INTRODUCTION
The commitment of hematopoietic stem cells to each cell lineage is strictly controlled. Specific extracellular stimuli and transcription factors play important roles in the development of B cells from hematopoietic stem cells. The transcription factors SPI1 (PU.1) and IKZF1 (Ikaros) work together during the early developmental stages, while TCF3 (E2A), Early B cell Factor 1 (EBF1) and Paired box 5 (PAX5) are crucial from pro-B cells to mature B cell stages [1]. PAX5 is one of the 9 members (PAX1 to PAX9) of the highly conserved paired-box (PAX) domain family of transcription factors [2, 3] characterized by a conserved PAIRED DNA-binding domain at the N-terminal part of the protein and involved in the regulation of tissue homeostasis.
During hematopoiesis, PAX5 is expressed from the pro-B cell stage and has to be turned off to allow plasma-cell transition [4]. PAX5 is crucial for the maintenance of the B lymphoid lineage identity [5, 6] and for suppression of alternative lineage choices [1, 7]. PAX5 also enhances the transcription of B cell specific genes and participates in the chromatin-remodeling of the immunoglobulin heavy chain (IGH) locus, ensuring its contraction during VDJ recombination [8]. At later stages, PAX5 regulates the IGH 3ʹ regulatory region (3ʹRR). The 3ʹRR is a 30 kb-long cis-acting regulation element of the immunoglobulin heavy chain (IGH) locus containing four enhancers in mice (hs1,2, hs3a, hs3b and hs4) with a strict B lineage specificity. They have been implicated in the late stages of B cell differentiation with a crucial role in class switch recombination (CSR) and somatic hypermutation (SHM) [9–12].
Pax5 homozygous inactivation in mouse leads to a blockade at the pro-B cell stage [6]. Pax5-deficient pro-B cells are able to transdifferentiate into other cell types, such as T, natural killer, and dendritic cells [1, 13] while mature B cells can dedifferentiate into lymphoid precursor cells upon Pax5 loss even at late stages of B cell differentiation as shown by Pax5 conditional inactivation [14].
In vertebrates, PAX5 expression is controlled by two distinct promoters: a distal P1a and a proximal P1b [15] which initiate transcription from two alternative 5′ first exons (exons 1A and 1B respectively) leading to the expression of two isoforms, PAX5A and PAX5B. The only structural difference between PAX5A and PAX5B consists in a short different N-terminal sequence. Exon 1A encodes 15 amino acids that differ totally from the 14 amino acids encoded by the exon 1B sequence [16]. Despite high sequence similarities between these two isoforms, they show different patterns of expression: PAX5B is transcribed in B cells, central nervous system and testis, while PAX5A, whose product is also named B cell–specific activator protein (BSAP) is restricted to the B cell lymphoid lineage [4]. However their respective roles in B cell differentiation have never been determined. Our work aims to detail the mechanisms that control the expression of PAX5A and PAX5B isoforms along B cell development and their effect on B cell differentiation.
RESULTS
Pax5 expression in B cell differentiation is independent of adjacent genes
The murine Pax5 gene encompasses a region of 392 kb of chromosome 4 from the end of its upstream neighbor gene, Melk, to the start of its downstream neighbor gene, Zcchc7 (Figure 1A). Pax5 has a reverse orientation compared to its two neighbors, from telomere to centromere (Figure 1A). The human PAX5 gene has a similar organization covering a slightly larger region of 444 kb on chromosome 9. In order to clarify the transcriptional activities within the Pax5 locus, quantitative RT-PCR (QPCR) was performed to measure the overall expression of Pax5 transcripts and its neighboring genes (Melk and Zcchc7), together with Abl1 as a widely expressed control gene and Ebf1 as a transcriptional target of Pax5. Their expression were measured in a series of murine B cell lines representing different stages of B cell differentiation (from the less to the most differentiated: Ba/F3, 70Z3, 38B9, 18.81, A20 and WEHI-231) along with murine primary tissues (T and B cells, Figure 1B). Since Pax5 expression is regulated by Ebf1, Pax5 expression is highly correlated to the expression of Ebf1, as expected (Pearson correlation, r2 = 0.85). In contrast, expression of Pax5 is independent of the expression of its two neighboring genes, Melk and Zcchc7 (Pearson correlation, r2 = 0.40 and r2 = 0.54 respectively), suggesting that the regulatory elements of Pax5 are not shared by Melk and Zcchc7.
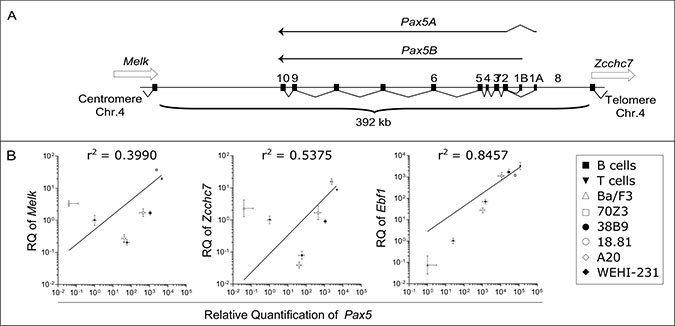
Figure 1: Expression of Pax5 isoforms is independent of the expression of neighboring genes. (A) Schematic organization of the genomic region of murine Pax5 gene. Pax5 is composed of 11 exons, the first two (exons 1A and 1B) being alternatively used to generate two isoforms (Pax5A and Pax5B respectively). Pax5 gene is flanked by Melk and Zcchc7 genes. (B) Correlation between Melk, Zcchc7 or Ebf1 expression and Pax5 expression. Quantitative PCR (QPCR) was performed at least as triplicate on Ba/F3, 70Z3, 38B9, 18.81, A20 and WEHI-231 cell lines and on B and T cells. Relative expressions (RQ) to Abl1 expression are expressed as mean with error bars representing RQMIN and RQMAX and constitute the acceptable error level for a 95% confidence interval according to Student’s t test. The square of the Pearson correlation (r2) is indicated for each comparison.
Pax5 isoforms are differentially expressed during B cell differentiation
Two major 5ʹ isoforms of Pax5 are expressed during B cell differentiation. Pax5A expression is driven by the promoter 1A and Pax5B by the promoter 1B [using alternative first exons (1A and 1B respectively, Figure 1A)]. We detailed the expression pattern of the two Pax5 isoforms during murine B cell differentiation using specific primers of these two isoforms on sorted B cell subsets from bone marrow. Pax5B has a low expression which does not vary during B cell differentiation. In contrast, Pax5A expression is strongly modulated during B cell differentiation with a higher expression in immature B cells (Figure 2A, left panel).
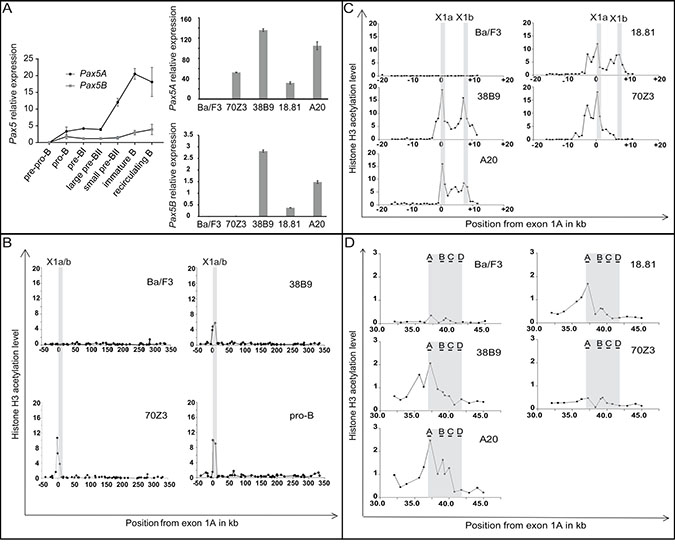
Figure 2: Correlation between Pax5 isoforms expression and chromatin acetylation of the Pax5 locus. (A) QPCR of Pax5A and Pax5B isoforms on B cells during differentiation and in various cell lines. Expression of Pax5 isoforms during medullar murine B cell differentiation was quantified on B cell subsets. Expression relative to Abl1 are presented as mean and standard deviation (left panel); Expression of Pax5A (upper right panel) or Pax5B (lower right panel) on Ba/F3,70Z3, 38B9, 18.81, A20 cell lines and pro-B cells from murine bone marrow are expressed relative to Gapdh expression as mean and standard deviation. (B–D) Chromatin Immunoprecipitation (ChIP) analysis of Pax5 locus on Ba/F3, 38B9, 70Z3 cell lines and murine pro-B sorted cells: QPCR was performed using primers covering the most conserved region (B) or located every 1 kb throughout promoter (C) or enhancer (D) regions. The increase over background was calculated by comparison with immunoprecipitates obtained using rabbit IgG antibody. Peaks of H3 acetylation are labeled according to their positions relative to the first nucleotide of Pax5 exon lA.
Expression of Pax5 isoforms is correlated with the histone H3 acetylation of their respective promoter
We defined the expression pattern of Pax5 isoforms on B cell lines. The 38B9, 18.81, A20 cell lines expressed both Pax5 isoforms. The early blocked 70Z3 B cell line expressed only Pax5A isoform. The Ba/F3 cell line did not express any of the two isoforms (Figure 2A, right panel).
Using the fact that B cell lines differentially expressed Pax5 isoforms, we detailed the transcriptional regulation within the 392 kb genomic region based on the acetylation of histone H3, a major mark of active regulatory elements [17] at low resolution over the 50 most conserved regions in vertebrates. We observed a peak of acetylation covering Pax5 promoters A and B in pro-B cells and the B cell lines expressing at least Pax5A in contrast to Ba/F3 (Figure 2B). We increased the resolution of the ChIP analysis to 1 kb over this region on five B cell lines (Figure 2C). The three cell lines expressing both Pax5 isoforms (38B9, 18.81 and A20) exhibited two acetylation peaks located upstream of exon 1A (defined as the location origin, 0 kb) and 1B (+7 kb). 70Z3 expressing only Pax5A isoform had peaks at 0 kb and -4kb, but none at the +7 kb while Ba/F3 displayed no acetylation peak (Figure 2C).
Decker et al. identified an enhancer region located in the intron 5–6 of Pax5 [24]. We analyzed the acetylation status of this region and showed that the acetylation is weak compared to the promoter regions. Interestingly, only the cell lines expressing both isoforms have an H3 acetylation mark on the enhancer region (Figure 2D).
Both Pax5 isoforms restore early B cell differentiation
We took advantage of the Pax5-/- cells which do not express any of the two isoforms and are subsequently blocked at a pro-B cell stage but still proliferating in presence of IL7. PAX5 isoforms were individually transduced in Pax5-/- pro-B cells and assessed for their ability to reinitiate B cell differentiation, using the expression of Cd19, a specific target of PAX5, and the appearance of IgMposIgΚpos B cells as hallmarks of B cell progression. Cells infected with the empty vector (MIE, only expressing eGFP) did not induce expression of Cd19. In contrast, both isoforms induced the expression of surface Cd19 in presence or absence of IL7 (Figure 3A, respectively upper and lower panels). Furthermore, both isoforms were also able to initiate the expression of IgM and IgΚ at the cell surface after removal of IL7 (Figure 3B, lower panel).
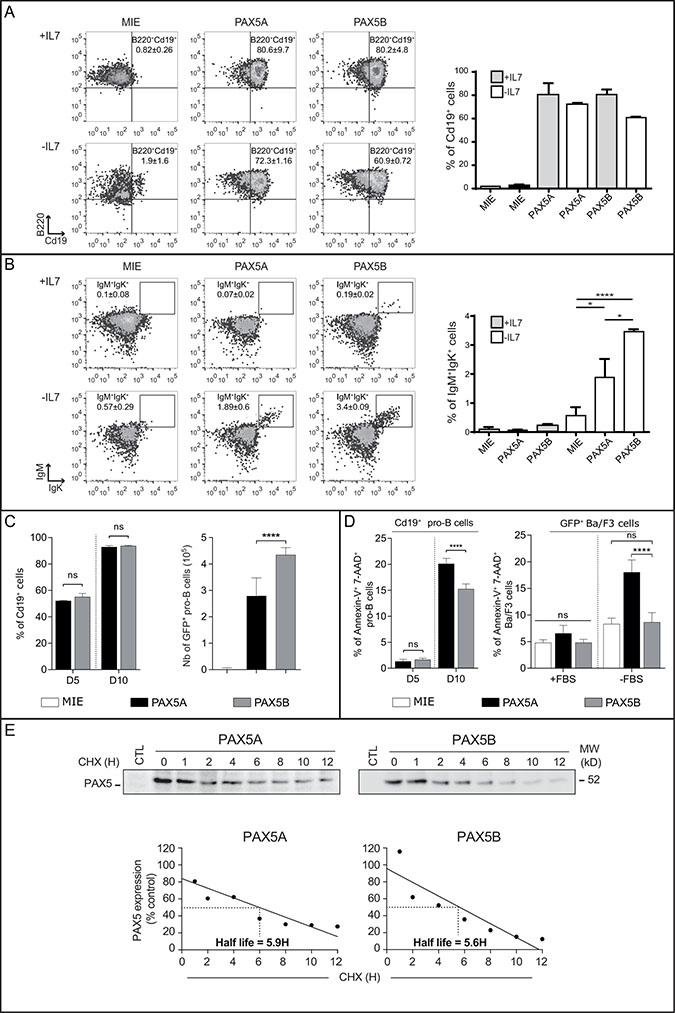
Figure 3: Both PAX5A and PAX5B can restore B cell differentiation program in complemented Pax5-/- pro-B cells. (A–B) Representative FACS analysis and statistics of differentiation of three independent infections of Pax5-/- pro-B cells. Cells were gated on FSC/SSC criteria, 7-AAD negative population (living cells) and eGFP-positive cells (infected cells) (not shown). Gated cells were analyzed for (A) the presence of B220 and Cd19 (the percentage of Cd19+ cells is indicated on the right panel graph) or (B) IgM and Igk membrane markers (the percentage of IgM+/Igk+ cells is indicated on the right panel graph) in two different conditions: with (+IL7, upper line) or without IL7 (-IL7, lower line) for three days. The first column show Pax5-/- cells infected with the eGFP-expressing vector (MIE), the second column with PAX5A retroviral construct and the third column with the PAX5B retroviral construct. The mean and the standard deviation of percentage of each population compared to the parental gate are indicated, *p < 0.05, **** p < 0.001. (C) Mean and standard deviation of percentage of Cd19+ cells (left panel); Absolute number of GFP+ Pax5-/- pro-B complemented with eGFP (MIE), PAX5A or PAX5B after 3 days of culture for three independent infections, ****p < 0.001 (right panel). (D) Percentage of 7-AADpos AnnexinVpos cells after 5 or 10 days in presence of IL7 (left panel) for three independent infections with PAX5A- or PAX5B-expressing vectors. ****p < 0.001. (E) Quantification of PAX5A and PAX5B proteins by Western-blot using an anti-Pax5 antibody after cycloheximide treatment for the indicated period of time on Ba/F3 infected cells demonstrating a similar half-life.
Transduced Pax5-/- cells were seeded at the same concentration and cultured for 3 days in presence of IL7. Although the percentage of Cd19pos cells is the same after transduction with PAX5A or PAX5B (Figure 3C, left panel), the absolute number of pro-B cells transduced by PAX5 isoforms was increased compared to the MIE with the most potent proliferation being driven by PAX5B (Figure 3C, right panel). Furthermore, after 10 days of culture in presence of IL7, Pax5-/- pro-B cells complemented with PAX5B have a quarter less in spontaneous cell death rate compared with PAX5A (Figure 3D, left panel). This difference of cell survival may explain the highest percentage of IgMposIgΚpos B cells after IL7 withdrawal with PAX5B complementation (Figure 3B).
We also stably transduced Ba/F3 cells with either MIE, PAX5A or PAX5B and stressed the cells by serum deprivation for 6 hours. Ba/F3 cells are less sensitive to serum deprivation when they express PAX5B compared to PAX5A whereas no difference was observed in 10% serum condition (Figure 3D, right panel). This difference is not linked to a difference of PAX5 isoform stability since half-life of the two proteins are comparable (Figure 3E). Consequently, the alternative use of PAX5 exon 1 has no qualitative effect on early B cell differentiation but may have an impact on cell death.
Both Pax5 isoforms induce a B cell specific program
After sorting transduced Pax5-/- pro-B cells, their respective transcriptomes were analyzed (Figure 4A–4C). Both PAX5A and PAX5B enhance the expression of B cell specific genes such as Cd19 and repress the expression of non-B cell specific markers such as Cd7 (a T cell marker) or Gp49a (a myeloid marker) (Figure 4D). Furthermore, out of the 24,184 genes, no expression differs more than 5-fold between PAX5A and PAX5B target genes (Figure 4C). By performing a Gene Ontology (GO) analysis [18, 19] on the genes whose expression is the most discriminant, we found a significant enrichment in genes implicated in the regulation of apoptosis (Supplementary Figure 1A, 1B). These data strongly suggest that the two PAX5 isoforms globally induce B cell differentiation by activating the same targets during early B cell differentiation and differences in their target gene expression may explain the differences observed in cell death rate and cell growth (Figure 3C and 3D).
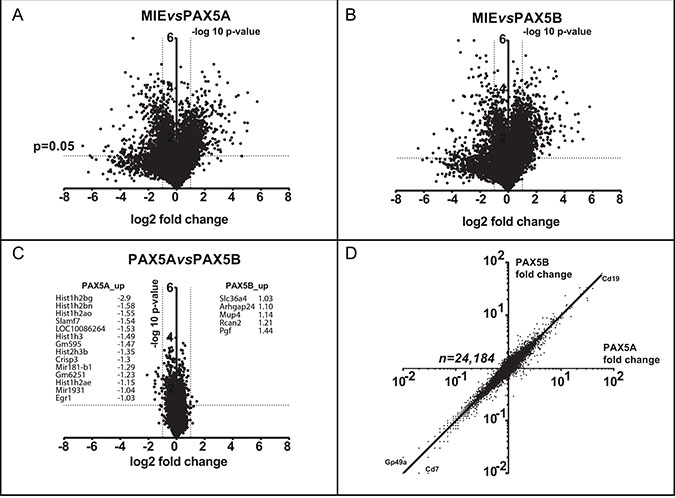
Figure 4: PAX5A and PAX5B drive a comparable transcriptomic early B cell program. (A–C) Volcano plots presenting the fold change in log2 on x axis and the log10 p-value of the false discovery rate permutation analysis on y axis of the transcriptome comparing eGFP expressing vector (MIE) to PAX5A (A), MIE to PAX5B (B) and PAX5A to PAX5B (C) showing the similar effect of PAX5A and PAX5B on early B cell differentiation restoration. For C, gene with a difference of expression >2 fold between PAX5A and PAX5B are listed. (D) Scatter graph with gene expression fold changes between PAX5A and MIE on x-axis and PAX5B and MIE on y-axis. The regression line shows the relationship between PAX5A and PAX5B impact on target gene expression. As example, Cd19 up-regulated gene and Gp49a and Cd7 down-regulated genes are indicated.
Pax5 isoforms have a distinct pattern of expression during late B cell differentiation
We also investigated the functional specificities of PAX5 isoforms during late B cell differentiation. We quantified Pax5 isoforms in murine sorted splenic resting B cells (Cd43neg B220pos), in vitro LPS-activated B cells (B220pos Cd138neg) and plasmablasts (B220pos Cd138pos). Overall, Pax5 expression is decreased after B cell activation (Figure 5A). In details, Pax5A was the most expressed isoform and downregulated in activated B cells and plasmablasts. In contrast, Pax5B was significantly upregulated in activated B cells and almost undetectable in plasmablasts. Therefore, the expression patterns of the two isoforms are distinct during late B cell differentiation.
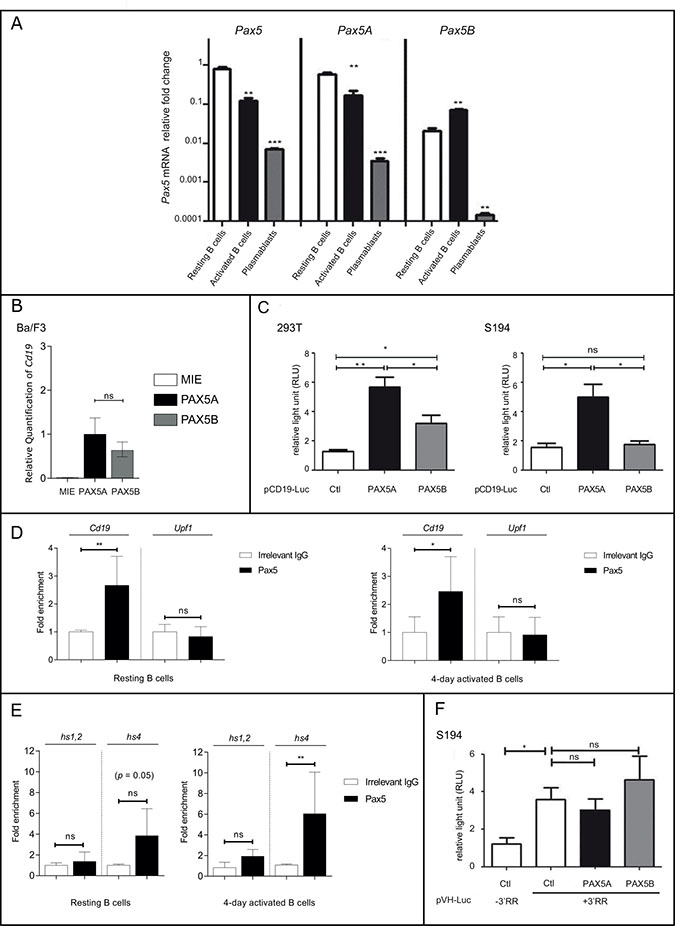
Figure 5: Pax5A and Pax5B show different activities during late B cell stages. (A) Expression by QPCR of total Pax5 or specific Pax5 isoforms in resting murine B cells, activated B cells and plasmablasts. Results are expressed as log scale with mean ± SEM of independent experiments. (B) Relative quantification of Cd19 transactivation in Ba/F3 after infection with either eGFP-expressing retrovirus (MIE) or MIE containing PAX5A or PAX5B (C) PAX5A or PAX5B trans-activity was determined using a Luciferase reporter (pCD19-Luc) assay in 293T cells (left panel) or S194 cells (right panel). Results are expressed as mean ± SEM of independent experiments. (D) ChIP was performed on Cd19 promoter (pCD19) and Upf1 intron 9 (as a PAX5 non-target) and using an irrelevant IgG or an anti-PAX5 antibody. Results are expressed as mean of fold change compared to IgG condition ± SEM of independent experiments. (E) ChIP was performed on the hypersensitive sites (hs)1,2 and hs4 elements of the 3ʹRR in resting splenic B cells or activated B cells (left and right panel respectively) using an irrelevant IgG or a anti-PAX5 antibody. Results are expressed as mean ± SEM of independent experiments. (F) A Luciferase expression B cell specific vector (pVH-luc) containing the IGH 3ʹRR minilocus (+3ʹRR) or not (–3ʹRR) was used in a reporter assay to determine the regulatory function of PAX5A and PAX5B in S194 cells. Results are expressed as mean of fold change compared to IgG condition ± SEM of independent experiments. Ctl: negative control, *p < 0.05; **p < 0.01; ***p < 0,001 (unpaired two-tailed t test).
Pax5B is inactive in plasma cells
The differential regulation of Pax5 isoforms at late B cell differentiation stages may reflect different properties of these isoforms. To assess this possibility, we evaluated the transactivation of Cd19 at an early B cell stage using transduced Ba/F3; both PAX5A and PAX5B can efficiently transactivate Cd19 (Figure 5B). We also evaluated the transactivation potential of PAX5A and PAX5B at later stage of B cell differentiation using S194, a plasmacytoma cell line, or in a non-lymphoid context using 293T, a human kidney cell line, using a luciferase reporter system containing the Cd19 promoter region. Compared to AID, a negative control, both PAX5A and PAX5B activate the Cd19 promoter (Figure 5C). The activity of PAX5B is significantly lower than PAX5A. In S194, only PAX5A displays transcriptional activity suggesting that PAX5B transactivation capacity on Cd19 promoter is altered in those cells (Figure 5C). To characterize the binding of Pax5 on the Cd19 promoter at late stages of the B cell differentiation, we precipitated chromatin (ChIP) using an anti-Pax5 antibody in splenic B cells (Figure 5D). We detected a specific binding of Pax5 to the Cd19 promoter sequence in resting B cells compared to the amplification of an irrelevant sequence (Upf1 intronic sequence) devoid of Pax5 binding sites (Figure 5D, left panel). After 4 days of LPS stimulation, we could detect the same enrichment of Pax5 on Cd19 promoter (Figure 5D, right panel).
Neither PAX5A nor PAX5B are able to silence the 3ʹRR in plasma cells
The 3ʹRR is an important region regulating the expression of the heavy immunoglobulin chain at late B cell differentiation stages. It has been reported that Pax5 could bind hypersensitive sites (hs)1,2 (3ʹαE) and hs4 elements ([20, 21]). A Pax5-specific ChIP using an antibody that does not discriminate between the two isoforms, identified a 3.8-fold enrichment at the hs4 element site in resting B cells (Figure 5E, left panel). Upon activation, Pax5 is enriched both at the hs4 and to a much lesser extend to hs1,2 sites (6-fold enrichment for hs4 and a 2.4-fold for hs1,2; Figure 5E, right panel).
We then analyzed whether both PAX5 isoforms have a similar transactivation activity on the 3ʹRR in S194 plasma cells (Figure 5F). PAX5 isoforms are physiologically downregulated in plasma cells, in contrast the 3ʹRR activity is increased. We evaluated the effect of the PAX5 isoforms on the 3ʹRR activity using a reporter assay depending on the VH promoter (pVH) active in the B cell lineage. Its transcription level is upregulated by the addition of a synthetic 3ʹRR containing the four enhancers in their palindromic structure (hs3a, hs1.2, hs3b, hs4). We did not observe any significant change upon the addition of both PAX5 isoforms, suggesting that 3ʹRR is insensitive to Pax5 (Figure 5F) despite the accumulation of Pax5 at the 3ʹRR after activation of B cells (Figure 5E, right panel).
DISCUSSION
PAX5 is a major regulator of B cell differentiation promoting the expression of B cell specific genes and repressing commitment to other cell lineages. Consequently, to allow an efficient B cell differentiation, the expression of PAX5 has to be tightly controlled. An alteration of PAX5 dosage induced B cell malignancies: its overexpression, secondary to the IGH-PAX5 rearrangement, is associated with B cell lymphomas [16] and loss of heterozygosity is associated with B cell progenitor acute lymphoblastic leukemia [22, 23]. The regulation of PAX5 expression, in vertebrates, is complex with two 5ʹ isoforms, PAX5A and PAX5B, initiated from two different promoters. These isoforms differ only by the sequence of their first coding exon encoding no specific protein motif. PAX5B isoform is related to PAX2, since the sequence encoded by their first exon are 65% homologous and is present as early as in the Fugu (Table 1). The PAX5A isoform appears later, with mammals and by extension B cells.
Table 1: Comparative analysis of exon 1A and exon 1B coding sequences during evolution
Species |
PAX5 coding exon 1A |
PAX5 coding exon 1B |
|---|---|---|
Human |
MDLEKNYPTPRTSRT |
MEIHCKHDPFASMH |
Chimpanzee |
MDLEKNYPTPRTSRT |
MEIHCKHDPFASMH |
Mouse |
MDLEKNYPTPRTIRT |
MEIHCKHDPFASMH |
Rat |
MDLEKNYPTPRTIRT |
MEIHCKHDPFASMH |
Rabbit |
MDLEKNYPTPRTGRT |
MEIHCKHDPFASMH |
Dog |
MDLEKNYPTPRSGRT |
MEIHCKHDPFASMH |
Pig |
MDLEKNYPTPRTGRT |
MEIHCKHDPFASMH |
Cow |
MDLEKNYPTPRTGRT |
MEIHCKHDPFASMH |
Xenopus |
No hits found |
MEIHCKHDPFAAMH |
Zebrafish |
No hits found |
MEIHCKHDPFAAMH |
Fugu |
No hits found |
MEIHCKHDPFAAMH |
C. elegans |
No hits found |
No hits found |
Fruitfly |
No hits found |
No hits found |
Lamprey |
No hits found |
No hits found |
The table shows the alignment of PAX5 exon 1A and exon 1B primary sequence in different species. The discrepancies between species are boxed.
Very little is known about the differential regulation of these two isoforms and their functional differences. The aim of this study was to determine if the presence of these two isoforms translate a functional difference or are merely a differential regulation of expression. In B cells, PAX5 isoforms are differentially expressed suggesting that different sets of transcription factors enhance or repress the activity of these promoters. ChIP analyses on cell lines and pro-B cells confirm a correlation of histone H3 acetylation on both promoter regions and expression of the two isoforms. Interestingly, we also detected histone H3 acetylation on the enhancer region [24] only when the two isoforms are expressed.
Functionally, PAX5A and PAX5B can act similarly in early B cell differentiation. Both are able to complement the invalidation of Pax5 in Pax5-/- cells and resume ex vivo the B cell differentiation until the Cd19pos/mIgΚpos stage demonstrating a similar capacity to transactivate or repress target genes during this process. This is shown by the transcriptome since no gene expression differs more than five times in Pax5-/- complemented by PAX5A or PAX5B. Nonetheless, we detected differences between the two isoforms. PAX5B confers a growth advantage of the rescued Pax5-/- cells compared to PAX5A-rescued cells. The physiological up-regulation of Pax5B during B cell activation may prevent apoptosis of activated B cells since suppression of PAX5B increases susceptibility of human acute lymphocytic leukemia REH cells to apoptotic death [25]. Interestingly GO term analysis on discriminant genes for which the expression is at least 1.5 fold higher for PAX5A or PAX5B condition shows that they are involved in cell death pathways.
As described by Revilla-i-Domingo et al., although Pax5 interacts with a similarly high number of genomic binding sites defining 8,000 target genes in pro-B and mature B cells, only 13 target genes are commonly activated and 18 target genes are commonly repressed in pro-B and mature B cells (cited from [26]) suggesting that Pax5A and Pax5B could have different transcriptional activities in early and late B cell differentiation stages.
At later stages of B cell differentiation, we observed that Pax5B and Pax5A are expressed simultaneously in primary B cells at different levels. After B cell activation, they exhibit a distinct pattern of expression: Pax5A decreases in activated B cells and plasmablasts while Pax5B increases in activated B cells and is virtually absent in plasmablasts.
PAX5A is able to transactivate Cd19 promoter in both early (Ba/F3) and late B cells (S194) and in a non-lymphoid 293T whereas PAX5B is able to transactivate Cd19 only at an early stage of B cell differentiation and in a non-lymphoid context. These results point out that PAX5B isoform cannot transactivate the Cd19 promoter in a late lymphoid context (S194) or may function in a different context as transcriptional repressor as proposed by Robichaud et al. The authors have shown that PAX5B negatively regulates CD19 expression in REH cell line since the specific suppression of PAX5B expression using ribozymes leads to an increase of CD19 expression [25].
PAX5 has been described as a negative regulator of the 3ʹRR activity. Nonetheless, we observed an increased binding of PAX5 to the 3ʹRR upon B cell activation. The accumulation of PAX5 binding on the 3ʹRR in activated B cells does not seem to reflect a role of PAX5 in the 3ʹRR activity regulation but rather a function in chromatin remodeling permitting accessibility to the IGH locus. Recently, it has been shown that PAX5 recruits PTIP, a molecule involved in class switch recombination [27, 28] which induces H3K4me3 and stabilizes the PAX5 binding on the IGH locus [28]. Alternatively, this could be explained by the fact that the activation of B cells results in the induction or increase of the expression of other factors counteracting PAX5 activity.
PAX5 isoforms are physiologically downregulated in plasma cells in contrast to the 3ʹRR activity which is strongly increased. We investigated whether these isoforms could modify the 3ʹRR activity. Our data showed that 3ʹRR is insensitive to a PAX5A or PAX5B regulation in the S194 plasmacytoma cell line, in agreement to the observation that an hs1,2-dependent transgene is not inhibited whereas Pax5 expression is still detected in activated B cells [29]. Consequently, repression of the 3ʹRR activity by Pax5 [20, 30, 31] is likely more complex.
Our results reveal only discrete differences of action between PAX5A and PAX5B functions and demonstrate that these two isoforms may be interchangeable in early B cell differentiation. Their very tight regulation may reflect different binding of alternative transcription factors according to the cellular context (testis, brain, B cells) leading to a fine control of the Pax5 dosage which is required for a zero-default B cell differentiation.
MATERIALS AND METHODS
Cell line culture
In this study, we used several B cell lines showing characteristics of various stages of B cell differentiation: Ba/F3 are pro-B cells (DSMZ ACC 300) [32], 38B9 are pro-B cells [33, 34], 70Z/3 are pre-B lymphoblast (ATCC® TIB-158™) [35], A20 is a BALB/c B cell lymphoma line derived from a spontaneous reticulum cell neoplasm found in an old BALB/cAnN mouse (ATCC® TIB-208™) [36], 18.81 is a Abelson-virus-transformed mouse lymphoid cell line with pre-B cell characteristic [37], WEHI-231 are immature B cells (ATCC® CRL-1702™) [38], S194 are mouse myeloma B cells, this line was derived from an IgA secreting mineral oil induced BALB/c myeloma. (ATCC® TIB-19™) [39]. Murine B cell lines S194, 38B9, A20, 18.81, and WEHI-231 were cultured in RPMI (Gibco), 10% FBS (fetal bovine serum, PAN Biotech), 2 mM L-glutamine, and 1 mM sodium pyruvate (Invitrogen). The same medium supplemented with 0.05 mM β-mercaptoethanol or 10 ng/ml recombinant murine IL3 (Peprotech) was used respectively to culture the murine cell line 70Z3 or Ba/F3. The Phoenix retrovirus producer line (Orbigen) was cultured in high glucose Dulbecco’s Modified Eagle’s Medium (DMEM; Gibco), 10% FBS (PAN Biotech). Murine OP9 bone marrow stromal cells were cultured in MEMα (Gibco), 20% FBS (PAN Biotech) when in monoculture. All the cells were cultured in presence of penicillin (100U/ml) and streptomycin (100U/ml) at 37° C, in 5% CO2 atmosphere.
Mice
Pax5± mice were kindly provided by Pr M. Busslinger [6]. Pax5-/- and Pax5+/+ (wild-type) mice were obtained by breeding two Pax5± mice. All mice were maintained on the C57BL/6 genetic background. The Pax5 genotype was determined using REDExtract-N-Amp Tissue PCR Kit according to the manufacturer’s instructions (Sigma-Aldrich) with the following oligonucleotides: 5ʹ-AGG ACA TGG AGG AGT GAA TCA G-3ʹ (primer1); 5ʹ-ACC CGA AGC TGC CTG GAG-3ʹ (primer2); and 5ʹ-AGG CGA TTA AGT TGG GTA ACG-3ʹ (primer 3). All animal experiments were carried out according to valid project licenses, which were approved and regularly controlled by French Authorities.
Purification and culture of B cell subsets
Pax5-deficient pro-B cells were harvested from Pax5-/- embryonic liver at E17.5. Cells were amplified on γ-irradiated (30 Gy) OP9-derived stromal cells in Iscove’s Modified Dulbecco’s Medium (IMDM; Gibco), supplemented with 5% FBS (Stemcell technologies), 100 U/ml murine IL7 (Peprotech), 0.05 mM β-mercaptoethanol (Sigma-Aldrich), 2 mM L-glutamine (Invitrogen), penicillin (100 U/ml) and streptomycin (100 U/ml). Wild-type pro-B cells were flushed from bone marrow. After red blood cell lysis, B cell fraction was enriched by B220 magnetics microbead sorting (Myltenyi Biotech). B cell fraction was subsequently incubated with B220, Kit (Cd117) and Igk antibodies and pro-B cells (B220pos kitpos and Igκneg) were sorted using a FACS-Vantage sorter (BD Biosciences). Splenic resting B cells were purified by negative selection using anti-CD43 microbeads (Miltenyi Biotech) and stimulated (0.5 × 106 cells/ml) by LPS (20 μg/ml; Sigma) for 2 to 4 days in RPMI supplemented with 10% FBS, sodium pyruvate, non-essential amino acids, β-mercaptoethanol, penicillin (100 U/ml) and streptomycin (100 U/ml) (Invitrogen).
B cell subsets were sorted on a MoFlo Astrios sorter (Beckman Coulter) after bone marrow flush of C57Bl/6 8 to 12 week-old mice as follows: pre-pro-B (B220+Cd19-), pro-B (B220+Cd19+IgL-IgD-Cd117+BP1-), pre-BI (B220+Cd19+IgL-IgD-Cd117+BP1+), large pre-BII (B220+Cd19+IgL-IgD-Cd117+, large FSC), small pre-BII (B220+Cd19+IgL-IgD-Cd117+, small FSC), immature B (B220+Cd19+IgL+IgD-), recirculating B (B220+Cd19+IgL+IgD+) and RNA extraction were performed on fresh samples.
For LPS stimulation B cells (B220pos Cd138neg) and plasmablasts (B220pos Cd138pos) were sorted 3 to 4 days after stimulation.
Quantitative RT-PCR
RNA was isolated using the AllPrep™ DNA/RNA Micro Kit (QIAGEN) and cDNA was synthesized using SuperScript® VILO™ cDNA Synthesis Kit (Invitrogen™) according to the manufacturer’s instructions. Quantitative PCR were performed on a LightCycler®480 II System (Roche). Quantitative SYBR Green PCR was performed to quantify relative mouse Cd19 cDNA expression in transduced Ba/F3 using LightCycler®480 SYBR Green I Master (Roche Diagnostics GmbH) according to the manufacturer’s instructions, and primers 10 μM primers for Cd19 (forward: 5ʹ-AAGGAACAGGTCCTCTGGGA-3ʹ and reverse: 5ʹ-GATTCAAACTGCTCCCCCGA-3ʹ) and 10 μM primers for Abl1 (forward: 5ʹ-GGAGAA GGTCTACGAGCTCAT-3ʹ and reverse: 5ʹ-ATCTGAGATA CTGGATTCCTGGA-3ʹ). Relative gene expression in cell lines or sorted bone marrow B cells was quantified using LightCycler®480 Probes Master (Roche Diagnostics GmbH) and the following TaqMan gene expression assays from Applied Biosystems™ Life Technologies: Mm00802038_m1 for Abl1, Mm00487680_m1 for Melk, Mm00435501_m1 for Pax5 (both isoforms), Mm00621713_m1 for Zcchc7, Mm00395519_m1 for Ebf1, Mm00435494 for specific expression of Pax5A. Primers/TaqMan probe mix specific for PAX5B transcripts was synthetized by Applied Biosystems (5ʹFAM-ATG CAT AGA CAT GGA GGA GTG AAT CAG CTT G-Tamra3ʹ as probe; 5ʹ-CAC TGT AAG CAC GAC CCG TT-3ʹ as forward primer; 5ʹ-AGT GGC CGT CCA TTC ACA AA-3ʹ as reverse primer). All QPCR programs were carried out as follows in a 20 μl volume: 5 min at 95° C, followed by 45 cycles of 10 s at 95° C, 10 s at annealing temperature of 60° C and 10 s at 72° C. All signals were quantified using either the ΔCt method or the Relative Quantification (ΔΔCt) study both including a normalization to the ΔCt values of Abl1 gene expression levels. Data were analyzed using the LC480 software (Roche Diagnostics) followed by statistical analyses.
ChIP experiment
CHromatin ImmunoPrecipitation (ChIP) assays were performed essentially as described previously [40] using anti-acetylated histone H3 or nonspecific rabbit immunoglobulin G (IgG) (06-599 and 12-370, respectively; Upstate Biotechnology, Lake Placid, NY). Oligonucleotide sequences are available on Supplementary Table 1. Murine splenic B cells were cultured (106 cells/ml) in presence of LPS (40 μg/ml) for 4 days. At day 3, dead cells were eliminated by Ficoll centrifugation (Lympholyte-Mammal, Cedarlane) and living cells were stimulated for an additional 24 h period. Chromatin preparations were performed on 5 × 107 to 108 LPS-stimulated B cells using standard fixation and sonication methods as described [41]. In all cases, 200 ng of chromatin was immunoprecipitated (ON, 4° C) with 1μg of anti-PAX5 mAb (mouse anti-Pax-5 mAb (A-11), Santa Cruz Biotechnology) or an irrelevant IgG antibody (purified mouse unlabeled IgG1, SouthernBiotech, ref.0102-01). Oligonucleotides used are given in Supplementary Table 1.
Retroviral particles production
Full-length human PAX5A and PAX5B cDNA were amplified, sequenced and inserted into the retroviral vector pMSCV-IRES-eGFP (MIE), a bicistronic vector allowing the expression of eGFP under the control of the long terminal repeat (LTR) promoter. Retroviral supernatants were produced by Phoenix retrovirus producer line (Orbigen) using Lipofectamine 2000 (Invitrogen) reagent for transfection and proceeding according to the manufacturer’s instructions. After overnight incubation of the transfection mix, cells were treated with 10 mM sodium butyrate (Millipore) and washed with PBS before pro-B cell medium was added. Viral supernatants were harvested after 24 h of incubation at 32° C, passed through a 0.45 μm filter, aliquoted and frozen at –80° C.
Pro-B culture, infection and differentiation
The in vitro pro-B cell differentiation/proliferation protocol used was adapted from Rolink et al. [42]. The pro-B Pax5-/- cells were isolated from Pax5-/- embryo fetal liver and cocultivated with irradiated OP9 cells in 24-well plates in Iscove’s Modified Dulbecco’s Medium (IMDM; Gibco), supplemented with 5% FBS (Stemcell Technologies), 2 ng/ml murine IL7 (Peprotech), 0.05 mM β-mercaptoethanol (Sigma-Aldrich), 2 mM L-glutamine (Invitrogen), penicillin (100 U/ml) and streptomycin (100 U/ml). Experiments were performed after an initial pro-B cell expansion on irradiated OP9 cells in the presence of 2 ng/ml IL7. Every 3 days, pro-B cells were harvested and propagated on fresh OP9 stromal cells.
The pro-B cells, cultured on irradiated OP9 cells in the presence of 2 ng/ml of IL7, are infected during exponential growth. Retroviral supernatant was diluted 1:3, supplemented with 4 μg/ml polybrene (Sigma) and 2 ng/ml of IL7 (Peprotech); spinoculation was performed at 1000 × g at 32° C for 90 min. After centrifugation, 1ml of fresh medium supplemented with IL7 was added and the cells were incubated overnight at 37° C. After infected pro-B cell expansion for few days, cells were washed out of IL-7 and plated in IMDM, 5% FBS, 0.05 mM β-mercaptoethanol and 2 mM L-glutamine on irradiated OP9-derived stromal cells with or without IL7. At day 2.5, single-cell suspensions were stained using standard protocols for flow cytometry using the antibodies listed above. Dead cells were excluded from FACS analysis as 7-amino-actinomycin D (7-AAD; BD Biosciences #559763)) positive (BD Biosciences). Stained cells were analyzed with a FACS Calibur flow cytometer (BD). Data were analyzed with the BD FACSDiva™ software (BD Biosciences) or FlowJo (TreeStar, Ashland, OR). All antibodies are rat monoclonal antibodies from BD biosciences: anti-B220 (clone RA3-6B2) conjugated with pacific blue, anti-CD19 (clone 1D3) conjugated with Alexa Fluor 700, anti-mouse Igk light chain (clone 187.1) conjugated with Phycoerythrin-Cyanin7, anti-IgM (clone R6-60.2) conjugated with Peridinin chlorophyll protein-cyanin5.5.
Ba/F3 cells were similarly transduced with empty MIE, MIE-PAX5A or MIE-PAX5B and then GFP-positive cells were isolated by fluorescence-activated cell sorting (FACS) in MoFlo Astrios (Beckman Coulter).
Cycloheximide treatment and Western blot
Stability assay was performed in the presence of cycloheximide (100 μg/mL, Sigma) for the indicated periods of time. The day before, 1.2 × 106 transduced Ba/F3 cells were incubated overnight at 37° C, 5% CO2. At each time point, treated cells were washed once with ice-cold Phosphate buffer saline (PBS) and then cell pellets were lysed in 40 μl of lysis buffer composed of 150 mM NaCl, 1 mM EDTA, 50 mM Tris, 1% Triton and protease inhibitor cocktail (cOmplete™, Roche). Protein concentrations of cell lysates were determined using the BCA1-Kit for Protein Determination (Sigma-Aldrich) to load 30 μg in every well. Samples were subjected to 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred to PVDF membranes. Blots were blocked 1 hour in the presence of 5% milk and incubated with 0.3 μg/mL of primary antibody overnight at +4° C (mouse anti-Pax-5 mAb (A-11), Santa Cruz Biotechnology). After incubation with secondary antibodies conjugated to horseradish peroxidase (Cell Signaling Technology), the immunoreactive bands were visualized by using the enhanced chemiluminescence lighting system (Amersham™ ECL™ Prime Western Blotting Detection Reagent, GE Healthcare) and data was acquired with Chemi-Smart 5000 and Bio-1D software (Vilber Lourmat). Bands were quantified by using ImageJ software.
Flow cytometry and cell growth
Transduced pro-B cells and Ba/F3 cells were stained with AlexaFluor700-conjugated anti-CD19 (1D3) antibody (BD Pharmingen). For cell-viability assays, cells were stained with Annexin-V-PE and 7-AAD (BD Biosciences #559763). Flow cytometric data were acquired on a Fortessa X20 or LSR II flow cytometer (BD Biosciences), and data were analyzed using FACSDiva (BD Biosciences) and FlowJo software (TreeStar).
Transcriptomic analysis
After 5 days in culture with 2 ng/ml of IL7, infected cells were sorted according to their positivity for GFP fluorescence with a FACSAria II (BD Biosciences) cell sorter. Total RNA from GFPpos sorted pro-B infected cells were extracted using the Trizol method according to manufacturer’s instruction (Invitrogen). RNA quality was assessed using the RNA 6000 Nano Assay on the Agilent 2400 Bioanalyser (Agilent Technologies, Massy, FRANCE). For each condition (PAX5A, PAX5B and MIE), three independent infection experiments were performed. RNA samples were purified and prepared according to the manufacturer’s protocol with the Affymetrix’s GeneChip Whole Transcript Sense Target Labeling Assay Kit (Affymetrix, UK) and hybridized on Affymetrix GeneChip Mouse Gene 2.0 ST arrays. Probe-signal intensities were normalized and summarized by the Robust Multiarray Average (RMA) method using Affymetrix’s Expression Console software.
The data discussed in this publication have been deposited in NCBI’s Gene Expression Omnibus [43] and are accessible through GEO Series accession number GSE104890 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE104890).
For the volcano plots, permutation analyses were performed with dChip software [44] allowing the calculation of false discovery rate p-value and the fold difference volcano plots were generated using GraphPad prism software.
The gene ontology analysis has been performed with the AutoCompare ZE software [45] using Gene Ontology database [18, 19], p-values have been calculated with the ZE test.
Luciferase reporter assay
Experiments were performed by cell transfections with 2μg DNA at 1:7:1 (pcDNA3-PAX5 vector: reporter vector: GFP-CMV) vector ratios. GFP-CMV was used to evaluate transfection efficiency. The S194 line was cultured in RPMI as above and for each condition 2 × 106 cells were electroporated according to the manufacturer’s instructions (Lonza). The 293T line was cultured in DMEM (with 10% FBS, sodium pyruvate, nonessential amino acids, penicillin and streptomycin) and subconfluent cells in 6-well dishes were transfected by lipofection with Exgen500 (Euromedex). Firefly luciferase-CD19-2(A ins) [46] was kindly provided by Pr M. Busslinger was used to measure PAX5 activity on the PAX5 binding site of the Cd19 promoter. PAX5 activity on the 3ʹRR was evaluated with firefly luciferase-pVH enhanced or not by the combination of core enhancers of 3ʹRR [47]. After 48 h transfection, cells were collected and split for GFP fluorescence analysis by flow cytometry and luminescence detection using the Luciferase Reporter Assay System and the 20/20n Luminometer (Promega). For each sample, luminescence was normalized to the measured GFP fluorescence.
Statistical analysis
Experiments have been performed at least as three independent conditions. All statistical analyses were performed using Excel (Microsoft) and GraphPad Prism software, version 7 (GraphPad Software). Statistical differences were determined using a 2-tailed Student’s t test or RM one-way ANOVA with Tukey’s correction or one-way ANOVA with Sidak’s correction for multiple comparisons or Pearson correlation, as mentioned in legends. All data are presented as mean ± SD or SEM. A p-value of less than 0.05 was considered statistically significant, the null hypothesis being rejected at the 0.05 level (***p < 0.0005, **p < 0.005, *p < 0.05).
ACKNOWLEDGMENTS
The authors thank “the Pôle Technologique of CRCT – Plateaux Cytométrie et Bioinformatique (INSERM U1037), Toulouse” for technical assistance. We also thank the Anexplo-Génotoul platform CREFRE INSERM - UMS 006 animal facility.
CONFLICTS OF INTEREST
The authors declare no financial or commercial conflict of interest.
GRANT SUPPORT
This work has been supported by INCa R11190BB-INCA 2011-131-PAX5, by ARC grant ARC SFI20101201888 and the FEDER grant CITTIL. CC, SP and NR have been supported by INCa grant R11190BB-INCA 2011-131-PAX5. The team is supported by association Laurette FUGAIN, la société française des cancers de l’enfant (SFCE), The association Capucine, “111 des Arts”, la Ligue régionale contre le cancer and the “région Occitanie”.
MC, ED, LD and CB designed the experiments. CC, SP, LJ, NR, NP, MD, BG, SH and SL performed the experiments. SM designed the FACS analysis strategy and performed the preliminary experiments. LD, SP, ED and CB wrote the manuscript.
REFERENCES
1. Nutt SL, Heavey B, Rolink AG, Busslinger M. Commitment to the B-lymphoid lineage depends on the transcription factor Pax5. Nature. 1999; 401:556–62. https://doi.org/10.1038/44076.
2. Lang D, Powell SK, Plummer RS, Young KP, Ruggeri BA. PAX genes: roles in development, pathophysiology, and cancer. Biochem Pharmacol. 2007; 73:1–14. https://doi.org/10.1016/j.bcp.2006.06.024.
3. Stapleton P, Weith A, Urbanek P, Kozmik Z, Busslinger M. Chromosomal localization of seven PAX genes and cloning of a novel family member, PAX-9. Nat Genet. 1993; 3:292–8. https://doi.org/10.1038/ng0493-292.
4. Adams B, Dorfler P, Aguzzi A, Kozmik Z, Urbanek P, Maurer-Fogy I, Busslinger M. Pax-5 encodes the transcription factor BSAP and is expressed in B lymphocytes, the developing CNS, and adult testis. Genes Dev. 1992; 6:1589–607.
5. Nutt SL, Thevenin C, Busslinger M. Essential functions of Pax-5 (BSAP) in pro-B cell development. Immunobiology. 1997; 198:227–35. https://doi.org/10.1016/S0171-2985(97)80043-5.
6. Urbanek P, Wang ZQ, Fetka I, Wagner EF, Busslinger M. Complete block of early B cell differentiation and altered patterning of the posterior midbrain in mice lacking Pax5/BSAP. Cell. 1994; 79:901–12.
7. Delogu A, Schebesta A, Sun Q, Aschenbrenner K, Perlot T, Busslinger M. Gene repression by Pax5 in B cells is essential for blood cell homeostasis and is reversed in plasma cells. Immunity. 2006; 24:269–81. https://doi.org/10.1016/j.immuni.2006.01.012.
8. Nutt SL, Urbanek P, Rolink A, Busslinger M. Essential functions of Pax5 (BSAP) in pro-B cell development: difference between fetal and adult B lymphopoiesis and reduced V-to-DJ recombination at the IgH locus. Genes Dev. 1997; 11:476–91.
9. Dunnick WA, Collins JT, Shi J, Westfield G, Fontaine C, Hakimpour P, Papavasiliou FN. Switch recombination and somatic hypermutation are controlled by the heavy chain 3ʹ enhancer region. J Exp Med. 2009; 206:2613–23. https://doi.org/10.1084/jem.20091280.
10. Rouaud P, Vincent-Fabert C, Saintamand A, Fiancette R, Marquet M, Robert I, Reina-San-Martin B, Pinaud E, Cogne M, Denizot Y. The IgH 3ʹ regulatory region controls somatic hypermutation in germinal center B cells. J Exp Med. 2013; 210:1501–7. https://doi.org/10.1084/jem.20130072.
11. Saintamand A, Rouaud P, Saad F, Rios G, Cogne M, Denizot Y. Elucidation of IgH 3ʹ region regulatory role during class switch recombination via germline deletion. Nat Commun. 2015; 6: 7084. https://doi.org/10.1038/ncomms8084.
12. Vincent-Fabert C, Fiancette R, Pinaud E, Truffinet V, Cogne N, Cogne M, Denizot Y. Genomic deletion of the whole IgH 3ʹ regulatory region (hs3a, hs1,2, hs3b, and hs4) dramatically affects class switch recombination and Ig secretion to all isotypes. Blood. 2010; 116:1895–8. https://doi.org/10.1182/blood-2010-01-264689.
13. Schaniel C, Bruno L, Melchers F, Rolink AG. Multiple hematopoietic cell lineages develop in vivo from transplanted Pax5-deficient pre-B I-cell clones. Blood. 2002; 99:472–8.
14. Cobaleda C, Jochum W, Busslinger M. Conversion of mature B cells into T cells by dedifferentiation to uncommitted progenitors. Nature. 2007; 449:473–7. https://doi.org/10.1038/nature06159.
15. Palmisano WA, Crume KP, Grimes MJ, Winters SA, Toyota M, Esteller M, Joste N, Baylin SB, Belinsky SA. Aberrant promoter methylation of the transcription factor genes PAX5 alpha and beta in human cancers. Cancer Res. 2003; 63:4620–5.
16. Busslinger M, Klix N, Pfeffer P, Graninger PG, Kozmik Z. Deregulation of PAX-5 by translocation of the Emu enhancer of the IgH locus adjacent to two alternative PAX-5 promoters in a diffuse large-cell lymphoma. Proc Natl Acad Sci U S A. 1996; 93:6129–34.
17. Delabesse E, Ogilvy S, Chapman MA, Piltz SG, Gottgens B, Green AR. Transcriptional regulation of the SCL locus: identification of an enhancer that targets the primitive erythroid lineage in vivo. Mol Cell Biol. 2005; 25:5215–25. https://doi.org/10.1128/MCB.25.12.5215-5225.2005.
18. Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000; 25:25–9. https://doi.org/10.1038/75556.
19. The Gene Ontology Consortium. Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 2017; 45: D331–D8. https://doi.org/10.1093/nar/gkw1108.
20. Michaelson JS, Singh M, Snapper CM, Sha WC, Baltimore D, Birshtein BK. Regulation of 3ʹ IgH enhancers by a common set of factors, including kappa B-binding proteins. J Immunol. 1996; 156:2828–39.
21. Chatterjee S, Ju Z, Hassan R, Volpi SA, Emelyanov AV, Birshtein BK. Dynamic changes in binding of immunoglobulin heavy chain 3ʹ regulatory region to protein factors during class switching. J Biol Chem. 2011; 286:29303–12. https://doi.org/10.1074/jbc.M111.243543.
22. Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD, Girtman K, Mathew S, Ma J, Pounds SB, Su X, Pui CH, Relling MV, et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature. 2007; 446:758–64. https://doi.org/10.1038/nature05690.
23. Familiades J, Bousquet M, Lafage-Pochitaloff M, Bene MC, Beldjord K, De Vos J, Dastugue N, Coyaud E, Struski S, Quelen C, Prade-Houdellier N, Dobbelstein S, Cayuela JM, et al. PAX5 mutations occur frequently in adult B-cell progenitor acute lymphoblastic leukemia and PAX5 haploinsufficiency is associated with BCR-ABL1 and TCF3-PBX1 fusion genes: a GRAALL study. Leukemia. 2009; 23:1989–98. https://doi.org/10.1038/leu.2009.135.
24. Decker T, Pasca di Magliano M, McManus S, Sun Q, Bonifer C, Tagoh H, Busslinger M. Stepwise activation of enhancer and promoter regions of the B cell commitment gene Pax5 in early lymphopoiesis. Immunity. 2009; 30:508–20. https://doi.org/10.1016/j.immuni.2009.01.012.
25. Robichaud GA, Perreault JP, Ouellette RJ. Development of an isoform-specific gene suppression system: the study of the human Pax-5B transcriptional element. Nucleic Acids Res. 2008; 36:4609–20. https://doi.org/10.1093/nar/gkn432.
26. Revilla IDR, Bilic I, Vilagos B, Tagoh H, Ebert A, Tamir IM, Smeenk L, Trupke J, Sommer A, Jaritz M, Busslinger M. The B-cell identity factor Pax5 regulates distinct transcriptional programmes in early and late B lymphopoiesis. EMBO J. 2012; 31:3130–46. https://doi.org/10.1038/emboj.2012.155.
27. Daniel JA, Santos MA, Wang Z, Zang C, Schwab KR, Jankovic M, Filsuf D, Chen HT, Gazumyan A, Yamane A, Cho YW, Sun HW, Ge K, et al. PTIP promotes chromatin changes critical for immunoglobulin class switch recombination. Science. 2010; 329:917–23. https://doi.org/10.1126/science.1187942.
28. Schwab KR, Patel SR, Dressler GR. Role of PTIP in class switch recombination and long-range chromatin interactions at the immunoglobulin heavy chain locus. Mol Cell Biol. 2011; 31:1503–11. https://doi.org/10.1128/MCB.00990-10.
29. Andersson T, Neurath MF, Grant PA, Pettersson S. Physiological activation of the IgH 3ʹ enhancer in B lineage cells is not blocked by Pax-5. Eur J Immunol. 1996; 26:2499–507. https://doi.org/10.1002/eji.1830261034.
30. Neurath MF, Strober W, Wakatsuki Y. The murine Ig 3ʹ alpha enhancer is a target site with repressor function for the B cell lineage-specific transcription factor BSAP (NF-HB, S alpha-BP). J Immunol. 1994; 153:730–42.
31. Singh M, Birshtein BK. NF-HB (BSAP) is a repressor of the murine immunoglobulin heavy-chain 3ʹ alpha enhancer at early stages of B-cell differentiation. Mol Cell Biol. 1993; 13:3611–22.
32. Palacios R, Henson G, Steinmetz M, McKearn JP. Interleukin-3 supports growth of mouse pre-B-cell clones in vitro. Nature. 1984; 309:126–31.
33. Alt F, Rosenberg N, Lewis S, Thomas E, Baltimore D. Organization and reorganization of immunoglobulin genes in A-MULV-transformed cells: rearrangement of heavy but not light chain genes. Cell. 1981; 27:381–90.
34. Alt FW, Yancopoulos GD, Blackwell TK, Wood C, Thomas E, Boss M, Coffman R, Rosenberg N, Tonegawa S, Baltimore D. Ordered rearrangement of immunoglobulin heavy chain variable region segments. EMBO J. 1984; 3:1209–19.
35. Paige CJ, Kincade PW, Ralph P. Murine B cell leukemia line with inducible surface immunoglobulin expression. J Immunol. 1978; 121:641–7.
36. Kim KJ, Kanellopoulos-Langevin C, Merwin RM, Sachs DH, Asofsky R. Establishment and characterization of BALB/c lymphoma lines with B cell properties. J Immunol. 1979; 122:549–54.
37. Rosenberg N, Baltimore D. The effect of helper virus on Abelson virus-induced transformation of lymphoid cells. J Exp Med. 1978; 147:1126–41.
38. Boyd AW, Goding JW, Schrader JW. The regulation of growth and differentiation of a murine B cell lymphoma. I. Lipopolysaccharide-induced differentiation. J Immunol. 1981; 126:2461–5.
39. Horibata K, Harris AW. Mouse myelomas and lymphomas in culture. Exp Cell Res. 1970; 60:61–77.
40. Forsberg EC, Downs KM, Christensen HM, Im H, Nuzzi PA, Bresnick EH. Developmentally dynamic histone acetylation pattern of a tissue-specific chromatin domain. Proc Natl Acad Sci U S A. 2000; 97:14494–9. https://doi.org/10.1073/pnas.97.26.14494.
41. Iacovoni JS, Caron P, Lassadi I, Nicolas E, Massip L, Trouche D, Legube G. High-resolution profiling of gammaH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 2010; 29:1446–57. https://doi.org/10.1038/emboj.2010.38.
42. Rolink AG. B-cell development and pre-B-1 cell plasticity in vitro. Methods Mol Biol. 2004; 271:271–81. https://doi.org/10.1385/1-59259-796-3:271.
43. Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002; 30:207–10.
44. Li C, Wong WH. Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc Natl Acad Sci U S A. 2001; 98:31–6. https://doi.org/10.1073/pnas.011404098.
45. Ycart B, Pont F, Fournie JJ. Curbing false discovery rates in interpretation of genome-wide expression profiles. J Biomed Inform. 2014; 47:58–61. https://doi.org/10.1016/j.jbi.2013.09.004.
46. Czerny T, Busslinger M. DNA-binding and transactivation properties of Pax-6: three amino acids in the paired domain are responsible for the different sequence recognition of Pax-6 and BSAP (Pax-5). Mol Cell Biol. 1995; 15:2858–71.
47. Chauveau C, Decourt C, Cogne M. Insertion of the IgH locus 3ʹ regulatory palindrome in expression vectors warrants sure and efficient expression in stable B cell transfectants. Gene. 1998; 222:279–85.

