INTRODUCTION
Breast cancer (BC) remains one of the most common diagnosed malignancies and the first leading cause of death from cancer in women worldwide [1]. For the year of 2017, it is estimated in the United States that approximately 252,710 female patients would be diagnosed with BC and 40,610 would die from it [2]. It has been well established that breast cancer is a heterogeneous disease in which the gene-expression profiles vary between individuals [3, 4]. Triple negative breast cancer (TNBC) is a subtype of breast cancer which lacks the expressions of estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor-2 (HER2). Although the incidence of TNBC only accounts for a small proportion (10–20%) of all BC [5], it has the worst prognosis and mortality risk than other types of breast cancer in the first three and five years [6–8]. However, for the the absence of ER, PR and HER2 expression, only a few adjuvant treatments, like conventional surgery, chemotherapy and radiotherapy, could provide benefits for patients with TNBC. Therefore, it is important to develop an accurate and practical prognostic predictive model to assist both short-term and long-term treatment decisions for TNBC patients.
Tumor staging system is frequently used to predict the prognosis of patients. The most commonly used staging system for breast cancer is the eighth edition of American Joint Committee on Cancer (8th AJCC) TNM classification [9]. The 8th TNM classification stratifies the patients according to the extent of tumor size (T stage), number of positive lymph nodes (N stage) and distant metastasis (M stage). Although staging system such as the AJCC system is helpful, it is more applicable to a general population rather than individual patients, with the survival of TNBC patients in the same stage may still vary greatly.
Currently, nomograms have been proposed as a reliable tool to quantify risk by incorporating and illustrating important factors for tumor prognosis [10]. Through incorporating significant factors, nomograms could provide a numerical probability of a clinical event, such as overall survival (OS), which is tailored to the profile of individual patient. In several types of cancers, such as intrahepatic cholangiocarcinoma [11–13], hepatocellular carcinoma [14], non-small-cell lung cancer [15] and gastric cancer [16], nomograms have been demonstrated to be more precise when compared with the traditional TNM staging systems. To the best of our knowledge, only one previous report by Dai et al [17] established a nomogram to predict the long-term survival for triple negative breast cancer patients, while this nomogram only involves 247 patients and mainly focused on the value of pre-therapeutic CEA and CA15-3 levels. In addition to this, the study also did not compare the nomogram with the traditional TNM staging systems. Therefore, the present study consisting of 604 TNBC patients was conducted, aiming to develop a simple and practical nomogram for patients with TNBC and compare the performance of this model with the currently available staging system.
RESULTS
Characteristic of the study subject
A total of 486 patients diagnosed with TNBC between August 2006 and July 2012 in Affiliated Union Hospital of Fujian Medical University were retrospectively enrolled in this study. The patients who had received prior radiotherapy or chemotherapy (n=58) and lost to follow up (n=24) were excluded. 404 TNBC patients were finally selected in the training cohort. For validation cohort, we enrolled 283 patients and a total of 200 patients between August 2012 and July 2014 were selected according to our inclusive and exclusive criteria. The baseline and clinicopathologic characteristics of the patients in training and validation cohort were listed in Table 1. For patients in the training cohort, the median age at diagnosis was 49.8 years (range, 29 to 65 years). The median OS was 54.6 months (range 4.0 to 131.2 months), and the 1-, 3-, and 5-year OS rates were 94.3%, 74.2% and 60.7%.
Table 1: Demographic and clinicopathologic features in the training and validation cohort of patients with triple negative breast cancer
Characteristics |
Training cohort (n=404) no. (%) |
Validation cohort (n=200) no. (%) |
|---|---|---|
Age, y (mean ± SD) |
49.8 ± 10.8 |
51.1 ± 10.9 |
BMI, kg/m2 (mean ± SD) |
22.6 ± 2.8 |
22.8 ± 3.1 |
Tumor size, cm (mean ± SD) |
2.6 ± 1.3 |
2.5 ± 1.2 |
No. of positive lymph nodes (mean ± SD) |
1.5 ± 3.7 |
1.3 ± 3.4 |
No. of examined lymph nodes (mean ± SD) |
20.6 ± 6.2 |
18.5 ± 7.9 |
Age at menarche |
|
|
≦16 |
329 (81.4) |
157 (78.5) |
>16 |
75 (18.6) |
43 (21.5) |
Menopausal status |
|
|
Premenopausal |
222 (55.0) |
99 (49.5) |
Postmenopausal |
178 (44.0) |
95 (47.5) |
Unnatural menopausea |
4 (1.0) |
6 (3.0) |
Age at first live birth |
|
|
≦25 |
237 (58.7) |
116 (58.0) |
>25 |
141 (34.9) |
60 (30.0) |
Nulliparas |
26 (6.4) |
24 (12.0) |
Family history of breast cancer |
|
|
No |
367 (90.8) |
186 (93.0) |
Yes |
37 (9.2) |
14 (7.0) |
No. of abortions |
|
|
≦1 |
310 (76.7) |
141 (70.5) |
>1 |
94 (23.3) |
59 (29.5) |
Tumor location |
|
|
Upper-outer quadrant |
226 (55.9) |
98 (49.0) |
Lower-outer quadrant |
61 (15.1) |
34 (17.0) |
Lower-inner quadrant |
23 (5.7) |
8 (4.0) |
Upper-inner quadrant |
71 (17.6) |
52 (26.0) |
Undefined |
23 (5.7) |
8 (4.0) |
Histopathological type |
|
|
Invasive ductal carcinoma |
353 (87.4) |
167 (83.5) |
Invasive lobular carcinoma |
5 (1.2) |
1 (0.5) |
Others |
46 (11.4) |
32 (16.0) |
Ki67 |
|
|
≦20% |
54 (13.4) |
26 (13.0) |
>20%,≦40% |
118 (29.2) |
35 (17.5) |
>40%, ≦60% |
139 (34.4) |
50 (25.0) |
(Continued) |
||
>60% |
93 (23.0) |
89 (44.5) |
Grade |
|
|
I-II |
209 (51.7) |
90 (45.0) |
III |
195 (48.3) |
110 (55.0) |
CEA (ng/ml) |
|
|
≦2.2 |
289 (71.5) |
158 (79.0) |
>2.2 |
115 (28.5) |
42 (21.0) |
CA125 (U/ml) |
|
|
≦17.5 |
356 (88.1) |
156 (78.0) |
>17.5 |
48 (11.9) |
44 (22.0) |
CA153 (U/ml) |
|
|
≦11.3 |
224 (55.4) |
120 (60.0) |
>11.3 |
180 (44.6) |
80 (40.0) |
a Unnatural menopause includes hysterectomy operation and other status.
Independent prognostic factors in the training cohort
We performed the univariate analyses to identify the factors which were correlated with the overall survival of TNBC patients. As shown in Table 2, family history of breast cancer, tumor location, number of positive lymph nodes, histological grade and Ki67 status were significantly associated with the OS. All three laboratory parameters including CEA, CA125 and CA153 were also indicated as significant factors that had impact on survival. The above variables were entered into multivariate Cox proportional hazard regression analyses and the results demonstrated that family history of breast cancer, tumor location, number of positive lymph nodes, histological grade, serum CEA, CA125 and CA153 were independent prognostic factors for OS. The detail results of multivariate analyses are displayed in Table 2. Kaplan-Meier curves for significant factors derived from univariate analysis were shown in Figure 1.
Table 2: Univariate and multivariate Cox regression analyses of overall survival for patients with TNBC in the training cohort
Variables |
Univariate analyses |
Multivariate analyses |
||
|---|---|---|---|---|
HR (95%CI) |
Log-rank p |
HR (95%CI) |
Log-rank p |
|
Factors selected |
||||
Family history of breast cancer |
1.61 (1.45-1.83) |
<0.001 |
1.59 (1.41-1.81) |
<0.001 |
Tumor location |
||||
Upper outer quadrant |
1.00 |
0.013 |
1.00 |
0.024 |
Lower outer quadrant |
1.02 (0.91-1.17) |
1.01 (0.90-1.12) |
||
Lower inner quadrant |
1.31 (1.23-1.52) |
1.33 (1.21-1.58) |
||
Upper inner quadrant |
1.24 (1.18-1.48) |
1.19 (1.10-1.58) |
||
No. of positive lymph nodes |
1.18 (1.13-1.22) |
<0.001 |
1.21 (1.14-1.28) |
<0.001 |
Grade |
||||
I-II |
1.00 |
<0.001 |
1.00 |
<0.001 |
III |
1.50 (1.29-1.68) |
1.48 (1.27-1.67) |
||
CEA (ng/ml) |
||||
≦2.2 |
1.00 |
<0.001 |
1.00 |
<0.001 |
>2.2 |
1.31 (1.18-1.49) |
1.28 (1.16-1.51) |
||
CA125 (U/ml) |
||||
≦17.5 |
1.00 |
<0.001 |
1.00 |
<0.001 |
>17.5 |
1.44 (1.24-1.68) |
1.42 (1.21-1.65) |
||
CA153 (U/ml) |
||||
≦11.3 |
1.00 |
0.002 |
1.00 |
0.003 |
>11.3 |
1.26 (1.10-1.45) |
1.24 (1.11-1.44) |
||
Factors not selected |
||||
Age |
1.00 (0.98-1.03) |
0.789 |
|
|
BMI |
0.97 (0.88-1.07) |
0.576 |
|
|
Age at menarche |
1.13 (0.97-1.32) |
0.111 |
|
|
Menopausal status |
0.72 (0.43-1.23) |
0.231 |
|
|
Age at first live birth |
0.96 (0.89-1.04) |
0.348 |
|
|
No. of abortions ( >1 vs. ≦1) |
0.82 (0.48-1.39) |
0.454 |
|
|
Tumor size |
1.13 (0.97-1.33) |
0.111 |
|
|
No. of examined lymph nodes |
0.99 (0.95-1.03) |
0.547 |
|
|
Histopathological type |
0.78 (0.68-1.09) |
0.301 |
|
|
Ki67 status |
1.32 (1.12-1.62) |
0.007 |
|
|
Abbreviations: CEA, carcinoembryonic antigen; CA125, cancer antigen 125; CA153, cancer antigen 153; HR, hazard ratio; CI, confidence interval.
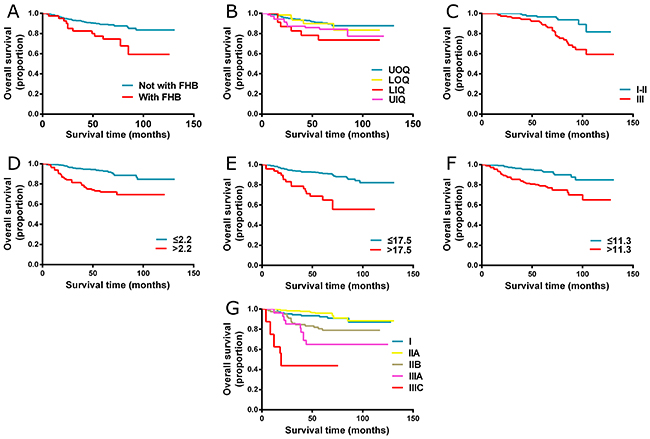
Figure 1: Kaplan-Meier curves for significant factors derived from univariate analysis. (A) Family history of breast cancer; (B) Tumor location; (C) Histological grade; (D) CEA; (E) CA125; (F) CA153; (G) AJCC 8th TNM staging system] Abbreviations: FHB, family history of breast cancer; UOQ, upper outer quadrant; LOQ, lower outer quadrant; LIQ, lower inner quadrant; UIQ, upper inner quadrant.
Prognostic nomogram for OS
All the significant factors identified by the Cox regression model were applied to establish the nomogram for OS. Each subtype within the variables was assigned a score on the point scale. By adding up the total score from all the variables and locating it into the total point scale, we could identify the probabilities of the outcomes by drawing a vertical line to the total score. As shown in Figure 2, the nomogram revealed that FHB (family history of breast cancer) had the largest contribution to the prognosis, followed by the histological grade and serum CA125 level. The C-index calculated in the training cohort for OS prediction was 0.76 (95%CI, 0.72-0.81), indicating the suitability of this new model for TNBC patients. The calibration plot on the probability of survival for 3 or 5 years OS showed an optimal agreement between the prediction by the nomogram and actual observations (Figure 3A and 3B).
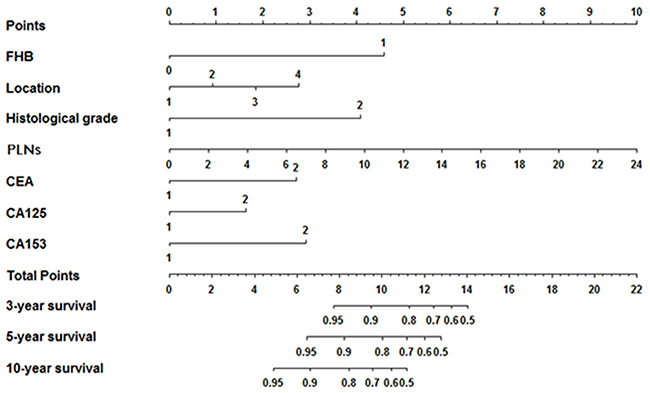
Figure 2: Prognostic nomogram for patients with triple negative breast cancer. Abbreviations: FHB, family history of breast cancer; PLNs, number of positive lymph nodes.
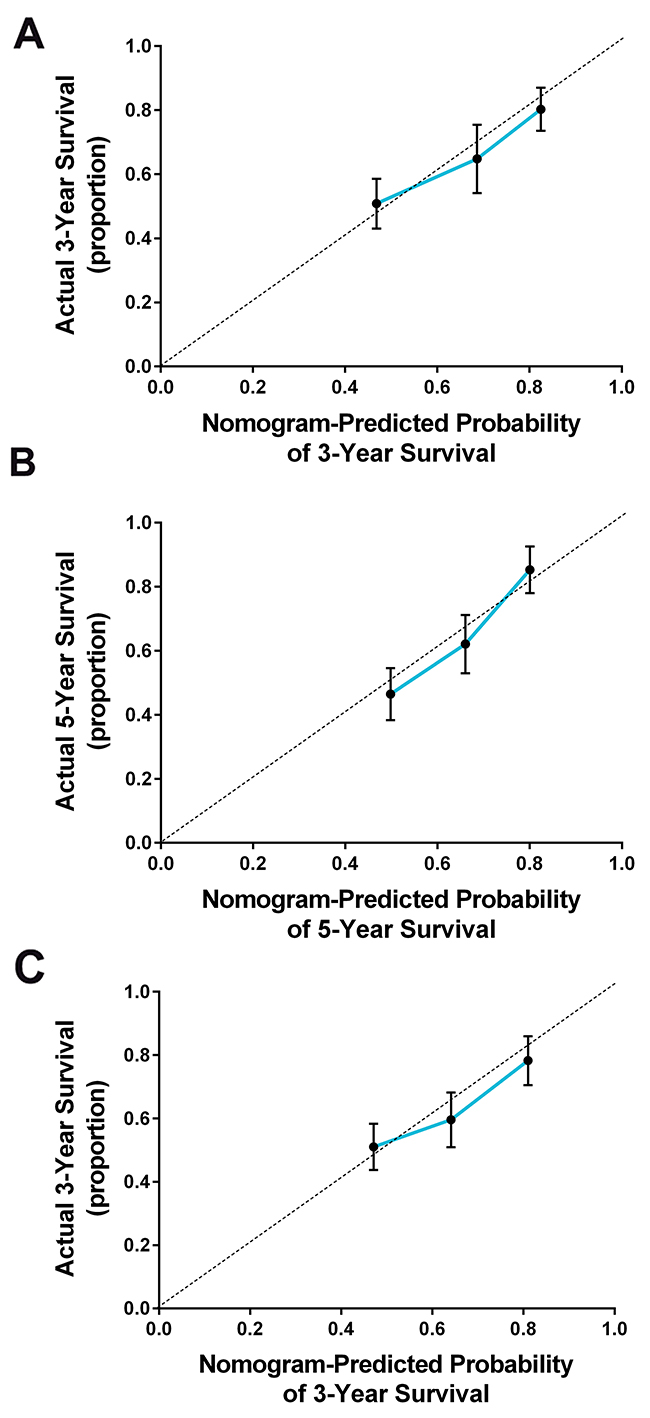
Figure 3: The calibration curve for predicting the overall survival for triple negative breast cancer patients at (A) 3 years and (B) 5 years in the training cohort and at (C) 3 years in the validation cohort. Nomogram-predicted probability of overall survival is plotted on the x-axis and the actual overall survival is plotted on the y-axis.
Comparison of predictive accuracy between nomogram and conventional staging system
We also compared the predictive accuracy between this nomogram and the 8th TNM staging system. As shown in Table 3, the performance of nomogram discrimination in the training cohort was 0.76 (95%CI, 0.72-0.81), which was significantly higher than the 8th TNM classification (0.66; 95%CI, 0.63-0.70, p<0.001), suggesting better accuracy in predicting OS than the conventional staging system.
Table 3: Comparison of the nomogram with TNM staging system
Models |
Training cohort |
Validation cohort |
||
|---|---|---|---|---|
C-index (95%CI) |
p |
C-index (95%CI) |
p |
|
Nomogram |
0.76 (0.72-0.81) |
<0.001 |
0.72 (0.69-0.76) |
0.002 |
8th TNM staging system |
0.66 (0.63-0.70) |
0.64 (0.60-0.67) |
||
Abbreviations: C-index, concordance index.
Validation of the predictive accuracy of the Nomogram for OS
An external validation was further performed in an independent cohort of 200 patients with TNBC. As shown in Figure 3C, the calibration curves also showed good agreement between the nomogram prediction and actual observation for the 3-year OS. In addition, the C-index of the nomogram in the validation cohort for predicting OS was 0.72 (95%CI, 0.69-0.76), which was also superior to the 8th TNM staging system (0.64; 95%CI, 0.60-0.67, p=0.002). These results again suggested that this nomogram was useful for predicting the survival of patients with TNBC.
DISCUSSION
Triple negative breast cancer is a subtype of breast cancer associated with a dismal prognosis [18]. Although the incidence of TNBC only accounts for a small proportion (10–20%) of all breast cancer [5], it has the highest risk of local relapse, distant metastasis and death than other types of BC, especially in the first three and five years [6–8]. TNBC represents an invasive phenotype of breast cancer with diverse molecular characterization and response to therapy [19, 20]. Due to the lack of estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor-2 (HER2) expression, only a few adjuvant treatments, like conventional surgery, chemotherapy and radiotherapy, may provide benefits for patients with TNBC. The 8th TNM classification staging system is the most widely used model to predict the clinical outcome of TNBC. However, the prognosis of some patients at same stage may still vary widely. Such heterogeneity might be mainly caused by the different biological and pathological behavior of TNBC. Therefore, it is important to develop an accurate and practical predictive model to assist physicians to make short-term and long-term treatment decisions for specific patients.
Nomogram is a pictorial representation of a complex mathematical formula which uses clinical and biological variables to determine a statistical predictive model and calculates the probability of a clinical event, such as tumor recurrence or death. Nomograms have been proved to be more accurate than the conventional staging systems for predicting the prognosis in several types of cancers [11–16, 21, 22]. In this study, we have developed a novel nomogram by incorporating the clinical and laboratory relevant prognostic factors to better predict the overall survival of TNBC patients. Through univariate analysis and subsequent multivariate regression analyses, we have identified family history of breast cancer, number of positive lymph nodes and histological grade as independent prognostic factors. These findings were in high concordance with previous studies on risk factors for breast cancer [23, 24]. Notably, tumor location was also indicated as an important prognostic factor for TNBC, with which some similar studies also supported that the tumor in lower inner quadrant (LIQ) showed a more unfavorable prognosis [25]. One of the possible hidden reasons is the potential internal mammary node metastasis. The nomogram also includes comprehensive laboratory parameters such as serum tumor biomarkers, which have not been involved in conventional staging systems. As shown in several studies [26–28], higher levels of preoperative serum tumor markers, such as CEA, CA125 and CA153 could represent tumor burden and have been suggested to be independent risk factors for the prognosis of breast cancer. Although these tumor markers have not been included in breast cancer staging systems, their role in increasing predictive performance has been validated in staging systems of other cancer types, such as the hepatocellular carcinoma (HCC) staging systems [29].
To the best of our knowledge, this is the first nomogram for predicting the survival of patients with TNBC which involves both clinical and laboratory relevant factors. The results indicated that this nomogram was superior to the existing staging system with higher C-index and optimal agreement between prognostic prediction and actual observation. Both physicians and patients could perform an individualized survival prediction through this scoring system. This nomogram could help physicians to identify subgroups of patients at different risk of poor survival and to decide who need intensive follow-up or additional treatment. However, several limitations in the current study should also be mentioned. Firstly, the nomogram was established based on the retrospective data obtained from only one single institution in China, although eligibility criteria were formulated to minimize the selective bias. Moreover, our nomogram failed to incorporate some recognized prognostic parameters (eg, vascular carcinoma embolus invasion) or important molecular factors (eg, P53 expression, BRCA1/2 mutation). In addition, relevant treatment conditions such as adjuvant chemotherapy or radiotherapy were also not included in this study. Further efforts on prospective multicenter data collection and incorporation of some other factors are also warranted to improve this model.
In conclusion, this proposed nomogram could provide accurate prediction for the prognosis of the patients with TNBC and was able to stratify patients into distinct prognostic groups. Further perspective cohort studies with larger sample size are also required to illustrate and improve the validity of this model in the therapeutic decision-making field for other ethnic patients with TNBC.
MATERIALS AND METHODS
Ethics statement
The study and consent procedure was approved by the Ethics Committee of Affiliated Union Hospital of Fujian Medical University (Fuzhou, China). All participants enrolled in this study provided their written informed consent.
Study patients and data collection
We conducted a retrospectively study between August 2006 and July 2012 on a training cohort of 486 patients and a prospectively study between August 2012 and July 2014 on a validation cohort of 283 patients. All patients underwent mastectomy or breast-conserving surgery at the Affiliated Union Hospital of Fujian Medical University. The inclusion criteria were as follows: age 20–65 years; no history of previous anticancer treatment; no history of other malignancies; and histopathologically diagnosed with triple negative breast cancer (ER, PR, and HER2 negative. HER2 evaluation was performed using IHC, it was considered as negative of scores were 0 and 1+. For score was 2+, the HER2 status was confirmed either as positive or negative according to the gene amplification ratio of the fluorescence in situ hybridization (FISH) by current standards. For FISH, the cut-off Her-2/CSP17 ratio was defined as 2.0 according to the ASCO guidelines). The exclusion criteria include tumors of uncertain origins or probable metastatic breast tumors; confirmed no metastasis in the preoperative examinations; lack of necessary information; and patients without follow-up data.
Each patient was interviewed face-to-face by a trained interviewer to obtain information on demographic factors (age, BMI, menstrual status, reproductive history, family history of breast cancer). All patients’ clinicopathological data was collected form the electronic medical record systems, including tumor size, lymph node status, tumor location, histopathological type, histological grade, and Ki67 status. The clinical staging of triple negative breast cancer was evaluated by the TNM staging system according to the AJCC 8th edition. Laboratory parameters were retrieved from hematologic tests which were performed at initial diagnosis and prior to any anti-cancer therapy which included carcinoembryonic antigen (CEA), cancer antigen 125 (CA125) and cancer antigen 153 (CA153). The optimal cut-off values for serum cancer biomarkers were determined by the minimum P value from log-rank X2 statistics using the X-tile 3.6.1 software (Yale University, New Haven, CT, USA) [30]. The cut-off values of CEA, CA125 and CA153 were 2.0 ng/ml, 17.5 U/ml and 11.3 U/ml, respectively (Figure 4).
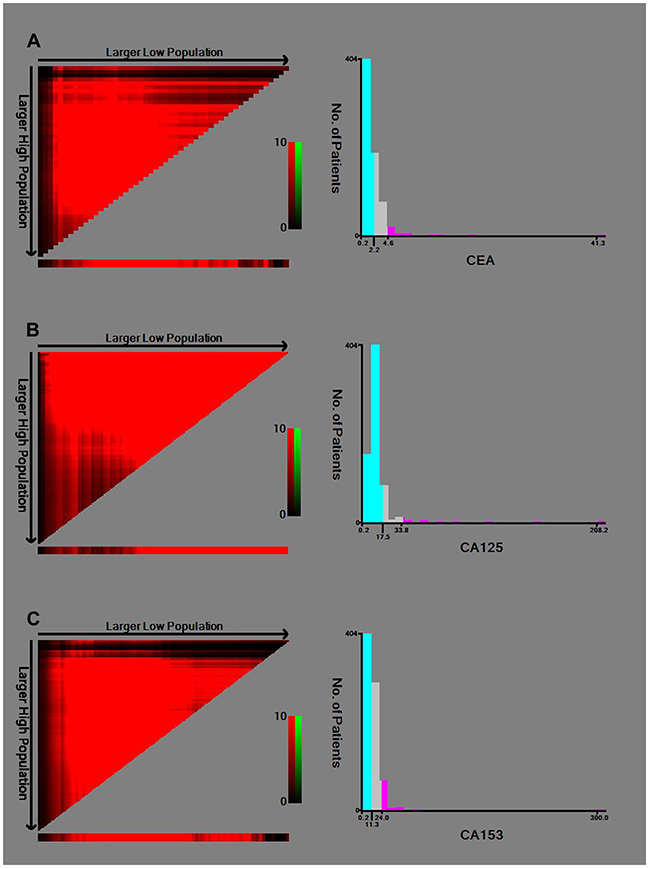
Figure 4: Identification of the optimal cut-off values for serum (A) CEA; (B) CA125; (C) CA153.
Follow-up
All patients were followed-up by telephones calls every 3 months during the first 2 years after treatment, and 6 months annually thereafter. At each follow-up, each patient’s status was carefully recorded. The primary observation endpoint of this study was overall survival (OS) and the last follow-up date was July 1, 2017. OS was defined as the interval between diagnosis and death from any cause or until the last follow-up date which is collected from the patients’ medical records or through direct contact with the patients or their families.
Statistical analysis
Continuous variables were presented as means ± standard deviation (SD) or medians and ranges, while frequencies and proportions were calculated for categorical variables. The Student’s t-test or non-parametric Mann–Whitney U test was performed for continuous data. The Pearson chi-square or Fisher’s exact tests were used to compare the differences in proportion between the groups. Kaplan-Meier method with log-rank test was assessed to depict the survival curves. Cox univariate and multivariate regression analyses were adopted to identify independent risk factors to predict mortality.
Nomogram was built based on the results of multivariate Cox regression analyses using R software (version 3.0.2, http://www.r-project.org/) with the RMS packages. Factors in multiple regression analyses were selected with a backward step-down process, while the smallest Akaike’s information criterion (AIC) was applied as a stopping rule [31]. The performance of the nomogram and TNM staging system for predicting survival were measured by the concordance index (C-index). The maximum C-index value is 1.0 which indicates a perfect prediction model, whereas 0.5 represents only half of the chance to correctly predict the outcome. Bootstraps with 1,000 resample were applied for validation to correct the C-index and explain the variance due to over-optimism. Comparisons between the nomogram and TNM staging system were performed with the rcorrp.cens package of Hmisc in R. Calibration curves of the nomogram for 3 year and 5 year overall survival (OS) were adopted to evaluate the agreement between the predicted survival and the observed survival. When externally validating the nomogram, the total points of each patient in the validation cohort were calculated according to the established nomogram, then the total points were used as a factor and applied into the Cox regression model, and finally, the C-index and calibration curve were derived based on the regression analysis. All statistical analyses were two-sided, and a P value of less than 0.05 was considered as a significance level.
Author contributions
WC and FFM conceived and designed the study. QW and LSP collected the data. ZW and LJX analyzed the data. LYX wrote the manuscript. All authors contributed toward data analysis, drafting and critically revising the paper, gave final approval of the version to be published, and agreed to be accountable for all aspects of the work.
CONFLICTS OF INTEREST
The authors have declared that no competing interests exist.
FUNDING
The study was supported by grants from Foundation for Young Talent Training Program of Fujian Provincial Health and Family Planning Commission (2015-ZQN-ZD-14) and Sci-Tech Key Program of Fujian Province (2016J01549).
REFERENCES
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108. https://doi.org/10.3322/caac.21262.
2. Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017; 67:7–30. https://doi.org/10.3322/caac.21387.
3. Sotiriou C, Neo SY, McShane LM, Korn EL, Long PM, Jazaeri A, Martiat P, Fox SB, Harris AL, Liu ET. Breast cancer classification and prognosis based on gene expression profiles from a population-based study. Proc Natl Acad Sci U S A. 2003; 100:10393–8. https://doi.org/10.1073/pnas.1732912100.
4. Ma XJ, Salunga R, Tuggle JT, Gaudet J, Enright E, McQuary P, Payette T, Pistone M, Stecker K, Zhang BM, Zhou YX, Varnholt H, Smith B, et al. Gene expression profiles of human breast cancer progression. Proc Natl Acad Sci U S A. 2003; 100:5974–9. https://doi.org/10.1073/pnas.0931261100.
5. Boyle P. Triple-negative breast cancer: epidemiological considerations and recommendations. Ann Oncol. 2012; 23:vi7–12. https://doi.org/10.1093/annonc/mds187.
6. Haffty BG, Yang Q, Reiss M, Kearney T, Higgins SA, Weidhaas J, Harris L, Hait W, Toppmeyer D. Locoregional relapse and distant metastasis in conservatively managed triple negative early-stage breast cancer. J Clin Oncol. 2006; 24:5652–7. https://doi.org/10.1200/JCO.2006.06.5664.
7. Dent R, Trudeau M, Pritchard KI, Hanna WM, Kahn HK, Sawka CA, Lickley LA, Rawlinson E, Sun P, Narod SA. Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res. 2007; 13:4429–34. https://doi.org/10.1158/1078-0432.CCR-06-3045.
8. Reis-Filho JS, Tutt AN. Triple negative tumours: a critical review. Histopathology. 2008; 52:108–18. https://doi.org/10.1111/j.1365-2559.2007.02889.x.
9. Amin MB, Edge SB, Greene FL, Byrd DR, Brookland RK, Washington MK, Gershenwald JE, Compton CC, Hess KR, Sullivan DC, Jessup JM, Brierley JD, Gaspar LE, et al. AJCC Cancer Staging Manual. 8th ed. New York: Springer; 2017.
10. Balachandran VP, Gonen M, Smith JJ, DeMatteo RP. Nomograms in oncology: more than meets the eye. Lancet Oncol. 2015; 16:e173–80. https://doi.org/10.1016/S1470-2045(14)71116-7.
11. Hyder O, Marques H, Pulitano C, Marsh JW, Alexandrescu S, Bauer TW, Gamblin TC, Sotiropoulos GC, Paul A, Barroso E, Clary BM, Aldrighetti L, Ferrone CR, et al. A nomogram to predict long-term survival after resection for intrahepatic cholangiocarcinoma: an Eastern and Western experience. JAMA Surg. 2014; 149:432–8. https://doi.org/10.1001/jamasurg.2013.5168.
12. Wang Y, Li J, Xia Y, Gong R, Wang K, Yan Z, Wan X, Liu G, Wu D, Shi L, Lau W, Wu M, Shen F. Prognostic nomogram for intrahepatic cholangiocarcinoma after partial hepatectomy. J Clin Oncol. 2013; 31:1188–95. https://doi.org/10.1200/JCO.2012.41.5984.
13. Yeh CN, Wang SY, Chen YY, Chen MH, Chiang KC, Cheng CT, Tsai CY, Wang CC, Yeh TS, Chen TC. A prognostic nomogram for overall survival of patients after hepatectomy for intrahepatic cholangiocarcinoma. Anticancer Res. 2016; 36:4249–58.
14. Wan G, Gao F, Chen J, Li Y, Geng M, Sun L, Liu Y, Liu H, Yang X, Wang R, Feng Y, Wang X. Nomogram prediction of individual prognosis of patients with hepatocellular carcinoma. BMC Cancer. 2017; 17:91. https://doi.org/10.1186/s12885-017-3062-6.
15. Liang W, Zhang L, Jiang G, Wang Q, Liu L, Liu D, Wang Z, Zhu Z, Deng Q, Xiong X, Shao W, Shi X, He J. Development and validation of a nomogram for predicting survival in patients with resected non-small-cell lung cancer. J Clin Oncol. 2015; 33:861–9. https://doi.org/10.1200/JCO.2014.56.6661.
16. Han DS, Suh YS, Kong SH, Lee HJ, Choi Y, Aikou S, Sano T, Park BJ, Kim WH, Yang HK. Nomogram predicting long-term survival after d2 gastrectomy for gastric cancer. J Clin Oncol. 2012; 30:3834–40. https://doi.org/10.1200/JCO.2012.41.8343.
17. Dai D, Chen B, Tang H, Wang B, Zhao Z, Xie X, Wei W. Nomograms for predicting the prognostic value of pre-therapeutic CA15-3 and CEA serum levels in TNBC patients. PLoS One. 2016; 11:e0161902. https://doi.org/10.1371/journal.pone.0161902.
18. O’Brien KM, Cole SR, Tse CK, Perou CM, Carey LA, Foulkes WD, Dressler LG, Geradts J, Millikan RC. Intrinsic breast tumor subtypes, race, and long-term survival in the Carolina Breast Cancer Study. Clin Cancer Res. 2010; 16:6100–10. https://doi.org/10.1158/1078-0432.CCR-10-1533.
19. Brouckaert O, Wildiers H, Floris G, Neven P. Update on triple-negative breast cancer: prognosis and management strategies. Int J Womens Health. 2012; 4:511–20. https://doi.org/10.2147/IJWH.S18541.
20. Rakha EA, El-Rehim DA, Paish C, Green AR, Lee AH, Robertson JF, Blamey RW, Macmillan D, Ellis IO. Basal phenotype identifies a poor prognostic subgroup of breast cancer of clinical importance. Eur J Cancer. 2006; 42:3149–56. https://doi.org/10.1016/j.ejca.2006.08.015.
21. Touijer K, Scardino PT. Nomograms for staging, prognosis, and predicting treatment outcomes. Cancer. 2009; 115:3107–11. https://doi.org/10.1002/cncr.24352.
22. Sternberg CN. Are nomograms better than currently available stage groupings for bladder cancer? J Clin Oncol. 2006; 24:3819–20. https://doi.org/10.1200/JCO.2006.07.1290.
23. Yuan N, Meng M, Liu C, Feng L, Hou L, Ning Q, Xin G, Pei L, Gu S, Li X, Zhao X. Clinical characteristics and prognostic analysis of triple-negative breast cancer patients. Mol Clin Oncol. 2014; 2:245–51. https://doi.org/10.3892/mco.2013.230.
24. Li CY, Zhang S, Zhang XB, Wang P, Hou GF, Zhang J. Clinicopathological and prognostic characteristics of triple- negative breast cancer (TNBC) in Chinese patients: a retrospective study. Asian Pac J Cancer Prev. 2013; 14:3779–84.
25. Hwang KT, Kim J, Kim EK, Jung SH, Sohn G, Kim SI, Jeong J, Lee HJ, Park JH, Oh S; Korean Breast Cancer Society. Poor prognosis of lower inner quadrant in lymph node-negative breast cancer patients who received no chemotherapy: a study based on nationwide korean breast cancer registry database. Clin Breast Cancer. 2017; 17:e169–e84. https://doi.org/10.1016/j.clbc.2016.12.011.
26. Fang C, Cao Y, Liu X, Zeng XT, Li Y. Serum CA125 is a predictive marker for breast cancer outcomes and correlates with molecular subtypes. Oncotarget. 2017; 8:63963–70. https://doi.org/10.18632/oncotarget.19246.
27. Wu SG, He ZY, Zhou J, Sun JY, Li FY, Lin Q, Guo L, Lin HX. Serum levels of CEA and CA15-3 in different molecular subtypes and prognostic value in Chinese breast cancer. Breast. 2014; 23:88–93. https://doi.org/10.1016/j.breast.2013.11.003.
28. Shao Y, Sun X, He Y, Liu C, Liu H. Elevated levels of serum tumor markers CEA and CA15-3 are prognostic parameters for different molecular subtypes of breast cancer. PLoS One. 2015; 10:e0133830. https://doi.org/10.1371/journal.pone.0133830.
29. Chevret S, Trinchet JC, Mathieu D, Rached AA, Beaugrand M, Chastang C. A new prognostic classification for predicting survival in patients with hepatocellular carcinoma. Groupe d’Etude et de Traitement du Carcinome Hepatocellulaire. J Hepatol. 1999; 31:133–41.
30. Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004; 10:7252–9. https://doi.org/10.1158/1078-0432.CCR-04-0713.
31. Harrell FE Jr, Lee KL, Mark DB. Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat Med. 1996; 15:361–87. https://doi.org/10.1002/(SICI)1097-0258(19960229)15:4<361::AID-SIM168>3.0.CO;2-4.

