INTRODUCTION
Ovarian cancer is the seventh most common cancer in women and is the eighth most frequent cause of cancer death among women [1]. The occurrence of ovarian cancer has been related to many factors, for example, age of menarche [2, 3], short or irregular cycles [2–6], age of menopause [7, 8], age of the first birth [2, 3, 9, 10], age of last pregnancy [7, 10, 11], the number of children [2, 3, 7, 10, 11], period of breastfeeding [12–16], oral contraceptives [17–21], intake of phytochemicals [22], genes BRCA1 or BRCA2 [23], etc.
As many diseases show clear patterns in their geographic and race distributions, ovarian cancer also has preference in these regards, for example, the women from East Asia have a very low incidence rate and the lowest mortality rate [1, 24]. Death due to ovarian cancer is more common in North America and Europe than in Africa and Asia [1]. High rates of epithelial ovarian cancer are reported in industrialized nations with the exception of Japan [24], which may be due to diet in those countries.
Ovarian cancer and its stem cell
As ovarian cancer is a high death-to-incidence disease, the role of cancer stem cells in ovarian cancer cannot be ignored because cancer stem cells are frequently resistant to chemotherapeutic and radiation treatments [25]. For example, drug resistance is due to the fact that specific therapies enrich cancer stem cells in residual pancreatic cancer treated with gemcitabine [26], colorectal cancer treated with cyclophosphamide [27], hepatocellular carcinoma treated with doxorubicin and fluorouracil [28] and lung cancer treated with cisplatin, doxorubicin, and methotrexate [29].
Ovarian cancer has a great degree of heterogeneity because it may arise from germ cell, stromal, or epithelial compartments [24]. Similarly, ovarian cancer is the best example of intra-tumor heterogeneity of cancer stem cell [30, 31]. It was hypothesized that ovarian cancer is driven and sustained by cancer stem cell as shown by CD44+/CD24- [32, 33], CD117 and CD133 [34, 35], especially ALDH1A1 [36–38]. Indeed, ALDH+/CD133+ ovarian cancer primary cells were defined as the top of hierarchical structure in ovarian cancer and as stem cell markers. This combination is a more multipotent phenotype than others including ALDH-/CD133+ [39]. Furthermore, it was reported that ALDH1 is better than CD133 in terms of identification of primary ovarian carcinoma-derived cells, which express stemness genes and are capable of self-renewal and tumor initiation [40].
As a marker of stem cells in both normal tissues and cancers [41], ALDH1 plays an extremely important role in ovarian cancer stem cells [25, 42] because studies show that ALDH1 activity is directly related to a subpopulation of ovarian cancer cells with cancer stem cell-like properties [36, 37, 39, 43–45]. Of ALDH1 isozymes (ALDH1A1, ALDH1A2 and ALDH1A3), expression of ALDH1A1 is prominent in cancer stem cell [42], for instance, in human breast cancer cell lines [46]. The role of ALDH1A1 in ovarian cancer stem cells, although it is not fully clear, is not far from its nominal role, i.e. its detoxifying role in terms of preventing the accumulation of reactive oxygen species and of reactive aldehydes. Additional role of ALDH1A1 is its regulatory function on ATP-binding cassette (ABC) drug transporters, which in turn leads to the resistance of ovarian cancer to chemotherapeutics [47].
It was found that ovarian cancer patients, who have high levels of type I receptor tyrosine kinase-like orphan receptor (ROR1), have stem cell-like gene-expression signatures, and their relapse rate and median survival are higher and shorter than other patients. The importance is that ROR1-positive (ROR1+) cells also expressed ALDH1 [48]. In ovarian cancer stem cells, ALDH1 have relatively high enzymatic activity [38, 49, 50], which no doubt intensifies detoxification of intracellular aldehydes as well as cytotoxic drugs [41, 51]. Accordingly, ALDH1 offers resistance to chemotherapeutics and radiation therapy [45]. Such role was confirmed by the finding that the knockdown of ALDH1A1 gene in ovarian cancer cell lines can restore the ovarian cancer cells’ sensitivity to chemotherapy in vitro [47] and xenograft models in mouse [38]. Therefore, the role of ALDH1 in ovarian cancer stem cells is far beyond the detoxification.
Regulation of ALDH1A1 in ovarian cancer stem cells is suggested at the transcriptional level through Wnt/β-catenin pathway [52]. Knockdown of ALDH1A1 in ovarian cancer cell line A2780 decreases regulators KLF4 and p21, which are the cell cycle checkpoints, and leads to an enhanced cell proliferation. This is the anti-proliferative role of ALDH1A1 because actively proliferating cells are more subject to cytotoxic drugs and so loss of ALDH1A1 contributes to the sensitization of ovarian carcinoma cells to chemotherapy [53] although a study showed the difference due to interplay between ALDH1A1 and the stemness-associated gene SOX2 [40]. Also, loss of ALDH1A1 triggered DNA damage suggesting that ALDH1A1 plays a genome-protecting role in ovarian cancer stem cells [53]. As a result, it is suggested that ALDH1A1 could be potential therapeutic target because a small-molecule ALDH1A1 inhibitor abolished sphere formation in ovarian cancer [52].
ALDH2
Apart from research in ALDH1 (ALDH1A1) in cancer stem cells, in fact, research interests in ALDH have been greatly increased recently [54], especially, ALDH2 [55], because ALDH2 is the most efficient enzyme for the metabolism of ethanol-derived acetaldehyde with the lowest Km [56]. ALDH has three different classes in mammals: class 1 (low Km, cytosolic), class 2 (low Km, mitochondrial), and class 3 (high Km, such as those expressed in tumors, stomach, and cornea). In all three classes, constitutive and inducible forms exist. ALDH1 and ALDH2 are the most important enzymes for aldehyde oxidation, and both are tetrameric enzymes composed of 54 kDa subunits.
The enzymatic reaction catalyzed by ALDH seems to be simple in humans, i.e. alcohol dehydrogenase (ADH) catalyzes ingested ethanol to acetaldehyde, and ALDH, mainly mitochondrial enzyme ALDH2, catalyzes acetaldehyde into acetate. Therefore, a single point mutation in ALDH2, termed ALDH2*2, wherein a lysine residue replaces a glutamate in the active site at position 487 of ALDH2 [57], causes facial flushing, headaches, nausea, dizziness, and cardiac palpitations in humans after alcohol consumption [58–61] because aldehydes cannot be fully detoxified [62]. Homozygous individuals with the mutant allele have almost no ALDH2 activity, and those heterozygous for the mutation have reduced activity. Thus, the mutation is partially dominant.
What is less known is that ALDH2 metabolizes numerous short-chain aliphatic aldehydes, aromatic and polycyclic aldehydes [63], environmental aldehydes such as acrolein in tobacco smoke and in car exhaust, and particularly endogenous aldehydic products from lipid peroxidation under oxidative stress, such as 4-hydroxy-2-nonenal (4-HNE) and malondialdehyde (MDA) [64, 65].
Consequently, many diseases and health problems can be related to ALDH2 mutation, whose carriers were associated to myocardial infarction [66], impaired myocardial function in rodents and humans [65, 67, 68], hypertension [69], blood pressure variation in East Asians [70], non-insulin-dependent diabetes mellitus due to the maternal ALDH2 [71, 72], a higher incidence of Alzheimer’s disease in Asian patients with the inactivating ALDH2*2 mutation [73–75], increase of esophageal cancer risk no matter of alcoholic beverage drinkers or not [76].
Intriguingly, the ALDH2*2 mutation exists mainly in 560 million East Asians [61, 77, 78] rather than the rest parts of the world [79]. This is because most Caucasians have both active ALDH1 and ADLH2, while approximately 50% of East Asians have active ALDH1 but not active ALDH2 due to its mutation. Therefore, flushing symptom is more popular for East Asians than for Caucasians, and the increased exposure to acetaldehyde in individual may be more susceptible to many types of cancer [55]. Many other studies from Japan, Taiwan, and China have overwhelmingly confirmed the significant association between ALDH2 enzyme deficiency and upper aerodigestive track (oropharyngolaryngeal, esophageal, stomach, colon and lung) cancer risk [80–86].
In broader sense, high ALDH expression is associated with a poor prognosis in acute myeloid leukaemia [87, 88], breast cancer [46, 89–91], early-stage lung cancer [92], head and neck squamous cell carcinoma [93], pancreas cancer [94], and prostate cancer [95]. Although ALDH2*2 mutation is associated with various diseases [55], there are exceptions, for example, liver cancer [96], Parkinson’s disease [97], and stroke [98, 99].
To go further, one can find the difference between male and female ALDH2*2 carriers with respect to different diseases. Chinese male ALDH2*2 carriers have a significantly higher incidence of acute coronary syndrome than noncarriers [100]. Also it demonstrated that the ALDH2*2 genotype is a risk factor for myocardial infarction in Japanese men [101]. On the other hand, ALDH2*2 genotype seemed to be a risk factor for non-insulin-dependent diabetes mellitus in women, but not in men [102]. Premenopausal females have a lower risk for cardiovascular disease, because female hearts have increased phosphorylation and activity of ALDH2 in ischemia and reperfusion injury, so ALDH2 activator is more effective in males than in females, and inhibition blocks the phosphorylation of ALDH2 in females, but had no effect in males [103]. A study including 2,200-plus Japanese between 40 and 70 years showed a clear higher level of serum lipid peroxides in female ALDH2*2 carriers after exclusion of alcohol drinking behavior [104].
If this ALDH2*2 mutation is considered so harmful [55], why it has been so popular in East Asia existing for 2000–3000 years [105]. What is the advantage in evolution to keep this mutation although unanswered hypotheses have been given [106, 107] ? In the past, ALDH2 mutation was considered as benign because it is a limiting factor for over drinking of alcoholic beverage [61, 108–110]. Another piece of evidence that shows the positive aspect of ALDH2 with clear difference between males and females is the ALDH2*2 genotype was associated with a longer life for male Koreans [111]. In addition, there is difference in ALDS2*2 carriers with certain disease in East Asia with respect to locality, for example, the number of Fanconi anemia patients with ALDH2*2 in Japan and Korea [112, 113] is far higher than in China or Taiwan, where the percentage of ALDH2*2 carriers is higher i.e., > 40% in Taiwan.
Over here, comparison between East Asian women with East Asian men is to indicate that ALDH2 plays different roles in East Asian women and men with respect to different diseases, so we could not simply say ALDH2 bad or good. On the other hand, one may wonder why we do not refer studies on East Asian women with Caucasian women. Actually, our theme begins to mention that East Asian women have a low incidence and mortality of ovarian cancers, which is the comparison between East Asian women with Caucasian women as well as women from the rest parts of world. Although a huge amount studies have been done to compare East Asian women with Caucasian women with respect to various diseases, there is no study conducted on ovarian cancer with respect to ALDH2. This is why we cannot conduct a meta-analysis to combine such studies from all the centers, whereas we can only propose a deduced hypothesis. Furthermore, one may also wonder who will conduct such a study to compare East Asian women with Caucasian women on ALDH2, simply because East Asian women generally do not have ALDH2.
Very interestingly and curiously, the relationship between ALDH2 and ovarian cancer has drawn little attention. Indeed, in transgenic mouse, the ALDH2*2 mutant subunits overexpressed particularly notably in cardiac and skeletal muscles [114] rather than ovary. Alcohol consumption does not appear to be related to ovarian cancer [115].
Taken all the references together, it is highly likely that the very low incidence rate and the lowest mortality rate of ovarian cancer in East Asian women is mainly due to the fact that many East Asian women are ALDH2*2 carriers. Therefore the ovarian stem cancer cells in East Asian women are more susceptible to environmental and endogenous aldehydic products at early stage of ovarian cancer and to chemotherapy at later stage of ovarian cancer, and these “toxic” substances could kill cancer stem cells readily. These are reasonable because ALDH cannot actively detoxify these substances in East Asian women who carry ALDH2*2. Indeed, although 4-HNE is capable to covalently bind to DNA as an important factor of carcinogenesis, it is also cytotoxic for cancer cells and can modulate their growth [116]. Thus ALDH1A1 inhibitor [41, 52] and eradication of ALDH high expression cells [117–121] were advocated although ALDH2 was not mention. From laboratory viewpoint, the determination of ALDH1A1 activity in live cells and of isolating ALDH1-positive cells with a fluorescence-based assay [122] is perhaps easier than determination of ALDH2, which is located in mitochondria although it is more active than ALDH1. Very strictly speaking, the study should be conducted in such a way that includes only East Asian women with ALDH2*2 versus ALDH2 with respect to their incidence and mortality of ovarian cancer. Therefore, we do not have direct experimental evidence to connect ALDH2 with ovarian cancer, but we can only deduce such a relationship using various pieces of knowledge in literature, which is the reason of why we propose our hypothesis and hope such hypothesis can stimulate more discussions and experiments. Fortunately, the transcriptomic data can provide experimental evidence for ALDH2 activity in cancer patients, which provide additional support for our literature evidence in the next section.
Another piece of evidence to support ALDH2’s role in ovarian cancer is that the risk of ovarian cancer goes up with age, and ALDH2*2 homozygous genotype was significantly reduced in females in the 60–70s age group versus 40–50s group in a study with more than 2,200 Japanese [104].
One may argue that our guess based on several paragraphs, however Newton’s guess on gravity was only based on the fact that an apple falls from tree. Therefore, the importance of guess is not dependent on how many paragraphs but on deduction. Moreover our guess is supported by fully referenced literature, which shows the ALDH’s role in cancer stem cells, as a component of ALDH, ALDH2 should play the same role as ALDH does although it was not on clinical routine screening now.
Evidence from cancer stem cells
Ideally the best way to test our reasoning and rationale is to compare the ALDH2 activity in ovarian cancer stem cells with the killing rate of chemotherapy for ovarian cancer stem cells or compare the ALDH2 activity in ovarian stem cells with the ovarian cancer incidence. Currently, this could be impossible. To take a step back, we analyze all the available transcriptomic data (GSE19713 [123], GSE23806 [124–126], GSE28799 [127], GSE35603 [128], GSE67966) of different cancer stem cells in Gene Expression Omnibus (GEO) [129, 130] using platform GPL570 [131] with ALDH1A1 and ALDH2 expression against 5-year overall survival data in different cancers documented in literature (Supplementary Table 1), if we consider ALDH1A1 and ALDH2 expression somewhat similar to their activities.
Figure 1 shows this analysis. In this figure, samples of eight types of cancers are further divided into parental tumor cell (PTC) and tumor stem-like cell (TSC), so there are 16 labels along x-axis. ALDH1A1 and ALDH2 expressions are presented as circles and triangles. As can be seen, the expression of ALDH1A1 and ALDH2 vary in different cancers, as well as between ovarian cancer PTC and TSC. ALDH expressions, especially the expression of ALDH1A1, are low in atypical teratoid/rhabdoid tumour and cancers from head and neck, breast, and prostate, but high in the rest of cancers. Moreover, the average 5-year overall survival (OS) is presented in this figure as square symbol for 8 different cancers because OS does not distinguish PTC from TSC.
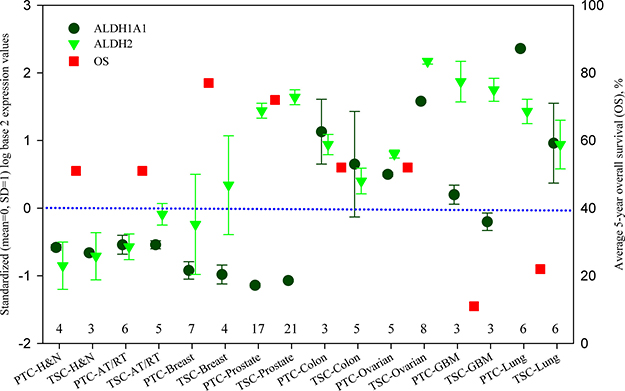
Figure 1: ALDH1A1 and ALDH2 expression of parental tumor cells and tumor stem-like cells, and 5-year overall survival in different cancers. ALDH1A1 and ALDH2 are presented by standardized (mean = 0, SD = 1) log base 2 expression values at the left axis. The data were calculated from the Series GSE19713, GSE23806, GSE28799, GSE35603 and GSE67966, and their sample numbers are listed above the x-axis. Average 5-year overall survival (OS) data are obtained from the literatures listed in Supplementary Table 1 and presented at the right axis. PTC: parental tumor cell, TSC: tumor stem-like cell, AT/RT: Atypical teratoid/rhabdoid tumour, GBM: glioblastoma, H & N: head and neck.
A general trend can be found in Figure 1, that is, OS is higher when the expressions of ALDH1A1 and ALDH2 are lower, whereas OS is lower when the expressions of ALDH1A1 and ALDH2 are higher. This feature is consistent with previous studies [46, 87, 88, 89–95] and can suggest the detoxification of ALDH on chemotherapy. Although prostate cancer has high ALDH2 expression with high OS, its ALDH1A1 expression is indeed very low. Because ALDH1A1 has an anti-proliferative role, a low activity of ALDH1A1 can promote cell proliferation and increase the sensitivity of cancer cells to chemotherapy [53]. Yet, great caution should be paid to OS because we could not stratify OS according to the treatments of chemotherapy, radiotherapy and surgery. Also the baseline of OS is unknown so OS in ovarian cancer apparently is relatively high.
Comparing the expression between ALDH1A1 and ALDH2, stronger ALDH2 expression can be found in the tumor stem-like cells of atypical teratoid/rhabdoid tumour (TSC-AT/RT), and in both parental and stem-like tumor cells of breast cancer, prostate cancer, ovarian cancer and glioblastoma (GBM). This observation is in good agreement with the knowledge that ALDH2 plays a more important role on detoxification [56]. It is notable to see that ALDH2 expression is at very high level in stem-like tumor cells of ovarian cancer, indicating that a high ALDH2 activity of stem cells renders significant influence on the morbidity and mortality of ovarian cancer. Therefore the population who lack ALDH2 would be more sensitive to chemotherapy and internal toxic substances such as 4-HNE. Along this thought of line, we would have expected to see a high OS in East Asian women with ovarian cancer because their ALDH2 activity would be zero in ALDH2 mutation carriers. Therefore, our hypothesis is supported by all the available transcriptomic data on ovarian cancer stem cells.
Final remark
In this review, we use the evidence from literature, transcriptomic data with average 5-year overall survival to suggest that the key factor that determines the low incidence and mortality of ovarian cancer in East Asian women is the ALDH2 mutation.
Interestingly, East Asia with its ALDH2*2 mutation seems to be the most economically active place in the world for quite a considerable periods in human history. This leads to the question of whether ALDH2*2 mutation is helpful to intelligent development.
Abbreviations
4-HNE - 4-hydroxy-2-nonenal, ABC - ATP-binding cassette, ADH - alcohol dehydrogenase, ALDH - aldehyde dehydrogenase, GEO - Gene Expression Omnibus, MDA - malondialdehyde, OS - average 5-year overall survival, PTC - parental tumor cell, ROR1 - type I receptor tyrosine kinase-like orphan receptor, TSC - tumor stem-like cell.
Author contributions
G. Wu designed this review and wrote this first draft, S.M. Yan analyzed the data, and both finalized this review.
CONFLICTS OF INTEREST
None.
FUNDING
This study was partly supported by National Natural Science Foundation of China (31460296 for SM Yan, and 31560315 for G Wu), and Special Funds for Building of Guangxi Talent Highland (for G Wu).
REFERENCES
1. World Health Organization. World Cancer Report. 2014; Map 5.12.3. 2014.
2. Brinton LA, Berman ML, Mortel R, Twiggs LB, Barrett RJ, Wilbanks GD, Lannom L, Hoover RN. Reproductive, menstrual, and medical risk factors for endometrial cancer: results from a case-control study. Am J Obstet Gynecol. 1992; 167:1317–1325.
3. Titus-Ernstoff L, Perez K, Cramer DW, Harlow BL, Baron JA, Greenberg ER. Menstrual and reproductive factors in relation to ovarian cancer risk. Br J Cancer. 2001; 84:714–721.
4. McPherson CP, Sellers TA, Potter JD, Bostick RM, Folsom AR. Reproductive factors and risk of endometrial cancer. The Iowa Women’s Health Study. Am J Epidemiol. 1996; 143:1195–1202.
5. Purdie DM, Bain CJ, Siskind V, Webb PM, Green AC. Ovulation and risk of epithelial ovarian cancer. Int J Cancer. 2003; 104:228–232.
6. Casagrande JT, Louie EW, Pike MC, Roy S, Ross RK, Henderson BE. “Incessant ovulation” and ovarian cancer. Lancet. 1979; 2:170–173.
7. Dossus L, Allen N, Kaaks R, Bakken K, Lund E, Tjonneland A, Olsen A, Overvad K, Clavel-Chapelon F, Fournier A, Chabbert-Buffet N, Boeing H, Schütze M, et al. Reproductive risk factors and endometrial cancer: the European Prospective Investigation into Cancer and Nutrition. Int J Cancer. 2010; 127:442–451.
8. Chiaffarino F, Pelucchi C, Parazzini F, Negri E, Franceschi S, Talamini R, Conti E, Montella M, La Vecchia C. Reproductive and hormonal factors and ovarian cancer. Ann Oncol. 2001; 12:337–341.
9. Henderson BE, Casagrande JT, Pike MC, Mack T, Rosario I, Duke A. The epidemiology of endometrial cancer in young women. Br J Cancer. 1983; 47:749–756.
10. Whittemore AS, Harris R, Itnyre J. Characteristics relating to ovarian cancer risk: collaborative analysis of 12 US case-control studies. II. Invasive epithelial ovarian cancers in white women. Collaborative Ovarian Cancer Group. Am J Epidemiol. 1992; 136:1184–1203.
11. Lambe M, Wuu J, Weiderpass E, Hsieh CC. Childbearing at older age and endometrial cancer risk (Sweden). Cancer Causes Control. 1999; 10:43–49.
12. Siskind V, Green A, Bain C, Purdie D. Breastfeeding, menopause, and epithelial ovarian cancer. Epidemiology. 1997; 8:188–191.
13. Salazar-Martinez E, Lazcano-Ponce EC, Gonzalez Lira-Lira G, Escudero-De los Rios P, Salmeron-Castro J, Hernandez-Avila M. Reproductive factors of ovarian and endometrial cancer risk in a high fertility population in Mexico. Cancer Res. 1999; 59:3658–3662.
14. Rosenblatt KA, Thomas DB. Prolonged lactation and endometrial cancer WHO Collaborative Study of Neoplasia and Steroid Contraceptives. Int J Epidemiol. 1995; 24:499–503.
15. Okamura C, Tsubono Y, Ito K, Niikura H, Takano T, Nagase S, Yoshinaga K, Terada Y, Murakami T, Sato S, Aoki D, Jobo T, Okamura K, et al. Lactation and risk of endometrial cancer in Japan: a case-control study. Tohoku J Exp Med. 2006; 208:109–115.
16. Newcomb PA, Trentham-Dietz A. Breast feeding practices in relation to endometrial cancer risk, USA. Cancer Causes Control. 2000; 11:663–667.
17. Weiderpass E, Adami HO, Baron JA, Magnusson C, Lindgren A, Persson I. Use of oral contraceptives and endometrial cancer risk (Sweden). Cancer Causes Control. 1999; 10:277–284.
18. Schlesselman JJ. Risk of endometrial cancer in relation to use of combined oral contraceptives A practitioner’s guide to meta-analysis. Hum Reprod. 1997; 12:1851–1863.
19. Maxwell GL, Schildkraut JM, Calingaert B, Risinger JI, Dainty L, Marchbanks PA, Berchuck A, Barrett JC, Rodriguez GC. Progestin and estrogen potency of combination oral contraceptives and endometrial cancer risk. Gynecol Oncol. 2006; 103:535–540.
20. La Vecchia C. Oral contraceptives and ovarian cancer: an update 1998–2004. Eur J Cancer Prev. 2006; 15:117–124.
21. Silverberg SG, Makowski EL. Endometrial carcinoma in young women taking oral contraceptive agents. Obstet Gynecol. 1975; 46:503–506.
22. Mohammadi V, Dehghani S, Larijani B, Azadbakht L. Ovarian cancer risk and nonisoflavone flavonoids intake: A systematic review of epidemiological studies. J Res Med. Sci 2016; 21:123.
23. Jayson GC, Kohn EC, Kitchener HC, Ledermann JA. Ovarian cancer. Lancet. 2014; 384:1376–1388.
24. Cramer DW. The epidemiology of endometrial and ovarian cancer. Hematol Oncol Clin North Am. 2012; 26:1–12.
25. Lupia M, Cavallaro U. Ovarian cancer stem cells: still an elusive entity? Mol Cancer. 2017; 16:64.
26. Mueller MT, Hermann PC, Witthauer J, Rubio-Viqueira B, Leicht SF, Huber S, Ellwart JW, Mustafa M, Bartenstein P, D’Haese JG, Schoenberg MH, Berger F, Jauch KW, et al. Combined targeted treatment to eliminate tumorigenic cancer stem cells in human pancreatic cancer. Gastroenterology. 2009; 137:1102–1113.
27. Dylla SJ, Beviglia L, Park IK, Chartier C, Raval J, Ngan L, Pickell K, Aguilar J, Lazetic S, Smith-Berdan S, Clarke MF, Hoey T, Lewicki J, et al. Colorectal cancer stem cells are enriched in xenogeneic tumors following chemotherapy. PLoS One. 2008; 3:e2428.
28. Ma S, Lee TK, Zheng BJ, Chan KW, Guan XY. CD133+ HCC cancer stem cells confer chemoresistance by preferential expression of the Akt/PKB survival pathway. Oncogene. 2008; 27:1749–1758.
29. Levina V, Marrangoni AM, DeMarco R, Gorelik E, Lokshin AE. Drug-selected human lung cancer stem cells: cytokine network, tumorigenic and metastatic properties. PLoS One. 2008; 3:e3077.
30. Tomao F, Papa A, Strudel M, Rossi L, Lo Russo G, Benedetti Panici P, Ciabatta FR, Tomao S. Investigating molecular profiles of ovarian cancer: an update on cancer stem cells. J Cancer. 2014; 5:301–310.
31. Boesch M, Zeimet AG, Reimer D, Schmidt S, Gastl G, Parson W, Spoeck F, Hatina J, Wolf D, Sopper S. The side population of ovarian cancer cells defines a heterogeneous compartment exhibiting stem cell characteristics. Oncotarget. 2014; 5:7027–7039. https://doi.org/10.18632/oncotarget.2053.
32. Gao Y, Foster R, Yang X, Feng Y, Shen JK, Mankin HJ, Hornicek FJ, Amiji MM, Duan Z. Up-regulation of CD44 in the development of metastasis, recurrence and drug resistance of ovarian cancer. Oncotarget. 2015; 6:9313–9326. https://doi.org/10.18632/oncotarget.3220.
33. Meng E, Long B, Sullivan P, McClellan S, Finan MA, Reed E, Shevde L, Rocconi RP. CD44+/CD24- ovarian cancer cells demonstrate cancer stem cell properties and correlate to survival. Clin Exp Metastasis. 2012; 29:939–948.
34. Stemberger-Papic S, Vrdoljak-Mozetic D, Ostojic DV, Rubesa-Mihaljevic R, Krigtofic I, Brncic-Fisher A, Kragevic M, Eminovic S. Expression of CD133 and CD117 in 64 serous ovarian cancer cases. Coll Antropol. 2015; 39:745–753.
35. Zhang J, Guo X, Chang DY, Rosen DG, Mercado-Uribe I, Liu J. CD133 expression associated with poor prognosis in ovarian cancer. Mod Pathol. 2012; 25:456–464.
36. Ayub TH, Keyver-Paik MD, Debald M, Rostamzadeh B, Thiesler T, Schroder L, Barchet W, Abramian A, Kaiser C, Kristiansen G, Kuhn W, Kubler K. Accumulation of ALDH1-positive cells after neoadjuvant chemotherapy predicts treatment resistance and prognosticates poor outcome in ovarian cancer. Oncotarget. 2015; 6:16437–16448. https://doi.org/10.18632/oncotarget.4103.
37. Sun Y, Jia X, Wu X. High expressions of Lgr5 and ALDH1 in primary epithelial ovarian cancer correlate with advanced tumor stage and grade as well as poor prognosis of the patients. Gynecol Obstet Invest. 2016; 81:162–168.
38. Landen CN Jr, Goodman B, Katre AA, Steg AD, Nick AM, Stone RL, Miller LD, Mejia PV, Jennings NB, Gershenson DM, Bast RC Jr, Coleman RL, Lopez-Berestein G, Sood AK. Targeting aldehyde dehydrogenase cancer stem cells in ovarian cancer. Mol Cancer Ther. 2010; 9:3186–3199.
39. Choi YJ, Ingram PN, Yang K, Coffman L, Iyengar M, Bai S, Thomas DG, Yoon E, Buckanovich RJ. Identifying an ovarian cancer cell hierarchy regulated by bone morphogenetic protein 2. Proc Natl Acad Sci USA. 2015; 112:E6882–E6888.
40. Ishiguro T, Sato A, Ohata H, Ikarashi Y, Takahashi RU, Ochiya T, Yoshida M, Tsuda H, Onda T, Kato T, Kasamatsu T, Enomoto T, Tanaka K, et al. Establishment and characterization of an in vitro model of ovarian cancer stem-like cells with an enhanced proliferative capacity. Cancer Res. 2016; 76:150–160.
41. Tomita H, Tanaka K, Tanaka T, Hara A. Aldehyde dehydrogenase 1A1 in stem cells and cancer. Oncotarget. 2016; 7:11018–11032. https://doi.org/10.18632/oncotarget.6920.
42. Alison MR, Lim SM, Nicholson LJ. Cancer stem cells: Problems for therapy? J Pathol. 2011; 223:147–161.
43. Wang Y, Cardenas H, Fang F, Condello S, Taverna P, Segar M, Liu Y, Nephew KP, Matei D. Epigenetic targeting of ovarian cancer stem cells. Cancer Res. 2014; 74:4922–4936.
44. Sharrow AC, Perkins B, Collector MI, Yu W, Simons BW, Jones RJ. Characterization of aldehyde dehydrogenase 1 high ovarian cancer cells: Towards targeted stem cell therapy. Gynecol Oncol. 2016; 142:341–348.
45. Liao J, Qian F, Tchabo N, Mhawech-Fauceglia P, Beck A, Qian Z, Wang X, Huss WJ, Lele SB, Morrison CD, Odunsi K. Ovarian cancer spheroid cells with stem cell-like properties contribute to tumor generation, metastasis and chemotherapy resistance through hypoxia-resistant metabolism. PLoS One. 2014; 9:e84941.
46. Charafe-Jauffret E, Ginestier C, Iovino F, Wicinski J, Cervera N, Finetti P, Hur MH, Diebel ME, Monville F, Dutcher J, Brown M, Viens P, Xerri L, et al. Breast cancer cell lines contain functional cancer stem cells with metastatic capacity and a distinct molecular signature. Cancer Res. 2009; 69:1302–1313.
47. Januchowski R, Wojtowicz K, Sterzynska K, Sosinska P, Andrzejewska M, Zawierucha P, Nowicki M, Zabel M. Inhibition of ALDH1A1 activity decreases expression of drug transporters and reduces chemotherapy resistance in ovarian cancer cell lines. Int J Biochem Cell Biol. 2016; 78:248–259.
48. Zhang S, Cui B, Lai H, Liu G, Ghia EM, Widhopf GF 2nd, Zhang Z, Wu CC, Chen L, Wu R, Schwab R, Carson DA, Kipps TJ. Ovarian cancer stem cells express ROR1, which can be targeted for anti-cancer-stem-cell therapy. Proc Natl Acad Sci USA. 2014; 111:17266–17271.
49. Penumatsa K, Edassery SL, Barua A, Bradaric MJ, Luborsky JL. Differential expression of aldehyde dehydrogenase 1a1 (ALDH1) in normal ovary and serous ovarian tumors. J Ovarian Res. 2010; 3:28.
50. Deng S, Yang X, Lassus H, Liang S, Kaur S, Ye Q, Li C, Wang LP, Roby KF, Orsulic S, Connolly DC, Zhang Y, Montone K, et al. Distinct expression levels and patterns of stem cell marker, aldehyde dehydrogenase isoform 1 (ALDH1), in human epithelial cancers. PLoS One. 2010; 5:e10277.
51. Yoshida A, Rzhetsky A, Hsu LC, Chang C. Human aldehyde dehydrogenase gene family. Eur J Biochem. 1998; 251:549–557.
52. Condello S, Morgan CA, Nagdas S, Cao L, Turek J, Hurley TD, Matei D. beta-Catenin-regulated ALDH1A1 is a target in ovarian cancer spheroids. Oncogene. 2015; 34:2297–2308.
53. Meng E, Mitra A, Tripathi K, Finan MA, Scalici J, McClellan S, Madeira da Silva L, Reed E, Shevde LA, Palle K, Rocconi RP. ALDH1A1 maintains ovarian cancer stem cell-like properties by altered regulation of cell cycle checkpoint and DNA repair network signaling. PLoS One. 2014; 9:e107142.
54. Jackson B, Brocker C, Thompson DC, Black W, Vasiliou K, Nebert DW, Vasiliou V. Update on the aldehyde dehydrogenase gene (ALDH) superfamily. Hum Genomics. 2011; 5:283–303.
55. Chen CH, Ferreira JC, Gross ER, Mochly-Rosen D. Targeting aldehyde dehydrogenase 2: new therapeutic opportunities. Physiol Rev. 2014; 94:1–34.
56. Eriksson CJ. Acetaldehyde metabolism in vivo during ethanol oxidation. Adv Exp Med Biol. 1977; 85A:319–341.
57. Takao A, Shimoda T, Kohno S, Asai S, Harda S. Correlation between alcohol-induced asthma and acetaldehyde dehydrogenase-2 genotype. J Allergy Clin Immunol. 1998; 101:576–580.
58. Wu G. Application of cross-impact analysis to the relationship between aldehyde dehydrogenase 2 and flushing. Alcohol Alcohol. 2000; 35:55–59.
59. Wu G. Application of queueing theory with Monte Carlo simulation to the study of the intake and adverse effects of ethanol. Alcohol Alcohol. 1998; 33:519–527.
60. Wu G. Frequency and Markov chain analysis of the amino acid sequence of human alcohol dehydrogenase a-chain. Alcohol Alcohol. 2000; 35:302–306.
61. Brooks PJ, Enoch MA, Goldman D, Li TK, Yokoyama A. The alcohol flushing response: an unrecognized risk factor for esophageal cancer from alcohol consumption. PLoS Med. 2009; 6:e50.
62. Wu G. Use of a five-compartment closed model to describe the effects of ethanol inhalation on the transport and elimination of injected pyruvate in the rat. Alcohol Alcohol. 1997; 32:555–561.
63. Klyosov AA. Kinetics and specificity of human liver aldehyde dehydrogenases toward aliphatic, aromatic, and fused polycyclic aldehydes. Biochemistry. 1996; 35:4457–4467.
64. Yoval-Sanchez B, Rodriguez-Zavala JS. Differences in susceptibility to inactivation of human aldehyde dehydrogenases by lipid peroxidation byproducts. Chem Res Toxicol. 2012; 25:722–729.
65. Chen CH, Sun L, Mochly-Rosen D. Mitochondrial aldehyde dehydrogenase and cardiac diseases. Cardiovasc Res. 2010; 88:51–57.
66. Bian Y, Chen YG, Xu F, Xue L, Ji WQ, Zhang Y. The polymorphism in aldehyde dehydrogenase-2 gene is associated with elevated plasma levels of high-sensitivity C-reactive protein in the early phase of myocardial infarction. Tohoku J Exp Med. 2010; 221:107–112.
67. Radovanovic S, Savic-Radojevic A, Pljesa-Ercegovac M, Djukic T, Suvakov S, Krotin M, Simic DV, Matic M, Radojicic Z, Pekmezovic T, Simic T. Markers of oxidative damage and antioxidant enzyme activities as predictors of morbidity and mortality in patients with chronic heart failure. J Cardiac Failure. 2012; 18:493–501.
68. Chen CH, Budas GR, Churchill EN, Disatnik MH, Hurley TD, Mochly-Rosen D. Activation of aldehyde dehydrogenase-2 reduces ischemic damage to the heart. Science. 2008; 321:1493–1495.
69. Chang YC, Chiu YF, Lee IT, Ho LT, Hung YJ, Hsiung CA, Quertermous T, Donlon T, Lee WJ, Lee PC, Chen CH, Mochly-Rosen D, Chuang LM. Common ALDH2 genetic variants predict development of hypertension in the SAPPHIRe prospective cohort: gene-environmental interaction with alcohol consumption. BMC Cardiovasc Disorders. 2012; 12:58.
70. Kato N, Takeuchi F, Tabara Y, Kelly TN, Go MJ, Sim X, Tay WT, Chen CH, Zhang Y, Yamamoto K, Katsuya T, Yokota M, Kim YJ, et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in east Asians. Nat Genet. 2011; 43:531–538.
71. Suzuki Y, Muramatsu T, Taniyama M, Atsumi Y, Kawaguchi R, Higuchi S, Hosokawa K, Asahina T, Murata C, Matsuoka K. Association of aldehyde dehydrogenase with inheritance of NIDDM. Diabetologia. 1996; 39:1115–1118.
72. Murata C, Taniyama M, Kuriyama S, Muramatsu T, Atsumi Y, Matsuoka K, Suzuki Y. Meta-analysis of three diabetes population studies: association of inactive ALDH2 genotype with maternal inheritance of diabetes. Diabetes Res Clin Pract. 2004; 66:S145–S147.
73. Wang B, Wang J, Zhou S, Tan S, He X, Yang Z, Xie YC, Li S, Zheng C, Ma X. The association of mitochondrial aldehyde dehydrogenase gene (ALDH2) polymorphism with susceptibility to late-onset Alzheimer’s disease in Chinese. J Neurol Sci. 2008; 268:172–175.
74. Kamino K, Nagasaka K, Imagawa M, Yamamoto H, Yoneda H, Ueki A, Kitamura S, Namekata K, Miki T, Ohta S. Deficiency in mitochondrial aldehyde dehydrogenase increases the risk for late-onset Alzheimer’s disease in the Japanese population. Biochem Biophys Res Commun. 2000; 273:192–196.
75. Hao PP, Chen YG, Wang JL, Wang XL, Zhang Y. Meta-analysis of aldehyde dehydrogenase 2 gene polymorphism and Alzheimer’s disease in East Asians. Can J Neurol Sci. 2011; 38:500–506.
76. Yokoyama A, Muramatsu T, Ohmori T, Higuchi S, Hayashida M, Ishii H. Esophageal cancer and aldehyde dehydrogenase-2 genotypes in Japanese males. Cancer Epidemiol Biomarkers Prev. 1996; 5:99–102.
77. Goedde HW, Agarwal DP, Fritze G, Meier-Tackmann D, Singh S, Beckmann G, Bhatia K, Chen LZ, Fang B, Lisker R. Distribution of ADH2 and ALDH2 genotypes in different populations. Hum Genet. 1992; 88:344–346.
78. Li H, Borinskaya S, Yoshimura K, Kal’ina N, Marusin A, Stepanov VA, Qin Z, Khaliq S, Lee MY, Yang Y, Mohyuddin A, Gurwitz D, Mehdi SQ, et al. Refined geographic distribution of the oriental ALDH2*504Lys (nee 487Lys) variant. Ann Hum Genet. 2009; 73:335–345.
79. Klyosov AA, Rashkovetsky LG, Tahir MK, Keung WM. Possible role of liver cytosolic and mitochondrial aldehyde dehydrogenases in acetaldehyde metabolism. Biochemistry. 1996; 35:4445–4456.
80. Yokoyama A, Omori T. Genetic polymorphisms of alcohol and aldehyde dehydrogenases and risk for esophageal and head and neck cancers. Jpn J Clin Oncol. 2003; 33:111–121.
81. Yokoyama A, Omori T. Genetic polymorphisms of alcohol and aldehyde dehydrogenases and risk for esophageal and head and neck cancers. Alcohol. 2005; 35:175–185.
82. Yin SJ, Liao CS, Lee YC, Wu CW, Jao SW. Genetic polymorphism and activities of human colon alcohol and aldehyde dehydrogenases: no gender and age differences. Alcohol Clin Exp Res. 1994; 18:1256–1260.
83. Muto M, Takahashi M, Ohtsu A, Ebihara S, Yoshida S, Esumi H. Risk of multiple squamous cell carcinomas both in the esophagus and the head and neck region. Carcinogenesis. 2005; 26:1008–1012.
84. Lee CH, Wu DC, Wu IC, Goan YG, Lee JM, Chou SH, Chan TF, Huang HL, Hung YH, Huang MC, Lai TC, Wang TN, Lan CC, et al. Genetic modulation of ADH1B and ALDH2 polymorphisms with regard to alcohol and tobacco consumption for younger aged esophageal squamous cell carcinoma diagnosis. Int J Cancer. 2009; 125:1134–1142.
85. Ding JH, Li SP, Cao HX, Wu JZ, Gao CM, Su P, Liu YT, Zhou JN, Chang J, Yao GH. Polymorphisms of alcohol dehydrogenase-2 and aldehyde dehydrogenase-2 and esophageal cancer risk in Southeast Chinese males. World J Gastroenterol. 2009; 15:2395–2400.
86. Asakage T, Yokoyama A, Haneda T, Yamazaki M, Muto M, Yokoyama T, Kato H, Igaki H, Tsujinaka T, Kumagai Y, Yokoyama M, Omori T, Watanabe H. Genetic polymorphisms of alcohol and aldehyde dehydrogenases, and drinking, smoking and diet in Japanese men with oral and pharyngeal squamous cell carcinoma. Carcinogenesis. 2007; 28:865–874.
87. Ran D, Schubert M, Pietsch L, Taubert I, Wuchter P, Eckstein V, Bruckner T, Zoeller M, Ho AD. Aldehyde dehydrogenase activity among primary leukemia cells is associated with stem cell features and correlates with adverse clinical outcomes. Exp Hematol. 2009; 37:1423–1434.
88. Cheung AM, Wan TS, Leung JC, Chan LY, Huang H, Kwong YL, Liang R, Leung AY. Aldehyde dehydrogenase activity in leukemic blasts defines a subgroup of acute myeloid leukemia with adverse prognosis and superior NOD/SCID engrafting potential. Leukemia. 2007; 21:1423–1430.
89. Charafe-Jauffret E, Ginestier C, Iovino F, Tarpin C, Diebel M, Esterni B, Houvenaeghel G, Extra JM, Bertucci F, Jacquemier J, Xerri L, Dontu G, Stassi G, et al. Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin Cancer Res. 2010; 16:45–55.
90. Tanei T, Morimoto K, Shimazu K, Kim SJ, Tanji Y, Taguchi T, Tamaki Y, Noguchi S. Association of breast cancer stem cells identified by aldehyde dehydrogenase 1 expression with resistance to sequential Paclitaxel and epirubicin-based chemotherapy for breast cancers. Clin Cancer Res. 2009; 15:4234–4241.
91. Morimoto K, Kim SJ, Tanei T, Shimazu K, Tanji Y, Taguchi T, Tamaki Y, Terada N, Noguchi S. Stem cell marker aldehyde dehydrogenase 1-positive breast cancers are characterized by negative estrogen receptor, positive human epidermal growth factor receptor type 2, and high Ki67 expression. Cancer Sci. 2009; 100:1062–1068.
92. Jiang F, Qiu Q, Khanna A, Todd NW, Deepak J, Xing L, Wang H, Liu Z, Su Y, Stass SA, Katz RL. Aldehyde dehydrogenase 1 is a tumor stem cell-associated marker in lung cancer. Mol Cancer Res. 2009; 7:330–338.
93. Chen YC, Chen YW, Hsu HS, Tseng LM, Huang PI, Lu KH, Chen DT, Tai LK, Yung MC, Chang SC, Ku HH, Chiou SH, Lo WL. Aldehyde dehydrogenase 1 is a putative marker for cancer stem cells in head and neck squamous cancer. Biochem Biophys Res Commun. 2009; 385:307–313.
94. Rasheed ZA, Yang J, Wang Q, Kowalski J, Freed I, Murter C, Hong SM, Koorstra JB, Rajeshkumar NV, He X, Goggins M, Iacobuzio-Donahue C, Berman DM, et al. Prognostic significance of tumorigenic cells with mesenchymal features in pancreatic adenocarcinoma. J Natl Cancer Inst. 2010; 102:340–351.
95. Li T, Su Y, Mei Y, Leng Q, Leng B, Liu Z, Stass SA, Jiang F. ALDH1A1 is a marker for malignant prostate stem cells and predictor of prostate cancer patients’ outcome. Lab Invest. 2010; 90:234–244.
96. Yokoyama A, Muramatsu T, Ohmori T, Yokoyama T, Okuyama K, Takahashi H, Hasegawa Y, Higuchi S, Maruyama K, Shirakura K, Ishii H. Alcohol-related cancers and aldehyde dehydrogenase-2 in Japanese alcoholics. Carcinogenesis. 1998; 19:1383–1387.
97. Fujii C, Harada S, Ohkoshi N, Hayashi A, Yoshizawa K. Study on Parkinson’s disease and alcohol drinking. Nihon Arukoru Yakubutsu Igakkai Zasshi. 1998; 33:683–691.
98. Yao CT, Cheng CA, Wang HK, Chiu SW, Chen YC, Wang MF, Yin SJ, Peng GS. The role of ALDH2 and ADH1B polymorphism in alcohol consumption and stroke in Han Chinese. Hum genomics. 2011; 5:569–576.
99. Nagasawa H, Wada M, Arawaka S, Kawanami T, Kurita K, Daimon M, Adachi M, Hosoya T, Emi M, Muramatsu M, Kato T. A polymorphism of the aldehyde dehydrogenase 2 gene is a risk factor for multiple lacunar infarcts in Japanese men: the Takahata Study. Eur J Neurol. 2007; 14:428–434.
100. Xu F, Chen YG, Xue L, Li RJ, Zhang H, Bian Y, Zhang C, Lv RJ, Feng JB, Zhang Y. Role of aldehyde dehydrogenase 2 Glu504lys polymorphism in acute coronary syndrome. J Cell Mol Med. 2011; 15:1955–1962.
101. Takagi S, Iwai N, Yamauchi R, Kojima S, Yasuno S, Baba T, Terashima M, Tsutsumi Y, Suzuki S, Morii I, Hanai S, Ono K, Baba S, et al. Aldehyde dehydrogenase 2 gene is a risk factor for myocardial infarction in Japanese men. Hypertens Res. 2002; 25:677–681.
102. Xu F, Chen Y, Lv R, Zhang H, Tian H, Bian Y, Feng J, Sun Y, Li R, Wang R, Zhang Y. ALDH2 genetic polymorphism and the risk of type II diabetes mellitus in CAD patients. Hypertension Res. 2010; 33:49–55.
103. Lagranha CJ, Deschamps A, Aponte A, Steenbergen C, Murphy E. Sex differences in the phosphorylation of mitochondrial proteins result in reduced production of reactive oxygen species and cardioprotection in females. Circ Res. 2012; 106:1681–1691.
104. Ohsawa I, Kamino K, Nagasaka K, Ando F, Niino N, Shimokata H, Ohta S. Genetic deficiency of a mitochondrial aldehyde dehydrogenase increases serum lipid peroxides in community-dwelling females. J Hum Genet. 2003; 48:404–409.
105. Luo HR, Wu GS, Pakstis AJ, Tong L, Oota H, Kidd KK, Zhang YP. Origin and dispersal of atypical aldehyde dehydrogenase ALDH2487Lys. Gene. 2009; 435:96–103.
106. Lin YP, Cheng TJ. Why can’t Chinese Han drink alcohol? Hepatitis B virus infection and the evolution of acetaldehyde dehydrogenase deficiency. Med Hypotheses. 2002; 59:204–207.
107. Goldman D, Enoch MA. Genetic epidemiology of ethanol metabolic enzymes: a role for selection. World Rev Nutr Diet. 1990; 63:143–160.
108. Yokoyama A, Muramatsu T, Ohmori T, Makuuchi H, Higuchi S, Matsushita S, Yoshino K, Maruyama K, Nakano M, Ishii H. Multiple primary esophageal and concurrent upper aerodigestive tract cancer and the aldehyde dehydrogenase-2 genotype of Japanese alcoholics. Cancer. 1996; 77:1986–1990.
109. Sinclair D. Deleterious pleiotropic effects of the atypical aldehyde dehydrogenase 2 (ALDH2) allele: comment on Luo et al., 2005. Biochem Genet. 2006; 44:385–390.
110. Luo HR, Israel Y, Tu GC, Eriksson CJ, Zhang YP. Genetic polymorphism of aldehyde dehydrogenase 2 (ALDH2) in a Chinese population: gender, age, culture, and genotypes of ALDH2. Biochem Genet. 2005; 43:223–227.
111. Park JW, Ji YI, Choi YH, Kang MY, Jung E, Cho SY, Cho HY, Kang BK, Joung YS, Kim DH, Park SC, Park J. Candidate gene polymorphisms for diabetes mellitus, cardiovascular disease and cancer are associated with longevity in Koreans. Exp Mol Med. 2009; 41:772–781.
112. Yagasaki H, Oda T, Adachi D, Nakajima T, Nakahata T, Asano S, Yamashita T. Two common founder mutations of the fanconi anemia group G gene FANCG/XRCC9 in the Japanese population. Hum Mutat. 2003; 21:555.
113. Park J, Chung NG, Chae H, Kim M, Lee S, Kim Y, Lee JW, Cho B, Jeong D, Park I. FANCA and FANCG are the major Fanconi anemia genes in the Korean population. Clin Genet. 2013; 84:271–275.
114. Endo J, Sano M, Katayama T, Hishiki T, Shinmura K, Morizane S, Matsuhashi T, Katsumata Y, Zhang Y, Ito H, Nagahata Y, Marchitti S, Nishimaki K, et al. Metabolic remodeling induced by mitochondrial aldehyde stress stimulates tolerance to oxidative stress in the heart. Circ Res. 2009; 105:1118–1127.
115. Hjartåker A, Meo MS, Weiderpass E. Alcohol and gynecological cancers: an overview. Eur J Cancer Prev. 2010; 19:1–10.
116. Gasparovic AC, Milkovic L, Sunjic SB, Zarkovic N. Cancer growth regulation by 4-hydroxynonenal. Free Radic Biol Med. 2017; 111:226–234.
117. Rausch V, Liu L, Kallifatidis G, Baumann B, Mattern J, Gladkich J, Wirth T, Schemmer P, Büchler MW, Zöller M, Salnikov AV, Herr I. Synergistic activity of sorafenib and sulforaphane abolishes pancreatic cancer stem cell characteristics. Cancer Res. 2010; 70:5004–5013.
118. Kallifatidis G, Rausch V, Baumann B, Apel A, Beckermann BM, Groth A, Mattern J, Li Z, Kolb A, Moldenhauer G, Altevogt P, Wirth T, Werner J, et al. Sulforaphane targets pancreatic tumour-initiating cells by NF-kappaB-induced antiapoptotic signalling. Gut. 2009; 58:949–963.
119. Cioce M, Gherardi S, Viglietto G, Strano S, Blandino G, Muti P, Ciliberto G. Mammosphere-forming cells from breast cancer cell lines as a tool for the identification of CSC-like- and early progenitor-targeting drugs. Cell Cycle. 2010; 9:2878–2887.
120. Li Y, Zhang T, Korkaya H, Liu S, Lee HF, Newman B, Yu Y, Clouthier SG, Schwartz SJ, Wicha MS, Sun D. Sulforaphane, a dietary component of broccoli/broccoli sprouts, inhibits breast cancer stem cells. Clin Cancer Res. 2010; 16:2580–2590.
121. Kakarala M, Brenner DE, Korkaya H, Cheng C, Tazi K, Ginestier C, Liu S, Dontu G, Wicha MS. Targeting breast stem cells with the cancer preventive compounds curcumin and piperine. Breast Cancer Res Treat. 2010; 122:777–785.
122. Aldefluor, Stem Cell Technologies, Durham, NC, USA.
123. Duhagon MA, Hurt EM, Sotelo-Silveira JR, Zhang X, Farrar WL. Genomic profiling of tumor initiating prostatospheres. BMC Genomics. 2010; 11:324.
124. Schulte A, Günther HS, Phillips HS, Kemming D, Martens T, Kharbanda S, Soriano RH, Modrusan Z, Zapf S, Westphal M, Lamszus K. A distinct subset of glioma cell lines with stem cell-like properties reflects the transcriptional phenotype of glioblastomas and overexpresses CXCR4 as therapeutic target. Glia. 2011; 59:590–602.
125. Günther HS, Schmidt NO, Phillips HS, Kemming D, Kharbanda S, Soriano R, Modrusan Z, Meissner H, Westphal M, Lamszus K. Glioblastoma-derived stem cell-enriched cultures form distinct subgroups according to molecular and phenotypic criteria. Oncogene. 2008; 27:2897–2909.
126. Zamykal M, Martens T, Matschke J, Günther HS, Kathagen A, Schulte A, Peters R, Westphal M, Lamszus K. Inhibition of intracerebral glioblastoma growth by targeting the insulin-like growth factor 1 receptor involves different context-dependent mechanisms. Neuro Oncol. 2015; 17:1076–1085.
127. Wang L, Mezencev R, Bowen NJ, Matyunina LV, McDonald JF. Isolation and characterization of stem-like cells from a human ovarian cancer cell line. Mol Cell Biochem. 2012; 363:257–268.
128. Yu YH, Chiou GY, Huang PI, Lo WL, Wang CY, Lu KH, Yu CC, Alterovitz G, Huang WC, Lo JF, Hsu HS, Chiou SH. Network biology of tumor stem-like cells identified a regulatory role of CBX5 in lung cancer. Sci Rep. 2012; 2:584.
129. Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002; 30:207–210.
130. Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Holko M, Yefanov A, Lee H, Zhang N, et al. NCBI GEO: archive for functional genomics data sets--update. Nucleic Acids Res. 2013; 41:D991–D995.
131. http://www.affymetrix.com/support/technical/byproduct.affx?product=hg-u133-plus.

