INTRODUCTION
Follicular lymphoma (FL) is a low-grade B cell lymphoma usually characterized by long life expectancy and heterogeneous outcome. About one third of FL patients alternate periods of disease remission with recurrent relapses. Some FLs undergo histologic transformation to a high-grade neoplasm, a transition associated with a more aggressive clinical course and poor survival [1].
The current evaluation of the risk of relapse/resistance to multiagent chemotherapy of FL is based on clinical parameters summarized in the Follicular Lymphoma International Prognostic Index (FLIPI) while no consistent biological marker has been identified that predicts the survival or the risk of relapse after drug treatments [2]. The majority of candidate biomarkers studied in FL patients focused on morphologic features, cytogenetic abnormalities, molecular aberrations, somatic mutations and altered protein expression [3–7]. Although the new discovered somatic mutations are promising markers of prognosis in FL, to date no single hallmark or single driver mutation can be considered predictive of relapse.
Gene expression studies have revealed that both survival and progression-free disease are significantly associated with non-neoplastic immune cell gene signatures [8, 9]. According to gene expression signatures, FLs split into two groups, i.e. T-cell-rich and dendritic cell-/macrophage-rich [10–14]. In FL, T-cells number, functionality and distribution may impact patient prognosis. PD1+ cells associated with shorter time to relapse of FLs [13]. High number of Foxp3+ cells predicted improved survival of FL patients [10]. However, the association of Foxp3+ cells with FL outcome was questioned in other studies [2, 13].
MiRNAs play critical regulatory roles in the immune system and are necessary for proper lineage decision [15]. MiRNA biogenesis is under tight temporal and spatial control and dysregulation is associated with many human diseases [16, 17]. A direct role of miRNAs in lymphomagenesis was demonstrated by knock-out miRNA mouse models with a direct impact on B-cell lymphoma/leukemia development [18]. In particular, miR-155, miR-150, miR-21, cluster miR-17-92 and miR-34a have pivotal role in the control of master regulator transcription factors essential in the B-cell biology [19]. MiR-16 expression discriminated FLs with and without BCL2 translocation [20]. MiRNA signatures in FLs were obtained from both the comparison with reactive lymph nodes (rLN) or from cells isolated from FL and rLNs [21–24]. Despite the potential relevance of miRNAs in FL biology, the association between the expression level of deregulated miRNAs and clinical parameters of FL patients has not been conclusively demonstrated yet.
The limited success of approaches applied so far for the identification of prognostic markers for FL patients has prompted us to consider the possible role of miRNA signatures. Since several miRNAs pointed to a role of immune cells, we have investigated the tumor microenvironment by means of immunohistochemistry. We have then investigated the possible association of markers typical of different immune cells with miRNA profiles and the clinical outcome of FL patients.
RESULTS
Increasing alterations of miRNAs expression from low to high grade FL
We performed miRNAs expression analysis by microarrays in 26 FLs and 12 reactive lymph nodes (rLN) as reference. In Figure 1, the heat map summarizes the differences in miRNA expression between FLs and rLNs at 10% FDR (false discovery rate), fold change >1.5. FLs clustered separately from rLNs except three cases. Both FLs and rLNs appeared heterogeneous and split apart into two groups. Seventeen miRNAs resulted upregulated and 12 downregulated in FL in comparison to rLNs (Figure 1). MiR-16 family four miRNAs, miR-15a, miR-15b, miR-16 and miR-195 were correlated to each other and upregulated in most of FLs.
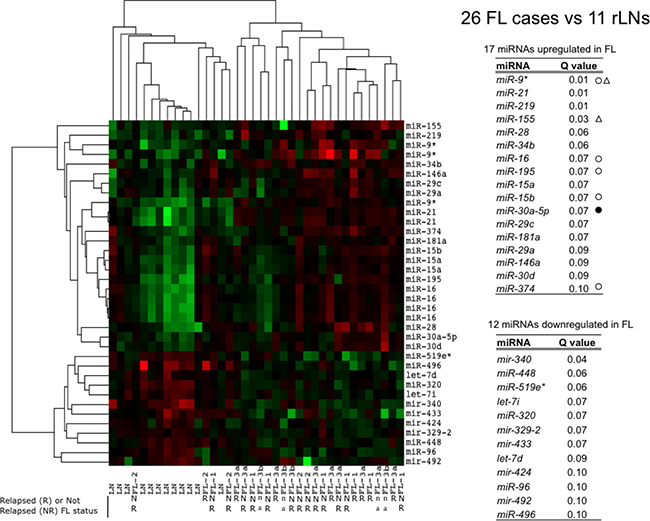
Figure 1: Profile of miRNAs differentially expressed between FLs and rLNs. The heat map describes the expression of 36 spots, corresponding to 29 single miRNAs, differentially expressed among 38 samples: 26 follicular lymphoma (FLs) and 12 reactive lymph nodes (rLNs) (FDR 10%). FL-1, FL-2, FL-3a, FL-3b are FL of grade 1, 2, 3a and 3b, respectively. Red, higher expression (log2, +4); green, lower expression (log2, -4). The tables report the list of 17 miRNAs upregulated in FLs and of 12 miRNAs downregulated in FL in comparison to rLNs and the corresponding Q values. White circles: upregulated in Rohele et al. [23]. Black circles: downregulated in Rohele et al. [23]. White triangles: upregulated in isolated FL cells in Wang et al. [24]. R, relapsed; NR, not relapsed; n.a. not available.
FLs distributed regardless of grade as compared to rLNs. However, when FLs of grade 1, 2 and 3a were separately compared with rLNs, FL grade 1 showed the upregulation of miR-21, miR-9* and miR-155 and the downregulation of miR-340; FL grade 3a showed the upregulation of five miRNAs and the downregulation of five miRNAs (Figure 2A and 2B). In FL grade 3, the increase of miR-219, miR-28 and miR-34a and the decrease of miR-329, miR-320, miR-519e* and miR-516-5p expression added to those altered in FL grade 1 (Figure 2C). FL grade 2 showed no significant differences in comparison to rLNs as well as the direct comparison of FL of different grade.
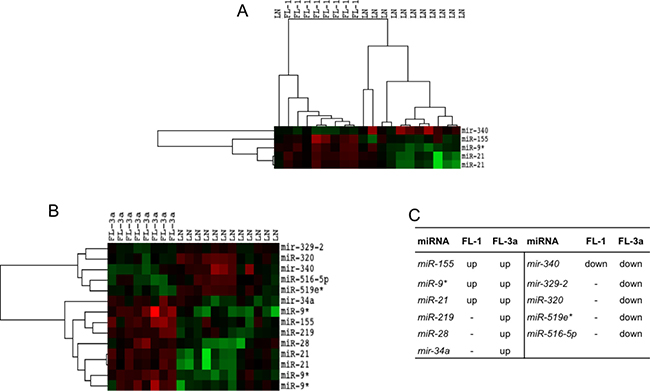
Figure 2: Profile of miRNAs differentially expressed between FL grade 1 or 3a and rLNs. FL-1 and FL-3a are follicular lymphoma (FL) of grade 1 and 3a, respectively. Red, higher expression (log2, +4); green, lower expression (log2, –4). (A) The heat maps describe the expression of 14 spots, corresponding to 11 single miRNAs, differentially expressed in 20 samples: eight FLs grade 1 and 12 reactive lymph nodes (rLNs) (FDR 10%). (B) The heat maps describe the expression of 5 spots, corresponding to 4 single miRNAs, differentially expressed among 20 samples: eight FLs grade 3a and 12 rLNs (FDR 10%). (C) The table reports the list of 11 miRNAs upregulated (up) or downregulated (down) in FL grade 1 and 3a in comparison to rLNs.
Expression of several miRNAs was validated by quantitative RT-PCR in rLN, FL1, FL3a and all FLs (Supplementary Figure 1). MiR-28 (P = 0.05), miR-9 (P = 0.002), miR-9* (P = 0.0001), miR-21 (P = 0.0007) and miR-195 (P = 0.05) were confirmed to be upregulated in FLs as compared to rLN. Mir-219 was not differentially expressed in FLs and rLN (P > 0.1). Downregulation of miR-320 (P = 0.002), miR-340 (P = 0.001) and let-7i(P = 0.05) in FLs was confirmed.
Ratio between expression of 5p and 3p forms of miR-9 and miR-21 is different in FLs compared to rLNs
We investigated whether the alteration of miRNA expression shown by FLs could imply coordinated changes of 5p/3p ratios. For most of miRNAs, no difference of 5p/3p relationships between FLs and rLNs could be observed whereas we observed a change of miR-9-2/miR-9* and miR-21/miR-21* in FLs with respect to rLNs (Figure 3A and 3B). No change of ratio was found for miR-9-1/miR-9* and miR-9-3/miR-9*.
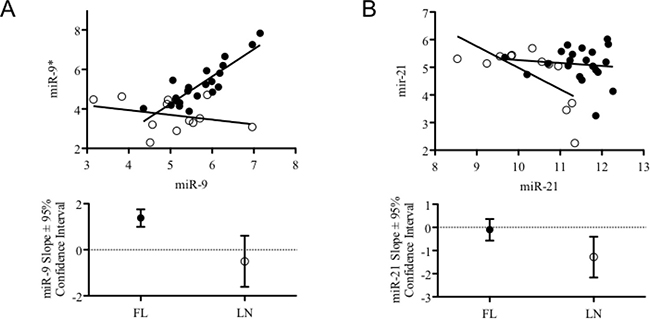
Figure 3: Combined expression levels and trend of 5p and 3p variants of miR-9-2 and miR-21 in FLs and rLNs. (A) Graph reports the expression level of miR-9-2 and miR-9* (from microarray data) in 21 follicular lymphomas (FLs) cases having clinical information (black circles) and in 12 reactive lymph nodes (rLNs) (empty circles). (B) Graph reports the expression level of miR-21 and miR-21* (from microarray data) in 21 FLs cases having clinical information (black circles) and in 12 rLNs (empty circles). Bottom graphs report slope and 95% confidence interval of the trend lines passing through the symbols of the two series represented in top graphs.
T-cells influence the pattern of miRNAs expressed in FLs
FL microenvironment is characterized by the presence of a significant number of T-cells, up to 50% of the total cells [25]. To evaluate whether the T-cell component affected the pattern of miRNA expressed in FLs, we compared miRNA expression profiles of FLs with those of two CD4+ and two CD8+ T-cell samples (Figure 4A and 4B). As expected, FLs and pure T-cells clustered separately and a large number of miRNA discriminated the two categories of samples. Differentially expressed miRNAs split in 3 main clusters. FLs in turn clustered into two groups independently from the FL grade. The same heatmap showing the complete list of differentially expressed miRNAs between FLs and T-cells is reported in Supplementary Figure 2.
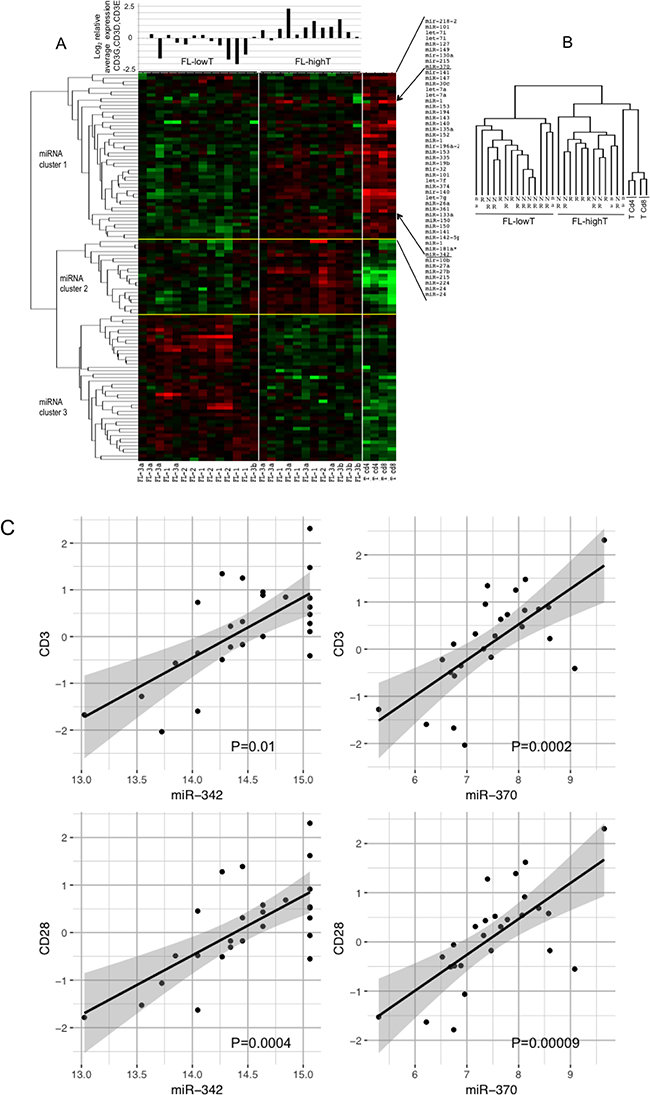
Figure 4: Profile of miRNAs differentially expressed among FLs, CD4+ T-cells and CD8+ T-cells. FL-HT, follicular lymphoma (FL) with high T-cell level. FL-LT, with low T-cell level. (A) The heat map describes the expression levels of miRNAs differentially expressed among 30 samples: 26 FLs, two CD4+ and two CD8+ T-cell samples (FDR 1%). FL-1, FL-2, FL-3a, FL-3b are FLs of grade 1, 2, 3a and 3b, respectively. Red, higher expression (log2, +4); green, lower expression (log2, -4). On the right of the heat map a partial list of miRNA differentially expressed among the three sets of samples, which are expressed at higher level in T-cells than in FLs (miRNA cluster 1). On the top of the heat map is the average mRNA expression level of three CD3 chains gamma, delta and epsilon for each of the 26 FL samples. Expression data were obtained by qRT-PCR. (B) Array tree of 30 samples based on the expression levels of miRNAs differentially expressed. (C) Correlation between the expression levels of miR-342 or miR-370 (from microrray analysis) and the expression level (log2) of CD3 and CD28 mRNAs by qRT-PCR in 26 FLs samples. P was calculated by Spearman’s correlation.
We firstly focused on miRNA cluster 1, which included miRNAs expressed at high level in both FLs and T-cells. To evaluate the possible influence of T-cells on miRNA expression profile of FLs, we measured the level of CD3E, CD3D and CD3G mRNA in FLs by qRT-PCR (Figure 4A). The two clusters of FLs were termed FL-LT (low T) and FL-HT (high T) (Figure 4A, miRNAs in cluster 1 listed on the right). Accordingly, by plotting the average expression level of the three CD3 chains upon each FL we found that the FL-HT group had a significantly higher level of CD3 than FL-LT (P < 0.01) (Figure 4A).
We searched for specific miRNAs representing a possible T-cell signature in FLs. To this aim, we correlated the levels of each miRNA with the average level of CD3 mRNA. MiR-342 (P = 0.01) and miR-370 (P = 0.0002) showed clear correlation with CD3 level (Figure 4C). Similarly, correlations were obtained between the mRNA level of CD28, another marker of T-cells, and miR-342 (P = 0.0004) or miR-370 (P = 0.0009) (Figure 4C).
Remaining miRNAs differentially expressed between FLs and T-cells discriminated cases belonging to FL-HT and FL-LT. Cluster 2 included miRNAs overexpressed in FL-HT group while cluster 3 included miRNAs overexpressed in FL-LT group (Figure 4A). To gain functional information about the cellular process potentially regulated by differentially expressed miRNAs, we performed pathway enrichment analysis using experimentally validated gene targets of miRNAs (Supplementary Tables 2 and 3). Target genes regulated by cluster 2 were mostly involved in T-cell and B-cell development, in cell cycle, and were under Treg (regulatory T-cells), NF-kB and MYC influence. Main pathways associated with cluster 3 included monocytes, macrophages, dendritic cells, Treg, follicular helper T-cells and B-cell receptor activation, as well as NF-kB, caspase and epithelial-mesenchymal transition.
Immuno-architectural pattern and total number of Foxp3+ and PD1+ cells but not CD68+ cells discriminate FLs from rLNs
Number and pattern distribution of regulatory T-cells (Foxp3+), suppressive and exhausted cytotoxic T-cells (PD1+), and macrophages (CD68+) were shown to variate considerably among FLs [10, 11, 13]. To evaluate these immune effector subsets in our FL series, we assessed Foxp3+, PD1+ and CD68+ cells by automated IHC in 24 FLs and in 11 rLNs as reference (Figure 5, Figure 6, Supplementary Figures 3–5).
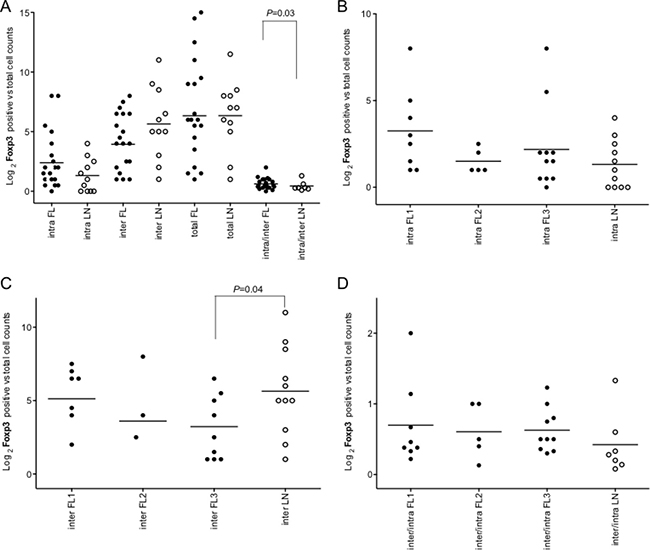
Figure 5: Intrafollicular, interfollicular and total Foxp3+ cell count in FLs and rLNs. White circles, follicular lymphomas (FLs); black circles, reactive lymph nodes (rLNs); black line, median. Intra, inter and total are intrafollicular, interfollicular and intrafollicular plus interfollicular Foxp3+ cells, respectively. Number of Foxp3+ cells in each FL and rLN case was the average number of cells in two consecutive slides. The average number of positive cells was normalized on the whole number of nuclei present in a slide. P values were calculated by t-test. (A) Foxp3+ cell counts in intrafollicular, interfollicular, total and interfollicular/intrafollicular ratio in FLs and rLN. (B) Intrafollicular Foxp3+ cell counts in FL grades and LNs. (C) Interfollicular Foxp3+ cell counts in FL grades and LNs. (D) Interfollicular/intrafollicular ratio Foxp3+ cell counts in FL grades and LNs.
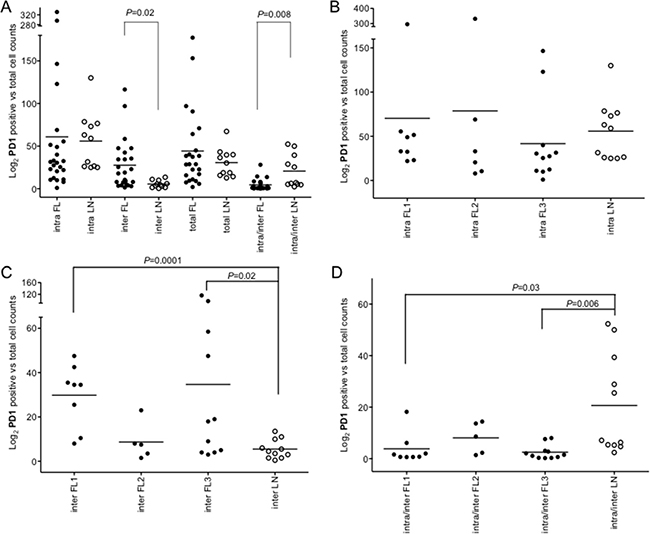
Figure 6: Intrafollicular, interfollicular and total PD1+ cell counts in FLs and rLNs. White circles, follicular lymphomas (FLs); black circles, reactive lymph nodes (rLNs); black line, median. Intra, inter and total are intrafollicular, interfollicular and intrafollicular plus interfollicular PD1+ cells, respectively. Number of PD1+ cells in each FL and rLN case was the average number of cells in two consecutive slides. The average number of positive cells was normalized on the whole number of nuclei present in a slide. P values were calculated by t-test. (A) PD1+ cell counts in intrafollicular, interfollicular, PD1+ total and interfollicular/intrafollicular ratio in FLs and rLNs. (B) Intrafollicular PD1+cell counts in FL grades and rLNs. (C) Interfollicular PD1+cell counts in FL grades and rLNs. (D) Interfollicular/intrafollicular ratio PD1+ cell counts in FL grades and rLNs.
The immuno-architectural structure of immune follicles was taken in account. Only intrafollicular/interfollicular Foxp3+ cells ratio was significantly higher in all FLs compared to rLNs (P = 0.03) (Figure 5A). Intrafollicular Foxp3+ cells were not significantly different in FLs of diverse grade with respect to rLNs (Figure 5B). Interfollicular Foxp3+ cells in FL grade 3 were significantly lower in number than in rLNs (P = 0.03) (Figure 5C). Intrafollicular/interfollicular Foxp3+ cells ratio was not significantly different between FL grade 1, 2 and 3 and rLNs (Figure 5D).
Interfollicular PD1+ cells were higher in FLs compared to rLNs (P = 0.02) and the intrafollicular/interfollicular ratio was higher in rLNs (P = 0.008) (Figure 6A). Intrafollicular PD1+ cells were not significantly different in FL grade 1, 2 and 3 compared to rLNs (Figure 6B), while interfollicular PD1+ cells were more evident in FL grade 1 (P = 0.0001) and FL grade 3 (P = 0.02) (Figure 6C). The intrafollicular/interfollicular PD1+ cells ratio was lower in grade 1 and grade 3 FL with respect to rLNs (P = 0.03 and P = 0.006, respectively) (Figure 6D).
In a minority of tumors, high density of macrophages infiltrating the tumor correlated with lower survival [2, 11, 26]. As expected according to morphology, we observed less intrafollicular and more interfollicular CD68+ cells in FL compared with rLNs, but both findings were not statistically significant. Total CD68+ cells did not change in the two sets (Supplementary Figure 3). A summary of Foxp3+, PD1+ and CD68+ total cell in FLs and rLNs, together with CD3 mRNA levels in FLs, is reported in Supplementary Figure 4. No correlation was found between Foxp3+ cells and CD3 mRNA levels in FLs.
Foxp3+CXCL13+ and Foxp3+PD1+ cells are rare in FL and rLN
In order to characterize the T-cells infiltrating the FL, we investigated the regulatory follicular T cells (TFR) cells, a subset of suppressive T-cells which together with follicular helper T-cells (TFH) cells control the GC reaction [27–29]. TFR cells share phenotypic characteristics with TFH cells and conventional Foxp3+ cells and express significant amounts of the prototypic TFH genes Cxcr5, CXCL13, Icos, Bcl6 and PD1 [28, 30]. We assessed slides double stained for Foxp3-CXCL13 as well as Foxp3-PD1 by IHC in FL and rLN samples. Foxp3+CXCL13+ cells were very rarely observed in FL and rLN (Supplementary Figure 6A and 6B). Foxp3+PD1+ cells were rare or less than 5% of Foxp3+ cells in the neoplastic GC (Supplementary Figure 6C and 6D). A significant number of Foxp3+PD1+ cells was observed in case T20 (Supplementary Figure 6D). Single-positive CXCL13 or PD1 cells represented a variable fraction mostly in the GC cells (Supplementary Figures 6A–6D).
CD8+ cells are reduced in FL in comparison to rLN
CD8+ cells are mainly involved in target cell killing and their increased infiltration was correlated to a better FL prognosis [31]. We assessed abundance and distribution of CD8+ cells in FL and rLN (Supplementary Figure 6E and 6F). The frequency of intratumoral CD8+ cells ranged from 1.5% to 56.1% of cells in FL patients (Supplementary Figure 7A). CD8+ cells were mostly interfollicular and in general less than 2% were intrafollicular. In FL cases T15 and T44, CD8+ cells infiltrated significantly also the GC compartment (Supplementary Figure 6E).
CD8+ cell counts were lower in FL compared to rLN (P = 0.05) (Supplementary Figure 7A). Moreover, CD8+ cell counts were not significantly different in relapsed and not relapsed FL. CD8+ cells correlated positively with PD1+ cells and CD68+ cells in FL (either P < 0.0001) but not in rLN (Supplementary Figure 7B and 7C).
We calculated the CD8+/Foxp3+ cell ratio in FL and rLN samples. This ratio did not discriminate FL from rLN and relapsed from not relapsed FLs.
Twenty-six miRNAs are differentially expressed between relapsed and not relapsed FLs
Twelve patients did not relapse after treatment, while nine FL patients relapsed. We compared the expression levels of miRNAs in relapsed and non-relapsed FLs. According to microarray data, 17 miRNAs were significantly upregulated and nine miRNAs were significantly downregulated in relapsed FLs (P < 0.05) (Table 1).
Table 1: MiRNAs up-regulated or down-regulated in relapsed versus not relapsed FLs and their reported experimentally validated gene targets
MiRNAs upregulated in relapsed vs not relapsed FL |
||
|---|---|---|
miRNA |
P value |
Gene targets* |
miR-376c |
0.0046 |
ACVR1C, IGF1R, GRB2, TGFBR1, TGFA |
mir-450-2 |
0.0102 |
- |
mir-431 |
0.0110 |
- |
miR-1 |
0.0123 |
MCL1, PTEN and other 17 targets |
miR-382 |
0.0126 |
BCL6, NFKB and other 22 targets |
miR-9 |
0.0145 |
BCL6, ETS, PRDM1 and other 42 targets |
miR-19b |
0.0166 |
MYCN, PTEN and other 21 targets |
miR-522 |
0.0190 |
SOX2, FXN |
miR-181a* |
0.0194 |
NANOG |
miR-101 |
0.0206 |
MYCN, EZH2, FOS, MCL1 and other 26 targets |
miR-320 |
0.0264 |
MCL1, PTEN and other 17 targets |
miR-526a |
0.0306 |
- |
miR-196a |
0.0328 |
NFKBIA, BACH1, 6 HOX genes and other 8 targets |
miR-383 |
0.0475 |
CCND1, VEGFA and other 5 targets |
miR-144 |
0.0492 |
NOTCH1, TGFB1, MTOR, PTEN and other 10 targets |
miR-184 |
0.0500 |
BCL2, AKT, MYC and other 5 targets |
miR-9* |
0.0500 |
RCOR1, ITGB1, GNAI1 |
MiRNAs downregulated in relapsed vs not relapsed FL |
||
miRNA |
P value |
Gene targets* |
miR-325 |
0.0022 |
- |
miR-302c* |
0.0072 |
- |
mir-330 |
0.0132 |
E2F1, VEGFA, CD44 and other 4 targets |
mir-376b |
0.0149 |
- |
mir-194-1 |
0.0180 |
ACVR2B, CDH2, SOCS2 and other 14 targets |
mir-106b |
0.0360 |
- |
miR-31 |
0.0363 |
RHOA, PPPR2A, LATS2, SATB2, FOXP3 and other 38 targets |
miR-410 |
0.0432 |
MET, MDM2 |
mir-491 |
0.0494 |
BCL2L1, CHD4, TAF10, MMP9, GIT1, SMAD3 |
*experimentally validated miRNA targets according with miRTarBase, strong evidence, reporter assay [74].
The expression of ten miRNAs correlates with Foxp3+ cell counts in FLs and rLNs
We searched for miRNAs correlating with Foxp3+, CD68+ or PD1+ cells in FLs and rLNs. Ten miRNAs showed correlation with Foxp3+ cells in FLs: directly, mir-325, miR-376b, mir-450-2 and miR-515-1 (P < 0.05) (Supplementary Figure 8A), and inversely miR-144, miR-302b, miR-325, mir-431, miR-432 and miR-490 (P < 0.05) (Supplementary Figure 8B). For eight of the ten miRNAs correlating with Foxp3+ cells in FLs, the relationship between miRNA level and Foxp3+ cells was not significantly different in FLs and rLNs. In contrast, the inverse correlation of miR-144 with Foxp3+ cells was lost in rLNs, while the direct correlation of miR-302b in FLs turned into inverse correlation in rLNs (P = 0.04) (Supplementary Figure 8A and 8B).
No correlation was observed between total CD68+ or PD1+ cells and miRNAs level in FLs and rLNs.
Foxp3+ cell counts are lower in relapsed FL
Clinico-pathological characteristics of FL patients were described in Table 2. Foxp3+, PD1+ and CD68+ cells were compared in relapsed and non-relapsed FLs. PD1+ and CD68+ cells were not significantly different between relapsed and non-relapsed FLs (P > 0.1). Instead, intrafollicular, interfollicular and total Foxp3+ cells were less represented in relapsed than in non-relapsed FLs (P = 0.01, P = 0.01, P = 0.003, respectively) (Figure 7A).
Table 2: FL patients, drug treatments, IHC markers and clinical status
FL |
Grade |
Age |
FLIPI |
Treatment |
IHC Bcl2 |
IHC Bcl6 |
Clinical status |
|---|---|---|---|---|---|---|---|
T15 |
1 |
52 |
Low |
No treatment |
2 |
2 |
NR |
T16 |
1 |
41 |
Low |
CHOP |
2 |
1 |
NR |
T17 |
1 |
50 |
Low |
RT |
2 |
1 |
R |
T18 |
1 |
48 |
Intermediate |
DHAP |
2 |
1 |
R |
T19 |
1 |
47 |
Intermediate |
CHOP + DHAP |
2 |
1 |
NR |
T20 |
1 |
76 |
High |
FluCy |
0 |
2 |
NR |
T21 |
1 |
51 |
Intermediate |
APO + DHAP |
1 |
1 |
NR |
T22 |
1 |
47 |
Low |
CHOP |
1 |
1 |
NR |
T23 |
2 |
37 |
Intermediate |
CHOP + DHAP |
2 |
2 |
NR |
T24 |
2 |
65 |
High |
CHOP |
2 |
1 |
R |
T25 |
2 |
60 |
High |
CHOP + DHAP |
1 |
1 |
NR |
T26 |
2 |
55 |
Intermediate |
CHOP |
2 |
1 |
NR |
T28 |
2 |
65 |
High |
CHOP + DHAP |
2 |
1 |
NR |
T30 |
3a |
70 |
High |
CHOP + RT |
1 |
1 |
NR |
T31 |
3a |
64 |
High |
CLB + CHOP |
0 |
2 |
R |
T33 |
3a |
n.a. |
n.a. |
n.a. |
2 |
1 |
n.a. |
T34 |
3a |
71 |
High |
CLB |
1 |
1 |
R |
T35 |
3a |
69 |
High |
CHOP |
2 |
1 |
NR |
T37 |
3a |
56 |
Low |
Surgery |
2 |
1 |
R |
T38 |
3a |
47 |
Low |
CHOP |
1 |
1 |
R |
T39 |
3a |
56 |
Intermediate |
APO + DHAP |
2 |
1 |
NR |
T40 |
3a |
67 |
Low |
RT |
2 |
1 |
R |
T42 |
3b |
n.a. |
n.a. |
n.a. |
1 |
1 |
n.a. |
T43 |
3b |
70 |
High |
Steroids |
1 |
1 |
n.a. |
T44 |
3b |
n.a. |
n.a. |
n.a. |
1 |
1 |
n.a. |
T45 |
3b |
77 |
Intermediate |
CLB |
0 |
1 |
NR |
n.a., not available; R, relapsed; NR, not relapsed. IHC: 0, absent; 1, 1+, weak stain; 2, 2+ strong stain; CHOP: cyclophosphamide, adriamycin, vincristine and prednisone-based regimen; APO: adriamycin, vincristine and prednisone-based regimen; DHAP: dexametasone, high dose-cytarabin and cisplatin-based regimen; FluCy: fludarabin and cyclophosphamide-based regimen; CLB: clorambucil; RT: radiotherapy.
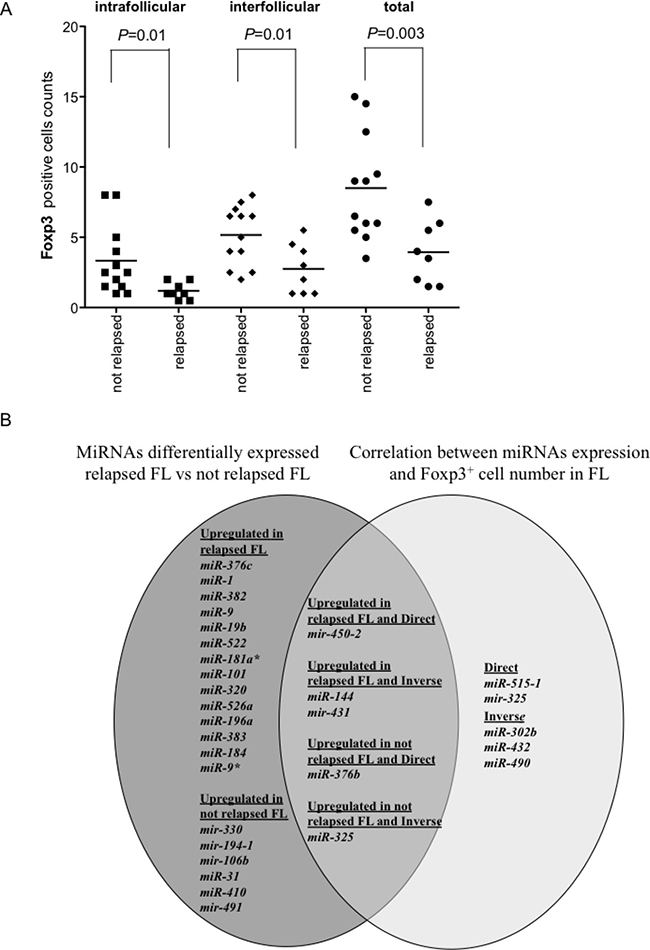
Figure 7: Foxp3+ cell counts in relapsed and not relapsed FLs and Venn diagram of miRNAs upregulated in relapsed or not relapsed FLs and of miRNAs correlated directly or inversely with Foxp3+ cell counts. Number of Foxp3+ cells in each follicular lymphoma (FL) was the average of cells in two consecutive slides. The average number of positive cells was normalized on the whole number of nuclei present in a slide. (A) Comparison of intrafollicular, interfollicular and total Foxp3+ cells in relapsed and not relapsed FLs. P values were calculated by t-test. (B) Venn diagram of miRNAs upregulated in relapsed or not relapsed FLs (dark grey) and the intersection (intermediate grey) with miRNAs correlated directly or inversely with Foxp3+ cell counts (light grey).
The comparison between miRNAs involved in FL relapse and miRNAs correlating with Foxp3+ cell counts in FLs was summarized by Venn diagram (Figure 7B). Five miRNAs intertwined the two pools. Mir-450-2 was upregulated in relapsed FLs and correlated positively with Foxp3+ cells. MiR-144 and mir-431 were upregulated in relapsed FLs and correlated inversely with Foxp3+ cells. MiR-376b was upregulated in not relapsed FLs and correlated directly with Foxp3+ cells. Finally, MiR-325 was upregulated in not relapsed FL and correlated inversely with Foxp3+ cells.
Foxp3+ cell counts associate with shorter time to relapse
PD1+ and CD68+ cells did not associate with time to relapse in FLs. Regarding Foxp3+ cells, FLs were subdivided in two categories according to the Foxp3+ cells, higher and lower than the median. FLs with lower Foxp3+ cells associated with a shorter time to relapse (P = 0.05) (Figure 8), with a median time to relapse of 2000 days.
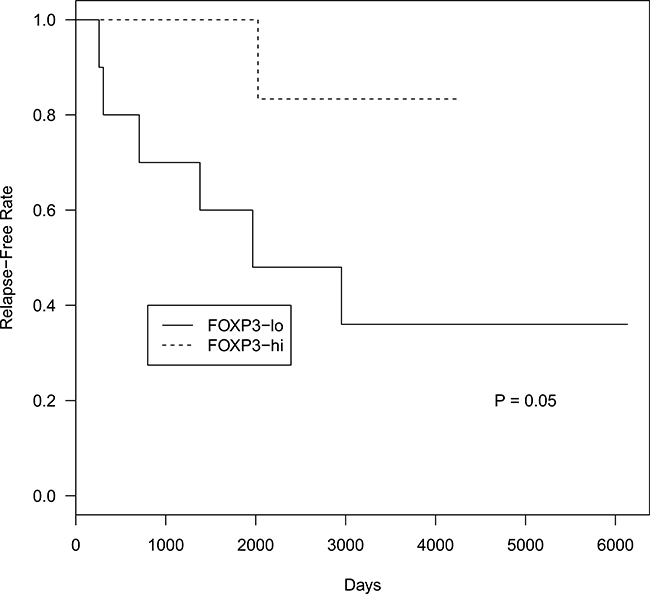
Figure 8: Time to relapse probability curves based on total Foxp3+ cell counts in FL patients. Top curve represents FL cases with total Foxp3+ cells higher than median. Bottom curve represents FL cases with total Foxp3+ cells lower than median.
DISCUSSION
The miRNA network supervises the cell proteome during lymphopoiesis and turned out essential in lymphomagenesis [32–34]. Although the importance of miRNA functions in the regulation of cellular processes is clear, both miRNAs deregulation and its possible use as a biomarker in FLs have been only partially addressed.
The tumor microenvironment provides supportive signaling for the proliferation of FL cells, as well as protection against attack from the immune system [12, 35, 36]. Several results have drawn attention to the host immune response in FLs [12]. FLs harbor up to 50% of T-cells [25]. Altered ratios between B-cells and other immune cells in FLs compared to rLNs would drive evident modifications of the level of molecules (i.e. mRNA and miRNAs) shared with lymphoma cells. Accordingly, two different gene expression signatures, which correlated with different outcomes, were identified in FLs [8]. A group included long surviving FLs expressing T-cell-derived genes; another group included FLs with worse prognosis and enriched of genes typical of dendritic cells and macrophages.
The molecular heterogeneity observed in our FLs and rLNs series by miRNA profiling (Figure 1) was clear in previous studies reporting the comparison of miRNA expression profiles in FLs and rLNs [21, 37, 38] but not in another study [23]. To test the hypothesis that the miRNA profile of FLs implies an immune signature, we compared FLs and pure T-cells, more abundant among the immune cells infiltrating the FLs. Similarly to what observed by gene expression data, FLs split into two clusters. The FL-HT cluster included miRNAs expressed at higher level in both FLs and T-cells (miRNA cluster 1). The higher level of CD3 mRNA in this group of FLs point to T-cells as main driver of this signature. These findings suggest that the clonal expansion of abnormal B-cells alters the immune response microenvironment, leaving a trace in the miRNA profile of FLs.
Being expressed at higher level in FL than in pure T-cells, both the cluster 2 miRNAs (overexpressed in FL-HT group) and the cluster 3 miRNAs (overexpressed in FL-LT group) could not be ascribed to a T-cell signature. Instead, miRNAs of the cluster 2 (mainly members of the tricistrons miR-199b/100-125b) were found expressed at higher level in NK cells, some myeloid cell lineages and in CD4+ rather than CD8+ T-cells [39]. Tumor cells suppress the antitumor immune response from helper T-cells, cytotoxic T lymphocytes, and macrophages by various mechanisms [40]. Consequently, the feature signatures of the two FL groups might recapitulate how the malignant B-cells alter the function of T-cells in the FL microenvironment in order to escape immunosurveillance.
Pathway enrichment analysis on target genes of miRNA cluster 2 showed a significant enrichment in pathways related to B-cells and T-cells activation and development as well as to cell cycle and control by MYC. For miRNA cluster 3, we found enrichment in pathways connected with mononocytes, macrophages, T helper and B-cell activation, as well as NF-kB, UV response and apoptosis. These results are reminiscent of the identification of two groups of FLs with different prognosis based on the gene expression signatures [8].
The two members of the cluster 1 miR-342 and miR-370 strongly correlated with two representative markers of T-cells CD3 and CD28. A connection between miR-342 and T-cells infiltrating the lymphomas was previously reported. Mir-342 was reported to be higher in T-cells than B-cells and in naive and Th17 CD4+ than in CD8+ T-cells [41, 42]. Moreover, MiR-342-3p expression associated the CD3+ T-cells in ALK- anaplastic large cell lymphoma [43]. Since B-cell lymphomas may display an increased CD4+/CD8+ T-cells ratio with respect to rLNs, it is possible that in FLs miR-342 represents a signature associated with CD4+ T-cells [35]. Abnormal number and activity of Th17 cells has been associated to autoimmune disorders and inflammation [44]. In B-cell lymphomas, altered B-cells might skew the balance of Treg cells and Th17 cells versus the increase of inhibitory microenvironment [45].
Various miRNAs reported to be differentially expressed in FLs play regulatory roles in lymphomagenesis [46]. We identified a set of 27 miRNAs differentially expressed between FLs and rLNs. Among these, miR-9*, miR-16, miR-195, miR-15b and miR-374, but not miR-30-5p, were previously found overexpressed in FL compared to rLNs [23]. Moreover, miR-9* and miR-155 have been found overexpressed also in B-cells purified from FLs with respect to those from rLNs [24]. Among 17 miRNAs upregulated in FLs, miR-29 family, miR-195, miR-15/16, miR-30 family, miR-146a are known to be downregulated as a consequence of MYC activity on their promoters [47]. Most of miRNAs upregulated and downregulated in FLs are equally modulated in germinal centre B-cells in comparison to naive B-cells (our unpublished data).
We identified a subset of seven miRNAs which discriminated grade 1 from grade 3a FLs. These data represent the first description of an increased alteration of miRNAs from low to high FL grade. The direct comparison between FLs of different grade did not return significant results, possibly in consequence of the low number of cases and the heterogeneity previously cited. Among the miRNAs altered in grade 3, miR-320 downregulation could be related to the reported increase of its gene target MCL1, which strongly associated with FL onset [48, 49].
MiRNA biogenesis is a complex process regulated at multiple stages [18]. The mechanisms responsible for the alteration of miRNA processing are not well known, in particular in lymphomas. Among all miRNAs here examined for the proportion 5p/3p, we found that miR-9-2/miR-9* and miR-21/miR-21* proportions changed in FLs with respect to rLNs. The peculiarity of miR-9 in FL was previously noticed. In fact, particularly high interindividual differences of MIRN9 and MIRN9* in FL was observed [23]. MiR-9 was also one of the 26 miRNAs which showed an altered proportion between different miRNA forms in primary effusion lymphoma [50]. Since miR-9-2 gene was reported to be methylated in normal CD19+ B-cells [51], the lymphomagenesis could imply changes of miR-9-2 methylation and of its expression. It is not yet demonstrated whether miR-9 alteration point to a etiologic field defect for FL genesis [52].
The level of specific miRNAs associated with clinical behaviour in our cohort of FL patients [23]. We could not associate miRNA expression and overall survival due to the low number of deaths in our series. Instead, we identified a set of 26 miRNA differentially expressed between relapsed and non-relapsed FL. None of the 26 miRNAs overlapped with those reported in the literature to be associated with FL outcome. Among miRNAs discriminating relapsed from non-relapsed FLs, several have been linked to drug resistance in lymphomas and other cancer types.
Malignant B-cells can attract Tregs or suppressive cells, thus influencing the patterns of miRNA expressed in FL tissues. Foxp3+ Tregs are relevant for B-cell development as they control the germinal centre response [28]. We observed a direct correlation of four miRNAs and an inverse correlation of six miRNAs with total Foxp3+ cells in FLs. Notably, miRNAs level and PD1+ or CD68+ cells did not show any correlation, thus supporting the idea that Foxp3+ cells are those mainly involved in setting of miRNAs level. MiR-144 and miR-302b level, which correlated inversely with Foxp3+ cells in FL, lost this trend in rLNs. MiR-144 was shown to target BCL6 in diffuse large B-cell lymphoma [53]. Since relapsed FLs showed higher level of miR-144 and lower number of Foxp3+ cells, it can be hypothesized that both events might cooperate in supporting the proliferation of FL cells.
In FL, Foxp3+ Treg cells might represent from less than 5% up to 25% of the germinal center T-cells [10, 28]. Treg cells might have significant clinical relevance in FL since they can serve as tumor-killing cells. Lower number of Tregs correlated with refractory disease, transformation, and aggressive histology of FL [10, 14, 54]. However, other studies failed to confirm this conclusion [55, 56]. Rather than Treg cells total content, the architectural distribution and pattern of Foxp3+ cells correlated with the outcome of patients [55]. Intrafollicular Foxp3+ cells pattern, but not a diffuse pattern, was found associated with poorer overall survival in FL patients. Our data confirmed the clinical relevance of both total number and pattern of Foxp3+ cells in FL patients. In our FL series, intrafollicular, interfollicular and total Foxp3+ cells were lower in relapsed than not relapsed FLs. Moreover, interfollicular Foxp3+ cells in grade 3 FLs, but not in FLs of inferior grade, were lower than in rLNs. Importantly, lower than median Foxp3+ cells associated with shorter time to relapse of FL patients. Taken together, our results and those reported by Farinha et al. [55] suggest that the immuno-architecture of the microenvironment around transformed B-cells impacts on patient outcome in terms of time to relapse and of survival of FL patients.
PD1 receptor was demonstrated to suppress cytokines signaling in lymphoma-infiltrating T-cells [57]. PD1 is expressed by intratumoral CD4+ follicular helper T-cells and from exhausted cytotoxic T-cells [58]. A diffuse pattern of PD1 staining associated with a shorter time before transformation and inferior overall survival [13, 58]. At variance with these authors, according to our data no association was established between the pattern of PD1 positivity and time of relapse.
TFH cells and TFR cells subtypes are recognized essential for the physiological regulation of humoral immunity. TFR cells are immune suppressive by limiting TFH cells action and GC B-cells number [28, 59, 60]. The specification of the two T-cell populations is driven by mechanisms operated by lineage-specific transcription factor as T-bet and BCL6 [61] and by mesenchymal stem cells [62]. It is believed that TFR cells arise from thymic regulatory T-cells. However, absence of Foxp3 expression in GC suppressive T-cells suggested also a possible origin of TFR from TFH cells [63]. TFR cells were reported to express significant amounts of the prototypic TFH genes CXCR5, CXCL13, ICOS, BCL6 and PD1 in mouse [28]. Foxp3+PD1+CXCL13+ TFR cells were detected in various mice and human settings [27–29, 64, 65]. Foxp3+PD1+ cells have been previously reported to be rare or absent in human FL, diffuse large B-cell lymphoma, Hodgkin’s disease, T-cell lymphoma normal tonsils and rLN [54, 66–69]. Small amount (1–4%) or absence of Foxp3+PD1+ cells were found in breast cancer tissues [70, 71], suggesting the possibility that the phenotype of TFR cells in lymphomas and cancer does not overlap those in other physiological contexts [62]. We observed clear staining for PD1 or CXCL13 in sparse cells within the neoplastic follicles of FL and the two signals were almost always independent from that of Foxp3. Only one FL cases showed a significant number of double stained cells. The general rarity of TFR cells in FL and rLN could be suggestive of a skewed T-cell differentiation in sustaining the immune activation.
The recruitment of CD8+ cells into the lymph nodes was found protective of a disease-free survival in FL patients [25]. We found that CD8+ cells distributed mainly in interfollicular areas and were in higher number in FL than in rLN. In our series CD8+ cell counts did not associate with relapse and not relapsed status of FL patients.
In conclusion, our data reveal new miRNAs involved in FL in comparison with rLN. The miRNA signature in FL was interpreted in light of the significant infiltration of immune cells typical of a quote of FLs. In this regard, lower infiltration by Foxp3+ cells is a feature of relapsed FLs and also associated with shorter time to relapse. High levels of miR-144 and mir-431, which are upregulated in relapsed FL and inversely correlated with Foxp3+ cell number, and of miR-376b, which is upregulated in non-relapsed FL and is directly correlated with Foxp3+ cells, might complement low Foxp3+ cell counts as a relapse risk predictor in FL patients. Multivariate analysis to assess combination of Foxp3+ cell number and miR-144, mir-431, miR-376b levels as markers of FL relapse was not possible in this context owing to the low number of events. The diagnostics implication of the combined measure of miRNAs and Foxp3 for the risk of relapse needs to be confirmed on a larger FL cohort.
MATERIALS AND METHODS
Samples
FL samples were obtained from patients accessing care in the Hematology section of the University of Verona Hospital compound (Table 1). All biopsy and blood samples were obtained after informed consent approved by Ethical Committee of AOUI of Verona (N. 1828, May 12, 2010 “Institution of cell and tissue collection for biomedical research in Onco-Hematology”). Twenty-six patients with FL grade 1 or 2 or 3 (subdivided in 3a and 3b) for whom fresh/frozen pre-treatment samples were available were included. Twenty-one patients were treated with first-line chemotherapy (cyclophosphamide or doxorubicin or etoposide or prednisone or clorambucil), radiotherapy or surgery. One patient received no chemotherapy and another one only radiotherapy. Clinical follow-up was available for 21 FL patients. The average follow-up of patients was of 13.5 years. Time to relapse was defined as the time interval between complete remission achievement and relapse. Control samples were 12 lymph nodes (rLNs) characterized by reactive follicular hyperplasia. CD4+ and CD8+ T-cells were isolated from blood of two healthy donors by Ficoll-Paque (Pharmacia, Germany) density gradient centrifugation, followed by positive immune affinity magnetic sorting.
Immunohistochemistry (IHC)
Number of Foxp3+, PD1+, CD68+ and CD8+ cells were quantified in whole-tissue sections of all samples using an automated scanning microscope and image analysis system (S.CORE Web Based Image Analysis, S.CO LifeScience, Germany). Double staining for Foxp3-PD1 and Foxp3-CXCL13 was performed using an automated stainer (Leica BondMax) following standard protocols, as previously described [72]. Ab for Foxp3 was revealed by horsereadish peroxidase (nuclear). The slides were incubated with a second Ab then revealed by alcaline phosphatase (cytoplasmic). For each slide, “intrafollicular”, “interfollicular” and total positive cells were determined. The number of positive cells in each FL and rLN sample was the average number of cells in two consecutive slides. The average number of positive cells used for subsequent analysis was normalized on the whole number of nuclei present in a slide.
Microarrays
MiRNAs labeling and hybridization were performed using 5 μg total RNA, as described [73]. MiRNAs were named according with the old nomenclature reported in miRBase (www.mirbase.org). MiR-X was the 5p or 3p form of a miRNA and mir-X is the 5p or 3p remaining form of MiR-X .
Quantitative RT-PCR
Validation of differentially expressed miRNAs and mRNA quantification was performed by quantitative RT-PCR as described in Malpeli et al. [34]. Taqman assays and oligonucleotide primers used are listed in Supplementary Table 1.
Data analysis
Microarrays data analyses were performed as described in Malpeli et al. [34]. Microarray data sets are available on the ArrayExpress under http://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-5844. To select significant differences among expressed gene or categories of samples, we performed using Anova tests. Correlation coefficients between expression of 5p and 3p miRNA forms were calculated using Deming linear regression. Pathway analysis on predicted miRNA targets was performed by Gene Set Enrichment Analysis (GSEA). Experimentally validated miRNA targets (strong evidences) taken from MirTarBase at http://mirtarbase.mbc.nctu.edu.tw [74] were submitted to GSEA http://software.broadinstitute.org/gsea/msigdb/index.jsp [75, 76]. More significant Immunological Signatures and Hallmark Gene Sets were considered.
A more extensive description of the methods for Nucleic acids, Quantitative RT-PCR, Immunohistochemistry, Microarrays and Data Analysis can be found in Supplementary Material.
Abbreviations
FL: follicular lymphoma; IHC: immunohistochemistry; rLN: reactive lymph node; FDR: false discovery rate; FL-HT: follicular lymphoma high T-cells; FL-LT: follicular lymphoma low T-cells; Treg: regulatory T-cells; TFR: regulatory follicular T-cells; TFH: follicular helper T-cells.
Author contributions
G.M. designed the study, performed molecular studies, interpreted data, and wrote the manuscript; S.B. designed the study, analyzed and interpreted data, wrote the manuscript; C.G. collected clinico-pathological data and revised the manuscript; S.Z., A.B., MT.S., PT.K. and CM.C. revised the manuscript; M.K. designed the study and revised the manuscript; A.S. designed the study, interpreted data and revised the manuscript; A.Z. performed pathological revision and wrote the manuscript. All authors have read the manuscript and approved the submission.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
This work was supported by Associazione Italiana Ricerca sul Cancro (AIRC grant n. 12182).
REFERENCES
1. Al-Tourah AJ, Gill KK, Chhanabhai M, Hoskins PJ, Klasa RJ, Savage KJ, Sehn LH, Shenkier TN, Gascoyne RD, Connors JM. Population-based analysis of incidence and outcome of transformed non-Hodgkin’s lymphoma. J Clin Oncol. 2008; 26:5165–5169.
2. Glas AM, Knoops L, Delahaye L, Kersten MJ, Kibbelaar RE, Wessels LA, van Laar R, van Krieken JH, Baars JW, Raemaekers J, Kluin PM, van’t Veer LJ, de Jong D. Gene-expression and immunohistochemical study of specific T-cell subsets and accessory cell types in the transformation and prognosis of follicular lymphoma. J Clin Oncol. 2007; 25:390–398.
3. Berg T, Thoene S, Yap D, Wee T, Schoeler N, Rosten P, Lim E, Bilenky M, Mungall AJ, Oellerich T, Lee S, Lai CK, Umlandt P, et al. A transgenic mouse model demonstrating the oncogenic role of mutations in the polycomb-group gene EZH2 in lymphomagenesis. Blood. 2014; 123:3914–3924.
4. Guo S, Chan JK, Iqbal J, McKeithan T, Fu K, Meng B, Pan Y, Cheuk W, Luo D, Wang R, Zhang W, Greiner TC, Chan WC. EZH2 mutations in follicular lymphoma from different ethnic groups and associated gene expression alterations. Clin Cancer Res. 2014; 20:3078–3086.
5. Lunning MA, Green MR. Mutation of chromatin modifiers; an emerging hallmark of germinal center B-cell lymphomas. Blood Cancer J. 2015; 5:e361.
6. Okosun J, Bodor C, Wang J, Araf S, Yang CY, Pan C, Boller S, Cittaro D, Bozek M, Iqbal S, Matthews J, Wrench D, Marzec J, et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat Genet. 2014; 46:176–181.
7. Pastore A, Jurinovic V, Kridel R, Hoster E, Staiger AM, Szczepanowski M, Pott C, Kopp N, Murakami M, Horn H, Leich E, Moccia AA, Mottok A, et al. Integration of gene mutations in risk prognostication for patients receiving first-line immunochemotherapy for follicular lymphoma: a retrospective analysis of a prospective clinical trial and validation in a population-based registry. Lancet Oncol. 2015; 16:1111–1122.
8. Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC, Fisher RI, Braziel RM, Rimsza LM, Grogan TM, Miller TP, LeBlanc M, Greiner TC, et al. Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N Engl J Med. 2004; 351:2159–2169.
9. Glas AM, Kersten MJ, Delahaye LJ, Witteveen AT, Kibbelaar RE, Velds A, Wessels LF, Joosten P, Kerkhoven RM, Bernards R, van Krieken JH, Kluin PM, van’t Veer LJ, et al. Gene expression profiling in follicular lymphoma to assess clinical aggressiveness and to guide the choice of treatment. Blood. 2005; 105:301–307.
10. Carreras J, Lopez-Guillermo A, Fox BC, Colomo L, Martinez A, Roncador G, Montserrat E, Campo E, Banham AH. High numbers of tumor-infiltrating FOXP3-positive regulatory T cells are associated with improved overall survival in follicular lymphoma. Blood. 2006; 108:2957–2964.
11. Farinha P, Masoudi H, Skinnider BF, Shumansky K, Spinelli JJ, Gill K, Klasa R, Voss N, Connors JM, Gascoyne RD. Analysis of multiple biomarkers shows that lymphoma-associated macrophage (LAM) content is an independent predictor of survival in follicular lymphoma (FL). Blood. 2005; 106:2169–2174.
12. Fend F, Quintanilla-Martinez L. Assessing the prognostic impact of immune cell infiltrates in follicular lymphoma. Haematologica. 2014; 99:599–602.
13. Smeltzer JP, Jones JM, Ziesmer SC, Grote DM, Xiu B, Ristow KM, Yang ZZ, Nowakowski GS, Feldman AL, Cerhan JR, Novak AJ, Ansell SM. Pattern of CD14+ follicular dendritic cells and PD1+ T cells independently predicts time to transformation in follicular lymphoma. Clin Cancer Res. 2014; 20:2862–2872.
14. Tzankov A, Meier C, Hirschmann P, Went P, Pileri SA, Dirnhofer S. Correlation of high numbers of intratumoral FOXP3+ regulatory T cells with improved survival in germinal center-like diffuse large B-cell lymphoma, follicular lymphoma and classical Hodgkin’s lymphoma. Haematologica. 2008; 93:193–200.
15. Musilova K, Mraz M. MicroRNAs in B-cell lymphomas: how a complex biology gets more complex. Leukemia. 2015; 29:1004–1017.
16. Blahna MT, Hata A. Smad-mediated regulation of microRNA biosynthesis. FEBS Lett. 2012; 586:1906–1912.
17. Davis BN, Hilyard AC, Lagna G, Hata A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature. 2008; 454:56–61.
18. Ha M, Kim VN. Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol. 2014; 15:509–524.
19. De Tullio G, De Fazio V, Sgherza N, Minoia C, Serrati S, Merchionne F, Loseto G, Iacobazzi A, Rana A, Petrillo P, Silvestris N, Iacopino P, Guarini A. Challenges and opportunities of microRNAs in lymphomas. Molecules. 2014; 19:14723–14781.
20. Leich E, Zamo A, Horn H, Haralambieva E, Puppe B, Gascoyne RD, Chan WC, Braziel RM, Rimsza LM, Weisenburger DD, Delabie J, Jaffe ES, Fitzgibbon J, et al. MicroRNA profiles of t(14;18)-negative follicular lymphoma support a late germinal center B-cell phenotype. Blood. 2011; 118:5550–5558.
21. Di Lisio L, Sanchez-Beato M, Gomez-Lopez G, Rodriguez ME, Montes-Moreno S, Mollejo M, Menarguez J, Martinez MA, Alves FJ, Pisano DG, Piris MA, Martinez N. MicroRNA signatures in B-cell lymphomas. Blood Cancer J. 2012; 2:e57.
22. Lawrie CH, Chi J, Taylor S, Tramonti D, Ballabio E, Palazzo S, Saunders NJ, Pezzella F, Boultwood J, Wainscoat JS, Hatton CS. Expression of microRNAs in diffuse large B cell lymphoma is associated with immunophenotype, survival and transformation from follicular lymphoma. J Cell Mol Med. 2009; 13:1248–1260.
23. Roehle A, Hoefig KP, Repsilber D, Thorns C, Ziepert M, Wesche KO, Thiere M, Loeffler M, Klapper W, Pfreundschuh M, Matolcsy A, Bernd HW, Reiniger L, et al. MicroRNA signatures characterize diffuse large B-cell lymphomas and follicular lymphomas. Br J Haematol. 2008; 142:732–744.
24. Wang W, Corrigan-Cummins M, Hudson J, Maric I, Simakova O, Neelapu SS, Kwak LW, Janik JE, Gause B, Jaffe ES, Calvo KR. MicroRNA profiling of follicular lymphoma identifies microRNAs related to cell proliferation and tumor response. Haematologica. 2012; 97:586–594.
25. Laurent C, Muller S, Do C, Al-Saati T, Allart S, Larocca LM, Hohaus S, Duchez S, Quillet-Mary A, Laurent G, Brousset P, Valitutti S. Distribution, function, and prognostic value of cytotoxic T lymphocytes in follicular lymphoma: a 3-D tissue-imaging study. Blood. 2011; 118:5371–5379.
26. Canioni D, Salles G, Mounier N, Brousse N, Keuppens M, Morchhauser F, Lamy T, Sonet A, Rousselet MC, Foussard C, Xerri L. High numbers of tumor-associated macrophages have an adverse prognostic value that can be circumvented by rituximab in patients with follicular lymphoma enrolled onto the GELA-GOELAMS FL-2000 trial. J Clin Oncol. 2008; 26:440–446.
27. Chung Y, Tanaka S, Chu F, Nurieva RI, Martinez GJ, Rawal S, Wang YH, Lim H, Reynolds JM, Zhou XH, Fan HM, Liu ZM, Neelapu SS, et al. Follicular regulatory T cells expressing Foxp3 and Bcl-6 suppress germinal center reactions. Nat Med. 2011; 17:983–988.
28. Linterman MA, Pierson W, Lee SK, Kallies A, Kawamoto S, Rayner TF, Srivastava M, Divekar DP, Beaton L, Hogan JJ, Fagarasan S, Liston A, Smith KG, et al. Foxp3+ follicular regulatory T cells control the germinal center response. Nat Med. 2011; 17:975–982.
29. Wollenberg I, Agua-Doce A, Hernandez A, Almeida C, Oliveira VG, Faro J, Graca L. Regulation of the germinal center reaction by Foxp3+ follicular regulatory T cells. J Immunol. 2011; 187:4553–4560.
30. Kim CH, Lim HW, Kim JR, Rott L, Hillsamer P, Butcher EC. Unique gene expression program of human germinal center T helper cells. Blood. 2004; 104:1952–1960.
31. Alvaro T, Lejeune M, Salvado MT, Lopez C, Jaen J, Bosch R, Pons LE. Immunohistochemical patterns of reactive microenvironment are associated with clinicobiologic behavior in follicular lymphoma patients. J Clin Oncol. 2006; 24:5350–5357.
32. Arrate MP, Vincent T, Odvody J, Kar R, Jones SN, Eischen CM. MicroRNA biogenesis is required for Myc-induced B-cell lymphoma development and survival. Cancer Res. 2010; 70:6083–6092.
33. Kuchen S, Resch W, Yamane A, Kuo N, Li Z, Chakraborty T, Wei L, Laurence A, Yasuda T, Peng S, Hu-Li J, Lu K, Dubois W, et al. Regulation of microRNA expression and abundance during lymphopoiesis. Immunity. 2010; 32:828–839.
34. Malpeli G, Barbi S, Zupo S, Tosadori G, Scardoni G, Bertolaso A, Sartoris S, Ugel S, Vicentini C, Fassan M, Adamo A, Krampera M, Scupoli MT, et al. Identification of microRNAs implicated in the late differentiation stages of normal B cells suggests a central role for miRNA targets ZEB1 and TP53. Oncotarget. 2017; 8:11809–11826. https://doi.org/10.18632/oncotarget.14683.
35. Daussy C, Damotte D, Molina TJ, Roussel M, Fest T, Varin A, Perrot JY, Ouafi L, Merle-Beral H, Julia P, Fridman WH, Sautes-Fridman C, Fisson S. CD4:CD8 T-cell ratio differs significantly in diffuse large B-cell lymphomas from other lymphoma subtypes independently from lymph node localization. Int Trends Immun. 2013; 1:45–48.
36. Kiaii S, Clear AJ, Ramsay AG, Davies D, Sangaralingam A, Lee A, Calaminici M, Neuberg DS, Gribben JG. Follicular lymphoma cells induce changes in T-cell gene expression and function: potential impact on survival and risk of transformation. J Clin Oncol. 2013; 31:2654–2661.
37. Arribas AJ, Campos-Martin Y, Gomez-Abad C, Algara P, Sanchez-Beato M, Rodriguez-Pinilla MS, Montes-Moreno S, Martinez N, Alves-Ferreira J, Piris MA, Mollejo M. Nodal marginal zone lymphoma: gene expression and miRNA profiling identify diagnostic markers and potential therapeutic targets. Blood. 2012; 119:e9–e21.
38. Goswami RS, Atenafu EG, Xuan Y, Waldron L, Reis PP, Sun T, Datti A, Xu W, Kuruvilla J, Good DJ, Lai R, Church AJ, Lam WS, et al. MicroRNA signature obtained from the comparison of aggressive with indolent non-Hodgkin lymphomas: potential prognostic value in mantle-cell lymphoma. J Clin Oncol. 2013; 31:2903–2911.
39. Emmrich S, Rasche M, Schoning J, Reimer C, Keihani S, Maroz A, Xie Y, Li Z, Schambach A, Reinhardt D, Klusmann JH. miR-99a/100~125b tricistrons regulate hematopoietic stem and progenitor cell homeostasis by shifting the balance between TGFbeta and Wnt signaling. Genes Dev. 2014; 28:858–874.
40. Kridel R, Sehn LH, Gascoyne RD. Pathogenesis of follicular lymphoma. J Clin Invest. 2012; 122:3424–3431.
41. Lawrie CH, Saunders NJ, Soneji S, Palazzo S, Dunlop HM, Cooper CD, Brown PJ, Troussard X, Mossafa H, Enver T, Pezzella F, Boultwood J, Wainscoat JS, et al. MicroRNA expression in lymphocyte development and malignancy. Leukemia. 2008; 22:1440–1446.
42. Merkerova M, Belickova M, Bruchova H. Differential expression of microRNAs in hematopoietic cell lineages. Eur J Haematol. 2008; 81:304–310.
43. Liu C, Iqbal J, Teruya-Feldstein J, Shen Y, Dabrowska MJ, Dybkaer K, Lim MS, Piva R, Barreca A, Pellegrino E, Spaccarotella E, Lachel CM, Kucuk C, et al. MicroRNA expression profiling identifies molecular signatures associated with anaplastic large cell lymphoma. Blood. 2013; 122:2083–2092.
44. Pene J, Chevalier S, Preisser L, Venereau E, Guilleux MH, Ghannam S, Moles JP, Danger Y, Ravon E, Lesaux S, Yssel H, Gascan H. Chronically inflamed human tissues are infiltrated by highly differentiated Th17 lymphocytes. J Immunol. 2008; 180:7423–7430.
45. Yang ZZ, Novak AJ, Ziesmer SC, Witzig TE, Ansell SM. Malignant B cells skew the balance of regulatory T cells and TH17 cells in B-cell non-Hodgkin’s lymphoma. Cancer Res. 2009; 69:5522–5530.
46. Kozloski GA, Lossos IS. LymphomiRs: microRNAs with regulatory roles in lymphomas. Curr Opin Hematol. 2015; 22:362–368.
47. Chang TC, Yu D, Lee YS, Wentzel EA, Arking DE, West KM, Dang CV, Thomas-Tikhonenko A, Mendell JT. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat Genet. 2008; 40:43–50.
48. Ruiz-Vela A, Aggarwal M, de la Cueva P, Treda C, Herreros B, Martin-Perez D, Dominguez O, Piris MA. Lentiviral (HIV)-based RNA interference screen in human B-cell receptor regulatory networks reveals MCL1-induced oncogenic pathways. Blood. 2008; 111:1665–1676.
49. Zhou P, Levy NB, Xie H, Qian L, Lee CY, Gascoyne RD, Craig RW. MCL1 transgenic mice exhibit a high incidence of B-cell lymphoma manifested as a spectrum of histologic subtypes. Blood. 2001; 97:3902–3909.
50. O’Hara AJ, Vahrson W, Dittmer DP. Gene alteration and precursor and mature microRNA transcription changes contribute to the miRNA signature of primary effusion lymphoma. Blood. 2008; 111:2347–2353.
51. Wang LQ, Kwong YL, Kho CS, Wong KF, Wong KY, Ferracin M, Calin GA, Chim CS. Epigenetic inactivation of miR-9 family microRNAs in chronic lymphocytic leukemia--implications on constitutive activation of NFkappaB pathway. Mol Cancer. 2013; 12:173.
52. Malpeli G. Potential etiologic of the epigenetic field defect in the diseases and in cancer. Edorium J Pathol. 2015; 2:10–13.
53. Wang H, Wang A, Hu Z, Xu X, Liu Z, Wang Z. A Critical Role of miR-144 in Diffuse Large B-cell Lymphoma Proliferation and Invasion. Cancer Immunol Res. 2016; 4:337–344.
54. Ahearne MJ, Bhuller K, Hew R, Ibrahim H, Naresh K, Wagner SD. Expression of PD-1 (CD279) and FoxP3 in diffuse large B-cell lymphoma. Virchows Arch. 2014; 465:351–358.
55. Farinha P, Al-Tourah A, Gill K, Klasa R, Connors JM, Gascoyne RD. The architectural pattern of FOXP3-positive T cells in follicular lymphoma is an independent predictor of survival and histologic transformation. Blood. 2010; 115:289–295.
56. Sweetenham JW, Goldman B, LeBlanc ML, Cook JR, Tubbs RR, Press OW, Maloney DG, Fisher RI, Rimsza LM, Braziel RM, Hsi ED. Prognostic value of regulatory T cells, lymphoma-associated macrophages, and MUM-1 expression in follicular lymphoma treated before and after the introduction of monoclonal antibody therapy: a Southwest Oncology Group Study. Ann Oncol. 2010; 21:1196–1202.
57. Myklebust JH, Irish JM, Brody J, Czerwinski DK, Houot R, Kohrt HE, Timmerman J, Said J, Green MR, Delabie J, Kolstad A, Alizadeh AA, Levy R. High PD-1 expression and suppressed cytokine signaling distinguish T cells infiltrating follicular lymphoma tumors from peripheral T cells. Blood. 2013; 121:1367–1376.
58. Yang ZZ, Grote DM, Ziesmer SC, Xiu B, Novak AJ, Ansell SM. PD-1 expression defines two distinct T-cell sub-populations in follicular lymphoma that differentially impact patient survival. Blood Cancer J. 2015; 5:e281.
59. Hilchey SP, Rosenberg AF, Hyrien O, Secor-Socha S, Cochran MR, Brady MT, Wang JC, Sanz I, Burack WR, Quataert SA, Bernstein SH. Follicular lymphoma tumor-infiltrating T-helper (T(H)) cells have the same polyfunctional potential as normal nodal T(H) cells despite skewed differentiation. Blood. 2011; 118:3591–3602.
60. Wallin EF, Jolly EC, Suchanek O, Bradley JA, Espeli M, Jayne DR, Linterman MA, Smith KG. Human T-follicular helper and T-follicular regulatory cell maintenance is independent of germinal centers. Blood. 2014; 124:2666–2674.
61. Oestreich KJ, Huang AC, Weinmann AS. The lineage-defining factors T-bet and Bcl-6 collaborate to regulate Th1 gene expression patterns. J Exp Med. 2011; 208:1001–1013.
62. Brady MT, Hilchey SP, Hyrien O, Spence SA, Bernstein SH. Mesenchymal stromal cells support the viability and differentiation of follicular lymphoma-infiltrating follicular helper T-cells. PLoS One. 2014; 9:e97597.
63. Aloulou M, Carr EJ, Gador M, Bignon A, Liblau RS, Fazilleau N, Linterman MA. Follicular regulatory T cells can be specific for the immunizing antigen and derive from naive T cells. Nat Commun. 2016; 7:10579.
64. Chen X, Fosco D, Kline DE, Meng L, Nishi S, Savage PA, Kline J. PD-1 regulates extrathymic regulatory T-cell differentiation. Eur J Immunol. 2014; 44:2603–2616.
65. Sage PT, Ron-Harel N, Juneja VR, Sen DR, Maleri S, Sungnak W, Kuchroo VK, Haining WN, Chevrier N, Haigis M, Sharpe AH. Suppression by TFR cells leads to durable and selective inhibition of B cell effector function. Nat Immunol. 2016; 17:1436–1446.
66. Bruneau J, Canioni D, Renand A, Marafioti T, Paterson JC, Martin-Garcia N, Gaulard P, Delfau MH, Hermine O, Macintyre E, Brousse N, Asnafi V. Regulatory T-cell depletion in angioimmunoblastic T-cell lymphoma. Am J Pathol. 2010; 177:570–574.
67. Carreras J, Lopez-Guillermo A, Roncador G, Villamor N, Colomo L, Martinez A, Hamoudi R, Howat WJ, Montserrat E, Campo E. High numbers of tumor-infiltrating programmed cell death 1-positive regulatory lymphocytes are associated with improved overall survival in follicular lymphoma. J Clin Oncol. 2009; 27:1470–1476.
68. Yamamoto R, Nishikori M, Kitawaki T, Sakai T, Hishizawa M, Tashima M, Kondo T, Ohmori K, Kurata M, Hayashi T, Uchiyama T. PD-1-PD-1 ligand interaction contributes to immunosuppressive microenvironment of Hodgkin lymphoma. Blood. 2008; 111:3220–3224.
69. Yang ZZ, Novak AJ, Ziesmer SC, Witzig TE, Ansell SM. CD70+ non-Hodgkin lymphoma B cells induce Foxp3 expression and regulatory function in intratumoral CD4+CD25 T cells. Blood. 2007; 110:2537–2544.
70. Chakraborty S, Panda AK, Bose S, Roy D, Kajal K, Guha D, Sa G. Transcriptional regulation of FOXP3 requires integrated activation of both promoter and CNS regions in tumor-induced CD8(+) Treg cells. Sci Rep. 2017; 7:1628.
71. Ghebeh H, Barhoush E, Tulbah A, Elkum N, Al-Tweigeri T, Dermime S. FOXP3+ Tregs and B7-H1+/PD-1+ T lymphocytes co-infiltrate the tumor tissues of high-risk breast cancer patients: Implication for immunotherapy. BMC Cancer. 2008; 8:57.
72. Zamo A, Malpeli G, Scarpa A, Doglioni C, Chilosi M, Menestrina F. Expression of TP73L is a helpful diagnostic marker of primary mediastinal large B-cell lymphomas. Mod Pathol. 2005; 18:1448–1453.
73. Liu CG, Calin GA, Meloon B, Gamliel N, Sevignani C, Ferracin M, Dumitru CD, Shimizu M, Zupo S, Dono M, Alder H, Bullrich F, Negrini M, et al. An oligonucleotide microchip for genome-wide microRNA profiling in human and mouse tissues. Proc Natl Acad Sci U S A. 2004; 101:9740–9744.
74. Chou CH, Chang NW, Shrestha S, Hsu SD, Lin YL, Lee WH, Yang CD, Hong HC, Wei TY, Tu SJ, Tsai TR, Ho SY, Jian TY, et al. miRTarBase 2016: updates to the experimentally validated miRNA-target interactions database. Nucleic Acids Res. 2016; 44:D239–247.
75. Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E, Ridderstrale M, Laurila E, Houstis N, Daly MJ, Patterson N, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003; 34:267–273.
76. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005; 102:15545–15550.

