INTRODUCTION
The survival of patients with metastatic melanoma has improved with treatment with BRAF and MEK inhibitors [1] or with immunotherapy such as anti-PD-1 or anti-CTLA-4 antibodies [2, 3]. The mutated BRAF oncogene represents a therapeutic target in metastatic melanoma, where a mutated NRAS oncogene is a biomarker of poor outcome [4] and resistance to treatment with BRAF inhibitors [5]. A prerequisite for safe clinical use of BRAF inhibitors is based on reliable molecular detection of activating BRAF mutations in routine clinical practice. Several methods to detect BRAF and NRAS mutations in formalin-fixed paraffin-embedded (FFPE) samples are currently available in molecular pathology laboratories worldwide. PCR-based techniques require a dedicated infrastructure, which is not always present in pathology laboratories. Moreover, guarantee of reliable and accurate molecular results can be obtained by continuous control of the pre-analytical steps and by a trained technical personnel working ideally in an ISO-accredited laboratory [6, 7]. As only patients whose tumors harbor the “druggable” mutation will benefit from a targeted treatment, there is strong need for reliable, fast, and easy-to-use detection of mutations. Furthermore, a personalized treatment scheme requires monitoring of the tumor’s genomic status. Liquid biopsy in metastatic melanoma has emerged as an alternative tool that is complementary to tumor biopsies for detection of “druggable” molecular alterations [8–10]. Many studies have demonstrated that circulating cell-free DNA (cfDNA) represents genetic information from the whole tumor genome and can provide evidence of the clonal evolution and tumor heterogeneity in several types of cancer, including melanoma [11–15]. The detection of the tumoral fraction of cfDNA (ctDNA) is challenging, notably because the relative yield of ctDNA varies significantly, and in some cases less than 1% of the total amount of cfDNA is obtained. The sensitivity and specificity also depends on respecting closely the pre-analytical steps, from sample collection to rapid handling in less than 6 hours [16, 17].
In this study, we evaluated the fully automated ready-to-use Idylla™ PCR-based system for identification of BRAF V600 and NRAS mutations in plasma samples from patients with metastatic melanoma at baseline and during the course of treatment. A comparison with pyrosequencing using matched tissue samples and with next-generation sequencing (NGS) with matched plasma specimens was also performed.
RESULTS
Study population
In this monocentric prospective study, samples from 19 patients with stage IV metastatic melanoma collected before and after treatment were assessed for BRAF and NRAS mutations. The main clinicopathological features of the included patients are highlighted in Table 1. To measure the metastatic tumor burden (TB), the number of metastatic sites was counted before and after therapy [range 1 to 5]. Sites of involvement, in order of frequency, were visceral (15/19; 79%), lymph node (7/19; 37%), subcutaneous (6/19; 32%), brain (5/19; 26%) and bone (2/19; 10%). At the first blood draw, all patients were naïve to treatment. Nine out of 19 (47%) patients received tyrosine kinase inhibitor therapy such as BRAF inhibitors (Zelboraf/Dabrafenib) or combined BRAF and MEK inhibitors (Mekinist); 9/19 (47%) patients received immunotherapy (Ipilimumab or Nivolumab) and one patient was treated with dacarbazine (5%). One patient with bone metastasis also received radiotherapy. 6 out of 19 (32%) patients were clinically disease-free after treatment.
Table 1: Main clinicopathological parameters
Demographical and clinical characteristics |
Overall |
|
|---|---|---|
19 (100%) |
||
Age at diagnostic |
||
(years) |
Median |
61.63 |
Range |
43–78 |
|
Gender |
Male |
16 (84%) |
Female |
3 (16%) |
|
Histology |
Superficial Spreading |
8 (42%) |
Nodular |
5 (26%) |
|
Acral lentiginous |
2 (11%) |
|
Other |
4 (21%) |
|
Breslow (mm) |
Median Range |
4.91 (0.35–10) |
Assessed |
17 (89%) |
|
Numbers of metastatic sites before therapy |
1 |
8 (42%) |
2 |
5 (26%) |
|
3 |
4 (21%) |
|
4 |
2 (10%) |
|
Visceral disease |
15 (79%) |
|
Non visceral disease |
4 (21 (%) |
|
Brain metastasis |
Yes |
3 (16%) |
No |
16 (82%) |
|
Treatment |
Targeted Therapy |
9 (47%) |
BRAF inhibitors |
2 (10%) |
|
Combine MEK and BRAF inhibitors |
7 (36%) |
|
Immunotherapy |
9 (47%) |
|
anti-CTLA4 |
8 (42%) |
|
anti-PD1 |
1 (5%) |
|
Chemotherapy |
1 (5%) |
|
LDH |
Assessed |
35 (100%) |
Increased |
6 (17%) |
|
Range |
504–1826 |
|
Treatment line |
1 |
19 |
Response at M2 |
Complete response |
3 (16%) |
Partial response |
5 (26%) |
|
Stabilization |
5 (26%) |
|
Progression |
6 (31%) |
|
Death before M2 |
2 (10%) |
|
Overall survival (month) |
Median |
56.4 |
Range |
[28.0–NA] |
|
Mutational status in tumor tissue |
BRAF |
10 (52%) |
p.V600E |
9 |
|
p.V600K |
1 |
|
NRAS |
5 (26%) |
|
p.Q61K |
2 |
|
p.Q61L |
1 |
|
p.Q61R |
1 |
|
pG12S |
1 |
|
Double Wild-Type |
4 (21%) |
|
Mutation status of cfDNA evaluated with the Idylla system
At baseline, 9 out of 19 (47%) patients harbored a plasmatic BRAFV600 mutation with a mean mutant allele frequency of 12.7% [range 0.36%–66.67%]. Two months after treatment the BRAFV600 mutant cfDNA was undetectable in all patients. For NRAS 2 out of 13 (15%) harbored a plasmatic mutation before and after treatment.
Mutation status in FFPE sample with pyrosequencing and comparison with Idylla in plasma
The pyrosequencing analysis of tumor tissue showed that 10 out of 19 (52%) patients had a BRAF mutation, 5 out 19 (26%) harbored a NRAS mutation, and 4 out of 19 (21%) patients were WT for both BRAF and NRAS genes (Table 1).
The agreement at baseline for both approaches was high for the BRAF mutation status, with a sensitivity of 80% and a specificity of 89% (κ = 0.69). Three cases out of 19 (16%) had discordant results. Two BRAFV600E mutations detected in FFPE samples were not detected in plasma, and one BRAFV600E mutation was detected in plasma (0.36% mutant allele frequency) but was not detected on the matched tissue specimen by pyrosequencing, Next Generation Sequencing (NGS) or immunohistochemistry. With regard to the NRAS mutation status, 3 FFPE mutations (Q61L; Q61K, G12S) were not detected in plasma. The genotype of matched FFPE samples determined by pyrosequencing showed a concordance rate of 84% (11/13) with a sensitivity of 79% and a specificity of 100%.
Next generation sequencing with plasma samples and comparison with tissue and Idylla™
The mutation status was assessed by NGS for all 35 plasma samples included in the study. An adequate amount of DNA was extracted from the plasma samples before and after treatment, however analysis failed for 3 samples. Nine mutations were detected prior to treatment (6 BRAFV600, 2 BRAF nonV600 and 1 NRASQ61K) and 4 subsequent to treatment (1 BRAF nonV600 and 3 NRAS) (Figure 1). There was a wide range of the mutant allele fraction (median = 15.1%, range 2.2 to 49%). When compared to the mutation status obtained with FFPE tumor samples, 14 out of 17 (82%) cases showed agreement for BRAFV600 (sensitivity 67%, and specificity 75%) and 14 out of 17(82%) for NRAS (sensitivity 40%, and specificity 100%). Twenty-seven out of 32 (84%) results were in agreement with the findings with Idylla (before and after treatment). When only the plasma NRAS and BRAFV600 mutation spectrum is considered, 5 cases are discordant. 3 BRAFV600 (#8, #14, #19) mutations detected with Idylla were absent with NGS, and 1 BRAFV600 (#21) and 1 NRAS (#7) mutation detected in NGS are not detected with Idylla. For these cases the mutant allelic fraction was low (mean 1.65 range [0.36–3.1]) near by the limit of detection of both methods. Finally, we did not find any KIT gene activating mutations or resistant mutations on PI3K, PTEN, AKT, CDKN2a or JAK2 genes [18, 19], although clonal evolution in melanoma has been described when on therapy [20, 21].
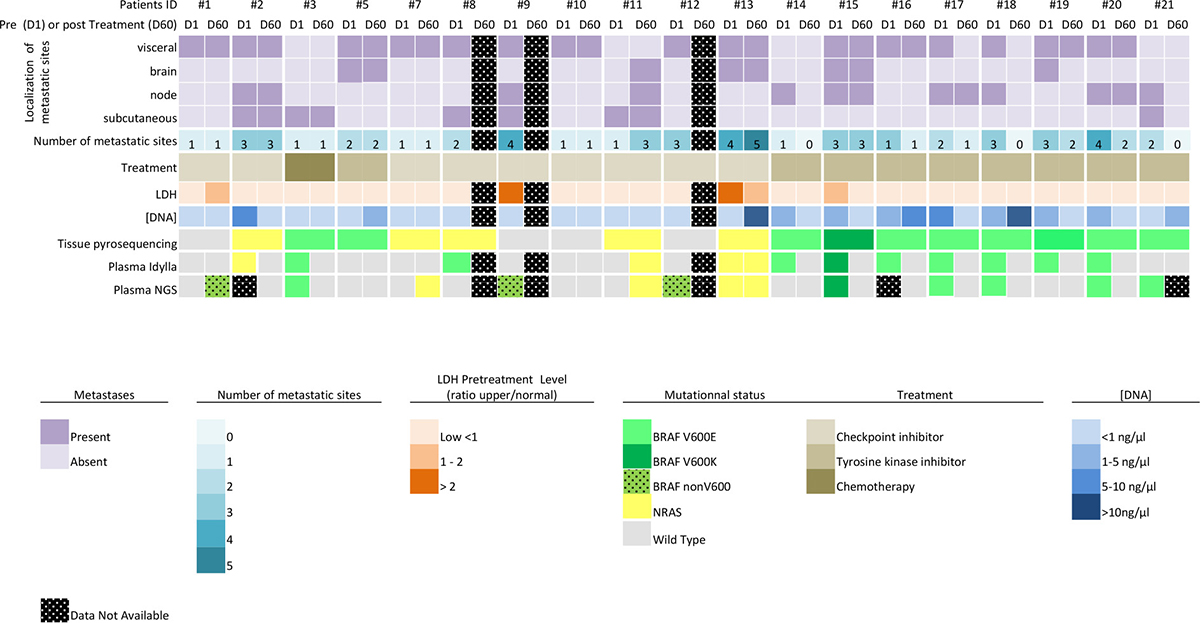
Figure 1: Overview of the results and clinical data.
Mutation status in cfDNA and clinico-pathological characteristics
Among the 3 conflicting cases Idylla BRAF prior to treatment, 2 BRAFV600E mutations detected in FFPE samples were not detected in plasma. For these patients the time between tissue analysis and plasma is 20 days and 3 months respectively. Both patients (#5 and #21) had a low cfDNA concentration (0.3 ng/μl and 0.8 ng/μl respectively), a normal LDH level, and a low metastatic TB (one brain metastasis and a residual pancreatic metastasis after surgical resection, and one lymph node metastasis and subcutaneous nodule). Both patients were still alive 3 years after the first blood draw when under combined targeted therapy. For the additional BRAFV600E detected in plasma (#8) the time between the tissue and the blood sampling is 6 months. When analyzing at the time elapsed between tissue and plasma analysis, we can see that the discrepancy observed in genotype is not linked with the time period in our cohort of naïve treatment patient. After targeted treatment (BRAF inhibitors with or without MEK inhibitors) the BRAFV600 mutant cfDNA was undetectable in all patients and 3 had no evidence of disease by imagery (CT-scan and TEP-Scan).
For Idylla NRAS, 3 patients (patients #7, #8, #11) harboring a NRAS mutation in tumor tissue did not show a NRAS mutation in the matched plasma samples. These patients also had a low metastatic TB and normal LDL level. Among them, 2 out of 13 (15%) harbored a plasmatic NRAS (Q61R and Q61K) mutation detected before treatment. After immunotherapy (anti-PD1 and anti CTLA4), one patient with metastatic progression had a persistent plasmatic NRAS Q61K mutation (patient #13), one NRAS Q61R mutation was undetectable in a patient with a partial response (patient #2) and a plasmatic NRAS Q61L mutation was identified in a patient with disease progression (lymph node and cerebral metastasis - patient #11) (Figure 1).
Using ultra-deep sequencing of cfDNA we discovered 3 new plasmatic BRAF mutations (two L597S #1; #9 and one K601E #12; mean allele fraction 26%– range [18–30%]; mean reads 887 – range [301–1998]) that were not investigated with pyrosequencing or Idylla. Two of these mutations were ever present on FFPE matched samples (SSM #12 and nodular #9 melanoma), they were detected retrospectively with the Sanger sequencing method, and one BRAF L597S mutation appeared de novo in an acral lentiginous melanoma after stabilization on immunotherapy (#1). In the latter case, it seems likely that we have identified an emerging clone under therapy because the time between plasma and tissue analysis did not exceed 2 months, the plasmatic allele frequency was 30% and the percentage of tumor cells in the tissue was 70% (well above the sensitivity threshold). Also case #8 remains inconsistent as neither the BRAF nor the NRAS mutation was detected with NGS.
Correlation of the cfDNA mutation status evaluated with Idylla with clinical outcome
BRAF and NRAS cfDNA mutations were mutually exclusive. The presence of a plasma mutation at baseline did not correlate statistically to OS (P = 0.25) and was not related to the level of LDH. Furthermore, when considering only the cohort of BRAF V600 mutated tumors with FFPE samples, the presence of a BRAF cfDNA mutation did not correlate either to OS (P = 0.23), or to the objective response rate (P = 0.22) or the number and location of metastases (P = 0.46).
As we could not assess the yield of cfDNA copies/mL we sought to see if the cfDNA concentration ([cfDNA]) correlated with tumor volume (number and location of metastatic sites), and OS. Measurement of [cfDNA] at baseline was performed for all patients. The mean concentration was 2.77 ng/μL (range 0.27–29 ng/μL). As shown in Figure 2, a correlation between [cfDNA] and the presence of a plasmatic mutation (P = 0.0031) was found. The correlation between [cfDNA] and the number or the localization of metastatic sites was not statistically significant. Nevertheless, there was a tendency to correlation between the number of metastatic sites and [cfDNA] (Figure 3). There was a tendency to a higher [cfDNA] for a visceral localization compared to a subcutaneous or brain localization, as previously described in other studies [11, 22, 23]. Moreover, there was no significant correlation between the baseline [cfDNA] or variation before and after treatment, Δ[cfDNA] and OS. The allele mutant fraction of BRAF cfDNA was not related to the number of metastatic sites, but there was a tendency towards a higher mutant fraction when a higher [cfDNA] (Spearman rho 0.62; P = 0.08) was found.
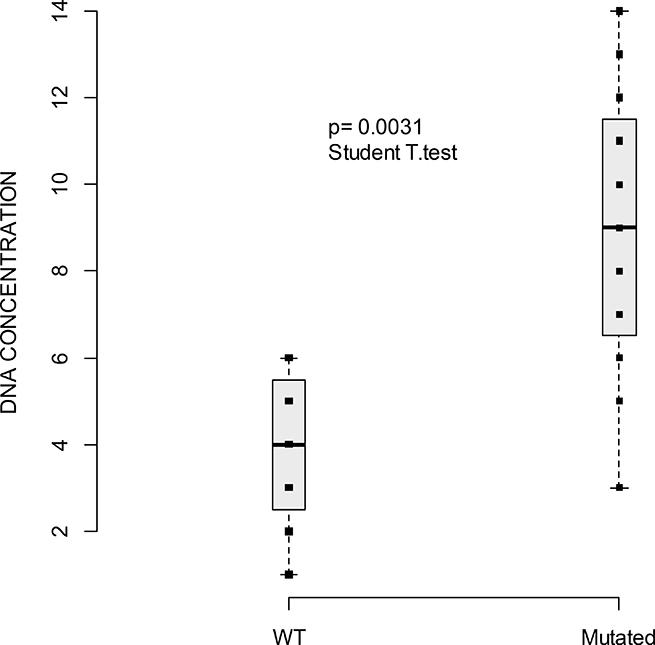
Figure 2: Boxplot of cfDNA concentration (measured with Qubit, ng/μl) and the presence of plasmatic mutations WT versus mutated. Horizontal line indicates median = 4 ng/μL (range: 1–6) for WT cfDNA [range 0.6 to 390.0 ng/μL] and median = 9 ng/μL (range: 3–14) for mutated cfDNA. Squares indicate the value of cfDNA. P-value of the Student T-test indicates a significant difference of cfDNA mean concentration between WT and mutated.
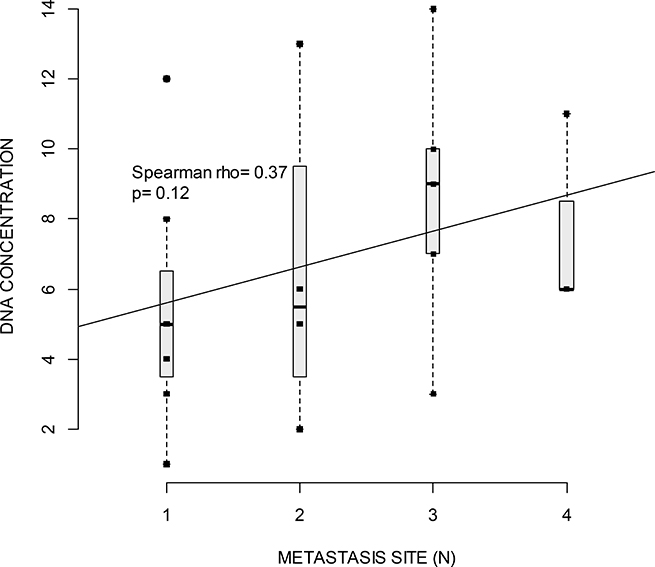
Figure 3: Boxplot of cfDNA concentration (measured with Qubit, ng/μl) and the number of metastatic sites. Horizontal lines indicate median for one, two, three or four metastatic sites. Squares indicate value of cfDNA. Spearman rho test indicates correlation between cfDNA and number of metastatic sites. The regression line show a non-significant positive correlation (p = 0.12) between the number of metastatic sites and cfDNA concentration.
DISCUSSION
Liquid biopsy is a non-invasive method that allows disease status monitoring during treatment of patients with cancer [24]. While detection of circulating melanoma cells in blood could have a prognostic impact for patients with metastatic melanoma, and despite a few interesting approaches [25, 26], these methods cannot assess the mutation status in clinical routine practice [27]. In contrast, the analysis of cfDNA can play a major role in personalized medicine, especially for cancers that can be treated with targeted therapies [28, 29]. Recently, Valpione et al. demonstrated that cfDNA could be a surrogate marker of TB and a prognostic factor for OS in patients with metastatic melanoma [30]. In our study, the number of metastatic sites and the [cfDNA] seemed to be relatively related but we did not find any significant correlation between the [cfDNA] at baseline or after treatment and the TB or OS, as previously described [31]. The lack of correlation between the TB and the [cfDNA] may probably be related to the number of metastatic sites that we added to our analysis to evaluate the TB. Precise evaluation of the TB by imaging is currently lacking in routine practice, probably because it is time consuming, and the RECIST 1.1 criteria that rely on “mono-dimensional” measurements of a maximum of 2 lesions per organ and 5 lesions in total, which is representative of all “involved organs”, does not seem satisfactory to accurately evaluate the relationship between the [cfDNA] and TB [30].
In our study, two BRAF V600E mutations detected on a FFPE sample were not detected in plasma in two patients with low TB and low [cfDNA]. This is consistent with recent reports that have demonstrated a lower [cfDNA] level in the presence of subcutaneous [23] or brain metastases [11, 22]. In patients with a cerebral metastasis, the analysis of the cerebrospinal fluid may be an alternative for detection of mutations [32]. Furthermore, in our study, a BRAF V600 mutation was detected in cfDNA with a small fraction of the mutant allele (0.36%) whereas the matched FFPE tissue sample was WT when using several methods. However, we could not reanalyze the sample as the cfDNA extracted with the Idylla™ device is not available after the experiment. Also the time between blood collection and biopsy was 6 months and even if it seems that it has a low risk impact on the discordant result, we can speculate that a minor emergent clone could have been identified. This would support the hypothesis that the cfDNA profile mirrored the genomic heterogeneity from multiple metastatic sites, which may not be detected with a single tissue biopsy specimen. Moreover, this tumor also harbored a NRAS Q61K mutation, which was detected on a subcutaneous biopsy FFPE sample and it seems likely, as previously mentioned for subcutaneous mutations [32], that the NRAS Q61K mutation of cfDNA was underrepresented in the blood, as it was not detected with either Idylla or NGS. Lastly, we cannot preclude a false positive result of the Idylla device in front of these elements add to the weak allelic frequency. Thus, we recommend that results with a low allele frequency should be interpreted carefully, be performed in duplicate or on serial analysis, and should always be compared with the total amount of cfDNA.
It is also interesting to monitor patients with a liquid biopsy for detection of mechanisms of resistance to treatment [5, 33, 34]. Although no resistance mutations were detected by NGS in plasma samples, probably due to the short time between initiation of the systemic therapy and the second blood draw, the persistence of the NRAS mutations in cfDNA suggests the presence of uncontrolled disease on therapy. Patients with NRAS mutations have a worse prognosis with a short OS, highlighting the need to identify these patients for more effective therapy or close surveillance [4].
In our study, the detection at baseline of a BRAF mutation in cfDNA was not predictive of response to treatment or OS. While baseline measurement of the BRAFV600 mutation in plasma with several methods, including Idylla™, has been shown to have prognostic value [10, 31]. In addition, study revealed that the increase in the mutant BRAF fraction preceded progression of the disease when detected with imagery giving a median interval of 25 days [35]. We did not find any correlation with OS or ORR with regard to the presence of a cfDNA mutation, the [cfDNA] or the mutant allele fraction. This may be in part due to the small size of the cohort, which also included tumors with NRAS mutations that are known to have a poorer prognosis.
Given that the sensitivity of detection of tumoral circulating DNA is low, the results should be interpreted with caution, ideally results should be confirmed with an orthogonal technique but it is sometimes unfeasible in routine manner. That’s why it is necessary to validate the limit of detection threshold (LOD) and to define the variation of the background noise in order to avoid any risk of false negatives and positives results. It is also recommended to use an internal validation control with mutations whose detection threshold is low (1 to 2%) close to the LOD. Interestingly, 2 uncommon exon 15 BRAF mutations (L597S; K501E) were identified by NGS in plasma specimens. These mutations are rare with a reported frequency of 1 to 4% [36, 37]. In our study, 17% (3/17) of patients harbored these mutations (11% L597S and 5% K501E). When considering only the cohort of double BRAF and NRAS WT tumors examined by pyrosequencing these mutations were present in 60% (3/5) of patients. Since they are known to be moderately sensitive to BRAF and MEK inhibitors [38], their presence at such a high rate in our population prompted us to perform a more comprehensive sequencing of “pan-negative” melanoma samples.
With the development of new PCR technologies (BEAMing, AS-PCR, ARMS, digital-PCR, NGS) and the improvement of the analytical sensitivity (0.005% to 0.1%) detection of cfDNA, even in early stage malignancies, has demonstrated potential clinical application [11, 39]. The clinical interest of cfDNA detection in breast, colon or lung cancers has been demonstrated for molecular profiling and early detection of resistance; disease monitoring; prognostication or detection of minimal residual disease [22, 40–42]. Recently, the Federal Drug Administration and the European Medicine Agency approved the plasmatic detection of the EGFR T790M resistant clone as a companion diagnostic test for second-line treatment of metastatic EGFR-mutant non-small cell cancer. Surprisingly, the published results for monitoring BRAF or NRAS mutant metastatic melanoma have not been adopted in routine clinical oncology. Calapre et al. [43] reviewed evidence of the clinical validity of cfDNA as a biomarker for clinical management in metastatic melanoma. CfDNA is an essential source of material for management of metastatic melanoma patients as it allows detection of somatic targeting mutations (BRAF, KIT), copy number variations [23, 44] and the tumor mutation load. In our opinion, we still have to determine the appropriate clinical context of cfDNA analysis.
We compared the three mains methods for cfDNA evaluation of melanoma patients that are available in our laboratory and proposed its clinical application (Table 2). The Idylla™ system offers a fast and easy-to-handle integrated “sample-to-result” approach with results available in less than 2 hours after blood, including plasma, preparation. It would be of particular interest for patients with a new metastatic melanoma, which often evolve rapidly and require urgent identification of the mutation status. Furthermore, the analysis of the BRAF and NRAS mutation status with cfDNA could be done in a single run, regardless of the mutation detected with the FFPE sample.
Table 2: Main technologies available at the Laboratory of Clinical and Experimental Pathology (University Côte d’Azur, Nice, France) and clinical applications
Method |
Mutation (Gene) |
Analytical sensitivity |
Material |
Turn around time |
Cost per patient tax-free (including plasma extraction) |
Proposal of clinical application |
|---|---|---|---|---|---|---|
Idylla™ assay (Biocartis, Mechelen, Belgium) |
Targeted mutation |
0.1 % |
1 ml plasma |
90 minutes |
115 € for BRAFV600 |
Fast identification of targeted mutation when tissue specimen not available / when polymetastatic tumor |
NGS |
Global mutational landscape |
0.1% |
20 ng DNA |
3 to 5 days (including data interpretation) |
543 € * |
Identification of targeted mutation when tissue specimen not available / when polymetastatic tumor |
Crystal digital-PCR |
Targeted mutation* |
0.3 copies DNA / μl |
10 ng DNA |
24 hours |
Approx. 114 € for BRAFV600E |
Longitudinal monitoring of residual disease, response to therapy, relapse |
When we compared NGS, which requires a complex workflow and a turnaround time of several days due to manual handling, to the Idylla™ system, the latter seems more effective in routine practice as it could also be implemented in a standard pathology laboratory. NGS has a more restricted indication, mainly during the course of the disease, to identify new biomarkers of resistance and potential therapeutic targets under the cover of a molecular medical and paramedical staff. Finally, digital-PCR allows precise quantification of tumor DNA and monitoring of response, especially in the era of checkpoint inhibitor therapy, with the need to distinguish between false “pseudo” progression and true progression and to oversee long-term responders. Thus, digital-PCR allows the study of the kinetics and variation of cfDNA over a longitudinal period with serial blood analyses.
The information obtained with cfDNA (e.g. somatic druggable mutations, copy number variation or the tumor mutation load) should not be ignored for the management of melanoma patients because it can provide a real-time and global assessment of the patient’s status [44, 45]. This analysis should be associated with imagery to reach the best practice [23] for patient care. Large prospective clinical studies are needed to evaluate the medical impact of cfDNA-guided decisions [43].
Our study holds a number of limitations, including the small size of the population and the limited number of longitudinal blood analyses. To our knowledge, we describe here, the first prospective study using the Idylla ctNRAS Assay and a NGS panel for cfDNA analysis in advanced melanoma patients before and after initiation of therapy. CfDNA is particularly interesting in polymetastatic patients as it could reflect genomic tumor heterogeneity.
In conclusion, we demonstrated that the detection of the BRAF and NRAS cfDNA status with the Idylla™ assay is feasible in a routine manner in a pathology laboratory, before and after systemic treatment of patients with metastatic melanoma. The assay reached acceptable concordance when compared with standard molecular analyses with matched tumor tissue. This approach could offer a good alternative to a surgical biopsy in fragile patients or patients with inaccessible metastatic sites to assess “druggable” molecular alterations.
MATERIALS AND METHODS
Patients and samples
Twenty-one consecutive treatment naïve patients with metastatic melanoma followed at the Department of Dermatology (Archet Hospital, Nice, France) were screened for the purpose of the study. 2 patients (#4; #6) were immediately excluded because the inclusion criteria were not met (e.g. both had choroidal melanoma, out of the scope of the study). Patients gave their informed consent and the study was conducted according to the Helsinki guidelines. All patients had a blood test at baseline and 16/19 (84%) had a blood test at their first evaluation two to three months (mean 81 days [range 63–94 days]) after initiation of treatment to evaluate the BRAF and NRAS status of cfDNA. The BRAF and NRAS status of FFPE blocks of corresponding tumors were available for all patients. Tumor specimens were selected by senior pathologists (EL, MI, CB, VH). The percentage of tumor cells (range 30%–80%, mean 65%) was evaluated independently by these pathologists, according to the French association for quality assurance in pathology (AFAQAP) procedures [46]. When several tissue samples were available the most recent resected tumor was selected. The time between tissue analysis and plasma collection ranged from less than 1 month to 38 months (mean 9.5 months). The main clinicopathological parameters are summarized in Table 1. Correlation of the results with the OS was evaluated. The therapeutic response was evaluated with RECIST1.1 criteria (Response Evaluation Criteria in Solid Tumors) on targeted lesions on CT imagery, the same day of the blood draw or within less than one month. The tumor burden (TB) was evaluated by addition of the number of metastatic sites (Table 1).
Circulating free DNA
Blood samples (10 mL) were collected in EDTA tubes and processed within 4 hours. The plasma was obtained by two rounds of centrifugation (10 minutes, 2000 g at 4° C) aliquoted into 1mL tubes and stored at –80° C until use. For the Idylla assay, DNA was extracted with the specific cartridge and on-board reagents, whereas for Ion Torrent PGM sequencing the DNA was extracted with the Kit QIAamp circulating Nucleic Acid (Qiagen, Hilden, Germany). The cell free DNA concentration was measured with the Qubit Fluorometric Quantitation (ThermoFisher Scientific) double strand (ds) DNA assay kit.
The BRAF and NRAS mutation status evaluated with Idylla
The Idylla™ platform (Biocartis, Mechelen, Belgium) is a fully cartridge-based automated platform processing with all reagents on-board [31]. The sensitivity was high, down to 0.1% mutant allele, according to the manufacturer’s instructions. The plasma was tested for BRAF and NRAS before and after treatment (when available) with the Idylla™ ctNRAS-BRAF Mutation Test (prototype). For the cfDNA assay, 1 ml of plasma was directly placed into the cartridge. After a 90-minute run and less than 1-minute hands-on time, all steps were automatically performed inside the cartridge and final reports were directly available on the console after an automatic on-board post-PCR curve analysis. As the real time PCR using allele specific primers separated into two chambers the results were given as “V600E/E2/D Mutation” or “V600K/R/M Mutation” or Wild Type. NRAS mutations were detected in the same way in codons 12, 13, 59, 61, 117, 146 (according to manufacturer’s guidelines). For the BRAF Assay, the BRAF mutant fraction was established on calculation of the ΔCq (CqMut-CqWT), but this was not possible for NRAS (Cq not available).
BRAF and NRAS Pyrosequencing
DNA extraction was performed with the QiAmp DNA FFPE Tissue Kit (Qiagen, Hilden, Germany) and pyrosequencing analysis was performed as previously described with the therascreen BRAF Pyro Kit (Qiagen) and the therascreen NRAS Pyro Kit (Qiagen) [47–49]. The platform of the LCEP, (Nice, France) has been accredited for molecular testing by pyrosequencing of exons 11 and 15 of the BRAF gene and for codon 12, 13 and 61 of the NRAS gene. The laboratory holds full COFRAC (“Comité Français d’Accréditation”) accreditation for these assays, according to the ISO 15189 norm (http://www.cofrac.fr). The analytical sensitivity of the pyrosequencing method was considered to be 5% according to the manufacturer (Qiagen). Pyrogram profile analyses and interpretation of the results were done blindly with PyromarkQ24 (Qiagen) by five independent assessors (EL, MI, VL, VH OB).
Next generation sequencing
Sequencing libraries were prepared from cfDNA using the Ion AmpliSeq™ Cancer Hotpost Panel v2, encompassing for of a broad set of 50 cancer-relevant genes (list of targeted exonic, intronic and regulatory regions available on https://www.ampliseq.com). According to manufacturer’s protocol; 10 ng of DNA for each sample was used as input for library preparation with the Ion AmpliSeqTM Library kit 2.0 (Thermo Fisher). The pooled barcoded libraries were processes on the Ion Chef™ System. The FASTQs sequencing data were aligned to the human genome using the Torrent Suit server. Analysis was then performed with two software VariantCaller (version 2.4.1) and Nextgene (2.4.1-UG001) by two pathologists and three bioinformaticians (VL, OB, CR).
Statistical analysis
Qualitative data are presented as absolute and relative frequencies and are compared according to the Chi2 test or Fisher exact test when necessary. Quantitative data are presented as mean, standard deviation, median and range and compared using the Student’s t-test or Wilcoxon test. Survival data are defined between the date of the metastasis diagnostic and the event onset date. Survival curves are compared with the Log-Rank test. Median follow-up was assessed using the reversed Kaplan–Meier method. Concordance between the baseline cfDNA BRAF and NRAS mutation status and the results in the reference tissue were assessed using the Cohen’s kappa coefficient (κ), if indicated, we also evaluated sensitivity and specificity. Correlation between cfDNA levels, the BRAF mutant fraction and the baseline tumor burden were assessed using Spearman correlation coefficients. All statistical analyses were two-sided and a p-value of less than 0.05 was considered as statistically significant. All calculations were made with R.3.4.3 software.
Author contributions
Elodie Long-Mira, Marius Ilie, Emmanuel Chamorey, Florence Leduff-Blanc, Henri Montaudié, Catherine Butori, Coraline Bence, Véronique Hofman, Jean-Philippe Lacour, Paul Hofman designed and supervised the study and wrote the manuscript; Virginie Tanga, Maryline Allégra, Virginie Lespinet, Olivier Bordone, Bonnetaud Christelle, and Renaud Schiappa designed and performed experiments.
All authors have reviewed and agreed to the information before submission.
CONFLICTS OF INTEREST
The authors declare that they have no competing interest.
FUNDING
This work was supported by a grant from the “Délégation à la Recherche Clinique et à l’Innovation” of University Hospital of Nice “2013”; the Cancéropole PACA; the “Conseil Départemental des Alpes-Maritimes”; the “Région Provence Alpes-Côte d’Azur”, France.
REFERENCES
1. Larkin J, Ascierto PA, Dréno B, Atkinson V, Liszkay G, Maio M, Mandalà M, Demidov L, Stroyakovskiy D, Thomas L, de la Cruz-Merino L, Dutriaux C, Garbe C, et al. Combined vemurafenib and cobimetinib in BRAF-mutated melanoma. N Engl J Med. 2014; 371:1867–76. https://doi.org/10.1056/NEJMoa1408868.
2. Read J. Recent advances in cutaneous melanoma: towards a molecular model and targeted treatment. Australas J Dermatol. 2013; 54:163–72. https://doi.org/10.1111/ajd.12013.
3. Robert C, Schachter J, Long GV, Arance A, Grob JJ, Mortier L, Daud A, Carlino MS, McNeil C, Lotem M, Larkin J, Lorigan P, Neyns B, et al, and KEYNOTE-006 investigators. Pembrolizumab versus Ipilimumab in Advanced Melanoma. N Engl J Med. 2015; 372:2521–32. https://doi.org/10.1056/NEJMoa1503093.
4. Jakob JA, Bassett RL Jr, Ng CS, Curry JL, Joseph RW, Alvarado GC, Rohlfs ML, Richard J, Gershenwald JE, Kim KB, Lazar AJ, Hwu P, Davies MA. NRAS mutation status is an independent prognostic factor in metastatic melanoma. Cancer. 2012; 118:4014–23. https://doi.org/10.1002/cncr.26724.
5. Girotti MR, Gremel G, Lee R, Galvani E, Rothwell D, Viros A, Mandal AK, Lim KH, Saturno G, Furney SJ, Baenke F, Pedersen M, Rogan J, et al. Application of Sequencing, Liquid Biopsies, and Patient-Derived Xenografts for Personalized Medicine in Melanoma. Cancer Discov. 2016; 6:286–99. https://doi.org/10.1158/2159-8290.CD-15-1336.
6. Long E, Hofman V, Ilie M, Lespinet V, Bonnetaud C, Bordone O, Gavric-Tanga V, Washetine K, Gaziello MC, Mauro V, Lassalle S, Selva E, Zahaf K, et al. [Setting up a department of molecular pathology in oncology in a pathology laboratory (LPCE, CHU de Nice)][Article in French]. Ann Pathol. 2013; 33:24–37. https://doi.org/10.1016/j.annpat.2012.12.003.
7. Long-Mira E, Washetine K, Hofman P. Sense and nonsense in the process of accreditation of a pathology laboratory. Virchows Arch. 2016; 468:43–49. https://doi.org/10.1007/s00428-015-1837-1.
8. Lipson EJ, Velculescu VE, Pritchard TS, Sausen M, Pardoll DM, Topalian SL, Diaz LA Jr. Circulating tumor DNA analysis as a real-time method for monitoring tumor burden in melanoma patients undergoing treatment with immune checkpoint blockade. J Immunother Cancer. 2014; 2:42. https://doi.org/10.1186/s40425-014-0042-0.
9. Gangadhar TC, Savitch SL, Yee SS, Xu W, Huang AC, Harmon S, Lieberman DB, Soucier D, Fan R, Black TA, Morrissette JJ, Salathia N, Waters J, et al. Feasibility of monitoring advanced melanoma patients using cell-free DNA from plasma. Pigment Cell Melanoma Res. 2018; 31:73–81. https://doi.org/10.1111/pcmr.12623.
10. Knol AC, Vallée A, Herbreteau G, Nguyen JM, Varey E, Gaultier A, Théoleyre S, Saint-Jean M, Peuvrel L, Brocard A, Quéreux G, Khammari A, Denis MG, Dréno B. Clinical significance of BRAF mutation status in circulating tumor DNA of metastatic melanoma patients at baseline. Exp Dermatol. 2016; 25:783–88. https://doi.org/10.1111/exd.13065.
11. Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, Bartlett BR, Wang H, Luber B, Alani RM, Antonarakis ES, Azad NS, Bardelli A, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014; 6:224ra24. https://doi.org/10.1126/scitranslmed.3007094.
12. Lebofsky R, Decraene C, Bernard V, Kamal M, Blin A, Leroy Q, Rio Frio T, Pierron G, Callens C, Bieche I, Saliou A, Madic J, Rouleau E, et al. Circulating tumor DNA as a non-invasive substitute to metastasis biopsy for tumor genotyping and personalized medicine in a prospective trial across all tumor types. Mol Oncol. 2015; 9:783–90. https://doi.org/10.1016/j.molonc.2014.12.003.
13. Santiago-Walker A, Gagnon R, Mazumdar J, Casey M, Long GV, Schadendorf D, Flaherty K, Kefford R, Hauschild A, Hwu P, Haney P, O’Hagan A, Carver J, et al. Correlation of BRAF Mutation Status in Circulating-Free DNA and Tumor and Association with Clinical Outcome across Four BRAFi and MEKi Clinical Trials. Clin Cancer Res. 2016; 22:567–74. https://doi.org/10.1158/1078-0432.CCR-15-0321.
14. Sanmamed MF, Fernández-Landázuri S, Rodríguez C, Zárate R, Lozano MD, Zubiri L, Perez-Gracia JL, Martín-Algarra S, González A. Quantitative cell-free circulating BRAFV600E mutation analysis by use of droplet digital PCR in the follow-up of patients with melanoma being treated with BRAF inhibitors. Clin Chem. 2015; 61:297–304. https://doi.org/10.1373/clinchem.2014.230235.
15. Ascierto PA, Minor D, Ribas A, Lebbe C, O’Hagan A, Arya N, Guckert M, Schadendorf D, Kefford RF, Grob JJ, Hamid O, Amaravadi R, Simeone E, et al. Phase II trial (BREAK-2) of the BRAF inhibitor dabrafenib (GSK2118436) in patients with metastatic melanoma. J Clin Oncol. 2013; 31:3205–11. https://doi.org/10.1200/JCO.2013.49.8691.
16. El Messaoudi S, Rolet F, Mouliere F, Thierry AR. Circulating cell free DNA: preanalytical considerations. Clin Chim Acta. 2013; 424:222–30. https://doi.org/10.1016/j.cca.2013.05.022.
17. Norton SE, Luna KK, Lechner JM, Qin J, Fernando MR. A new blood collection device minimizes cellular DNA release during sample storage and shipping when compared to a standard device. J Clin Lab Anal. 2013; 27:305–11. https://doi.org/10.1002/jcla.21603.
18. Shi H, Hugo W, Kong X, Hong A, Koya RC, Moriceau G, Chodon T, Guo R, Johnson DB, Dahlman KB, Kelley MC, Kefford RF, Chmielowski B, et al. Acquired resistance and clonal evolution in melanoma during BRAF inhibitor therapy. Cancer Discov. 2014; 4:80–93. https://doi.org/10.1158/2159-8290.CD-13-0642.
19. Zaretsky JM, Garcia-Diaz A, Shin DS, Escuin-Ordinas H, Hugo W, Hu-Lieskovan S, Torrejon DY, Abril-Rodriguez G, Sandoval S, Barthly L, Saco J, Homet Moreno B, Mezzadra R, et al. Mutations Associated with Acquired Resistance to PD-1 Blockade in Melanoma. N Engl J Med. 2016; 375:819–29. https://doi.org/10.1056/NEJMoa1604958.
20. Shain AH, Yeh I, Kovalyshyn I, Sriharan A, Talevich E, Gagnon A, Dummer R, North J, Pincus L, Ruben B, Rickaby W, D’Arrigo C, Robson A, Bastian BC. The Genetic Evolution of Melanoma from Precursor Lesions. N Engl J Med. 2015; 373:1926–36. https://doi.org/10.1056/NEJMoa1502583.
21. Siroy AE, Boland GM, Milton DR, Roszik J, Frankian S, Malke J, Haydu L, Prieto VG, Tetzlaff M, Ivan D, Wang WL, Torres-Cabala C, Curry J, et al. Beyond BRAF(V600): clinical mutation panel testing by next-generation sequencing in advanced melanoma. J Invest Dermatol. 2015; 135:508–15. https://doi.org/10.1038/jid.2014.366.
22. De Mattos-Arruda L, Mayor R, Ng CK, Weigelt B, Martínez-Ricarte F, Torrejon D, Oliveira M, Arias A, Raventos C, Tang J, Guerini-Rocco E, Martínez-Sáez E, Lois S, et al. Cerebrospinal fluid-derived circulating tumour DNA better represents the genomic alterations of brain tumours than plasma. Nat Commun. 2015; 6:8839. https://doi.org/10.1038/ncomms9839.
23. Wong SQ, Raleigh JM, Callahan J, Vergara IA, Ftouni S, Hatzimihalis A, Colebatch AJ, Li J, Semple T, Doig K, Mintoff C, Sinha D, Yeh P, et al. Circulating Tumor DNA Analysis and Functional Imaging Provide Complementary Approaches for Comprehensive Disease Monitoring in Metastatic Melanoma. JCO Precision Oncology. 2017; 1–14. https://doi.org/10.1200/po.16.00009.
24. Arneth B. Update on the types and usage of liquid biopsies in the clinical setting: a systematic review. BMC Cancer. 2018; 18:527. https://doi.org/10.1186/s12885-018-4433-3.
25. Long E, Ilie M, Bence C, Butori C, Selva E, Lalvée S, Bonnetaud C, Poissonnet G, Lacour JP, Bahadoran P, Brest P, Gilson E, Ballotti R, et al. High expression of TRF2, SOX10, and CD10 in circulating tumor microemboli detected in metastatic melanoma patients. A potential impact for the assessment of disease aggressiveness. Cancer Med. 2016; 5:1022–30. https://doi.org/10.1002/cam4.661.
26. Khoja L, Lorigan P, Dive C, Keilholz U, Fusi A. Circulating tumour cells as tumour biomarkers in melanoma: detection methods and clinical relevance. Ann Oncol. 2015; 26:33–39. https://doi.org/10.1093/annonc/mdu207.
27. Hofman V, Ilie M, Long-Mira E, Giacchero D, Butori C, Dadone B, Selva E, Tanga V, Passeron T, Poissonnet G, Emile JF, Lacour JP, Bahadoran P, Hofman P. Usefulness of immunocytochemistry for the detection of the BRAF(V600E) mutation in circulating tumor cells from metastatic melanoma patients. J Invest Dermatol. 2013; 133:1378–81. https://doi.org/10.1038/jid.2012.485.
28. Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, Thornton K, Agrawal N, Sokoll L, Szabo SA, Kinzler KW, Vogelstein B, Diaz LA Jr. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008; 14:985–90. https://doi.org/10.1038/nm.1789.
29. Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011; 11:426–37.
30. Valpione S, Gremel G, Mundra P, Middlehurst P, Galvani E, Girotti MR, Lee RJ, Garner G, Dhomen N, Lorigan PC, Marais R. Plasma total cell-free DNA (cfDNA) is a surrogate biomarker for tumour burden and a prognostic biomarker for survival in metastatic melanoma patients. Eur J Cancer. 2018; 88:1–9. https://doi.org/10.1016/j.ejca.2017.10.029.
31. Janku F, Huang HJ, Claes B, Falchook GS, Fu S, Hong D, Ramzanali NM, Nitti G, Cabrilo G, Tsimberidou AM, Naing A, Piha-Paul SA, Wheler JJ, et al. BRAF Mutation Testing in Cell-Free DNA from the Plasma of Patients with Advanced Cancers Using a Rapid, Automated Molecular Diagnostics System. Mol Cancer Ther. 2016; 15:1397–404. https://doi.org/10.1158/1535-7163.MCT-15-0712.
32. Ballester LY, Glitza Oliva IC, Douse DY, Chen MM, Lan C, Haydu LE, Huse JT, Roy-Chowdhuri S, Luthra R, Wistuba II, Davies MA. Evaluating Circulating Tumor DNA From the Cerebrospinal Fluid of Patients With Melanoma and Leptomeningeal Disease. J Neuropathol Exp Neurol. 2018; 77:628–35. https://doi.org/10.1093/jnen/nly046.
33. Gray ES, Rizos H, Reid AL, Boyd SC, Pereira MR, Lo J, Tembe V, Freeman J, Lee JH, Scolyer RA, Siew K, Lomma C, Cooper A, et al. Circulating tumor DNA to monitor treatment response and detect acquired resistance in patients with metastatic melanoma. Oncotarget. 2015; 6:42008–18. https://doi.org/10.18632/oncotarget.5788.
34. Tsao SC, Weiss J, Hudson C, Christophi C, Cebon J, Behren A, Dobrovic A. Monitoring response to therapy in melanoma by quantifying circulating tumour DNA with droplet digital PCR for BRAF and NRAS mutations. Sci Rep. 2015; 5:11198. https://doi.org/10.1038/srep11198.
35. Schreuer M, Meersseman G, van Den Herrewegen S, Jansen Y, Seremet T, Bott A, Chevolet I, Wilgenhof S, Maertens G, Neyns B. Applications for quantitative measurement of BRAF V600 mutant cell-free tumor DNA in the plasma of patients with metastatic melanoma. Melanoma Res. 2016; 26:157–63. https://doi.org/10.1097/CMR.0000000000000224.
36. Forbes SA, Bindal N, Bamford S, Cole C, Kok CY, Beare D, Jia M, Shepherd R, Leung K, Menzies A, Teague JW, Campbell PJ, Stratton MR, Futreal PA. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 2011; 39:D945–50. https://doi.org/10.1093/nar/gkq929.
37. Voskoboynik M, Mar V, Mailer S, Colebatch A, Fennessy A, Logan A, Hewitt C, Cebon J, Kelly J, McArthur G. Clinicopathological characteristics associated with BRAF(K601E) and BRAF(L597) mutations in melanoma. Pigment Cell Melanoma Res. 2016; 29:222–28. https://doi.org/10.1111/pcmr.12450.
38. Dahlman KB, Xia J, Hutchinson K, Ng C, Hucks D, Jia P, Atefi M, Su Z, Branch S, Lyle PL, Hicks DJ, Bozon V, Glaspy JA, et al. BRAF(L597) mutations in melanoma are associated with sensitivity to MEK inhibitors. Cancer Discov. 2012; 2:791–97. https://doi.org/10.1158/2159-8290.CD-12-0097.
39. Alix-Panabières C, Pantel K. Clinical Applications of Circulating Tumor Cells and Circulating Tumor DNA as Liquid Biopsy. Cancer Discov. 2016; 6:479–91. https://doi.org/10.1158/2159-8290.CD-15-1483.
40. Knebel FH, Bettoni F, Shimada AK, Cruz M, Alessi JV, Negrão MV, Reis LF, Katz A, Camargo AA. Sequential liquid biopsies reveal dynamic alterations of EGFR driver mutations and indicate EGFR amplification as a new mechanism of resistance to osimertinib in NSCLC. Lung Cancer. 2017; 108:238–41. https://doi.org/10.1016/j.lungcan.2017.04.004.
41. Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, Dunning MJ, Gale D, Forshew T, Mahler-Araujo B, Rajan S, Humphray S, Becq J, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013; 368:1199–209. https://doi.org/10.1056/NEJMoa1213261.
42. Garcia-Murillas I, Schiavon G, Weigelt B, Ng C, Hrebien S, Cutts RJ, Cheang M, Osin P, Nerurkar A, Kozarewa I, Garrido JA, Dowsett M, Reis-Filho JS, et al. Mutation tracking in circulating tumor DNA predicts relapse in early breast cancer. Sci Transl Med. 2015; 7:302ra133. https://doi.org/10.1126/scitranslmed.aab0021.
43. Calapre L, Warburton L, Millward M, Ziman M, Gray ES. Circulating tumour DNA (ctDNA) as a liquid biopsy for melanoma. Cancer Lett. 2017; 404:62–69. https://doi.org/10.1016/j.canlet.2017.06.030.
44. Chan KC, Jiang P, Zheng YW, Liao GJ, Sun H, Wong J, Siu SS, Chan WC, Chan SL, Chan AT, Lai PB, Chiu RW, Lo YM. Cancer genome scanning in plasma: detection of tumor-associated copy number aberrations, single-nucleotide variants, and tumoral heterogeneity by massively parallel sequencing. Clin Chem. 2013; 59:211–24. https://doi.org/10.1373/clinchem.2012.196014.
45. Leary RJ, Sausen M, Kinde I, Papadopoulos N, Carpten JD, Craig D, O’Shaughnessy J, Kinzler KW, Parmigiani G, Vogelstein B, Diaz LA Jr, Velculescu VE. Detection of chromosomal alterations in the circulation of cancer patients with whole-genome sequencing. Sci Transl Med. 2012; 4:162ra154. https://doi.org/10.1126/scitranslmed.3004742.
46. Lhermitte B, Egele C, Weingertner N, Ambrosetti D, Dadone B, Kubiniek V, Burel-Vandenbos F, Coyne J, Michiels JF, Chenard MP, Rouleau E, Sabourin JC, Bellocq JP. Adequately defining tumor cell proportion in tissue samples for molecular testing improves interobserver reproducibility of its assessment. Virchows Arch. 2017; 470:21–27. https://doi.org/10.1007/s00428-016-2042-6.
47. Ilie MI, Lassalle S, Long-Mira E, Bonnetaud C, Bordone O, Lespinet V, Lamy A, Sabourin JC, Haudebourg J, Butori C, Guevara N, Peyrottes I, Sadoul JL, et al. Diagnostic value of immunohistochemistry for the detection of the BRAF(V600E) mutation in papillary thyroid carcinoma: comparative analysis with three DNA-based assays. Thyroid. 2014; 24:858–66. https://doi.org/10.1089/thy.2013.0302.
48. Ilie M, Long-Mira E, Funck-Brentano E, Lassalle S, Butori C, Lespinet-Fabre V, Bordone O, Gay A, Zahaf K, Poissonnet G, Lacour JP, Bahadoran P, Ballotti R, et al. Immunohistochemistry as a potential tool for routine detection of the NRAS Q61R mutation in patients with metastatic melanoma. J Am Acad Dermatol. 2015; 72:786–93. https://doi.org/10.1016/j.jaad.2015.01.012.
49. Lade-Keller J, Rømer KM, Guldberg P, Riber-Hansen R, Hansen LL, Steiniche T, Hager H, Kristensen LS. Evaluation of BRAF mutation testing methodologies in formalin-fixed, paraffin-embedded cutaneous melanomas. J Mol Diagn. 2013; 15:70–80. https://doi.org/10.1016/j.jmoldx.2012.08.003.

