INTRODUCTION
Lung cancer is the most common cancer and the leading cause of cancer-related mortality around the world. In 2015, a total of 733,300 patients (509,300 men and 224,000 women) were diagnosed with lung cancer in China [1]. Non-small-cell lung cancer (NSCLC) and small cell lung cancer (SCLC) are two main types of lung cancers; of these, NSCLC accounts for 80–85% of all lung cancers [2]. Unfortunately, about 53% patients of NSCLC are diagnosed at an advanced stage (III b-IV) and, therefore, have a poor prognosis [3].
Programmed cell death 1 ligand 1 (PD-L1; also referred to as B7-H1), which belongs to B7 family, is an inhibitory cell surface molecule. PD-L1 expressed on NSCLC cells is shown to inhibit T cell proliferation and activation by combining with programmed cell death 1 (PD-1) receptor. Blockade of PD-1/PD-L1 represents a novel target for immunotherapy [4]. In a recent clinical trial, Keynote-024, [5]. progression free survival (PFS) in the pembrolizumab group was 4.3 months longer than that in the chemotherapy group (10.3 vs. 6.0 months; hazard ratio [HR]: 0.50, 95% confidence interval [CI], 0.37–0.68). Based on the promising results, anti-PD-1 /PD-L1 agents, nivolumab, pembrolizumab and atezolizumab were approved by FDA for treatment of advanced NSCLCs in the past few years [6–9].
Driver gene alternations including epidermal growth factor receptor (EGFR), anaplastic lymphoma kinase (ALK), and Kirsten rat sarcoma viral oncogene homolog (KRAS) have been verified in a large proportion of NSCLCs. For example, 45% of never smokers and 7% of patients with tobacco-associated lung cancer harbor EGFR mutations. At least 30% of East Asian patients with NSCLC have EGFR mutation as compared to <10% of Caucasian patients [10–12]. ALK rearrangements containing exon 20 coding for the tyrosine domain occur in 3.0% to 11.5% of patients with NSCLC [13]. KRAS mutations have been reported in 15–30% of patients with NSCLC, and approximately 97% of mutations are point mutations located in codons 12 or 13 of exon [14].
Gainor et al. reported an objective response rate (ORR) of 3.6% in EGFR mutation or ALK rearrangement cohort treated with pembrolizumab, while the ORR in EGFR/ALK wild type cohort was 23.3%. Similarly, in Checkmate 057 and Keynote-010, neither nivolumab nor pembrolizumab showed an obvious superiority over docetaxel with respect to overall survival (OS) in the EGFR mutation cohorts [15]. In contrast to EGFR/ALK, KRAS mutation was associated with better outcomes (versus docetaxel) with respect to OS in Checkmate 057 (HR: 0.52, 95% CI, 0.29–0.95). PD-L1 expression in tumor issues is considered as a predictor of immunotherapy outcomes, and its’ relationship with driver gene status has attracted increasing attention in recent years.
Results of preclinical studies indicate a positive correlation between EGFR, ALK and KRAS mutations and higher PD-L1 levels, although the underlying mechanisms are not clear. EGFR mutation may upregulate PD-L1 expression through PI3K, MAPK, STAT3 and NF-κB signaling pathway [16, 17]. Similarly, PI3K, MAPK, STAT3 and HIF-1α signaling pathways may be involved in upregulation of PD-L1 expression in ALK mutated cell lines [18, 19]. Treatment with tyrosine kinase inhibitors (TKIs), gefitinib and crizotinib was shown to decrease PD-L1 expression levels in vitro [17]. Co-mutation of STK11/LKB1 and KRAS was shown to downregulate PD-L1 expression level in NSCLC cell lines; in contrast, co-mutation of tp53 and KRAS was shown an upregulating effect [20, 21]. However, the association between PD-L1 expression and driver gene status in clinical tumor specmens is not well characterized [22]. This meta-analysis was aimed at further evaluation of the association between PD-L1 expression and driver gene mutation in order to explain variable outcomes of immunotherapy in patients with NSCLCs harboring various driver gene status.
RESULTS
Search result and selection of studies
The MOOSE guidelines were followed for this meta-analysis of observational studies [23]. Flow diagram showing study selection procedure is presented (Figure 1). Twenty-five publications and one meeting abstract (2015–2017) with a combined study population of 7541 patients were finally included. Due to the higher prevalence of driver gene (especially EGFR and ALK) in East-Asian population, 19 studies were conducted on East-Asian population, while 8 studies were conducted on Caucasian or American population. Quality assessment using STROBE checklist showed a 57.5% median total score (Supplementary Table 1); one study was classified as “high-quality”; one study was classified as “low-quality”; and the remaining studies were classified as “moderate-quality” (Table 1). The percentage of tumor cells staining positive for PD-L1 was calculated for each specimen. The optimal cutoff level is still controversial; 5% tumor cells staining positive is the most common cutoff level used in clinical trials. Baseline population characteristics including median age, gender, tumor stage, histological types and smoking status are summarized in Table 1. For clinical specimens, 1598 (32.67%), 99 (3.25%) and 628 (19.83%) were reported harboring EGFR, ALK and KRAS mutations, respectively (Supplementary Table 2).
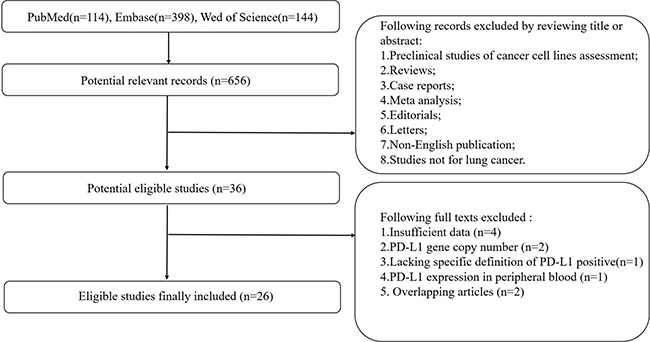
Figure 1: Flow diagram showing selection of studies.
Table 1: Basic characteristics of studies included in the meta-analysis
First author |
Year of publication |
No of patients |
Ethnicity |
Median age(ys) |
Gender |
Smoking status |
Stage |
Histology |
Antibody |
Cutoff |
Quality |
|---|---|---|---|---|---|---|---|---|---|---|---|
Zhang [33] |
2014 |
143 |
East-Asian |
<60 |
Male (41%) Female (59%) |
Smoker (34%) Non-Smoker (66%) |
I (46%) II–III (54%) |
ADCa (100%) |
PoAbe |
40% |
M1 |
Yang [34] |
2014 |
163 |
East-Asian |
≥60 |
Male (33%) Female (67%) |
Smoker (19%) Non-Smoker (81%) |
I (100%) |
ADC (100%) |
PoAb |
5% |
M |
Incecoo [35] |
2014 |
123 |
Other |
≥60 |
Male (54%) Female (46%) |
Smoker (60%) Non-Smoker (30%) |
NRg |
SCCb (66%) ADC (18%) Other (15%) |
PoAb |
5% |
M |
Cooper [36] |
2015 |
678 |
Other |
≥60 |
Male (70%) Female (30%) |
NR |
I (50%) II–III (50%) |
SCC (41%) ADC (40%) Other (19%) |
McAbf |
50% |
M |
Schmidt [37] |
2015 |
321 |
Other |
Other |
Male (78%) Female (22%) |
Smoker (81%) Non-Smoker (20%) |
I (58%), II (26%) III (16%) |
SCC (46%) ADC (39%) Other (15%) |
McAb |
5% |
M |
Kim [38] |
2015 |
331 |
East-Asian |
≥60 |
Male (96%) Female (4%) |
Smoker (95%) Non-Smoker (5%) |
I (40%) II (36%) III (24%) |
SCC (100%) |
McAb |
10% |
M |
Chang [39] |
2015 |
66 |
East-Asian |
<60 |
Male (38%) Female (62%) |
Smoker (12%) Non-Smoker (88%) |
I (32%) II (24%) III–IV (21%) |
LELCc (100%) |
PoAb |
5% |
M |
Koh [40] |
2015 |
497 |
East-Asian |
≥60 |
Male (46%) Female (54%) |
Smoker (40%) Non-Smoker (60%) |
NR |
ADC (100%) |
McAb |
5% |
M |
Tang [41] |
2015 |
170 |
East-Asian |
<60 |
Male (55%) Female (45%) |
Smoker (34%) Non-Smoker (66%) |
III B (5%) IV (95%) |
ADC (85%) Other (15%) |
McAb |
5% |
M |
Omori [42] |
2015 |
95 |
East-Asian |
≥60 |
Male (63%) Female (37%) |
Smoker (74%) Non-Smoker (26%) |
I (32%) II (24%) III (21%) |
SCC (16%) ADC (78%) Other (6%) |
McAb |
1% |
M |
Yang2 [43] |
2016 |
105 |
East-Asian |
≥60 |
Male (85%) Female (15%) |
Smoker (75%) Non-Smoker (15%) |
I (100%) |
SCC (100%) |
PoAb |
5% |
M |
Andreas [44] |
2016 |
436 |
Other |
≥60 |
Male (54%) Female (46%) |
Smoker (89%) Non-Smoker (10%) |
I (47%) II (13%) III (33%) IV (7%) |
ADC (100%) |
McAb |
1% |
M |
Ameratunga [45] |
2016 |
527 |
Other |
≥60 |
Male (69%) Female (31%) |
Smoker (90%) Non-Smoker (7%) |
NR |
SCC (34%) ADC (55%) Other (11%) |
McAb |
5% 50% |
M |
Song [46] |
2016 |
385 |
East-Asian |
<60 |
Male (51%) Female (49%) |
Smoker (39%) Non-Smoker (61%) |
I (31%) II (21%) III (48%) |
ADC (100%) |
McAb |
5% |
M |
Inamura [47] |
2016 |
268 |
East-Asian |
≥60 |
Male (53%) Female (47%) |
Smoker (58%) Non-Smoker (42%) |
I (56%) II–III (44%) |
ADC (100%) |
McAb |
1%, 5% |
M |
Jia [48] |
2016 |
55 |
East-Asian |
≥60 |
Male (53%) Female (47%) |
Smoker (31%) Non-Smoker (69%) |
NR |
SDPLCd (100%) |
McAb |
10% |
M |
Inoue [49] |
2016 |
654 |
East-Asian |
≥60 |
Male (68%) Female (32%) |
Smoker 68%) Non-Smoker (30%) |
I (64%) II (17%) III (19%) |
SCC (27%) ADC (66%) Other (11%) |
McAb |
5% |
M |
Ji [50] |
2016 |
100 |
East-Asian |
≥60 |
Male (51%) Female (49%) |
Smoker (26%) Non-Smoker (74%) |
I 42%) II (27%) III (31%) |
ADC (100%) |
PoAb |
5% |
M |
Huynh [51] |
2016 |
261 |
Other |
≥60 |
Male (35%) Female (65%) |
Smoker (78%) Non-Smoker (22%) |
I (77%) II (13%) III (8%) IV (2%) |
ADC (100%) |
McAb |
5% |
M |
Mori [52] |
2016 |
296 |
East-Asian |
≥60 |
Male (50%) Female (50%) |
Smoker (52%) Non-Smoker (48%) |
NR |
ADC (100%) |
McAb |
Score 50 |
M |
Takada [53] |
2016 |
417 |
East-Asian |
≥60 |
Male (49%) Female (51%) |
Smoker (48%) Non-Smoker (52%) |
I (73%) II–III (27%) |
ADC (100%) |
McAb |
1%5% |
M |
Dong [54] |
2017 |
13 |
East-Asian |
≥60 |
Male (92%) Female (8%) |
Smoker (52%) Non-Smoker (48%) |
III (23%) IV (77%) |
SCC (23%) ADC (69%) Other (8%) |
McAb |
5%, 50% |
M |
Rangachari [55] |
2017 |
71 |
Other |
Other |
Male (45%) Female (55%) |
Smoker (68%) Non-Smoker (32%) |
I–III (25%) IV (75%) |
ADC (100%) |
McAb |
50% |
L2 |
Tsao [56] |
2017 |
982 |
Other |
Other |
Male (73%) Female (27%) |
NR |
I (46%) II (37%) III (8%) IV (2%) |
SCC (45%) ADC (41%) Other (14%) |
McAb |
1%, 25% 50% |
H3 |
Chen [57] |
2017 |
65 |
East-Asian |
≥60 |
Male (57%) Female (43%) |
Smoker (52%) Non-Smoker (48%) |
I + II (51%) III (49%) |
ADC (100%) |
NR |
5% |
M |
Cho [58] |
2017 |
319 |
East-Asian |
≥60 |
Male (39%) Female (61%) |
Smoker (36%) Non-Smoker (64%) |
I (63%) II (13%) III (19%) IV (5%) |
ADC (97%) Other (3%) |
McAb |
1%, 50% |
M |
Note: a: “ADC” represents adenocarcinoma; b: “SCC” represents squamous-cell carcinoma; c: “LELC” represents lymphoepithelioma-like carcinoma; d: “SDPLC” represents synchronous double primary lung cancer, and 2 cases of SCLC were excluded; e: “PoAb” represents “polyclonal antibody”; f: “McAb” represents “monoclonal antibody”; g: “NR” represents “not reported”; 1: “M” represents moderate quality; 2: “L” represents low quality; 3: “H” represents high quality.
EGFR mutation versus EGFR wild type
Twenty-four studies with 4891 clinical specimens were included. A high level of heterogeneity was observed among the included studies (Q = 93.67, I2 = 75.4%, p = 0.000). Using a random effect model, we found a statistically significant negative correlation between PD-L1 expression and EGFR mutation in NSCLCs (OR = 0.64, 95% CI = 0.45–0.91, p = 0.014) (Figure 2A). On sensitivity analysis, respective pooled ORs remained significantly stable after sequential elimination of individual studies (Figure 3A). In order to detect the origin of heterogeneity and to assess the influence of different population characteristics, we set up a series of subgroups according to histological type, cutoff level for PD-L1 positivity, population characteristics (median age, ethnicity, stage, gender, smoking status), types of primary antibodies and study quality. Meta regression was performed to examine the influence of each parameter on heterogeneity. However, neither subgroup analysis nor meta regression could identify any parameter which was responsible for high heterogeneity (Table 2). We also explored potential differences of PD-L1 expression among diverse EGFR mutation status [exon 19 deletion (Del19) and codon 858 mutation in exon 21 (L858R)]; however, no evidence was found to validate our hypothesis (Figure 4A). Furthermore, pooled data showed a significant negative correlation between PD-L1 and EGFR mutation status when the cutoff level for PD-L1 positivity was 1% and 50%. Similar results were found in East-Asian, elder/female/smoker dominant groups (Table 2). In terms of potential publication bias, statistical results of Begg’s test and Egger’s test (Begg’s p = 0.126 and Egger’s p = 0.033) were inconsistent (Figure 5A and 5B). However, no trimming was performed during the trim and fill procedure, which indicated that dissymmetry of funnel plots may have resulted from other parameters rather than publication bias.
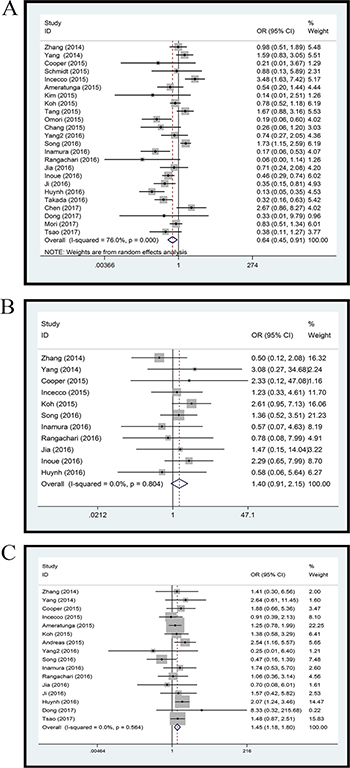
Figure 2: Forest plot. (A) EGFR mutation versus EGFR wild type; (B) ALK mutation versus ALK wild type; (C) KRAS mutation versus KRAS wild type.
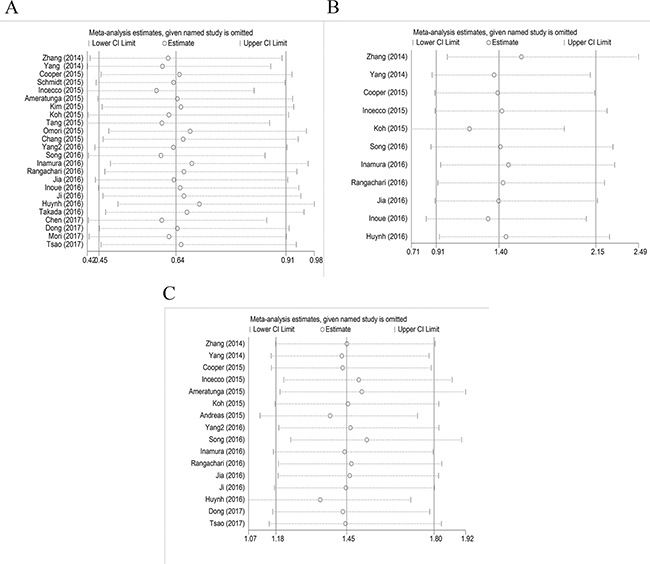
Figure 3: Sensitivity analysis. (A) EGFR mutation versus EGFR wild type; (B) ALK mutation versus ALK wild type; (C) KRAS mutation versus KRAS wild type.
Table 2: Association between PD-L1 expression and driver gene status among different clinicopathological conditions
Driver gene |
Subgroup |
Studies |
OR |
95% CI |
p |
Heterogeneity test |
|||
|---|---|---|---|---|---|---|---|---|---|
Q |
I2 |
p1 |
p2 |
||||||
EGFR |
|||||||||
Median age(years) |
|||||||||
≥60 |
16 |
0.56 |
0.35–0.89 |
0.013 |
64.05 |
76.6% |
0.000 |
0.109 |
|
<60 |
4 |
1.18 |
0.70–2.00 |
0.526 |
6.84 |
56.1% |
0.077 |
||
Gender |
|||||||||
Male > Female |
16 |
0.68 |
0.41–1.12 |
0.129 |
62.45 |
76.0% |
0.000 |
0.720 |
|
Male ≤ Female |
8 |
0.55 |
0.33–0.94 |
0.030 |
29.33 |
76.1% |
0.000 |
||
Smoking status |
|||||||||
Smoker > non-Smoker |
13 |
0.52 |
0.29–0.96 |
0.035 |
52.76 |
77.3% |
0.000 |
0.129 |
|
Smoker ≤ non-Smoker |
9 |
0.82 |
0.53–1.27 |
0.367 |
31.64 |
74.7% |
0.000 |
||
Cutoff |
|||||||||
1% cutoff |
3 |
0.35 |
0.22–0.55 |
0.000 |
1.25 |
0.0% |
0.536 |
0.440 |
|
5% cutoff |
16 |
0.68 |
0.43–1.08 |
0.104 |
83.17 |
82.0% |
0.000 |
||
10% cutoff |
2 |
0.54 |
0.15–1.88 |
0.332 |
1.16 |
13.6% |
0.282 |
||
50% cutoff |
5 |
0.33 |
0.14–0.81 |
0.015 |
2.31 |
0.0% |
0.679 |
||
Race |
|||||||||
East-Asian |
17 |
0.68 |
0.48–0.98 |
0.037 |
59.26 |
73.0% |
0.000 |
0.711 |
|
Other |
7 |
0.45 |
0.14–1.48 |
0.188 |
53.85 |
75.4% |
0.000 |
||
Primary antibody |
|||||||||
PoAba |
6 |
0.93 |
0.46–1.90 |
0.852 |
21.55 |
76.8% |
0.001 |
0.150 |
|
MoAbb |
17 |
0.50 |
0.33–0.75 |
0.001 |
61.91 |
74.2% |
0.000 |
||
Historical type |
|||||||||
ADCc |
12 |
0.69 |
0.43–1.10 |
0.119 |
60.32 |
81.8% |
0.000 |
0.833 |
|
SCCd |
2 |
0.55 |
0.15–2.02 |
0.365 |
1.23 |
18.4% |
0.268 |
||
Stage |
|||||||||
I–II |
12 |
0.73 |
0.45–1.19 |
0.207 |
42.67 |
74.2% |
0.000 |
0.525 |
|
III–IV |
2 |
1.38 |
0.71–2.69 |
0.344 |
0.95 |
0.0% |
0.400 |
||
Quality |
|||||||||
Me |
22 |
0.67 |
0.47–0.97 |
0.033 |
91.05 |
76.9% |
0.000 |
0.187 |
|
ALK |
|||||||||
Median age(years) |
|||||||||
≥60 |
8 |
1.74 |
0.99–3.07 |
0.056 |
3.32 |
0.0% |
0.854 |
/ |
|
<60 |
2 |
0.95 |
0.37–2.43 |
0.968 |
1.30 |
23.2% |
0.254 |
||
Gender |
|||||||||
Male > Female |
6 |
1.39 |
0.77–2.49 |
0.272 |
1.45 |
0.0% |
0.919 |
/ |
|
Male ≤ Female |
5 |
1.41 |
0.74–2.68 |
0.301 |
4.69 |
14.7% |
0.321 |
||
Smoking status |
|||||||||
Smoker > non-Smoker |
5 |
1.23 |
0.58–2.57 |
0.591 |
2.04 |
0.0% |
0.728 |
/ |
|
Smoker ≤ non-Smoker |
5 |
1.51 |
0.84–2.70 |
0.168 |
3.82 |
0.0% |
0.432 |
||
Cutoff |
|||||||||
5% cutoff |
7 |
1.62 |
0.99–2.63 |
0.054 |
3.45 |
0.0% |
0.750 |
/ |
|
50% cutoff |
2 |
1.08 |
0.17–6.96 |
0.569 |
0.32 |
0.0% |
0.569 |
||
Race |
|||||||||
East-Asian |
7 |
1.52 |
0.93–2.47 |
0.095 |
5.06 |
0.0% |
0.536 |
/ |
|
Other |
4 |
1.02 |
0.39–2.66 |
0.962 |
0.66 |
0.0% |
0.883 |
||
Historical type |
|||||||||
ADC |
7 |
1.30 |
0.78–2.17 |
0.138 |
3.33 |
0.0% |
0.650 |
/ |
|
KRAS |
|||||||||
Median age (years) |
|||||||||
≥60 |
12 |
1.58 |
1.23–2.04 |
0.000 |
8.51 |
0.0% |
0.667 |
/ |
|
<60 |
2 |
0.71 |
0.25–2.00 |
0.517 |
1.32 |
24.0% |
0.251 |
||
Gender |
|||||||||
Male > Female |
12 |
1.34 |
1.04–1.72 |
0.025 |
10.38 |
0.0% |
0.497 |
/ |
|
Male ≤ Female |
4 |
1.77 |
1.19–2.62 |
0.004 |
1.81 |
0.0% |
0.613 |
||
Smoking status |
|||||||||
Smoker > non-Smoker |
8 |
1.35 |
1.03–1.77 |
0.032 |
7.8 |
10.3% |
0.350 |
/ |
|
Smoker ≤ non-Smoker |
6 |
1.16 |
0.70–1.19 |
0.576 |
4.52 |
0.0% |
0.477 |
||
Cutoff |
|||||||||
1% cutoff |
2 |
1.84 |
1.23–2.74 |
0.003 |
0.88 |
0.0% |
0.347 |
/ |
|
5% cutoff |
10 |
1.35 |
1.00–1.86 |
0.049 |
10.21 |
11.9% |
0.333 |
||
50% cutoff |
5 |
1.32 |
0.93–1.89 |
0.13 |
0.83 |
0.0% |
0.935 |
||
Race |
|||||||||
East-Asian |
9 |
1.24 |
0.78–1.96 |
0.358 |
7.15 |
0.0% |
0.520 |
/ |
|
Other |
7 |
1.54 |
1.20–1.96 |
0.001 |
5.69 |
0.0% |
0.459 |
||
Historical type |
|||||||||
ADC |
8 |
1.69 |
1.20–2.39 |
0.003 |
7.65 |
8.5% |
0.364 |
/ |
|
SCC |
2 |
0.51 |
0.09–3.07 |
0.464 |
0.26 |
0.0% |
0.609 |
||
Note: a: “PoAb” represents “polyclonal antibody”; b: “McAb” represents “monoclonal antibody”; c: “ADC” represents adenocarcinoma; d: “SCC” represents squamous-cell carcinoma; e: “M” represents moderate quality; 1: p for heterogeneity within each subgroup; 2: p for heterogeneity between subgroups with meta-regression analysis.
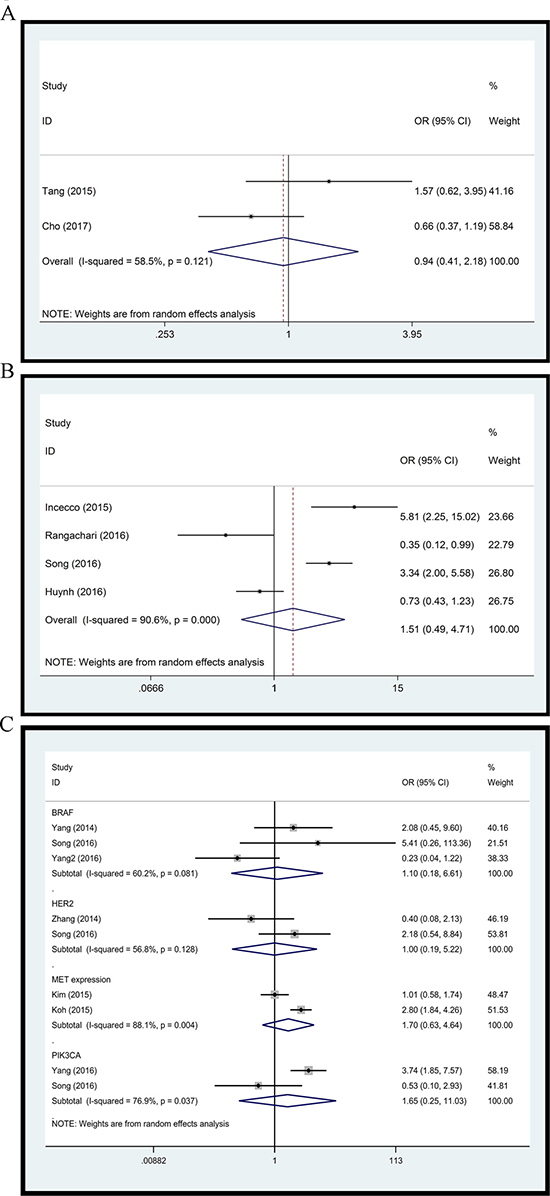
Figure 4: Forest plot. (A) L858R versus Del19; (B) EGFR/ALK/KRAS mutation versus triple wild type; (C) BRAF, HER2, PIK3CA status and MET expression.
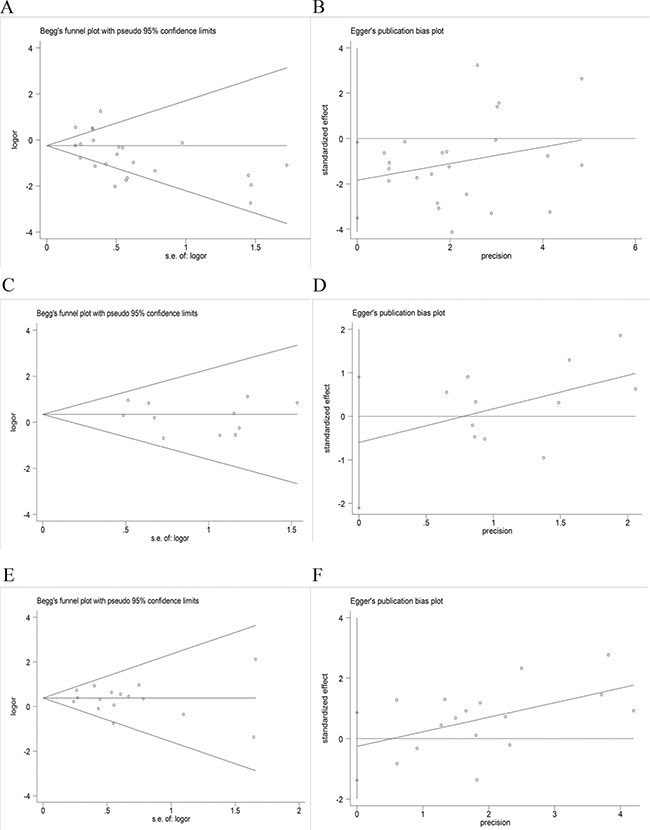
Figure 5: Assessment of publication bias. (A and B) Begg’s test and Egger’s test of EGFR mutation versus EGFR wild type; (C and D) Begg’s test and Egger’s test of ALK mutation versus ALK wild type; (E and F) Begg’s test and Egger’s test of KRAS mutation versus KRAS wild type.
ALK mutation versus ALK wild type
Eleven studies with 3050 clinical specimens were included. No obvious heterogeneity was observed among these studies (Q = 6.13, I2 = 0.0%, p = 0.804). Using a fixed effect model, no statistically significant difference of PD-L1 expression was found among NSCLCs with different ALK status (OR: 1.40, 95% CI, 0.91–2.15, p = 0.131) (Figure 2B). On sensitivity analysis, the studies by Koh et al. and Zhang et al. showed a significant influence on the pooled ORs (Figure 3B). On subgroup analysis, no statistically significant correlation between PD-L1 expression and ALK status was observed (Table 2). The results of Begg’s and Egger’s test (Begg’s p = 0.876 and Egger’s p = 0.389) (Figure 5C and 5D) indicated a lack of potential publication bias in this respect.
KRAS mutation versus KRAS wild type
Sixteen studies with 3167 clinical specimens were included. No obvious heterogeneity was found among these studies (Q = 13.50, I2 = 0.0%, p = 0.566). Pooled analysis using a fixed effects model revealed a statistically significant positive correlation between PD-L1 expression and KRAS mutation in NSCLCs (OR: 1.45, 95% CI, 1.18–1.80, p = 0.001) (Figure 2C). Respective pooled ORs were stable after sequential elimination of one study at a time (Figure 3C). Statistically significant positive correlation was also found with 1% cutoff, 5% cutoff, non-East-Asian ethnicity, lung adenocarcinoma and elder/smoker dominant groups (Table 2). The results of Begg’s test and Egger’s test showed no existence of potential publication bias (Begg’s p = 0.822 and Egger’s p = 0.628) (Figure 5E and 5F).
EGFR/ALK/KRAS mutation versus triple wild type
To further explore the difference in PD-L1 expression between NSCLCs with and without EGFR/ALK/KRAS mutation, we performed a comparison of specimens with at least 1 EGFR/ALK/KRAS mutation and those with triple wild type. Four studies with 840 clinical specimens were included. As compared to triple wild type NSCLCs, those with EGFR/ALK/KRAS mutation did not show any significant differences of PD-L1 expression level with random effect model (OR: 1.51, 95% CI, 0.49–4.71, p = 0.475) (Figure 4B). A high level of heterogeneity was found between these studies (Q = 31.85, I2 = 90.6%, p = 0.000).
Other
Several studies reported the relationship between PD-L1 expression and status of other driver genes, including BRAF, HER2, PIK3CA status and MET expression. We assessed these associations as well. Pooled ORs did not show any significant correlation of BRAF (OR: 1.10, 95% CI, 0.18–6.61, p = 0.913), HER2 (OR: 1.00, 95% CI, 0.19–5.22, p = 0.996), MET expression (OR: 1.70, 95% CI, 0.63–4.64, p = 0.298), and PIK3CA (OR: 1.65, 95% CI, 0.25–11.03, p = 0.604) status with PD-L1 expression (Figure 4C).
DISCUSSION
Clinical efficacy and safety of immune checkpoint inhibitors targeting PD-1/PD-L1 has helped improve prognosis of patients with NSCLCs. There is an urgent need to maximize the clinical benefit of these emerging therapies by optimizing management strategies. The best predictor of the response to PD-1/PD-L1 inhibitors is yet to be determined. A popular hypothesis was that poor treatment outcomes of anti PD-1/PD-L1 therapies may be due to low rates of concurrent CD8+ tumor infiltrating lymphocytes (TILs) and lower PD-L1 expression in the tumor microenvironment (TME) [15].
At cellular level, PD-L1 expression appeared to be associated with strong activation of driver gene (EGFR/ALK) pathways. Up-regulation of PD-L1 level was observed in cell lines with EGFR mutation or ALK rearrangement; further, treatment of EGFR/ALK-TKIs also induced down-regulation of PD-L1 expression [24]. However, the results of preclinical studies are not consistent and have shown lower response rates to anti PD-1/PD-L1 therapies in EGFR/ALK-initiated NSCLCs. The present meta-analysis revealed lower PD-L1 expression in NSCLCs with EGFR mutation. Heterogeneity, which may result from different baseline clinical characteristics of the study population or different investigators for IHC assessment, may influence the accuracy of the conclusions. Besides, we further compared PD-L1 expression levels between exon 19 deletion(del19) and codon 858 mutation in exon 21(L858R), which were the two most common forms of EGFR mutation. Although no significant difference in PD-L1 expression level was found between different EGFR mutated forms owing to limited data, the possibility that heterogeneity resulted from different EGFR mutated forms could not be excluded; additional studies are recommended. In addition, other biological factors like infection may also contribute to heterogeneity [25]. It is reasonable to speculate that lower expression level of PD-L1 in NSCLCs with EGFR mutation may lead to a lower response rate to PD-1/PD-L1 blockade. However, PD-L1 expression level did not differ by EGFR status in the subgroup of patients with 5% cutoff level for PD-L1 positivity, although 5% cutoff has been frequently used in clinical trials. Moreover, the association was found in East-Asian patients but not in patients of other ethnic origin. This result indicates that EGFR mutation may also have potentially influenced the tumor microenvironment. However, no significant correlation between ALK status and PD-L1 positivity was found in this meta-analysis. Studies by Koh et al. and Zhang et al. obviously influenced the pooled OR. However, pooled ORs were not statistically significant even after elimination of these 2 studies from the analysis.
Prior to introduction of PD1/PD-L1 blockade agents, therapeutic options for NSCLCs with KRAS mutation were extremely limited because the molecular diversity of KRAS-mutant tumors is much greater than that of tumors initiated by other driver genes [26]. The molecular diversity of KRAS-mutant tumors resulted in variable response with the same therapeutic strategy [27]. KRAS mutation was associated with better outcomes of PD1/PD-L1 blockade, which is consistent with the observed correlation of KRAS mutation with higher PD-L1 expression in tumor specimens. Although molecular diversity existed, heterogeneity was not present as expected. The positive correlation was more obvious in the smoker dominant group. Higher levels of PD-L1 expression in tumor samples may explain better outcomes of PD-1/PD-L1 inhibitor therapy in patients with KRAS mutation. Since KRAS mutations are linked to somatic mutational burden, smoking history and expression of neoantigens, our conclusion is in agreement with the emerging concept that tumors with a high mutational burden are more likely to benefit from immunotherapy [28]. Interestingly, driver gene (EGFR/ALK/KRAS) initiated tumors did not show higher levels of PD-L1 as compared to that in triple wild type NSCLCs. This indicated that driver gene (EGFR/ALK/KRAS) mutations may not fully reflect the mutational burden. Furthermore, studies of the relationship of PD-L1 expression and other driver genes like HER2, MET, BRAF and PIK3CA in NSCLCs have been rare. Although none of these genes were found to be correlated with PDL1 expression in our study, the influence of their status on tumor microenvironment deserves further investigation.
This study has a number of limitations. First, whether IHC is an ideal method to assess PD-L1 expression level is debatable. Partial PD 1/PD-L1 blocking response was seen in patient samples who were judged as “PD-L1 negative” by IHC [6, 29]. Second, TME is under dynamic regulation and PD-L1 expression status may alter in different stages of tumors and during treatment. Due to limited data, we could not perform further analysis. Co-mutation of driver gene exists in a certain proportion of NSCLCs, [30]. and we were not able to evaluate whether PD-L1 expression had impact on various co-mutation status. Moreover, the meta-analysis is based on retrospective studies, which may have introduced an element of bias.
CONCLUSIONS
In conclusion, we found that PD-L1 expression levels in NSCLCs with EGFR mutation were lower than those with EGFR wild type. As compared to the KRAS wild type tumors, those with KRAS mutation tend to express higher levels of PD-L1. ALK, BRAF, HER2, PIK3CA and MET expression status were not significantly associated with PD-L1 expression. As compared to triple negative NSCLCs, driver gene (EGFR/ALK/KRAS) mutations did not alter the level of PD-L1.
METHOD
Search strategy
A search for relevant studies was conducted on PubMed, Web of Science and Embase databases. Studies published up to April 26, 2017 were eligible for inclusion. The combinations of key words used are listed in Table 3. Two independent investigators (Bo Lan and Chengxi Ma) were responsible for the assessment of the included studies, and a third investigator (Chengyan Zhang) was involved in resolving any disagreements between the first two investigators.
Table 3: Key words used for literature search
#1 |
((((“Antigens, CD274”[Mesh]) OR B7 H1[Title/Abstract]) OR CD274[Title/Abstract]) OR PD L1[Title/Abstract]) OR Programmed Cell Death 1 Ligand 1 [Title/Abstract] |
#2 |
((“Receptor, Epidermal Growth Factor”[Mesh]) OR Epidermal Growth Factor Receptor) OR EGFR |
#3 |
(((CD246) OR anaplastic lymphoma kinase) OR ALK) OR EML4-ALK fusion |
#4 |
((Kirsten Ras) OR Kirsten rat sarcoma viral oncogene homolog) OR KRAS |
#5 |
(((((((((“Lung Neoplasms”[Mesh]) OR Pulmonary Neoplasms[Title/Abstract]) OR Lung Neoplasms[Title/ Abstract]) OR Pulmonary cancer[Title/Abstract]) OR Lung cancer[Title/Abstract])) OR Lung adenocarcinoma [Title/Abstract]) OR Pulmonary adenocarcinoma[Title/Abstract]) OR Lung squamous carcinoma[Title/Abstract]) OR Pulmonary squamous carcinoma[Title/ Abstract] |
#6 |
#2 OR #3 OR #4 |
#1 AND #6 AND #5 |
Selection criteria
Eligible studies were required to meet the following criteria to be included: 1) Patients were diagnosed with NSCLC by histopathological methods; 2) Expression of PD-L1 in tumor tissues was evaluated by immunohistochemistry and the specific cutoff for PD-L1 positive expression was reported; 3) At least one driver gene was involved in the study; 4) Distribution of driver gene in participants was reported; 5) Sufficient data to form an 2 × 2 table was reported: number of PD-L1 positive specimens with or without driver genes mutation, and number of PD-L1 negative specimens with or without driver genes mutation.
Exclusion criteria were as follows: 1) Studies not related to NSCLC; 2) Studies on patients with metastatic lung cancer; 3) PD-L1 expression was not evaluated in tumor tissues; 4) Other methods were used for evaluation of expression of PD-L1 such as ELISA, gene copy number or RNA expression level; 5) Studies that included patients <18 years of age or those that included patients with more than one cancer; 6) Non-English publication; 7) Overlapping studies; 8) Editorials, letters, reviews, case reports, duplicate publications, and expert opinions.
Quality assessment
In order to assess the quality of each included study, three investigators (Bo Lan, Shoujie Chai and Pingli Wang) scored the individual studies according to the Strengthening the Reporting of Observational studies in Epidemiology-Molecular Epidemiology (STROBE-ME) statement [31].(supplement). Quality of each included study was appraised on several aspects and each aspect contained several items. We scored the possible values (0, 1) for each item by consensus at a meeting of all involved investigators. Final scores were displayed as percentages of maximal score. According to percentages of final score (Supplementary Table 1), quality of included studies was classified as: “high-quality” (100%–75%); “moderate-quality” (74%–50%); “low-quality” (49%–25%); “very low-quality” (24%–0%).
Data extraction
Data from the included studies was extracted independently by two investigators (Bo Lan and Chengxi Ma). A third investigator (Chengyan Zhang) was responsible for resolving any disagreement between the first two investigators. The following information were collected: last name of first author, year of publication, ethnicity, median age, gender, stage, smoking status, histological type, primary PD-L1 antibody used for IHC assessment, cutoff level for PD-L1 positive, status of driver genes and PD-L1 expression (Table 1).
Statistical analysis
All data analysis was performed using software STATA version 14.0 (Stata Corp). Pooled odds ratios (OR) of PD-L1 positive with 95% CI for driver gene status were calculated by fixed or random effect model and forest plots generated. I2 and p statistics were selected as measures of heterogeneity. P value < 0.05 or I2 > 0.0% were considered indicative of existence of significant heterogeneity. I2 ≤ 25.0% indicated low heterogeneity while 25% and 75% were defined as the thresholds for moderate and high heterogeneity, respectively. I2 ≤ 50.0% was considered indicative of an acceptable level of heterogeneity [32]. A random effect model was used when significant heterogeneity was found; otherwise, fixed effect model was used. Sensitivity analysis was performed by sequential omission of one study at a time to explore the effect of individual studies on the pooled OR. Subgroup analysis using random effect model was performed to detect association of PD-L1 expression and driver gene status with different population characteristics (median age, cutoff value, ethnicity, histological type, stage, gender and smoking status), as well as to assess the source of heterogeneity. “Cutoff” was defined as the score or percentage of positive staining tumor cells used for classification of a specimen as “PD-L1 positive” in each study. In the event of an unacceptable level of heterogeneity, meta regression with Knapp-Hartung method was used to assess the effect of median age, cutoff level, ethnicity, histological type, stage, gender, smoker and quality of publication. Publication bias was evaluated by Begg’s rank correlation test and Egger’s linear regression test. P < 0.05 was considered indicative of statistically significant difference. Once significant publication bias was observed, trim and fill procedure was performed to evaluate the number of missing publications and the pooled ORs with 95% CI adjusted.
ACKNOWLEDGMENTS AND FUNDING
We thank Jiyong Jing for his professional advice. The research was fund by grants from the National Natural Science Foundation of China (No. 81472170), the Health Department of Zhejiang Province (No.2016135484), the Key National Research and Developmental Plan of China (No.2016YFC0902300), the Major Science and Technology Special Project of Zhejiang Province (No. 2015C03043), the Natural Science Foundation of Zhejiang Province (No. LY13H010001).
CONFLICTS OF INTEREST
There are no potential conflicts of interest to report.
REFERENCES
1. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA: A Cancer Journal for Clinicians. 2016; 66:115–32. https://doi.org/10.3322/caac.21338.
2. Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA. Non-Small Cell Lung Cancer: Epidemiology, Risk Factors, Treatment, and Survivorship. Mayo Clinic Proceedings. 2008; 83:584–94. https://doi.org/10.4065/83.5.584.
3. Miller KD, Siegel RL, Lin CC, Mariotto AB, Kramer JL, Rowland JH, Stein KD, Alteri R, Jemal A. Cancer treatment and survivorship statistics, 2016. CA: A Cancer Journal for Clinicians. 2016; 66:271–89. https://doi.org/10.3322/caac.21349.
4. Wang J, Yuan R, Song W, Sun J, Liu D, Li Z. PD-1, PD-L1 (B7-H1) and Tumor-Site Immune Modulation Therapy: The Historical Perspective. Journal of Hematology & Oncology. 2017; 10: 34. https://doi.org/10.1186/s13045-017-0403-5.
5. Reck M, Rodríguez-Abreu D, Robinson AG, Hui R, Csőszi T, Fülöp A, Gottfried M, Peled N, Tafreshi A, Cuffe S, O’Brien M, Rao S, Hotta K, et al. Pembrolizumab versus Chemotherapy for PD-L1-Positive Non-Small-Cell Lung Cancer. New England Journal of Medicine. 2016; 375:1823–33. https://doi.org/10.1056/NEJMoa1606774.
6. Brahmer J, Reckamp KL, Baas P, Crinò L, Eberhardt WEE, Poddubskaya E, Antonia S, Pluzanski A, Vokes EE, Holgado E, Waterhouse D, Ready N, Gainor J, et al. Nivolumab versus Docetaxel in Advanced Squamous-Cell Non-Small-Cell Lung Cancer. The New England journal of medicine. 2015; 373:123–35. https://doi.org/10.1056/NEJMoa1504627.
7. Borghaei H, Paz-Ares L, Horn L, Spigel DR, Steins M, Ready NE, Chow LQ, Vokes EE, Felip E, Holgado E, Barlesi F, Kohlhäufl M, Arrieta O, et al. Nivolumab versus Docetaxel in Advanced Nonsquamous Non-Small-Cell Lung Cancer. New England Journal of Medicine. 2015; 373:1627–39. https://doi.org/doi:10.1056/NEJMoa1507643.
8. Rizvi NA, Mazières J, Planchard D, Stinchcombe TE, Dy GK, Antonia SJ, Horn L, Lena H, Minenza E, Mennecier B, Otterson GA, Campos LT, Gandara DR, et al. Activity and safety of nivolumab, an anti-PD-1 immune checkpoint inhibitor, for patients with advanced, refractory squamous non-small-cell lung cancer (CheckMate 063): a phase 2, single-arm trial. The Lancet Oncology. 16:257–65. https://doi.org/10.1016/S1470-2045(15)70054-9.
9. Fehrenbacher L, Spira A, Ballinger M, Kowanetz M, Vansteenkiste J, Mazieres J, Park K, Smith D, Artal-Cortes A, Lewanski C, Braiteh F, Waterkamp D, He P, et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. The Lancet. 387:1837–46. https://doi.org/10.1016/S0140-6736(16)00587-0.
10. Johnson BE, Jänne PA. Epidermal growth factor receptor mutations in patients with non-small cell lung cancer. Cancer Research. 2005; 65:7525–9. https://doi.org/10.1158/0008-5472.CAN-05-1257.
11. Chang YL, Wu CT, Lin SC, Hsiao CF, Jou YS, Lee YC. Clonality and Prognostic Implications of p53 and Epidermal Growth Factor Receptor Somatic Aberrations in Multiple Primary Lung Cancers. Clinical Cancer Research. 2007; 13:52–8. https://doi.org/10.1158/1078-0432.ccr-06-1743.
12. Wu CT, Lin MW, Hsieh MS, Kuo SW, Chang YL. New aspects of the clinicopathology and genetic profile of metachronous multiple lung cancers. Annals of Surgery. 2013; 259:1018. https://doi.org/10.1097/SLA.0000000000000385.
13. Gou LY, Niu FY, Wu YL, Zhong WZ. Differences in driver genes between smoking-related and non-smoking-related lung cancer in the Chinese population. Cancer. 2015; 121:3069–79. https://doi.org/10.1002/cncr.29531.
14. Bircan S, Baloglu H, Kucukodaci Z, Bircan A. EGFR and KRAS mutations in Turkish non-small cell lung cancer patients: a pilot study. Medical Oncology. 2014; 31:87. https://doi.org/10.1007/s12032-014-0087-4.
15. Gainor JF, Shaw AT, Sequist LV, Fu X, Azzoli CG, Piotrowska Z, Huynh TG, Zhao L, Fulton L, Schultz KR, Howe E, Farago AF, Sullivan RJ, et al. EGFR Mutations and ALK Rearrangements Are Associated with Low Response Rates to PD-1 Pathway Blockade in Non-Small Cell Lung Cancer: A Retrospective Analysis. Clinical Cancer Research. 2016; 22:4585–93. https://doi.org/10.1158/1078-0432.ccr-15-3101.
16. Akca H, Tani M, Hishida T, Matsumoto S, Yokota J. Activation of the AKT and STAT3 pathways and prolonged survival by a mutant EGFR in human lung cancer cells. Lung Cancer. 2006; 54:25–33. https://doi.org/10.1016/j.lungcan.2006.06.007.
17. Lin K, Cheng J, Yang T, Li Y, Zhu B. EGFR-TKI down-regulates PD-L1 in EGFR mutant NSCLC through inhibiting NF-κB. Biochemical and Biophysical Research Communications. 2015; 463:95–101. https://doi.org/10.1016/j.bbrc.2015.05.030.
18. Koh J, Jang JY, Keam B, Kim S, Kim MY, Go H, Kim TM, Kim DW, Kim CW, Jeon YK, Chung DH. EML4-ALK enhances programmed cell death-ligand 1 expression in pulmonary adenocarcinoma via hypoxia-inducible factor (HIF)-1α and STAT3. Oncoimmunology. 2016; 5:e1108514. https://doi.org/10.1080/2162402X.2015.1108514.
19. Ota K, Azuma K, Kawahara A, Hattori S, Iwama E, Tanizaki J, Harada T, Matsumoto K, Takayama K, Takamori S, Kage M, Hoshino T, Nakanishi Y, et al. Induction of PD-L1 Expression by the EML4–ALK Oncoprotein and Downstream Signaling Pathways in Non-Small Cell Lung Cancer. Clinical Cancer Research. 2015; 21:4014–21. https://doi.org/10.1158/1078-0432.ccr-15-0016.
20. Skoulidis F, Byers LA, Diao L, Papadimitrakopoulou VA, Tong P, Izzo J, Behrens C, Kadara H, Parra ER, Canales JR, Zhang J, Giri U, Gudikote J, et al. Co-occurring genomic alterations define major subsets of KRAS - mutant lung adenocarcinoma with distinct biology, immune profiles, and therapeutic vulnerabilities. Cancer discovery. 2015; 5:860–77. https://doi.org/10.1158/2159-8290.CD-14-1236.
21. Chen Z, Cheng K, Walton Z, Wang Y, Ebi H, Shimamura T, Liu Y, Tupper T, Ouyang J, Li J, Gao P, Woo MS, Xu C, et al. A murine lung cancer co-clinical trial identifies genetic modifiers of therapeutic response. Nature. 2012; 483:613–7. https://doi.org/10.1038/nature10937.
22. Chen N, Fang W, Zhan J, Hong S, Tang Y, Kang S, Zhang Y, He X, Zhou T, Qin T, Huang Y, Yi X, Zhang L. Upregulation of PD-L1 by EGFR Activation Mediates the Immune Escape in EGFR-Driven NSCLC: Implication for Optional Immune Targeted Therapy for NSCLC Patients with EGFR Mutation. Journal of Thoracic Oncology. 2015; 10:910–23. https://doi.org/10.1097/JTO.0000000000000500.
23. Stroup DF, Berlin JA, Morton SC, Olkin I, Williamson GD, Rennie D, Moher D, Becker BJ, Sipe TA, Thacker SB. Meta-analysis of observational studies in epidemiology: A proposal for reporting. JAMA. 2000; 283:2008–12. https://doi.org/10.1001/jama.283.15.2008.
24. Hong S, Chen N, Fang W, Zhan J, Liu Q, Kang S, He X, Liu L, Zhou T, Huang J, Chen Y, Qin T, Zhang Y, et al. Upregulation of PD-L1 by EML4-ALK fusion protein mediates the immune escape in ALK positive NSCLC: Implication for optional anti-PD-1/PD-L1 immune therapy for ALK-TKIs sensitive and resistant NSCLC patients. Oncoimmunology. 2016; 5:e1094598. https://doi.org/10.1080/2162402X.2015.1094598.
25. Shaabani N, Duhan V, Khairnar V, Gassa A, Ferrer-Tur R, Häussinger D, Recher M, Zelinskyy G, Liu J, Dittmer U, Trilling M, Scheu S, Hardt C, et al. CD169(+) macrophages regulate PD-L1 expression via type I interferon and thereby prevent severe immunopathology after LCMV infection. Cell Death & Disease. 2016; 7:e2446. https://doi.org/10.1038/cddis.2016.350.
26. Stinchcombe TE, Johnson GL. MEK Inhibition in Non-Small Cell Lung Cancer. Lung cancer (Amsterdam, Netherlands). 2014; 86:121–5. https://doi.org/10.1016/j.lungcan.2014.09.005.
27. Ihle NT, Byers LA, Kim ES, Saintigny P, Lee JJ, Blumenschein GR, Tsao A, Liu S, Larsen JE, Wang J, Diao L, Coombes KR, Chen L, et al. Effect of KRAS Oncogene Substitutions on Protein Behavior: Implications for Signaling and Clinical Outcome. JNCI Journal of the National Cancer Institute. 2012; 104:228–39. https://doi.org/10.1093/jnci/djr523.
28. Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, Lee W, Yuan J, Wong P, Ho TS, Miller ML, Rekhtman N, Moreira AL, et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science (New York, NY). 2015; 348:124–8. https://doi.org/10.1126/science.aaa1348.
29. Weber JS, D’Angelo SP, Minor D, Hodi FS, Gutzmer R, Neyns B, Hoeller C, Khushalani NI, Miller WH Jr, Lao CD, Linette GP, Thomas L, Lorigan P, et al. Nivolumab versus chemotherapy in patients with advanced melanoma who progressed after anti-CTLA-4 treatment (CheckMate 037): a randomised, controlled, open-label, phase 3 trial. The Lancet Oncology. 16:375–84. https://doi.org/10.1016/S1470-2045(15)70076-8.
30. Qu Y, Che N, Zhao D, Zhang C, Su D, Zhou L, Zhang L, Wang C, Zhang H, Wei L. The clinicopathological significance of ALK rearrangements and KRAS and EGFR mutations in primary pulmonary mucinous adenocarcinoma. Tumor Biology. 2015; 36:6417–24. https://doi.org/10.1007/s13277-015-3331-4.
31. Gallo V, Egger M, McCormack V, Farmer PB, Ioannidis JPA, Kirsch-Volders M, Matullo G, Phillips DH, Schoket B, Stromberg U, Vermeulen R, Wild C, Porta M, et al. STrengthening the Reporting of OBservational studies in Epidemiology–Molecular Epidemiology (STROBE-ME): An extension of the STROBE statement. European Journal of Clinical Investigation. 2012; 42:1–16. https://doi.org/10.1111/j.1365-2362.2011.02561.x.
32. Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. British Medical Journal. 2003; 327:557–60. https://doi.org/10.1136/bmj.327.7414.557.
33. Zhang Y, Wang L, Li Y, Pan Y, Wang R, Hu H, Li H, Luo X, Ye T, Sun Y, Chen H. Protein expression of programmed death 1 ligand 1 and ligand 2 independently predict poor prognosis in surgically resected lung adenocarcinoma. OncoTargets and therapy. 2014; 7:567–73. https://doi.org/10.2147/OTT.S59959.
34. Yang CY, Lin MW, Chang YL, Wu CT, Yang PC. Programmed cell death-ligand 1 expression in surgically resected stage I pulmonary adenocarcinoma and its correlation with driver mutations and clinical outcomes. European Journal of Cancer. 2014; 50:1361–9. https://doi.org/10.1016/j.ejca.2014.01.018.
35. D’Incecco A, Andreozzi M, Ludovini V, Rossi E, Capodanno A, Landi L, Tibaldi C, Minuti G, Salvini J, Coppi E, Chella A, Fontanini G, Filice ME, et al. PD-1 and PD-L1 expression in molecularly selected non-small-cell lung cancer patients. British Journal of Cancer. 2015; 112:95–102. https://doi.org/10.1038/bjc.2014.555.
36. Cooper WA, Tran T, Vilain RE, Madore J, Selinger CI, Kohonen-Corish M, Yip P, Yu B, O’Toole SA, McCaughan BC, Yearley JH, Horvath LG, Kao S, et al. PD-L1 expression is a favorable prognostic factor in early stage non-small cell carcinoma. Lung Cancer. 2015; 89:181–8. https://doi.org/10.1016/j.lungcan.2015.05.007.
37. Schmidt LH, Kümmel A, Görlich D, Mohr M, Bröckling S, Mikesch JH, Grünewald I, Marra A, Schultheis AM, Wardelmann E, Müller-Tidow C, Spieker T, Schliemann C, et al. PD-1 and PD-L1 Expression in NSCLC Indicate a Favorable Prognosis in Defined Subgroups. PLoS ONE. 2015; 10:e0136023. https://doi.org/10.1371/journal.pone.0136023.
38. Kim MY, Koh J, Kim S, Go H, Jeon YK, Chung DH. Clinicopathological analysis of PD-L1 and PD-L2 expression in pulmonary squamous cell carcinoma: Comparison with tumor-infiltrating T cells and the status of oncogenic drivers. Lung Cancer. 2015; 88:24–33. https://doi.org/10.1016/j.lungcan.2015.01.016.
39. Chang YL, Yang CY, Lin MW, Wu CT, Yang PC. PD-L1 is highly expressed in lung lymphoepithelioma-like carcinoma: A potential rationale for immunotherapy. Lung Cancer. 2015; 88:254–9. https://doi.org/10.1016/j.lungcan.2015.03.017.
40. Koh J, Go H, Keam B, Kim MY, Nam SJ, Kim TM, Lee SH, Min HS, Kim YT, Kim DW, Jeon YK, Chung DH. Clinicopathologic analysis of programmed cell death-1 and programmed cell death-ligand 1 and 2 expressions in pulmonary adenocarcinoma: comparison with histology and driver oncogenic alteration status. Mod Pathol. 2015; 28:1154–66. https://doi.org/10.1038/modpathol.2015.63.
41. Tang Y, Fang W, Zhang Y, Hong S, Kang S, Yan Y, Chen N, Zhan J, He X, Qin T. The association between PD-L1 and EGFR status and the prognostic value of PD-L1 in advanced non-small cell lung cancer patients treated with EGFR-TKIs. Oncotarget. 2015; 6:14209–19. https://doi.org/https://doi.org/10.18632/oncotarget.3694.
42. Omori S. Changes in PD-L1 expression in non-small cell lung cancer by immunohistochemical analysis. Journal of clinical oncology. 2015; 33.
43. Yang CY, Lin MW, Chang YL, Wu CT, Yang PC. Programmed cell death-ligand 1 expression is associated with a favourable immune microenvironment and better overall survival in stage I pulmonary squamous cell carcinoma. European Journal of Cancer. 2016; 57:91–103. https://doi.org/10.1016/j.ejca.2015.12.033.
44. Scheel AH, Ansén S, Schultheis AM, Scheffler M, Fischer RN, Michels S, Hellmich M, George J, Zander T, Brockmann M, Stoelben E, Groen H, Timens W, et al. PD-L1 expression in non-small cell lung cancer: Correlations with genetic alterations. Oncoimmunology. 2016; 5:e1131379. https://doi.org/10.1080/2162402X.2015.1131379.
45. Ameratunga M, Asadi K, Lin X, Walkiewicz M, Murone C, Knight S, Mitchell P, Boutros P, John T. PD-L1 and Tumor Infiltrating Lymphocytes as Prognostic Markers in Resected NSCLC. PLOS ONE. 2016; 11:e0153954. https://doi.org/10.1371/journal.pone.0153954.
46. Song Z, Yu X, Cheng G, Zhang Y. Programmed death-ligand 1 expression associated with molecular characteristics in surgically resected lung adenocarcinoma. Journal of Translational Medicine. 2016; 14:188. https://doi.org/10.1186/s12967-016-0943-4.
47. Inamura K, Yokouchi Y, Sakakibara R, Kobayashi M, Subat S, Ninomiya H, Nagano H, Nomura K, Okumura S, Ishikawa Y. Relationship of tumor PD-L1 expression with EGFR wild-type status and poor prognosis in lung adenocarcinoma. Japanese Journal of Clinical Oncology. 2016; 46:935–41. https://doi.org/10.1093/jjco/hyw087.
48. Jia X, Zhang L, Wu W, Zhang W, Wu C. Driver Mutation Analysis and PD-L1 Expression in Synchronous Double Primary Lung Cancer. Applied Immunohistochemistry & Molecular Morphology Aimm. 2016: 1. https://doi.org/10.1097/PAI.0000000000000412.
49. Inoue Y, Yoshimura K, Mori K, Kurabe N, Kahyo T, Mori H, Kawase A, Tanahashi M, Ogawa H, Inui N, Funai K, Shinmura K, Niwa H, et al. Clinical significance of PD-L1 and PD-L2 copy number gains in non-small-cell lung cancer. Oncotarget. 2016; 7:32113–28. https://doi.org/10.18632/oncotarget.8528.
50. Ji M, Liu Y, Li Q, Li X, Ning Z, Zhao W, Shi H, Jiang J, Wu C. PD-1/PD-L1 expression in non-small-cell lung cancer and its correlation with EGFR/KRAS mutations. Cancer Biology & Therapy. 2016; 17:407–13. https://doi.org/10.1080/15384047.2016.1156256.
51. Huynh TG, Morales-Oyarvide V, Campo MJ, Gainor JF, Bozkurtlar E, Uruga H, Zhao L, Gomez-Caraballo M, Hata AN, Mark EJ, Lanuti M, Engelman JA, Mino-Kenudson M. Programmed Cell Death Ligand 1 Expression in Resected Lung Adenocarcinomas: Association with Immune Microenvironment. Journal of Thoracic Oncology. 2016; 11:1869–78. https://doi.org/10.1016/j.jtho.2016.08.134.
52. Mori S, Motoi N, Ninomiya H, Matsuura Y, Nakao M, Mun M, Okumura S, Nishio M, Morikawa T, Ishikawa Y. High expression of programmed cell death 1 ligand 1 in lung adenocarcinoma is a poor prognostic factor particularly in smokers and wild-type epidermal growth-factor receptor cases. Pathology International. 2017; 67:37–44. https://doi.org/10.1111/pin.12489.
53. Takada K, Okamoto T, Shoji F, Shimokawa M, Akamine T, Takamori S, Katsura M, Suzuki Y, Fujishita T, Toyokawa G, Morodomi Y, Okano S, Oda Y, et al. Clinical Significance of PD-L1 Protein Expression in Surgically Resected Primary Lung Adenocarcinoma. Journal of Thoracic Oncology. 2016; 11:1879–90. https://doi.org/10.1016/j.jtho.2016.06.006.
54. Dong ZY, Zhong W, Zhang XC, Su J, Xie Z, Liu SY, Tu HY, Chen HJ, Sun YL, Zhou Q, Yang J, Yang X, Lin JX, et al. Potential Predictive Value of TP53 and KRAS Mutation Status for Response to PD-1 Blockade Immunotherapy in Lung Adenocarcinoma. Clinical Cancer Research. 2017. https://doi.org/10.1158/1078-0432.ccr-16-2554.
55. Rangachari D, VanderLaan PA, Shea M, Le X, Huberman MS, Kobayashi SS, Costa DB. Correlation between Classic Driver Oncogene Mutations in EGFR, ALK, or ROS1 and 22C3–PD-L1 ≥ 50% Expression in Lung Adenocarcinoma. Journal of Thoracic Oncology. 2017; 12:878–83. https://doi.org/10.1016/j.jtho.2016.12.026.
56. Tsao MS, Le Teuff G, Shepherd FA, Landais C, Hainaut P, Filipits M, Pirker R, Le Chevalier T, Graziano S, Kratze R, Soria JC, Pignon JP, Seymour L, et al. PD-L1 protein expression assessed by immunohistochemistry is neither prognostic nor predictive of benefit from adjuvant chemotherapy in resected non-small cell lung cancer. Annals of Oncology. 2017; 28:882–9. https://doi.org/10.1093/annonc/mdx003.
57. Chen J, Li H, Pang R, Huang J. Altered status of programmed death-ligand 1 after recurrence in resected lung adenocarcinoma patients. OncoTargets and therapy. 2017; 10:2003–7. https://doi.org/10.2147/OTT.S127498.
58. Cho JH, Zhou W, Choi YL, Sun JM, Choi H, Kim TE, Dolled-Filhart M, Emancipator K, Rutkowski MA, Kim J. Retrospective Molecular Epidemiology Study of PD-L1 Expression in Patients with EGFR-Mutant Non-small Cell Lung Cancer. J Korean Cancer Assoc. 2017. https://doi.org/10.4143/crt.2016.591.

