INTRODUCTION
Circular RNA (circRNA) is a novel kind of endogenous noncoding RNA that form circular structures without 5′-3′ polarities and polyadenylated tails [1]. In 2017, two reports identified that circRNAs had the translation function [2, 3]. CircRNAs are produced from pre-mRNA back-splicing and according to the circRNA biogenesis, intronic circRNA, exonic circRNA, sense overlapping circRNA, antisense circRNA and intergenic circRNA are the five types that have been reported [4–7]. CircRNAs are enriched, conserved and stable in mammalian cells [7, 8]. Recently, it was reported that circRNAs function as microRNA (miRNA) sponges that naturally sequester and competitively suppress miRNA activity [9]. It was demonstrated that circRNAs are involved in the development of several types of diseases, such as atherosclerosis, myocardial Infarction and neuron injury [10–12]. Indeed, there has been a special outpouring of concern for immense amounts of research shown that circRNAs are associated with many human cancers, such as hepatocellular carcinoma [13], esophageal squamous cell carcinoma [14], human gastric cancer [15], colorectal cancer [16]. Our previous study showed that circRNA alterations are involved in resistance to avian leucosis virus subgroup-J-induced tumor formation in chickens [17], however, no reports showed that circRNAs play a key role in chicken cancer which was induced by avian leucosis virus subgroup-J (ALV-J).
The avian leucosis virus belongs to the Alpharetrovirus genus of the Retrvoviridae family and induces sarcoma and leucosis in chickens [18]. Up to now, there are seven subgroups of ALV in chickens: ALV-A, -B, -C, -D, -E, -J and –K [19, 20]. ALV-J was first detected in meat-type chickens in 1987 as a new subgroup [21] and has spread to many countries such as America [22], UK [23], China [24] and so on. Clinical infection is associated with immune tolerance, high mortality, delayed growth, inducing different kinds of tumors [25, 26]. ALV-J can be transmitted vertically and horizontally, and horizontal transmission of ALV-J is more efficient than for other ALV subgroups [27]. ALV-J has caused great economic losses in the poultry industry worldwide [28]. Developed countries and China have made efforts to eradicate ALV-J from chicken breeding flocks in the poultry industry, unfortunately, an effective vaccine against ALV-J infection is not currently available [29, 30]. Breeding ALV-J-resistance chickens may be a new technique to reduce the severe economic cost which is caused by ALV-J and we have successfully cultivated ALV-J-resistance chickens [31].
Recently, we have identified that circRNAs alterations are involved in resistance to ALV-J-induced tumor formation in chickens [17], however, no reports showed that circRNAs aberrant expression also plays a key role in ALV-J-induced tumor formation. High-throughput RNA sequencing (RNA-seq) has been widely used to infer host gene expression in response to virus infection [32]. Herein, we confirm that enriched and differentially expressed circRNAs by RNA-seq in tumor liver samples of ALV-J-susceptible chickens. Further biological function prediction analysis was also conducted to develop a better understanding of the circRNAs function in ALV-J-induced tumor chickens.
RESULTS
Identification of differentially upregulated circRNAs in tumor liver samples of ALV-J-susceptible chickens
CircRNAs were sequenced in three tumor liver samples of ALV-J-susceptible chickens and three normal liver samples of ALV-J-resistance chickens. A total of 13419 circRNAs were detected after RNA-seq. Among those 13419 circRNAs, 2822 differentially expressed circRNAs (fold change ≥ 2, P ≤ 0.05, FDR < 0.05) were detected in both tumor liver samples and normal liver samples by Volcano Plot, including 2808 differentially upregulated circRNAs that were identified in tumor liver samples (Figure 1A). Scatter plot analysis made the variation of circRNAs expressions of two groups be visualized in these plots (Figure 1B). Hierarchical clustering analysis of randomly selected 24 differentially expressed circRNAs clearly screened them into two groups (Figure 2). After comparison with the genome sequence of chicken, total five types of circRNAs including exonic, intronic, sense overlapping, antisense and intergenic circRNAs were found, comprising 40.62%, 6.86%, 21.36%, 13.62% and 17.53%, respectively (Figure 3).
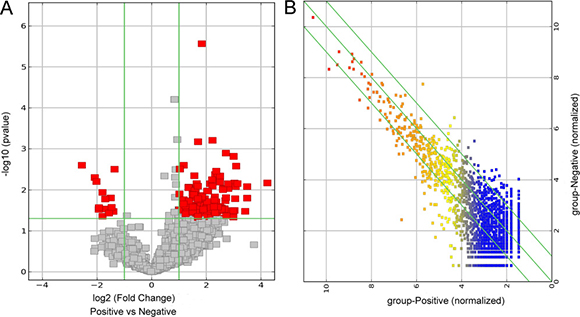
Figure 1: Bioinformatics analysis of differentially expressed circRNAs in tumor livers of ALV-J-susceptible chickens and normal livers of ALV-J-resistance chickens. (A) Volcano plot of differentially expressed circRNAs. (B) Scatter plot of differentially expressed circRNAs.
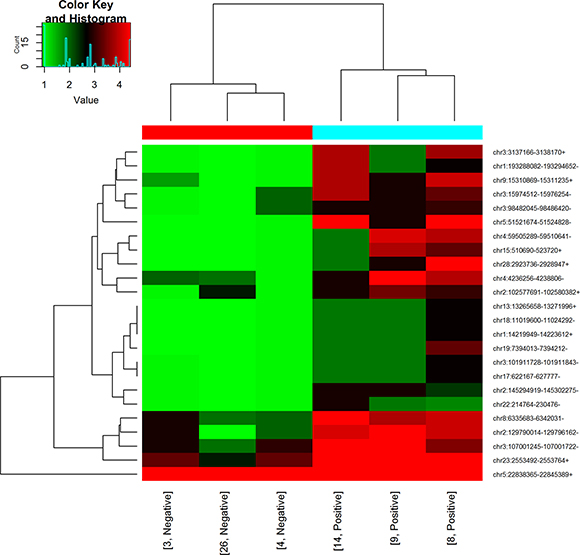
Figure 2: Hierarchical cluster analysis of circRNAs. circRNA expression profile of six RNA-seq samples was showed in red (upregulated expression) and green (downregulated expression) scale. The dendrogram shows the relationships among the expression levels of samples.
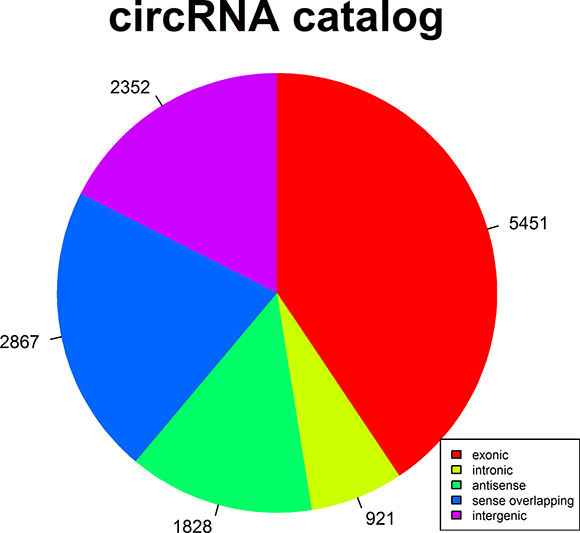
Figure 3:The percentage of five different types of circRNAs in chickens.
Validation of differentially expressed circRNAs using qRT-PCR
To validate the accuracy of RNA-seq data, a total of 4 differentially expressed circRNAs (chr21:4172114-4172476-, chr19:6585504-6585731+, chr9:22282452-22289593- and chr13:15110945-15122920-) were randomly selected and the quantitative real-time reverse transcription-polymerase chain reaction (qRT-PCR) was carried out in 3 tumor liver samples and 3 normal liver samples. QRT-PCR results of those selected differentially expressed circRNAs were consisted with RNA-seq data (Figure 4).
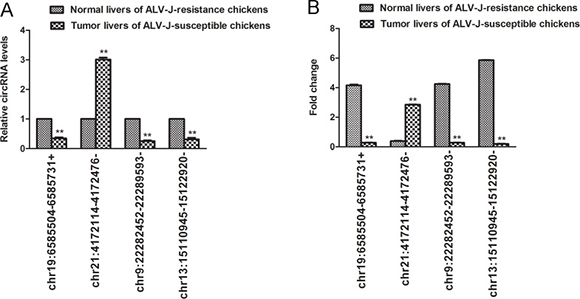
Figure 4: Validation of the expression level of four selected differentially expressed circRNAs by qRT-PCR. (A) Relative circRNA levels as indicated by RNA-seq; (B) Fold change for circRNA levels as indicated by qRT-PCR. Bars are mean ± SD (n = 3); **P < 0.05.
MiRNA response elements analysis of differentially upregulated circRNAs in tumor liver samples
For better understanding the detailed original information of circRNAs, 24 upregulated circRNAs in tumor liver samples of ALV-J-susceptible chickens were selected and ranked by fold change (FC), the 24 circRNAs were annotated in the gallus genome and the names of the best-predicted transcript for each of this 24 circRNAs were shown in Table 1.
Table 1: The 24 significantly upregulated circRNAs in tumor livers of ALV-J-susceptible chickens
CircRNA ID |
Fold Change |
P-value |
txStart |
txEnd riginal |
chrom |
Parent Gene |
|---|---|---|---|---|---|---|
chr15:510690-523720+ |
5.14 |
0.026 |
510689 |
523720 |
Chr15 |
MAPK1 |
chr5:51521674-51524828- |
11.26 |
0.008 |
51521673 |
51524828 |
Chr5 |
AKT1 |
chr28:2923736-2928947+ |
5.54 |
0.048 |
2923735 |
2928947 |
Chr28 |
STk11 |
chr4:4236256-4238806- |
4.25 |
0.022 |
4236255 |
4238806 |
Chr4 |
MAP7D3 |
chr18:11019600-11024292- |
2.83 |
0.026 |
11019599 |
11024292 |
Chr18 |
GRB2 |
chr3:3137166-3138170+ |
4.49 |
0.028 |
3137165 |
3138170 |
Chr3 |
MERTK |
chr1:14219949-14223612+ |
2.83 |
0.026 |
14219948 |
14223612 |
Chr1 |
PIK3CG |
chr2:129790014-129796162- |
4.95 |
0.015 |
129790013 |
129796162 |
Chr2 |
LRP12 |
chr19:73940137394212- |
3.24 |
0.044 |
7394012 |
7394212 |
Chr19 |
RPS6KB1 |
chr4:59505289-59510641- |
6.34 |
0.030 |
59505288 |
59510641 |
Chr4 |
EIF4E |
chr23:2553492-2553764+ |
5.00 |
0.004 |
2553491 |
2553764 |
Chr23 |
PTPRU |
chr9:15310869-15311235+ |
5.33 |
0.005 |
15310868 |
15311235 |
Chr9 |
AP2M1 |
chr17:622167-627777- |
2.25 |
0.017 |
622166 |
627777 |
Chr17 |
RABL6 |
chr3:101911728-101911843- |
2.25 |
0.017 |
101911727 |
101911843 |
Chr3 |
APOB |
chr22:214764-230476- |
2.25 |
0.029 |
214763 |
230476 |
Chr22 |
ANTXR1 |
chr5:2283836522845389+ |
2.84 |
0.026 |
22815788 |
22883775 |
Chr5 |
AMBRA1 |
chr3:98482045-98486420- |
3.05 |
0.011 |
98482044 |
98486420 |
Chr3 |
NBAS |
chr2:102577691-102580382+ |
3.05 |
0.038 |
102577690 |
102580382 |
Chr2 |
CABLES1 |
chr3:107001245-107001722- |
3.10 |
0.027 |
107001244 |
107001722 |
Chr3 |
CTSB |
chr13:13265658-13271996+ |
2.83 |
0.026 |
13265657 |
13271996 |
Chr13 |
MAPK9 |
chr2:145294919-145302275- |
3.24 |
0.001 |
145294918 |
145302275 |
Chr2 |
PTK2 |
chr1:193288082193294652- |
3.57 |
0.033 |
193288081 |
193294652 |
Chr1 |
UVRAG |
chr3:15974512-15976254- |
4.08 |
0.011 |
15974511 |
15976254 |
Chr3 |
SOS1 |
chr8:6335683-6342031- |
6.60 |
0.001 |
6350257 |
6368257 |
Chr8 |
RALGPS2 |
The homology of chicken circRNAs
Comparing chicken circRNAs with those of mouse and humans, homology analysis results showed that 8564 (63.82%) chicken circRNAs were homologous with humans, 8278 (61.69%) were homologous with the mouse. Importantly, a total of 8024 chicken circRNAs (59.80%) showed homology with mouse and humans (Figure 5).
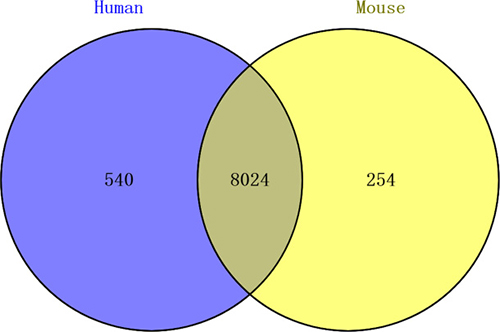
Figure 5: Chicken circRNAs shared homology with mouse and humans.
MiRNA binding sites prediction and circRNA-targeted miRNA-gene network identification
Because that circRNAs as an active miRNA sponge regulator, we then listed top 5 possible miRNA targets for each circRNA which was up-regulated in tumor liver samples of ALV-J-susceptible chickens in Table 2. CircRNA-targeted miRNA-gene network was constructed using Targetscan and miRanda. CircRNAs contain many miRNA response elements (MREs) [33], we then selected the top 5 predicted miRNA targets of 3 randomly differentially upregulated circRNAs (chr5: 51521674-51524828, chr3: 3137166-3138170 +, chr4: 59505289-59510641-) in tumor liver samples, meanwhile, the top 10 target genes for each miRNA were collected. Finally, the integrated circRNA-targeted miRNA-gene network was determined (Figure 6).
Table 2: Predicted miRNAs for 24 differentially upregulated circRNAs in tumor livers of ALV-J-susceptible chickens
CircRNA ID |
miRNA |
miRNA |
miRNA |
miRNA |
miRNA |
|---|---|---|---|---|---|
chr15:510690-523720+ |
gga-miR-6640-5p |
gga-miR-106-3p |
gga-miR-1551-5p |
gga-miR-1688 |
gga-miR-1793 |
chr5:51521674-51524828- |
gga-miR-6675-3p |
gga-miR-7463-5p |
gga-miR-1643-5p |
gga-miR-1698 |
gga-miR-456-5p |
chr28:2923736-2928947+ |
gga-miR-20a-3p |
gga-miR-6665-5p |
gga-miR-1730-3p |
gga-miR-6646-3p |
gga-miR-6623-3p |
chr4:4236256-4238806- |
gga-miR-1651-3p |
gga-miR-7450-5p |
gga-miR-1600 |
gga-miR-1748 |
gga-miR-1641 |
chr18:11019600-11024292- |
gga-miR-6573-5p |
gga-miR-6619-5p |
gga-miR-1573 |
gga-miR-187-3p |
gga-miR-1785 |
chr3:3137166-3138170+ |
gga-miR-7449-3p |
gga-miR-6663-5p |
gga-miR-1586 |
gga-miR-1803 |
gga-miR-1464 |
chr1:14219949-14223612+ |
gga-miR-1627-5p |
gga-miR-1799 |
gga-miR-6545-5p |
gga-miR-1753 |
gga-miR-6552-3p |
chr2:129790014-129796162- |
gga-miR-1806 |
gga-miR-6678-3p |
gga-miR-1565 |
gga-miR-211 |
gga-miR-204 |
chr19:7394013-7394212- |
gga-miR-2128 |
gga-miR-1662 |
gga-miR-1811 |
gga-miR-184-5p |
gga-miR-1598 |
chr4:59505289-59510641- |
gga-miR-757 |
gga-miR-29b-1-5p |
gga-miR-1705 |
gga-miR-1804 |
gga-miR-1762 |
chr23:2553492-2553764+ |
gga-miR-6698-3p |
gga-miR-20b-5p |
gga-miR-17-5p |
gga-miR-106-5p |
gga-miR-6587-5p |
chr9:15310869-15311235+ |
gga-miR-6578-3p |
gga-miR-6700-3p |
gga-miR-6552-5p |
gga-miR-7473-5p |
gga-miR-6582-3p |
chr17:622167-627777- |
gga-miR-140-5p |
gga-miR-1744-3p |
gga-miR-1654 |
gga-miR-302b-5p |
gga-miR-1815 |
chr3:101911728-101911843- |
gga-miR-6551-3p |
gga-miR-7450-3p |
gga-miR-30e-5p |
gga-miR-30c-5p |
gga-miR-30b-5p |
chr22:214764-230476- |
gga-miR-148a-5p |
gga-miR-6584-5p |
gga-miR-1611 |
gga-miR-205b |
gga-miR-1626-3p |
chr5:22838365-22845389+ |
gga-miR-7448-5p |
gga-miR-6679-5p |
gga-miR-4732-5p |
gga-miR-1762 |
gga-miR-200b-3p |
chr3:98482045-98486420- |
gga-miR-7450-5p |
gga-miR-6673-3p |
gga-miR-6655-5p |
gga-miR-6577-5p |
gga-miR-7464-5p |
chr2:102577691-102580382+ |
gga-miR-1754-5p |
gga-miR-6673-3p |
gga-miR-148a-5p |
gga-miR-7470-5p |
gga-miR-1662 |
chr3:107001245-107001722- |
gga-miR-383-3p |
gga-miR-6607-5p |
gga-miR-1756b |
gga-miR-1665 |
gga-miR-1608 |
chr13:13265658-13271996+ |
gga-miR-190a-5p |
gga-miR-365-1-5p |
gga-miR-7483-5p |
gga-miR-217-3p |
gga-miR-146b-3p |
chr2:145294919-145302275- |
gga-miR-29b-1-5p |
gga-miR-29b-2-5p |
gga-miR-1656 |
gga-miR-1716 |
gga-miR-9-5p |
chr1:193288082-193294652- |
gga-miR-1564-3p |
gga-miR-7479-3p |
gga-miR-1416-3p |
gga-miR-1731-5p |
gga-miR-9-5p |
chr3:15974512-15976254- |
gga-miR-19b-5p |
gga-miR-19a-5p |
gga-miR-23b-5p |
gga-miR-1710 |
gga-miR-6577-5p |
chr8:6335683-6342031- |
gga-miR-7481-3p |
gga-miR-1685-5p |
gga-miR-7462-5p |
gga-miR-1645 |
gga-miR-7483-3p |
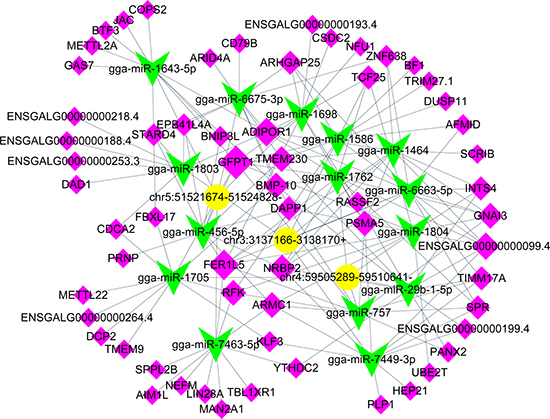
Figure 6: circRNA-miRNA-gene network for three selected upregulated circRNAs in tumor livers of ALV-J-susceptible chickens. Yellow represents circRNAs, green represents miRNAs and pink represents gene.
Functional annotations for miRNA target genes
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of the predicted miRNA target genes which were predicted to be targeted by the 24 differentially upregulated circRNAs in tumor liver samples of ALV-J-susceptible chickens showed that those miRNA target genes were involved in cysteine and methionine metabolism pathway (Figure 7). Furthermore, Gene Ontology (GO) annotation of biological process (BP) of those miRNA target genes participated in mRNA metabolic process, cell development, mRNA catabolic process, phospholipid metabolic process, regulation of cell projection organization, regulation of cellular component size, regulation of cellular component organization, regulation of transmembrane transporter activity, regulation of cell projection assembly and mRNA processing (Figure 8A). GO annotation of cellular component (CC) of those miRNA target genes participated in cell junction and collagen trimer (Figure 8B), while for GO annotation of molecular function (MF) of those miRNA target genes participated in nucleoside phosphate binding, nucleotide binding, small molecule binding, DNA helicase activity, adenyl nucleotide binding, adenyl ribonucleotide binding, binding, cytoskeletal protein binding, ATP binding and RNA binding (Figure 8C).
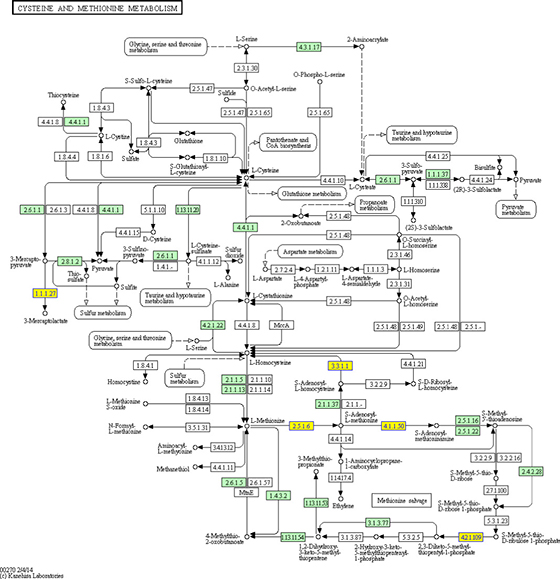
Figure 7: KEGG pathway analysis for the miRNA target genes of 24 upregulated circRNAs in tumor livers of ALV-J-susceptible chickens. The yellow boxes indicate the genes targeted by differentially expressed circRNAs.
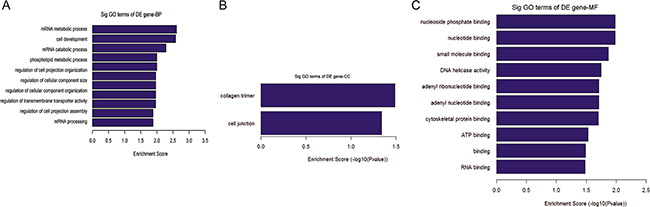
Figure 8: Go analysis of the predicted miRNA target genes. Go analysis of the predicted miRNA target genes which were predicted to be targeted by those 24 upregulated circRNAs in tumor liver samples of ALV-J-susceptible chickens according to the values in the enrichment score under the theme of BP, CC and MF.
DISCUSSION
CircRNA expression has the cell-type-specific, tissue-specific or developmental-stage specific manner [34, 35]. To date, several studies have proved that circRNAs may regulate gene expression by manipulating miRNAs through its miRNA sponges function [36], and circRNAs also compete with mRNAs for miRNA binding in the cytoplasm and interfere in gene regulation [8]. MiRNAs control a large set of biological processes through direct interaction with their target mRNA, so circRNA sponge activity will also affect these pathways [37, 38]. Importantly, circRNAs have become the new markers for diseases including cancers [10, 39].
RNA-seq was used for discovering new transcript and genes expression levels including circRNAs expression in many cancer research [40, 41, 42]. ALV-J mainly induces myelocytomatosis in chickens and can cause other reproduction problems in the poultry industry worldwide [43]. In the present study, we want to explore whether circRNAs are involved in ALV-J-induced tumor formation or not. Interestingly, totally 13419 circRNAs were detected using RNA-seq in tumor liver samples of ALV-J-susceptible chickens and normal liver samples of ALV-J-resistance chickens, totally 2822 differentially expressed circRNAs (fold change ≥ 2, P ≤ 0.05, FDR < 0.05) were detected in both of tumor liver samples and normal liver samples, while 2808 differentially upregulated circRNAs were identified in tumor liver samples, the different expression patterns implied circRNAs may play a key role in tumor formation which is induced by ALV-J. QRT-PCR was a necessary tool to valid the RNA-seq data in many studies [44, 45]. Here, in our study, we confirmed the differentially expressed circRNAs from RNA-seq data by qRT-PCR. As expected, qRT-PCR results of randomly selected 4 differentially expressed circRNAs had the same expression level trend compared with that from RNA-seq data, then we were sure that the RNA-seq results were reliable.
For better understanding the function of miRNA target genes, the additional functional analysis was conducted using the GO annotation and KEGG pathway. Statistical analysis of miRNA target genes was used to determine which GO terms or KEGG pathways were significantly enriched in the tumor livers samples as compared to the normal livers samples. These enriched categories along with the number of miRNA target genes identified for biological process, cellular component, molecular function three aspects. Interestingly, GO term of biological process and KEGG pathway analysis both showed that miRNA target genes are involved in the metabolic process, that implied metabolism make an important function in tumor liver formation induced by ALV-J. We suspected that maybe those up-regulated circRNAs changed the status and then caused the physical disorder and finally result in tumor formation. We also conducted homology analysis of circRNAs among chickens, mouse and humans to explore whether differentially expressed circRNAs of tumorigenesis in chickens may also involve in tumorigenesis in humans. It is worth mentioning that homology analysis results showed that 63.82% and 61.69% of chicken circRNAs were homologous with humans and mouse respectively, while a total of 59.80% chicken circRNAs showed homology with mouse and humans, that implies those differentially expressed circRNAs of tumor livers in chickens may also involve in tumor formation in humans.
In recent years, circRNA-miRNA-gene regulatory axes were confirmed that involved in pathogenic processes or cancer [46, 47]. In order to show a more intuitionistic result, three up-regulated circRNAs (chr5: 51521674-51524828-, chr3: 3137166-3138170+, chr4: 59505289-59510641-) in tumor liver samples of ALV-J-susceptible chickens were selected and five target miRNAs and 10 miRNA target genes were showed in the circRNA-miRNA-gene network which provides a foundation for further pathogenic mechanism research.
In conclusion, our study for the first time reveals that circRNAs are involved in ALV-J-induced tumor formation in chickens. We totally identified 2808 differentially expressed upregulated circRNAs in tumor livers of ALV-J-susceptible chickens. Based on miRNA sponge function, top 5 miRNA targets for 24 selected upregulated circRNAs in ALV-J-induced tumor livers were listed according to the match score. We also constructed the circRNA-miRNA-gene network, furthermore, GO terms and KEGG pathway analyses to point out that metabolism plays a key role in circRNAs alterations of ALV-J-induced tumor livers. Homology analysis results showed that totally 59.80% chicken circRNAs showed homology with mouse and humans, that opens a scenario for the investigation of the role of those differentially expressed chicken circRNA in human tumors. However, more underlying mechanism research still needs to be done to reveal which circRNAs alteration are directly involved in ALV-J-induced tumor formation in chickens.
MATERIALS AND METHODS
Ethics statement
All experiments were carried out in strict accordance with the recommendations of the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The use of animals in this study was approved by South China Agricultural University Committee of Animal Experiments (approval ID: 201004152).
RNA-seq specimens
Tumor livers of ALV-J-susceptible chickens (the model of ALV-J-induced tumor chickens) and normal livers of ALV-J-resistance chickens (the model of ALV-J-infected negative chickens) in the F3 generation at 20 weeks after challenge with ALV-J NX0101 strain which induced myeloid tumors were collected for RNA-seq [31]. In each group, three livers were collected as three duplicates.
RNA library synthesis and RNA-seq
Total RNAs were extracted from tumor livers and normal livers using TRizol reagent (Invitrogen, Karlsruhe, Germany) according to the manufacturer’s instructions. The purity and concentration of the RNA samples were determined by assessing the OD260/280 using a NanoDrop ND-2000 instrument (Thermo, Waltham, MA, USA). To enrich circRNAs, RNase R (Epicentre, Madison, WI) was used to digest the extracted total RNAs to remove linear RNAs [48]. RNA libraries were constructed from the treated RNAs using the TruSeq Stranded Total RNA Library Prep Kit (Illumina, San Diego, CA, USA) according to the manufacturer’s instructions. Libraries were controlled for quality and quantity according to the BioAnalyzer 2100 system (Agilent Technologies, Inc., Santa Clara, CA, USA) and then was subjected to paired-end sequencing for 150 cycles by HiSeq 4000 Sequencer according to the manufacturer’s instructions (Illumina, SanDiego, CA). The paired-end reads were harvested and subject to quality control using Q30 [49]. After 3′ adaptor-trimming and the removal of low quality reads using Cutadapt software [50], the high quality trimmed reads were used for analysis of circRNAs. The differential expressions of circRNAs between the two groups were detected through fold-change filtering and Student’s t-test. CircRNAs exhibiting fold changes ≥ 2.0 and P-values ≤ 0.05 were selected as significantly differentially expressed circRNAs.
Quantitative real-time PCR
QRT-PCR was achieved using PrimeScript RT Reagent Kit (Perfect Real Time; TaKaRa, Osaka, Japan) on a Mx3005P real-time PCR System (Stratagene, La Jolla, CA) with SYBR Green qPCR SMix (ROX; Roche) following the manufacturer’s instructions. Divergent primers, rather than the more commonly used convergent primers, of chr21:4172114-4172476-, chr19:6585504-6585731+, chr9:22282452-22289593- and chr13:15110945-15122920- were designed for further test the accuracy of RNA-seq. Primers for above four randomly selected differentially expressed circRNAs and β-actin were synthesized by Sangon Biotech (Shanghai, China) and were listed in Table 3. The 2−ΔΔCt method was used to analyze the qRT-PCR results [46].
Table 3: The primers were used for qRT-PCR in this study
CircRNA |
Primers |
|---|---|
chr21:4172114-4172476- |
F: GGGCTTGTTTCTGGTTTGGGTTAG |
chr19:6585504-6585731+ |
F: GGTGGACTCCTTCTGGATGTTGTAGT |
chr9:22282452-22289593- |
F: CTGATCCCGAGACATCTAATGTTTCCT |
chr13:15110945-15122920- |
F: CAGGGGTCTGCCAGCACATAGT |
β-actin |
R: GCTCAAGGAGATGGGTTTTCCG |
Bioinformatics analysis
CircRNA and miRNA interactions were predicted using customized Arraystar miRNA target prediction software based on TargetScan and miRanda [51, 52]; the top five putative target miRNAs were identified for upregulated circRNAs in tumor liver samples of ALV-J-susceptible chickens. The putative target genes of these miRNAs were identified using Targetscan [51]. Cytoscape software was used to construct the circRNA-miRNA-gene networks [53]. Homology analysis of chicken circRNA was conducted using blastn (v2.2.27) from the alignment between human and mouse [54]. Gene ontology and KEGG pathway analysis were performed for the differentially expressed circRNA-associated genes [55, 56].
Statistical analysis
Data were processed using GraphPad Prism (version 5.0) and are expressed as the mean ± SE. The Student’s t-test was used to assess differences among groups, where P ≤ 0.05 was considered to show significant differences between the groups.
Abbreviations
circRNA, circular RNA; RNA sequencing, RNA-seq; ALV-J, subgroup J avian leucosis virus; miRNA, microRNA; qRT-PCR, quantitative real-time polymerase chain reaction; FC, fold change; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; BP, biological process; CC, cellular component; MF, molecular function.
ACKNOWLEDGMENTS
The authors would like to thank Professor Huanmin Zhang (USDA, Agriculture Research Service, Avian Disease and Oncology Laboratory) for providing guidance on experimental design and thank Professor Aijun Li for providing guidance on data analysis.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
GRANT SUPPORT
This study was supported by the National Natural Science Foundation of China (Grant Nos. 31672564, 31472217), Construction project of modern agricultural science and technology innovation alliance in Guangdong Province (2016LM1112) , International Science and technology cooperation project of Guangdong Province (2016A050502042) and the Natural Science Foundation of Guangdong Province (Grant No. S2013030013313).
REFERENCES
1. Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015; 12:381–388.
2. Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E, Shenzis S, Samson M, Dittmar G, et al. Translation of CircRNAs. Mol Cell. 2017; 66:9–21.
3. Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, Fatica A, Santini T, Andronache A, Wade M, Laneve P, Rajewsky N, Bozzoni I. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol Cell. 2017; 66:22–37.
4. Zhang Y, Xue W, Li X, Zhang J, Chen S, Zhang JL, Yang L, Chen LL. The Biogenesis of Nascent Circular RNAs. Cell Rep. 2016; 15:611–624.
5. Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, Zhu S, Yang L, Chen LL. Circular intronic long noncoding RNAs. Mol Cell. 2013; 51:792–806.
6. Danan M, Schwartz S, Edelheit S, Sorek R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 2012; 40:3131–3142.
7. Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013; 495:333–38.
8. Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013; 19:141–57.
9. Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J. Natural RNA circles function as efficient microRNA sponges. Nature 2013; 495:384–388.
10. Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet. 2010; 6:e1001233.
11. Geng HH, Li R, Su YM, Xiao J, Pan M, Cai XX, Ji XP. The Circular RNA Cdr1as Promotes Myocardial Infarction by Mediating the Regulation of miR-7a on Its Target Genes Expression. PLoS One. 2016; 11:e0151753.
12. Lin SP, Ye S, Long Y, Fan Y, Mao HF, Chen MT, Ma QJ. Circular RNA expression alterations are involved in OGD/R-induced neuron injury. Biochem Biophys Res Commun. 2016; 471:52–56.
13. Qin M, Liu G, Huo X, Tao X, Sun X, Ge Z, Yang J, Fan J, Liu L, Qin W. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016; 16:161–169.
14. Xia W, Qiu M, Chen R, Wang S, Leng X, Wang J, Xu Y, Hu J, Dong G, Xu PL, Yin R. Circular RNA has_circ_0067934 is upregulated in esophageal squamous cell carcinoma and promoted proliferation. Sci Rep. 2016; 6:35576.
15. Shao Y, Li J, Lu R, Li T, Yang Y, Xiao B, Guo J. Global circular RNA expression profile of human gastric cancer and its clinical significance. Cancer Med. 2017; 6:1173–1180.
16. Xiong W, Ai YQ, Li YF, Ye Q, Chen ZT, Qin JY, Liu QY, Wang H, Ju YH, Li WH, Li YF. Microarray Analysis of Circular RNA Expression Profile Associated with 5-Fluorouracil-Based Chemoradiation Resistance in Colorectal Cancer Cells. Biomed Res Int. 2017; 2017:8421614.
17. Zhang X, Yan Y, Lei X, Li A, Zhang H, Dai Z, Li X, Chen W, Lin W, Chen F, Ma J, Xie Q. Circular RNA alterations are involved in resistance to avian leukosis virus subgroup-J-induced tumor formation in chickens. Oncotarget. 2017; 8:34961–70. https://doi.org/10.18632/oncotarget.16442.
18. Cui Z, Sun S, Zhang Z, Meng S. Simultaneous endemic infections with subgroup J avian leukosis virus and reticuloendotheliosis virus in commercial and local breeds of chickens. Avian Pathol. 2009; 38:443–448.
19. Lin W, Li X, Dai Z, Zhang X, Chang S, Zhao P, Zhang H, Chen F, Xie Q. Molecular epidemiology of J-subgroup avian leukosis virus isolated from meat-type chickens in southern China between 2013 and 2014. Arch Virol. 2016; 161:3039–3046.
20. Li X, Lin W, Chang S, Zhao P, Zhang X, Liu Y, Chen W, Li B, Shu D, Zhang H, Chen F, Xie Q. Isolation, identification and evolution analysis of a novel subgroup of avian leukosis virus isolated from a local Chinese yellow broiler in South China. Arch Virol. 2016; 161:2717–2725.
21. Payne LN. HPRS-103: a retorovirus strikes back. The emergence of J avian leukosis virus. Avian Pathol. 1998; 27:36–45.
22. Zavala G, Cheng S, Jackwood MW. Molecular epidemiology of avian leucosis virus subgroup J and evolutionary history of its 3’ untranslated region. Avian Dis. 2007; 51:942–953.
23. Venugopal K, Howes K, Flannery DM, Payne LN. Subgroup J avian leukosis virus infection in turkeys: induction of rapid onset tumours by acutely transforming virus strain 966. Avian Pathol. 2000; 29:319–25.
24. Ji J, Li H, Zhang H, Xie Q, Chang S, Shang H, Ma J, Bi Y. Complete genome sequence of an avian leukosis virus isolate associated with hemangioma and myeloid leukosis in egg-type and meat-type chickens. J Virol. 2012; 86:10907–08.
25. Payne LN. Retrovirus-induced disease in poultry. Poult Sci. 1998; 77:1204–1212.
26. Sironi G, Manarolla G, Pisoni G, Recordati C, Rampin T. Myotropic avian leukosis virus subgroup J infection in a chicken. J Vet Med B Infect Dis Vet Public Health. 2006; 53:347–349.
27. Lin Y, Xia J, Zhao Y, Wang F, Yu S, Zou N, Wen X, Cao S, Huang Y. Reproduction of hemangioma by infection with subgroup J avian leukosis virus: the vertical transmission is more hazardous than the horizontal way. Virol J. 2013; 10:97.
28. Shen Y, Cai L, Wang Y, Wei R, He M, Wang S, Wang G, Cheng Z. Genetic mutations of avian leukosis virus subgroup J strains extended their host Range. J Gen Virol. 2014; 95:691–699.
29. Nakamura K, Ogiso M, Tsukamoto K, Hamazaki N, Hihara H, Yuasa N. Lesions of bone and bone marrow in myeloid leukosis occuring naturally in adult broiler breeders. Avian Dis. 2000; 44:215–221.
30. Li Y, Liu X, Yang Z, Xu C, Liu D, Qin J, Dai M, Hao J, Feng M, Huang X, Tan L, Cao W, Liao M. The MYC, TERT, and ZIC1 genes are common targets of viral integration and transcriptional deregulation in avian leukosis virus subgroup J-induced myeloid leukosis. J Virol. 2014; 88:3182–3191.
31. Zhang X, Yan Z, Li X, Lin W, Dai Z, Yan Y, Lu P, Chen W, Zhang H, Chen F, Ma J, Xie Q. GADD45β, an anti-tumor gene, inhibits avian leukosis virus subgroup J replication in chickens. Oncotarget. 2016; 7:68883–93. https://doi.org/10.18632/oncotarget.12027.
32. Hamzić E, Kjærup RB, Mach N, Minozzi G, Strozzi F, Gualdi V, Williams JL, Chen J, Wattrang E, Buitenhuis B, Juul-Madsen HR, Dalgaard TS. RNA sequencing-based analysis of the spleen transcriptome following infectious bronchitis virus infection of chickens selected for different mannose-binding lectin serum concentrations. BMC Genomics. 2016; 17:82.
33. Qu S, Yang X, Li X, Wang J, Gao Y, Shuang R, Sun W, Dou K, Li H. Circular RNA: A new star of noncoding RNAs. Cancer Lett. 2015; 365:141–148.
34. Yao T, Chen Q, Fu L, Guo J. Circular RNAs: biogenesis, properties, roles, and their relationships with liver diseases. Hepatol Res. 2017; 47:497–504.
35. Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO. Cell-type specific features of circular RNA expression. PLoS Genet. 2013; 9:e1003777.
36. Liu Q, Zhang X, Hu X, Dai L, Fu X, Zhang J, Ao Y. Circular RNA Related to the Chondrocyte ECM Regulates MMP13 Expression by Functioning as a MiR-136 ‘Sponge’ in Human Cartilage Degradation. Sci Rep. 2016; 6:22572.
37. Hutvagner G, Simard MJ. Argonaute proteins: key players in RNA silencing. Nat Rev Mol Cell Biol. 2008; 9:22–32.
38. Kulcheski FR, Christoff AP, Margis R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J Biotechnol. 2016; 238:42–51.
39. Xuan L, Qu L, Zhou H, Wang P, Yu H, Wu T, Wang X, Li Q, Tian L, Liu M, Sun Y. Circular RNA: a novel biomarker for progressive laryngeal cancer. Am J Transl Res. 2016; 8:932–939.
40. Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008; 5:621–28.
41. Nagalakshmi U, Wang Z, Waern K, Shou C, Raha D, Gerstein M, Snyder M. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science. 2008; 320:1344–1349.
42. Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G, Liang L, Gu J, He X, et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun. 2016; 7:11215.
43. Gao Y, Yun B, Qin L, Pan W, Qu Y, Liu Z, Wang Y, Qi X, Gao H, Wang X. Molecular Epidemiology of Avian Leukosis Virus Subgroup J in Layer Flocks in China. J Clin Microbiol. 2012; 50:953–960.
44. Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J, Chen D, Gu J, He X, Huang S. Circular RNA is enriched and stable in exosomes: a promising biomarker for call diagnosis. Cell Res. 2015; 25:981–984.
45. Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, Mesteri I, Grunt TW, Zeillinger R, Pils D. Correlation of circular RNA abundance with proliferation - exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015; 5:8057.
46. Cao S, Wei D, Li X, Zhou J, Li W, Qian Y, Wang Z, Li G, Pan X, Lei D. Novel circular RNA expression profiles reflect progression of patients with hypopharyngeal squamous cell carcinoma. Oncotarget. 2017; 8:45367–79. https://doi.org/10.18632/oncotarget.17488.
47. Rong D, Sun H, Li Z, Liu S, Dong C, Fu K, Tang W, Cao H. An emerging function of circRNA-miRNAs-mRNA axis in human diseases. Oncotarget. 2017; 8:73271–81. https://doi.org/10.18632/oncotarget.19154.
48. Shang X, Li G, Liu H, Li T, Liu J, Zhao Q, Wang C. Comprehensive Circular RNA Profiling Reveals That hsa_circ_0005075, a New Circular RNA Biomarker, Is Involved in Hepatocellular Crcinoma Development. Medicine (Baltimore). 2016; 95:e3811.
49. Ewing B, Hillier L, Wendl MC, Green P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 1998; 8:175–185.
50. Martin M. Cutadapt removes adapter sequences from high-throughput sequencingreads. EMBnet J. 2001; 17:10–12.
51. Enright AJ, John B, Gaul U, Tuschl T, Sander C, Marks DS. MicroRNA targets in Drosophila. Genome Biol. 2003; 5:R1.
52. Pasquinelli AE. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet. 2012; 13:271–82.
53. Smoot ME, Ono K, Ruscheinski J, Wang PL, Ideker T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics. 2011; 27:431–432.
54. Zhao W, Cheng Y, Zhang C, You Q, Shen X, Guo W, Jiao Y. Genome-wide identification and characterization of circular RNAs by high throughput sequencing in soybean. Sci Rep. 2017; 7:5636.
55. Xing Z, Chu C, Chen L, Kong X. The use of Gene Ontology terms and KEGG pathways for analysis and prediction of oncogenes. Biochim Biophys Acta. 2016; 1860:2725–2734.
56. Kanehisa M, Sato Y, Morishima K. BlastKOALA and GhostKOALA: KEGG Tools for Functional Characterization of Genome and Metagenome Sequences. J Mol Biol. 2016; 428:726–731.

