INTRODUCTION
Cancer is one of the ever dreadful disease, and a leading cause of death, and a severe public health problem worldwide [1]. Various projection based studies reported that the worldwide burden of cancer will rise more rapidly with rising population, aging and changes in the lifestyle [2]. It is predicted that there will be more than 20 million new cancer cases globally by the year 2025 [3]. The increasing incidence and mortality rate due to cancer during the last two decades have posed a big challenge to the clinicians and scientists. Also, the precise mechanism of carcinogenesis has not been fully deciphered yet. In the recent years, the growing number of studies reported that the initiation of cancer is a complex process that includes the results of genetic susceptibility and various environmental factors [4]. Therefore, the identification of genetic risk factors that contribute to the substantial burden of the disease in general population is warranted for the development of broad therapeutic strategies for cancer prevention.
Recent genomewide association studies (GWAS), and other case-control studies have revealed that single-nucleotide polymorphisms (SNPs) are the most common forms of human genetic variations, have important role in defining susceptibility to cancer [5, 6]. This clearly suggests the SNPs can be used as a promising biomarker for the evaluation of individual genetic background for cancer prognosis, and signifies an interesting field of cancer research.
Major histocompatibility complex (MHC) is a set of cell surface glycoproteins that bind with the intracellularly processed peptides from pathogens, and present them on the surface of tumor cells to cytotoxic T lymphocytes (CTL). Therefore, MHC plays a key role in the initiation of CTL linked inflammation and anticancer immune response [7]. MHC restricted immune response has been tested to eliminate cancer cells, and the expression of MHC antigens may be important for the host immune response against cancer [8].
The low molecular weight polypeptide 7 (LMP7, also called as PSMB8) gene is located in the class II region of the MHC locus on chromosome 6 [9]. This gene encodes peptide forming the large components of the proteasome complex engaged in the degradation of cytosolic proteins and the generation of antigenic peptides [10, 11]. The components of the proteasome are thought to participate in the proteolytic digestion of cancer-relevant antigen peptides derived from endogenous or exogenous antigenic proteins, and play central role in homoeostasis of cellular proteins and regulation of cellular processes that are important in cancer initiation and progression [12].
Single-nucleotide polymorphism (SNPs) is one of the most common forms of genetic alterations. Several studies from TCGA and COSMIC were perused to analyze the association between the LMP7 gene mutations and risk of developing cancer including human breast and colorectal cancers [13], human glioblastoma multiforme [14], pancreatic cancer [15], melanoma [16], and human colon and rectum cancer [17]. Various polymorphic residues have also been identified in LMP7 gene [18]. Previous studies have reported that LMP7–145 (Gln > Lys, C > A, rs2071543) gene polymorphism result in functional alternation, and ultimately deteriorate the capacity of antigen processing [19]. Thus, an abnormal expression of low molecular peptide (LMP) subunits attributes to many disease phenotypes and malignant tumors.
In view of the crucial restrictive role of LMP7 gene in antigen processing and presentation, this gene is a promising candidate for carcinogenesis susceptibility. In the last few years, a number of case-control studies have investigated the impact of this gene polymorphism on cancer risk in various populations; but the reported results varied across studies and remain inconclusive [20–27]. Inconsistency in the findings of previous studies could be possibly attributed to small sample size, and low statistical power. Lately, Burton et al. (2009) reported that large sample size is always good, and necessary to study the genetic associations with complex diseases [28]. Therefore, in this study, a meta-analysis was performed by pooling all the eligible published studies to determine the more precise association and understanding the possible role of LMP7–145 (C > A) gene polymorphism as genetic marker for cancer progression. Further, the quality of the included studies was checked by performing Newcastle Ottawa Scale (NOS) analysis. Also, type-I statistical errors, i.e., publication bias and random errors occurred due to sparse data were minimized by performing Trial Sequential Analysis (TSA) for quantifying the statistical reliability of the data included in the cumulative meta-analysis with the threshold of statistical significance. Overall, a meta-analysis is a powerful statistical tool for analyzing cumulative data from different studies, in which the individual sample sizes are small and the statistical power is low, and it provides precise and robust conclusion [29].
RESULTS
Literature search and meta-analysis databases
The confounding conclusions from previous studies regarding the role of LMP7–145 (C > A) gene polymorphism as genetic marker for cancer susceptibility and progression [20–27] impelled us to perform their meta-analysis in order to understand the precise association between this polymorphism and cancer risk. For meta-analysis a literature search following stringent criteria, as stated in the methodology section, was adopted. A total of eight studies regarding LMP7 –145 (C > A) gene polymorphism and cancer risk were found eligible for inclusion in this meta-analysis. All the retrieved articles were examined carefully by reading their titles and abstracts, and the full texts of the potentially germane publications were further reviewed for their appropriateness for this meta-analysis. Published studies either showing LMP polymorphism to predict survival in cancer patients or considering LMP variants as an indicators for response-to-therapy were disqualified straightaway. Likewise, studies related to the investigation of the levels of LMP mRNA or protein expression or pertinent review articles were also disqualified. In this meta-analysis, only case-control or cohort design clinical studies having frequency of all the three genotypes were included. In addition to the online database search, the references listed in the primary retrieved articles were also examined for other relevant potential articles. The major characteristics of all the eight studies included in this meta-analysis, i.e., distribution of genotypes, minor allele frequency (MAF) in controls and cases have been given in Table 1 and Table 2. All the eight studies included in this meta-analysis were appraised for the quality score according to the NOS analysis, and almost all the studies (< 80%) scored 5 stars or more, and suggested a moderate to good quality (Table 3). The needful information related to the selection and inclusion of the studies for this meta-analysis has been given in Figure 1 (PRISMA 2009 Flow Diagram).
Table 1: Main characteristics of all the 8 studies included in the present meta-analysis
First author, (Year) Ref. No. |
Country |
Ethnicity |
Type of cancer |
Type of study |
Controls |
Cases |
Methods |
Association |
|---|---|---|---|---|---|---|---|---|
Ma et al. (2015) [20] |
China |
Asian |
Gastric |
HB |
502 |
502 |
PCR-RFLP |
Yes |
Mehta et al. (2015) [21] |
Indonesia |
Asian |
Cervical |
PB |
173 |
201 |
TaqMan SNP Genotyping Assay |
No |
Song et al. (2014) [22] |
China |
Asian |
Ovarian |
HB |
338 |
235 |
PCR-RFLP |
Yes |
Ozbas et al. (2013) [23] |
Turkey |
Asian |
Hematological malignancy |
HB |
130 |
132 |
PCR-RFLP |
Yes |
Fellerhoff et al. (2011) [24] |
Germany |
Caucasian |
Colon |
HB |
165 |
174 |
ARMS-PCR |
Yes |
Deshpande et al. (2008) [25] |
USA |
Caucasian |
Cervical |
PB |
224 |
134 |
Sequencing |
No |
Mehta et al. (2007) [26] |
Netherlands |
Caucasian |
Cervical |
HB |
124 |
127 |
TaqMan SNP |
Yes |
Cao et al. (2005) [27] |
China |
Asian |
Esophageal |
HB |
357 |
265 |
Sequencing |
Yes |
HB: Hospital base; PB: Population base.
Table 2: Genotypic distribution of LMP7 -145 C > A gene polymorphism included in this meta-analysis
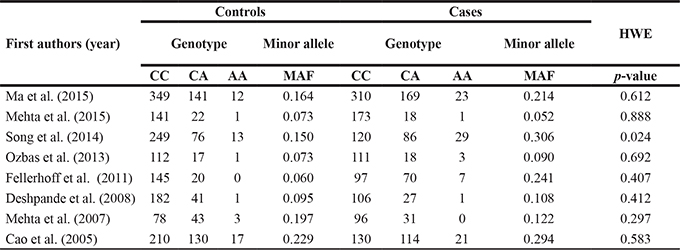
MAF: Minor allele frequency, HWE: Hardy weinberg equilibrium.
Table 3: Quality assessment for all the studies included in this meta-analysis according to the Newcastle-Ottawa scale
First author (Year) |
Quality indicators |
||
|---|---|---|---|
Selection |
Comparability |
Exposure |
|
Ma et al. (2015) |
**** |
* |
** |
Mehta et al. (2015) |
** |
* |
** |
Song et al. (2014) |
*** |
* |
** |
Ozbas et al. (2013) |
*** |
* |
** |
Fellerhoff et al. (2011) |
*** |
* |
** |
Deshpande et al. (2008) |
**** |
* |
** |
Mehta et al. (2007) |
*** |
* |
** |
Cao et al. (2005) |
**** |
* |
** |
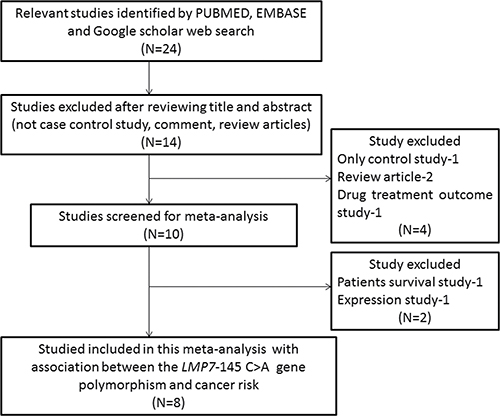
Figure 1: PRISMA (preferred reporting items for systematic reviews and meta-analyses) flow-diagram outlining the identification and selection of studies for inclusion/exclusion of the relevant studies in this meta-analysis.
Detection of publication bias
The Begg’s funnel plot and Egger’s test were performed to appraise the publication bias among the clinical studies of LMP7 –145 (C > A) gene polymorphism included in this meta-analysis. No evidence of publication bias was observed by the appearance of the shape of funnel plots and the results of Egger’s test in all the five genetic models of LMP7 –145 (C > A) (Table 4, Supplementary Information: Supplementary Figure 1) gene polymorphism.
Table 4: Statistics to test publication bias and heterogeneity of LMP7 -145 C > A gene polymorphism
Comparisons |
Egger’s regression analysis |
Heterogeneity analysis |
Model used for the meta-analysis |
|||||
|---|---|---|---|---|---|---|---|---|
Intercept |
95% Confidence Interval |
p-value |
Q-value |
Pheterogeneity |
I2 (%) |
|||
Overall risk |
||||||||
A vs C |
−1.27 |
−8.25 to 5.70 |
0.67 |
55.96 |
0.01 |
87.49 |
Random |
|
AA vs CC |
−0.47 |
−2.48 to 1.52 |
0.57 |
10.64 |
0.15 |
34.26 |
Fixed |
|
AC vs CC |
−1.16 |
−8.29 to 5.95 |
0.70 |
43.99 |
0.01 |
84.09 |
Random |
|
AA + AC vs CC |
−1.42 |
−9.07 to 6.22 |
0.66 |
53.57 |
0.01 |
86.93 |
Random |
|
AA vs AC + CC |
−0.35 |
−2.09 to 1.39 |
0.63 |
8.090 |
0.32 |
13.51 |
Fixed |
|
Asian risk |
||||||||
A vs C |
−1.98 |
−11.03 to 7.06 |
0.53 |
18.83 |
0.01 |
78.76 |
Random |
|
AA vs CC |
−0.55 |
−4.18 to 3.07 |
0.66 |
4.210 |
0.37 |
5.020 |
Fixed |
|
AC vs CC |
−2.04 |
−9.80 to 5.71 |
0.46 |
12.54 |
0.01 |
68.11 |
Random |
|
AA + AC vs CC |
−2.22 |
−11.34 to 6.90 |
0.49 |
16.86 |
0.01 |
76.22 |
Random |
|
AA vs AC + CC |
−0.33 |
−3.42 to 2.75 |
0.75 |
2.980 |
0.56 |
0.01 |
Fixed |
|
Caucasian risk |
||||||||
A vs C |
164.81 |
143.45 to 186.16 |
0.00 |
36.90 |
0.01 |
94.58 |
Random |
|
AA vs CC |
−26.74 |
−590.48 to 536.99 |
0.65 |
6.200 |
0.04 |
67.77 |
Random |
|
AC vs CC |
170.25 |
−2043.43 to 2383.94 |
0.50 |
31.42 |
0.01 |
93.63 |
Random |
|
AA+AC vs CC |
154.88 |
−2017.17 to 2326.93 |
0.53 |
36.67 |
0.01 |
94.54 |
Random |
|
AA vs AC+CC |
−25.34 |
−525.35 to 474.67 |
0.63 |
4.940 |
0.08 |
59.58 |
Fixed |
|
Evaluation of heterogeneity
Q-test and I2 statistics were employed to test heterogeneity among the selected genetic association studies of LMP7–145 (C > A) gene polymorphism and cancer susceptibility. Significant heterogeneity was observed in three models of LMP7–145 (C > A). Thus, random effects model was used to synthesize the data (Table 4).
Association of LMP7–145 C > A polymorphism and overall cancer susceptibility
A total of eight studies were qualified for inclusion in this analysis to assess the overall association between LMP7−145 (C > A) polymorphism and cancer risk. Pooled analysis of total 1770 different cancer cases and 2013 controls demonstrated that homozygous AA genotype was significantly associated with 2.6 folds increased risk of overall cancer as compared to the CC genotype (AA vs CC: p = 0.001; OR = 2.602, 95% CI = 1.780 to 3.803). The recessive genetic model also demonstrated OR of 2.21 (AA vs AC+CC: p = 0.001; OR = 2.216, 95% CI = 1.525 to 3.221), indicating that individuals with AA genotype had almost 2 folds higher risk of cancer than those with the AC and CC genotypes. However, other genetic models, allelic (A vs C: p = 0.076; OR = 1.409, 95% CI = 0.964 to 2.060), heterozygous (AC vs CC: p = 0.118; OR = 1.378, 95% CI = 0.922 to 2.061), dominant model (AA+AC vs CC: p = 0.094; OR = 1.441, 95% CI = 0.939 to 2.210) did not show any association (Figure 2).
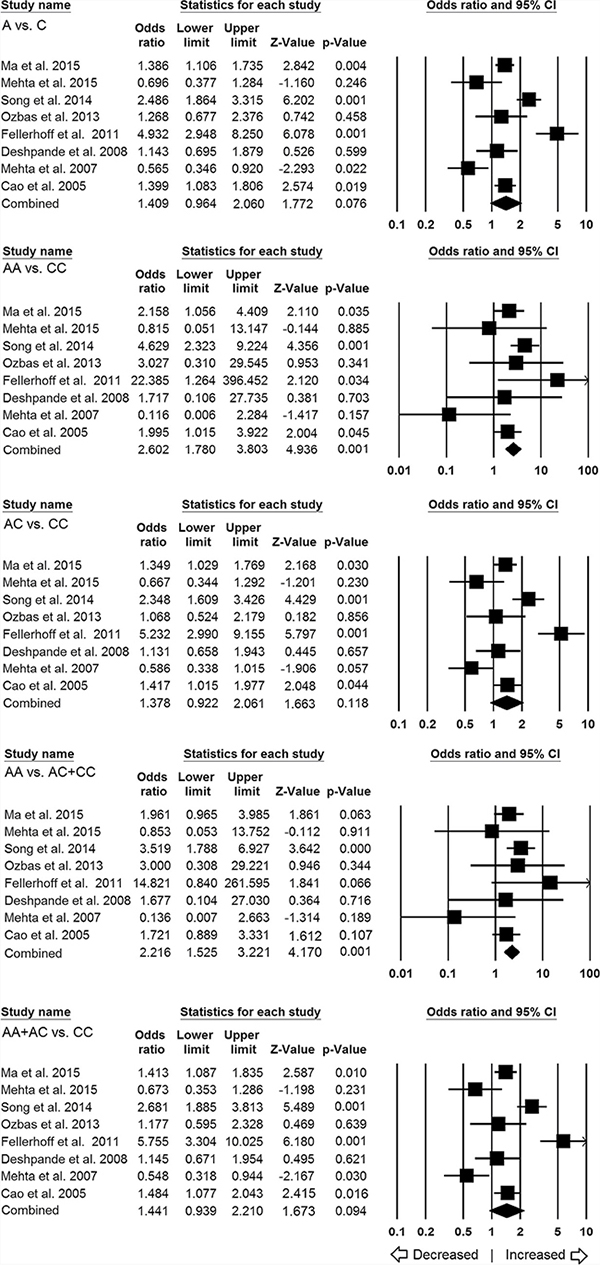
Figure 2: Association of LMP7–145 C > A polymorphism and cancer susceptibility in overall population. Forest plot of ORs with 95% CI of cancer risk associated with the LMP7 –145 C > A gene polymorphism for overall population. Note: Black square represents the value of OR and the size of the square indicates the inverse proportion relative to its variance. Horizontal line is the 95% CI of OR.
In ethnicity-wise subgroup analysis, a total of five studies were found qualified for Asian population and comprised of 1500 controls and 1335 different cancer cases. Heterogeneity was observed in three genetic models, so random effects model was applied to generate ORs and 95% CIs (Table 4). The results of Asian subgroup analysis suggested 2.2 folds statistically significant increased risk of cancer with homozygous AA (AA vs CC: p = 0.001; OR = 2.659, 95% CI = 1.800 to 3.928) and recessive genetic models (AA vs AC+CC: p = 0.001; OR = 2.256, 95% CI = 1.537 to 3.312) in comparison with wild type CC genotype. In addition, variant allele also revealed 1.4 folds increased risk of developing cancer (A vs C: p = 0.038; OR = 1.422, 95% CI = 1.020 to 1.983). Marginally significant risk was also observed for the dominant (AA+AC vs CC: p = 0.055; OR = 1.437, 95% CI = 0.993 to 2.079) genetic model, whereas, heterozygous (AC vs CC: p = 0.072; OR = 1.359, 95% CI = 0.973 to 1.897) model failed to show any risk (Figure 3).
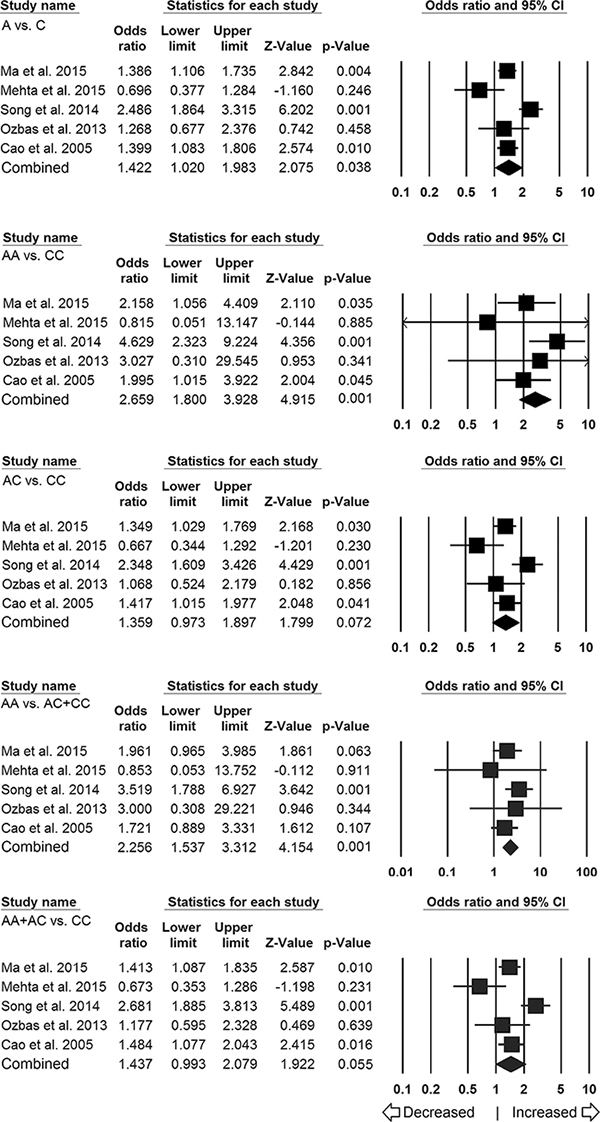
Figure 3: Association of LMP7–145 C > A polymorphism and cancer susceptibility in Asian subpopulation. Forest plot of ORs with 95% CI of cancer risk associated with the LMP7–145 C > A gene polymorphism for Asian subgroup population. Note: Black square represents the value of OR and the size of the square indicates the inverse proportion relative to its variance. Horizontal line is the 95% CI of OR.
For ethnicity-wise subgroup analysis of Caucasian population, only three studies (including 513 controls and 435 different cancer cases) were qualified and included for this analysis. Heterogeneity was observed in four genetic models; hence, random effects model was applied to generate pooled ORs and corresponding 95% CI values (Table 4). No obvious relevance of LMP7–145 (C > A) variant with cancer susceptibility was observed in all the genetic models (Figure 4), i.e., allelic (A vs C: p = 0.544; OR = 1.468, 95% CI = 0.425 to 5.074), homozygous (AA vs CC: p = 0.728; OR = 1.681, 95% CI = 0.090 to 31.375), heterozygous (AC vs CC: p = 0.520; OR = 1.512, 95% CI = 0.429 to 5.326), dominant model (AA+AC vs CC: p = 0.535; OR = 1.532, 95% CI = 0.398 to 5.891) and recessive (AA vs AC+CC: p = 0.584; OR = 1.589, 95% CI = 0.303 to 8.340).
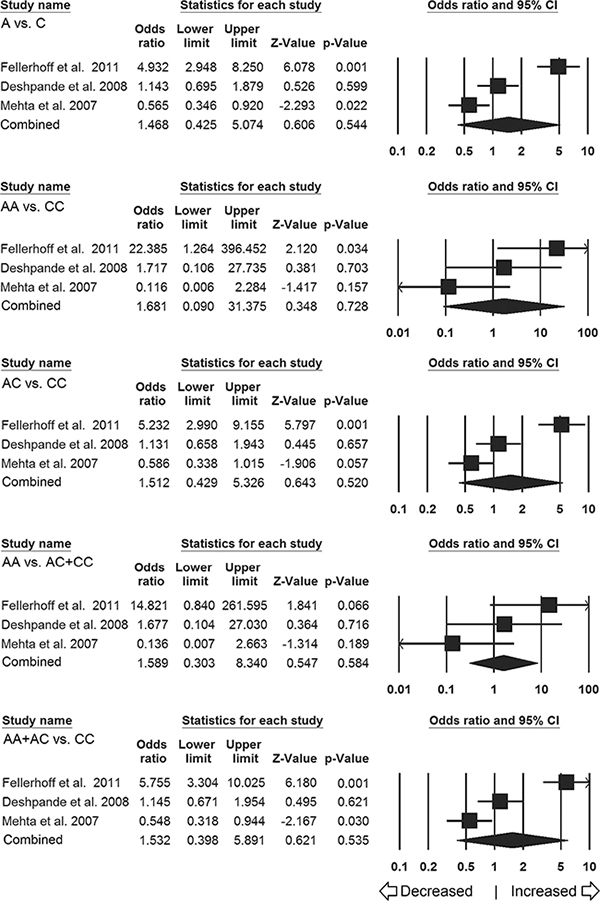
Figure 4: Association of LMP7 –145 C > A polymorphism and cancer susceptibility in caucasian subpopulation. Forest plot of ORs with 95% CI of cancer risk associated with the LMP7 –145 C > A gene polymorphism for Caucasian subgroup population. Note: Black square represents the value of OR and the size of the square indicates the inverse proportion relative to its variance. Horizontal line is the 95% CI of OR.
Sensitivity analysis for LMP7–145 C > A gene polymorphism and cancer susceptibility
To evaluate the impact of individual study on the risk of overall cancer, we performed leave-one-out sensitivity analysis and calculated the pooled ORs. The ORs recomputed after excluding each single study did not show any difference from their primary values, and assured the stability of the overall results (Supplementary Figure 2). Furthermore, the estimated pooled ORs in Asian (Supplementary Figure 3) and Caucasian (Supplementary Figure 4) subgroup populations did not change, which showed that the results of subgroup analysis were also stable.
Trial sequential analysis of LMP7–145 C > A gene polymorphism and cancer risk
The results of TSA were consistent with the conventional meta-analysis in the case of LMP7 –145 C > A gene polymorphism as the cumulative Z curve crossed with TSA monitoring boundary confirming that no further relevant trials are necessary. TSA analysis using recessive genetic (AA vs AC+CC) model, for e.g., showed that meta-analysis had enough number of studies (required sample size = 3611, achieved 80% power) for a significant conclusion, as it crossed the O’Breien-Fleming boundary (Figure 5A). Similarly, the cumulative Z curve crossed with TSA monitoring boundary before reaching the required information size in Asian subjects, confirming that LMP7 –145 C > A polymorphism is associated with increased cancer risk and further relevant trials are unnecessary (Figure 5B). Whereas, for Caucasian population, the cumulative Z curve failed to cross the trial monitoring boundary before reaching the required power and indicated that the cumulative evidence is insufficient and further trials are necessary (Figure 5C).
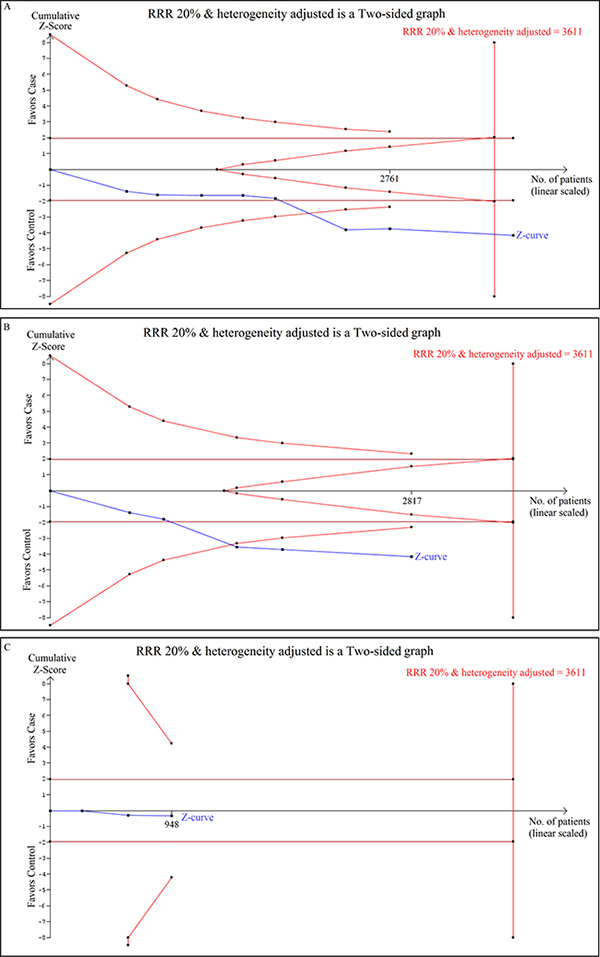
Figure 5: Trial sequential analysis of LMP7–145 C > A gene polymorphism and cancer risk. Trial sequence analysis of all studies on LMP7 –145 C > A gene polymorphism based on recessive genetic model. (A) In overall cancer risk (B) Cancer risk among Asian population (C) Cancer risk among Caucasian population.
DISCUSSION
Despite innumerable cutting edge improvements in diagnostic and therapeutic techniques, cancer remains the most lethal disease all over the world. Genetic variants that are important factor in immunogenesis are gradually being recognized as clues to individual’s susceptibility for cancer, exploring why individuals with shared environmental exposures do not always share cancer related morbidity and mortality.
The LMP7 protein is considered as catalytic subunit of the immunoproteasome, and can be induced with interferon-γ ensuing distinct subunit composition and altered catalytic characteristics [30]. As being important subunits of the immunoproteasome, the proteins of LMP7 have significant roles in antigen presentation and therefore they have played a suspected factor for a large variety of autoimmune diseases including cancer. Endogenous antigen presentation is a pivotal mechanism for the recognition of virally infected cells, maintenance of self-tolerance, and the surveillance of newly arising tumors by the immune system [31]. It is commonly anticipated that the functions of antigen processing and transport pathway might be impacted by the structural differences encoded by TAP (transporter associated with antigen processing) and LMP alleles, and immunoproteasome advances the quality and quantity of the generated class-I ligands. In the human host’s protective immunity, LMP/TAP system may perhaps detects tumor antigens and plays a key role in the immune surveillance via MHC-I molecule and CTL [32]. The genes that encode above said proteins are polymorphic in nature, hence it is possible that a particular genotype/allele of LMP7 –145 C > A polymorphism affect the functional alternations that may lead to the production of an insufficient peptide level, which may allow cancer cell to escape immune processing and lead to cancer development. Earlier studies already proved that individual studies with a low sample size may have not sufficient statistical power to identify a small risk factor. Hence, it is quite rational to appraise the precise association of LMP7 –145 C > A gene variant to understand the contribution in overall cancer risk.
To the best of our knowledge, this is the very first meta-analysis evaluating the association between LMP7 –145 C > A gene polymorphism and overall cancer risk for obtaining a precise conclusion. In the present study, we pooled all eight qualified case-control studies retrieved from different online web-databases that supported the key role of LMP7 −145 C > A polymorphism in overall cancer risk. More specifically, the AA genotype of LMP7 −145 C > A gene polymorphism had significantly greater risk of developing cancer in comparison with the wild type CC genotype. Subgroup analysis by ethnicity also demonstrated a similar trend of an increased risk of overall cancer in Asian population. Whereas, no association of LMP7 −145 C > A gene polymorphism with cancer risk was observed in Caucasian population.
In addition, the potential association of LMP7 −145 C > A polymorphism with overall cancer risk was confirmed by the Trial Sequential analysis, which further strengthen the conclusion that LMP7 −145 C > A polymorphism confers an increased risk of cancer. Overall, the pooled analysis suggested that genetic polymorphisms in the LMP7 gene may affect its expression level and play a crucial role in the failure of immunosurveillance and contribute to cancer progression. This is in line with previously published reports where they observed low protein expression and protein down regulation of LMP7 in several malignancies [31, 33, 34].
Although, the etiology of cancer is polygenic in nature and the roles of LMP7 polymorphism in the risk of developing cancer are quite diverse. Hence, a single genetic polymorphism is usually inadequate to clarify the susceptibility of this complex disease possibly because of significant heterogeneity in the clinical features and acquired genetic alterations.
This meta-analysis has certain limitations that must be discussed for better understanding and for the future studies, for e.g., first, heterogeneity was present in some genetic models and thus the random-effects model was applied to obtain the broader CI values. Second, due to non-availability of the original data for each included study, we failed to adjust the pooled ORs with respect to subjects’ age, sex, and environmental factors.
Despite above mentioned minor limitations, the current study has some major advantages, like, in order to check the credibility of the findings, we performed the publication bias analysis and the generated funnel plots suggested no obvious publication bias among the selected studies. This suggests that the results of the present meta-analysis are statically robust. Sensitivity analysis also indicated that no single study yield obvious impact on the pooled ORs and corresponding CIs. The credibility of all the eight included studies in terms of their quality was also checked using NOS scale. Additionally, in order to reconfirm the results of meta-analysis, TSA was performed and the results of TSA suggested that the conclusions were robust. Moreover, explicit criteria for the study examination and inclusion, strict data extraction, and exhaustive pooled analysis were applied to draw satisfactory and reliable conclusion.
In summary, the overall results of the present meta-analysis suggested that individuals with LMP7−145 C > A genetic polymorphism have increased cancer risk. These results will improve our understanding of the role of LMP7−145 C > A genetic polymorphism and assist in identification of at-risk individuals. Furthermore, this increasing knowledge could imply for other aspects of cancer management, including prevention, screening, and therapeutic treatment.
Also, future larger studies comprising other relevant factors along with integrative network modules analysis are warranted to clarify the potential role of LMP7 genetic variant in cancer risk.
MATERIALS AND METHODS
Identification and eligibility of the relevant studies
A systematic search was performed using PubMed (Medline), Google Scholar and EMBASE web-databases to retrieve compatible and peer reviewed research articles for this meta-analysis. Last search was updated on January 2017 with the combination of following keywords: ‘low molecular weight polypeptide 7 OR LMP7 OR PSMB8 gene (polymorphism OR mutation OR variant) AND cancer susceptibility OR risk. The search was limited on published studies that had been conducted in human subjects only. All the retrieve articles were examined by reading their titles and abstracts, and all the published studies matching with the above said eligibility criteria were selected for this meta-analysis. We also did manual search of the reference list from the retrieve articles for other eligible pertinent studies.
Inclusion and exclusion criteria
All the published articles included in the current meta-analysis had to meet all the given criteria: a) it must evaluated the association between LMP7 −145 C > A polymorphisms and cancer risk, b) used a case-control design, c) recruited histologically confirmed cancer patients and healthy controls, d) have available genotype frequency in cases and controls, e) and must be published in the English language. In addition to above, when the same patient/subject populations appeared in several publications, only the most recent one or the complete study was included in this meta-analysis.
The major reasons for study exclusion for this pooled analysis were overlapping of the data, case-only studies, review articles, and lack of genotype frequencies or number not reported. The information related to the selection (inclusion/ exclusion) of the studies is appended as Figure 1 in the form of PRISMA 2009 Flow Diagram.
Data extraction
For each retrieved article, the methodological quality assessment and data extraction were independently summarized in duplicate by two independent investigators (RKM & SAD) using a standard procedure. Data-collection form was used to ensure the accuracy of the collected data by strictly following the inclusion/ exclusion criteria as stated above. The major characteristic summarized from the retrieved articles included, the name of the first author, year of publication, the country of origin, the number of cases and controls, type of cancer, type of study, association/no-association status, methods of genotyping and genotype frequencies for the cases and controls. The cases related with disagreement on any item of the data from the collected studies were fully debated with the investigators in the presence of adjudicator (SH) to attain a final consensus.
Quality assessment by Newcastle-Ottawa scale (NOS) criteria
The methodological quality assessment of the included studies was also done separately by two independent investigators (RKM & SAD) using the NOS criteria. The NOS criteria majorly included three aspects: (1) subject selection: 0–4 points; (2) comparability of subjects: 0−2 points; (3) clinical outcomes: 0−3 points. The studies that were awarded 5 stars or more can be considered as of moderate to high quality [35]. The assessment was performed independently by two investigators (RKM & SAD, as stated above) and the inconformity was resolved by a discussion or consultation if necessary (with the help of adjudicator SH).
Statistical analysis
The potential association between studied LMP7 single nucleotide polymorphism and cancer susceptibility was assessed by computing crude Odds Ratios (ORs) and the corresponding 95% CI. The pooled ORs were estimated for the allele contrast, log-additive, dominant, and recessive models [36]. Heterogeneity assumptions between the studies across the eligible comparisons were performed by the chi-square-based Q-test [37]. Heterogeneity was considered significant when p-value < 0.05 to avoid underestimation of the presence of heterogeneity. A fixed effect model (if p > 0.05) [38] or a random effect model (if p < 0.05) [39] was used for pooling the results. Moreover, I2 statistics was also applied to efficiently test the heterogeneity among the selected studies [40]. Hardy-Weinberg equilibrium (HWE) in the controls was estimated via Chi-square test. Likewise, the Funnel plot asymmetry was estimated by Egger’s linear regression test, which is a type of linear regression approach to measure the funnel plot asymmetry on the natural logarithm scale of the ORs. The significance of the intercept was determined by the t-test (p-value <0.05 was considered as representation of statistically significant publication bias) [41]. All the statistical calculations were done by using Comprehensive Meta-Analysis (CMA) Version2 software program (Biostat, NJ, USA). All the p-values were two sided, and the statistical significance was considered as p-value less than 0.05 for this meta-analysis.
Trial sequential analysis (TSA)
According to Cochrane handbook, meta-analyses are considered to be the best if all the eligible trials are included in the analysis. However, it may not be sufficient evidence. Occasionally, the meta-analysis may lead to systematic errors (bias) or random errors (play of chance). In order to minimize errors, a novel statistical analysis based software program, TSA (Trial sequential analysis tool from Copenhagen Trial Unit, Center for Clinical Intervention Research, Denmark) has been introduced, which estimates the required information size, and an adjusted threshold for the statistical significance, and estimates the power of the current conclusion [42–44]. If the TSA monitoring boundary crossed with Z curve before the required information size is reached, robust evidence might have been confirmed, and further trails are unnecessary. In contrast, if the Z curve does not cross monitoring boundaries and the required information size has not been reached, it is necessary to continue doing trials. In the present investigation, Trial Sequential Analysis (Version 0.9, http://www.ctu.dk/tsa/) software program was used for the TSA analysis.
ACKNOWLEDGMENTS
The authors are grateful to the Deanship of Scientific Research, Jazan University, Jazan-45142, Saudi Arabia, for providing the infrastructural support and dry-lab facility for this research study.
Author contributions
Conceived and designed the study and experiments: RKM SAD AJ MW ML AKP BNM NA SH. Performed the experiments: RKM SAD AKP AJ MW. Analyzed the data: RKM SAD MW ML AKP BNM NA. Contributed reagents/materials/analysis tools: SAD AJ MW ML BNM NA SH. Wrote the paper: RKM SAD AKP SH. All authors reviewed the manuscript.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
No specific financial support was available for this research work.
REFERENCES
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016; 66:7–30.
2. Shin HR, Carlos MC, Varghese C. Cancer control in the Asia Pacific region: current status and concerns. Jpn J Clin Oncol. 2012; 42:867–81.
3. Bray F, Znaor A, Cueva P, Korir A, Swaminathan R, Ullrich A, Wang SA, Parkin DM. Planning and developing population-based cancer registration in low- and middle income settings. IARC Technical Publication No. 43. IARC, Lyon. 2015
4. Pharoah PD, Dunning AM, Ponder BA, Easton DF. Association studies for finding cancer-susceptibility genetic variants. Nat Rev Cancer. 2004; 4:850–60.
5. Chung CC, Magalhaes WC, Gonzalez-Bosquet J, Chanock SJ. Genome-wide association studies in cancer--current and future directions. Carcinogenesis. 2010; 31:111–20.
6. Savas S, Liu G. Genetic variations as cancer prognostic markers: review and update. Hum Mutat. 2009; 30:1369–77.
7. Heemels MT, Ploegh H. Generation, translocation, and presentation of MHC class I-restricted peptides. Annu Rev Biochem. 1995; 64:463–91.
8. Suzuki K, Sahara H, Okada Y, Yasoshima T, Hirohashi Y, Nabeta Y, Hirai I, Torigoe T, Takahashi S, Matsuura A, Takahashi N, Sasaki A, Suzuki M, et al. Identification of natural antigenic peptides of a human gastric signet ring cell carcinoma recognized by HLA-A31-restricted cytotoxic T lymphocytes. J Immunol. 1999; 163:2783–91.
9. Ortiz-Navarrete V, Seelig A, Gernold M, Frentzel S, Kloetzel PM, Hämmerling GJ. Subunit of the ‘20S’ proteasome (multicatalytic proteinase) encoded by the major histocompatibility complex. Nature. 1991; 353:662–4.
10. Früh K, Gossen M, Wang K, Bujard H, Peterson PA, Yang Y. Displacement of housekeeping proteasome subunits by MHC-encoded LMPs: a newly discovered mechanism for modulating the multicatalytic proteinase complex. EMBO J. 1994; 13:3236–44.
11. Dick LR, Aldrich C, Jameson SC, Moomaw CR, Pramanik BC, Doyle CK, DeMartino GN, Bevan MJ, Forman JM, Slaughter CA. Proteolytic processing of ovalbumin and beta-galactosidase by the proteasome to a yield antigenic peptides. J Immunol. 1994; 152:3884–94.
12. Ho YK, Bargagna-Mohan P, Wehenkel M, Mohan R, Kim KB. LMP2-specific inhibitors: chemical genetic tools for proteasome biology. Chem Biol. 2007; 14:419–30.
13. Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, Barber TD, Mandelker D, Leary RJ, Ptak J, Silliman N, Szabo S, Buckhaults P, Farrell C, et al. The consensus coding sequences of human breast and colorectal cancers. Science. 2006; 314:268–74.
14. Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P, Mankoo P, Carter H, Siu IM, Gallia GL, Olivi A, McLendon R, Rasheed BA, et al. An integrated genomic analysis of human glioblastoma multiforme. Science 2008; 321:1807–12.
15. Jones S, Zhang X, Parsons DW, Lin JC, Leary RJ, Angenendt P, Mankoo P, Carter H, Kamiyama H, Jimeno A, Hong SM, Fu B, Lin MT, et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science. 2008; 321 :1801–6.
16. Wei X, Walia V, Lin JC, Teer JK, Prickett TD, Gartner J, Davis S, Stemke-Hale K, Davies MA, Gershenwald JE, Robinson W, Robinson S, Rosenberg SA, et al. Exome sequencing identifies GRIN2A as frequently mutated in melanoma. Nature genetics. 2011; 43:442–6.
17. Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
18. Lim JK, Hunter J, Fernandez-Vina M, Mann DL. Characterization of LMP polymorphism in homozygous typing cells and a random population. Hum Immunol. 1999; 60:145–51.
19. Dosenko V, Mykhal’chuk DV, Zahoriĭ V, Khaĭtovych MV, Moĭbenko OO. Allelic polymorphism of genes encoding catalytic immunoproteasome subunits and its functional meaning. Fiziol Zh. 2005; 51:3–10.
20. Ma X, Yang C, Tang R, Xu Z, Zhang Z, Wang Y, Zhang J, Yang LI. Association between LMP2 and LMP7 gene polymorphisms and the risk of gastric cancer: A case-control study. Oncol Lett. 2015; 10:509–517.
21. Mehta AM, Spaans VM, Mahendra NB, Osse EM, Vet JN, Purwoto G, Surya IG, Cornian S, Peters AA, Fleuren GJ, Jordanova ES. Differences in genetic variation in antigen-processing machinery components and association with cervical carcinoma risk in two Indonesian populations. Immunogenetics. 2015; 67:267–75.
22. Song L, Ma N, Han L, Yan H, Yan B, Yuan Z, Cao B. Association between LMP2/LMP7 genetic variability and the metastasis risk of ovarian cancer in Chinese women in Beijing. Hum Immunol. 2014; 75:239–44.
23. Ozbas-Gerceker F, Bozman N, Kok S, Pehlivan M, Yilmaz M, Pehlivan S, Oguzkan-Balci S. Association of an LMP2 polymorphism with acute myeloid leukemia and multiple myeloma. Asian Pac J Cancer Prev. 2013; 14:6399–402.
24. Fellerhoff B, Gu S, Laumbacher B, Nerlich AG, Weiss EH, Glas J, Kopp R, Johnson JP, Wank R. The LMP7-K allele of the immunoproteasome exhibits reduced transcript stability and predicts high risk of colon cancer. Cancer Res. 2011; 71:7145–54.
25. Deshpande A, Wheeler CM, Hunt WC, Peyton CL, White PS, Valdez YE, Nolan JP. Variation in HLA class I antigen-processing genes and susceptibility to human papillomavirus type 16-associated cervical cancer. J Infect Dis. 2008; 197:371–81.
26. Mehta AM, Jordanova ES, van Wezel T, Uh HW, Corver WE, Kwappenberg KM, Verduijn W, Kenter GG, van der Burg SH, Fleuren GJ. Genetic variation of antigen processing machinery components and association with cervical carcinoma. Genes Chromosomes Cancer. 2007; 46:577–86.
27. Cao B, Tian X, Li Y, Jiang P, Ning T, Xing H, Zhao Y, Zhang C, Shi X, Chen D, Shen Y, Ke Y. LMP7/TAP2 gene polymorphisms and HPV infection in esophageal carcinoma patients from a high incidence area in China. Carcinogenesis. 2005; 26:1280–4.
28. Burton PR, Hansell AL, Fortier I, Manolio TA, Khoury MJ, Little J, Elliott P. Size matters: just how big is BIG?: Quantifying realistic sample size requirements for human genome epidemiology. Int J Epidemiol. 2009; 38:263–73.
29. Panagiotou OA, Willer CJ, Hirschhorn JN, Ioannidis JP. The power of meta-analysis in genome-wide association studies. Annu Rev Genomics Hum Genet. 2013; 14:441–65.
30. Liu Y, Luo YR, Lu X, Qiu XT, Zhou JP, Gong YF, Ding XD, Zhang Q. Association analysis of polymorphisms of porcine LMP2 and LMP7 genes with haematological traits. Mol Biol Rep. 2011; 38:4455–60.
31. Atkins D, Breuckmann A, Schmahl GE, Binner P, Ferrone S, Krummenauer F, Störkel S, Seliger B. MHC class I antigen processing pathway defects, ras mutations and disease stage in colorectal carcinoma. Int J Cancer. 2004; 109:265–73.
32. Xu C, Qi S, Gao L, Cui H, Liu M, Yang H, Li K, Cao B. Genetic polymorphisms of LMP/TAP gene and hepatitis B virus infection risk in the Chinese population. J Clin Immunol. 2007; 27:534–41.
33. Kageshita T, Hirai S, Ono T, Hicklin DJ, Ferrone S. Down-regulation of HLA class I antigen-processing molecules in malignant melanoma: association with disease progression. Am J Pathol. 1999; 154:745–54.
34. Zheng F, Hasim A, Anwer J, Niyaz M, Sheyhidin I. LMP gene promoter hypermethylation is a mechanism for its down regulation in Kazak’s esophageal squamous cell carcinomas. Mol Biol Rep. 2013; 40:2069–75
35. Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol. 2010; 25:603–5.
36. Woolf B. On estimating the relation between blood group and disease. Ann Hum Genet. 1955; 19:251–3.
37. Wu R, Li B. A multiplicative-epistatic model for analyzing interspecific differences in outcrossing species. Biometrics. 1999; 55:355–65.
38. Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959; 22:719–48.
39. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986;7:177–88.
40. Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003; 327:557–60.
41. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629–34.
42. Wetterslev J, Thorlund K, Brok J, Gluud C. Trial sequential analysis may establish when firm evidence is reached in cumulative meta-analysis. J Clin Epidemiol. 2008; 61:64–75.
43. Turner RM, Bird SM, Higgins JP. The impact of study size on meta-analyses: examination of underpowered studies in Cochrane reviews. PLoS One. 2013; 8:e59202.
44. Brok J, Thorlund K, Wetterslev J, Gluud C. Apparently conclusive meta-analyses may be inconclusive--Trial sequential analysis adjustment of random error risk due to repetitive testing of accumulating data in apparently conclusive neonatal meta-analyses. Int J Epidemiol. 2009; 38:287–98.

