada K, Fukayama M, Seto Y. Use of droplet digital PCR for quantitative and automatic analysis of the HER2 status in breast cancer patients. Breast Cancer Res Treat. 2017; 162:11–18. https://doi.org/10.1007/s10549-016-4092-5. [PubMed].
52. Wang X, Wu Y, Song X, Sun C, Wu C, Feng H. Human epidermal growth factor receptor 2 amplification detection by droplet digital polymerase chain reaction in formalin-fixed paraffin-embedded breast and gastric cancer samples. J Cancer Res Ther. 2017; 13:730–34. https://doi.org/10.4103/jcrt.JCRT_587_17. [PubMed].
53. de Bazelaire C, Coffin A, Cohen S, Scemama A, de Kerviler E. Biopsies in oncology. Diagn Interv Imaging. 2014; 95:647–57. https://doi.org/10.1016/j.diii.2014.04.016. [PubMed].
54. Lefebvre C, Bachelot T, Filleron T, Pedrero M, Campone M, Soria JC, Massard C, Lévy C, Arnedos M, Lacroix-Triki M, Garrabey J, Boursin Y, Deloger M, et al. Mutational Profile of Metastatic Breast Cancers: A Retrospective Analysis. PLoS Med. 2016; 13:e1002201. https://doi.org/10.1371/journal.pmed.1002201. [PubMed].
55. Furet E, El Bouchtaoui M, Feugeas JP, Miquel C, Leboeuf C, Beytout C, Bertheau P, Le Rhun E, Bonneterre J, Janin A, Bousquet G. Increased risk of brain metastases in women with breast cancer and p16 expression in metastatic lymph-nodes. Oncotarget. 2017; 8:37332–41. https://doi.org/10.18632/oncotarget.16953. [PubMed].
56. NCCN NCCN. Genetic/familial high-risk assessmentfor: breast and ovarian (Version 2.2019).
57. UNICANCER. Quels gènes analyser face à un risque de cancer héréditaire du sein et de l’ovaire?
58. UNICANCER. Recommandations de prévention ou dépistage des personnes porteuses d’une mutation d’un des gènes analysés dans le panel GGC-UNICANCER.
59. Bahassi M, Stambrook PJ. Next-generation sequencing technologies: breaking the sound barrier of human genetics. Mutagenesis. 2014; 29:303–10. https://doi.org/10.1093/mutage/geu031. [PubMed].
60. Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012; 490:61–70. https://doi.org/10.1038/nature11412. [PubMed].
61. Gonzalez-Angulo AM, Timms KM, Liu S, Chen H, Litton JK, Potter J, Lanchbury JS, Stemke-Hale K, Hennessy BT, Arun BK, Hortobagyi GN, Do KA, Mills GB, Meric-Bernstam F. Incidence and outcome of BRCA mutations in unselected patients with triple receptor-negative breast cancer. Clin Cancer Res. 2011; 17:1082–89. https://doi.org/10.1158/1078-0432.CCR-10-2560. [PubMed].
62. Peto J, Collins N, Barfoot R, Seal S, Warren W, Rahman N, Easton DF, Evans C, Deacon J, Stratton MR. Prevalence of BRCA1 and BRCA2 gene mutations in patients with early-onset breast cancer. J Natl Cancer Inst. 1999; 91:943–49. https://doi.org/10.1093/jnci/91.11.943. [PubMed].
63. Rigakos G, Razis E. BRCAness: finding the Achilles heel in ovarian cancer. Oncologist. 2012; 17:956–62. https://doi.org/10.1634/theoncologist.2012-0028. [PubMed].
64. Leidy J, Khan A, Kandil D. Basal-like breast cancer: update on clinicopathologic, immunohistochemical, and molecular features. Arch Pathol Lab Med. 2014; 138:37–43. https://doi.org/10.5858/arpa.2012-0439-RA. [PubMed].
65. Turner N, Tutt A, Ashworth A. Hallmarks of ‘BRCAness’ in sporadic cancers. Nat Rev Cancer. 2004; 4:814–19. https://doi.org/10.1038/nrc1457. [PubMed].
66. Robson M, Im SA, Senkus E, Xu B, Domchek SM, Masuda N, Delaloge S, Li W, Tung N, Armstrong A, Wu W, Goessl C, Runswick S, Conte P. Olaparib for Metastatic Breast Cancer in Patients with a Germline BRCA Mutation. N Engl J Med. 2017; 377:523–33. https://doi.org/10.1056/NEJMoa1706450. [PubMed].
67. Zimmer AS, Gillard M, Lipkowitz S, Lee JM. Update on PARP Inhibitors in Breast Cancer. Curr Treat Options Oncol. 2018; 19:21. https://doi.org/10.1007/s11864-018-0540-2. [PubMed].
68. Michailidou K, Hall P, Gonzalez-Neira A, Ghoussaini M, Dennis J, Milne RL, Schmidt MK, Chang-Claude J, Bojesen SE, Bolla MK, Wang Q, Dicks E, Lee A, et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat Genet. 2013; 45:353–61, 361e1–2. https://doi.org/10.1038/ng.2563. [PubMed].
69. Michailidou K, Beesley J, Lindstrom S, Canisius S, Dennis J, Lush MJ, Maranian MJ, Bolla MK, Wang Q, Shah M, Perkins BJ, Czene K, Eriksson M, et al. Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat Genet. 2015; 47:373–80. https://doi.org/10.1038/ng.3242. [PubMed].
70. Moffa AB, Ethier SP. Differential signal transduction of alternatively spliced FGFR2 variants expressed in human mammary epithelial cells. J Cell Physiol. 2007; 210:720–31. https://doi.org/10.1002/jcp.20880. [PubMed].
71. Ahmadiyeh N, Pomerantz MM, Grisanzio C, Herman P, Jia L, Almendro V, He HH, Brown M, Liu XS, Davis M, Caswell JL, Beckwith CA, Hills A, et al. 8q24 prostate, breast, and colon cancer risk loci show tissue-specific long-range interaction with MYC. Proc Natl Acad Sci U S A. 2010; 107:9742–46. https://doi.org/10.1073/pnas.0910668107. [PubMed].
72. Antoniou AC, Pharoah PD, McMullan G, Day NE, Ponder BA, Easton D. Evidence for further breast cancer susceptibility genes in addition to BRCA1 and BRCA2 in a population-based study. Genet Epidemiol. 2001; 21:1–18. https://doi.org/10.1002/gepi.1014. [PubMed].
73. Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Gräf S, Ha G, Haffari G, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012; 486:346–52. https://doi.org/10.1038/nature10983. [PubMed].
74. Filipits M, Rudas M, Jakesz R, Dubsky P, Fitzal F, Singer CF, Dietze O, Greil R, Jelen A, Sevelda P, Freibauer C, Müller V, Jänicke F, et al. A new molecular predictor of distant recurrence in ER-positive, HER2-negative breast cancer adds independent information to conventional clinical risk factors. Clin Cancer Res. 2011; 17:6012–20. https://doi.org/10.1158/1078-0432.CCR-11-0926. [PubMed].
75. Ma XJ, Salunga R, Dahiya S, Wang W, Carney E, Durbecq V, Harris A, Goss P, Sotiriou C, Erlander M, Sgroi D. A five-gene molecular grade index and HOXB13:IL17BR are complementary prognostic factors in early stage breast cancer. Clin Cancer Res. 2008; 14:2601–08. https://doi.org/10.1158/1078-0432.CCR-07-5026. [PubMed].
76. Wang Y, Klijn JG, Zhang Y, Sieuwerts AM, Look MP, Yang F, Talantov D, Timmermans M, Meijer-van Gelder ME, Yu J, Jatkoe T, Berns EM, Atkins D, Foekens JA. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet. 2005; 365:671–79. https://doi.org/10.1016/S0140-6736(05)17947-1. [PubMed].
77. Krijgsman O, Roepman P, Zwart W, Carroll JS, Tian S, de Snoo FA, Bender RA, Bernards R, Glas AM. A diagnostic gene profile for molecular subtyping of breast cancer associated with treatment response. Breast Cancer Res Treat. 2012; 133:37–47. https://doi.org/10.1007/s10549-011-1683-z. [PubMed].
78. Weigelt B, Glas AM, Wessels LF, Witteveen AT, Peterse JL, van’t Veer LJ. Gene expression profiles of primary breast tumors maintained in distant metastases. Proc Natl Acad Sci U S A. 2003; 100:15901–05. https://doi.org/10.1073/pnas.2634067100. [PubMed].
79. Wang C, Iakovlev VV, Wong V, Leung S, Warren K, Iakovleva G, Arneson NC, Pintilie M, Miller N, Youngson B, McCready DR, Done SJ. Genomic alterations in primary breast cancers compared with their sentinel and more distal lymph node metastases: an aCGH study. Genes Chromosomes Cancer. 2009; 48:1091–101. https://doi.org/10.1002/gcc.20711. [PubMed].
80. Desouki MM, Liao S, Huang H, Conroy J, Nowak NJ, Shepherd L, Gaile DP, Geradts J. Identification of metastasis-associated breast cancer genes using a high-resolution whole genome profiling approach. J Cancer Res Clin Oncol. 2011; 137:795–809. https://doi.org/10.1007/s00432-010-0937-1. [PubMed].
81. Craig DW, O’Shaughnessy JA, Kiefer JA, Aldrich J, Sinari S, Moses TM, Wong S, Dinh J, Christoforides A, Blum JL, Aitelli CL, Osborne CR, Izatt T, et al. Genome and transcriptome sequencing in prospective metastatic triple-negative breast cancer uncovers therapeutic vulnerabilities. Mol Cancer Ther. 2013; 12:104–16. https://doi.org/10.1158/1535-7163.MCT-12-0781. [PubMed].
82. Lee JY, Park K, Lim SH, Kim HS, Yoo KH, Jung KS, Song HN, Hong M, Do IG, Ahn T, Lee SK, Bae SY, Kim SW, et al. Mutational profiling of brain metastasis from breast cancer: matched pair analysis of targeted sequencing between brain metastasis and primary breast cancer. Oncotarget. 2015; 6:43731–42. https://doi.org/10.18632/oncotarget.6192. [PubMed].
83. Onstenk W, Sieuwerts AM, Weekhout M, Mostert B, Reijm EA, van Deurzen CH, Bolt-de Vries JB, Peeters DJ, Hamberg P, Seynaeve C, Jager A, de Jongh FE, Smid M, et al. Gene expression profiles of circulating tumor cells versus primary tumors in metastatic breast cancer. Cancer Lett. 2015; 362:36–44. https://doi.org/10.1016/j.canlet.2015.03.020. [PubMed].
84. McBryan J, Fagan A, McCartan D, Bane FT, Varešlija D, Cocchiglia S, Byrne C, Bolger J, McIlroy M, Hudson L, Tibbitts P, Gaora PO, Hill AD, Young LS. Transcriptomic Profiling of Sequential Tumors from Breast Cancer Patients Provides a Global View of Metastatic Expression Changes Following Endocrine Therapy. Clin Cancer Res. 2015; 21:5371–79. https://doi.org/10.1158/1078-0432.CCR-14-2155. [PubMed].
85. Lang JE, Scott JH, Wolf DM, Novak P, Punj V, Magbanua MJ, Zhu W, Mineyev N, Haqq CM, Crothers JR, Esserman LJ, Tripathy D, van ’t Veer L, Park JW. Expression profiling of circulating tumor cells in metastatic breast cancer. Breast Cancer Res Treat. 2015; 149:121–31. https://doi.org/10.1007/s10549-014-3215-0. [PubMed].
86. Kimbung S, Johansson I, Danielsson A, Veerla S, Egyhazi Brage S, Frostvik Stolt M, Skoog L, Carlsson L, Einbeigi Z, Lidbrink E, Linderholm B, Loman N, Malmström PO, et al. Transcriptional Profiling of Breast Cancer Metastases Identifies Liver Metastasis-Selective Genes Associated with Adverse Outcome in Luminal A Primary Breast Cancer. Clin Cancer Res. 2016; 22:146–57. https://doi.org/10.1158/1078-0432.CCR-15-0487. [PubMed].
87. Fumagalli D, Wilson TR, Salgado R, Lu X, Yu J, O’Brien C, Walter K, Huw LY, Criscitiello C, Laios I, Jose V, Brown DN, Rothé F, et al. Somatic mutation, copy number and transcriptomic profiles of primary and matched metastatic estrogen receptor-positive breast cancers. Ann Oncol. 2016; 27:1860–66. https://doi.org/10.1093/annonc/mdw286. [PubMed].
88. Petrucelli N, Daly MB, Pal T. BRCA1- and BRCA2-Associated Hereditary Breast and Ovarian Cancer. In: Adam MP (ed.) (ed.), Ardinger HH (ed.) (ed.), Pagon RA (ed.) (ed.), Wallace SE (ed.) (ed.), Bean LJ (ed.) (ed.), Stephens K (ed.) (ed.), Amemiya A (ed.) (ed.), editors. GeneReviews((R)). (Seattle (WA)); 1993.
89. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *113705 : 08/17/2018.
90. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: * 600185: 11/28/2017.
91. Viassolo V, Ayme A, Chappuis PO. Cancer du sein : risque génétique. Imag Femme. 2016; 26:95–104. https://doi.org/10.1016/j.femme.2016.04.009.
92. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *610355: 02/04/2016.
93. Schneider K, Zelley K, Nichols KE, Garber J. Li-Fraumeni Syndrome. In: Adam MP (ed.) (ed.), Ardinger HH (ed.) (ed.), Pagon RA (ed.) (ed.), Wallace SE (ed.) (ed.), Bean LJ (ed.) (ed.), Stephens K (ed.) (ed.), Amemiya A (ed.) (ed.), editors. GeneReviews((R)). (.Seattle (WA)); 1993.
94. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *191170 : 11/07/2018.
95. Kaurah P, Huntsman DG. Hereditary Diffuse Gastric Cancer. In: Adam MP (ed.) (ed.), Ardinger HH (ed.) (ed.), Pagon RA (ed.) (ed.), Wallace SE (ed.) (ed.), Bean LJ (ed.) (ed.), Stephens K (ed.) (ed.), Amemiya A (ed.) (ed.), editors. GeneReviews((R)). (Seattle (WA)); 1993.
96. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *192090: 10/18/2018.
97. Eng C. PTEN Hamartoma Tumor Syndrome. In: Adam MP (ed.) (ed.), Ardinger HH (ed.) (ed.), Pagon RA (ed.) (ed.), Wallace SE (ed.) (ed.), Bean LJ (ed.) (ed.), Stephens K (ed.) (ed.), Amemiya A (ed.) (ed.), editors. GeneReviews((R)). (Seattle (WA)); 1993.
98. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: 601728: 06/28/2018.
99. Mehta PA, Tolar J. Fanconi Anemia. In: Adam MP (ed.) (ed.), Ardinger HH (ed.) (ed.), Pagon RA (ed.) (ed.), Wallace SE (ed.) (ed.), Bean LJ (ed.) (ed.), Stephens K (ed.) (ed.), Amemiya A (ed.) (ed.), editors. GeneReviews((R)). (Seattle (WA)); 1993.
100. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *602774: 12/04/2013.
101. Sopik V, Akbari MR, Narod SA. Genetic testing for RAD51C mutations: in the clinic and community. Clin Genet. 2015; 88:303–12. https://doi.org/10.1111/cge.12548. [PubMed].
102. Thompson ER, Boyle SE, Johnson J, Ryland GL, Sawyer S, Choong DY, ConFab K, Chenevix-Trench G, Trainer AH, Lindeman GJ, Mitchell G, James PA, Campbell IG. Analysis of RAD51C germline mutations in high-risk breast and ovarian cancer families and ovarian cancer patients. Hum Mutat. 2012; 33:95–99. https://doi.org/10.1002/humu.21625. [PubMed].
103. Pelttari LM, Kiiski JI, Ranta S, Vilske S, Blomqvist C, Aittomäki K, Nevanlinna H. RAD51, XRCC3, and XRCC2 mutation screening in Finnish breast cancer families. Springerplus. 2015; 4:92. https://doi.org/10.1186/s40064-015-0880-3. [PubMed].
104. Kleibl Z, Kristensen VN. Women at high risk of breast cancer: molecular characteristics, clinical presentation and management. Breast. 2016; 28:136–44. https://doi.org/10.1016/j.breast.2016.05.006. [PubMed].
105. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *602954: 12/04/2013.
106. Kohlmann W, Gruber SB. Lynch Syndrome. In: Adam MP (ed.) (ed.), Ardinger HH (ed.) (ed.), Pagon RA (ed.) (ed.), Wallace SE (ed.) (ed.), Bean LJ (ed.) (ed.), Stephens K (ed.) (ed.), Amemiya A (ed.) (ed.), editors. (GeneReviews((R)). (Seattle (WA)); 1993.
107. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *120436 : 04/20/2018.
108. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *609309 : 10/05/2016.
109. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *600678: 01/24/2018.
110. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *600259: 12/27/2016.
111. Online Mendelian Inheritance in Man OJHU, Baltimore, MD. MIM Number: *185535: 12/18/2017.

