INTRODUCTION
Lung cancer is a leading cause of cancer-related death worldwide, with a 5 year survival rate of about 15% [1]. Non-small cell lung cancer (NSCLC) accounts for approximately 80% of lung cancers, whose ravages has intensely hijacked welfare of human beings for the tremendous burden on health and economy [2, 3]. However, the complexity of the pathogenic mechanisms involved means that effective therapies are difficult to design; therefore, a deeper understanding of the molecular pathways underlying such malignancy is essential [4, 5].
Cancer possesses several malignant hallmarks caused by disorganized cellular signaling networks, owing to the dysfunction of critical genes in cellular processes [6]. Recently, attributing to unique physiochemical properties like flexibility in recognizing ability and prevalence in distributing, long non-coding RNAs (lncRNAs) have emerged to have pivotal function in cancer biology mainly via regulating oncogenes or tumor suppressor genes [7–10]. Moreover, a myriad of studies highlighted the relationship between lncRNAs and NSCLC, and suggested that lncRNAs could play a momentous role in modulating malignant progression, presented to be a novel breakpoint to handle NSCLC [11, 12].
However, a single study involved a single lncRNA was not a strong platform on which to base an evaluation of the clinical value of lncRNAs in NSCLC. Therefore, we undertook a systematic review of the relationship between lncRNAs and the clinicopathological characteristics of NSCLC. We found that expression of lncRNA was associated with prognosis, tumor size, and lymph node metastasis. Furthermore, lncRNAs in serum may have utility as a diagnostic biomarker for NSCLC. Last, we summarized the molecular mechanisms about these lncRNAs in NSCLC. Taken together, these findings highlighted the clinical value of lncRNAs in NSCLC and promoted research into how lncRNAs modulate carcinogenesis of NSCLC.
RESULTS
Data extraction
Up to 392 publications were identified. After excluding duplicates, the titles and abstracts of 137 studies were reviewed and irrelevant studies were excluded. Full text versions of the remaining 124 studies were examined, and those with insufficient data were excluded. Next, 59 studies were subjected to quality assessment. Finally, 37 studies were subjected to systematic review (Figure 1).
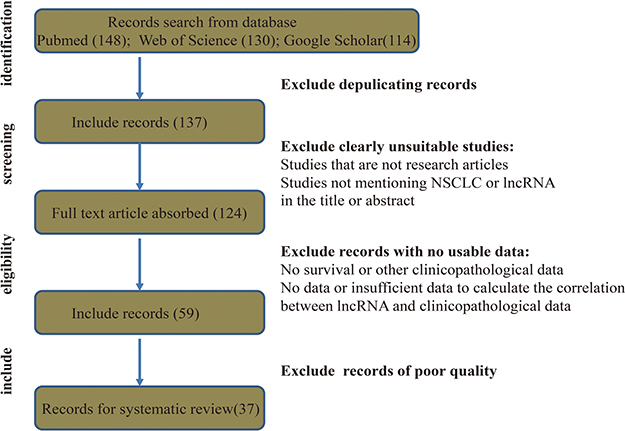
Figure 1: Flow diagram showing the study collection process.
LncRNA expression is associated with NSCLC prognosis
First, we constructed a Table, including author name, publication year, cohort and country, lncRNA type, data type, sample size (exhibiting whole numbers of patients and numbers of groups with high or low expression based on the original classification), Expression file (relative expression of tumor tissues compared to corresponding normal tissues), hazard ratios (HRs), and 95% confidence intervals (CIs) (Supplementary Table 1). After carefully reviewing the medical records with respect to lncRNA and NSCLC, lncRNAs were divided into two groups: oncogenic lncRNAs and tumour suppressor lncRNAs. Next, a separate meta-analysis was performed to estimate the relationship between these two types of lncRNAs and overall survival (OS) of patients diagnosed with NSCLC. We found that 24 lncRNAs (FOXD2-AS1 [13], BCAR4 [14], SNHG1 [15], SBF2-AS1 [16], H19 [17], LINC01133 [18], LINC00342 [19], PVT1 [20–22], ZFAS1 [23], LINC00511 [24], NEAT1 [25], UCA1 [26, 27], AGAP2-AS1 [28], ATB [29], HIT [30], LOC146880 [31], ENST00000439577 [31], XIST [32], ANRIL [33, 34], AFAP1-AS1 [35], MVIH [36], CARLo-5 [37], Sox2ot [38], HOTAIR [39]) were oncogenic (HR, 1.18; 95% Cl, 1.14–1.22; Figure 2A), whereas eight (TUG1 [40, 41], CASC2 [42], PANDAR [43], HMlincRNA717 [44], BANCR [45], SPRY4-IT1 [46], MEG3 [47], GAS6-AS1 [48]) were tumour-suppressor lncRNAs(HR, 0.54; 95% Cl, 0.44–0.66; Figure 2B).
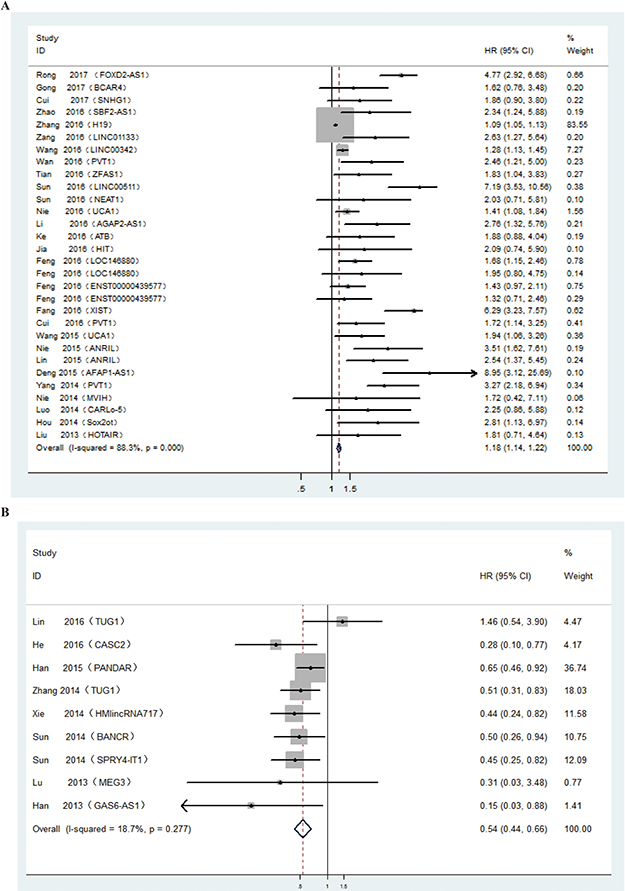
Figure 2: (A) Forest plot showing the hazard ratios and 95% confidence intervals for OS according to the type of oncogenic lncRNAs. (B) Forest plot showing the hazard ratios and 95% confidence intervals for OS according to the type of tumour suppressor lncRNAs.
The type of lncRNA affects the malignant clinicopathologic characteristics of NSCLC
We performed another meta-analysis to further investigate the correlation between lncRNA expression and malignant clinicopathologic characteristics of NSCLC. We first summarized the relationship between each lncRNA and clinicopathologic characteristics such as age, gender, smoking history, histological type, tumor size, and lymph node metastasis (Supplementary Table 2). We then calculated the pooled odds ratios (ORs) and 95% CIs according to the different clinicopathological features. We found that lncRNA expression showed no correlation with age [13–15, 17, 18, 20–30, 32–34, 36, 42, 44–46, 48–53], gender [13–18, 20–30, 32–36, 39, 42, 44–46, 48–53], smoking history [13, 17, 18, 20, 23–26, 28, 29, 32, 33, 35, 36, 42, 45, 46, 48, 49, 51, 53], or histological type [14–18, 20, 24, 25, 28, 29, 32–34, 36, 39, 42, 44–46, 48, 49, 52, 53], but that it showed a close relationship with tumor size [13–18, 20–29, 32–34, 36, 42, 44–46, 48–53] and lymph node metastasis [13–18, 20–29, 32–36, 42, 44–46, 49–53]; i.e., high expression of oncogenic lncRNAs was associated with poorer pathological parameters, while high expression of tumour-suppressor lncRNAs showed the opposite. The pooled ORs and 95% CIs of the relationship between lncRNA expression and clinicopathological characteristics were summarized in Table 1. The corresponding forest plots were shown in Supplementary Figure 1.
Table 1: Pooled odds ratios and 95% confidence intervals for clinical characteristics of lncRNAs in NSCLC
Tumor promoter |
Tumor suppressor |
|||
|---|---|---|---|---|
OR |
95% Cl |
OR |
95% CI |
|
Age |
||||
cut off, 60 |
0.95 |
0.72–1.25 |
0.85 |
0.54–1.34 |
cut off, 65 |
0.93 |
0.69–1.25 |
0.85 |
0.55–1.32 |
Gender |
||||
W/M |
1.13 |
0.94–1.35 |
0.92 |
0.67–1.27 |
Smoking history |
||||
Y/N |
1.16 |
0.79–1.70 |
1.26 |
0.86–1.84 |
Histological type |
||||
Ade/Squ |
0.94 |
0.74–1.18 |
1.27 |
0.91–1.77 |
Tumor size |
||||
cut off, 3 cm |
2.74 |
1.66–4.52 |
0.33 |
0.14–0.75 |
cut off, 5 cm |
2.04 |
1.09–3.79 |
0.28 |
0.18–0.45 |
Lymph node metastasis |
||||
Y/N |
3.3 |
2.42–4.49 |
0.37 |
0.26–0.52 |
W, women; M, men; Y, yes; N, no; Ade, adenocarcinoma; Squ, squamous cell carcinoma.
LncRNA may have utility as a serum biomarker for NSCLC
To investigate the diagnostic utility of lncRNA for NSCLC, we first identified studies (six studies [54–59]) reporting that ten lncRNAs may be useful as serum markers of NSCLC. We summarized the trial name, lncRNA type, data type, area under the curve (AUC), p values, 95% CIs, and sensitivity and specificity in Supplementary Table 3. Next, we estimated the pooled indices for these diagnostic reference values, while Spearman correlation coefficient ruled out the threshold effect (r = 0.104 , p = 0.713). The results were as follows: sensitivity, 0.81 [0.72–0.87]; specificity, 0.83 [0.73–0.90]; positive likelihood ratio, 4.8 [3.0–7.8]; negative likelihood ratio, 0.23 [0.16–0.34]; and overall diagnostic OR, 21 [11–39]. The forest plot was shown in Figure 3A. The summary receiver operator characteristic curve (SROC) AUC value was 0.89 (0.86–0.91), indicating that lncRNAs have a high diagnostic accuracy (Figure 3B).
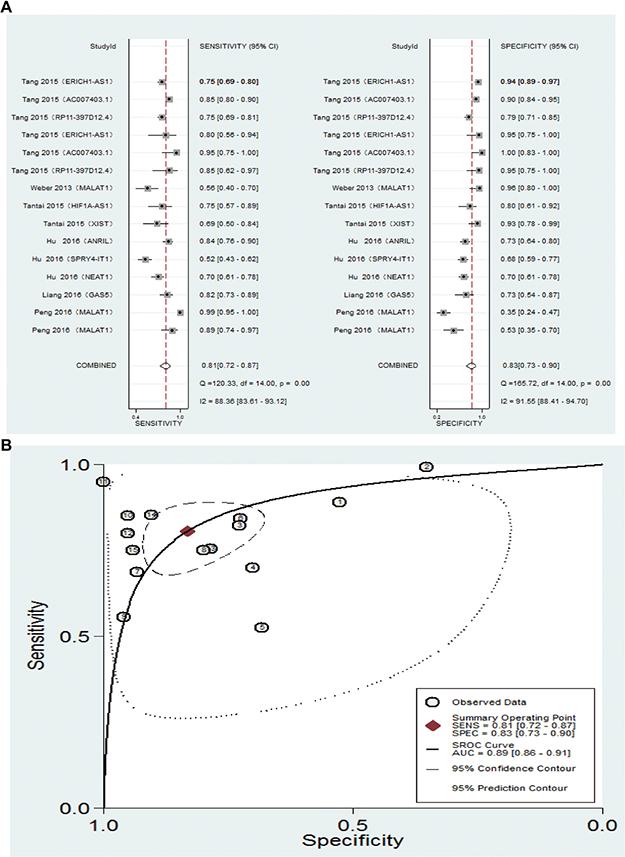

Figure 3: (A) Forest plot showing the sensitivity and specificity of serum lncRNAs for a diagnosis of NSCLC. (B) Summary Receiver operating characteristics curve for lncRNAs.
LncRNAs affect malignant progression of NSCLC by regulating signaling pathways via gene expression modulation
To further examine the link between lncRNAs and NSCLC, we examined their effect on the malignant phenotype. We found that lncRNAs regulated cell proliferation, apoptosis, and metastasis. Some lncRNAs affected all three of these cancer hallmarks, whereas some affected only one (Figure 4A). Furthermore, we found that lncRNAs regulated expression of key proteins within signal networks or biological processes (e.g., P53, Wnt/β-catenin signaling, cell cycle checkpoints, epithelial mesenchymal transition (EMT), and extracellular matrix (ECM) remodeling), all of which regulate the malignant characteristics of cancer cells (Figure 4B). In addition, we examined how lncRNAs regulate gene expression; we found that lncRNAs function at epigenetic, transcriptional, post-transcriptional, and post-translational levels to modulate gene expression (Figure 4C). Taken together, the data suggested that lncRNAs functioned at every step of gene expression to mediate levels of key proteins. This in turn could impact molecular signaling pathways, leading to promotion or inhibition of malignant progression.
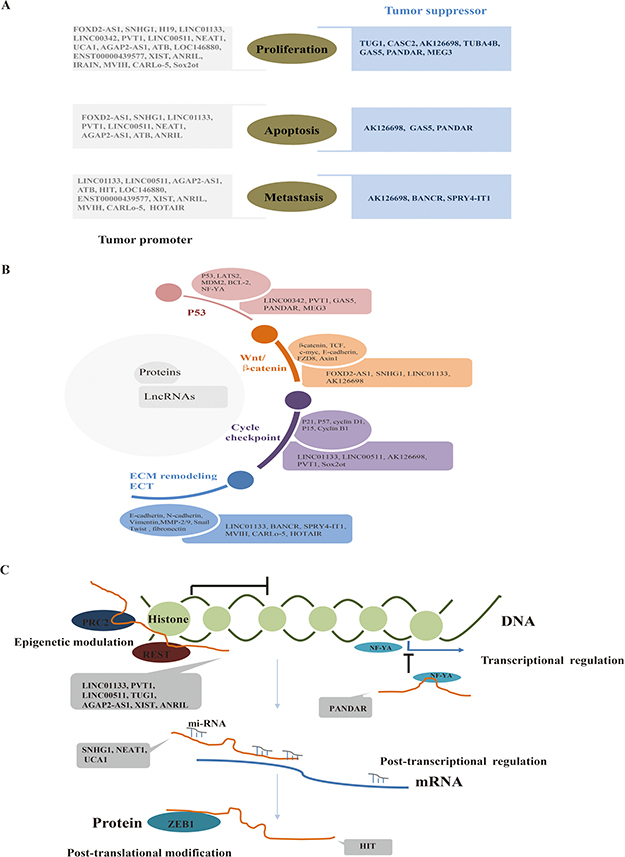
Figure 4: (A) LncRNAs function as tumor promoters or suppressors by regulating malignant characteristics. (B) LncRNAs regulate classical signaling pathways or biological processes critical for NSCLC by regulating critical proteins. (C) LncRNAs regulate almost every step of gene expression.
DISCUSSION
Here, we systematically reviewed 37 eligible records to estimate the clinical utility of lncRNAs as diagnostic and prognostic indicators for NSCLC. We found that lncRNAs may be useful biomarkers for NSCLC. Further, we summarized the molecular mechanisms underlying involvement of lncRNAs in malignancy of NSCLC. Thus, lncRNAs are deserving of much attention (alongside proteins and DNA) in terms of their role in the pathogenesis of NSCLC.
Almost 70% NSCLC were diagnosed in an advanced state, which means limitation in medical intervention and poor prognosis, where efficient biomarker for predicting urgently needed [1, 5]. LncRNA is an emerging star in cancer research for its powerful function in modulating cellular processes and numerous studies have revealed the specific lncRNAs in NSCLC prognosis and diagnosis [7, 10, 11, 60, 61]. In our research, we systematically reviewed the function of lncRNAs in NSCLC. First, we found that lncRNAs had a close relationship with clinicopathological features of NSCLC, which consists of survival state, tumor size and lymph node metastasis. What we found proved the potency of lncRNAs in evaluating the clinical stage of the tumor and telling the prognosis of NSCLC, which may help us handle such malignancy in a novel perspective not the traditional invasive way. Further, we found that lncRNAs had high diagnostic efficiency in NSCLC, which may provide a convenient, economic and microinvasive approach to screen the NSCLC via this liquid biopsy.
Moreover, we deeply investigated the molecular underpinning of lncRNAs associated with NSCLC. As we know, NSCLC is a disease caused and promoted by defects in genes that activate or inactivate signal networks inside and outside the cells [62], where lncRNAs could regulate almost every step of gene expression, whose aberrance could trigger cellular chaos, facilitating malignancy indeed [8, 10]. In this manuscript, we found lncRNAs could regulate a series of critical tumor associated genes like P53, cyclinD1, β-catenin, E-cadherin and etc. involving in signaling networks like P53 pathway, cycle checkpointing group, Wnt/ β-catenin signal transduction cascades and EMT, ECM remodeling systems, which could modulate proliferation, survival and metastasis. Further, we focused on the gene expression modulation function of lncRNAs and we found lncRNAs involved in NSCLC could participate in epigenetic, transcriptional, post-transcriptional and post-translational modification via its interaction with chromatin, DNA, RNA, and protein.
However, limitations still exist,which could cause bias and heterogeneity. For example, insufficient samples, limited publication of negative results and language restrictions all could cause bias [63–65]; Next, due to inherent weakness in observational trial, group characters difference, trial plan difference and evaluation means difference all could cause higher heterogeneity [64, 66]; Last but not least, the complex function of lncRNAs, high heterogeneity of NSCLC, and unknown mechanisms in carcinogenesis all could affect our deducation [4, 61]. Where a more comprehensive and deeper research consisting of sufficient samples, multi-centers, higher quality studies with unified criteria, and etc. should be explored in the future work.
In conclusion, lncRNAs orchestrate gene expression at almost every step, resulting in activation or suppression of oncogenes and tumor suppressor genes, and ultimately affecting malignant progression via regulating signal networks. In NSCLC, such aberrant lncRNAs may have utility as novel diagnostic and prognostic indicators, which may point the way to new preventative and therapeutic options.
MATERIALS AND METHODS
Date sources and search strategy
PubMed, Web of Science, and Google Scholar were probed using the search term “ lncRNA OR long non-coding RNA OR lncRNAs OR long non-coding RNAs and NSCLC OR non-small cell lung cancer ”. The time window started at the time of database inception and extended to 1 April 2017. The language was limited to English.
Inclusion and exclusion criteria
The inclusion criteria were as follows: (1) all patients diagnosed with NSCLC; (2) all studies used well-established methods of measuring and analyzing lncRNA (e.g., qPCR); (3) studies investigated the association between lncRNA and cancer prognosis with OS as a definitive outcome or other clinicopathological parameters like tumor size, lymph node metastasis and etc.. . Exclusion criteria were as follows: (1) reviews, case reports, or meeting presentations; (2) unqualified patients and grouping methods; (3) insufficient data to accurately calculate HRs, ORs and corresponding 95% CIs; (4) where duplicate publications or the same research was published in different formats, only the latest article was included.
Databases were examined by two independent investigators, and studies were selected based on title, abstract, and/or full text. Any disagreements were discussed and resolved by consensus.
Date extraction and quality assessment
The date of each study was extracted, along with the name of the first author, the publication date, the number of samples, lncRNA examination methods and expression patterns, prognostic outcome (HR and 95% CI) and clinicopathological parameters. All extractions were performed by two independent investigators, and any disagreement was resolved by consensus. Furthermore, the quality of each study was assessed using the Newcastle-Ottawa Scale criteria or Quality Assessment for Diagnostic Accuracy Studies based on the type of clinical trial. This was again carried out by two independent investigators, and any disagreement was resolved by consensus. The assessment criteria included cohort selection, comparability, expose, and outcome.
Statistical analysis
Data were analyzed using STATA software, version 12.0 (STATA Corporation, College Station, Texas, USA). HRs and corresponding 95% CIs were used to estimate the correlation between lncRNA expression and clinical prognosis of NSCLC. HRs, ORs, and sensitivity and specificity (and their respective 95% CIs) were collected from the article (if available) or calculated based on available information (calculations were carried out according to previously reported methods [67, 68]). Heterogeneity was examined using the Q test and the I2 test. Heterogeneity was considered statistically significant at a p value < 0.05 or an I2 value > 50% (when a random-effect model was applied). Otherwise, a fixed-effect model was used.
Abbreviations
Non-small cell lung cancer (NSCLC); long non-coding RNAs (lncRNAs); hazard ratios (HRs); confidence intervals (CIs); overall survival (OS); odds ratios (ORs) epithelial mesenchymal transition (EMT); and extracellular matrix (ECM); Polycomb Repressive Complex 2 (PRC2); RE1 Silencing Transcription Factor (REST); Nuclear Transcription Factor Y Subunit Alpha (NF-YA); ZEB1 (Zinc Finger E-Box Binding Homeobox 1).
Author contributions
Mr. Yanlu Xiong put forward this idea; Mr. Yanlu Xiong and Mr. Tao Wang processed data and wrote this manuscript; Mr. Mingxing Wang , Mr. Jinbao Zhao , Mr. Xiaofei Li, Mr. Yongsheng Zhou, Mr. Jiabao Liu and Mr. Zhipei Zhang helped the data collected and analysed; Mr. Lintao Jia and Mr. Yong Han assessed this progress and designed this plan.
ACKNOWLEDGMENTS
We thank the staff at the Department of Thoracic Surgery, Tangdu Hospital, Fourth Military Medical University, Xi’an, China, and the State Key Laboratory of Cancer Biology, Department of Biochemistry and Molecular Biology, Fourth Military Medical University, Xi’an, China, for technical support.
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.
FUNDING
This study was funded by the Special Clinical Research Fund from Wu JiePing Medical Foundation (320.6750.15095), and the Natural Science and Technology Foundation Research Key Project of Shaanxi Province (2015JZ 024), China.
REFERENCES
1. Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011; 61:69–90. https://doi.org/10.3322/caac.20107.
2. Goldstraw P, Ball D, Jett JR, Le Chevalier T, Lim E, Nicholson AG, Shepherd FA. Non-small-cell lung cancer. Lancet. 2011; 378:1727–40. https://doi.org/10.1016/s0140-6736(10)62101-0.
3. Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA. Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc. 2008; 83:584–94. https://doi.org/10.4065/83.5.584.
4. Chen Z, Fillmore CM, Hammerman PS, Kim CF, Wong KK. Non-small-cell lung cancers: a heterogeneous set of diseases. Nat Rev Cancer. 2014; 14:535–46. https://doi.org/10.1038/nrc3775.
5. Thomas A, Liu SV, Subramaniam DS, Giaccone G. Refining the treatment of NSCLC according to histological and molecular subtypes. Nat Rev Clin Oncol. 2015; 12:511–26. https://doi.org/10.1038/nrclinonc.2015.90.
6. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011; 144:646–74. https://doi.org/10.1016/j.cell.2011.02.013.
7. Evans JR, Feng FY, Chinnaiyan AM. The bright side of dark matter: lncRNAs in cancer. J Clin Invest. 2016; 126:2775–82. https://doi.org/10.1172/jci84421.
8. Geisler S, Coller J. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol. 2013; 14:699–712. https://doi.org/10.1038/nrm3679.
9. Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011; 1:391–407. https://doi.org/10.1158/2159-8290.cd-11-0209.
10. Schmitt AM, Chang HY. Long Noncoding RNAs in Cancer Pathways. Cancer Cell. 2016; 29:452–63. https://doi.org/10.1016/j.ccell.2016.03.010.
11. Chen J, Wang R, Zhang K, Chen LB. Long non-coding RNAs in non-small cell lung cancer as biomarkers and therapeutic targets. J Cell Mol Med. 2014; 18:2425–36. https://doi.org/10.1111/jcmm.12431.
12. Wei MM, Zhou GB. Long Non-coding RNAs and Their Roles in Non-small-cell Lung Cancer. Genomics Proteomics Bioinformatics. 2016; 14:280–8. https://doi.org/10.1016/j.gpb.2016.03.007.
13. Rong L, Zhao R, Lu J. Highly expressed long non-coding RNA FOXD2-AS1 promotes non-small cell lung cancer progression via Wnt/beta-catenin signaling. Biochem Biophys Res Commun. 2017; 484:586–91. https://doi.org/10.1016/j.bbrc.2017.01.141.
14. Gong J, Zhang H, He L, Wang L, Wang J. Increased Expression of Long Non-Coding RNA BCAR4 Is Predictive of Poor Prognosis in Patients with Non-Small Cell Lung Cancer. Tohoku J Exp Med. 2017; 241:29–34. https://doi.org/10.1620/tjem.241.29.
15. Cui Y, Zhang F, Zhu C, Geng L, Tian T, Liu H. Upregulated lncRNA SNHG1 contributes to progression of non-small cell lung cancer through inhibition of miR-101-3p and activation of Wnt/beta-catenin signaling pathway. Oncotarget. 2017; 8:17785–17794. https://doi.org/10.18632/oncotarget.14854.
16. Zhao QS, Li L, Zhang L, Meng XW, Li LL, Ge XF, Li ZP. Over-expression of lncRNA SBF2-AS1 is associated with advanced tumor progression and poor prognosis in patients with non-small cell lung cancer. Eur Rev Med Pharmacol Sci. 2016; 20:3031–4.
17. Zhang E, Li W, Yin D, De W, Zhu L, Sun S, Han L. c-Myc-regulated long non-coding RNA H19 indicates a poor prognosis and affects cell proliferation in non-small-cell lung cancer. Tumour Biol. 2016; 37:4007–15. https://doi.org/10.1007/s13277-015-4185-5.
18. Zang C, Nie FQ, Wang Q, Sun M, Li W, He J, Zhang M, Lu KH. Long non-coding RNA LINC01133 represses KLF2, P21 and E-cadherin transcription through binding with EZH2, LSD1 in non small cell lung cancer. Oncotarget. 2016; 7:11696–707. https://doi.org/10.18632/oncotarget.7077.
19. Wang L, Chen Z, An L, Wang Y, Zhang Z, Guo Y, Liu C. Analysis of Long Non-Coding RNA Expression Profiles in Non-Small Cell Lung Cancer. Cell Physiol Biochem. 2016; 38:2389–400. https://doi.org/10.1159/000445591.
20. Wan L, Sun M, Liu GJ, Wei CC, Zhang EB, Kong R, Xu TP, Huang MD, Wang ZX. Long Noncoding RNA PVT1 Promotes Non-Small Cell Lung Cancer Cell Proliferation through Epigenetically Regulating LATS2 Expression. Mol Cancer Ther. 2016; 15:1082–94. https://doi.org/10.1158/1535-7163.mct-15-0707.
21. Cui D, Yu CH, Liu M, Xia QQ, Zhang YF, Jiang WL. Long non-coding RNA PVT1 as a novel biomarker for diagnosis and prognosis of non-small cell lung cancer. Tumour Biol. 2016; 37:4127–34. https://doi.org/10.1007/s13277-015-4261-x.
22. Yang YR, Zang SZ, Zhong CL, Li YX, Zhao SS, Feng XJ. Increased expression of the lncRNA PVT1 promotes tumorigenesis in non-small cell lung cancer. Int J Clin Exp Pathol. 2014; 7:6929–35.
23. Tian FM, Meng FQ, Wang XB. Overexpression of long-noncoding RNA ZFAS1 decreases survival in human NSCLC patients. Eur Rev Med Pharmacol Sci. 2016; 20:5126–31.
24. Sun CC, Li SJ, Li G, Hua RX, Zhou XH, Li DJ. Long Intergenic Noncoding RNA 00511 Acts as an Oncogene in Non-small-cell Lung Cancer by Binding to EZH2 and Suppressing p57. Mol Ther Nucleic Acids. 2016; 5:e385. https://doi.org/10.1038/mtna.2016.94.
25. Sun C, Li S, Zhang F, Xi Y, Wang L, Bi Y, Li D. Long non-coding RNA NEAT1 promotes non-small cell lung cancer progression through regulation of miR-377-3p-E2F3 pathway. Oncotarget. 2016; 7:51784–814. https://doi.org/10.18632/oncotarget.10108.
26. Nie W, Ge HJ, Yang XQ, Sun X, Huang H, Tao X, Chen WS, Li B. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett. 2016; 371:99–106. https://doi.org/10.1016/j.canlet.2015.11.024.
27. Wang HM, Lu JH, Chen WY, Gu AQ. Upregulated lncRNA-UCA1 contributes to progression of lung cancer and is closely related to clinical diagnosis as a predictive biomarker in plasma. Int J Clin Exp Med. 2015; 8:11824–30.
28. Li W, Sun M, Zang C, Ma P, He J, Zhang M, Huang Z, Ding Y, Shu Y. Upregulated long non-coding RNA AGAP2-AS1 represses LATS2 and KLF2 expression through interacting with EZH2 and LSD1 in non-small-cell lung cancer cells. Cell Death Dis. 2016; 7:e2225. https://doi.org/10.1038/cddis.2016.126.
29. Ke L, Xu SB, Wang J, Jiang XL, Xu MQ. High expression of long non-coding RNA ATB indicates a poor prognosis and regulates cell proliferation and metastasis in non-small cell lung cancer. Clin Transl Oncol. 2017; 19:599–605. https://doi.org/10.1007/s12094-016-1572-3.
30. Jia X, Wang Z, Qiu L, Yang Y, Wang Y, Chen Z, Liu Z, Yu L. Upregulation of LncRNA-HIT promotes migration and invasion of non-small cell lung cancer cells by association with ZEB1. Cancer Med. 2016; 5:3555–63. https://doi.org/10.1002/cam4.948.
31. Feng N, Ching T, Wang Y, Liu B, Lin H, Shi O, Zhang X, Zheng M, Zheng X, Gao M, Zheng ZJ, Yu H, Garmire L, et al. Analysis of Microarray Data on Gene Expression and Methylation to Identify Long Non-coding RNAs in Non-small Cell Lung Cancer. Sci Rep. 2016; 6:37233. https://doi.org/10.1038/srep37233.
32. Fang J, Sun CC, Gong C. Long noncoding RNA XIST acts as an oncogene in non-small cell lung cancer by epigenetically repressing KLF2 expression. Biochem Biophys Res Commun. 2016; 478:811–7. https://doi.org/10.1016/j.bbrc.2016.08.030.
33. Nie FQ, Sun M, Yang JS, Xie M, Xu TP, Xia R, Liu YW, Liu XH, Zhang EB, Lu KH, Shu YQ. Long noncoding RNA ANRIL promotes non-small cell lung cancer cell proliferation and inhibits apoptosis by silencing KLF2 and P21 expression. Mol Cancer Ther. 2015; 14:268–77. https://doi.org/10.1158/1535-7163.mct-14-0492.
34. Lin L, Gu ZT, Chen WH, Cao KJ. Increased expression of the long non-coding RNA ANRIL promotes lung cancer cell metastasis and correlates with poor prognosis. Diagn Pathol. 2015; 10:14. https://doi.org/10.1186/s13000-015-0247-7.
35. Deng J, Liang Y, Liu C, He S, Wang S. The up-regulation of long non-coding RNA AFAP1-AS1 is associated with the poor prognosis of NSCLC patients. Biomed Pharmacother. 2015; 75:8–11. https://doi.org/10.1016/j.biopha.2015.07.003.
36. Nie FQ, Zhu Q, Xu TP, Zou YF, Xie M, Sun M, Xia R, Lu KH. Long non-coding RNA MVIH indicates a poor prognosis for non-small cell lung cancer and promotes cell proliferation and invasion. Tumour Biol. 2014; 35:7587–94. https://doi.org/10.1007/s13277-014-2009-7.
37. Luo J, Tang L, Zhang J, Ni J, Zhang HP, Zhang L, Xu JF, Zheng D. Long non-coding RNA CARLo-5 is a negative prognostic factor and exhibits tumor pro-oncogenic activity in non-small cell lung cancer. Tumour Biol. 2014; 35:11541–9. https://doi.org/10.1007/s13277-014-2442-7.
38. Hou Z, Zhao W, Zhou J, Shen L, Zhan P, Xu C, Chang C, Bi H, Zou J, Yao X, Huang R, Yu L, Yan J. A long noncoding RNA Sox2ot regulates lung cancer cell proliferation and is a prognostic indicator of poor survival. Int J Biochem Cell Biol. 2014; 53:380–8. https://doi.org/10.1016/j.biocel.2014.06.004.
39. Liu XH, Liu ZL, Sun M, Liu J, Wang ZX, De W. The long non-coding RNA HOTAIR indicates a poor prognosis and promotes metastasis in non-small cell lung cancer. BMC Cancer. 2013; 13:464. https://doi.org/10.1186/1471-2407-13-464.
40. Lin PC, Huang HD, Chang CC, Chang YS, Yen JC, Lee CC, Chang WH, Liu TC, Chang JG. Long noncoding RNA TUG1 is downregulated in non-small cell lung cancer and can regulate CELF1 on binding to PRC2. BMC Cancer. 2016; 16:583. https://doi.org/10.1186/s12885-016-2569-6.
41. Zhang EB, Yin DD, Sun M, Kong R, Liu XH, You LH, Han L, Xia R, Wang KM, Yang JS, De W, Shu YQ, Wang ZX. P53-regulated long non-coding RNA TUG1 affects cell proliferation in human non-small cell lung cancer, partly through epigenetically regulating HOXB7 expression. Cell Death Dis. 2014; 5:e1243. https://doi.org/10.1038/cddis.2014.201.
42. He X, Liu Z, Su J, Yang J, Yin D, Han L, De W, Guo R. Low expression of long noncoding RNA CASC2 indicates a poor prognosis and regulates cell proliferation in non-small cell lung cancer. Tumour Biol. 2016; 37:9503–10. https://doi.org/10.1007/s13277-016-4787-6.
43. Han L, Zhang EB, Yin DD, Kong R, Xu TP, Chen WM, Xia R, Shu YQ, De W. Low expression of long noncoding RNA PANDAR predicts a poor prognosis of non-small cell lung cancer and affects cell apoptosis by regulating Bcl-2. Cell Death Dis. 2015; 6:e1665. https://doi.org/10.1038/cddis.2015.30.
44. Xie X, Liu HT, Mei J, Ding FB, Xiao HB, Hu FQ, Hu R, Wang MS. LncRNA HMlincRNA717 is down-regulated in non-small cell lung cancer and associated with poor prognosis. Int J Clin Exp Pathol. 2014; 7:8881–6.
45. Sun M, Liu XH, Wang KM, Nie FQ, Kong R, Yang JS, Xia R, Xu TP, Jin FY, Liu ZJ, Chen JF, Zhang EB, De W, et al. Downregulation of BRAF activated non-coding RNA is associated with poor prognosis for non-small cell lung cancer and promotes metastasis by affecting epithelial-mesenchymal transition. Mol Cancer. 2014; 13:68. https://doi.org/10.1186/1476-4598-13-68.
46. Sun M, Liu XH, Lu KH, Nie FQ, Xia R, Kong R, Yang JS, Xu TP, Liu YW, Zou YF, Lu BB, Yin R, Zhang EB, et al. EZH2-mediated epigenetic suppression of long noncoding RNA SPRY4-IT1 promotes NSCLC cell proliferation and metastasis by affecting the epithelial-mesenchymal transition. Cell Death Dis. 2014; 5:e1298. https://doi.org/10.1038/cddis.2014.256.
47. Lu KH, Li W, Liu XH, Sun M, Zhang ML, Wu WQ, Xie WP, Hou YY. Long non-coding RNA MEG3 inhibits NSCLC cells proliferation and induces apoptosis by affecting p53 expression. BMC Cancer. 2013; 13:461. https://doi.org/10.1186/1471-2407-13-461.
48. Han L, Kong R, Yin DD, Zhang EB, Xu TP, De W, Shu YQ. Low expression of long noncoding RNA GAS6-AS1 predicts a poor prognosis in patients with NSCLC. Med Oncol. 2013; 30:694. https://doi.org/10.1007/s12032-013-0694-5.
49. Wu Y, Lyu H, Liu H, Shi X, Song Y, Liu B. Downregulation of the long noncoding RNA GAS5-AS1 contributes to tumor metastasis in non-small cell lung cancer. Sci Rep. 2016; 6:31093. https://doi.org/10.1038/srep31093.
50. Lu Y, Zhou X, Xu L, Rong C, Shen C, Bian W. Long noncoding RNA ANRIL could be transactivated by c-Myc and promote tumor progression of non-small-cell lung cancer. Onco Targets Ther. 2016; 9:3077–84. https://doi.org/10.2147/ott.s102658.
51. Fu X, Li H, Liu C, Hu B, Li T, Wang Y. Long noncoding RNA AK126698 inhibits proliferation and migration of non-small cell lung cancer cells by targeting Frizzled-8 and suppressing Wnt/beta-catenin signaling pathway. Onco Targets Ther. 2016; 9:3815–27. https://doi.org/10.2147/ott.s100633.
52. Shi X, Sun M, Liu H, Yao Y, Kong R, Chen F, Song Y. A critical role for the long non-coding RNA GAS5 in proliferation and apoptosis in non-small-cell lung cancer. Mol Carcinog. 2015; 54 Suppl 1:E1–e12. https://doi.org/10.1002/mc.22120.
53. Feng J, Sun Y, Zhang EB, Lu XY, Jin SD, Guo RH. A novel long noncoding RNA IRAIN regulates cell proliferation in non small cell lung cancer. Int J Clin Exp Pathol. 2015; 8:12268–75.
54. Peng H, Wang J, Li J, Zhao M, Huang SK, Gu YY, Li Y, Sun XJ, Yang L, Luo Q, Huang CZ. A circulating non-coding RNA panel as an early detection predictor of non-small cell lung cancer. Life Sci. 2016; 151:235–42. https://doi.org/10.1016/j.lfs.2016.03.002.
55. Liang W, Lv T, Shi X, Liu H, Zhu Q, Zeng J, Yang W, Yin J, Song Y. Circulating long noncoding RNA GAS5 is a novel biomarker for the diagnosis of nonsmall cell lung cancer. Medicine (Baltimore). 2016; 95:e4608. https://doi.org/10.1097/md.0000000000004608.
56. Hu X, Bao J, Wang Z, Zhang Z, Gu P, Tao F, Cui D, Jiang W. The plasma lncRNA acting as fingerprint in non-small-cell lung cancer. Tumour Biol. 2016; 37:3497–504. https://doi.org/10.1007/s13277-015-4023-9.
57. Tantai J, Hu D, Yang Y, Geng J. Combined identification of long non-coding RNA XIST and HIF1A-AS1 in serum as an effective screening for non-small cell lung cancer. Int J Clin Exp Pathol. 2015; 8:7887–95.
58. Tang Q, Ni Z, Cheng Z, Xu J, Yu H, Yin P. Three circulating long non-coding RNAs act as biomarkers for predicting NSCLC. Cell Physiol Biochem. 2015; 37:1002–9. https://doi.org/10.1159/000430226.
59. Weber DG, Johnen G, Casjens S, Bryk O, Pesch B, Jockel KH, Kollmeier J, Bruning T. Evaluation of long noncoding RNA MALAT1 as a candidate blood-based biomarker for the diagnosis of non-small cell lung cancer. BMC Res Notes. 2013; 6:518. https://doi.org/10.1186/1756-0500-6-518.
60. Yang H, Han Y, Wu L, Wu C. Long Non-Coding RNA Expression Signature Hallmarks Promising Efficacy in Identification of Human Non-Small Cell Lung Cancer: a Meta-Analysis Study. Clin Lab. 2017; 63:1447–56. https://doi.org/10.7754/Clin.Lab.2017.170325.
61. Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet. 2016; 17:47–62. https://doi.org/10.1038/nrg.2015.10.
62. Park SJ, More S, Murtuza A, Woodward BD, Husain H. New Targets in Non-Small Cell Lung Cancer. Hematol Oncol Clin North Am. 2017; 31:113–29. https://doi.org/10.1016/j.hoc.2016.08.010.
63. DerSimonian R, Laird N. Meta-analysis in clinical trials revisited. Contemp Clin Trials. 2015; 45:139–45. https://doi.org/10.1016/j.cct.2015.09.002.
64. Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002; 21:1539–58. https://doi.org/10.1002/sim.1186.
65. Delgado-Rodriguez M, Sillero-Arenas M. [Biases in meta-analysis] [Article in Spanish]. Med Clin (Barc). 1999; 112 Suppl 1:43–50.
66. Black N. Why we need observational studies to evaluate the effectiveness of health care. BMJ. 1996; 312:1215–8.
67. Guyot P, Ades AE, Ouwens MJ, Welton NJ. Enhanced secondary analysis of survival data: reconstructing the data from published Kaplan-Meier survival curves. BMC Med Res Methodol. 2012; 12:9. https://doi.org/10.1186/1471-2288-12-9.
68. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials. 2007; 8:16. https://doi.org/10.1186/1745-6215-8-16.

