INTRODUCTION
The mammalian transcription factor FoxM1 is dominantly overexpressed and plays critical role in tumorigenesis, proliferation, and metastasis, as well as drug resistance in a broad range of human cancer types, such as lung, gastric, and breast cancers [1, 2].
A plenty of studies, including an insight computational analysis, have demonstrated that elevated FoxM1 expression is a major predictor of adverse outcomes across a variety of human malignancies, indicating the oncogenic activity of FoxM1 in cancer. Recently, it was identified that FoxM1 acts as a critical regulator of mammary differentiation with significant implications for the development of aggressive breast cancers [3]. Elevated expression of FoxM1 in breast cancer correlates with an undifferentiated tumor phenotype and a negative clinical outcome. Abdeljaoued, S. and colleagues also reported that overexpression of FoxM1 is an adverse prognostic factor in male breast cancer [4].
More interestingly, accumulating evidences indicated that FoxM1 was significantly involved in drug resistance that compromised the efficacy of transtuzumab, tamoxifen and taxanes in the treatment of breast cancer. Furthermore, FoxM1-targeted therapy could effectively restrain tumor development of cancer [4–7]. There is increasing awareness that development of FoxM1 inhibitor is a promising strategy for breast cancer therapy.
Nevertheless, viewpoints on the role of FoxM1 in breast cancer were mostly from basic studies, and lacking support from clinical data. In the current study, we carried out a data-mining process in a variety of public databases with clinical information to evaluate the potent function and prognostic value of FoxM1 expression in breast cancer, with attempt of providing informative clues for future development of FoxM1-targeted therapy and prognostic prediction in breast cancer.
RESULTS
FoxM1was significantly overexpressed in breast cancer comparing with normal breast tissue
Hitherto, expression of FoxM1 had been identified in a number of human cancers, including hematological malignancies and solid tumors (Figure 1A). ONCOMINE analysis revealed that FoxM1 mRNA expression was significantly higher in a wide variety of datasets in different cancer types than corresponding normal samples, especially in sarcoma, lung cancer and breast cancer.
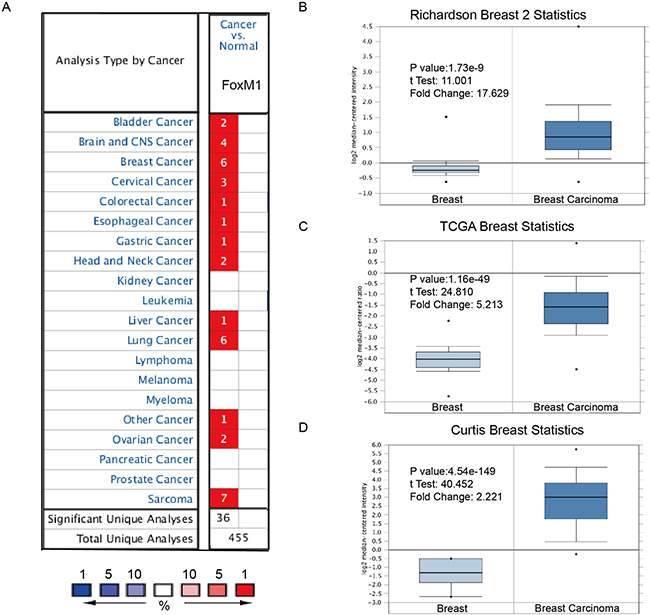
Figure 1: FoxM1 was significantly overexpressed in breast cancer comparing with normal breast tissue. (A) FoxM1 mRNA expressions (cancer vs. normal tissue) analyzed with ONCOMINE database. The graphic demonstrated the numbers of datasets with statistically significant mRNA over-expression (red) or down-expression (blue) of the target gene. The p value threshold is 0.01. The number in each cell represents the number of analyses that meet the threshold within those analysis and cancer types. The gene rank was analyzed by percentile of target gene in the top of all genes measured in each research. Cell color is determined by the best gene rank percentile for the analyses within the cell. (B–D) Comparison of FoxM1 mRNA expression in Richardson’s study (B), TCGA breast statistics (C), and Curtis’s study (D). Box plots derived from gene expression data in ONCOMINE comparing expression of a specific GATA family member in normal and BC tissue. The p value was set up at 0.01 and fold change was defined as 2.
In a dataset from Richardson’s study [5], FoxM1 was 17.629-fold elevated in breast cancer samples as compared with normal tissues (p = 1.73e-9) (Figure 1B). Another dataset with 593 samples that derived from the Cancer Genome Atlas (TCGA) database showed that FoxM1 transcripts were 5.213-fold elevated in breast cancer samples as compared with normal tissues (Figure 1C). Consistently, in another dataset from Curtis’s study [6], FoxM1 was 2.221-fold increase in cancer VS. normal samples (p = 0.001).
Expression of FoxM1 was distinct in different molecular subtypes of BC patients
In bc-GenExMiner, the Welch’s test was performed to compare the mRNA expression of FoxM1 between groups of patients, according to different clinicopathological parameters. Higher FoxM1 mRNA levels were found in BC patients with ER (−) than ER (+) (Figure 2A). Similarly, increasing expression of FoxM1 mRNA was found in BC patients with PR (−) than PR (+) (Figure 2B). However, there was no significantly different between BC patients with HER-2 (+) and HER-2 (−) (Figure 2C).
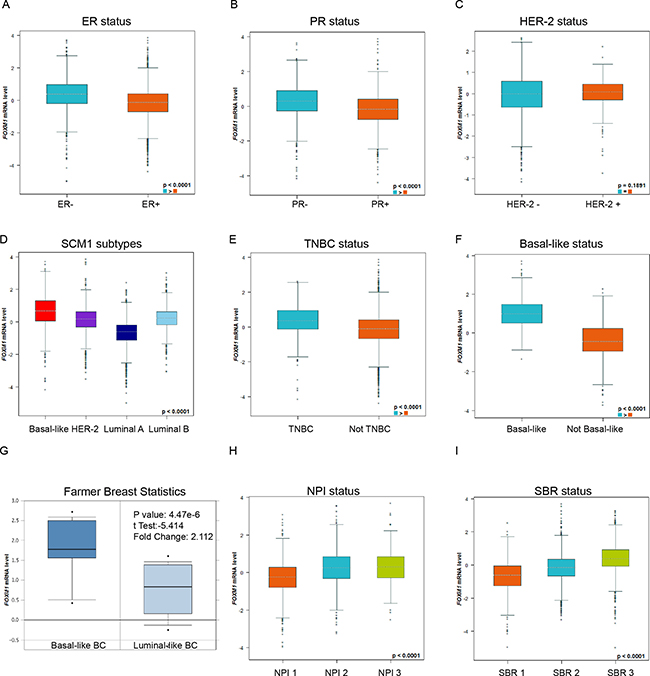
Figure 2: Expression of FoxM1 was distinct in different molecular subtypes of BC patients. Global significant different between groups was assessed by Welch’s test to generate p value, along with Dunnett-Tukey-Kramer’s tests for pairwise comparison when a global significant difference exists (p < 0.05). (A) The mRNA expression level of FoxM1 in BC patients with ER (−) and ER (+). (B) The mRNA expression level of FoxM1 in BC patients with PR (−) and PR (+). (C) The mRNA expression level of FoxM1 in BC patients with HER-2 (−) and HER-2(+). (D) The mRNA expression level of FoxM1 in a variety of breast cancer subtype. (E) The mRNA expression level of FoxM1 in TNBC or NOT TNBC patients. (F) The mRNA expression level of FoxM1 in Basal-like or NOT Basal-like breast cancer subtype. (G) The mRNA expression level of FoxM1 in Basal-like or Luminal-like breast cancer subtype. (H) The mRNA expression level of FoxM1 in different Nottingham Prognostic Index(NPI) BC subgroups. (I) The mRNA expression level of FoxM1 in different Scarff Bloom & Richardson grade status (SBR) BC subgroups.
For molecular subtypes analysis, the expression of FoxM1 in Basal-like and HER-2 subtypes was significantly higher than Luminal A and Luminal B subtypes of BC (Dunnett-Tukey-Kramer’s Tests, p < 0.0001) (Figure 2D, all of the groups comparison were showed in Supplementary Table 1). Consistently, higher FoxM1 mRNA levels were found in Basal-like or Triple-negative breast cancer (TNBC) patients (Figure 2E–2F). Another dataset from Farmer’s study [7] also showed that FoxM1 was 2.112-fold increase in Basal-like BC VS. Luminal-Like BC samples (p = 4.47e-6) (Figure 2G).
In Nottingham Prognostic Index (NPI), higher NPI level was associated with the enriched mRNA level of FoxM1 (Dunnett-Tukey-Kramer’s Tests, p < 0.0001) (Figure 2H, all of the group comparisons were showed in Supplementary Table 2). In Scarff Bloom & Richardson grade status (SBR) criterion, more advanced SBR grade was relevant to higher mRNA level of FoxM1 (Dunnett-Tukey-Kramer’s Tests, p < 0.0001) (Figure 2H, all of the group comparisons were showed in Supplementary Table 3).
Higher expression of FoxM1 correlated with low expression of ESR1, ERBB2, FoxA1 and GATA3
In bc-GenExMiner, the Welch’s test was performed to compare the mRNA expression of FoxM1 between groups of patients, according to mRNA expression of ESR1, ERBB2, and typical Luminal epithelial biomarkers FoxA1 and GATA3. Gene correlation targeted analysis showed that higher expression of FoxM1 correlated with low expression of ESR1 (Figure 3A–3B, r = −0.21, p < 0.001), ERBB2 (Figure 3C–3D, r = −0.03, p = 0.0238), FoxA1 (Figure 3E–3F, r = −0.32, p < 0.001) and GATA3 (Figure 3G–3H, r = –0.31, p < 0.001). Correlation map for all patients among FoxM1, ESR1, ERBB2, FoxA1 and GATA3 were showed (Figure 3I).
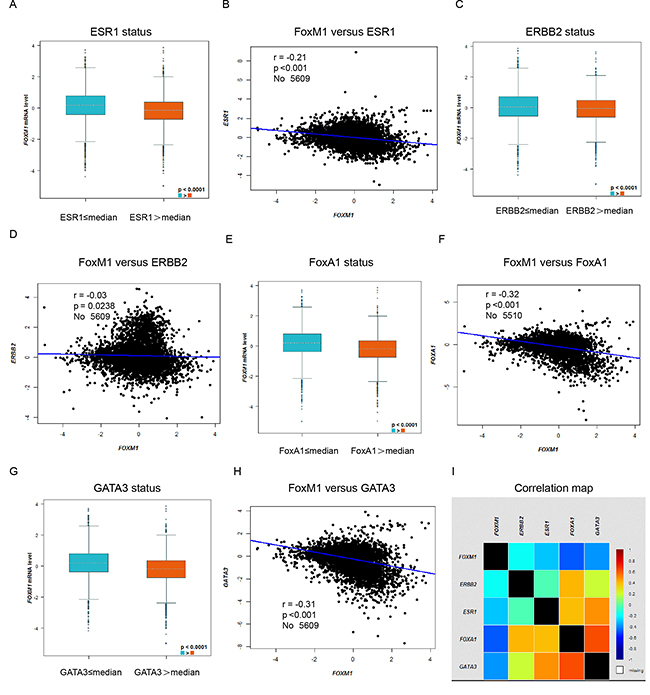
Figure 3: Higher expression of FoxM1 correlated with low expression of ESR1, ERBB2, FoxA1 and GATA3. In bc-GenExMiner, the Welch’s test was performed to compare the mRNA expression of FoxM1 between groups of patients, according to mRNA expression of different genes. (A) The mRNA expression level of FoxM1 according to mRNA expression level of ESR1. (B) Gene correlation targeted analysis between FoxM1 and ESR1. (C) The mRNA expression level of FoxM1 according to mRNA expression level of ERBB2. (D) Gene correlation targeted analysis between FoxM1 and ERBB2. (E) The mRNA expression level of FoxM1 according to mRNA expression level of FoxA1. (F) Gene correlation targeted analysis between FoxM1 and FoxA1. (G) The mRNA expression level of FoxM1 according to mRNA expression level of GATA3. (H) Gene correlation targeted analysis between FoxM1 and GATA3. (I) Correlation map for all patients among FoxM1, ESR1, ERBB2, FoxA1 and GATA3.
Elevated FoxM1 expression predicted shorter DMFS in BC patients, especially in the ER positive and Luminal A, Luminal B subtypes of BC patients
We next assessed the prognostic value of FoxM1 for distant metastasis in patients with BC. Analysis from bc-GenExMiner showed that FoxM1 mRNA high expression was associated with shorter DMFS in all BC patients (HR = 1.86, p < 0.0001) (Figure 4A). Sub-analysis indicated that FoxM1 mRNA high expression was correlated to shorter DMFS in BC patients with ER positive tumors (HR = 2.19, p < 0.0001) (Figure 4B), but not in ER negative tumors (HR = 0.98, p = 0.8623) (Figure 4C).
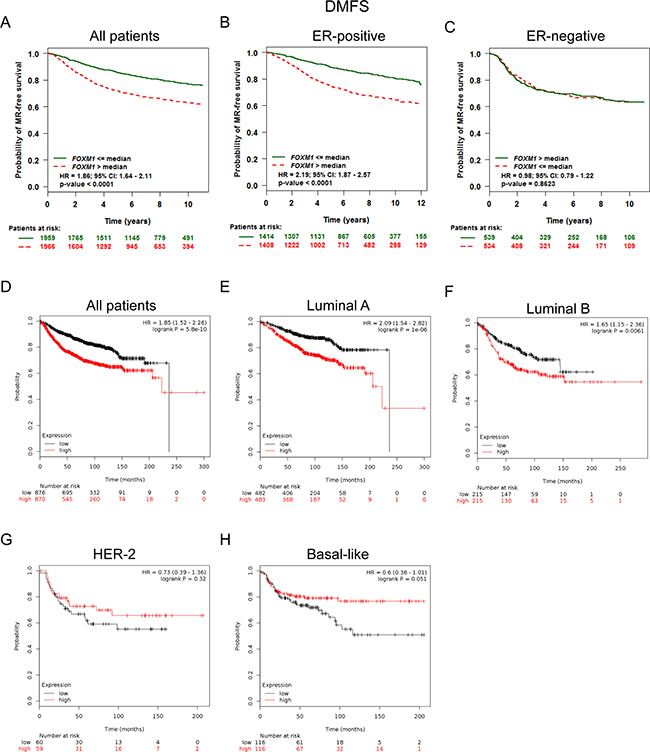
Figure 4: Elevated FoxM1 expression predicted shorter DMFS in BC patients, especially in the ER positive and Luminal A, Luminal B subtypes of BC patients. (A) Analysis from bc-GenExMiner showed that FoxM1 mRNA expression was associated with DMFS in all BC patients (B–C) FoxM1 mRNA expression was correlated to DMFS in BC patients with ER positive tumors, but not in ER negative tumors. (D) The Kaplan-Meier plotter survival analysis showed that FoxM1 mRNA expression was correlated to DMFS in all BC patients. (E–H) The Kaplan-Meier plotter survival analysis showed that FoxM1 mRNA expression was correlated to DMFS in different subtypes of BC patients.
The Kaplan-Meier plotter survival analysis also showed that FoxM1 mRNA high expression was correlated to shorter DMFS in all BC patients (HR = 1.85, p = 5.8e–10) (Figure 4D), specifically in BC patients with Luminal A tumors (HR = 2.09, p = 1.0e–6) and Luminal B tumors (HR = 1.65, p = 0.0061) (Figure 4E–4F). However, there was no significant difference in DMFS in BC patients either with HER-2 positive (HR = 0.73, p = 0.32) tumors or Basal-like (HR = 0.6, p = 0.051) between high and low FoxM1 mRNA expression (Figure 4G–4H).
Elevated FoxM1 expression predicted poor survival in breast cancer patients, especially in the ER (+), PR (+) subgroups and BC patients received adjuvant chemotherapy only or treated with tamoxifen only
FoxM1 mRNA high expression was significantly correlated with shorter RFS in all BC patients (HR = 1.67, p < 1.0e-16) (Figure 5A). FoxM1 mRNA high expression was correlated to shorter RFS in BC patients with ER positive tumors (HR = 1.9, p = 3.1e-13) (Figure 5B), and PR positive tumors (HR = 2.66, p = 1.3e-07) (Figure 5C).
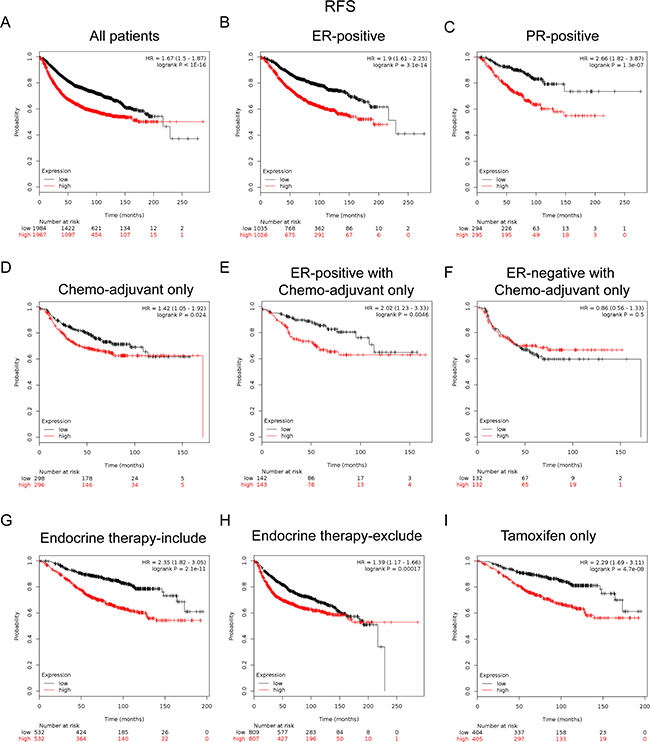
Figure 5: Elevated FoxM1 expression predicted poor survival in breast cancer patients, especially in the ER(+), PR(+) subgroups and BC patients received adjuvant chemotherapy only or treated with tamoxifen only. (A) FoxM1 mRNA level was significantly associated with RFS in all BC patients. (B–C) FoxM1 mRNA expression was correlated to RFS in BC patients with ER (+) and PR (+) tumors. (D) High mRNA level of FoxM1 was significantly associated with shorter RFS in BC patients who have received chemotherapy adjuvant only. (E–F) High mRNA level of FoxM1 was significantly associated with shorter RFS in ER (+) subgroup of BC patients who have received chemotherapy adjuvant only but not in ER (−) BC patients. (G–H) High mRNA level of FoxM1 was significantly associated with shorter RFS in BC patients who have received therapies that include or exclude endocrine therapy. (I) High mRNA level of FoxM1 was associated with shorter RFS in BC patients who have received tamoxifen only sub-group.
Notably, for the respond to treatments, the results demonstrated that FoxM1 high mRNA expression was significantly correlated to shorter RFS in patients who have received adjuvant chemotherapy only (HR = 1.42, p = 0.024) (Figure 5D), especially in the ER-positive subgroup (HR = 2.02, p = 0.0046) (Figure 5E), but not in the ER-negative subgroup (HR = 0.86, p = 0.5) (Figure 5F). These results indicated a potential role of FoxM1 in contribution to chemoresistance in BC, which may even involve in the expression of ER.
Of noteworthy, the results demonstrated that FoxM1 high mRNA expression was significantly correlated to shorter RFS in patients who have received treatments included endocrine therapy (HR = 2.35, p = 2.1e–11) (Figure 5G) or excluded endocrine therapy (HR = 1.39, p = 0.00017) (Figure 5H), especially in patients who treated with tamoxifen only (HR = 2.29, p = 4.7e-8) (Figure 5I), indicating a potential role of FoxM1 in contribution to tamoxifen resistance in breast cancer.
DISCUSSION
Discovery of therapeutic targets and development of novel targeted agents has changed the landscape in the battle field against breast cancer in the past few decades [8]. A variety of master genes, including ER, human epidermal growth factor receptor-2 (HER-2), and cyclin dependent kinase (CDK)4/6 have been identified to be involved in the development and progression of BC [9, 10]. Scientists have developed targeted agents against these genes or proteins that yielded successful results in the treatment of BC [11].
Myriad of studies have demonstrated that FoxM1 overexpressed in multiple cancers types [12–16], including breast cancer [4]. Recently, Li and colleagues provided a systematic review of prognostic value of FoxM1 in solid tumors, which demonstrated that elevated FoxM1 expression was associated with poor survival in most solid tumors. FoxM1 was a potential biomarker for prognosis prediction and a promising therapeutic target in human solid tumors [2]. Through analysis of a broad spectrum of pooled datasets, we found that FoxM1was significantly overexpressed in breast cancer comparing with normal breast tissue. Even more, the expression and prognostic value of FoxM1 in BC patients were highly associated with molecular subtypes and expressions of ER/PR, which imply the potentiality of FoxM1 acting as a master gene with important functions in breast cancer.
Several studies proposed that FoxM1 may promote distant metastasis through inducing of epithelial–mesenchymal transition (EMT) [17–19]. Xue and colleagues demonstrated that FoxM1 promoted typical EMT cellular pathway TGF-beta-dependent cancer metastasis via sustained activation of SMAD3/SMAD4 [20]. Wang and colleagues showed that increased FoxM1 expression is a target for metformin in the suppression of EMT in prostate cancer [21]. Yang and colleagues reported Slug to be the downstream target though which FoxM1 stimulating the processing of EMT [22]. Our data analysis agrees with those studies that elevated FoxM1 expression predicted shorter DMFS in BC patients, especially in patients with ER positive and Luminal A, Luminal B subtypes of BC. Also, our analysis on the correlation between FoxM1 and FoxA1, as well as GATA3, showed that FoxM1 gene was negatively associated with the classical luminal epithelial regulation gene GATA3 and FoxA1. These results indicated that FoxM1 was a negative regulator of breast epithelial phenotype. Therefore, FoxM1 might be a key driver and potential predictor of distant metastasis in BC patients.
More recently, increasing evidences have demonstrated a causal link between FoxM1 and chemoresistance. It has been reported that FoxM1expression associated with the chemotherapy resistance [23–29], especially in ER positive subgroup. Park and colleagues reported that FoxM1 mediates Dox resistance in breast cancer by enhancing DNA repair [30]. Khongkow and colleagues reported that paclitaxel targets FoxM1 to regulate KIF20A in mitotic catastrophe and dysfunction of FoxM1 expression will lead to breast cancer paclitaxel resistance [31]. Our analysis also indicated that elevated FoxM1 expression predicted worse survival in breast cancer patients with adjuvant chemotherapy only, particularly in the ER-positive subgroup. These results further confirmed the perspective that FoxM1 was a significant contributor of chemoresistance in breast cancer [32]. Although the exact mechanism are still unknown, some contributing reason could be self-explanatory. For example, the expression of FoxM1 in different breast cancer subtypes, as our data showed, was correlated to different expression patterns across different breast cancer subtypes. Moreover, the expression of FoxM1 was associated with tumor pathological grade. Particularly, high-level expressed in TNBC indicating that higher FoxM1 is an adverse prognostic factor and may involve in the chemoresistance of BC [33]. Evidence partly showed by our study suggested that FoxM1 was a pivotal driver of EMT, which was characterized with property of chemoresistance to BC. This perspective was supported by recent studies with results indicating that EMT contributes to chemoresistance of cancer [34, 35].
Of noteworthy, our results demonstrated that FoxM1 high mRNA expression was significantly correlated to shorter RFS in patients who have received endocrine therapy, especially in patients who treated with tamoxifen only, indicating a potential role of FoxM1 in contribution to tamoxifen resistance in breast cancer. Many studies revealed that high level of FoxM1 was associated with endocrine therapy sensitivity and resistance, especially tamoxifen resistance [36–39]. The possible reason is that the gene level FoxM1 is negatively related to the ESR1 expression, as our data showed, as well as negatively related to the ER expression in protein level. Genome-wide mapping of FoxM1 binding conducted by Sanders and colleagues also reveals co-binding of FoxM1 with estrogen receptor alpha in breast cancer cells [40]. Similarly, in our survival analysis, it was found that FoxM1 was associated with poor prognosis and early recurrence in patients with ER (+), PR (+), which implied that FoxM1 was actively involved in ER signaling pathway. This result agrees with the notion in the study by Millour and colleagues, proposing that FOXM1 is a transcriptional target of ER alpha and plays a critical role in breast cancer endocrine sensitivity and resistance [41].
A number of literatures reported that FoxM1was closely related to the expression of HER-2 [42, 43], and even some studies have documented that FoxM1has a direct control overHER-2 and may be the mediator of Herceptin resistance [44, 45]. Conversely, study by Francis, R and colleagues demonstrated that FoxM1was a downstream target and marker of HER-2 overexpression in breast cancer. However, our analysis showed that the correlation of FoxM1 and HER-2 expression remained uncertain. For one reason, the expression correlation coefficient r = –0.03 in the gene level of FoxM1 and ERBB2 gene is different from the perspective in an in vitro study by Kambach and colleagues [46]. Another reason is that the expression of FoxM1 in HER-2 (+) and HER-2 group (−) was not statistically difference. Finally, HER-2 subtypes in DMFS analysis of FoxM1 instead of the high expression of this seemingly contradictory phenomenon associated with good prognosis is warrant future research.
In conclusion, the oncogenic transcription factor FoxM1 is overexpressed in breast cancer versus normal controls. FoxM1 plays diverse roles in different molecular subtypes of breast cancer, which might be underlying the diverse mechanism of tumorigenesis and genetic background, as well as orchestrating with other cofactors in various tumor contexts. These evidences suggest that FoxM1 is an attractive prognostic prediction biomarker and promising therapeutic target for breast cancer.
MATERIALS AND METHODS
Oncomine analysis
The mRNA levels of FoxM1 in different type of cancers were determined through analysis in ONCOMINE database(www.oncomine.org), which is a publicly accessible online cancer microarray database to facilitate discovery from genome-wide expression analyses.
In this study, students’ t-test was used to generate a p-value for comparison between cancer specimens and normal control datasets. The fold change was defined as 2 and p value was set up at 0.01as described in our previous study [47]. Significant correlations can be found in an array of BC researches, as showed in typical figures.
Breast Cancer Gene-Expression Miner v4.0 analysis
Breast Cancer Gene-Expression Miner v4.0 (bcGenExMiner v4.0) consisted 36 annotated genomic datasets and three statistical mining functions [48, 49]. The expression module was added on March 29th, 2016, comparing the expression of a target gene according to clinical criteria, such as hormonal receptors, nodal status, and so on. The prognostic module assessed the prognostic impact of candidate genes in human BC and provided potential prognostic markers for BC. The correlation module. computed the correlation between genes or identified clusters of correlated co-expressed genes located in the same chromosomal region.
The Kaplan-Meier plotter survival analysis
Prognostic values of FoxM1was further assessed by displaying the relapse-free survival (RFS) and DMFS using the Kaplan-Meier plotter (www.kmplot.com). The log-rank p was calculated and shown on the webpage [50].
Author contributions
Xiao-Feng Lu, De Zeng and Hao-Yu Lin conceived and designed the project. Xiao-Feng Lu, De Zeng, Shu-Ming Sun, Chun-Fa Chen and Wei-Quan Liang performed data-mining process and prepared the figures and tables. Xiao-Feng Lu and De Zeng analyzed and interpreted the data. Xiao-Feng Lu, De Zeng and Hao-Yu Lin wrote the manuscript. Hao-Yu Lin approved the final version to be submitted.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
This study is partly supported by Medical Scientific Research Foundation of Guangdong Province, China (NO. A2016368), Natural Science Foundation of Guangdong Province, China (NO. 2015A030313429), Youth Innovative Talent Project of Colleges and Universities in Guangdong Province, China (NO. 2016KQNCX051), and National Natural Science Foundation of China (No. 81602345).
REFERENCES
1. Gartel AL. FOXM1 in Cancer: Interactions and Vulnerabilities. Cancer Res. 2017; 77:3135–3139.
2. Li L, Wu D, Yu Q, Li L, Wu P. Prognostic value of FOXM1 in solid tumors: a systematic review and meta-analysis. Oncotarget. 2017; 8:32298–308. https://doi.org/10.18632/oncotarget.15764.
3. Carr JR, Kiefer MM, Park HJ, Li J, Wang Z, Fontanarosa J, DeWaal D, Kopanja D, Benevolenskaya EV, Guzman G, Raychaudhuri P. FoxM1 regulates mammary luminal cell fate. Cell Reports. 2012; 1:715–729.
4. Abdeljaoued S, Bettaieb I, Nasri M, Adouni O, Goucha A, El Amine O, Boussen H, Rahal K, Gamoudi A. Overexpression of FOXM1 Is a Potential Prognostic Marker in Male Breast Cancer. Oncol Res Treat. 2017; 40:167–172.
5. Richardson AL, Wang ZC, De Nicolo A, Lu X, Brown M, Miron A, Liao X, Iglehart JD, Livingston DM, Ganesan S. X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell. 2006; 9:121–132.
6. Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Gräf S, Ha G, Haffari G, et al, and METABRIC Group. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012; 486:346–352.
7. Farmer P, Bonnefoi H, Becette V, Tubiana-Hulin M, Fumoleau P, Larsimont D, Macgrogan G, Bergh J, Cameron D, Goldstein D, Duss S, Nicoulaz AL, Brisken C, et al. Identification of molecular apocrine breast tumours by microarray analysis. Oncogene. 2005; 24:4660–4671.
8. Maruthanila VL, Elancheran R, Kunnumakkara AB, Kabilan S, Kotoky J. Recent development of targeted approaches for the treatment of breast cancer. Breast Cancer. 2017; 24:191–219.
9. Mayank, Jaitak V. Drug target strategies in breast cancer treatment: recent developments. Anticancer Agents Med Chem. 2014; 14:1414–27.
10. de la Vega M, Diaz-Canton E, Alvarez RH. Novel targeted agents for the treatment of advanced breast cancer. Future Med Chem. 2012; 4:893–914.
11. Zurrida S, Veronesi U. Milestones in breast cancer treatment. Breast J. 2015; 21:3–12.
12. Rinaldetti S, Wirtz RM, Worst TS, Eckstein M, Weiss CA, Breyer J, Otto W, Bolenz C, Hartmann A, Erben P. FOXM1 predicts overall and disease specific survival in muscle-invasive urothelial carcinoma and presents a differential expression between bladder cancer subtypes. Oncotarget. 2017; 8:47595–606. https://doi.org/10.18632/oncotarget.17394.
13. Ito T, Kohashi K, Yamada Y, Maekawa A, Kuda M, Furue M, Oda Y. Prognostic significance of forkhead box M1 (FoxM1) expression and antitumour effect of FoxM1 inhibition in melanoma. Histopathology. 2016; 69:63–71.
14. Okada K, Fujiwara Y, Takahashi T, Nakamura Y, Takiguchi S, Nakajima K, Miyata H, Yamasaki M, Kurokawa Y, Mori M, Doki Y. Overexpression of forkhead box M1 transcription factor (FOXM1) is a potential prognostic marker and enhances chemoresistance for docetaxel in gastric cancer. Ann Surg Oncol. 2013; 20:1035–1043.
15. Liu D, Zhang Z, Kong CZ. High FOXM1 expression was associated with bladder carcinogenesis. Tumour Biol. 2013; 34:1131–1138.
16. He SY, Shen HW, Xu L, Zhao XH, Yuan L, Niu G, You ZS, Yao SZ. FOXM1 promotes tumor cell invasion and correlates with poor prognosis in early-stage cervical cancer. Gynecol Oncol. 2012; 127:601–610.
17. Gottimukkala KP, Jangid R, Patta I, Sultana DA, Sharma A, Misra-Sen J, Galande S. Regulation of SATB1 during thymocyte development by TCR signaling. Mol Immunol. 2016; 77:34–43.
18. Arora R, Yates C, Gary BD, McClellan S, Tan M, Xi Y, Reed E, Piazza GA, Owen LB, Dean-Colomb W. Panepoxydone targets NF-kB and FOXM1 to inhibit proliferation, induce apoptosis and reverse epithelial to mesenchymal transition in breast cancer. PLoS One. 2014; 9:e98370.
19. Wang Z, Zhang S, Siu TL, Huang S. Glioblastoma multiforme formation and EMT: role of FoxM1 transcription factor. Curr Pharm Des. 2015; 21:1268–1271.
20. Xue J, Lin X, Chiu WT, Chen YH, Yu G, Liu M, Feng XH, Sawaya R, Medema RH, Hung MC, Huang S. Sustained activation of SMAD3/SMAD4 by FOXM1 promotes TGF-beta-dependent cancer metastasis. J Clin Invest. 2014; 124:564–579.
21. Wang Y, Yao B, Wang Y, Zhang M, Fu S, Gao H, Peng R, Zhang L, Tang J. Increased FoxM1 expression is a target for metformin in the suppression of EMT in prostate cancer. Int J Mol Med. 2014; 33:1514–1522.
22. Yang C, Chen H, Tan G, Gao W, Cheng L, Jiang X, Yu L, Tan Y. FOXM1 promotes the epithelial to mesenchymal transition by stimulating the transcription of Slug in human breast cancer. Cancer Lett. 2013; 340:104–112.
23. Karunarathna U, Kongsema M, Zona S, Gong C, Cabrera E, Gomes AR, Man EP, Khongkow P, Tsang JW, Khoo US, Medema RH, Freire R, Lam EW. OTUB1 inhibits the ubiquitination and degradation of FOXM1 in breast cancer and epirubicin resistance. Oncogene. 2016; 35:1433–1444.
24. Nestal de Moraes G, Delbue D, Silva KL, Robaina MC, Khongkow P, Gomes AR, Zona S, Crocamo S, Mencalha AL, Magalhaes LM, Lam EW, Maia RC. FOXM1 targets XIAP and Survivin to modulate breast cancer survival and chemoresistance. Cell Signal. 2015; 27:2496–2505.
25. Myatt SS, Kongsema M, Man CW, Kelly DJ, Gomes AR, Khongkow P, Karunarathna U, Zona S, Langer JK, Dunsby CW, Coombes RC, French PM, Brosens JJ, Lam EW. SUMOylation inhibits FOXM1 activity and delays mitotic transition. Oncogene. 2014; 33:4316–4329.
26. Kwok JM, Peck B, Monteiro LJ, Schwenen HD, Millour J, Coombes RC, Myatt SS, Lam EW. FOXM1 confers acquired cisplatin resistance in breast cancer cells. Mol Cancer Res. 2010; 8:24–34.
27. Rajamanickam S, Panneerdoss S, Gorthi A, Timilsina S, Onyeagucha B, Kovalskyy D, Ivanov D, Hanes MA, Vadlamudi RK, Chen Y, Bishop AJ, Arbiser JL, Rao MK. Inhibition of FoxM1-Mediated DNA Repair by Imipramine Blue Suppresses Breast Cancer Growth and Metastasis. Clin Cancer Res. 2016; 22:3524–3536.
28. Hamurcu Z, Ashour A, Kahraman N, Ozpolat B. FOXM1 regulates expression of eukaryotic elongation factor 2 kinase and promotes proliferation, invasion and tumorgenesis of human triple negative breast cancer cells. Oncotarget. 2016; 7:16619–35. https://doi.org/10.18632/oncotarget.7672.
29. Dai J, Yang L, Wang J, Xiao Y, Ruan Q. Prognostic Value of FOXM1 in Patients with Malignant Solid Tumor: A Meta-Analysis and System Review. Dis Markers. 2015; 2015:352478.
30. Park YY, Jung SY, Jennings NB, Rodriguez-Aguayo C, Peng G, Lee SR, Kim SB, Kim K, Leem SH, Lin SY, Lopez-Berestein G, Sood AK, Lee JS. FOXM1 mediates Dox resistance in breast cancer by enhancing DNA repair. Carcinogenesis. 2012; 33:1843–1853.
31. Khongkow P, Gomes AR, Gong C, Man EP, Tsang JW, Zhao F, Monteiro LJ, Coombes RC, Medema RH, Khoo US, Lam EW. Paclitaxel targets FOXM1 to regulate KIF20A in mitotic catastrophe and breast cancer paclitaxel resistance. Oncogene. 2016; 35:990–1002.
32. Wilson MS, Brosens JJ, Schwenen HD, Lam EW. FOXO and FOXM1 in cancer: the FOXO-FOXM1 axis shapes the outcome of cancer chemotherapy. Curr Drug Targets. 2011; 12:1256–1266.
33. Hamurcu Z, Kahraman N, Ashour A, Ozpolat B. FOXM1 transcriptionally regulates expression of integrin beta1 in triple-negative breast cancer. Breast Cancer Res Treat. 2017; 163:485–493.
34. Fischer KR, Durrans A, Lee S, Sheng J, Li F, Wong ST, Choi H, El Rayes T, Ryu S, Troeger J, Schwabe RF, Vahdat LT, Altorki NK, et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature. 2015; 527:472–476.
35. Zheng X, Carstens JL, Kim J, Scheible M, Kaye J, Sugimoto H, Wu CC, LeBleu VS, Kalluri R. Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature. 2015; 527:525–530.
36. Yang N, Wang C, Wang Z, Zona S, Lin SX, Wang X, Yan M, Zheng FM, Li SS, Xu B, Bella L, Yong JS, Lam EW, Liu Q. FOXM1 recruits nuclear Aurora kinase A to participate in a positive feedback loop essential for the self-renewal of breast cancer stem cells. Oncogene. 2017; 36:3428–3440.
37. Bergamaschi A, Madak-Erdogan Z, Kim YJ, Choi YL, Lu H, Katzenellenbogen BS. The forkhead transcription factor FOXM1 promotes endocrine resistance and invasiveness in estrogen receptor-positive breast cancer by expansion of stem-like cancer cells. Breast Cancer Res. 2014; 16:436.
38. Yau C, Wang Y, Zhang Y, Foekens JA, Benz CC. Young age, increased tumor proliferation and FOXM1 expression predict early metastatic relapse only for endocrine-dependent breast cancers. Breast Cancer Res Treat. 2011; 126:803–810.
39. Bergamaschi A, Christensen BL, Katzenellenbogen BS. Reversal of endocrine resistance in breast cancer: interrelationships among 14-3-3zeta, FOXM1, and a gene signature associated with mitosis. Breast Cancer Res. 2011; 13:R70.
40. Sanders DA, Ross-Innes CS, Beraldi D, Carroll JS, Balasubramanian S. Genome-wide mapping of FOXM1 binding reveals co-binding with estrogen receptor alpha in breast cancer cells. Genome Biol. 2013; 14:R6.
41. Millour J, Constantinidou D, Stavropoulou AV, Wilson MS, Myatt SS, Kwok JM, Sivanandan K, Coombes RC, Medema RH, Hartman J, Lykkesfeldt AE, Lam EW. FOXM1 is a transcriptional target of ERalpha and has a critical role in breast cancer endocrine sensitivity and resistance. Oncogene. 2010; 29:2983–2995.
42. Francis RE, Myatt SS, Krol J, Hartman J, Peck B, McGovern UB, Wang J, Guest SK, Filipovic A, Gojis O, Palmieri C, Peston D, Shousha S, et al. FoxM1 is a downstream target and marker of HER2 overexpression in breast cancer. Int J Oncol. 2009; 35:57–68.
43. Bektas N, Haaf A, Veeck J, Wild PJ, Luscher-Firzlaff J, Hartmann A, Knuchel R, Dahl E. Tight correlation between expression of the Forkhead transcription factor FOXM1 and HER2 in human breast cancer. BMC Cancer. 2008; 8:42.
44. Peake BF, Nahta R. Resistance to HER2-targeted therapies: a potential role for FOXM1. Breast Cancer Manag. 2014; 3:423–431.
45. Carr JR, Park HJ, Wang Z, Kiefer MM, Raychaudhuri P. FoxM1 mediates resistance to herceptin and paclitaxel. Cancer Res. 2010; 70:5054–5063.
46. Kambach DM, Sodi VL, Lelkes PI, Azizkhan-Clifford J, Reginato MJ. ErbB2, FoxM1 and 14-3-3zeta prime breast cancer cells for invasion in response to ionizing radiation. Oncogene. 2014; 33:589–598.
47. Lin HY, Zeng D, Liang YK, Wei XL, Chen CF. GATA3 and TRPS1 are distinct biomarkers and prognostic factors in breast cancer: database mining for GATA family members in malignancies. Oncotarget. 2017; 8:34750–61. https://doi.org/10.18632/oncotarget.16160.
48. Jezequel P, Frenel JS, Campion L, Guerin-Charbonnel C, Gouraud W, Ricolleau G, Campone M. bc-GenExMiner 3.0: new mining module computes breast cancer gene expression correlation analyses. Database (Oxford). 2013; 2013:bas060.
49. Jezequel P, Campone M, Gouraud W, Guerin-Charbonnel C, Leux C, Ricolleau G, Campion L. bc-GenExMiner: an easy-to-use online platform for gene prognostic analyses in breast cancer. Breast Cancer Res Treat. 2012; 131:765–775.
50. Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010; 123:725–731.

