INTRODUCTION
MicroRNAs (miRNAs) are small, endogenous, noncoding, 18–24 nucleotide RNAs that can regulate gene expression by base pairing with the 3′-untranslated regions (UTRs) of target messenger RNAs [1, 2]. MiRNAs have been reported to contribute the regulation of a diverse range of genetic processes including development, apoptosis and differentiation. The expressions of circulating miRNAs are stable, reproducible and consistent among individuals of the same species [3]. MiRNAs are receiving increasing attention for their potential as diagnostic and therapeutic targets [4–6]. Many studies have reported that serum miRNAs have been identified as fingerprints for numerous diseases and cancers [7, 8].
Liver cirrhosis is a pathological condition of liver that results from sustained wound healing in response to various causes of chronic liver injury, including chronic hepatitis B (CHB), chronic hepatitis C (CHC), autoimmune hepatitis and alcoholic hepatitis [9]. The exact prevalence of cirrhosis worldwide is unknown. More than one million deaths worldwide were attributed to cirrhosis in 2010, although these figures are probably heavily under-reported. The total worldwide prevalence of cirrhosis has been estimated to be approximately 1% with significant regional variation due to the presence of viral hepatitis, metabolic syndrome and alcohol consumption [10]. Cirrhosis results in 170,000 deaths per year in Europe and 33,539 deaths per year in the USA. The main causes in these countries are infection with hepatitis C virus, alcohol misuse and non-alcoholic liver disease. The numbers in Europe and the USA are even higher than those in most Asian and African countries where chronic viral hepatitis B and C are common [11]. The prevalence of cirrhosis is difficult to assess and probably higher than reported because the initial stages are asymptomatic, so the disorder is undiagnosed.
Liver biopsy is considered as the gold standard for evaluating fibrosis [12]. However, it is limited by the need for hospital admission and sedation in children. It is not widely accepted by patients due to its limitations, especially the serious risks that include bleeding. The accuracy of liver histology assessment has been challenged because of sampling errors because, for example, the liver specimens are small or fragmented [13]. Advanced imaging technologies, including magnetic resonance imaging (MRI), computed tomography (CT), and transient elastography (TE) may also be used to detect liver fibrosis, but the use of these methods in countries such as China is limited by their high costs and lack of availability in many medical institutions. The diagnostic accuracy of CT, MRI and TE may be influenced by many factors, including obesity, ascites, acute inflammation, liver congestion, and elevated portal vein pressure [14, 15]. Many non-invasive markers for assessing liver cirrhosis are applied in clinical practice but are not sufficiently accurate because of low sensitivities and specificities [16]. Noninvasive fibrosis indices, such as the APRI and FIB-4, are associated with lower costs, do not require particular expertise in their interpretation, and can be performed in an outpatient setting. However, these indices have limited use in distinguishing patients who have chronic hepatitis from those who have developed cirrhosis [17].
In the past, an overwhelming amount of data supporting a role for miRNAs in the development and progression of chronic liver diseases into liver cirrhosis and finally hepatocellular carcinoma (HCC) has been presented [18, 19]. Various miRNAs have been reported to be correlated with liver cirrhosis and could be used as novel non-invasive biomarkers, including miRNA122, miRNA181b, miRNA29, etc. [20–23]. Therefore, we conducted a systematic and comprehensive meta-analysis of all eligible studies to explore the overall diagnostic values of serum miRNAs as promising biomarkers of liver cirrhosis.
RESULTS
Search results and characteristics of the eligible studies
We searched 430 records of which 92 records were duplicates. After a primary screening of the titles and abstracts, 304 records were excluded. Further articles were excluded after review. Seven eligible articles with 627 patients were included in our meta-analysis [33–39] (Figure 1). The characteristics of all 8 of the included publications are provided in Table 1. The included publications were published between 2013 and 2016. All 1045 described subjects were included. The results of the QUADAS-2 study quality assessments are presented in Figure 2. The majority of all articles included in the current meta-analysis satisfied most of the items in the QUADAS-2, which suggest that the overall quality of included studies was moderate-high.
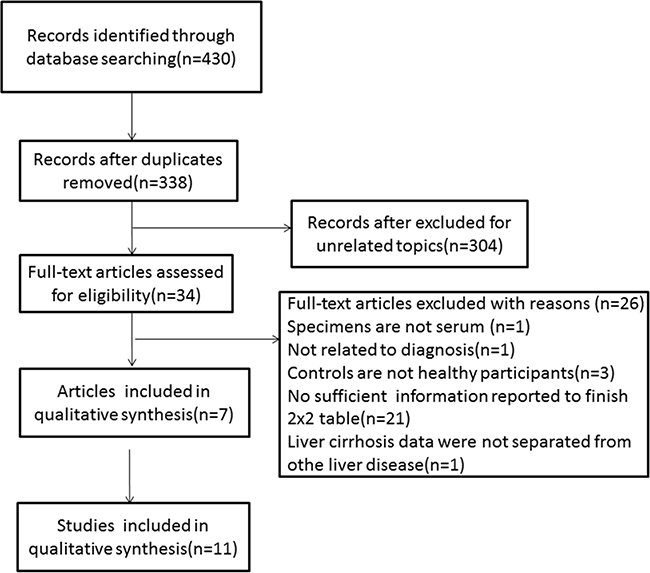
Figure 1: Flow diagram of study inclusion and exclusion for meta-analysis.
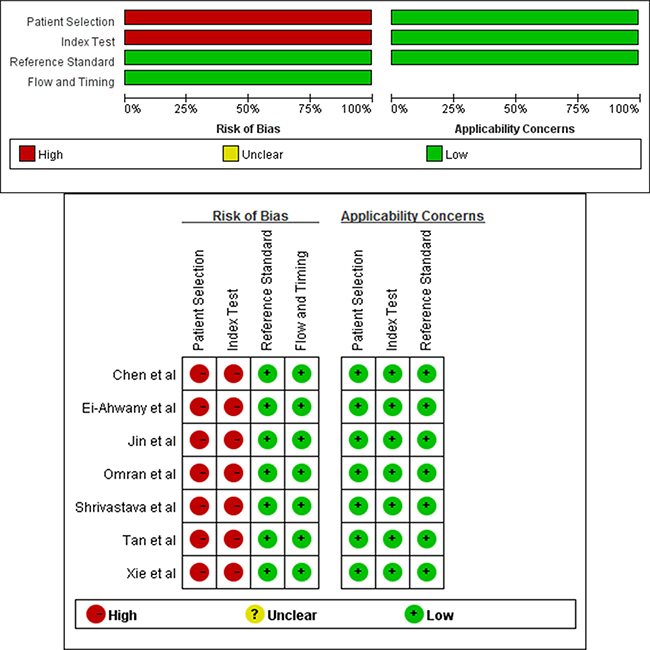
Figure 2: Details of QUADAS-2 quality assessment of each included study (QUADAS-2 tool).
Table 1: Characteristics of eligible studies
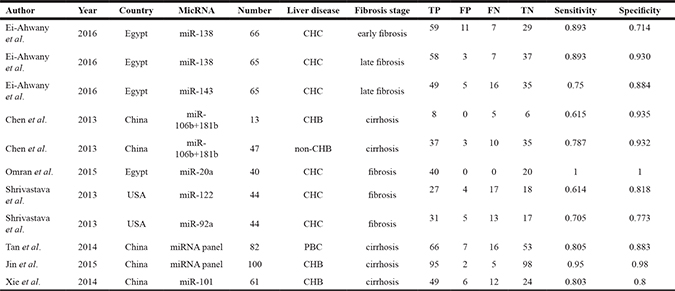
Abbreviations: PBC: Primary Biliary Cirrhosis, CHC: Chronic hepatitis C, CHB: Chronic hepatitis B, TP: true positive, FP, false positive, TN: true negative, FN, false negative.
Diagnostic accuracy of the serum miRNAs in the detection of liver cirrhosis
The sensitivity was calculated with the random effects model, and the pooled sensitivity was 0.83 (95% confidence interval [CI]: 0.80–0.86; Figure 3A). The pooled diagnostic specificity was 0.89 (95% CI: 0.86–0.92) based on the random effects model (Figure 3B). The pooled PLR was 6.41 (95% CI: 3.93–10.44) based on the random effects model (Figure 3C). The pooled NLR was 0.22 (95% CI: 0.14–0.33) based on the random effects model (Figure 3D). The DOR was pooled in a random effects model, and the pooled DOR was 35.18 (95% CI: 15.90–77.81; Figure 4A). The area under the ROC curve was 0.93 (95% CI: 0.91–0.95) (Figure 4B). To evaluate the clinical utility of the index test, a Fagan’s nomogram was created to predict the increasing inerrability of a positive diagnosis using the value of the test, and it was used to estimate the post-test probabilities (Figure 4C). The PLR was 8, the NLR was 0.18, and the post-test probabilities were 67 and 4, respectively. These data demonstrate that circulating miRNAs can be assayed with high diagnostic accuracy and specificity.
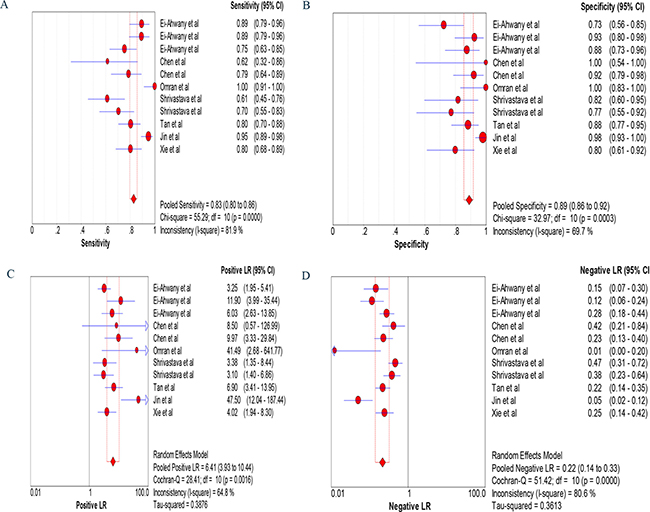
Figure 3: Forest plots of sensitivity (A), specificity (B), positive likelihood ratio specificity (C) and negative likelihood ratio specificity (D). The width of the horizontal line represents the 95% CI of each study; square proportional means the weight of every study. The weight is evaluated by the sample size and is presented as percent of total. The diamond represents pooled sensitivity, specificity, positive likelihood ratio specificity, negative likelihood ratio specificity and 95% CI.
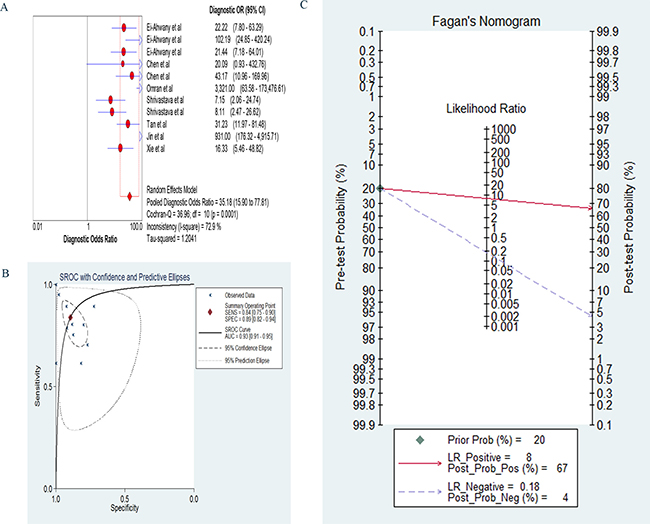
Figure 4: (A) The forest of diagnostic odds ratio specificity; (B) The pooled receiver operating characteristic curve: each X mark represents a study and AUC is the area under the curve; (C) Fagan’s Nomogram for calculation of post-test probabilities.
Statistical heterogeneity of the included studies
Differences in cut-off values lead to the threshold effect. When there is a threshold effect, an inverse correlation is present between the sensitivity and specificity. The Spearman correlation coefficient was −0.318, and the p value was 0.340 (P > 0.05), which indicated that there was no significant threshold effect.
Publication bias and sensitivity analysis
A Deek’s funnel plot was constructed, and the asymmetry test was performed to explore any potential publication bias in this meta-analysis. No significant publication bias was discovered (P = 0.239 > 0.05; Figure 5A). The sensitivity analysis is presented in Figure 5B; this analysis was accomplished by excluding studies one by one. The data were stable and not significantly different.
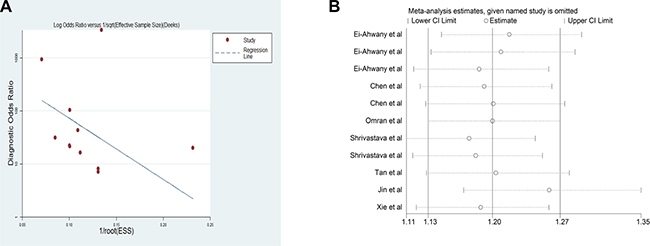
Figure 5: (A) Deek’s funnel plot indicates no significant publication bias (p = 0.239 > 0.05); (B) Sensitivity analysis plot of meta-analysis. Every row represents an included study. The width of the horizontal line represents the 95% CI for each study. The vertical bar on both sides represents the lowest and highest values of 95% CI.
Sub-group analysis
The sub-group analysis is presented in Table 2. The sample size (>60 or ≤60), miRNA type (single or combined) and liver disease (CHC or others) are displayed. The data suggested that the combined miRNAs exhibited greater diagnostic accuracy than the single miRNAs. The results were less accurate for the CHC cirrhosis patients than those with liver cirrhosis resulting from other diseases.
Table 2: Detail information of subgroup analysis
Subgroup |
Patients |
Sensitivity |
Specificity |
PLR |
NLR |
DOR |
AUC |
|---|---|---|---|---|---|---|---|
Total |
685 |
0.83 (0.80–0.86) |
0.89 (0.86–0.92) |
6.41 (3.93–10.44) |
0.22 (0.14–0.33) |
35.18 (15.90–77.81) |
0.93 (0.91–0.95) |
Sample size |
|||||||
>60 |
503 |
0.856 (0.820–0.888) |
0.890 (0.850–0.923) |
7.105 (3.640–13.868) |
0.166 (0.103–0.266) |
46.702 (17.539–124.35) |
0.93 (0.91–0.95) |
≤60 |
182 |
0.761 (0.693–0.802) |
0.889 (0.814–0.941) |
5.460 (2.500–11.924) |
0.323 (0.184–0.567) |
22.865 (5.789–90.313) |
0.94 (0.91–0.96) |
MiRNA type |
|||||||
single |
385 |
0.813 (0.770–0.851) |
0.841 (0.785–0.887) |
4.512 (2.944–6.916) |
0.234 (0.144–0.382) |
22.307 (9.778–50.889) |
0.92 (0.89–0.94) |
combined |
300 |
0.851 (0.880–0.894) |
0.941 (0.900–0.969) |
12.417 (4.756–32.420) |
0.189 (0.086–0.417) |
75.885 (14.895–386.60) |
0.95 (0.93–0.97) |
Liver disease |
|||||||
CHC |
241 |
0.815 (0.768–0.856) |
0.848 (0.788–0.896) |
4.790 (2.797–8.203) |
0.225 (0.123–0.411) |
24.874 (9.058–68.286) |
0.93 (0.90–0.95) |
Other |
444 |
0.842 (0.796–0.881) |
0.923 (0.881–0.951) |
9.222 (3.974–21.400) |
0.202 (0.112–0.366) |
52.854 (14.231–196.29) |
0.94 (0.90–0.95) |
DISCUSSION
Currently, circulating miRNAs are attracting increasing attention for their high stability and great potential as biomarkers for various diseases and cancers [40–42]. Liver cirrhosis is the advanced stage of chronic liver diseases. Liver biopsy is not widely accepted by patients due to its limitations. Many non-invasion biomarkers have been used in the detection of liver cirrhosis, including the APRI and FIB-4. However, the diagnostic accuracies are inconsistent [43–45]. Previous studies have reported miRNAs as biomarkers of liver cirrhosis [46, 47]. This study is the first meta-analysis to assess the diagnostic value of circulating miRNAs in identifying liver cirrhosis. This meta-analysis was conducted with multiple searching strategies performed by independent reviewers according the inclusion and exclusion criteria.
Eleven studies from 7 articles with 1045 subjects (627 patients and 418 healthy controls) were included in this meta-analysis. The 11 included studies exhibited moderate or high sensitivity and specificity, as the sensitivities ranged from 0.614 to 1, and the specificities ranged from 0.714 to 1. The results of our meta-analysis indicated that circulating miRNAs presented satisfactory pooled sensitivity and specificity. The pooled sensitivity was 0.83 (95% CI, 0.80–0.86), and the pooled specificity was 0.89 (95% CI, 0.86–0.92); thus, circulating miRNAs represent a promising diagnostic marker for liver cirrhosis. We also used the SROC curve and the corresponding AUC to estimate the overall diagnostic performance in the meta-analysis. The evaluation criteria can be divided into 3 levels of accuracy: low (AUC: 0.5–0.7), moderate (AUC: 0.7–0.9), and high (AUC: 0.9–1) [48]. In this meta-analysis, the AUC value for liver cirrhosis was 0.93, which indicated a high level of overall accuracy.
To further evaluate the diagnostic accuracy, we analyzed the DOR, PLR and NLR. The DOR represents the discrimination ability of a diagnostic test and ranges from 0 to infinity; the greater DORs indicate greater discriminative abilities. In this meta-analysis, the DOR value was 35.18, which indicates that the overall accuracy of the circulating miRNAs for the diagnosis of liver cirrhosis was credible. The LRs indicate the amount by which the odds of disease increase or decrease with a positive or negative test result [49]. The probability of a true positive and the value of the PLR exhibit a direct ratio when the test is positive. Higher NLR values indicate higher probabilities of false-negatives when the test is negative. When the PLR>10 or the NLR<0.1, the likelihood of diagnosis or exclusion of a disease increases significantly [49]. Nevertheless, the PLR of 6.41 (95% CI, 3.93–10.44) and the NLR of 0.22 (95% CI: 0.14–0.33) indicated that patients with liver cirrhosis have a ~6.41-fold higher chance of testing positive based on the circulating miRNAs than the controls, and 22% of individuals with liver cirrhosis will have a negative result.
LRs and post-test probabilities are correlations for clinicians because they provide information about the likelihood of a patient with a positive or negative test actually exhibiting liver cirrhosis [25, 49]. From the Fagan’s nomogram, we found that, when a pre-test probability of 20% was specified, the post-test probability of positivity increased to 67% with a positive likelihood ratio of 8, and the post-test probability of negativity decreased to 4% with a negative likelihood ratio was 0.18. These outcomes suggest a stable value of circulating miRNAs in the diagnosis of liver cirrhosis.
The threshold effect is one of the causes of heterogeneity in diagnostic accuracy studies. We did not find obvious heterogeneity as a result of heterogeneity in our meta-analysis. Therefore, we performed a meta-regression to examine the effects of sample size, liver disease type and whether single or combined miRNAs were utilized. The results revealed that sample size was a potential source of heterogeneity in this meta-analysis. The sub-group analysis revealed that the diagnostic sensitivity, specificity, PLR, DOR and AUC for liver cirrhosis in the subgroup of combined miRNAs were greater than the corresponding values in the single miRNA subgroup. These findings indicate that combined miRNAs have higher diagnostic value than single miRNAs. Additionally, the sensitivity analysis and the Deek’s funnel plot revealed that there were no outliers or a significant publication bias (p = 0.239).
This meta-analysis has several limitations. First, despite extensive literature search were performed, the number of included studies and sample sizes were small, which may restrict our ability to evaluate the accuracy of circulating miRNAs in detecting liver cirrhosis. Therefore, more large-scale, well-designed and multi-center clinical researches should be performed before the application of circulating miRNAs for the diagnosis of patients with liver cirrhosis. Second, there was significant heterogeneity in this meta-analysis; the Spearman correlation coefficient data indicated that the heterogeneity was not due to a threshold effect. Thus, the heterogeneity may have primarily been due to the small sample sizes. We attempted to establish a subgroup analysis for the disease stages, but because one study did not discriminate fibrosis and cirrhosis, this was difficult. Future studies should be designed to evaluate the heterogeneity. Finally, the demographics of the studies were limited to three countries. We have reviewed records from other countries, but these studies did not meet the inclusion criteria. Thus, it remains unknown whether these findings may be applicable to other parts of the world.
In conclusion, this meta-analysis demonstrated that circulating miRNAs can serve as potential biomarkers of liver cirrhosis. However, further large-scale studies are needed to confirm our analyses.
MATERIALS AND METHODS
Searching strategy and study selection
We reviewed publications in PubMed, EMBASE and the Cochrane Library up to July 1, 2017. We used the following search terms: (‘serum’ or ‘plasma’ or ‘circulating’ or ‘blood’) and (‘microRNA’ or ‘miRNA’) and (‘liver fibrosis’ or ‘liver cirrhosis’) and (‘biomarker’ or ‘diagnosis’). Only studies published in English were included.
Inclusion and exclusion criteria
The inclusion criteria for the primary studies were as follows: (1) the study was a diagnostic study using serum miRNA; (2) the subjects included liver cirrhosis patients and healthy controls; and (3) sufficient information was available to construct 2 × 2 tables that consisted of the true positives (TPs), false positives (FPs), true negatives (TNs) and false negatives (FNs). Articles were excluded if the miRNAs were not detected using serum samples, if there was insufficient information reported to create a 2 x 2 table, and if the control groups were not healthy participants. The studies included in our meta-analysis were independently assessed by two investigators. All of the selected studies were managed using EndNote X7.
Data extraction and quality assessment
The first author’s name, year of publication, country, number of patients, miRNA type, liver disease, fibrosis stage and detection methods were collected from the eligible studies. Then, 2 × 2 tables that displayed the TP, TN, FP, and FN were created. The Quality Assessment of Diagnostic Accuracy Studies-2 (QUADAS-2) was used to evaluate the diagnostic accuracy qualities using the patient selection, index test, reference standard, and flow and timing [24].
Statistical analysis
For the diagnostic meta-analysis, the accuracy indicators included the pooled sensitivity (SEN), pooled specificity (SPE), positive likelihood ratio (PLR), negative likelihood ratio (NLR), diagnostic odds ratio (DOR), and their 95% confidence intervals (CIs) were calculated using the random-effects model [25, 26]. The PLR was based on the odds of positive test results for liver cirrhosis patients, and the NLR reflected the odds of positive results for those without cirrhosis. The DOR was the outcome of the combination of the PLR and NLR (DOR = PLR/NLR) [27]. Simultaneously, the summary receiver operator characteristic (SROC) curve was created, and the area under the SROC curve (AUC) was calculated. The analysis of the diagnostic accuracy was pursuant to a SROC curve and the AUC of the SROC [28, 29]. The heterogeneity was measured with the I2 and Q-test, and a P < 0.05 and an I2 > 50% indicated the existence of significant heterogeneity among studies. If heterogeneity was detected, the random effects model was employed; otherwise, the fixed effects model was used. Meta-regression was used to detect the potential heterogeneity among the included studies [30]. Additionally, the Spearman correlation coefficient was used to verify if the heterogeneity in the meta-analysis could be explained by a threshold effect. A threshold effect was defined as a positive correlation (P < 0.05) [31]. Publication bias was investigated using Deek’s funnel plot [32]. Sensitivity analysis was accomplished by excluding the studies one by one.
The data analyses were performed using the Meta-Disc statistical software version 1.4 (XI Cochrane Colloquium, Barcelona, Spain) and STATA software (version 12.0, STATA Corp, MIDAS module). Quality assessment was managed by Review Manager 5.3 (Cochrane Collaboration, Copenhagen, Denmark). A p value < 0.05 was considered statistically significant.
Author contributions
Conceptualization: Weifeng Liang, Lifei Yu. Formal analysis: LiweiGuo, Weiyan Li. Investigation: LiweiGuo, Weiyan Li. Methodology: Lei Zheng, Huanhuan Zhou. Validation: Weifeng Liang, Lifei Yu. Writing-original draft: Lei zheng, Huanhuan Zhou, Liyang Hu. Writing-review & editing: Weifeng Liang, Lifei Yu.
ACKNOWLEDGMENTS
We are grateful to everyone who helped complete this study successfully.
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
GRAND SUPPORT
This study was supported by National Science and Technology Major Project of the Ministry of Science and Technology of China (2017ZX10203202, 2017ZX10203202-003, 2014ZX10004006, 2013ZX10002005), and Social Development Project of Hangzhou Technology Bureau (20160533B11).
REFERENCES
1. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004; 116:281–297.
2. Ambros V. The functions of animal microRNAs. Nature. 2004; 431:350–355.
3. Harfe BD. MicroRNAs in vertebrate development. Curr Opin Genet Dev. 2005; 15:410–415.
4. Kotaki R, Koyama-Nasu R, Yamakawa N, Kotani A. miRNAs in Normal and Malignant Hematopoiesis. Int J Mol Sci. 2017; 18:1495.
5. Inamura K. Diagnostic and Therapeutic Potential of MicroRNAs in Lung Cancer. Cancers (Basel). 2017; 9:49.
6. Inamura K. Major Tumor Suppressor and Oncogenic Non-Coding RNAs: Clinical Relevance in Lung Cancer. Cells. 2017; 6:12.
7. Weiland M, Gao XH, Zhou L, Mi QS. Small RNAs have a large impact: circulating microRNAs as biomarkers for human diseases. RNA Biol. 2012; 9:850–859.
8. Wang J, Zhang KY, Liu SM, Sen S. Tumor-associated circulating microRNAs as biomarkers of cancer. Molecules. 2014; 19:1912–1938.
9. Schuppan D, Afdhal NH. Liver cirrhosis. Lancet. 2008; 371:838–851.
10. Management of liver cirrhosis. Lancet. 2014; 383:1694.
11. Tsochatzis EA, Bosch J, Burroughs AK. Liver cirrhosis. Lancet. 2014; 383:1749–1761.
12. Baranova A, Lal P, Birerdinc A, Younossi ZM. Non-invasive markers for hepatic fibrosis. BMC Gastroenterol. 2011; 11:91.
13. Bedossa P, Dargere D, Paradis V. Sampling variability of liver fibrosis in chronic hepatitis C. Hepatology. 2003; 38:1449–1457.
14. Tsochatzis EA, Gurusamy KS, Ntaoula S, Cholongitas E, Davidson BR, Burroughs AK. Elastography for the diagnosis of severity of fibrosis in chronic liver disease: a meta-analysis of diagnostic accuracy. J Hepatol. 2011; 54:650–659.
15. Yin M, Talwalkar JA, Glaser KJ, Venkatesh SK, Chen J, Manduca A, Ehman RL. Dynamic postprandial hepatic stiffness augmentation assessed with MR elastography in patients with chronic liver disease. AJR Am J Roentgenol. 2011; 197:64–70.
16. Stasi C, Milani S. Evolving strategies for liver fibrosis staging: Non-invasive assessment. World J Gastroenterol. 2017; 23:191–196.
17. Gitlin N. Hepatitis B: diagnosis, prevention, and treatment. Clin Chem. 1997; 43:1500–1506.
18. Lemoinne S, Thabut D, Housset C, Moreau R, Valla D, Boulanger CM, Rautou PE. The emerging roles of microvesicles in liver diseases. Nat Rev Gastroenterol Hepatol. 2014; 11:350–361.
19. Loosen SH, Schueller F, Trautwein C, Roy S, Roderburg C. Role of circulating microRNAs in liver diseases. World J Hepatol. 2017; 9:586–594.
20. Roy S, Benz F, Luedde T, Roderburg C. The role of miRNAs in the regulation of inflammatory processes during hepatofibrogenesis. Hepatobiliary Surg Nutr. 2015; 4:24–33.
21. Cermelli S, Ruggieri A, Marrero JA, Ioannou GN, Beretta L. Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic fatty liver disease. PLoS One. 2011; 6:e23937.
22. Salvoza NC, Klinzing DC, Gopez-Cervantes J, Baclig MO. Association of Circulating Serum miR-34a and miR-122 with Dyslipidemia among Patients with Non-Alcoholic Fatty Liver Disease. PLoS One. 2016; 11:e0153497.
23. Roderburg C, Urban GW, Bettermann K, Vucur M, Zimmermann H, Schmidt S, Janssen J, Koppe C, Knolle P, Castoldi M, Tacke F, Trautwein C, Luedde T. Micro-RNA profiling reveals a role for miR-29 in human and murine liver fibrosis. Hepatology. 2011; 53:209–218.
24. Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, Leeflang MM, Sterne JA, Bossuyt PM, and QUADAS-2 Group. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011; 155:529–536.
25. Glas AS, Lijmer JG, Prins MH, Bonsel GJ, Bossuyt PM. The diagnostic odds ratio: a single indicator of test performance. J Clin Epidemiol. 2003; 56:1129–1135.
26. Jaeschke R, Guyatt GH, Sackett DL. Users' guides to the medical literature. III. How to use an article about a diagnostic test. B. What are the results and will they help me in caring for my patients? The Evidence-Based Medicine Working Group. JAMA. 1994; 271:703–707.
27. Fischer JE, Bachmann LM, Jaeschke R. A readers' guide to the interpretation of diagnostic test properties: clinical example of sepsis. Intensive Care Med. 2003; 29:1043–1051.
28. Moses LE, Shapiro D, Littenberg B. Combining independent studies of a diagnostic test into a summary ROC curve: data-analytic approaches and some additional considerations. Stat Med. 1993; 12:1293–1316.
29. Rutter CM, Gatsonis CA. A hierarchical regression approach to meta-analysis of diagnostic test accuracy evaluations. Stat Med. 2001; 20:2865–2884.
30. Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003; 327:557–560.
31. Dinnes J, Deeks J, Kirby J, Roderick P. A methodological review of how heterogeneity has been examined in systematic reviews of diagnostic test accuracy. Health Technol Assess. 2005; 9:1–113, iii.
32. Deeks JJ, Macaskill P, Irwig L. The performance of tests of publication bias and other sample size effects in systematic reviews of diagnostic test accuracy was assessed. J Clin Epidemiol. 2005; 58:882–893.
33. El-Ahwany E, Nagy F, Zoheiry M, Shemis M, Nosseir M, Taleb HA, El Ghannam M, Atta R, Zada S. Circulating miRNAs as Predictor Markers for Activation of Hepatic Stellate Cells and Progression of HCV-Induced Liver Fibrosis. Electron Physician. 2016; 8:1804–1810.
34. Chen YJ, Zhu JM, Wu H, Fan J, Zhou J, Hu J, Yu Q, Liu TT, Yang L, Wu CL, Guo XL, Huang XW, Shen XZ. Circulating microRNAs as a Fingerprint for Liver Cirrhosis. PLoS One. 2013; 8:e66577.
35. Omran AA, Osman KS, Kamel HM, Abdel-Naem EA, Hasan DE. MicroRNA-122 as a Novel Non-Invasive Marker of Liver Fibrosis in Hepatitis C Virus Patients. Clin Lab. 2016; 62:1329–1337.
36. Shrivastava S, Petrone J, Steele R, Lauer GM, Di Bisceglie AM, Ray RB. Up-regulation of circulating miR-20a is correlated with hepatitis C virus-mediated liver disease progression. Hepatology. 2013; 58:863–871.
37. Tan Y, Pan T, Ye Y, Ge G, Chen L, Wen D, Zou S. Serum microRNAs as potential biomarkers of primary biliary cirrhosis. PLoS One. 2014; 9:e111424.
38. Jin BX, Zhang YH, Jin WJ, Sun XY, Qiao GF, Wei YY, Sun LB, Zhang WH, Li N. MicroRNA panels as disease biomarkers distinguishing hepatitis B virus infection caused hepatitis and liver cirrhosis. Sci Rep. 2015; 5:15026.
39. Xie Y, Yao Q, Butt AM, Guo J, Tian Z, Bao X, Li H, Meng Q, Lu J. Expression profiling of serum microRNA-101 in HBV-associated chronic hepatitis, liver cirrhosis, and hepatocellular carcinoma. Cancer Biol Ther. 2014; 15:1248–1255.
40. Armand-Labit V, Pradines A. Circulating cell-free microRNAs as clinical cancer biomarkers. Biomol Concepts. 2017; 8:61–81.
41. Pritchard CC, Kroh E, Wood B, Arroyo JD, Dougherty KJ, Miyaji MM, Tait JF, Tewari M. Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer Prev Res (Phila). 2012; 5:492–97.
42. Di Meo A, Bartlett J, Cheng Y, Pasic MD, Yousef GM. Liquid biopsy: a step forward towards precision medicine in urologic malignancies. Mol Cancer. 2017; 16:80.
43. Xu H, Kong W, Liu L, Chi X, Wang X, Wu R, Gao X, Wang H, Qu L, Qi Y, Pan Y, Niu J. Accuracy of M2BPGi, compared with Fibro Scan®, in analysis of liver fibrosis in patients with hepatitis C. BMC Gastroenterol. 2017; 17:62.
44. Li Q, Lu C, Li W, Huang Y, Chen L. Impact of age on the diagnostic performances and cut-offs of APRI and FIB-4 for significant fibrosis and cirrhosis in chronic hepatitis B. Oncotarget. 2017; 8:45768–45776. https://doi.org/10.18632/oncotarget.17470.
45. Li Q, Ren X, Lu C, Li W, Huang Y, Chen L. Evaluation of APRI and FIB-4 for noninvasive assessment of significant fibrosis and cirrhosis in HBeAg-negative CHB patients with ALT≤2 ULN: A retrospective cohort study. Medicine (Baltimore). 2017; 96:e6336.
46. Riazalhosseini B, Mohamed R, Apalasamy YD, Langmia IM, Mohamed Z. Circulating microRNA as a marker for predicting liver disease progression in patients with chronic hepatitis B. Rev Soc Bras Med Trop. 2017; 50:161–166.
47. Krauskopf J, de Kok TM, Schomaker SJ, Gosink M, Burt DA, Chandler P, Warner RL, Johnson KJ, Caiment F, Kleinjans JC, Aubrecht J. Serum microRNA signatures as "liquid biopsies" for interrogating hepatotoxic mechanisms and liver pathogenesis in human. PLoS One. 2017; 12:e0177928.
48. Swets JA. Measuring the accuracy of diagnostic systems. Science. 1988; 240:1285–1293.
49. Gallagher EJ. Clinical utility of likelihood ratios. Ann Emerg Med. 1998; 31:391–397.

