INTRODUCTION
The incidence rate of Anal squamous cell carcinoma (ASCC) ranges from 1.0 to 2.5 per 100,000 population in a lot of developed countries, which is an rare malignancy of the anal canal and perianal skin area [1]. The annual incidence increases by about 2%, especially in females [2–4].
Even though many risk factors for the development of ASCC have been identified, ASCC is known to be strongly linked with a small double-stranded DNA virus, the human papillomavirus (HPV), known for its role in the development of head and neck squamous cell carcinoma (HNSCC), cervical cancer and other gynaecological cancers [5–7]. The prevalence in ASCC for high-risk HPV (HR-HPV) types, which are associated with carcinogenesis, ranges from 70 percent to 100 percent, depending on the sensitivity of the method used for HPV detection and the studied population [8–10].
One study reported that HPV-positive cervical cancer patients receiving radiotherapy have significantly better survival [11]. Other studies for HNSCC also shown that HPV-positive patients had a better prognosis than those HPV-negative [12–15]. Anus can be infected with these viruses the same way as the oral cavity, pharynx, and tonsils do; it is assumed that histological similarities of squamous epithelia between anus and the head and neck suggest clinical features.
The value for prognostic role of the HPV status has already been studied in ASCC patients. However, the sole detection of HPV DNA may misclassify cancers as being associated with HPV, since it does not prove the overexpression of viral oncogenes and thus the transformation induced by HPV [16]. As a result, additional biomarkers are being developed to refine the identification of HPV-associated tumors to achieve clinically acceptable accuracy. On this point, the cyclin-dependent kinase inhibitor 2A (CDKN2A), better known as p16INK4a (p16), appeared to be the best validated candidate because of its association with high-risk HPV infection. This cyclin-dependent kinase inhibitor is normally repressed by a phospho-retinoblastoma protein (pRB)/transcription factor E2F complex, but this suppression is inhibited by the high-risk E7 oncoprotein of HPV, resulting in an overexpression of p16INK4a [17, 18]. According to this, several studies have indicated that p16INK4a detection by immunohistochemistry (IHC) is predictive for a significantly improved response to treatment with RT/CRT and a more favourable prognosis in patients with HPV-associated malignancies [19–21].
Therefore, we conducted the meta-analysis to assess the combined effects of HPV status and p16 expression on overall survival (OS) and disease-free survival (DFS)/ disease-specific survival (DSS/relapse-free survival (RFS)/progression-free survival (PFS) in patients of ASCC.
RESULTS
Study selection
86 citations were generated from the search strategy. Through the literature search and browsing titles and/or abstracts, 17 papers were identified as potentially relevant (Figure 1). 13 of these 17 articles were subsequently excluded from our study due to several reasons, including 1 was review, 1 was case report, and 11 that did not provide HRs or CIs. A total of 4 prospective studies was included in the final meta-analysis [16, 22–24].
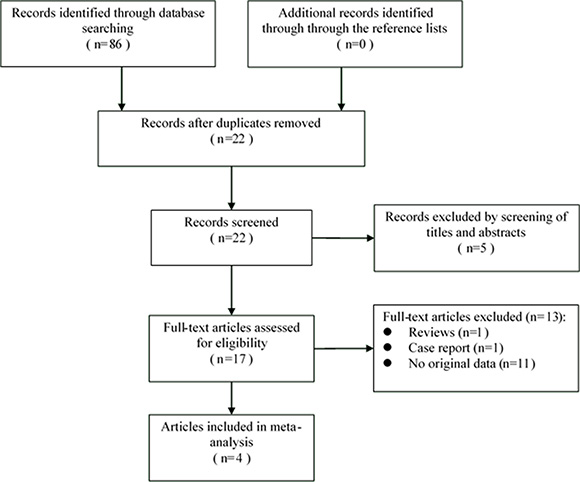
Figure 1: The PRISMA flow diagram of systematic literature search.
Characteristics of the selected studies
Table 1 summarized the each characteristics of the included 4 studies. The total number of patients included in themeta-analysis was 398, involingsin OS and DFS/DSS/RFS/PFS in both 4 studies. HRs and 95% CIs were directly extracted from the survival curve of two studies [16, 23]. One study [22] did not providespecific data for follow-up. The median follow-up period for all included studies ranged from 40 to 54 months. The prevalence of HPV varied from 67.9% to 95.8%, and the prevalence of p16 varied from 65.3% to 90.7%.
Table 1: Characteristics of the included studies
First author |
Year |
Period of |
Country |
Study design |
Stage |
No. of patients |
Genotype (s) |
HPV + ve |
|---|---|---|---|---|---|---|---|---|
Koerber SA |
2014 |
2000–2011 |
Germany |
Retrospective |
I-III |
90 |
24 types* |
75 (83.3) |
Rödel F |
2014 |
NA |
Germany |
Prospective |
I-IV |
95 |
28 types# |
91 (95.8) |
Meulendijks D |
2015 |
2003–2011 |
Netherlands |
Prospective |
I-III |
107 |
28 types# |
93 (86.9) |
Mai S |
2015 |
1990–2012 |
Germany |
Prospective |
I-III |
106 |
24 types* |
72 (67.9) |
Table 1: Characteristics of the included studies (continued)
First author |
p16 + ve |
Age, y |
Treatment |
DNA/p16 |
Median follow-up |
Survival |
Hazard |
|---|---|---|---|---|---|---|---|
Koerber SA |
75 (83.3) |
55 (22–94) |
CRT or R |
PCR/IHC |
48.6 (2.8–169.1) |
OS/PFS |
SC |
Rödel F |
62 (65.3) |
NA |
CRT |
PCR/IHC |
40 (1–264) |
OS/CSS |
SC |
Meulendijks D |
97 (90.7) |
60 (34–86) |
CRT or R |
PCR/IHC |
NA |
OS/DFS |
Adjusted |
Mai S |
74 (69.8) |
59.5(31–86) |
CRT |
PCR/IHC |
54 (5–205) |
OS/DFS |
Unadjusted |
Abbreviations: HPV + ve, human papillomavirus positive; p16 + ve, p16 positive; R, radiotherapy; CRT, chemo-radiotherapy; PCR, polymerase chain reaction; IHC, immunohistochemistry; OS, overall survival; DFS, disease-free survival; CSS, cancer-specific survival; RFS, relapse-free survival; PFS, progression-free survival; SC, survival curve; NA, not available.
* Including 15 high-risk types (HPV 16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 68, 73, 82), 3putative high-risk types (HPV 26,53,66), and 6 low-risk types(HPV 6, 11, 42, 43, 44, 70); # Including 20 high-risk types (HPV 16, 18, 26, 31, 33, 35, 39,45, 51, 52, 53, 56, 58, 59, 66, 68, 69, 70, 73, 82) and 8 low-risk types (HPV 6, 11,40, 43, 44, 54, 71, 74).
Quality assessment
The mean quality score of individual studies was 71.02% (range from 56.82% to 79.55%), indicating that the studies included in this meta-analysis were with relatively high quality (Table 2).
Table 2: Methodological assessments of the studies included in the meta-analysis
First author |
Global |
Scientific |
Laboratory |
Generalizability |
Results |
|---|---|---|---|---|---|
Koerber SA |
75.00 |
8 |
10 |
10 |
5 |
Rödel F |
72.23 |
9 |
10 |
8 |
5 |
Meulendijks D |
79.55 |
7 |
12 |
10 |
6 |
Mai S |
56.82 |
6 |
6 |
8 |
5 |
Results of the meta-analysis
The heterogeneity test, in particular, had a very low degree of heterogeneity among included studies, so that the pooled HR was obtained by a fixed effects model.
Overall survival
The HR pooled from the 3 individual effect estimates comparing HPV+/p16+ to HPV+/p16- cancers was 0.47 (95% CI: 0.19–1.17), which was not significantly correlated with OS. The meta-analysis showed a significant association for OS comparing HPV+/p16+ to HPV-/p16+ cancers the HR (95% CI) being 0.29 (95% CI: 0.13–0.64) from the 3 individual effect estimates. Compared with HPV-/p16- cancers, patients with HPV +/p16+ cancers had significant improved OS (HR = 0.30, 95% CI: 0.17–0.51), and patients with HPV+/p16- cancers had no significant improved OS (HR = 0.65, 95% CI: 0.21–2.04) (Figure 2).
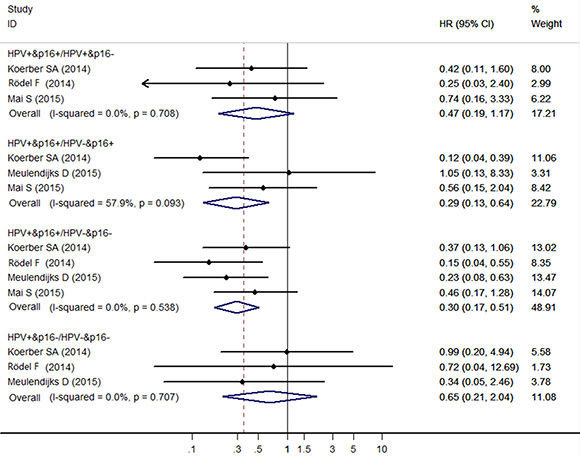
Figure 2: Forest plot for the association between HPV/p16 status and OS in ASCC patients.
DFS/DSS/RFS/PFS
The meta-analysis showed no significant association for DFS/DSS/RFS/PFS comparing HPV+/p16+ to HPV-/p16+ cancers the HR (95% CI) being 0.44 (95% CI: 0.16–1.22) from the 3 individual effect estimates. Compared with HPV-/p16+ cancers, patients with HPV+/p16+ cancers had significant improved DFS/DSS/RFS/PFS (HR = 0.31, 95% CI: 0.14–0.65), and patients with HPV+/p16- cancers had no significant improved DFS/DSS/RFS/PFS (HR = 0.94, 95% CI: 0.41–2.15). Compared with HPV-/p16- cancers, patients with HPV+/p16+ cancers had significant improved DFS/DSS/RFS/PFS (HR = 0.23, 95% CI: 0.14–0.36), patients with HPV+/p16- cancers had no significant improved DFS/DSS/RFS/PFS (HR=0.63, 95% CI: 0.32–1.24), and patients with HPV-/p16+ cancers had no significant improved DFS/DSS/RFS/PFS (HR = 0.88, 95% CI: 0.45–1.69) (Figure 3).
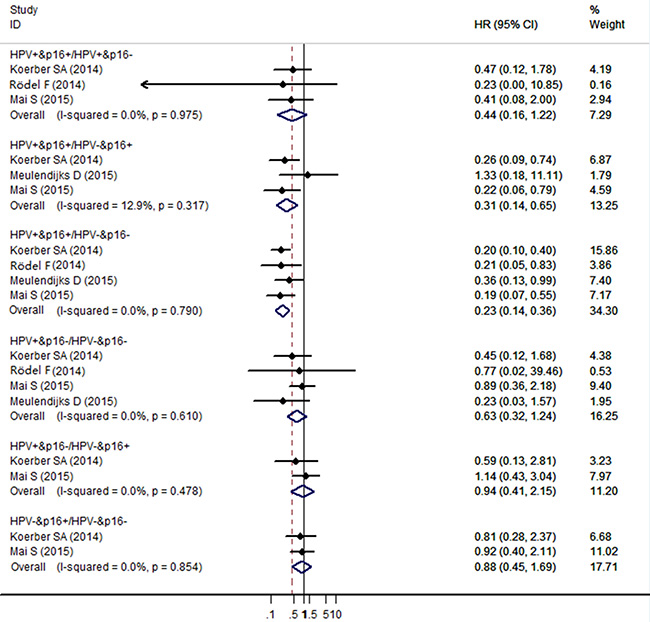
Figure 3: Forest plot for the association between HPV/p16 status and DFS/DSS/RFS/PFS in ASCC patients.
DISCUSSION
This is the first systematic review of the prognostic impact of HPV status together with p16 expression in ASCCs. In this study, we find that HPV and p16 status are strong predictors for OS and DFS/DSS/RFS/PFS. Pooled effect estimates among included studies demonstrated that ASCC patients with HPV-/p16- or HPV-/p16+ was considerably inferior compared with those patients with HPV+/p16+, suggesting that, to evaluating factors contributing to ASCC, HPV and p16 status especially HPV status, may be of therapeutic or prognostic utility.
In the late 1980s, it’s the first time to reported the relationship between HPV infection and the incidence of ASCC [25–27]. After then, the association between HPV infection and ASCC prognosis have been reported by a cumulative number of studies. However, the resulting studies were inconsistent. Since overexpression of p16 is not entirely limited to HPV-related transformation, it is important to analyze the combined status of HPV-DNA/p16 in order to more accurately identify cancers associated with HPV. S prognosis [28, 29]. However, data is limited in the assessment of the coefficient of exposure to HPV-DNA and p16 expression on the survival in patients of ASCC, which is of interest to us. CC.We showed that HPV+/p16+ ASCCs had a 70% reduction in OS and a 77% reduction in DFS/DSS/RFS/PFS, in comparison to HPV-/p16- ASCCs. HPV+/p16+ patients seemed to be sufficiently treated with current radiation therapy doses concomitantly with standard chemotherapy.
It is important to note that HPV-/p16+ patients do not have a comparably good prognosis as HPV+/p16+ patients, which is probably because the HPV-/p16+ tumors are not induced by HPV. Cancers associated HPV often have a viral sequence integrated into cancer cells genome. E6 and E7 are two early structural genes of the HPV. Through inactivation of p53 and the retinoblastoma protein (pRb), a negative regulator of the cyclin-dependent kinase inhibitor p16 and thus leads to upregulation of p16 [12, 30], the E6 and E7 proteins contribute to the genetic instability. Interfering changes in TP53 were reported in HPV-/p16- tumours (80%) and HPV-/p16+ tumours (33%), compared only sporadically to HPV+/p16+ tumours (6%) [22]. It is not surprising that TP53 mutations are only sporadically found in HPV+ tumours, as the HPV oncoprotein E6 inhibits p53 function by targeting it for ubiquitination and degradation. An additional mutation in TP53 would, therefore, not be necessary for these tumours to evolve. The apparent lower frequency of TP53 mutations in HPV-/p16+ tumours could be explained by aberrations in other tumour suppressor proteins, which could be investigated in future studies. The loss of p53 function was related to resistance to radiotherapy [31–33]. It is therefore conceivable that patients with HPV- tumours have a lower treatment response level due to a higher frequency of disrupted p53 function (via TP53 mutations). Other tests, such as HPV E6/E7 mRNA tests, can be evaluated in future studies in comparison with combined HPV DNA/p16 detection. However, HPV DNA and p16 tests are easy to perform and widely used, and are therefore accepted candidate prognostic markers.
The strengths including: a) the current analysis is the first to study the prognostic impact of HPV status together with p16 expression in ASCCs, and b) we used a strict inclusion and exclusion criterions, fully outcomes of interest (OS and DFS/DSS/RFS/PFS) and an advanced meta-analysis of HR for survival. The limitations including: a) a comparison of all combined HPV-DNA- and p16- subgroups in this meta-analysis is limited due to the few studies included with HPV-/p16+ and HPV+/p16- cancers. b) only English studies were included in the meta-analysis, which might resulting in language bias, and c) the adjusted HRs was only reported in one study, which might introduce residual confounding caused by other prognostic factors.
MATERIALS AND METHODS
Literature search strategy
A systematic search up to 31 May 2017 was conducted in MEDLINE (via PubMed) and Excerpta Medica database (EMBASE) to identify relevant articles. Search terms included ‘‘human papillomavirus OR HPV”, “p16 OR CDKN2 OR INK4A”, ‘‘anal cancer OR anal neoplasms OR anal carcinoma’’ combined with “prognosis OR prognostic OR survival”. Additional relevant references cited in retrieved articles were also evaluated.
Inclusion and exclusion criteria
All papers were reviewed by two authors (X.T. and H.Q.) independently. Uncertainties and discrepancies were resolved by consensus after discussing with a senior researcher (G.S.). All studies included in the final meta-analysis satisfied the following criteria: (a) patients were pathologically diagnosed as ASCC; (b) OS or DFS/DSS/RFS/PFS as the outcome of interest; (c) reported HRestimates with their corresponding 95% CI (or sufficient data to calculate of these effect measure), and (d) English articles. If the study was reported in duplication, the one published earlier or provided more detailed information was included. Review articles and editorials were included if they contained original data. Abstracts were excluded.
Quality assessment
The quality of each study was evaluated in accordance with the revised ELCWP scoring scale described by Steel [34]. Each item was assessed using an ordinal scale (possible values: 2, 1, 0). The overall score evaluated several dimensions of the methodology, grouped into four main categories: (1) scientific design: 0–10; (2) laboratory methodology: 0–14; (3) generalizability: 0–12; (4) results analysis: 0–8. The total scores ranged from 0 to 44. The final scores were expressed as percentages, ranging from 0% to 100%, higher values indicated a better methodological quality.
Data extraction
Two of the authors (X.T. and H.Q.) performed the data extraction from each article and discrepancies were resolved by consensus. For studies meeting our inclusion criteria, a standardized data extraction form was used to extract the following data: the first author’s name, year of publication, country of origin, study design, period of enrollment, the length of follow-up, characteristics of the studied population (sample size, age, stage of disease and treatment method), HPV detection methods, p16 detection methods, and HR estimates for OS or DFS/DSS/RFS/PFS with corresponding 95% CIs. When data for HR was not available, we extracted the total numbers of observed deaths and the numbers of patients in each group to calculate HR. [35]. Data were extracted by Engauge Digitizer version 4.1 (http://digitizer.sourceforge.net/) from the graphical survival plots when data were only available as Kaplan-Meier curves, [36]. then the estimation of the HR was performed by the described method [35].
Statistical analysis
The HR with 95% CI was used to compute the pooled HPV status combined p16 expression and the OS or DFS/DSS/RFS/PFS in ASCC patients. A fix-effect or random-effect model was used to pool the data, based on the Mantel–Haenszel method [37]. and the DerSimonian and Laird method, [38]. respectively. These two models provide similar results when between-studies heterogeneity is absent; otherwise, random-effect model is more appropriate.
Cochrane Q test (P < 0.10 indicated a high level of statistical heterogeneity) and I2 (values of 25%, 50% and 75% corresponding to low, moderate and high degrees of heterogeneity, respectively) was used to assess the heterogeneity between eligible studies, which test total variation across studies that was attributable to heterogeneity rather than to chance [39].
All statistical analyses were performed using STATA version 12 for Windows (StataCorp LP, College Station, TX, USA). A two-tailed P < 00.05 was considered statistically significant.
CONCLUSIONS
In conclusion, concomitant detection of HPV-DNA and p16 expression represents a prognostic marker in ASCC patients. Escalating treatment options for HPV-/p16- cancers and de-escalating therapy for ASCCs with HPV+/p16+ could be considered with the order to generate better outcome and fewer side effects related to treatment. An overexpression of p16 without HPV infection seem to have an unfavorable prognosis and may therefore require intensification of treatment for ASCCs. In this rule, prospective trials are mandatory to further determine the predictive joint-role of HPV-DNA and p16 expression in ASCC patients.
Author contributions
All authors have made substantial contributions to the conception and design of the study. G.S. and X.D. led protocol design, search, data extraction, statistical analysis, and manuscript drafting. X.T. and H.Q contributed to search, data extraction, and manuscript modifications. G.S., H.Z. and E.Z. contributed to quality assessment and revision of the manuscript. X.D. contributed to data interpretation and revision of the manuscript. All authors have reviewed and approved the final version.
CONFLICTS OF INTEREST
None.
FUNDING
This research was supported by the Shandong key research and development program under Fund number: 2017GSF218034.
REFERENCES
1. van der Zee RP, Richel O, de Vries HJ, Prins JM. The increasing incidence of anal cancer: can it be explained by trends in risk groups? The Netherlands journal of medicine. 2013; 71:401–411.
2. Johnson LG, Madeleine MM, Newcomer LM, Schwartz SM, Daling JR. Anal cancer incidence and survival: the surveillance, epidemiology, and end results experience, 1973–2000. Cancer. 2004; 101:281–288.
3. Bilimoria KY, Bentrem DJ, Rock CE, Stewart AK, Ko CY, Halverson A. Outcomes and prognostic factors for squamous-cell carcinoma of the anal canal: analysis of patients from the National Cancer Data Base. Diseases of the colon and rectum. 2009; 52:624–631.
4. Yao JN, Zhang XX, Zhou HN, Li YL, Xu HR, Wang CF, Chen LD, Gao B, Cheng P, Zhang LF. Human papillomavirus related anal squamous cell carcinoma survival: a systematic review and meta-analysis. Translational cancer research. 2017; 6:463–473.
5. Ciapponi A, Bardach A, Glujovsky D, Gibbons L, Picconi MA. Type-specific HPV prevalence in cervical cancer and high-grade lesions in Latin America and the Caribbean: systematic review and meta-analysis. PloS one. 2011; 6:e25493.
6. Kreimer AR, Clifford GM, Boyle P, Franceschi S. Human papillomavirus types in head and neck squamous cell carcinomas worldwide: a systematic review. Cancer epidemiology, biomarkers & prevention. 2005; 14:467–475.
7. Stelzer MK, Pitot HC, Liem A, Schweizer J, Mahoney C, Lambert PF. A mouse model for human anal cancer. Cancer prevention research. 2010; 3:1534–1541.
8. Daling JR, Madeleine MM, Johnson LG, Schwartz SM, Shera KA, Wurscher MA, Carter JJ, Porter PL, Galloway DA, McDougall JK. Human papillomavirus, smoking, and sexual practices in the etiology of anal cancer. Cancer. 2004; 101:270–280.
9. Hoots BE, Palefsky JM, Pimenta JM, Smith JS. Human papillomavirus type distribution in anal cancer and anal intraepithelial lesions. International journal of cancer. 2009; 124:2375–2383.
10. Valmary-Degano S, Jacquin E, Pretet JL, Monnien F, Girardo B, Arbez-Gindre F, Joly M, Bosset JF, Kantelip B, Mougin C. Signature patterns of human papillomavirus type 16 in invasive anal carcinoma. Human pathology. 2013; 44:992–1002.
11. Harima Y, Sawada S, Nagata K, Sougawa M, Ohnishi T. Human papilloma virus (HPV) DNA associated with prognosis of cervical cancer after radiotherapy. International journal of radiation oncology, biology, physics. 2002; 52:1345–1351.
12. Reimers N, Kasper HU, Weissenborn SJ, Stutzer H, Preuss SF, Hoffmann TK, Speel EJ, Dienes HP, Pfister HJ, Guntinas-Lichius O, Klussmann JP. Combined analysis of HPV-DNA, p16 and EGFR expression to predict prognosis in oropharyngeal cancer. International journal of cancer. 2007; 120:1731–1738.
13. Syrjanen S. HPV infections and tonsillar carcinoma. Journal of clinical pathology. 2004; 57:449–455.
14. Kong CS, Narasimhan B, Cao H, Kwok S, Erickson JP, Koong A, Pourmand N, Le QT. The relationship between human papillomavirus status and other molecular prognostic markers in head and neck squamous cell carcinomas. International journal of radiation oncology, biology, physics. 2009; 74:553–561.
15. Weinberger PM, Yu Z, Haffty BG, Kowalski D, Harigopal M, Brandsma J, Sasaki C, Joe J, Camp RL, Rimm DL, Psyrri A. Molecular classification identifies a subset of human papillomavirus--associated oropharyngeal cancers with favorable prognosis. Journal of clinical oncology. 2006; 24:736–747.
16. Koerber SA, Schoneweg C, Slynko A, Krug D, Haefner MF, Herfarth K, Debus J, Sterzing F, von Knebel Doeberitz M, Prigge ES, Reuschenbach M. Influence of human papillomavirus and p16(INK4a) on treatment outcome of patients with anal cancer. Radiotherapy and oncology. 2014; 113:331–336.
17. Lassen P, Eriksen JG, Hamilton-Dutoit S, Tramm T, Alsner J, Overgaard J. Effect of HPV-associated p16INK4A expression on response to radiotherapy and survival in squamous cell carcinoma of the head and neck. Journal of clinical oncology. 2009; 27:1992–1998.
18. Yugawa T, Kiyono T. Molecular mechanisms of cervical carcinogenesis by high-risk human papillomaviruses: novel functions of E6 and E7 oncoproteins. Reviews in medical virology. 2009; 19:97–113.
19. Palefsky JM, Giuliano AR, Goldstone S, Moreira ED Jr, Aranda C, Jessen H, Hillman R, Ferris D, Coutlee F, Stoler MH, Marshall JB, Radley D, Vuocolo S, et al. HPV vaccine against anal HPV infection and anal intraepithelial neoplasia. The New England journal of medicine. 2011; 365:1576–1585.
20. Rischin D, Young RJ, Fisher R, Fox SB, Le QT, Peters LJ, Solomon B, Choi J, O’Sullivan B, Kenny LM, McArthur GA. Prognostic significance of p16INK4A and human papillomavirus in patients with oropharyngeal cancer treated on TROG 02.02 phase III trial. Journal of clinical oncology. 2010; 28:4142–4148.
21. Yhim HY, Lee NR, Song EK, Kwak JY, Lee ST, Kim JH, Kim JS, Park HS, Chung IJ, Shim HJ, Hwang JE, Kim HR, Nam TK, et al. The prognostic significance of tumor human papillomavirus status for patients with anal squamous cell carcinoma treated with combined chemoradiotherapy. International journal of cancer. 2011; 129:1752–1760.
22. Meulendijks D, Tomasoa NB, Dewit L, Smits PH, Bakker R, van Velthuysen ML, Rosenberg EH, Beijnen JH, Schellens JH, Cats A. HPV-negative squamous cell carcinoma of the anal canal is unresponsive to standard treatment and frequently carries disruptive mutations in TP53. British journal of cancer. 2015; 112:1358–1366.
23. Rodel F, Wieland U, Fraunholz I, Kitz J, Rave-Frank M, Wolff HA, Weiss C, Wirtz R, Balermpas P, Fokas E, Rodel C. Human papillomavirus DNA load and p16INK4a expression predict for local control in patients with anal squamous cell carcinoma treated with chemoradiotherapy. International journal of cancer. 2015; 136:278–288.
24. Mai S, Welzel G, Ottstadt M, Lohr F, Severa S, Prigge ES, Wentzensen N, Trunk MJ, Wenz F, von Knebel-Doeberitz M, Reuschenbach M. Prognostic Relevance of HPV Infection and p16 Overexpression in Squamous Cell Anal Cancer. International journal of radiation oncology, biology, physics. 2015; 93:819–827.
25. Beckmann AM, Daling JR, Sherman KJ, Maden C, Miller BA, Coates RJ, Kiviat NB, Myerson D, Weiss NS, Hislop TG, Beagrie M, McDougall JK. Human papillomavirus infection and anal cancer. International journal of cancer. 1989; 43:1042–1049.
26. Ostrow RS, Manias DA, Fong WJ, Zachow KR, Faras AJ. A survey of human cancers for human papillomavirus DNA by filter hybridization. Cancer. 1987; 59:429–434.
27. Palmer JG, Scholffield JH, Coates PJ, Shepherd NA, Jass JR, Crawford LV, Northover JMA. Anal cancer and human papillomaviruses. Diseases of the Colon & Rectum. 1989; 32:1016–1022.
28. Deng Z, Hasegawa M, Aoki K, Matayoshi S, Kiyuna A, Yamashita Y, Uehara T, Agena S, Maeda H, Xie M, Suzuki M. A comprehensive evaluation of human papillomavirus positive status and p16INK4a overexpression as a prognostic biomarker in head and neck squamous cell carcinoma. International journal of oncology. 2014; 45:67–76.
29. Salazar CR, Anayannis N, Smith RV, Wang Y, Haigentz M Jr, Garg M, Schiff BA, Kawachi N, Elman J, Belbin TJ, Prystowsky MB, Burk RD, Schlecht NF. Combined P16 and human papillomavirus testing predicts head and neck cancer survival. International journal of cancer. 2014; 135:2404–2412.
30. Lewis JS Jr, Thorstad WL, Chernock RD, Haughey BH, Yip JH, Zhang Q, El-Mofty SK. p16 positive oropharyngeal squamous cell carcinoma:an entity with a favorable prognosis regardless of tumor HPV status. The American journal of surgical pathology. 2010; 34:1088–1096.
31. Lu C, El-Deiry WS. Targeting p53 for enhanced radio- and chemo-sensitivity. Apoptosis. 2009; 14:597–606.
32. Skinner HD, Sandulache VC, Ow TJ, Meyn RE, Yordy JS, Beadle BM, Fitzgerald AL, Giri U, Ang KK, Myers JN. TP53 disruptive mutations lead to head and neck cancer treatment failure through inhibition of radiation-induced senescence. Clinical cancer research. 2012; 18:290–300.
33. Kimple RJ, Smith MA, Blitzer GC, Torres AD, Martin JA, Yang RZ, Peet CR, Lorenz LD, Nickel KP, Klingelhutz AJ, Lambert PF, Harari PM. Enhanced radiation sensitivity in HPV-positive head and neck cancer. Cancer research. 2013; 73:4791–4800.
34. Steels E, Paesmans M, Berghmans T, Branle F, Lemaitre F, Mascaux C, Meert AP, Vallot F, Lafitte JJ, Sculier JP. Role of p53 as a prognostic factor for survival in lung cancer: a systematic review of the literature with a meta-analysis. The European respiratory journal. 2001; 18:705–719.
35. Parmar MK, Torri V, Stewart L. Extracting summary statistics to perform meta-analyses of the published literature for survival endpoints. Statistics in medicine. 1998; 17:2815–2834.
36. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials. 2007; 8:16.
37. Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959; 22:719–748.
38. DerSimonian R, Kacker R. Random-effects model for meta-analysis of clinical trials: an update. Contemporary clinical trials. 2007; 28:105–114.
39. Higgins JPT, Thompson SG. Quantifying heterogeneity in a meta-analysis. Statistics in medicine. 2002; 21:1539–1558.

