INTRODUCTION
Esophageal cancer is the sixth leading cause of cancer-related mortality and the eighth most common cancer worldwide [1]. An estimated 455,800 new esophageal cancer cases and 400,200 deaths occurred in 2012 worldwide [2]. The two major types are esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EADC). Smoking and drinking are well-known environmental risk factors for ESCC, whereas obesity and chronic gastroesophageal refluxing are main EADC risk factors. However, only a subset of individuals exposed to those environmental risk factors develop EC, suggesting a role of host susceptibility factors. Some studies have suggested that genetic polymorphisms might explain individual differences in susceptibility to esophageal cancer [3, 4].
Glutathione-S-transferases (GSTs) are important phase II biotransformation enzymes that catalyzing the nucleophilic addition of glutathione to several hazardous xenobiotics, including phase I electrophilic and carcinogenic metabolites [5]. However, these enzymes can also activate certain chemicals that target cellular proteins and DNA to elicit detrimental carcinogenic effects through genotoxic and non-genotoxic mechanisms. One of important enzymes in GSTs family is GSTT1. GSTT1 is genetically polymorphic, and deletion polymorphism of the GSTT1 loci (null genotype) results in the loss of functional activity. Several studies have found that GSTT1 null genotype is strongly associated with susceptibility to a number of cancers, such as colorectal, renal and oral cancers et al. [6–8]. Previous studies have been published to estimate the association between GSTT1 null genotype and the risk of esophageal cancer, but the results are inconsistent [9–11].
To date, several meta-analysis studies have reported the association between null GSTT1 genotype and the risk of esophageal cancer. However, the results of these studies remain outdated and incomprehensive [12–14]. In the last 4 years, many case-control studies were published to estimate this association. Thus, to obtain a conclusive result about this association, we performed current meta-analysis that includes all recent publications to review and summarize the association between the GSTT1 polymorphism and the risk of esophageal cancer.
RESULTS
Characteristics
In total, 100 articles were retrieved. Figure 1 summarized the selecting process. Finally, a total of 30 studies with 4482 cases and 6681 controls met the inclusion criteria [9–11, 15–41]. Among them, 18 were from Asians, 10 were from Caucasians, and 2 were from Africans. There were 21 studies focused on the risk of ESCC with 3272 cases and 5535 controls, and 8 studies focused on the risk of EADC with 646 cases and 1908 controls. Characteristics of included studies and the distribution of GSTT1 polymorphism are summarized in Table 1 and Table 2, respectively.
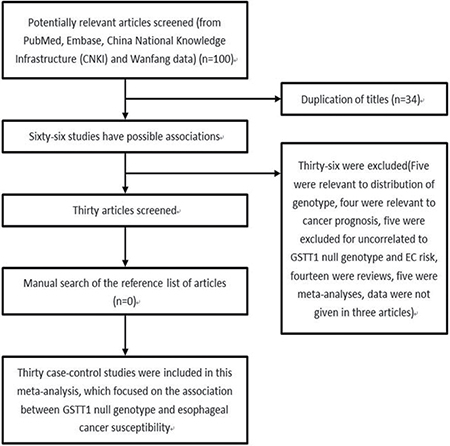
Figure 1: Flow chart shows studies included procedure for meta-analysis.
Table 1: Characteristics of the individual studies included in the meta-analysis
Study |
Year |
Country |
Ethnicity |
Sample size |
Genotype method |
|---|---|---|---|---|---|
Makhdoomi MA |
2014 |
India |
Asian |
492/492 |
multiplex PCR |
Sharma A |
2013 |
India |
Asian |
315/436 |
multiplex PCR |
Dura P |
2013 |
Netherlands |
Caucasian |
432/591 |
PCR |
Talukdar FR |
2013 |
India |
Asian |
112/130 |
PCR |
Zhang L |
2013 |
China |
Asian |
138/170 |
PCR |
Djansugurova LB |
2013 |
Kazakhstan |
Caucasian |
107/96 |
PCR |
Gao P |
2012 |
China |
Asian |
40/80 |
PCR |
Matejcic M |
2011 |
South Africa |
African |
528/876 |
PCR |
Malik MA |
2010 |
India |
Asian |
135/195 |
multiplex PCR |
Li D |
2010 |
South Africa |
African |
238/280 |
PCR |
Moaven O |
2010 |
Iran |
Asian |
148/136 |
PCR |
Liu R |
2010 |
China |
Asian |
97/97 |
multiplex PCR |
Ji R |
2010 |
China |
Asian |
189/216 |
multiplex PCR |
Zendehdel K |
2009 |
Sweden |
Caucasian |
172/470 |
multiplex PCR |
Zhang WL |
2009 |
China |
Asian |
88/72 |
PCR |
Deng J |
2008 |
China |
Asian |
87/162 |
PCR |
Rossini A |
2007 |
Brazil |
Caucasian |
125/252 |
multiplex PCR |
Wideroff L |
2007 |
USA |
Caucasian |
67/208 |
PCR |
Casson AG |
2006 |
Canada |
Caucasian |
56/95 |
multiplex PCR |
Jain M |
2006 |
India |
Asian |
100/137 |
multiplex PCR |
Yin LH |
2005 |
China |
Asian |
106/106 |
PCR |
Roth MJ |
2004 |
China |
Asian |
131/454 |
PCR |
Abbas A |
2004 |
French |
Caucasian |
70/115 |
multiplex PCR |
Wang LD |
2003 |
China |
Asian |
62/38 |
multiplex PCR |
Casson AG |
2003 |
Canada |
Caucasian |
45/45 |
multiplex PCR |
Ribeiro Pinto LF |
2003 |
Brazil |
Caucasian |
32/67 |
PCR |
Gao CM |
2002 |
China |
Asian |
141/223 |
multiplex PCR |
Tan W |
2000 |
China |
Asian |
150/150 |
multiplex PCR |
van Lieshout EM |
1999 |
Netherlands |
Caucasian |
34/247 |
PCR |
Lin DX |
1998 |
China |
Asian |
45/45 |
multiplex PCR |
Table 2: Distribution of GSTT1 null genotype among cases and controls
Study |
Year |
Control |
EC |
ESCC |
EADC |
||||
|---|---|---|---|---|---|---|---|---|---|
Present |
Null |
Present |
Null |
Present |
Null |
Present |
Null |
||
Makhdoomi MA |
2014 |
367 |
125 |
306 |
186 |
306 |
186 |
/ |
/ |
Sharma A |
2013 |
373 |
63 |
233 |
82 |
233 |
82 |
/ |
/ |
Dura P |
2013 |
463 |
128 |
335 |
97 |
87 |
18 |
248 |
79 |
Talukdar FR |
2013 |
92 |
38 |
66 |
46 |
66 |
46 |
/ |
/ |
Zhang L |
2013 |
90 |
80 |
62 |
76 |
62 |
76 |
/ |
/ |
Djansugurova LB |
2013 |
35 |
61 |
19 |
88 |
19 |
88 |
/ |
/ |
Gao P |
2012 |
55 |
25 |
18 |
22 |
18 |
22 |
/ |
/ |
Matejcic M |
2011 |
648 |
228 |
375 |
153 |
375 |
153 |
/ |
/ |
Malik MA |
2010 |
146 |
49 |
110 |
25 |
/ |
/ |
/ |
/ |
Li D |
2010 |
178 |
102 |
125 |
113 |
125 |
113 |
/ |
/ |
Moaven O |
2010 |
105 |
31 |
112 |
36 |
112 |
36 |
/ |
/ |
Liu R |
2010 |
57 |
40 |
34 |
63 |
34 |
63 |
/ |
/ |
Ji R |
2010 |
122 |
94 |
91 |
98 |
91 |
98 |
/ |
/ |
Zendehdel K |
2009 |
394 |
76 |
150 |
22 |
70 |
7 |
80 |
15 |
Zhang LW |
2009 |
39 |
33 |
31 |
57 |
31 |
57 |
/ |
/ |
Deng J |
2008 |
75 |
87 |
36 |
51 |
/ |
/ |
/ |
/ |
Rossini A |
2007 |
192 |
60 |
110 |
15 |
110 |
15 |
/ |
/ |
Wideroff L |
2007 |
173 |
35 |
59 |
8 |
/ |
/ |
59 |
8 |
Casson AG |
2006 |
80 |
15 |
42 |
14 |
/ |
/ |
42 |
14 |
Jain M |
2006 |
100 |
37 |
72 |
28 |
56 |
20 |
6 |
3 |
Yin LH |
2005 |
55 |
51 |
60 |
46 |
/ |
/ |
/ |
/ |
Roth MJ |
2004 |
211 |
243 |
54 |
77 |
54 |
77 |
/ |
/ |
Abbas A |
2004 |
85 |
30 |
56 |
14 |
31 |
13 |
25 |
1 |
Wang LD |
2003 |
18 |
20 |
28 |
34 |
25 |
34 |
/ |
/ |
Casson AG |
2003 |
33 |
12 |
37 |
8 |
/ |
/ |
37 |
8 |
Rebeiro |
2003 |
52 |
15 |
26 |
6 |
/ |
/ |
/ |
/ |
Gao CM |
2002 |
104 |
119 |
67 |
74 |
/ |
/ |
/ |
/ |
Tan W |
2000 |
91 |
59 |
90 |
60 |
90 |
60 |
/ |
/ |
van Lieshout EM |
1999 |
198 |
49 |
28 |
6 |
11 |
2 |
17 |
4 |
Lin DX |
1998 |
22 |
23 |
26 |
19 |
/ |
/ |
/ |
/ |
Abbreviations: EC: esophageal cancer; ESCC: esophageal squamous cell carcinoma; EADC: esophageal adenocarcinoma.
Quantitative synthesis
Table 3 showed the main result of the association between GSTT1 null genotype and the risk of esophageal cancer. Overall, there was a significant correlation of GSTT1 null genotype with esophageal cancer risk (OR = 1.20; 95% CI 1.04–1.40; P < 0.05; Table 3 and Figure 2). In a subgroup analysis by ethnicity, GSTT1 null genotype was correlated with a significantly increased risk of esophageal cancer among Asians (OR = 1.33; 95% CI 1.12–1.58; P < 0.05; Table 3 and Figure 2), but not among Caucasians and Africans (OR = 0.91; 95% CI 0.65–1.26; P > 0.05 for Caucasians and OR = 1.32; 95% CI 0.98–1.77; P > 0.05 for Africans; Table 3 and Figure 2).
Table 3: Meta-analysis of the GSTT1 null genotype and esophageal cancer risk
Number of Study |
OR (95%CI) |
P |
P (Q-test) |
I-squared |
|
|---|---|---|---|---|---|
Total |
30 |
1.20 (1.04–1.40) |
0.014 |
0.000 |
60.2 |
Ethnicity |
|||||
Asians |
18 |
1.33 (1.12–1.58) |
0.001 |
0.003 |
54.9 |
Caucasians |
10 |
0.91 (0.65–1.26) |
0.569 |
0.01 |
58.3 |
Africans |
2 |
1.32 (0.98–1.77) |
0.070 |
0.157 |
50.0 |
ESCC |
21 |
1.34 (1.12–1.61) |
0.001 |
0.000 |
62.6 |
Ethnicity |
|||||
Asians |
13 |
1.54 (1.30–1.82) |
0.000 |
0.097 |
35.7 |
Caucasians |
6 |
0.87 (0.48–1.57) |
0.640 |
0.002 |
73.5 |
Africans |
2 |
1.32 (0.98–1.77) |
0.070 |
0.157 |
50.0 |
EADC |
8 |
0.98 (0.71–1.35) |
0.911 |
0.240 |
23.7 |
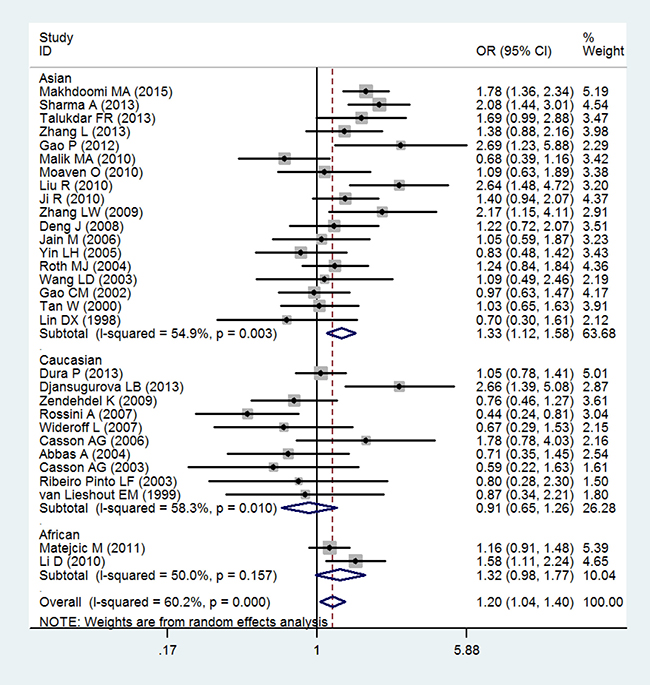
Figure 2: Forest plot of the association of GSTT1 null genotype with esophageal cancer risk.
In the analysis by histological type, GSTT1 null genotype were correlated with a significantly increased risk of ESCC (OR = 1.34; 95% CI 1.12–1.61; P < 0.05; Table 3 and Figure 3). Moreover, in a subgroup analysis by ethnicity, GSTT1 null genotype was correlated with a significantly increased risk of ESCC among Asians (OR = 1.54; 95% CI 1.30–1.82; P < 0.05; Table 3 and Figure 3), but not among Caucasians and Africans (OR = 0.87; 95% CI 0.48–1.57; P > 0.05 for Caucasians and OR = 1.32; 95% CI 0.98–1.77; P > 0.05 for Africans; Table 3 and Figure 3). In addition, there is no significant correlation of GSTT1 null genotype with the risk of EADC (OR = 0.98; 95% CI 0.71–1.35; P > 0.05; Table 3 and Figure 4).
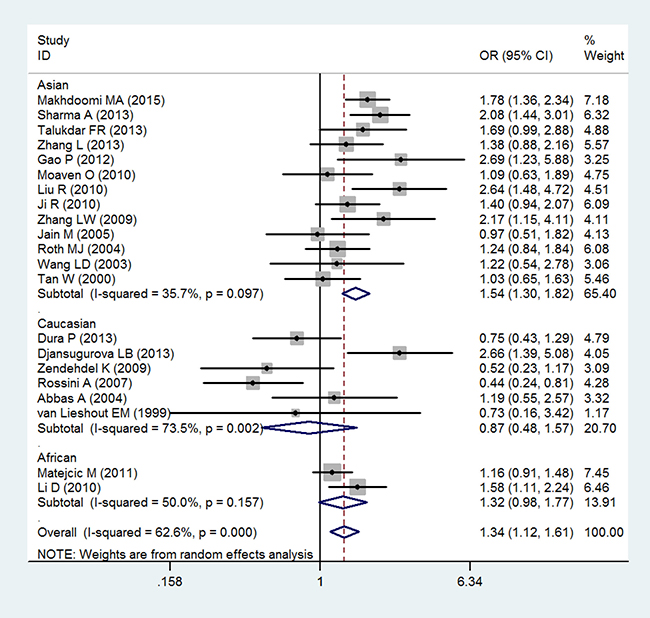
Figure 3: Forest plot of the association of GSTT1 null genotype with esophageal squamous cell carcinoma risk.
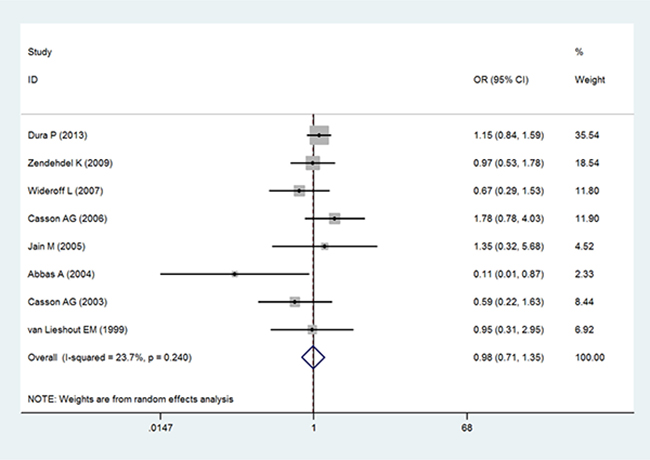
Figure 4: Forest plot of the association of GSTT1 null genotype with esophageal adenocarcinoma risk.
Test for publication bias, sensitivity analyses, and heterogeneity
Publication bias was assessed by both the Begg’s funnel plot and the Egger’s test. The shape of the Begg’s funnel plot did not reveal any evidence of obvious asymmetry (Figure 5). Egger’s test further suggested no evidence of publication bias (P = 0.210). Thus, there was no obvious publication bias in this meta-analysis.
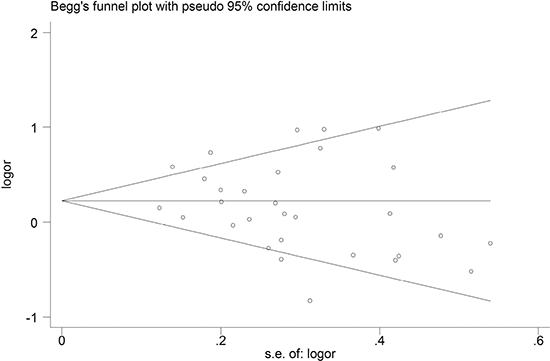
Figure 5: Begg's funnel plot analysis of GSTT1 polymorphism with esophageal cancer risk.
The sensitivity analysis was conducted to test the influence of an individual data on the pooled ORs and CIs by eliding a study in turn. Our findings suggested that the present meta-analysis results were relatively robust and stable (Figure 6).
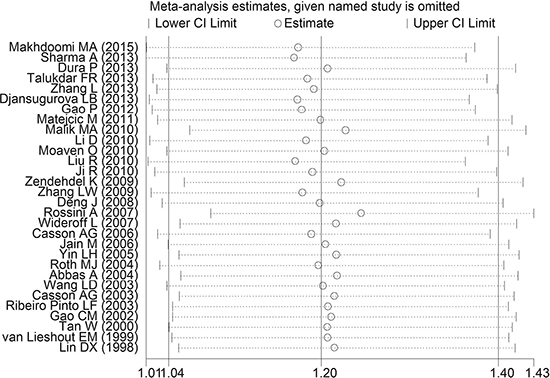
Figure 6: One-way sensitivity analysis of GSTT1 polymorphism with esophageal cancer risk.
As showed in Figure 2 and Figure 3, heterogeneity was significant in overall and in some subgroups. Thus, we measured the sources of heterogeneity by subgroup analysis. The results showed that Caucasians may lead to the major source of heterogeneity.
DISCUSSION
Many studies suggest genetic variants play important roles in individual susceptibility to esophageal cancer [3, 42]. In decades, epidemiological studies have been performed to assess the association of GSTT1 null genotype with the risk of esophageal cancer. However, the results were inconsistent [34, 36]. Previous meta-analyses also investigated the association [12–14], the findings should be interpreted with very cautions. In Weng et al. study, 11 studies included in the meta-analysis were in Chinese Han population. Although the results showed a significant association between GSTT1 null genotype and esophageal cancer risk, the single Chinese Han population limited the power of the statistical analysis [12]. Also In Yi et al. study, 15 studies included in the meta-analysis were in Asian population. Although the results showed a significant association between GSTT1 null genotype and esophageal cancer risk, the single Asian population limited the power of the statistical analysis [13]. In Cai et al. study, a total of 24 studies were used. Adjusted ORs with corresponding 95% CIs were reported in 9 studies. In the overall analysis there was no significant association between GSTT1 null genotype and esophageal cancer risk. However, meta-analysis of adjusted ORs showed a significant association between GSTT1 null genotype and esophageal cancer risk [14]. Because of the lack of available data, subgroup analysis by histological type was not performed in these studies [12–14]. Thus we conducted a comprehensive meta-analysis, to investigate not only the strength of association between GSTT1 null genotype and the risk of esophageal cancer, but also the association of GSTT1 null genotype with the risk of different histological types.
This meta-analysis, including 30 case-control studies with 4482 cases and 6681 controls, identified the association between GSTT1 null genotype and esophageal cancer risk. GSTT1 null genotype significantly increased overall esophageal cancer risk. In a subgroup analysis by ethnicity, GSTT1 null genotype was correlated with a significantly increased risk of esophageal cancer among Asians. In the analysis by histological type, GSTT1 null genotype was correlated with a significantly increased risk of ESCC, particularly in Asians. However, there was no significant correlation of GSTT1 null genotype with the risk of EADC. To date, this is the first meta-analysis concerning the association of GSTT1 null genotype with the risk of different histological types.
GSTT1, encodes an enzyme that plays a crucial role in the detoxification of a variety of endogenous or exogenous carcinogens. It is located on 22q11.23 with 8146 base pairs, 5 exons and 4 introns in all [5]. GSTT1 is genetically polymorphic, and GSTT1 null genotype results in the loss of functional activity [12]. Our results demonstrated that GSTT1 null genotype significantly increased overall esophageal cancer risk.
Since the results from meta-analysis can be affected by histological types, a subgroup analysis was carried out regarding different histological type for the GSTT1 null genotype. GSTT1 null genotype was correlated with a significantly increased risk of ESCC among Asians, but not among Caucasians and Africans. However, there was no significant correlation of GSTT1 null genotype with the risk of EADC. All results should be interpreted with caution. Only two African studies were recruited in the current meta-analysis, which may restrict statistical power to detect a real assessment in Africans. More large scale studies are needed to verify the results. Subgroup analyses were also performed regarding ethnicity for the GSTT1 null genotype. GSTT1 null genotype was correlated with a significantly increased risk of esophageal cancer among Asians, but not among Caucasians and Africans. The results were more robust on histological type of ESCC. This meta-analysis confirmed the mutual effect of GSTT1 null genotype in different populations to the risk of esophageal cancer. Possible explanations include: (1) significance of these enzymes may vary with the ethnicity genetic backgrounds, environmental exposures and histological types. The regional difference in the frequency of esophageal cancer is probably due to genetic polymorphism and variable exposure to environmental factors; (2) GSTs metabolize a variety of overlapping substrates and individuals lacking GSTT1 can also metabolize the carcinogens by other alternative GST enzymes. Furthermore, there was only one study concerning the association between GSTT1 null genotype and EADC on Asians. Thus, we failed to evaluate the potential role of GSTT1 null genotype in EADC risk in Asians due to the lack of available data to date. More case-control studies on the GSTT1 null genotype are encouraged, especially in Asians, for a better understanding the role of GSTT1 null genotype in the EADC development.
Some limitations must be acknowledged in the current meta-analysis. First, significant heterogeneity was observed between publications for GSTT1 null genotype. Potential sources of heterogeneity include the study design, publication year, ethnicity, country, histological type, sample size, and so on. When subgroup analyses were carried out according to ethnicity and histological type, this heterogeneity was reduced or removed in some subgroups, implying different effects on histological types and ethnic populations. These findings should be interpreted with very cautions. Second, our findings were based on unadjusted ORs and CIs, whereas a more precisely investigation could be performed if the sufficient individual data were available. Third, only two African studies were recruited in the current meta-analysis, the results in African population should be interpreted with caution. More large scale studies are needed to verify the results in Africans. Finally, due to lack of uniform individual-level data, further stratified analysis to measure any interactions between gene–gene variation and gene-metabolic traits was not performed.
In conclusion, our meta-analysis findings demonstrated that GSTT1 null genotype significantly increased esophageal cancer risk, particularly in Asians. In addition, GSTT1 null genotype was correlated with a significantly increased risk of ESCC, particularly among Asians. However, more studies are warranted to confirm or refute these correlations, particularly with respect to gene-gene and gene-environment interactions.
MATERIALS AND METHODS
Study selection
Pubmed, Embase, China National Knowledge Infrastructure (CNKI) and Wanfang databases (the search was updated in March 31, 2017) were searched using the following terms: ‘glutathione S-transferase T1’ or ‘GSTT1’, ‘polymorphism’ or ‘variant’, and ‘esophageal’ or ‘esophagus’, and ‘cancer’ or ‘carcinoma’ or ‘tumor’ or ‘malignancy’. The literature search was limited to English or Chinese articles. Additional publications were identified by a manual search based on references of retrieved studied or reviews.
Inclusion and exclusion criteria
The selection criteria were: (1) in a case-control study design, (2) studies that evaluated the relationship between the GSTT1 null genotype polymorphism and esophageal cancer, (3) usable data on genotype frequency. Thus, reports without usable data, reviews, comments and duplicated publications were excluded.
Data extraction
The data were collected by two independent reviewers. The extracted information contained: first author, year of publication, country of origin, ethnicity, number of cases and controls, genotyping method and characteristics of cases and controls. When come to conflicting assessment, disagreements were settled through a discussion among all authors.
Statistical analysis
The strength of correlation between GSTT1 null genotype and the susceptibility of esophageal cancer was assessed by the crude odds ratios (ORs) with 95% confidence intervals (CIs). A P < 0.05 (two-tailed) was considered as statistical significance. A Chi-square-based I2 test was used to detect heterogeneity [43] and an I2 < 25% indicates low heterogeneity, 25% ≤ I2 ≤ 50% indicates moderate heterogeneity, and I2 > 50% indicates large heterogeneity [44]. When I2 > 50% or P < 0.10 (two-sided), the random-effects model (the DerSimonian-Laird method) was utilized to pool the data [45], otherwise the fixed-effects model (the Mantel-Haenszel method) was used [46]. Subgourp analyses were conducted according to different ethnicity to identify the specific effects of heterogeneity. Publication bias was assessed by Begg’s funnel plot and Egger’s test [47]. Sensitivity analyses were conducted by one-way method. All statistical analyses were performed using STATA version 12.0 software (Stata Corporation, College Station, TX, USA).
Abbreviations
Glutathione-S-Transferase T1, GSTT1; China National Knowledge Infrastructure, CNKI; odds ratios, ORs; confidence intervals, CIs; esophageal squamous cell carcinoma, ESCC; esophageal adenocarcinoma, EADC.
Author contributions
FH, NZ and LZ designed the study. All authors were involved in article search, extraction, analysis and interpretation of data. FH drafted the manuscript, which was critically revised by NZ and LZ. All authors read and approved the final manuscript.
ACKNOWLEDGMENTS
We thank Dr. Haiyong Gu for his invaluable help. This study is supported in part by Changzhou High-Level Medical Talents Training Project (No: 2016CZBJ043) and The Applied Basic Research Programs of ChangZhou (No: CJ20159030).
CONFLICTS OF INTEREST
The authors report no conflicts of interest.
FUNDING
Changzhou High-Level Medical Talents Training Project (No: 2016CZBJ043). The Applied Basic Research Programs of ChangZhou (No: CJ20159030).
REFERENCES
1. Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011; 61:69–90. https://doi.org/10.3322/caac.20107.
2. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108. https://doi.org/10.3322/caac.21262.
3. Wang LD, Zhou FY, Li XM, Sun LD, Song X, Jin Y, Li JM, Kong GQ, Qi H, Cui J, Zhang LQ, Yang JZ, Li JL, et al. Genome-wide association study of esophageal squamous cell carcinoma in Chinese subjects identifies susceptibility loci at PLCE1 and C20orf54. Nat Genet. 2010; 42:759–63. https://doi.org/10.1038/ng.648.
4. Wu C, Kraft P, Zhai K, Chang J, Wang Z, Li Y, Hu Z, He Z, Jia W, Abnet CC, Liang L, Hu N, Miao X, et al. Genome-wide association analyses of esophageal squamous cell carcinoma in Chinese identify multiple susceptibility loci and gene-environment interactions. Nat Genet. 2012; 44:1090–7. https://doi.org/10.1038/ng.2411.
5. Hayes JD, Strange RC. Glutathione S-transferase polymorphisms and their biological consequences. Pharmacology. 2000; 61:154–66.
6. Xu D, Yan S, Yin J, Zhang P. Null genotype of GSTT1 contributes to colorectal cancer risk in Asian populations: evidence from a meta-analysis. Asian Pac J Cancer Prev. 2011; 12:2279–84.
7. Zhang ZJ, Hao K, Shi R, Zhao G, Jiang GX, Song Y, Xu X, Ma J. Glutathione S-transferase M1 (GSTM1) and glutathione S-transferase T1 (GSTT1) null polymorphisms, smoking, and their interaction in oral cancer: a HuGE review and meta-analysis. Am J Epidemiol. 2011; 173:847–57. https://doi.org/10.1093/aje/kwq480.
8. Cheng HY, You HY, Zhou TB. Relationship between GSTM1/GSTT1 null genotypes and renal cell carcinoma risk: a meta-analysis. Ren Fail. 2012; 34:1052–7. https://doi.org/10.3109/0886022X.2012.708380.
9. Sharma A, Das BC, Sehgal A, Mehrotra R, Kar P, Sardana S, Phukan R, Mahanta J, Purkayastha J, Saxena S, Kapur S, Chatterjee I, Sharma JK. GSTM1 and GSTT1 polymorphism and susceptibility to esophageal cancer in high- and low-risk regions of India. Tumour Biol. 2013; 34:3249–57. https://doi.org/10.1007/s13277-013-0897-6.
10. Talukdar FR, Ghosh SK, Laskar RS, Mondal R. Epigenetic, genetic and environmental interactions in esophageal squamous cell carcinoma from northeast India. PLoS One. 2013; 8:e60996. https://doi.org/10.1371/journal.pone.0060996.
11. Makhdoomi MA, Shah IA, Bhat GA, Amin S, Lone MM, Islami F, Dar NA. Association between GSTM1 and GSTT1 polymorphisms and esophageal squamous cell carcinoma: results from a case-control study in Kashmir, India. Tumour Biol. 2015; 36:2613–9. https://doi.org/10.1007/s13277-014-2882-0.
12. Weng Y, Fei B, He P, Cai M. Glutathione-S-transferase T1 polymorphism is associated with esophageal cancer risk in Chinese Han population. Asian Pac J Cancer Prev. 2012; 13:4403–7.
13. Yi SM, Li GY. Null genotype of GSTT1 contributes to esophageal cancer risk in Asian populations: evidence from a meta-analysis. Asian Pac J Cancer Prev. 2012; 13:4967–71.
14. Cai Y, Wang J. Significant association of glutathione S-transferase T1 null genotype with esophageal cancer risk: a meta-analysis. Mol Biol Rep. 2013; 40:2397–403. https://doi.org/10.1007/s11033-012-2320-6.
15. Lin DX, Tang YM, Peng Q, Lu SX, Ambrosone CB, Kadlubar FF. Susceptibility to esophageal cancer and genetic polymorphisms in glutathione S-transferases T1, P1, and M1 and cytochrome P450 2E1. Cancer Epidemiol Biomarkers Prev. 1998; 7:1013–8.
16. van Lieshout EM, Roelofs HM, Dekker S, Mulder CJ, Wobbes T, Jansen JB, Peters WH. Polymorphic expression of the glutathione S-transferase P1 gene and its susceptibility to Barrett’s esophagus and esophageal carcinoma. Cancer Res. 1999; 59:586–9.
17. Tan W, Song N, Wang GQ, Liu Q, Tang HJ, Kadlubar FF, Lin DX. Impact of genetic polymorphisms in cytochrome P450 2E1 and glutathione S-transferases M1, T1, and P1 on susceptibility to esophageal cancer among high-risk individuals in China. Cancer Epidemiol Biomarkers Prev. 2000; 9:551–6.
18. Gao CM, Takezaki T, Wu JZ, Li ZY, Liu YT, Li SP, Ding JH, Su P, Hu X, Xu TL, Sugimura H, Tajima K. Glutathione-S-transferases M1 (GSTM1) and GSTT1 genotype, smoking, consumption of alcohol and tea and risk of esophageal and stomach cancers: a case-control study of a high-incidence area in Jiangsu Province, China. Cancer Lett. 2002; 188:95–102.
19. Casson AG, Zheng Z, Chiasson D, MacDonald K, Riddell DC, Guernsey JR, Guernsey DL, McLaughlin J. Associations between genetic polymorphisms of Phase I and II metabolizing enzymes, p53 and susceptibility to esophageal adenocarcinoma. Cancer Detect Prev. 2003; 27:139–46.
20. Ribeiro Pinto LF, Teixeira Rossini AM, Albano RM, Felzenszwalb I, de Moura Gallo CV, Nunes RA, Andreollo NA. Mechanisms of esophageal cancer development in Brazilians. Mutat Res. 2003; 544:365–73.
21. Wang LD, Zheng S, Liu B, Zhou JX, Li YJ, Li JX. CYP1A1, GSTs and mEH polymorphisms and susceptibility to esophageal carcinoma: study of population from a high- incidence area in north China. World J Gastroenterol. 2003; 9:1394–7.
22. Abbas A, Delvinquiere K, Lechevrel M, Lebailly P, Gauduchon P, Launoy G, Sichel F. GSTM1, GSTT1, GSTP1 and CYP1A1 genetic polymorphisms and susceptibility to esophageal cancer in a French population: different pattern of squamous cell carcinoma and adenocarcinoma. World J Gastroenterol. 2004; 10:3389–93.
23. Roth MJ, Abnet CC, Johnson LL, Mark SD, Dong ZW, Taylor PR, Dawsey SM, Qiao YL. Polymorphic variation of Cyp1A1 is associated with the risk of gastric cardia cancer: a prospective case-cohort study of cytochrome P-450 1A1 and GST enzymes. Cancer Causes Control. 2004; 15:1077–83.
24. Yin LH, Pu YP, Song YH, Hu X, Liu YZ, Kai HT. Polymorphisms of susceptible genes for esophageal cancer risk in Huaian population in Jiangsu province. Tumor; 2005. p. 04.
25. Casson AG, Zheng Z, Porter GA, Guernsey DL. Genetic polymorphisms of microsomal epoxide hydroxylase and glutathione S-transferases M1, T1 and P1, interactions with smoking, and risk for esophageal (Barrett) adenocarcinoma. Cancer Detect Prev. 2006; 30:423–31. https://doi.org/10.1016/j.cdp.2006.09.005.
26. Jain M, Kumar S, Rastogi N, Lal P, Ghoshal UC, Tiwari A, Pant MC, Baiq MQ, Mittal B. GSTT1, GSTM1 and GSTP1 genetic polymorphisms and interaction with tobacco, alcohol and occupational exposure in esophageal cancer patients from North India. Cancer Lett. 2006; 242:60–7. https://doi.org/10.1016/j.canlet.2005.10.034.
27. Rossini A, Rapozo DC, Soares Lima SC, Guimaraes DP, Ferreira MA, Teixeira R, Kruel CD, Barros SG, Andreollo NA, Acatauassu R, Matos HJ, Albano RM, Pinto LF. Polymorphisms of GSTP1 and GSTT1, but not of CYP2A6, CYP2E1 or GSTM1, modify the risk for esophageal cancer in a western population. Carcinogenesis. 2007; 28:2537–42. https://doi.org/10.1093/carcin/bgm222.
28. Wideroff L, Vaughan TL, Farin FM, Gammon MD, Risch H, Stanford JL, Chow WH. GST, NAT1, CYP1A1 polymorphisms and risk of esophageal and gastric adenocarcinomas. Cancer Detect Prev. 2007; 31:233–6. https://doi.org/10.1016/j.cdp.2007.03.004.
29. Deng J, Guo RL, Yue HW, Huang ZG, Ma YX. A Case-control Study of the Polymorphisms of Phase I and Phase II Metabolic Genes and Esophageal Carcinoma Susceptibility. PJCCPVD; 2008. p. 07.
30. Zendehdel K, Bahmanyar S, McCarthy S, Nyren O, Andersson B, Ye W. Genetic polymorphisms of glutathione S-transferase genes GSTP1, GSTM1, and GSTT1 and risk of esophageal and gastric cardia cancers. Cancer Causes Control. 2009; 20:2031–8. https://doi.org/10.1007/s10552-009-9399-7.
31. Zhang LW, Sheyhidin I, Wu MB, Zhang Z, Li DS, Liu Z. Study on relations between genetic polymorphisms in CYP2C19, GSTT1 and risk of Kazakh’s esophageal cancer in Xinjiang. Journal of Practical Oncology. 2009; 03.
32. Ji R, Wu J, Zhou YN, Zhang B, Zhang ZY, Yang Z. Relationship between CYP1A1, GSTM1 and GSTT1 genetic polymorphisms and susceptibility of esophageal cancer in Wuwei, Gansu province. Journal of Lanzhou University (Medical Sciences). 2010; 02.
33. Li D, Dandara C, Parker MI. The 341C/T polymorphism in the GSTP1 gene is associated with increased risk of oesophageal cancer. BMC Genet. 2010; 11:47. https://doi.org/10.1186/1471-2156-11-47.
34. Liu R, Yin L, Pu Y, Li Y, Liang G, Zhang J, Li X. Functional alterations in the glutathione S-transferase family associated with enhanced occurrence of esophageal carcinoma in China. J Toxicol Environ Health A. 2010; 73:471–82. https://doi.org/10.1080/15287390903523394.
35. Malik MA, Upadhyay R, Mittal RD, Zargar SA, Mittal B. Association of xenobiotic metabolizing enzymes genetic polymorphisms with esophageal cancer in Kashmir Valley and influence of environmental factors. Nutr Cancer. 2010; 62:734–42. https://doi.org/10.1080/01635581003605904.
36. Moaven O, Raziee HR, Sima HR, Ganji A, Malekzadeh R, A’Rabi A, Abdollahi A, Memar B, Sotoudeh M, Naseh H, Nekoui N, Razavipour A, Gholamin M, et al. Interactions between Glutathione-S-Transferase M1, T1 and P1 polymorphisms and smoking, and increased susceptibility to esophageal squamous cell carcinoma. Cancer Epidemiol. 2010; 34:285–90. https://doi.org/10.1016/j.canep.2010.03.009.
37. Matejcic M, Li D, Prescott NJ, Lewis CM, Mathew CG, Parker MI. Association of a deletion of GSTT2B with an altered risk of oesophageal squamous cell carcinoma in a South African population: a case-control study. PLoS One. 2011; 6:e29366. https://doi.org/10.1371/journal.pone.0029366.
38. Gao P, Tian Y, Ye XF, Ge J, Zhang D, Xu WD. Study of CTPIA1, GSTT1, GSTM1 polymorphisms and susceptibility on esophageal carcinoma in Ningxia Hui nationality. Ningxia Med J. 2012; 03.
39. Djansugurova LB, Perfilyeva AV, Zhunusova GS, Djantaeva KB, Iksan OA, Khussainova EM. The determination of genetic markers of age-related cancer pathologies in populations from Kazakhstan. Front Genet. 2013; 4:70. https://doi.org/10.3389/fgene.2013.00070.
40. Dura P, Salomon J, Te Morsche RH, Roelofs HM, Kristinsson JO, Wobbes T, Witteman BJ, Tan AC, Drenth JP, Peters WH. No role for glutathione S-transferase genotypes in Caucasian esophageal squamous cell or adenocarcinoma etiology: an European case-control study. BMC Gastroenterol. 2013; 13:97. https://doi.org/10.1186/1471-230X-13-97.
41. Zhang L, Ma W, Li Y, Jiang YZ, Ma GY. Genetic polymorphisms of cytochrome P450 and glutathione S-transferase in relation to human esophageal carcinoma. Chin J Publ Health. 2013; 10:3.
42. Wu C, Li D, Jia W, Hu Z, Zhou Y, Yu D, Tong T, Wang M, Lin D, Qiao Y, Zhou Y, Chang J, Zhai K, et al. Genome-wide association study identifies common variants in SLC39A6 associated with length of survival in esophageal squamous-cell carcinoma. Nat Genet. 2013; 45:632–8. https://doi.org/10.1038/ng.2638.
43. Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002; 21:1539–58. https://doi.org/10.1002/sim.1186.
44. Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003; 327:557–60. https://doi.org/10.1136/bmj.327.7414.557.
45. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7:177–88.
46. Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959; 22:719–48.
47. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629–34.

