INTRODUCTION
Stroke is a major cause of death and disability worldwide as well as a major burden on public health worldwide [1]. Within the globe in 2010, roughly 10% of the 52769700 deaths and about 4% of the 2490385000 disability adjusted life years (DALYs) were due to stroke, with developing countries becoming more severe than in developed countries [2]. Globally, stroke is projected to be the fourth most common cause of premature death and disability by the year 2020 [2]. Stroke is a multifactorial disease associated with a variety of factors. In previous studies, hypertension, diabetes, obesity, smoking and advanced age were identified as risk factors for stroke [3]. Through the study of twins and family history, family inheritance is also closely related with stroke [4]. Simultaneously, Genetics as a risk factor for stroke have also been confirmed, and some genetic polymorphisms such as MTHFR, ApoE [5], P-selectin and interleukin-4 gene [6] have been shown to be associated with the risk of stroke.
Telomere, as a protective structure at the end of chromosome, plays a crucial role in maintaining the stability of the genome. The shortening of telomere length is related to cell senescence and apoptosis. Many studies have confirmed that telomere length is associated with stroke risk, shortening telomere length increases the risk of stroke [7–9]. Genetic polymorphisms of telomere-related genes such as ACYP2, TSPYL6, and TERT have been reported to be associated with stroke [10, 11]. Regulator of telomere elongation helicase 1(RTEL1)belongs to the superfamily 2 (SF2) helicase [12], is an essential helicase that plays an important role in maintaining telomere length and genomic stability [12, 13] RTEL1 gene, located in chromosome 20q13.3, plays an important role in DNA repair and apoptosis [14]. The single nucleotide polymorphism (SNP) of RTEL1 gene has been confirmed to be associated with telomere-related diseases such as Hoyeraal Hreidarsson Syndrome [15], glioma [16], lung cancer [17] and atopic dermatitis [18]. However, so far, no studies have investigated the association between SNPs in the RTEL1 gene and the risk of stroke. In this case-control study, we genotyped five SNPs in RTEL1 gene: rs6089953, rs6010620, rs6010621, rs4809324 and rs2297441, and performed a comprehensive association analysis to identify SNPs associated with the risk of stroke in Chinese population.
RESULTS
Participant characteristics
A total of 400 patients with stroke and 395 healthy individuals were enrolled in the study. The demographic characteristics of the participants are shown in Table 1.The Pearson’s test showed that there was no statistical difference in sex distribution between case group and control group (p > 0.05). The mean age of the participants was 66.83 years in the case group and 48.67 years in the control group.
Table 1: Demographic characteristics of the patients with stroke and control individuals
Characteristic |
Cases (N = 400) |
Controls (N = 395) |
P-value |
|
|---|---|---|---|---|
gender |
female |
137 (34.3%) |
152 (38.5%) |
0.215 |
male |
263 (65.8%) |
243 (61.5%) |
||
Mean age ± SD |
66.83 ± 11.637 |
48.67 ± 11.059 |
< 0.001 |
SD: Standard deviation.
P value of gender was calculated by Welch’s t test
P value of age was calculated by Pearson’s χ2 test.
Association between SNPs in the RTEL1 gene and Stroke
The detailed data of the SNP of the RTEL1 gene and the associations between various SNPs and stroke risk are shown in Table 2. We performed the Hardy Weinberg equilibrium test on the 5 SNPs investigated, and found that all 5 SNPs were in Hardy Weinberg equilibrium in the control subjects (p > 0.05). Rs2297441 was associated with an increased risk of stroke in an allele model(odds ratio [OR] = 1.24, 95% confidence interval [95% CI] = 1.01–1.52, p = 0.043).While in the other four SNPs(Rs6089953, Rs6010620, Rs6010621, Rs4809324), no associations were observed between the alleles and stroke risk.
Table 2: Basic information of candidate SNPs in this study
SNP-ID |
Band |
Position |
Role |
Alleles A/B |
HWE |
MAF |
OR |
95% CI |
P value |
||
|---|---|---|---|---|---|---|---|---|---|---|---|
Case |
Control |
||||||||||
rs6089953 |
20q13.33 |
62291008 |
Intron |
G/A |
0.457 |
0.326 |
0.282 |
1.23 |
1.00 |
1.53 |
0.054 |
rs6010620 |
20q13.33 |
62309839 |
Intron |
G/A |
0.900 |
0.308 |
0.276 |
1.17 |
0.94 |
1.45 |
0.166 |
rs6010621 |
20q13.33 |
62310872 |
Intron |
G/T |
0.801 |
0.301 |
0.272 |
1.15 |
0.93 |
1.43 |
0.200 |
rs4809324 |
20q13.33 |
62318220 |
Intron |
C/T |
1.000 |
0.150 |
0.141 |
1.08 |
0.82 |
1.43 |
0.577 |
rs2297441 |
20q13.33 |
62327582 |
Upstream |
A/G |
1.000 |
0.383 |
0.334 |
1.24 |
1.01 |
1.52 |
0.043 |
SNPs: Single nucleotide polymorphisms; MAF: Minor allele frequency; HWE: Hardy-Weinberg equilibrium; OR: Odds ratio; CI: Confidence interval. A :Minor alleles. B: Major alleles.
We further assessed the association between each SNP and stroke risk using the four models in the unconditional logistic regression analysis: genotype, additive, dominant, and recessive models (Tables 3 and 4). Rs6089953 was associated with an increased risk of stroke under the genotype model (odds ratio [OR] = 1.862, 95% confidence interval [95% CI] = 1.123–3.085, p = 0.016) (Table 3). Rs2297441 was associated with an increased risk of stroke in an additive model (OR = 1.234, 95% CI = 1.005, p = 0.045 , Rs6089953,Rs6010620 and Rs6010621 were associated with an increased risk of stroke in the recessive model (Rs6089953:OR = 1.825, 95% CI = 1.121–2.969, p = 0.01546; Rs6010620:OR = 1.64, 95% CI = 1.008–2.669, p = 0.04656;Rs6010621:OR = 1.661, 95% CI = 1.014–2.722, p = 0.04389). We then evaluated the relationship between SNP haplotypes and stroke, no positive results were observed (Table 5, Figure 1).All of the above results were obtained by the Wald test.
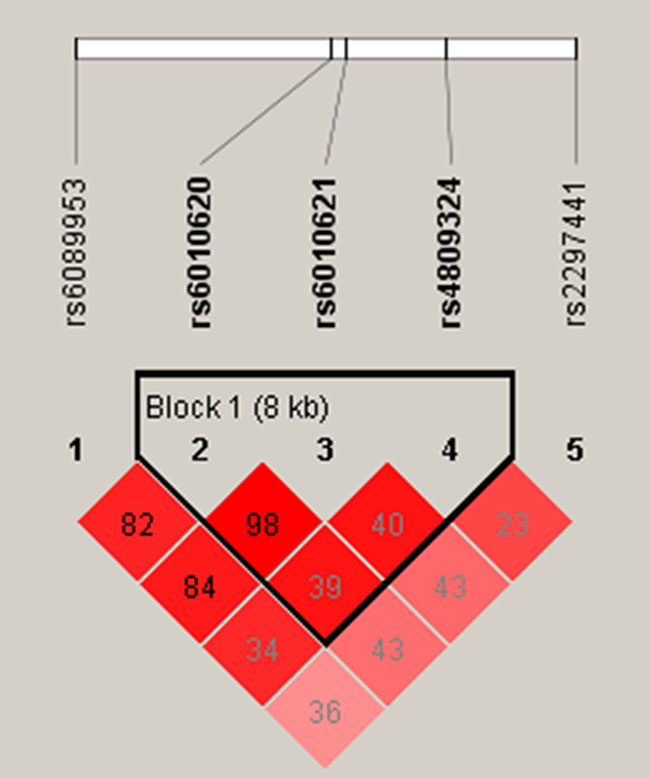
Figure 1: Haplotype block map for all the SNPs of the RTEL1 gene.
Table 3: Single loci association with stroke
SNP-ID |
Dominant |
Recessive |
Additive |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
OR |
95% CI |
P |
OR |
95% CI |
P |
OR |
95% CI |
P |
||||
rs6089953 |
1.16 |
0.88 |
1.54 |
0.289 |
1.83 |
1.12 |
2.97 |
0.015 |
1.23 |
0.99 |
1.52 |
0.057 |
rs6010620 |
1.09 |
0.83 |
1.44 |
0.544 |
1.64 |
1.01 |
2.67 |
0.047 |
1.16 |
0.94 |
1.43 |
0.176 |
rs6010621 |
1.07 |
0.81 |
1.41 |
0.640 |
1.66 |
1.01 |
2.72 |
0.044 |
1.15 |
0.93 |
1.42 |
0.210 |
rs4809324 |
1.03 |
0.75 |
1.41 |
0.876 |
1.87 |
0.74 |
4.73 |
0.188 |
1.08 |
0.82 |
1.42 |
0.583 |
rs2297441 |
1.27 |
0.96 |
1.68 |
0.100 |
1.42 |
0.94 |
2.16 |
0.099 |
1.23 |
1.01 |
1.52 |
0.045 |
SNPs: Single nucleotide polymorphisms; OR: Odds ratio. CI: Confidence interval. P value was calculated by Wald test. *p < 0.05 indicates statistical significant.
Table 4: The association between the single-nucleotide polymorphisms and stroke in Genotype model
SNP-ID |
Genotype |
Case (N) |
Control (N) |
OR |
95% CI |
P |
|
|---|---|---|---|---|---|---|---|
rs6089953 |
GG |
49 |
28 |
1.86 |
1.12 |
3.09 |
0.016 |
GA |
163 |
166 |
1.05 |
0.78 |
1.40 |
0.771 |
|
AA |
188 |
200 |
- |
- |
- |
0.051 |
|
rs6010620 |
GG |
46 |
29 |
1.63 |
0.99 |
2.70 |
0.056 |
GA |
154 |
160 |
0.99 |
0.74 |
1.33 |
0.954 |
|
AA |
200 |
206 |
- |
- |
- |
0.138 |
|
rs6010621 |
GG |
45 |
28 |
1.64 |
0.98 |
2.73 |
0.058 |
GT |
151 |
159 |
0.97 |
0.72 |
1.30 |
0.830 |
|
TT |
204 |
208 |
- |
- |
- |
0.128 |
|
rs4809324 |
CC |
13 |
7 |
1.85 |
0.73 |
4.71 |
0.196 |
CT |
94 |
97 |
0.97 |
0.70 |
1.34 |
0.835 |
|
TT |
292 |
291 |
- |
- |
- |
0.412 |
|
rs2297441 |
AA |
60 |
44 |
1.56 |
1.00 |
2.44 |
0.050 |
AG |
184 |
176 |
1.20 |
0.89 |
1.61 |
0.242 |
|
GG |
153 |
175 |
- |
- |
- |
0.130 |
|
OR: odd ratio; CI: confidence interval; p value was calculated by Wald test. *p < 0.05 indicates statistical significance.
Table 5: Haplotype frequency and their association with stroke in case and control subjects
SNPs |
Haplotype |
Frequency% |
OR |
95% CI |
Wald |
P |
||
|---|---|---|---|---|---|---|---|---|
case |
control |
|||||||
rs6010620|rs6010621|rs4809324 |
||||||||
GGC |
0.1454 |
0.1367 |
1.072 |
0.8113 |
1.415 |
0.4872 |
0.6261 |
|
GGT |
0.1554 |
0.1354 |
1.172 |
0.8875 |
1.548 |
1.119 |
0.2631 |
|
ATT |
0.688 |
0.7203 |
0.8612 |
0.6966 |
1.065 |
-1.381 |
0.1672 |
|
SNPs: SNPs forming the haplotype
Wald: Z statistics of wald test for measuring odds ratio departure from 1
P: P value calculated by Wald test. P-value < 0.05 indicates statistical significance.
DISCUSSION
In this study, we investigated the associations between SNPs in the RTEL1 gene and stroke risk in the Chinese population. Among the five SNPs we selected, four SNPs (rs6089953, rs6010620, rs6010621 and rs2297441) may be associated with the risk of stroke in the Chinese Han population.
Stroke is a multifactorial disease associated with a variety of factors. Among them, the growth of age and stroke risk is positively correlated. Vascular cell senescence contributes to endothelial dysfunction [19], which can cause atherosclerotic plaque progression [20], further resulting in stroke [21]. It has been demonstrated in previous studies that telomere length shortening is associated with stroke [7–9]. The telomere length decreases continuously with aging due to the decrease of telomerase activity [22].The length of telomeres is also affected by many factors, including Eating habits and lifestyles [23], body mass index (BMI)and smoking [24],which genetic factors in different races occupied 36% -84% [25]. RTEL1 is the essential helicase for telomere maintenance and the regulation of homologous recombination, and plays an important role in maintaining telomere length [26]. The RTEL1 gene, as an important gene for maintaining telomere length, has been confirmed to be associated with telomere-related diseases such as Hoyeraal Hreidarsson Syndrome, glioma, lung cancer and atopic dermatitis [15–18], But so far, no studies have investigated the association between SNPs in the RTEL1 gene and the risk of stroke. Based on our experimental results, we conclude that perhaps the SNPs in the RTEL1 gene can cause cell apoptosis by influencing the length of telomere, and then causes endothelial dysfunction, and finally affects stroke.
Recent studies have reported that the genetic polymorphisms of telomere-related genes such as ACYP2,TSPYL6,and TERT are associated with stroke by affecting telomere length [10, 11], including rs11125529, rs12615793, rs843711, rs11896604, rs843706 within both ACYP2 and TSPYL6, rs17045754 in ACYP2 gene, and rs2736122, rs2853668 in TERT gene. In our experimental results, We found that four SNPs (Rs2297441, Rs6089953, Rs6010620, and Rs6010621) were associated with the risk of stroke in this population. It may be possible to show that the mechanism of the SNPs in the RTEL1 gene impact on stroke may be the same as other telomere - related genes. Because in previous studies there was a lack of research and reporting on the association between the SNPs in the RTEL1 gene and Stroke, we have no way to compare with other data. The exact mechanism for explaining the relationship between the SNPs in the RTEL1 gene and stroke cannot be determined from our study alone, we should do further functional experiments to verify.
There are several limitations of this study. First, the sample size is not large enough, Second, the clinical subtypes of stroke were not considered in this study. In addition, some influencing factors such as eating habits, living environment, and Smoking history are not considered. These limitations will be addressed in future research.
MATERIALS AND METHODS
Study participants
A total of 400 patients with stroke and 395 healthy individuals were enrolled in the study. The characteristics of the participants are shown in Table 1. All participants are genetically unrelated Han Chinese. The stroke status of each case was determined by Magnetic Resonance Imaging and their clinical records as confirmed by two clinicians. Healthy controls without history of stroke and neurological impairments were recruited from healthcare center of our hospital. This study was conducted in accordance with the Chinese Department of Health and Human Services regulations for the protection of human research subjects. Informed consent was obtained from all participants.
SNP selection and genotyping
We selected five SNPs from previously reported RTEL1 gene polymorphisms, and matched SNPs with MAF > 5% in the HapMap Asian population selected for association analysis [16, 27, 28]. Venous blood samples (5 mL) were collected from each Participant during a laboratory examination. DNA was extracted from whole blood samples using the Gold Mag-Mini Whole Blood Genomic DNA Purification Kit (version 3.0; TaKaRa, Japan) [29]. The DNA concentration was measured by spectrometry (DU530 UV/VIS spectrophotometer, Beckman Instruments, Fullerton, CA, USA). The Sequenom MassARRAY Assay Design 3.0 software (Sequenom, Inc, San Diego, CA, USA) was used to design the multiplexed SNP Mass EXTEND assay. Genotyping was performed using a Sequenom MassARRAY RS1000 (Sequenom, Inc.) according to the manufacturerfacturerufa [30]. The SequenomTyper 4.0 Software 4.Sequenom, Inc.) was used to manage and analyze the data [31]. Based on these results, the following five SNPs were selected: rs6089953, rs6010620, rs6010621, rs4809324, and rs2297441. The SNP data are shown in Table 3.
Statistical analysis
Pearson’s χ2 test and Welch’s t test were used to evaluate the differences in the demographic characteristics between the cases and controls [32]. The Hardy-Weinberg equilibrium of each SNP was assessed in order to compare the expected frequencies of the genotypes in the control patients. All of the minor alleles were regarded as risk alleles for stroke susceptibility. To evaluate associations between the SNPs and risk of stroke in the four models (genotype, dominant, recessive, and additive), ORs and 95% CIs were calculated using unconditional logistic regression analysis [33]. In multivariate analyses, unconditional logistic regression was used to assess the association between each SNP and the risk of stroke [33]. Linkage disequilibrium analysis and SNP haplotypes were analyzed using the Haploview software package (version 4.2) and the SHEsi software platform (http://www.nhgg.org/analysis/) [34]. All statistical analyses were performed using the SPSS version 17.0 statistical package (SPSS, Chicago, IL, USA) and Microsoft Excel. A p < 0.05 was considered statistically significant and all statistical tests were two-sided.
CONCLUSIONS
In summary, we have identified four new associations between the SNPs (Rs6089953, Rs6010620, Rs6010621 and Rs2297441) in RTEL1 gene and stroke. Our results suggest that these SNPs may serve as molecular markers of stroke susceptibility for primary prevention of stroke during clinical treatment. At the same time, this study provides a new idea for future research on association between single nucleotide polymorphisms and Stroke.
ACKNOWLEDGMENTS AND FUNDING
This work was supported by Natural Science Foundation of Hainan Province (Grant Number: 817334). The authors are also grateful to the patients and control individuals for their participation in the study. We thank the clinicians and hospital staff who contributed to sample and data collection for this study.
CONFLICTS OF INTEREST
The authors declare that there are no conflicts of interest.
REFERENCES
1. Bonita R. Epidemiology of stroke. Lancet. 1992; 339:342–344.
2. Feigin VL, Forouzanfar MH, Krishnamurthi R, Mensah GA, Myles C, Bennett DA, Moran AE, Sacco RL, Anderson L. Global and regional burden of stroke during 1990–2010: findings from the Global Burden of Disease Study 2010. Lancet. 2014; 383:245–254.
3. O’Donnell MJ, Xavier D, Liu L, Zhang H, Chin SL, Raomelacini P, Rangarajan S, Islam S, Pais P, Mcqueen MJ. Risk factors for ischaemic and intracerebral haemorrhagic stroke in 22 countries (the INTERSTROKE study): a case-control study. Lancet. 2010; 376:112–123.
4. Dichgans M. Genetics of ischaemic stroke. Lancet Neurol. 2007; 6:149–161.
5. Banerjee I, Gupta V, Ganesh S. Association of gene polymorphism with genetic susceptibility to stroke in Asian populations: a meta-analysis. J Hum Genet. 2007; 52:205–219.
6. Zee RY, Cook NR, Cheng S, Reynolds R, Erlich HA, Lindpaintner K, Ridker PM. Polymorphism in the P-selectin and interleukin-4 genes as determinants of stroke: a population-based, prospective genetic analysis. Hum Mol Genet. 2004; 13:389–396.
7. Sen A, Marsche G, Freudenberger P, Schallert M, Toeglhofer AM, Nagl C, Schmidt R, Launer LJ, Schmidt H. Association Between Higher Plasma Lutein, Zeaxanthin, and Vitamin C Concentrations and Longer Telomere Length: Results of the Austrian Stroke Prevention Study. Journal of the American Geriatrics Society. 2014; 62:222–229.
8. Jiang X, Dong M, Cheng J, Huang S, He Y, Ma K, Tang B, Guo Y. Decreased leukocyte telomere length (LTL) is associated with stroke but unlikely to be causative. Plos One. 1932; 8:e68254-e68254.
9. Zhang W, Chen Y, Wang Y, Liu P, Zhang M, Zhang C, Hu FB, Hui R. Short telomere length in blood leucocytes contributes to the presence of atherothrombotic stroke and haemorrhagic stroke and risk of post-stroke death. Clinical Science. 2013; 125:27–36.
10. Liang Y, Zhang R, Zhang S, Ji G, Shi P, Yang T, Liu F, Feng J, Li C, Guo D. Association of ACYP2 and TSPYL6 Genetic Polymorphisms with Risk of Ischemic Stroke in Han Chinese Population. Mol Neurobiol. 2016:1–8.
11. Bressler J, Franceschini N, Demerath EW, Mosley TH, Folsom AR, Boerwinkle E. Sequence variation in telomerase reverse transcriptase ( TERT ) as a determinant of risk of cardiovascular disease: the Atherosclerosis Risk in Communities (ARIC) study. BMC Med Genet. 2015; 16:1–10.
12. Uringa EJ, Lisaingo K, Pickett HA, Brind’Amour J, Rohde JH, Zelensky A, Essers J, Lansdorp PM. RTEL1 contributes to DNA replication and repair and telomere maintenance. Mol Biol Cell . 2012; 23:2782–2792.
13. Barber LJ, Youds JL, Ward JD, Mcilwraith MJ, O’Neil NJ, Petalcorin MIR, Martin JS, Collis SJ, Cantor SB, Auclair M. RTEL1 Maintains Genomic Stability by Suppressing Homologous Recombination. Cell. 2008; 135:261–271.
14. Vannier JB, Pavicickaltenbrunner V, Petalcorin MI, Ding H, Boulton SJ. RTEL1 dismantles T loops and counteracts telomeric G4-DNA to maintain telomere integrity. Cell. 2012; 149:795–806.
15. Ballew BJ, Joseph V, De S, Sarek G, Vannier JB, Stracker T, Schrader KA, Small TN, O’Reilly R, Manschreck C. A Recessive Founder Mutation in Regulator of Telomere Elongation Helicase 1, RTEL1, Underlies Severe Immunodeficiency and Features of Hoyeraal Hreidarsson Syndrome. Plos Genet. 2013; 9:535–554.
16. Pei S, Zhao F, Liu J, Fu Q, Shang P. Association between regulator of telomere elongation helicase 1 polymorphism and susceptibility to glioma. Int J Clin Exp Med. 2014; 8:690–697.
17. Yan S, Xia R, Jin T, Ren H, Yang H, Li J, Yan M, Zhu Y, Chen M. RTEL1 polymorphisms are associated with lung cancer risk in the Chinese Han population. Oncotarget. 2016; 7:70475–70480. https://doi.org/10.18632/oncotarget.12297.
18. Li P, Wu B, Guo B, Zhang X, Li Y, Cheng H, Ding Y, Wang J, Zuo X, Yuan X. Genetic variants rs2393903 at 10q21.2 and rs6010620 at 20q13.33 are associated with clinical features of atopic dermatitis in the Chinese Han population. Journal of Dermatological Science. 2013; 72:64–66.
19. Minamino T, Miyauchi H, Yoshida T, Ishida Y, Yoshida H, Komuro I. Endothelial Cell Senescence in Human Atherosclerosis Role of Telomere in Endothelial Dysfunction. Circulation. 2002; 105:1541–1544.
20. Bürrig KF. The endothelium of advanced arteriosclerotic plaques in humans. Arteriosclerosis & Thrombosis A Journal of Vascular Biology. 1991; 11:1678–1689.
21. Parmar JP, Rogers WJ, Mugler JP, Baskurt E, Altes TA, Nandalur KR, Stukenborg GJ, Phillips CD, Hagspiel KD, Matsumoto AH. Magnetic Resonance Imaging of Carotid Atherosclerotic Plaque in Clinically Suspected Acute Transient Ischemic Attack and Acute Ischemic Stroke. Circulation. 2010; 122:2031–2038.
22. Atzmon G, Cho M, Cawthon RM, Budagov T, Katz M, Yang X, Siegel G, Bergman A, Huffman DM, Schechter CB. Evolution in health and medicine Sackler colloquium: Genetic variation in human telomerase is associated with telomere length in Ashkenazi centenarians. Proceedings of the National Academy of Sciences. 2010; 107:1710–1717.
23. Cassidy A, De VI, Liu Y, Han J, Prescott J, Hunter DJ, Rimm EB. Associations between diet, lifestyle factors, and telomere length in women. Am J Clin Nutr . 2010; 91:1273–1280.
24. Valdes AM, Andrew T, Gardner JP, Kimura M, Oelsner E, Cherkas LF, Aviv A, Spector TD. Obesity, cigarette smoking, and telomere length in women. Lancet. 2005; 366:662–664.
25. Njajou OT, Cawthon RM, Damcott CM, Wu SH, Ott S, Garant MJ, Blackburn EH, Mitchell BD, Shuldiner AR, Hsueh WC. Telomere length is paternally inherited and is associated with parental lifespan. P Natl Acad Sci Usa. 2007; 104:12135–12139.
26. Uringa EJ, Youds JL, Lisaingo K, Lansdorp PM, Boulton SJ. RTEL1: an essential helicase for telomere maintenance and the regulation of homologous recombination. Nucleic Acids Res. 2011; 39:1647–1655.
27. Fahmideh MA, Lavebratt C, Schüz J, Röösli M, Tynes T, Grotzer MA, Johansen C, Kuehni CE, Lannering B, Prochazka M. CCDC26, CDKN2BAS, RTEL1 and TERT Polymorphisms in pediatric brain tumor susceptibility. Carcinogenesis. 2015; 36:876–882.
28. Ren Z, Zhu J, Gu H, Liu R, Chen S, Rong G, Sun B. Decoy receptor 3 polymorphisms are not associated with the risk of esophageal cancer in a Chinese population. Biomarkers. 2014; 19:340–344.
29. Carracedo A, Sánchez-Diz P. Forensic DNA Typing Protocols. Methods in Molecular Biology. 2005; 297:1–12.
30. Gabriel S, Ziaugra L, Tabbaa D. SNP genotyping using the Sequenom MassARRAY iPLEX platform. 2009; Chapter 2:2.12.11–12.12.16.
31. Thomas RK, Baker AC, Debiasi RM, Winckler W, Laframboise T, Lin WM, Wang M, Feng W, Zander T, Macconaill L. High-throughput oncogene mutation profiling in human cancer. Nat Genet. 2007; 39:347–351.
32. Adamec C. Example of the use of the nonparametric test. Test X2 for comparison of 2 independent examples. Ceskoslovenské Zdravotnictví. 1964; 12:613–619.
33. Bland JM, Altman DG. The odds ratio. BMJ. 2000; 320:381–382.
34. Yong Y, Lin HE. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Research. 2006; 15:97–98.

