INTRODUCTION
Congenital heart diseases (CHDs) are a large group of prevalent and complex anatomic malformations of the heart, with the incidence of about 7.5% in newborns [1], and about 1% of the CHD patients would require clinical intervention [2]. The most common types of CHDs include tetralogy of Fallot, ventricular septal defects, pulmonary stenosis, patent ductusar teriosus, and mitral valve insufficiency [3, 4]. CHDs are often complicated with malignant arrhythmias or heart failure, which may lead to death [5]. Although many chromosomal variants have been identified for their associations with the illness [6–8], much remains unknown about the relationships between the genetic abnormalities and CHDs.
The mammalian heart is one of the first formed organs during embryogenesis. In the process, the human embryonic stem (HES) cells differentiate to various cell types of the ectoderm, endoderm and mesoderm, and cardiomyocytes are generated in the mesoderm [9]. The mammalian heart consists of several cell types [10], including myocardium cells, cardiac neural crest (NC) cells, aorticopulmonary septum cells and membranous ventricular septum cells [11]. These cardiac progenitors interact to form the heart and many gene defects or variants in the cells have been reported for their relevance with the heart malformations, such as muscle segment homeobox 1 (MSX1) [1, 12, 13]. In a previous study, we found that variants in the MSX1 gene are closely associated with the risk of ventricular septal defect (VSD, a common type of CHD) [14]. Additionally, our recent work has also demonstrated the associations of variants in other genes with the risk of CHD, such as those of the STX18 gene [15]. STX18 is involved in several cell processes, such as transporting vesicle membrane fusion with target compartments in the cellular activities [16] and forming complexes with cell cycle-related proteins [17, 18]. These findings suggest that genes associated with cellular genesis processes may affect the development of all embryonic tissues, including cardiac progenitor cells and therefore might be associated with the pathogenesis of CHDs. For example, the kinetochore protein ZW10 plays key roles in the cell cycle, especially at the cell division stage, so we postulated the potential involvement of ZW10 in CHD.
ZW10 ensures proper chromosome segregation and participates in turning off spindle checkpoint activity, preventing cells from prematurely exiting mitosis [19–22]. When inactive ZW10 and colchicine exist, the cells of Drosophila melanogaster showed chromosome missegregation and abnormal separation of sisterchromatids [20]. Similar phenomena were observed in HeLa cells when anti-ZW10 antibody was applied [21, 19, 23, 24]. Based on the documented key roles of ZW10 in the cellular processes, we postulate its association with CHD. To validate this postulation, we compared the gene sequences between 300 Chinese Han CHD patients and 600 controls and found that the variation rs2459976 in the ZW10 gene was associated with the risk of CHD.
RESULTS
Patients
Clinical diagnosis of all the participants was confirmed at the Second or the Fourth Affiliated Hospital of Harbin Medical University. The participants had no history or manifestations of any other systemic abnormalities. We also established that their mothers did not take medicines or attract infections during gestation, because these factors had been found to be associated with heart malformation in pregnancy [25, 26].
A total of 300 CHD patients and 600 unrelated controls were recruited for this study, and there were no statistical differences in age or gender composition between the two groups (Table 1). The 300 CHD patients contained 122 with ventricular septal defects (VSD), 109 with atrial septal defects (ASD), 46 with patent ductus arteriosus (PDA),11 with tetralogy of Fallot, and 12 with other types of congenital heart defects.
Table 1: General information of the study population
Parameter |
CHD |
Control |
F |
t |
P |
95% CI-Up |
95% CI-Low |
|---|---|---|---|---|---|---|---|
Sample (n) |
300 |
600 |
- |
- |
- |
- |
- |
Male/Female (n) |
134/166 |
260/340 |
- |
- |
0.704 |
- |
- |
Age (years) |
15.42 ± 17.00 |
14.02 ± 9.31 |
185.530 |
1.581 |
0.114 |
−0.33561 |
3.11994 |
Data are shown as mean ± SD; between the two groups, there were no statistical differences of the age and gender composition.
ZW10 gene analysis
We sequenced the ZW10 gene to test the hypothesis that germline common variants in ZW10 may confer susceptibility to CHD. We identified rs2885987 and rs11605483 near the 3′UTR, rs2271261 and rs2271796 within the intron region, and rs2459976 near the 5′UTR of the gene (Figure 1). The genetic heterozygosity of the rs11605483(T/T, 0.95; C/T, 0.05) and rs2271796 (T/T, 0.99; C/T, 0.01) loci was very low. On the other hand, the genetic heterozygosity of rs2885987,rs2271261 and rs2459976 in the gene ZW10 was considerably high, so we focused on the latter three variants for further analysis.

Figure 1: Schematic diagramoflocations of ZW10 gene SNPsrs2885987, rs11605483, rs2271261, rs2271796 and rs2459976.
Statistical analysis of SNPs rs2885987, rs2271261 and rs2459976
We conducted analyses on the three SNPs and found that rs2459976 was associated with the risk of CHD in the Chinese Han population (Chi-square value: 7.153; P value: 0.028), but rs2885987 and rs2271261 were not (Tables 2 and 3). The Hardy-Weinberg equilibrium test for the CHD and controls were conducted and it was in line with the equilibrium (Table 4).
Table 2: The genotypes and allele frequencies of SNPs rs2459976, rs2271261 and rs2885987 in 300 Chinese Han CHD patients and 600 non-CHD controls
SNP |
Group |
Genotype frequency (%) |
Allele frequency (%) |
||||
|---|---|---|---|---|---|---|---|
rs2459976 |
Genotype |
C/C |
C/T |
T/T |
C |
T |
|
CHD |
300 |
167 (55.67) |
108 (36.00) |
25 (8.33) |
442 (73.67) |
158 (26.33) |
|
ASD |
109 |
57 (52.29) |
39 (35.78) |
13 (11.93) |
153 (70.18) |
65 (29.82) |
|
VSD |
122 |
67 (54.92) |
47 (38.52) |
8 (6.56) |
181 (74.18) |
63 (25.82) |
|
Controls |
600 |
281 (46.83) |
271 (45.17) |
48 (8.00) |
833 (69.42) |
367 (30.58) |
|
rs2271261 |
Genotype |
T/T |
T/A |
A/A |
T |
A |
|
CHD |
300 |
223 (74.33) |
69 (23.00) |
8 (2.67) |
515 (85.83) |
85 (14.17) |
|
ASD |
109 |
73 (66.97) |
35 (32.11) |
1 (0.92) |
181 (83.03) |
37 (16.97) |
|
VSD |
122 |
88 (72.13) |
29 (23.78) |
5 (4.10) |
205 (84.02) |
39 (15.98) |
|
Controls |
600 |
460 (76.67) |
133 (22.17) |
7 (1.17) |
1053 (87.75) |
147 (12.25) |
|
rs2885987 |
Genotype |
T/T |
T/G |
G/G |
T |
G |
|
CHD |
300 |
230 (76.67) |
67 (22.33) |
3 (1.00) |
527 (87.83) |
73 (12.17) |
|
ASD |
109 |
80 (73.39) |
28 (25.69) |
1 (0.92) |
188 (86.24) |
30 (13.76) |
|
VSD |
122 |
99 (81.15) |
21 (17.21) |
2 (1.64) |
219 (89.75) |
25 (10.25) |
|
Controls |
600 |
478 (79.67) |
111 (18.50) |
11 (1.83) |
1067 (88.92) |
133 (11.08) |
|
Table 3: Associations of rs2459976, rs2271261 and rs2885987 variants in ZW10 with risk of CHD in Chinese Han populations
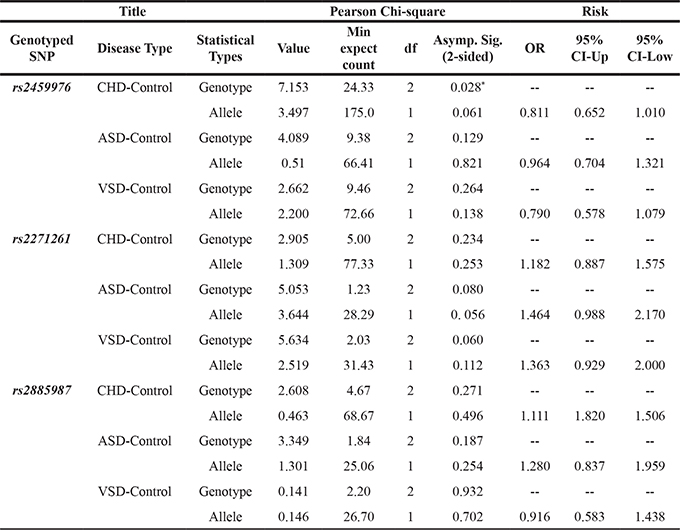
*:Statistically significant
Table 4: The CHD and controls groups were in line with Hardy-Weinberg equilibrium
Group |
Genotype |
H-W equilibrium Testing |
|||
|---|---|---|---|---|---|
SNP |
Test |
Homo/Hetero/Homozygote |
0 (HET) |
E (HET) |
P |
rs2459976 |
All |
73/379/448 |
0.4211 |
0.4132 |
0.6283 |
Affected |
25/108/167 |
0.3600 |
0.3880 |
0.2331 |
|
Unaffected |
48/271/281 |
0.4517 |
0.4246 |
0.1488 |
|
rs2271261 |
All |
15/202/683 |
0.2244 |
0.2246 |
1.0000 |
Affected |
8/69/223 |
0.2300 |
0.2432 |
0.3417 |
|
Unaffected |
7/133/460 |
0.2217 |
0.2150 |
0.5694 |
|
rs2885987 |
All |
14/178/708 |
0.1978 |
0.2027 |
0.5087 |
Affected |
3/67/230 |
0.2233 |
0.2137 |
0.5927 |
|
Unaffected |
11/111/478 |
0.1850 |
0.1971 |
0.1439 |
|
The experiment-wide significance threshold of the variant rs2459976 in ZW10 gene was 0.026. The Haploview software was used to conduct LD analysis of the variants rs2885987, rs2271261 and rs2459976, and the results were consistent with the data from the HapMap CHB population (Figure 2). The genotype frequencies in the CHD and control groups were further analyzed by three genetic models, including trend, dominant and recessive models, in addition to chi-square and fisher tests (Table 5). All results indicated that the variant rs2459976 was associated with the risk of CHD.
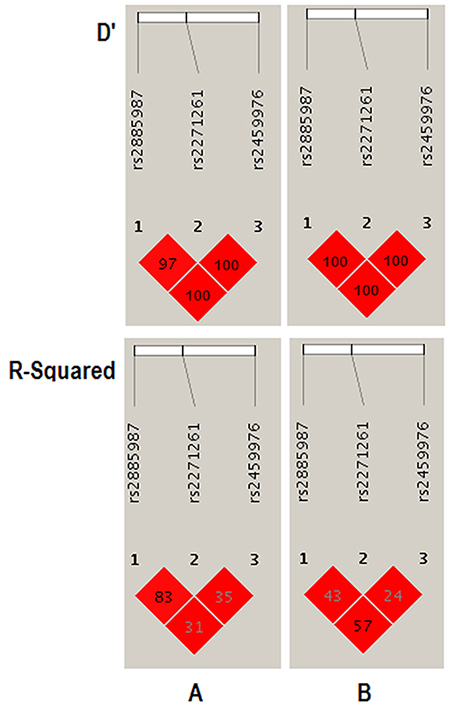
Figure 2: LD analysis of the variants rs2885987, rs2271261 and rs2459976 in the ZW10 gene. The LD plots were generated using the Haploview software v4.2. (A) Data analysis between CHD patients and controls from the present study of variants in ZW10 gene; (B) Data from HapMap CHB of variants in ZW10 gene. The data from the HapMap CHB and this work were consistent.
Table 5: SNP rs2459976 within ZW10 gene associated with the risk of congenital heart diseases in Chinese Han populations
Disease Type |
Value |
Trend model |
Dominant model |
Recessive model |
|---|---|---|---|---|
CHD-Control |
ChisQ |
3.565 |
6.242 |
0.030 |
P |
0.0590 |
0.0125* |
0.8629 |
|
ASD-Control |
ChisQ |
0.053 |
1.102 |
1.809 |
P |
0.8181 |
0.2937 |
0.1787 |
|
VSD-Control |
ChisQ |
2.324 |
2.654 |
0.295 |
P |
0.1274 |
0.1033 |
0.5871 |
*:Statistically significant.
DISCUSSION
In this study, we analyzed the transcribed regions and splicing sites of the kinetochore protein gene ZW10 in a large cohort of CHD patients and controls, finding that the variant rs2459976 in the ZW10 gene was associated with the risk of CHD in the Chinese Han population. Further work is needed to determine whether the kinetochore protein ZW10 may be involved in the CHD etiology.
The mammalian heart is a complex organ and its formation involves many genes with strict temporal, spatial and sequential expression [3]. Additionally, there are several cell types with distinct lineage origins in the mammalian heart [10, 11], interactions of which are necessary in the process of the cardiac development [1]. In previous studies, we found that variants in the STX18 and MSX1 genes are associated with CHDs, probably due to the regulating effects of their protein products on cell cycle [14, 15]. STX18 also transports proteins between the cellular endoplasmic reticulum (ER) and Golgi [27]. On the other hand, MSX1 is an active gene in the HES cells: MSX1 is up-expressed when the HES cells are co-cultured with PA6 cells [28] and is lower expressed when the HES cells are treated with dopamine [29]. In the embryonic development, HES cells differentiate to various cell types including cardiomyocytes in the mesoderm [9]. These findings suggest that any defects in highly expressed genes or factors in embryonic cells, such as HES cardiac progenitor cells, may affect the development of the heart.
The kinetochore protein ZW10 is expressed consistently through out the cell cycle in the cardiac progenitor cells during embryonic development [30]. It co-localizes with the ER, participates in the ER and Golgi trafficking, takes part in the mitotic checkpoint and interacts with the dynactin subunit dynamitin [20, 24]. Additionally, ZW10 associates with specific interphase structures and regulates transport movement of Golgi, endosome, and lysosome [31]. In the present work, we found that the variant rs2459976 in the ZW10 gene was associated with the risk of CHD.
Inmembrane budding and fusion, the N-terminal fragment of ZW10 interacts with STX18 and participates in STX18 mediated membrane sorting [24].The C-terminal end of ZW10 contains part of the dynamitin-interacting region, by which ZW10 anchors dynein to membranous organelles [31]. Whereas the C-terminal region of ZW10 exhibits weak interactions with dynamitin [32], the N-terminal region of ZW10 interacts with dynamitinin in amutually exclusive manner [32, 33]. The variant rs2459976 is located within the region between exons 1 and 2 of the ZW10 gene, which encodes the N-terminal region of ZW10 protein. The locations of the variants rs2271261 and rs2885987 are far from the N-terminal region of ZW10 protein, and we did not find any statistical significance between the two variants and CHD.
The 3′UTR and 5′UTR regions contain functional sequences that regulate the expression of many genes [34, 35] and may be associated with disease pathogenesis [36–39]. In a previous study, we identified variants near or within the 3′UTR or 5′UTR of genes that are associated with the risk of CHD [14, 40]. All these previous findings demonstrate the important roles of 5′UTR and 3′UTR on the functions of genes and their involvement in diseases. Although results in the current study support the possible involvement of the kinetochore protein gene ZW10 in the etiology of CHD, further validation is necessary to distinguish its roles in the CHD pathogenesis from associations just as a by stander.
In conclusion, we compared the ZW10 gene sequences between 300 Chinese Han CHD patients and 600 controls, and revealed a correlation between rs2459976 and the risk of CHD in the Chinese Han population.
MATERIALS AND METHODS
The study populations
We gathered 300 CHD patients and 600 normal controls for this study, from the Second and the Fourth Affiliated Hospitals of Harbin Medical University, Harbin, China; specimens of 260 of the 300 CHD patients were overlapping with those used for another study [3, 40]. Although we collected the cohort from two hospitals, the patients and controls were ethnically matched and their relatives were excluded from this study. All the CHD patients and normal controls were given comprehensive physical, electrocardiogram and ultrasonic echocardiogram examinations. None of the patients showed any other cardiac or systemic abnormalities, and the normal controls did not show any defects in the heart or other body parts. For this work, we obtained a written informed consent from each participant or their parents on behalf of minors, and the Ethics Committee of the Harbin Medical University approved this work. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
DNA analysis
Genomic DNA was extracted from the peripheral blood leukocytes of the participants as detailed in previous studies [15, 41]. The human ZW10 gene is located on 11q23.2. To determine the SNP genotypes in the gene, we amplified the exons and splicing sites, 3′UTR and 5′UTR of the ZW10 gene using the polymerase chain reaction (PCR) method (Supplementary Table 1), and sequenced the products by the Sanger technique using standard protocols [42]. After that, the genotypes were determined using PCR and gene sequencing methods [3, 43].
SNP genotyping analysis
We determined genotypes of rs2885987, rs11605483, rs2271261, rs2271796 and rs2459976 in the ZW10 gene (Figure 1) on 300 CHD patients and 600 normal controls, and two independent researchers conducted the measurements. The overall CHD statistical analysis was conducted according to the types of CHD and sample sizes.
Statistical analysis
The statistical analyses and Hardy-Weinberg equilibrium tests of the CHD and control populations were conducted by the software and methods as previously reported [15]. The experiment-wide significance threshold, matrix of linkage-disequilibrium (LD) correlation for the markers and haplotype diagram of LD structure were calculated using the online software SNPSpD (http://neurogenetics.qimrberghofer.edu.au/SNPSpD/) [44] and Haploview software (http://www.broadinstitute.org/scientific-community/science/programs/medical-and-population-genetics/haploview/haploview) [45], respectively.
Patient consent
Informed consent was obtained from all individual participants included in the study.
Ethics approval
This work has been approved by the Ethics Committee of Harbin Medical University.
Abbreviations
CHD, Congenital heart disease; ASD, atrial septal defect; VSD, ventricular septal defect; HES, human embryonic stem; LD, linkage-disequilibrium.
Author contributions
Conceptualization: FF L, SL L; methodology: FF L, SL L, CY S, JX Z, XQ L; software: FF L, SL L, S C; formal analysis: FF L, SL L, CY S,S C; investigation: CY S, S C, R C, S S, Y H; resources: CY S, S C, R C, S S, Y H, JX Z, XQ L; data curation: FF L, SL L, JX Z, XQ L; writing: FF L, SL L; funding acquisition: FF L, SL L, CY S.
ACKNOWLEDGMENTS
The authors thank the patients and their families for cooperation and participation in this study, and the physicians for collecting the specimens and conducting clinical examinations.
CONFLICTS OF INTEREST
All the authors have declared that no competing interests exist.
FUNDING
This work was supported by grants from Chinese Postdoctoral Science Foundation (2016M601460) and Health and Family Planning Commission of HeilongjiangprovinceFoundation(2016-189) to FFL, grants of National Natural Science Foundation of China to SLL (NSFC81271786, 81030029, 81671980),and Heilongjiang Postdoctoral Fund (LBH-Z16160), Beijing Municipal Science and technology project (Z161100000116090). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
REFERENCES
1. Hoffman JI, Kaplan S. The incidence of congenital heart disease. J Am Coll Cardiol. 2002; 39:1890–900.
2. Hoffman JI, Kaplan S, Liberthson RR. Prevalence of congenital heart disease. Am Heart J. 2004; 147:425–39.
3. Deng X, Zhou J, Li FF, Yan P, Zhao EY, Hao L, Yu KJ, Liu SL. Characterization of Nodal/TGF-Lefty Signaling Pathway Gene Variants for Possible Roles in Congenital Heart Diseases. PLoS One. 2014; 9:e104535.
4. Edwards JE. Survey of operative congenital heart disease. A review. Am J Pathol. 1976; 82:407–36.
5. van der Bom T, Zomer AC, Zwinderman AH, Meijboom FJ, Bouma BJ, Mulder BJ. The changing epidemiology of congenital heart disease. Nat Rev Cardiol. 2011; 8:50–60.
6. Bruneau BG. The developmental genetics of congenital heart disease. Nature. 2008; 451:943–8.
7. Richards AA, Garg V. Genetics of congenital heart disease. Curr Cardiol Rev. 2010; 6:91–7.
8. Pierpont ME, Basson CT, Benson DW Jr, Gelb BD, Giglia TM, Goldmuntz E, McGee G, Sable CA, Srivastava D, Webb CL. Genetic basis for congenital heart defects: current knowledge: a scientific statement from the American Heart Association Congenital Cardiac Defects Committee, Council on Cardiovascular Disease in the Young: endorsed by the American Academy of Pediatrics. Circulation. 2007; 115:3015–38.
9. Schuldiner M, Benvenisty N. Factors controlling human embryonic stem cell differentiation. Methods Enzymol. 2003; 365:446–61.
10. Chen YH, Ishii M, Sun J, Sucov HM, Maxson RE Jr. Msx1 and Msx2 regulate survival of secondary heart field precursors and post-migratory proliferation of cardiac neural crest in the outflow tract. Dev Biol. 2007; 308:421–37.
11. Cheng G, Litchenberg WH, Cole GJ, Mikawa T, Thompson RP, Gourdie RG. Development of the cardiac conduction system involves recruitment within a multipotent cardiomyogenic lineage. Development. 1999; 126:5041–9.
12. Han J, Ishii M, Bringas P Jr, Maas RL, Maxson RE Jr, Chai Y. Concerted action of Msx1 and Msx2 in regulating cranial neural crest cell differentiation during frontal bone development. Mech Dev. 2007; 124:729–45.
13. Ishii M, Han J, Yen HY, Sucov HM, Chai Y, Maxson RE Jr. Combined deficiencies of Msx1 and Msx2 cause impaired patterning and survival of the cranial neural crest. Development. 2005; 132:4937–50.
14. Li FF, Han Y, Shi S, Li X, Zhu XD, Zhou J, Shao QL, Li XQ, Liu SL. Characterization of Transcriptional Repressor Gene MSX1 Variations for Possible Associations with Congenital Heart Diseases. PLoS One. 2015; 10:e0142666.
15. Li X, Shi S, Li FF, Cheng R, Han Y, Diao LW, Zhang Q, Zhi JX, Liu SL. Characterization of soluble N-ethylmaleimide-sensitive factor attachment protein receptor gene STX18 variations for possible roles in congenital heart diseases. Gene. 2017; 598:79–83.
16. Hong W. SNAREs and traffic. Biochim Biophys Acta. 2005; 1744:493–517.
17. Bassett T, Harpur B, Poon HY, Kuo KH, Lee CH. Effective stimulation of growth in MCF-7 human breast cancer cells by inhibition of syntaxin18 by external guide sequence and ribonuclease P. Cancer Lett. 2008; 272:167–75.
18. Famulski JK, Vos L, Sun X, Chan G. Stable hZW10 kinetochore residency, mediated by hZwint-1 interaction, is essential for the mitotic checkpoint. J Cell Biol. 2008; 180:507–20.
19. Arasaki K, Uemura T, Tani K, Tagaya M. Correlation of Golgi localization of ZW10 and centrosomal accumulation of dynactin. Biochem Biophys Res Commun. 2007; 359:811–6.
20. Karess R. Rod-Zw10-Zwilch: a key player in the spindle checkpoint. Trends Cell Biol. 2005; 15:386–92.
21. Williams BC, Karr TL, Montgomery JM, Goldberg ML. The Drosophila l(1)zw10 gene product, required for accurate mitotic chromosome segregation, is redistributed at anaphase onset. J Cell Biol. 1992; 118:759–73.
22. Chan GK, Jablonski SA, Starr DA, Goldberg ML, Yen TJ. Human Zw10 and ROD are mitotic checkpoint proteins that bind to kinetochores. Nat Cell Biol. 2000; 2:944–7.
23. Vallee RB, Varma D, Dujardin DL. ZW10 function in mitotic checkpoint control, dynein targeting and membrane trafficking: is dynein the unifying theme? Cell Cycle. 2006; 5:2447–51.
24. Hirose H, Arasaki K, Dohmae N, Takio K, Hatsuzawa K, Nagahama M, Tani K, Yamamoto A, Tohyama M, Tagaya M. Implication of ZW10 in membrane trafficking between the endoplasmic reticulum and Golgi. EMBO J. 2004; 23:1267–78.
25. van Driel LM, Smedts HP, Helbing WA, Isaacs A, Lindemans J, Uitterlinden AG, van Duijn CM, de Vries JH, Steegers EA, Steegers-Theunissen RP. Eight-fold increased risk for congenital heart defects in children carrying the nicotinamide N-methyltransferase polymorphism and exposed to medicines and low nicotinamide. Eur Heart J. 2008; 29:1424–31.
26. Kebed KY, Bishu K, Al Adham RI, Baddour LM, Connolly HM, Sohail MR, Steckelberg JM, Wilson WR, Murad MH, Anavekar NS. Pregnancy and Postpartum Infective Endocarditis: A Systematic Review. Mayo Clin Proc. 2014; 89:1143–52.
27. Hay JC, Chao DS, Kuo CS, Scheller RH. Protein interactions regulating vesicle transport between the endoplasmic reticulum and Golgi apparatus in mammalian cells. Cell. 1997; 89:149–58.
28. Chan AA, Hertsenberg AJ, Funderburgh ML, Mann MM, Du Y, Davoli KA, Mich-Basso JD, Yang L, Funderburgh JL. Differentiation of human embryonic stem cells into cells with corneal keratocyte phenotype. PLoS One. 2013; 8:e56831.
29. Belinsky GS, Sirois CL, Rich MT, Short SM, Moore AR, Gilbert SE, Antic SD. Dopamine receptors in human embryonic stem cell neurodifferentiation. Stem Cells Dev. 2013; 22:1522–40.
30. Starr DA, Williams BC, Li Z, Etemad-Moghadam B, Dawe RK, Goldberg ML. Conservation of the centromere/kinetochore protein ZW10. J Cell Biol. 1997; 138:1289–301.
31. Varma D, Dujardin DL, Stehman SA, Vallee RB. Role of the kinetochore/cell cycle checkpoint protein ZW10 in interphase cytoplasmic dynein function. J Cell Biol. 2006; 172:655–62.
32. Inoue M, Arasaki K, Ueda A, Aoki T, Tagaya M. N-terminal region of ZW10 serves not only as a determinant for localization but also as a link with dynein function. Genes Cells. 2008; 13:905–14.
33. Arasaki K, Taniguchi M, Tani K, Tagaya M. RINT-1 regulates the localization and entry of ZW10 to the syntaxin 18 complex. Mol Biol Cell. 2006; 17:2780–8.
34. Lou F, Ma HN, Xu L, Chen M, Zhu YB. Two polymorphisms of CD44 3′UTR weaken the binding of miRNAs and associate with naso-pharyngeal carcinoma in a Chinese population. Eur Rev Med Pharmacol Sci. 2014; 18:2444–52.
35. Fujimuro T, Matsui T, Nitanda Y, Matta T, Sakumura Y, Saito M, Kohno K, Nakahata Y, Bessho Y. Hes7 3′UTR is required for somite segmentation function. Sci Rep. 2014; 4:6462.
36. Chen S, He Y, Ding J, Jiang Y, Jia S, Xia W, Zhao J, Lu M, Gu Z, Gao Y. An insertion/deletion polymorphism in the 3′ untranslated region of beta-transducin repeat-containing protein (betaTrCP) is associated with susceptibility for hepatocellular carcinoma in Chinese. Biochem Biophys Res Commun. 2010; 391:552–6.
37. Christensen BC, Moyer BJ, Avissar M, Ouellet LG, Plaza SL, McClean MD, Marsit CJ, Kelsey KT. A let-7 microRNA-binding site polymorphism in the KRAS 3′ UTR is associated with reduced survival in oral cancers. Carcinogenesis. 2009; 30:1003–7.
38. Tosetto E, Casarin A, Salviati L, Familiari A, Lieske JC, Anglani F. Complexity of the 5′UTR region of the CLCN5 gene: eleven 5′UTR ends are differentially expressed in the human kidney. BMC Med Genomics. 2014; 7:41.
39. Orom UA, Nielsen FC, Lund AH. MicroRNA-10a binds the 5′UTR of ribosomal protein mRNAs and enhances their translation. Mol Cell. 2008; 30:460–71.
40. Li FF, Zhou J, Zhao DD, Yan P, Li X, Han Y, Li XS, Wang GY, Yu KJ, Liu SL. Characterization of SMAD3 Gene Variants for Possible Roles in Ventricular Septal Defects and Other Congenital Heart Diseases. PLoS One. 2015; 10:e0131542.
41. Li FF, Deng X, Zhou J, Yan P, Zhao EY, Liu SL. Characterization of human bone morphogenetic protein gene variants for possible roles in congenital heart disease. Mol Med Rep. 2016; 14:1459–64.
42. Tan ZX, Li FF, Qu YY, Liu J, Liu GR, Zhou J, Zhu YL, Liu SL. Identification of a known mutation in Notch 3 in familiar CADASIL in China. PLoS One. 2012; 7:e36590.
43. Li FF, Yan P, Zhao ZX, Liu Z, Song DW, Zhao XW, Wang XS, Wang GY, Liu SL. Polymorphisms in the CHIT1 gene: Associations with colorectal cancer. Oncotarget. 2016; 7:39572–81. https://doi.org/10.18632/oncotarget.91389138.
44. Nyholt DR. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am J Hum Genet. 2004; 74:765–9.
45. Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005; 21:263–5.

