INTRODUCTION
Transformer 2β (Tra2β) belongs to the serine/arginine-rich (SR)-like protein splicing factor family. It contains two SR domains and an RNA recognition motif (RRM), and it functions as a sequence-specific pre-mRNA splicing enhancer [1, 2]. Overexpression of Tra2β protein is associated with several types of cancer, including those in breast, ovary, and colon. It appears to accelerate tumor growth and metastasis, and induces drug resistance [3–5]. The human TRA2B gene consists of 10 exons and 9 introns, and produces five mRNA isoforms (TRA2β1 to 5) through alternative splicing [6]. A functional full-length Tra2β protein is translated from TRA2β1 mRNA lacking exon 2, a region that encodes multiple premature termination codons (PTCs) [6]. The TRA2β4 mRNA isoform (referred to here as TRA2β4) contains exon 2 and is not translated to a functional protein [6]. Aberrant splicing occurs in many genetic and acquired diseases, including cancer [7–10]. PTC-containing mRNA splice variants are usually degraded through nonsense-mediated mRNA decay (NMD). However, we previously reported that Hu antigen R regulated alternative splicing of TRA2β pre-mRNA to selectively produce TRA2β4 in human colon cancer cells under oxidative stress [11]. In addition, transcribed TRA2β4 was retained preferentially within the nucleus and was resistant to RNA surveillance NMD [12]. The accumulated TRA2β4 in the nucleus regulates gene expression (including CDKN1A) by occupying transcriptional factor Sp1, resulting in the promotion of cell growth by interrupting p21-related cellular senescence [12].
Intriguingly, the human TRA2B gene contains a 419-bp genomic segment with perfect human-to-rodent sequence identity, a segment termed the “ultraconserved region” (UCR) [13]. This UCR spans exon 2 (276 bp) and its neighboring introns. Among the five isoforms generated from TRA2B, only TRA2β4 contains a complete version of exon 2 (Figure 1A) [6]. There are 481 described UCRs, and more than half of them do not encode a protein [14]. However, 68% of UCRs are transcribed, a group that constitutes a novel category of functional RNAs, transcribed-UCRs (T-UCRs) [14]. Genome-wide expression profiling revealed the distinct signatures of T-UCRs in human leukemia and carcinomas and an association with tumorigenesis [15]. Importantly, the expression levels of several T-UCRs show a tissue-specific pattern that is altered in human diseases, especially in cancer [12, 16–20]. However, understanding of the regulatory mechanisms in T-UCR expression remains unclear.
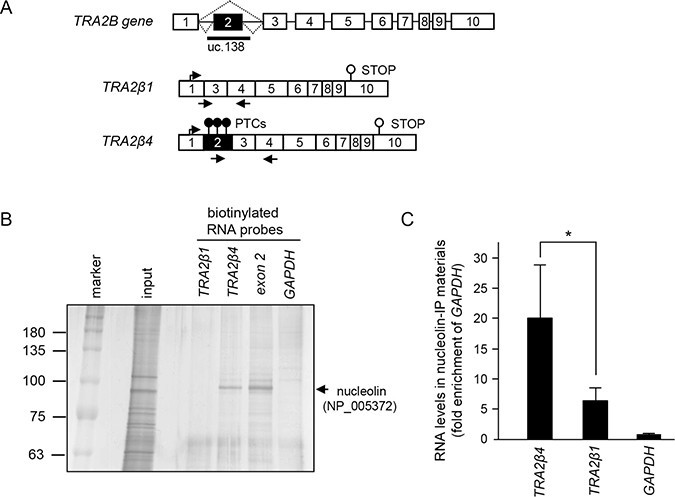
Figure 1: TRA2β4 interacts with nucleolin. (A) Schematic diagram of the human TRA2B gene. Exons are indicated by open boxes and Arabic numbers. Filled boxes denote the ultraconserved exon 2. Two major splice variants TRA2β1 and TRA2β4 are generated from the TRA2B gene and the use of each exon is shown. Black arrows show the specific primers used to detect each of the transcripts. PTCs; premature stop codons. (B) After biotinylated RNA pull-down assays using biotinylated exon 2 probes, the purified proteins were resolved by SDS-PAGE and visualized by silver staining. According to the analysis of the separated ~100 kDa protein by mass spectrometry, nucleolin (NP_005372) was one of RNA-binding proteins in the precipitated materials. (C) Cell lysates from HCT116 were subjected to a RIP assay using an anti-nucleolin antibody. Immunoprecipitated RNAs were quantified by qPCR. Data are shown as enrichment relative to GAPDH. Values are means ± s.d. (n = 6). *Significantly different by unpaired Student’s t-test (p < 0.05).
Recently, several lines of evidence have suggested the important role of RNA-binding proteins (RBPs) in the regulation of the functional activities of long non-coding RNAs (lncRNAs) [21, 22]. For example, the bromodomain protein Brd4 regulates a lncRNA called HOTAIR, which is essential for glioblastoma proliferation [23]. Accordingly, we focused on the association between RBPs and a UCR-containing TRA2β4 in the nucleus and examined whether post-transcriptional regulation impacted aberrant expression of T-UCR.
Distinct functional non-coding RNAs retained within the mammalian cell nucleus are now referred to as “nuclear-retained regulatory RNAs”, and they are suggested to play structural roles or act as riboregulators [24, 25]. Although the majority of lncRNAs reside in the nucleus, only a portion have been functionally characterized. In this study, using biotinylated RNA pull-down assays followed by liquid chromatographic-mass spectrometric (LS/MS) analysis, we identified nucleolin as a TRA2β4-binding protein. Nucleolin is a multifunctional protein that is located primarily in the nucleus. It plays significant roles in the metabolism and function of RNA, including transcription, ribosome assembly, mRNA turnover and translation, and it participates in the regulation of cell fate [26–28]. By interacting with target mRNAs that typically contain AU-rich and/or G-rich elements, nucleolin regulates their expression state [27, 29]. We suggest here a potential role of nucleolin in nuclear localization of TRA2β4 that facilitates abnormal growth of T-UCR-bearing cancer cells.
RESULTS
Association between TRA2β4 and nucleolin
The human TRA2B gene contains an ultraconserved region (uc.138) that spans exon 2 and its neighboring introns. The TRA2β mRNA isoform that contains exon 2 (TRA2β4) is produced by alternative splicing (Figure 1A). Although exon 2 contains multiple PTCs, TRA2β4 resides preferentially in the nuclei of HCT116 cells and could escape from the NMD surveillance pathway [12]. To reveal the mechanism supporting the expression of TRA2β4 in colon cancer cells, we first searched for nuclear RNA-binding protein(s) that specifically interacted with and retained TRA2β4 in nuclei. Here, we employed biotinylated RNA pull-down analyses with the biotinylated exon 2 probe followed by LS/MS analysis. The isolated proteins with molecular masses around 100 kDa preferentially contained RNA-binding proteins including nucleolin and splicing factors, such as heterogeneous nuclear ribonucleoprotein (hnRNP) UL 1, hnRNPR, hnRNPU, and hnRNPQ (Figure 1B and Supplementary Table 1). These hnRNPs likely participate in alternative splicing reaction of exon 2. We were particularly interested in nucleolin (NP_005372), a nuclear protein related to cell survival, proliferation, and invasion of cancer cells [30]. To confirm the binding between nucleolin and endogenous TRA2β4, ribonucleoprotein immunoprecipitation (RIP) analysis was carried out with an anti-nucleolin antibody and specific primers for TRA2B transcripts. TRA2β4 was more abundantly precipitated in nucleolin-IP materials compared with TRA2β1 (Figure 1C). Taken together, TRA2β4 was likely associated with nucleolin in the nucleus of HCT116 cells.
Identification of nucleolin-binding sites in TRA2β4
To determine the binding sites of TRA2β4 exon 2 that were responsible for its association with nucleolin, we prepared a series of biotinylated RNA fragments (F1- F5) that encoded various lengths of TRA2B exon 2 (Figure 2A). Biotinylated RNA pull-down assays showed more specific association of nucleolin with fragment F2 compared with that of F1 (Figure 2B, upper panel). Then, fragment F2 was further divided into three parts (F3-F5). The putative binding site(s) was likely present in both F3 and F4 (Figure 2B, lower panel). As shown in Figure 2C, the common sequences between F3 and F4 (475–494) contain a G-rich element (485-GGGG-488), which was reported to be essential for binding with nucleolin [31]. The introduction of two-point mutations (mt; 485-GGGG-488 to 485-GGAA-488) into exon 2 F2 significantly reduced its interaction with nucleolin (Figure 2D). We also generated constructs encoding FLAG-tagged full length TRA2β4 (pCMV- TRA2β4) or two point-mutated TRA2β4 (pCMV- TRA2β4 mt) (Figure 2E). These plasmids were transfected into HCT116 cells and then RIP assays were carried out with an anti-nucleolin antibody. Using primer sets to detect endogenous FLAG-tagged TRA2β4, we confirmed that mutagenesis in exon 2 decreased the interaction between TRA2β4 and nucleolin (Figure 2E).
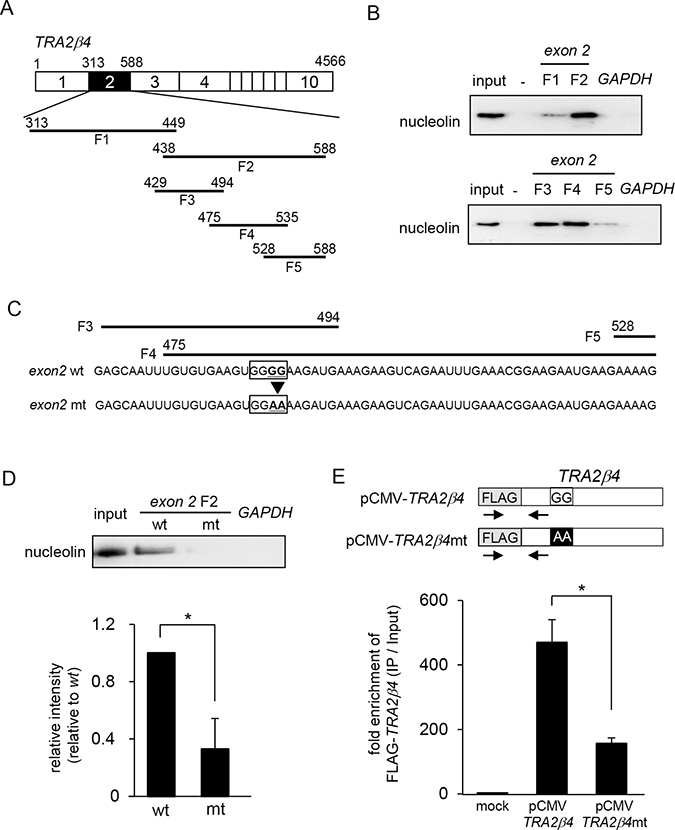
Figure 2: Identification of nucleolin-binding sites in TRA2β4. (A) Schema of the fragments of TRA2β4 that were used for in vitro binding assays. (B) A biotinylated RNA pull-down assay was carried out using lysates prepared from HCT116 cells and the biotinylated RNA fragments. (C) The RNA sequence in exon 2 F3 to F5. The introduction of two-point mutations (mt; 485-GGGG-488 to 485-GGAA-488) in TRA2β4 exon 2. (D) A biotinylated RNA pull-down assay was carried out using biotinylated RNA fragments with (mt) or without (wt) mutations. (E) The constructs coding FLAG-tagged full length TRA2β4 (pCMV- TRA2β4) or the mutated TRA2β4 (pCMV- TRA2β4 mt) were generated. These plasmids were transfected into HCT116 cells and then RIP assays were carried out with an anti-nucleolin antibody. The exogenous TRA2β4 in IP materials was detected using the primers shown as arrows. Each value represents the mean ± s.d. from 3 independent experiments. *Significantly different by unpaired Student’s t-test (p < 0.05).
TRA2β4 recognition domain in nucleolin
To further verify the specific association between nucleolin and TRA2β4, we generated a construct encoding FLAG-tagged nucleolin. Nucleolin includes 3 major domains: an acidic N-terminal region (NT), RNA binding domains (RBDs), and a glycine/arginine-rich (GAR) domain (Figure 3A). Nucleolin binds to the pre-ribosomal RNA processing complex through the NT (1–269 aa). The central region of nucleolin contains four tandem RRMs (307–647 aa), an RBD, which mediates the association with mRNAs. An arginine-glycine-glycine (RGG) repeat in the GAR domain (649–698 aa) in the C-terminal region of nucleolin plays an important role for interaction with target mRNAs as well as other proteins [32].
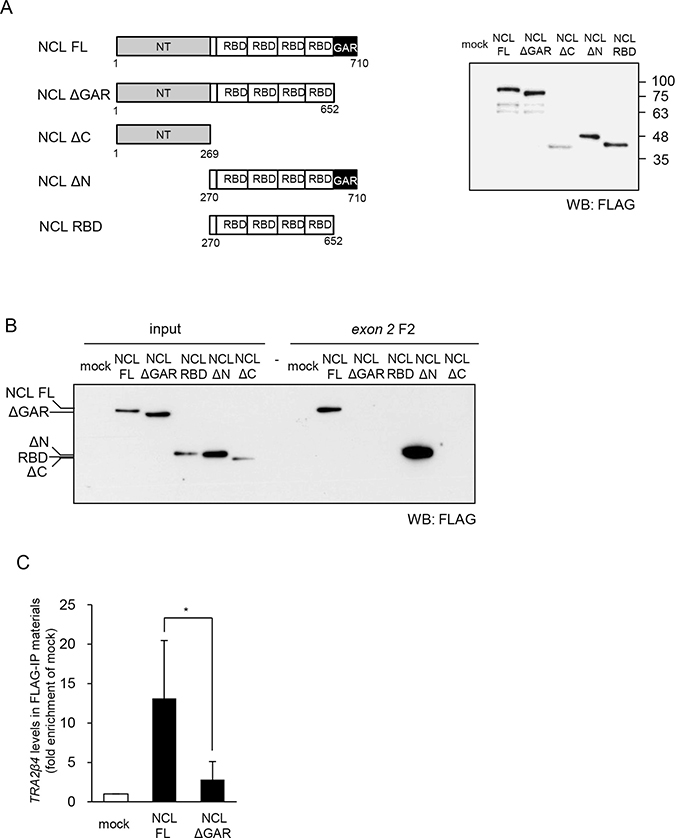
Figure 3: Characterization of TRA2β4 recognition domain in nucleolin. (A) The scheme of constructs encoding full-length nucleolin (NCL FL), lacking the GAR (NCL ΔGAR), lacking the C-terminus (NCL ΔC), lacking the N-terminal domain (NT) (NCL ΔN), and RNA-binding domains (RBDs) only (NCL RBD). Each plasmid was transfected into HCT116 cells for 48 hours, and then whole-cell lysates were subjected to Western blotting using an anti-FLAG antibody. (B) A biotinylated RNA pull-down assay was carried out using biotinylated RNA fragment F2 and lysates prepared from HCT116 cells that were transfected with each plasmid. The bound nucleolin was detected by Western blotting using anti-FLAG antibody. (C) The plasmids bearing full-length or GAR-deficient nucleolin were transfected into HCT116 cells and then a RIP assay was carried out with an anti-FLAG antibody. TRA2β4 in IP materials was detected by qPCR. Each value represents the mean ± s.d. from 3 independent experiments. *Significantly different by unpaired Student’s t-test (p < 0.05).
We first produced 5 different constructs expressing full-length or truncated nucleolin proteins: (1) pCMV-FLAG-tagged full-length nucleolin (NCL FL), (2) nucleolin lacking GAR (NCL ΔGAR), (3) NT only (NCL ΔC), (4) RBDs and GAR (NCL ΔN), and (5) RBDs (NCL RBD) (Figure 3A). Cell lysates were prepared from HCT116 cells overexpressing one of these constructs and subjected to pull-down assays using biotinylated exon 2 F2. As shown in Figure 3B, full-length nucleolin was detected in the complex with biotinylated exon 2 using an anti-FLAG antibody. The association of nucleolin with exon 2 did not require its N-terminal domain nor tandem RBDs of nucleolin, whereas the presence of the c-terminal domain seemed to be essential for the association with TRA2β4 exon 2 (Figure 3B). Finally, we confirmed that the association between nucleolin and TRA2β4 exon 2 disappeared when GAR was deleted from nucleolin (Figure 3B). To further validate the association, we asked whether the deletion of GAR affected the interaction between nucleolin and endogenous TRA2β4. Using an anti-FLAG antibody, RIP was employed to detect the association between TRA2β4 and either wild-type or GAR-deficient nucleolin. The RIP assay showed that the GAR-deleted nucleolin mutant could not bind to TRA2β4 (Figure 3C).
Reduction of TRA2β4 levels after nucleolin knockdown
We next examined whether the specific interaction between TRA2β4 and nucleolin affected TRA2β4 expression. A reduction of nucleolin levels using siRNAs targeting the coding region (NCL #1) or 3′-UTR (NCL #2) of NCL mRNA significantly decreased TRA2β4 levels in HCT116 cells (Figure 4A and 4B). To elucidate the mechanisms for the TRA2β4 decline following nucleolin knockdown, we measured TRA2β4 stability in control or nucleolin siRNA-transfected cells. Nucleolin knockdown significantly accelerated the degradation rate of TRA2β4, whereas it did not affect the stability of TRA2β1 mRNA (Figure 4C). After downregulation of endogenous nucleolin using NCL #2 siRNA targeting its 3′-UTR, nucleolin expression was rescued by transfection with plasmids encoding a FLAG-tagged coding region of nucleolin with or without GAR (Figure 4D). Overexpression of full-length nucleolin completely rescued TRA2β4 expression, whereas GAR-deficient nucleolin had no effect on TRA2β4 levels (Figure 4E). We also confirmed that the replacement of the full-length, but not GAR-deficient nucleolin, blocked the accelerated degradation of TRA2β4 in HCT116 cells treated with the nucleolin 3′-UTR-targeting siRNA (Figure 4F). Hence, we hypothesize that nucleolin may influence TRA2β4 stability via association through the GAR domain.
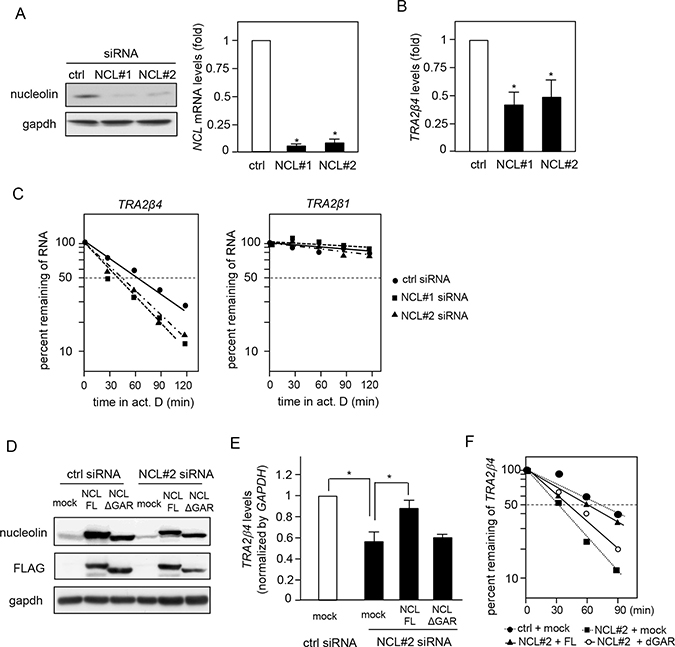
Figure 4: Effect of nucleolin on TRA2β4 stability. (A) HCT116 cells were treated with siRNAs for 48 h as follows: 10 nM control, NCL#1 (targeting the coding region of nucleolin), or NCL #2 (targeting 3′-UTR). Nucleolin levels were analyzed by Western blotting and qPCR. (B) The levels of TRA2β4 in control or nucleolin-silenced cells were measured by qPCR. (C) HCT116 cells were transfected with control, NCL #1, or NCL #2 siRNAs for 48 h and then incubated in the presence of 2 μg/mL actinomycin D for the indicated times. TRA2β1 and TRA2β4 levels were measured by qPCR and plotted on a logarithmic scale to calculate the time required for each RNA to reach one-half of its initial abundance (50%, dashed line). (D) After downregulation of endogenous nucleolin using an siRNA targeting its 3′-UTR (NCL#2), nucleolin expression was rescued by transfection with plasmids encoding a FLAG-tagged coding region of nucleolin with or without GAR. (E) The levels of TRA2β4 in nucleolin-rescued cells were measured by qPCR. (F) TRA2β4 levels in nucleolin-rescued cells after incubation with 2 μg/mL actinomycin D for the indicated times were measured by qPCR. The results were plotted on a logarithmic scale to calculate the time required for each RNA to reach one-half of its initial abundance (50%, dashed line).
Regulation of nuclear localization of TRA2β4 by nucleolin
To test whether nucleolin regulated the nuclear localization of TRA2β4, we first examined the subcellular localization of nucleolin. We prepared HCT116 cells that overexpressed FLAG-tagged full-length nucleolin, or one of the truncated nucleolins (Figure 3A). As shown in Figure 5A, full-length nucleolin mainly resided in nuclei, whereas the deletion of GAR (ΔGAR, ΔC, RBD) led to nucleolin’s loss of its nuclear retention ability. After NMD was inactivated by treatment with 100 μg/mL cycloheximide, we measured the amounts of TRA2β4 and TRA2β1 mRNAs in the cytoplasmic or nuclear fraction. As shown in Figure 5B, silencing of nucleolin decreased the nuclear content of TRA2β4 without changing the subcellular localization of TRA2β1 mRNA. The purities of the extracted nuclear and cytoplasmic fractions were confirmed by Western blotting using antibodies for cytosolic (α-tubulin) or nuclear proteins (hnRNP C1/C2) and by RT-PCR using primers targeting GAPDH pre-mRNA as a nuclear marker (Figure 5C).
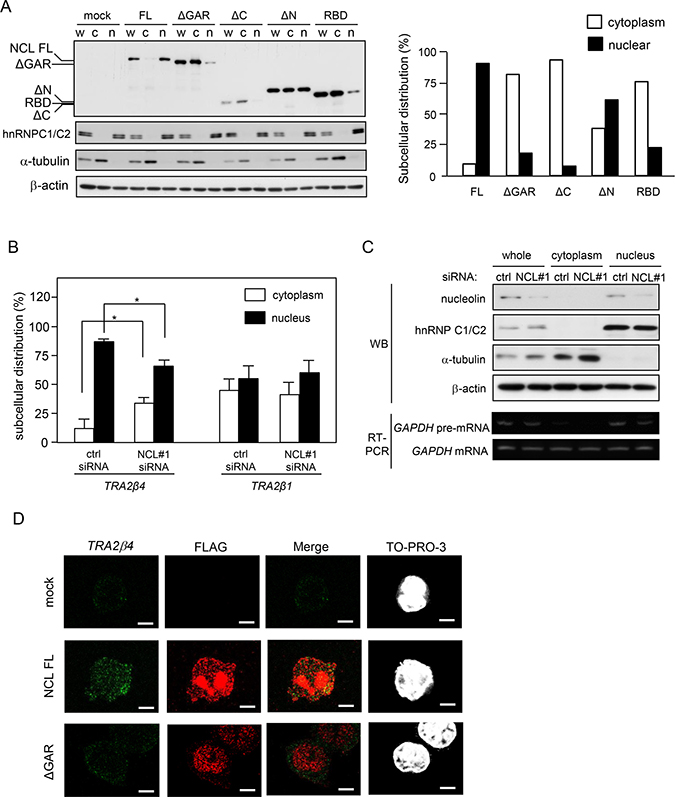
Figure 5: Nucleolin is essential for nuclear localization of TRA2β4. (A) After transfection with the truncated nucleolin, the subcellular localization was measured by Western blotting. FLAG signals were quantified by densitometry. (B) After NMD was inhibited with cycloheximide treatment (100 μg/mL), the amounts of TRA2β4 and TRA2β1 mRNAs in the cytoplasmic or nuclear fraction were analyzed by qPCR. (C) Purities of the extracted nuclear and cytoplasmic fractions were confirmed by Western blotting using antibodies for cytosolic (α-tubulin) or nuclear proteins (hnRNP C1/C2) and by RT-PCR using primers targeting GAPDH pre-mRNA as a nuclear marker. (D) After a 24-h transfection of full-length or ΔGAR nucleolin, subcellular localization of TRA2β4 (green) and nucleolin (red) was examined by RNA-fluorescence in situ hybridization (RNA-FISH) using a probe that specifically hybridized to exon 2 (313-588 nt) and anti-FLAG antibody. Nucleoli were counterstained with TO-PRO-3. Scale bars, 5 μM.
In a previous study, we demonstrated that TRA2β4 was predominantly located in the nucleic fraction compared with TRA2β1 mRNA [12]. In this study, RNA FISH staining for TRA2β4 was employed with immunostaining of FLAG-tagged full-length nucleolin or ΔGAR overexpressed in HCT116 cells. TRA2β4 signals were detectable in the nucleus of mock-transfected, control cells (Figure 5D, upper panel). Overexpression of full-length nucleolin increased TRA2β4 signals in the nucleus (Figure 5D, middle panel), and the overexpressed nucleolin partially co-localized with TRA2β4. However, in ΔGAR-overexpressing cells, TRA2β4 signals were diffusely distributed both in the nucleus and the cytoplasm. ΔGAR signals did not co-localize with TRA2β4 signals (Figure 5D, lower panels).
Roles of nucleolin and TRA2β4 in cell growth and resistance to anticancer drug
Several reports have shown that the inhibition of nucleolin with siRNAs or anti-nucleolin agents inhibits cell growth [33–36]. In fact, knockdown of nucleolin with NCL siRNA #1 and #2 significantly inhibited HCT116 cell growth (Figure 6A). The growth inhibition with NCL siRNA #2, which targeted 3′-UTR of NLC mRNA, was reversed by overexpression of a full-length nucleolin, but not by that of a GAR deficient nucleolin (ΔGAR) (Figure 6B). Knockdown of nucleolin induces apoptosis via caspase 3 activation in response to oxidative stress or anticancer agents such as doxorubicin and etoposide [37, 38]. When HCT116 cells were treated with NCL siRNA #2 and exposed to etoposide for 24 h, caspases 3/7 were activated (Figure 6C) and cell numbers were decreased (Figure 6D). In these endogenous nucleolin-silenced cells, overexpression of full-length nucleolin, but not that of ΔGAR, could block the activation of caspases 3/7 (Figure 6C) and rescue the etoposide-induced cell loss (Figure 6D). To further confirm the importance of the nucleolin-mediated TRA2β4 expression for the resistance to etoposide, we rescued TRA2β4 in nucleolin knockdown cells. As shown in Figure 6E, overexpression of TRA2β4 significantly inhibited the etoposide-induced activation of caspases 3/7 in nucleolin knockdown cells. These results suggested that nucleolin seemed to be necessary for the tumorigenic property of TRA2β4.
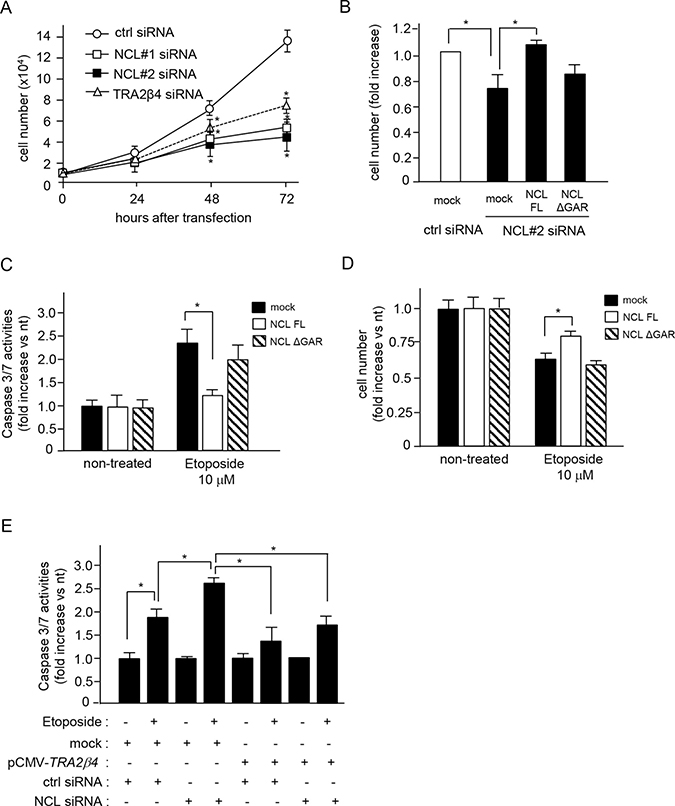
Figure 6: Association between nucleolin and TRA2β4 is essential for cell growth and susceptibility to an anticancer drug. (A) HCT116 cells (1 × 104 cells) were seeded in 35-mm dishes and transfected with the indicated siRNA. Subsequently, growing cells were harvested and counted at the indicated times. Values are means ± s.d. from 4 independent experiments. *Significantly different by unpaired Student’s t-test compared with control siRNA-treated cells (p < 0.05). (B) After transfection with control (ctrl) or NCL siRNA #2 for 12 h, the cells were treated with plasmids encoding a FLAG-tagged full-length or ΔGAR nucleolin. After a 48-h transfection, these cells were harvested and counted. Values are means ± s.d. from 4 independent experiments. *Significantly different by unpaired Student’s t-test (p < 0.05). (C, D) HCT116 cells were treated with the indicated siRNAs and plasmids for 24 h, and then cells were exposed to 10 μM etoposide for 24 h. Subsequently, growing cells were harvested and counted or used for measurement of caspase 3/7 activities. Values are means ± s.d. from 4 independent experiments. *Significantly different by unpaired Student’s t-test (p < 0.05). (E) After HCT116 cells were treated with the indicated siRNAs and plasmids for 24 h, the cells were exposed to 10 μM etoposide for 24 h. Subsequently, growing cells were harvested and caspase 3/7 activities were measured. Values are means ± s.d. from 3 independent experiments. *Significantly different by unpaired Student’s t-test (p < 0.05).
As previously reported [12], TRA2β4 siRNA significantly inhibited growth of HCT116 cells (Supplementary Figure 1A, 1B and Figure 6A) without changing either protein or mRNA levels of nucleolin as well as Tra2β (Supplementary Figure 1C–1E). In the previous study, we found that TRA2β4 accelerated cell growth by downregulating p21 expression through an inhibition of Sp1-binding to the CDKN1A promoter [12]. We therefore tested whether nucleolin modulated the TRA2β4/p21 pathway. Chromatin immunoprecipitation (ChIP) assays indicated that nucleolin did not directly associate with the p21/CDKN1A promoter (Supplementary Figure 2A). In addition, silencing of Sp1 had no effect on nucleolin binding to the CDKN1A promoter, suggesting that nucleolin did not compete with Sp1 for binding to the promoter of CDKN1A (Supplementary Figure 2A). The reduction of endogenous nucleolin with NCL siRNA #2 caused the decline of TRA2β4 levels (Figure 4E) and resulted in a corresponding increase of p21/CDKN1A mRNA levels in mock-transfected, control cells (Supplementary Figure 2B). Overexpression of full length nucleolin, but not ΔGAR, could block this increase of p21/CDKN1A mRNA levels (Supplementary Figure 2B). These results suggest that nucleolin may play an important role in the TRA2β4/p21-mediated regulation of cell growth. The GAR domain may be crucial for this function.
Gene expression signatures in TRA2β4- or nucleolin-silenced cells
Finally, gene expression signatures in TRA2β4- or nucleolin-silenced cells were examined to understand the underlying molecular mechanism and to assess the ability of nucleolin to impact nuclear-retained TRA2β4-mediated gene expression (NCBI Gene Expression Omnibus #GSE112758). As shown in Figure 7A, microarray analysis showed that NCL siRNA #1-treated cells differentially expressed 2,493 genes (≥ 1.5-fold), compared with control siRNA-treated cells. TRA2β4-silenced cells altered the expression of 3,044 genes (≥ 1.5-fold). Interestingly, a total of 630 genes were commonly changed in the same direction between TRA2β4- and nucleolin-silenced cells. Ingenuity Pathway Analysis (QIAGEN Bioinformatics) was employed to identify biological functions relevant to the 630 genes. As shown in Figure 7B, the enriched molecular and cellular functions of genes regulated in both nucleolin- and TRA2β4-silenced cells included 1) Cellular Movement (p = 1.35E-07), 2) Cell Death and Survival (p = 3.92E-07), and Cell Cycle (p = 2.00E-06). Moreover, the top disease associated with the 630 genes was Cancer (p = 1.88E-12). These results suggest that TRA2β4 may be one of the important targets for facilitating abnormal cell growth.
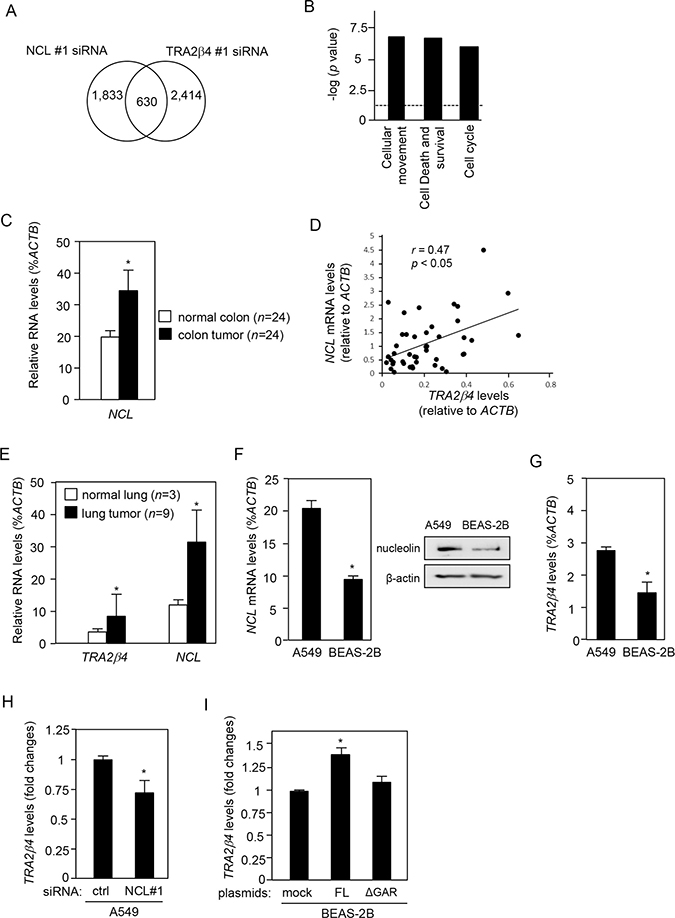
Figure 7: Nucleolin regulates TRA2β4 expression in colon and breast cancer cells. (A) Microarray analysis showed that NCL siRNA #1-treated cells differentially expressed 2,493 genes (≥ 1.5-fold), compared with control siRNA-treated cells. TRA2β4-silenced cells altered the expression of 3,044 genes (≥ 1.5-fold). A total of 630 genes were commonly changed in the same direction between TRA2β4- and nucleolin-silenced cells. (B) Commonly regulated 630 genes were subjected to Ingenuity Pathway Analysis (QIAGEN Bioinformatics) to identify biologically relevant functions. (C) Using colon adenocarcinoma TissueScan Tissue qPCR arrays (HCRT103, OriGene), NCL and ACTB mRNAs were measured by qPCR. ACTB mRNA was used as an endogenous quality control. *Significantly different by paired Student’s t-test (p < 0.05). (D) The correlation between TRA2β4 and NCL expression in a colon cancer cDNA array was analyzed by determining the Pearson’s correlation coefficient. (E) Expression levels of TRA2β4, NCL, and ACTB mRNAs were measured by qPCR using lung cancer cDNA libraries (CSRT101, OriGene). ACTB mRNA was used as an endogenous quality control. Values are means ± s.d. *Significantly different by unpaired Student’s t-test (p < 0.05). (F) Both protein and mRNA levels of nucleolin in lung carcinoma (A549) and lung epithelial cells (BEAS-2B) were analyzed by qPCR or Western blotting. Values are means ± s.d. from 3 independent experiments. *Significantly different by unpaired Student’s t-test (p < 0.05). (G) The levels of TRA2β4 expression were measured by qPCR in A549 and BEAS-2B cells. Values are means ± s.d. from 4 independent experiments. *Significantly different by unpaired Student’s t-test (p < 0.05). (H) After A549 cells were treated with 10 nM of the indicated siRNA for 48 h, changes in the expression levels of TRA2β4 were measured by qPCR. GAPDH mRNA was used as an endogenous quality control. Values are means ± s.d. from 4 independent experiments. *Significantly different by unpaired Student’s t-test compared with control (ctrl) siRNA-treated cells (p < 0.05). (I) After BEAS-2B cells were transfected with mock or plasmids encoding nucleolin with/without GAR for 48 h, changes in the expression level of TRA2β4 were measured by qPCR. GAPDH mRNA was used as an endogenous quality control. Values are means ± s.d. from 3 independent experiments. *Significantly different by unpaired Student’s t-test compared with mock-transfected cells (p < 0.05).
Nucleolin-dependent expression of TRA2β4 in colon and lung cancer cells
Using a paired colon cancer cDNA panel prepared from 24 patients (HCRT103; OriGene, Rockville, MD, USA), we measured NCL mRNA levels. Compared with paired normal colon tissues, colon cancers expressed significantly higher levels of NCL mRNA (Figure 7C). TRA2β4 was overexpressed in human colon cancer cell lines and colon carcinomas [12]. We examined the association between the expression of NCL mRNA and that of TRA2β4, and we found that the TRA2β4 levels were positively correlated with those of NCL in both normal and colon cancer tissues (r = 0.47, Pearson’s correlation coefficient) (Figure 7D).
We also examined the expression of TRA2β4 and NCL mRNA in lung cancer cells. Expression levels of TRA2β4 and NCL mRNAs were upregulated in lung cancer cDNA libraries (CSRT101, OriGene) compared with normal lung tissues (Figure 7E). Human lung cancer cell line A549 cells expressed higher amounts of NCL mRNA and nucleolin protein than those in lung epithelial BEAS-2B cells (Figure 7F). At the same time, A549 cells expressed significantly higher amounts of TRA2β4 compared with BEAS-2B cells (Figure 7G). Nucleolin knockdown also decreased in TRA2β4 levels in A549 cells (Figure 7H). In contrast, overexpression of full-length (but not GAR-deficient nucleolin) in BEAS-2B cells significantly increased TRA2β4 expression (Figure 7I). Thus, the nucleolin-mediated regulation of TRA2β4 expression was not unique to HCT116 colon cancer cells.
DISCUSSION
About 70% of the 481 UCRs in the human genome are transcribed and appear to be functional RNAs, T-UCRs [13]. For each T-UCR, one corresponding to the sense genomic sequence is named ultraconserved (Uc.) ‘+’, and the other complementary sequence is Uc. ‘-’. Recent studies have shown that T-UCRs function as regulators in multiple pathways such as pri-miRNA processing, transcriptional regulation, translation, and chromatin modification [20, 39, 40]. Dysregulated expression of distinct T-UCRs may be directly linked to the development of various diseases. For instance, decreased expression of Uc.173 in the hippocampus is associated with lead-induced neuronal apoptosis by inter-regulation of miRNAs [41]. Uc.261 is overexpressed in intestinal mucosa of active Crohn’s disease and is suggested to participate in the damage to tight junctions in inflammation [42]. In addition, differential expression of T-UCRs has been observed in several types of cancer [14, 16, 18, 19]. Uc.190, Uc.233 and Uc.270 levels are increased in human pancreatic adenocarcinoma and knockdown of each of them reduced the proliferation of MIA PaCa-2 pancreatic cancer cells [19]. Uc. 8+ works as a natural decoy for miR-596 and cooperates in the promotion and development of bladder cancer [20]. Carotenuto et al. reported that Uc. 158- was activated by the Wnt/β-catenin pathway in liver cancer and drove cellular growth and migration, possibly by modulating miR-193b expression [18]. One of 5 transcripts generated from the human TRA2B gene, TRA2β4 is categorized as a T-UCR (Uc.138+). TRA2β4 is overexpressed in human gastric cancer cells and colon cancer cells and facilitates their growth [11, 12, 43]. Knockdown of TRA2β4 in colon cancer cells leads to cellular senescence through p21 induction, meaning that TRA2β4 is one of the key molecules that causes aberrant cell cycle regulation of cancer cells [12].
The molecular mechanisms of T-UCR expression are not fully understood. Several studies have demonstrated that T-UCR expression is altered at the transcriptional level in cancer cells. A batch of T-UCRs (Uc.160+, Uc.285+, Uc.346+) undergo specific CpG island hypermethylation-associated silencing in colon cancer cells [44]. Suppression of Uc.160+ expression by DNA hypermethylation was also observed in human gastric cancer cells [17]. Tumor-related activation of transcription factor Egr1 caused induction of genes encoding T-UCRs in pancreatic adenocarcinoma [19]. Recent studies have suggested that functional RNAs, including lncRNAs, are also regulated in a post-transcriptional manner. LncRNAs are reported to exert their function by various mechanisms, such as interacting with regulatory proteins to modulate their processing and metabolism and directing localization within cellular compartments. A novel lncRNA, colorectal neoplasia differentially expressed (CRNDE), promotes cell proliferation and metastasis of colorectal carcinoma [45]. The turnover of CRNDE is dependent on its associating protein, heterogeneous nuclear ribonucleoprotein U-like 2 (hnRNPUL2) [45]. On the other hand, it is not fully understood whether RNA-binding proteins are involved in the regulation of T-UCR expression at the post-transcriptional levels. In colon cancer cells, TRA2β4 preferentially resides in the nucleus and could escape RNA degradation in the cytoplasm [12]. Non-coding RNAs retained within the nucleus are referred to as nuclear-retained regulatory RNAs, and their structural roles are recognized as riboregulators [46]. To elucidate the mechanism for nuclear retention of TRA2β4, we characterized TRA2β4-associated nuclear proteins.
This study shows for the first time that a nuclear RNA-binding protein, nucleolin, is a possible regulator of TRA2β4 expression. Nucleolin interacted with TRA2β4 via a G-rich sequence (485-GGGG-488) in exon 2. This interaction is likely crucial for nuclear retention and the resultant stabilization of TRA2β4. Overexpression of full-length nucleolin significantly increased the stability of TRA2β4 and accelerated cell proliferation. A significant part of overexpressed nucleolin co-localized with TRA2β4 in the nucleus. In contrast, nucleolin knockdown decreased TRA2β4 levels by accelerating its degradation. Nucleolin is composed of three main structural domains: the N-terminus, the central region, and the C-terminal domain [26]. The C-terminus is the arginine/glycine-rich (GAR) domain and interacts with nucleic acids [28]. Importantly, GAR-deficient nucleolin did not modulate nuclear localization of TRA2β4, resulting in accelerated decay of TRA2β4 in the cytoplasm. GAR-deficient nucleolin could not induce TRA2β4-mediated tumorigenic effects such as accelerated cell growth and anticancer drug resistance. Finally, we confirmed the positive correlation between TRA2β4 and nucleolin expression levels in colon cancer cells.
Nucleolin is a clinically important protein as it possesses oncogenic properties. In fact, nucleolin is overexpressed in a number of malignant tumors, including cancers of the breast, lung, stomach, pancreas, cervix, prostate as well as colorectal cancers, melanomas and leukemias [28, 47–52]. For instance, nucleolin binds to a receptor tyrosine kinase (ErbB2) and promotes its activation, resulting in enhancement of tumorigenicity of breast cancer [52]. In gastric cancer, Yang et al. demonstrated that nucleolin mediated the epithelial-mesenchymal transition via upregulation of the Erk1/2 and Akt pathways [50]. Moreover, nucleolin stabilizes the mRNA of anti-apoptotic protein bcl-2 [53]. Recent studies have proposed that nucleolin is a novel target for anticancer therapy as demonstrated by the effects of antagonistic molecules [33, 54, 55]. Nucleolin is a multifunctional protein that is mainly found in the nucleolus, but it is also observed in the nucleoplasm and cytoplasm. Nucleoplasmic nucleolin is involved in the regulation of translation and stability of oncogenic mRNAs [33, 56]. Interestingly, nucleolin may protect cancer cells from senescence by controlling intracellular localization of the telomerase complex [57].
In the previous study, we showed that TRA2β4 accelerated cancer cell growth by downregulating p21 expression through an inhibition of Sp1 binding to the CDKN1A promoter [12]. In this study, we found that nucleolin did not compete with Sp1 for binding to the promoter of CDKN1A. We also assessed the ability of nucleolin to impact nuclear-retained TRA2β4-mediated gene expression indirectly by measuring changes in gene expression in nucleolin- or TRA2β4-silenced cells. Both manipulated cells commonly changed expression of a large number of genes (630 genes). IPA ranked “Cancer” as the top disease associated with the 630 genes. Of course, nucleolin exerts its oncogenic properties by interacting with many targets as described above. TRA2β4 may be one of the important targets for facilitating abnormal growth of cancer cells. The GAR-domain may be essential for the activity of tumorigenic T-UCR TRA2β4 retained within the nucleus. Further studies are needed to reveal the pathological significance of the nucleolin-mediated aberrant expression of TRA2β4.
MATERIALS AND METHODS
Cell culture and transfection
Cells of the human colorectal carcinoma line (HCT116) were cultured in McCoy’s 5A medium (Thermo Fisher Scientific, Waltham, MA) supplemented with 5% (v/v) heat-inactivated fetal bovine serum (FBS) and antibiotics (penicillin and streptomycin). Human lung cell lines (BEAS-2B and A549) were maintained in RPMI-1640 medium (Thermo Fisher Scientific) supplemented with 10% FBS. These cells were cultured at 37° C in 5% CO2. Twenty-four h prior to transfection, 1 × 104 cells were cultured in 6-well plates and then treated with 10 nM control or nucleolin (NCL) mRNA-targeting siRNA (Santa Cruz, CA, USA) using Lipofectamine RNAiMAX (Thermo Fisher Scientific). Expression plasmids encoding the full-length or truncated nucleolin were transfected with Lipofectamine 3000 according to the manufacturer’s protocol (Thermo Fisher Scientific).
Apoptosis was assessed by measuring caspase-3/7 activities using the caspase-Glo 3/7 assay kit (Promega, Fitchburg, WI, USA) according to the manufacturer’s protocol.
Western blotting
Whole cell lysates were prepared using RIPA (Thermo Fisher Scientific) with a complete protease inhibitor cocktail (Roche, Mannheim, Germany). After HCT116 cells were incubated in cytosolic lysis buffer (10 mM Tris-HCl, pH 7.4, containing 100 mM NaCl, 2.5 mM MgCl2, and 40 μg/ml digitonin) for 10 min at 4° C, the resultant lysates were centrifuged at 2,060 g for 8 min at 4° C, and the supernatants were collected as cytosolic extracts. The pellets were washed twice with the lysis buffer and then lysed with RIPA buffer. After centrifugation at 10,000 g for 10 min at 4° C, the resultant supernatants were collected as nuclear extracts, as previously described [12]. Ten micrograms of the extracted proteins were separated by sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis and then transferred to a polyvinylidene fluoride membrane (Bio-Rad, Hercules, CA, USA). After blocking with 5% nonfat dry milk (Cell Signaling Technology, Danvers, MA, USA) for 1 h at room temperature, the membrane was incubated with anti-nucleolin (C23) (Santa Cruz), anti-FLAG (Sigma-Aldrich, St. Louis, MO, USA), anti-hnRNPC1/C2 (Santa Cruz), anti-α-tubulin (Santa Cruz), anti-β-actin (Abcam, Cambridge, UK), or anti-glyceraldehyde 3-phosphate dehydrogenase (Gapdh) (Santa Cruz) antibody overnight at 4° C. Following incubation with an appropriate secondary antibody for 1 h at room temperature, the bound antibodies were detected with Pierce Western Blotting Substrate (Thermo Fisher Scientific). The intensities of the bound antibodies were quantified by using ImageJ software (National Institutes of Health, USA).
Biotinylated RNA pull-down analysis
RNA fragments were generated by PCR using specific primers containing the T7 RNA polymerase promoter sequence (CTAATACG ACTCACTATAGGGAGA [T7]) (Supplementary Table 2). Biotinylated transcripts were synthesized in vitro using the MAXIscript Kit (Thermo Fisher Scientific) and the Biotin-11-CTP (PerkinElmer, CT, USA). The synthesized transcripts were then purified with ssDNA/RNA Clean and Concentrator (Zymo Research, CA, USA). After HCT116 cells were lysed with RIPA buffer, whole-cell lysates were prepared by centrifugation at 10,000 g for 10 min at 4° C and used for the biotinylated RNA pull-down assay. Eighty μg of whole-cell lysates were incubated with 2.5 μg of the biotinylated transcripts in TENT buffer (10 mM Tris-HCl buffer, pH 8.0, containing 1 mM EDTA, 250 mM NaCl, and 0.5% Triton X-100) for 30 min at room temperature [5, 11]. After adding Dynabeads M-280 Streptavidin (Thermo Fisher Scientific), the mixture was incubated for 1 h at room temperature. The bound proteins were pulled down using a magnetic particle separator and analyzed by Western blotting or liquid chromatography-mass spectrometry analysis (nanoLC-MS/MS, CapLC, Q-Tof).
Real-time quantitative PCR (qPCR)
Total RNA was extracted from cells using the RNAiso Plus (Takara, Tokyo, Japan) and precipitated in the presence of Dr. GenTLE Precipitation Carrier (Takara). Isolated RNAs were reverse-transcribed using ReverTra Ace qPCR RT Master Mix (Toyobo, Osaka, Japan). TRA2β1, TRA2β4, NCL, ACTB, and GAPDH mRNA levels were measured using specific primer sets and SYBR Green Master Mix (Thermo Fisher Scientific) (Supplementary Table 2). SiRNA-treated cells were incubated with 2 μg/mL actinomycin D for the indicated times and then the amounts of TRA2β4 and GAPDH mRNAs in the cells were measured by qPCR using 18S rRNA for normalization.
Ribonucleoprotein immunoprecipitation (RIP) analysis
Twenty μl of Protein A Sepharose (Sigma-Aldrich) and 3 μg of the anti-nucleolin antibody were mixed in 1 ml of NT-2 buffer (50 mM Tris HCl; 150 mM NaCl; 1 mM MgCl2; 0.05% Nonidet P-40) [27]. The mixture was rotated overnight at 4° C and then washed 3 times with ice-cold NT-2 buffer. HCT116 cells were lysed with lysis buffer (pH 7.5) consisting of 25 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, 1% (v/v) Nonidet P-40, 5% (v/v) glycerol, and 100 U/ml RNase inhibitor (Nacalai Tesque, Tokyo, Japan) [11]. After these lysates were rotated for 2 h at 4° C with the pre-coated protein A Sepharose, they were treated with TURBO DNase (Thermo Fisher Scientific) for 10 min at 37° C, and then with 0.1% SDS and proteinase K solution (Wako, Osaka, Japan) for 10 min at 55° C. Finally, bound RNAs were extracted using a phenol-chloroform mixture and then subjected to qPCR.
Plasmids
Full-length human NCL mRNA was cloned into the pCMV- FLAG vector (Agilent Technology, CA, USA). In brief, a fragment encoding a full-length nucleolin (pCMV-NCL FL; amino acid (aa) 1 to 710) was amplified by PCR using the following primer set: 5′-GGATCCATGGTGAAGCTCGCGAAGGCAGG-3′ (forward) and 5′-GAATTCCTATTCAAACTTCGTCTTCTTTCCT-3′ (reverse). The amplified fragment was subcloned into pCMV- FLAG vector using BamHI and EcoRI sites (these sequences are underlined). The truncated constructs were subcloned using the following primers: a construct for the deletion of GAR (pCMV-NCL ΔGAR; aa 1 to 652); 5′-GGATCCATGGTGAAGCTCGCGAAGGCAGG-3′ and 5′-GAATTCGCCACCTTCACCCTTAGGTTTGGC-3′; N-terminus of nucleolin (pCMV-NCL ΔC, aa 1 to 269), 5′-GGATCCATGGTGAAGCTCGCGAAGGCAGG-3′ and 5′-GAATTCCTCCTCCTCTTCTTCCTCCTCCTC A-3′; C-terminus of nucleolin (pCMV-NCL ΔN, aa 270 to 710), 5′-GGATCCGAAGAGCCTGTCAAA GAAGCACCT-3′ and 5′-GAATTCCTATTCAAACTTCG TCTTCTTTCCT-3′; RNA-binding domains (pCMV-NCL RBD; aa 270 to 652), 5′-GGATCCGAAGAGCCT GTCAAAGAAGCACCT-3′ and 5′-GAATTCGCCACC TTCACCCTTAGGTTTGGC-3′.
RNA fluorescence in situ hybridization (RNA-FISH) and immunofluorescence staining
To detect TRA2β4, sequences complementary to exon 2 (313–588 nt) were transcribed in vitro using a FITC RNA labeling kit (Roche). HCT116 cells were fixed in 4% paraformaldehyde for 15 min and then permeabilized with 0.5% Triton X-100 for 15 min at room temperature. These cells were incubated with a 1 μg/mL FITC-labeled RNA probe in hybridization buffer (1 × Denhardt’s solution containing 100 μg/mL tRNA, 0.01% Tween-20, and 5% dextran sulfate) at 55° C for 16 h. After washing, the cells were incubated overnight with anti-FITC polyclonal antibody (Thermo Fisher Scientific) and anti-FLAG monoclonal antibody (Sigma-Aldrich) at 4° C. After washing with TBST, the cells were treated with fluorophore-conjugated secondary antibodies (Alexa Fluor 488, 555: Thermo Fisher Scientific) for 1 h at room temperature and mounted with Vectashield (Vector Laboratories).
Gene expression profiling
Total RNAs were extracted from cells using RNeasy kit (Qiagen) according to the manufacturer’s protocol. Purified RNA quality was assessed by Agilent 2100 Bioanalyzer using an RNA 6000 Nano Labchip kit (Agilent Technologies, Santa Clara, CA, USA). RNA samples with 9.0 RNA integrity number (RIN) were used to measure mRNA expression profiles using a human mRNA microarray (SurePrint G3 Human; Agilent). The expression data were analyzed using GeneSpring 14.9 (Agilent).
Statistical analysis
All statistical analyses were performed with the SPSS 21.0 software package (SPSS Inc., Chicago, IL, USA). Results are expressed as means ± SD. Significant differences between two groups were estimated by two-tailed Student’s t-test. Non-parametric data were analyzed using a Wilcoxon-Mann-Whitney U test when comparing two groups. Pearson’s correlation analysis was used to estimate the relationship between the expression levels of NCL and TRA2β4. *p < 0.05 was considered statistically significant.
Author contributions
This study was conceived and supervised by YS, YK, and KR; YS and YK performed most of the experiments. YS performed the biotinylated RNA pull-down assays in Figures 1, 2, and 3. KK, YA, MH and KF contributed with biochemical and microscope analysis. YS, YK, KN and KR wrote the manuscript.
ACKNOWLEDGMENTS AND FUNDING
This research was supported by research grants (#15K15294, #26713027) from the Ministry of Education, Culture, Sports, Science and Technology, Japan (Y.K) and Takeda Science Foundation. This study was supported by Support Center for Advanced Medical Sciences, Institute of Biomedical Sciences, Tokushima University Graduate School.
CONFLICTS OF INTEREST
The authors declare that they have no conflict of interest.
REFERENCES
1. Dauwalder B, Amaya-Manzanares F, Mattox W. A human homologue of the Drosophila sex determination factor transformer-2 has conserved splicing regulatory functions. Proc Natl Acad Sci U S A. 1996; 93:9004–9. https://doi.org/10.1073/pnas.93.17.9004.
2. Tsuda K, Someya T, Kuwasako K, Takahashi M, He F, Unzai S, Inoue M, Harada T, Watanabe S, Terada T, Kobayashi N, Shirouzu M, Kigawa T, et al. Structural basis for the dual RNA-recognition modes of human Tra2-β RRM. Nucleic Acids Res. 2011; 39:1538–53. https://doi.org/10.1093/nar/gkq854.
3. Watermann DO, Tang Y, Zur Hausen A, Jäger M, Stamm S, Stickeler E. Splicing factor Tra2-β1 is specifically induced in breast cancer and regulates alternative splicing of the CD44 gene. Cancer Res. 2006; 66:4774–80. https://doi.org/10.1158/0008-5472.CAN-04-3294.
4. Gabriel B, Zur Hausen A, Bouda J, Boudova L, Koprivova M, Hirschfeld M, Jager M, Stickeler E. Significance of nuclear hTra2-beta1 expression in cervical cancer. Acta Obstet Gynecol Scand. 2009; 88:216–21. https://doi.org/10.1080/00016340802503021.
5. Kuwano Y, Nishida K, Kajita K, Satake Y, Akaike Y, Fujita K, Kano S, Masuda K, Rokutan K. Transformer 2β and miR-204 regulate apoptosis through competitive binding to 3′ UTR of BCL2 mRNA. Cell Death Differ. 2015; 22:815–25. https://doi.org/10.1038/cdd.2014.176.
6. Nayler O, Cap C, Stamm S. Human transformer-2-beta gene (SFRS10): complete nucleotide sequence, chromosomal localization, and generation of a tissue-specific isoform. Genomics. 1998; 53:191–202. https://doi.org/10.1006/geno.1998.5471.
7. Hertel KJ. Combinatorial control of exon recognition. J Biol Chem. 2008; 283:1211–5. https://doi.org/10.1074/jbc.R700035200.
8. Srebrow A. The connection between splicing and cancer. J Cell Sci. 2006; 119:2635–41. https://doi.org/10.1242/jcs.03053.
9. Licatalosi DD, Darnell RB. RNA processing and its regulation: global insights into biological networks. Nat Rev Genet. 2010; 11:75–87. https://doi.org/10.1038/nrg2673.
10. Venables JP, Klinck R, Bramard A, Inkel L, Dufresne-Martin G, Koh CS, Gervais-Bird J, Lapointe E, Froehlich U, Durand M, Gendron D, Brosseau JP, Thibault P, et al. Identification of alternative splicing markers for breast cancer. Cancer Res. 2008; 68:9525–31. https://doi.org/10.1158/0008-5472.CAN-08-1769.
11. Akaike Y, Masuda K, Kuwano Y, Nishida K, Kajita K, Kurokawa K, Satake Y, Shoda K, Imoto I, Rokutan K. HuR Regulates Alternative Splicing of the TRA2 Gene in Human Colon Cancer Cells under Oxidative Stress. Mol Cell Biol. 2014; 34:2857–73. https://doi.org/10.1128/MCB.00333-14.
12. Kajita K, Kuwano Y, Satake Y, Kano S, Kurokawa K, Akaike Y, Masuda K, Nishida K, Rokutan K. Ultraconserved region-containing Transformer 2β4 controls senescence of colon cancer cells. Oncogenesis. 2016; 5: e213. https://doi.org/10.1038/oncsis.2016.18.
13. Bejerano G, Pheasant M, Makunin I, Stephen S, Kent WJ, Mattick JS, Haussler D. Ultraconserved elements in the human genome. Science. 2004; 304:1321–5. https://doi.org/10.1126/science.1098119.
14. Scaruffi P. The Transcribed-Ultraconserved Regions: A Novel Class of Long Noncoding RNAs Involved in Cancer Susceptibility. Sci World J. 2011; 11:340–52. https://doi.org/10.1100/tsw.2011.35.
15. Calin GA, Liu CG, Ferracin M, Hyslop T, Spizzo R, Sevignani C, Fabbri M, Cimmino A, Lee EJ, Wojcik SE, Shimizu M, Tili E, Rossi S, et al. Ultraconserved Regions Encoding ncRNAs Are Altered in Human Leukemias and Carcinomas. Cancer Cell. 2007; 12:215–29. https://doi.org/10.1016/j.ccr.2007.07.027.
16. Peng JC, Shen J, Ran ZH. Transcribed ultraconserved region in human cancers. RNA Biol. 2013; 10:1771–7. https://doi.org/10.4161/rna.26995.
17. Honma R, Goto K, Sakamoto N, Sekino Y, Sentani K, Oue N, Yasui W. Expression and function of Uc.160+, a transcribed ultraconserved region, in gastric cancer. Gastric Cancer. 2017; 20:960–969. https://doi.org/10.1007/s10120-017-0714-9.
18. Carotenuto P, Fassan M, Pandolfo R, Lampis A, Vicentini C, Cascione L, Paulus-Hock V, Boulter L, Guest R, Quagliata L, Hahne JC, Ridgway R, Jamieson T, et al. Wnt signalling modulates transcribed-ultraconserved regions in hepatobiliary cancers. Gut. 2017; 66:1268–1277. https://doi.org/10.1136/gutjnl-2016-312278.
19. Jiang J, Azevedo-Pouly AC, Redis RS, Lee EJ, Gusev Y, Allard D, Sutaria DS, Badawi M, Elgamal OA, Lerner MR, Brackett DJ, Calin GA, Schmittgen TD. Globally increased ultraconserved noncoding RNA expression in pancreatic adenocarcinoma. Oncotarget. 2016; 7:53165–53177. https://doi.org/10.18632/oncotarget.10242.
20. Olivieri M, Ferro M, Terreri S, Durso M, Romanelli A, Avitabile C, De Cobelli O, Messere A, Bruzzese D, Vannini I, Marinelli L, Novellino E, Zhang W, et al. Long non-coding RNA containing ultraconserved genomic region 8 promotes bladder cancer tumorigenesis. Oncotarget. 2016; 7:20636–54. https://doi.org/10.18632/oncotarget.7833.
21. Hua WF, Zhong Q, Xia TL, Chen Q, Zhang MY, Zhou AJ, Tu ZW, Qu C, Li MZ, Xia YF, Wang HY, Xie D, Claret FX, et al. RBM24 suppresses cancer progression by upregulating miR-25 to target MALAT1 in nasopharyngeal carcinoma. Cell Death Dis. 2016; 7252:1–12. https://doi.org/10.1038/cddis.2016.252.
22. Noh JH, Kim KM, Abdelmohsen K, Yoon JH, Panda AC, Munk R, Kim J, Curtis J, Moad CA, Wohler CM, Indig FE, de Paula W, Dudekula DB, et al. HuR and GRSF1 modulate the nuclear export and mitochondrial localization of the lncRNA RMRP. Genes Dev. 2016; 30:1224–39. https://doi.org/10.1101/gad.276022.115.
23. Pastori C, Kapranov P, Penas C, Peschansky V, Volmar CH, Sarkaria JN, Bregy A, Komotar R, St Laurent G, Ayad NG, Wahlestedt C. The Bromodomain protein BRD4 controls HOTAIR, a long noncoding RNA essential for glioblastoma proliferation. Proc Natl Acad Sci U S A. 2015; 112:8326–31. https://doi.org/10.1073/pnas.1424220112.
24. Quinodoz S, Guttman M. Long noncoding RNAs: An emerging link between gene regulation and nuclear organization. Trends Cell Biol. 2014; 24:651–63. https://doi.org/10.1016/j.tcb.2014.08.009.
25. Singh DK, Prasanth KV. Functional insights into the role of nuclear-retained long noncoding RNAs in gene expression control in mammalian cells. Chromosome Res. 2013; 21:695–711.
26. Ginisty H, Sicard H, Roger B, Bouvet P. Structure and functions of nucleolin. J Cell Sci. 1999; 112:761–72. https://doi.org/10.1093/emboj/17.5.1476.
27. Abdelmohsen K, Tominaga K, Lee EK, Srikantan S, Kang MJ, Kim MM, Selimyan R, Martindale JL, Yang X, Carrier F, Zhan M, Becker KG, Gorospe M. Enhanced translation by Nucleolin via G-rich elements in coding and non-coding regions of target mRNAs. Nucleic Acids Res. 2011; 39:8513–30. https://doi.org/10.1093/nar/gkr488.
28. Jia W, Yao Z, Zhao J, Guan Q, Gao L. New perspectives of physiological and pathological functions of nucleolin (NCL). Life Sci. 2017; 186:1–10. https://doi.org/10.1016/j.lfs.2017.07.025.
29. Sengupta TK, Bandyopadhyay S, Fernandes DJ, Spicer EK. Identification of Nucleolin as an AU-rich Element Binding Protein Involved in bcl-2 mRNA Stabilization. J Biol Chem. 2004; 279:10855–63. https://doi.org/10.1074/jbc.M309111200.
30. Berger CM, Gaume X, Bouvet P. The roles of nucleolin subcellular localization in cancer. Biochimie. 2015; 113:78–85. https://doi.org/10.1016/j.biochi.2015.03.023.
31. Woo HH, Baker T, Laszlo C, Chambers SK. Nucleolin Mediates MicroRNA-directed CSF-1 mRNA Deadenylation but Increases Translation of CSF-1 mRNA. Mol Cell Proteomics. 2013; 12:1661–77. https://doi.org/10.1074/mcp.M112.025288.
32. Thandapani P, O’Connor TR, Bailey TL, Richard S. Defining the RGG/RG Motif. Mol Cell. 2013; 50:613–23. https://doi.org/10.1016/j.molcel.2013.05.021.
33. Palmieri D, Richmond T, Piovan C, Sheetz T, Zanesi N, Troise F, James C, Wernicke D, Nyei F, Gordon TJ, Consiglio J, Salvatore F, Coppola V, et al. Human anti-nucleolin recombinant immunoagent for cancer therapy. Proc Natl Acad Sci U S A. 2015; 112:9418–23. https://doi.org/10.1073/pnas.1507087112.
34. Wise JF, Berkova Z, Mathur R, Zhu H, Braun FK, Tao RH, Sabichi AL, Ao X, Maeng H, Samaniego F. Nucleolin inhibits Fas ligand binding and suppresses Fas-mediated apoptosis in vivo via a surface nucleolin-Fas complex. Blood. 2013; 121:4729–39. https://doi.org/10.1182/blood-2012-12-471094.
35. Pichiorri F, Palmieri D, De Luca L, Consiglio J, You J, Rocci A, Talabere T, Piovan C, Lagana A, Cascione L, Guan J, Gasparini P, Balatti V, et al. In vivo NCL targeting affects breast cancer aggressiveness through miRNA regulation. J Exp Med. 2013; 210:951–68. https://doi.org/10.1084/jem.20120950.
36. Yamada T, Park CS, Shen Y, Rabin KR, Lacorazza HD. G0S2 inhibits the proliferation of K562 cells by interacting with nucleolin in the cytosol. Leuk Res. 2014; 38:210–7. https://doi.org/10.1016/j.leukres.2013.10.006.
37. Jain N, Zhu H, Khashab T, Ye Q, George B, Mathur R, Singh RK, Berkova Z, Wise JF, Braun FK, Wang X, Patel K, Xu-Monette ZY, et al. Targeting nucleolin for better survival in diffuse large B-cell lymphoma. Leukemia. 2018; 32:663–674. https://doi.org/10.1038/leu.2017.215.
38. Zhang B, Wang H, Jiang B, Liang P, Liu M, Deng G, Xiao X. Nucleolin/C23 is a negative regulator of hydrogen peroxide-induced apoptosis in HUVECs. Cell Stress Chaperones. 2010; 15:249–57. https://doi.org/10.1007/s12192-009-0138-5.
39. Liz J, Portela A, Soler M, Gómez A, Ling H, Michlewski G, Calin GA, Guil S, Esteller M. Regulation of pri-miRNA processing by a long noncoding RNA transcribed from an ultraconserved region. Mol Cell. 2014; 55:138–147. https://doi.org/10.1016/j.molcel.2014.05.005.
40. Mestdagh P, Fredlund E, Pattyn F, Rihani A, Van Maerken T, Vermeulen J, Kumps C, Menten B, De Preter K, Schramm A, Schulte J, Noguera R, Schleiermacher G, et al. An integrative genomics screen uncovers ncRNA T-UCR functions in neuroblastoma tumours. Oncogene. 2010; 29:3583–92. https://doi.org/10.1038/onc.2010.106.
41. Nan A, Zhou X, Chen L, Liu M, Zhang N, Zhang L, Luo Y, Liu Z, Dai L, Jiang Y. A transcribed ultraconserved noncoding RNA, Uc.173, is a key molecule for the inhibition of lead-induced neuronal apoptosis. Oncotarget. 2016; 7:112–124. https://doi.org/10.18632/oncotarget.6590.
42. Qian XX, Peng JC, Xu AT, Zhao D, Qiao YQ, Wang TR, Shen J, Ran ZH. Noncoding Transcribed Ultraconserved Region (T-UCR) uc.261 Participates in Intestinal Mucosa Barrier Damage in Crohn’s Disease. Inflamm Bowel Dis. 2016; 12:2840–2852.
43. Takeo K, Kawai T, Nishida K, Masuda K, Teshima-Kondo S, Tanahashi T, Rokutan K. Oxidative stress-induced alternative splicing of transformer 2beta (SFRS10) and CD44 pre-mRNAs in gastric epithelial cells. Am J Physiol Cell Physiol. 2009; 297: C330–8. https://doi.org/10.1152/ajpcell.00009.2009.
44. Lujambio A, Portela A, Liz J, Melo S, Rossi S, Spizzo R, Croce C, Calin G, Esteller M. CpG island hypermethylation-associated silencing of non-coding RNAs transcribed from ultraconserved regions in human cancer. Oncogene. 2010; 29:6390–6401. https://doi.org/10.1038/onc.2010.361.
45. Jiang H, Wang Y, Ai M, Wang H, Duan Z, Wang H, Zhao L, Yu J, Ding Y, Wang S. Long noncoding RNA CRNDE stabilized by hnRNPUL2 accelerates cell proliferation and migration in colorectal carcinoma via activating Ras/MAPK signaling pathways. Cell Death Dis. 2017; 8: e2862–12. https://doi.org/10.1038/cddis.2017.258.
46. Prasanth KV, Spector DL. Eukaryotic regulatory RNAs: an answer to the 'genome complexity' conundrum. Genes Dev. 2007; 21:11–42. https://doi.org/10.1101/gad.1484207.
47. Wu DM, Zhang P, Liu RY, Sang YX, Zhou C, Xu GC, Yang JL, Tong AP, Wang CT. Phosphorylation and changes in the distribution of nucleolin promote tumor metastasis via the PI3K/Akt pathway in colorectal carcinoma. FEBS Lett. 2014; 588:1921–9. https://doi.org/10.1016/j.febslet.2014.03.047.
48. Gilles ME, Maione F, Cossutta M, Carpentier G, Caruana L, Di Maria S, Houppe C, Destouches D, Shchors K, Prochasson C, Mongelard F, Lamba S, Bardelli A, et al. Nucleolin targeting impairs the progression of pancreatic cancer and promotes the normalization of tumor vasculature. Cancer Res. 2016; 76:7181–8193. https://doi.org/10.1158/0008-5472.CAN-16-0300.
49. Yan J, Zhang Y, Ren C, Shi W, Chen L. Involvement of nuclear protein C23 in activation of EGFR signaling in cervical cancer. Tumour Biol. 2016; 37:905–10. https://doi.org/10.1007/s13277-015-3889-x.
50. Yang Y, Yang C, Zhang J. C23 protein meditates bone morphogenetic protein-2-mediated EMT via up-regulation of Erk1/2 and Akt in gastric cancer. Med Oncol. 2015; 32:76. https://doi.org/10.1007/s12032-015-0547-5.
51. Tsou JH, Chang KY, Wang WC, Tseng JT, Su WC, Hung LY, Chang WC, Chen BK. Nucleolin regulates c-Jun/Sp1-dependent transcriptional activation of cPLA2α in phorbol ester-treated non-small cell lung cancer A549 cells. Nucleic Acids Res. 2008; 36:217–27. https://doi.org/10.1093/nar/gkm1027.
52. Wolfson E, Goldenberg M, Solomon S, Frishberg A, Pinkas-Kramarski R. Nucleolin-binding by ErbB2 enhances tumorigenicity of ErbB2- positive breast cancer. Oncotarget. 2016; 7:65320–65334. https://doi.org/10.18632/oncotarget.11323.
53. Otake Y, Soundararajan S, Sengupta TK, Kio EA, Smith JC, Pineda-Roman M, Stuart RK, Spicer EK, Fernandes DJ. Overexpression of nucleolin in chronic lymphocytic leukemia cells induces stabilization of bcl2 mRNA. Hematology. 2009; 109: 3069–75. https://doi.org/10.1182/blood-2006-08-043257.
54. Chen Z, Xu XH. Roles of nucleolin: Focus on cancer and anti-cancer therapy. Saudi Med J. 2016; 37:1312–1318. https://doi.org/10.15537/smj.2016.12.15972.
55. Soundararajan S, Chen W, Spicer EK, Courtenay-Luck N, Fernandes DJ. The nucleolin targeting aptamer AS1411 destabilizes Bcl-2 messenger RNA in human breast cancer cells. Cancer Res. 2008; 68:2358–65. https://doi.org/10.1158/0008-5472.CAN-07-5723.
56. Pickering BF, Yu D, Van Dyke MW. Nucleolin protein interacts with microprocessor complex to affect biogenesis of microRNAs 15a and 16. J Biol Chem. 2011; 286:44095–103. https://doi.org/10.1074/jbc.M111.265439.
57. Khurts S, Masutomi K, Delgermaa L, Arai K, Oishi N, Mizuno H, Hayashi N, Hahn WC, Murakami S. Nucleolin interacts with telomerase. J Biol Chem. 2004; 279:51508–15. https://doi.org/10.1074/jbc.M407643200.

